Abstract
Arabidopsis thaliana is well established as a model plant in modern plant biology. However, remarkably few details are known about plastidial promoters in Arabidopsis. Here, we report on the identification and analyses of sequences at transcription start sites of selected genes. The genes encoded by the plastome of higher plants are transcribed by a plastid-encoded (PEP) and a nuclear-encoded RNA plastid polymerase (NEP). To discriminate between NEP and PEP promoters we compared the 5′-ends of transcripts from chlorophyll-deficient Arabidopsis plants, which were grown on prokaryotic translation inhibitor spectinomycin to inhibit biosynthesis of PEP, with those of untreated plants. Using 5′-RACE combined with enzymatic treatment of RNAs to recognize primary and secondary 5′-ends, we unambiguously identified transcription initiation sites of the Arabidopsis accD, atpB, atpI, rpoB, rps4, rps15, and ycf1 genes. Comparison of plastidial promoters from tobacco and Arabidopsis revealed a high diversity, which may also apply to other plants. Furthermore, the diversity in individual promoter usage in different plants suggests that there are species-specific solutions for attaining control over gene expression in plastids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chloroplasts, as the sites of photosynthesis, have a central role in the metabolism of plant cells (Gray 1993; López-Juez and Pyke 2005). Although many genes of the genome of the ancestral cyanobacteria have been transferred into the nucleus, plastomes still harbor genes for products that function primarily in photosynthesis and gene expression. Examples are the rpo genes that encode homologues of the bacterial RNA polymerase core subunits, which are complemented with nuclear encoded σ-factors to form the plastid-encoded plastid RNA polymerase (PEP; Shiina et al. 2005; Liere and Börner 2007). In surprising contrast to their bacterial ancestors, this RNA polymerase is not sufficient to transcribe all genes encoded by the rather small plastomes of higher plants. The existence of a second, plastid-localized, nuclear-encoded transcription activity (NEP, nuclear-encoded plastid RNA polymerase) was established by analyzing mutant and transplastomic plants, respectively, that transcribe plastid genes despite the lack of PEP (reviewed in Hess and Börner 1999; Weihe 2004; Liere and Börner 2007). These studies revealed three classes of plastidial genes. Generally, genes encoding components for the photosynthetic apparatus have PEP promoters (class I), whereas non-photosynthesis-related genes have promoters for both RNA polymerases (class II); few genes are transcribed exclusively from an NEP promoter (class III; Maliga 1998).
Since PEP has evolved from a eubacterial-type RNA polymerase, it recognizes promoters which contain -35 (TTGaca) and −10 (TAtaaT) consensus sequences typical of σ70-type E. coli promoters (Reznikov et al. 1985; for reviews see Gruissem and Tonkyn 1993; Link 1994; Hess and Börner 1999; Weihe and Börner 1999; Liere and Maliga 2001; Weihe 2004). NEP transcription activity is represented by an RNA polymerase similar to RNA polymerases of the T7 phage type (Lerbs-Mache 1993; Chang et al. 1999; Liere and Maliga 1999; Liere et al. 2004). Genes encoding organellar phage-type RNA polymerases (RpoT) have been found in several higher plant genomes (Liere and Börner 2007). One organellar phage-type RNA polymerase is targeted to plastids (RpoTp), a second is targeted into mitochondria (RpoTm), whereas a third type is found exclusively in dicots, and encodes an enzyme dually targeted into both mitochondria and plastids (RpoTmp; Weihe 2004; Khan 2005). Most NEP promoters consist of a core sequence (YRTA; type-Ia), similar to promoters of plant mitochondria (Liere and Maliga 1999, 2001; Weihe and Börner 1999; Binder and Brennicke 2003; Kühn et al. 2005). A subclass of NEP promoters shares a GAA-box motif upstream of the YRTA-motif (type-Ib; Kapoor and Sugiura 1999). Type-II NEP promoters, represented by dicot clpP promoters, lack these motifs and possess crucial sequences located downstream of the transcription initiation site (Weihe and Börner 1999; Liere and Maliga 2001). Furthermore, the existence of additional non-consensus NEP promoters (Pc) has been reported for the rrn operon in spinach, mustard, and Arabidopsis, and for the internal promoters of certain tRNAs (reviewed in Liere and Börner 2007).
Although Arabidopsis is well established as a model plant in modern plant biology, remarkably few details are known about its plastidial promoters. In addition to the PEP promoters PpsbA-77 (Liere et al. 1995; Shen et al. 2001), PrbcL-179, PpsbD-256, PpsbD-541, PtrnE-UUC, PtrnV-UAC (Hanaoka et al. 2003), PpsbD-946 (Hoffer and Christopher 1997; Hanaoka et al. 2003), PpsaA-188 (Fey et al. 2005), PpsaJ-37 (Nagashima et al. 2004), and Prrn16-112 (Sriraman et al. 1998b), only two NEP promoters, PclpP-58 (type-II; Sriraman et al. 1998a), and Prrn16-139 (Pc; Sriraman et al. 1998b) have been identified (promoters are specified by the name of the gene and the position of the initiating nucleotide with respect to the start of the coding sequence or mature rRNA). Comparison of transcription in various plant species revealed striking differences in promoter usage for some plastome-encoded genes (e.g. rrn16; reviewed in Liere and Börner 2007). The use of different promoters and types of RNA polymerases in expression of plastidial genes is not yet fully understood. Furthermore, identification of NEP promoters in Arabidopsis is an important prerequisite to analyze promoter specificities of the plastid-localized RpoTp and RpoTmp enzymes. Therefore, we investigated promoter usage using 5′-RACE combined with enzymatic treatment of RNAs to discriminate between primary (formed by transcription initiation) and secondary 5′-ends (formed by nucleolytic processing steps; Bensing et al. 1996; Kühn et al. 2005). Mutants or transplastomic plants lacking PEP activity have not been described in Arabidopsis. We therefore used leaves from chlorophyll-deficient Arabidopsis plants grown on spectinomycin as the material in our experiments (Zubko and Day 1998). Since the core subunits of PEP are synthesized on plastid ribosomes, treatment with spectinomycin, which inhibits plastidial translation (Moazed and Noller 1987), should lead to PEP-deficiency. Therefore, comparison of transcription initiation sites used in plastids of spectinomycin-treated and untreated material should allow for discrimination between NEP and PEP promoters. Here, we report on the analyses of sequences at transcription start sites of selected plastome-encoded genes. Although some plastidial promoter sequences are highly homologous across plant species, highly diverse promoter sequences and types indicate species-specific solutions in gaining control over plastidial gene expression.
Materials and methods
Plant material
Arabidopsis thaliana (ecotype Columbia) was grown on MS medium under a light–dark cycle of 8 h/16 h at 20°C. After 14 days, the seedlings were exposed to a light regime of 16 h light/8 h darkness. The leaves were harvested after 21 days for RNA isolation. Germinating Arabidopsis seeds on MS medium containing 500 mg ml−1 spectinomycin dihydrochloride (Duchefa) generated chlorophyll-deficient plants, which were harvested after growing for 21 days on the same medium (Zubko and Day 1998). The light intensity was 150 μE s−1 m−2.
RNA isolation and mapping of transcription initiation sites
The 5′-ends of organellar primary transcripts carry triphosphates, while processed transcripts have monophosphates at their 5′-ends. The latter can be selectively ligated to an RNA oligonucleotide (heswa-001; 5′-GAUAUGCGCGAAUUCCUGUAGAACGAACACUAGAAGAAA-3′), to which a forward primer (P1a; 5′-CGAATTCCTGTAGAACGAACACTAGAAG-3′) anneals in a subsequent 5′-RACE step. 5′-ends of primary transcripts are ligated only after dephosphorylation through tobacco acid pyrophosphatase (TAP; Epicentre Biotechnologies). Thus, 5′-RACE yields products from TAP-treated RNA for both primary and processed transcripts, whereas without TAP treatment, products resulting from primary transcript termini are absent. Primary transcript 5′-ends are therefore identified by comparison of 5′-RACE products obtained from TAP-treated (+T) and non-treated RNA (-T).
RNA was extracted from 21-day-old leaves and isolated plastids (Gruissem et al. 1986) using TRIzol (Invitrogen) following the manufacturer’s protocol and subjected to TAP-treatment and 5′-RACE as detailed in Kühn et al. (2005). Starting from the coding regions, reverse gene-specific primers for cDNA synthesis and PCR were designed to span intergenic regions upstream of the tested genes in intervals of about 100–300 bp (Fig. S1 and S2). Relevant gene-specific primers are listed in Table 1. Adaptor primer P1a served as forward primer in PCR reactions. PCR reactions were analyzed on agarose gels. Products of interest were excised, purified, and ligated into pDrive (Qiagen). Bacterial clones containing the plasmid insert were identified by colony PCR with vector-specific primers, and PCR products were purified and sequenced. Obtained sequences were aligned and analyzed for heswa-linker/plastome sequence junctions which mark transcript 5′-ends.
Results
PEP-deficient Arabidopsis plants
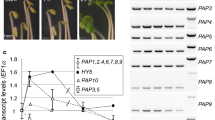
With few exceptions, most NEP promoters are actively used, and therefore mapped, only in plastids of non-green tissues with no or reduced PEP activity (Shiina et al. 2005; Liere and Börner 2007). Hence, most NEP promoters in dicots were characterized in transplastomic tobacco plants lacking one of the PEP core components (Δrpo; Hajdukiewicz et al. 1997; Serino and Maliga 1998). Thus far, plastid transformation to knock out rpo genes in Arabidopsis is not routinely feasible (Sikdar et al. 1998). To map Arabidopsis NEP promoters we blocked plastidial protein synthesis using the antibiotic spectinomycin, which generated chlorophyll-deficient plants lacking PEP (Zubko and Day 1998). On spectinomycin containing media Arabidopsis plants germinated and grew for about 10 days until their further development stopped (Fig. 1a). RNA isolated from these plants showed drastically reduced levels of plastidial ribosomal RNA (Fig. 1b), which indicated a severe loss in their translational competence. The deficiency in translation resulted in low or lacking PEP activity as evident from reduction or loss of transcripts generated by PEP (Fig. 2, psaA, psbA, rrn16; Fig. 4, atpI).
Chlorophyll-deficient Arabidopsis. Arabidopsis seeds germinated on MS medium containing spectinomycin generated chlorophyll-deficient plants (a). RNA isolated from 21 days old plants showed drastically reduced levels of plastidial ribosomal RNA (b, lane 1), when compared to the wild type (lane 2). Positions of ribosomal RNAs are indicated on the right
Identification of psaA, psbA, clpP, and rrn16 promoters in Arabidopsis plastids. Arabidopsis RNA isolated from green (+Tg) and white leaf tissue (+Tw) 5′-ligated to an RNA-linker after TAP-treatment and, as a control, untreated RNA (-T) were subjected to RT-PCR with psaA (a), psbA (b), clpP (c), and rrn16 (d) gene-specific primers (Table 1). Products were separated on agarose gels alongside molecular weight markers; sizes are indicated in base pairs. Please note that the PCR fragments contain additional 31 bp of the heswa-linker that was ligated to the RNA. Arrows mark bands corresponding to primary transcript 5′-ends and labeled with the position in respect to the translation initiation site or mature rRNA (+1). Chromatograms below display sequences at the ligation sites of cloned 5′-RACE products. Additionally, sequences of the promoter regions with putative promoter motifs marked in bold face letters are given. Transcription initiation sites are underlined and marked by bent arrows
Identification of transcript initiation sites of plastidial genes
To precisely determine transcript initiation sites of the psaA, psbA, atpB, atpI, clpP, rrn16, rps4, rps15, accD, rpoB, and ycf1 genes we used a previously described 5′-RACE technique selectively detecting 5′-ends of primary transcripts (Bensing et al. 1996; Miyagi et al. 1998; Kühn et al. 2005). To distinguish between PEP and NEP generated transcripts, we compared initiation sites of RNA isolated from green (+Tg) and chlorophyll-deficient spectinomycin-treated tissue (+Tw). To prove the concept, we mapped the transcription initiation sites of known PEP and NEP promoters. Analysis of psaA transcripts revealed a distinct PCR fragment of 305 bp, which was only amplified from TAP-treated RNA isolated from green tissue (+Tg) but neither from TAP-treated RNA isolated from white (+Tw) nor from untreated RNA (-T, Fig. 2a). This was therefore concluded to correspond to a primary PEP transcript. As described by Fey et al. (2005), sequence analysis placed the native psaA 5′-end at position −188 with respect to the translation initiation site (+1). Similarly, we located the exact 5′-end of native psbA transcripts at position −77 (Fig. 2b), consistent with earlier reports (Liere et al. 1995; Shen et al. 2001). Interestingly, a weak PCR signal was also detected in +Tw samples. Sequence analysis revealed the same initiation site PpsbA-77 as detected in +Tg samples (data not shown). This suggests that some residual PEP transcripts are still present in spectinomycin-treated plants, only detectable by the sensitive PCR method. As expected from data on clpP expression published by Sriraman et al. (1998a), we were able to detect transcript 5′-ends from the strong type-II PclpP-58 NEP promoter in both +Tg and +Tw assays (Fig. 2c). However, although a PCR fragment of the size expected for a transcript initiated from the PEP promoter PclpP-115 has been detected (Fig. 2c, +Tg, asterisk), we did not succeed in determining the sequence. As described for rrn16 (Sriraman et al. 1998b), transcripts initiated at the PEP promoter Prrn16-112 were mapped in +Tg, as well as transcripts initiated at the Pc promoter Prrn16-139 in +Tw samples (Fig. 2d). Therefore, we concluded that we were able to distinguish between NEP and PEP promoters in our assays.
The tobacco atpB gene is transcribed from at least four promoters (Hajdukiewicz et al. 1997). In Arabidopsis, however, only two initiation sites were found (Fig. 3a). One clusters at positions −520/−517/−515 strongly detectable in green RNA samples (panel atpB-a; +Tg) and is preceded by eukaryotic −35/−10 promoter consensus motifs of PEP promoters (Fig. 5a). As observed in the psbA assays, frail residual transcripts originating from this initiation site were also detectable in RNAs of white leaf tissue (panel atpB-a; +Tw). Using a different primer (P3atpB-b), we detected a PCR fragment in both RNAs placing a native RNA 5′-end at position −318 (Fig. 3a, panel atpB-b). This initiation site is preceded by a CATA sequence showing similarity to the YRTA-consensus motif of type-I NEP promoters (Fig. 5b). These data indicate that atpB is transcribed by both NEP and PEP in green tissue.
Arabidopsis atpB, rps4, and ycf1 genes are transcribed from both NEP and PEP promoters. Arabidopsis RNA isolated from green (+Tg) and white leaf tissue (+Tw) 5′-ligated to an RNA-linker after TAP-treatment and, as a control, untreated RNA (-T) were subjected to RT-PCR with atpB (a), rps4 (b), and ycf1 (c) gene-specific primers (Table 1). In case of atpB two primers were used, P3atpB-a in atpB-a and P3atpB-b in atpB-b. PCR-fragments representing the different transcript 5′-ends of PCR-fragments of ycf1 in green and white tissue are marked with the letters ‘g’ and ‘w’. Products were separated on agarose gels alongside molecular weight markers; sizes are indicated in base pairs. Arrows mark bands corresponding to primary transcript 5′-ends and labeled with the position in respect to the translation initiation site or mature rRNA (+1). Chromatograms below display sequences at the ligation sites of cloned 5′-RACE products. Additionally, sequences of the promoter regions with putative promoter motifs marked in bold face letters are given. Transcription initiation sites are underlined and marked by bent arrows
Transcription initiation sites with upstream-located PEP (Fig. 5a) and NEP promoter elements (Fig. 5b) were also found upstream of rps4 (Fig. 3b). However, while the PEP initiation site Prps4-123 was only detectable in green tissue (+Tg), transcripts initiated from Prps4-151 were only evident in RNA of white leaf tissue (+Tw), indicating that rps4 is transcribed by PEP but not by NEP in green tissue.
Characterization of ycf1 5′-ends revealed three transcription start sites at positions −34, −39, and −104 (Fig. 3c). The Pycf1-104 initiation site was detected in both RNA samples (+Tg and +Tw; 255 bp). Different 5′-ends, however, were identified by sequence analyses of the smaller PCR fragments (190 bp in +Tw; 185 bp in +Tg). The 190-bp fragment, which was amplified in white leaves, revealed an initiation site at position −39 (Pycf1-39; in Fig. 3c marked with P-39w). However, the 185-bp fragment detected in green leaves placed a 5′-end at position −34 (Pycf1-34; in Fig. 3c indicated as P-34g). Sequences upstream of Pycf1-39 and Pycf1-104 show typical CATA motifs of type-I NEP promoters. The Pycf1-39 promoter sequence is highly conserved to the sole ycf1 promoter in tobacco (NtPycf1-41; Hajdukiewicz et al. 1997) and was classified into the group of type Ib promoters (Fig. 5b). Pycf1-34 active in green leaves, however, displays typical −35/−10 motifs of PEP promoters upstream of its initiation site, suggesting a polymerase switch from NEP to PEP comparing white and green tissue in this distinct ycf1 upstream region.
Analysis of RNAs synthesized from the accD gene (Fig. 4a) revealed two PCR fragments of 170 and 240 bp which were amplified from TAP-treated RNA from white leaves (+Tw) and untreated cellular RNA (-T) indicating processed RNA 5-ends. Sequence analysis of two additional bands of 94 and 174 bp in size detected in +Tw but neither in +Tg, nor in −T samples placed native accD 5′-ends at positions −172 and −252. CATA (PaccD-252) and TAAA (PaccD-172) sequence motifs are located immediately upstream, showing similarity to the YRTA-consensus motif of type-I NEP promoters. In case of PaacD-172, an ATAAGAA-motif found immediately upstream allowed sub-classification of this promoter into the group of type-Ib NEP promoters, whereas PaccD-252 belongs to the group of type-Ia NEP promoters (Fig. 5b). Interestingly, the processing site yielding the 240-bp PCR fragment indicates transcriptional initiation further upstream. However, TAP assays spanning the rbcL-accD intergenic region did not reveal further 5′-ends of primary transcripts suggesting additional initiation of transcription within or upstream of the rbcL gene (data not shown).
Arabidopsis accD, rpoB, rps15, and atpI genes are transcribed from NEP or PEP promoters. Arabidopsis RNA isolated from green (+Tg) and white leaf tissue (+Tw) 5′-ligated to an RNA-linker after TAP-treatment and, as a control, untreated RNA (-T) were subjected to RT-PCR with accD (a), rpoB (b), rps15 (c), and atpI (d) gene-specific primers (Table 1). Products were separated on agarose gels alongside molecular weight markers; sizes are indicated in base pairs. Arrows mark bands corresponding to primary transcript 5′-ends and labeled with the position in respect to the translation initiation site or mature rRNA (+1). Chromatograms below display sequences at the ligation sites of cloned 5′-RACE products. Additionally, sequences of the promoter regions with putative promoter motifs marked in bold face letters are given. Transcription initiation sites are underlined and marked by bent arrows
Synopsis of PEP (a) and type-Ia, type-Ib, type-II NEP (b), and Pc promoters (c). Conserved -35/-10 boxes, YRTA motifs and GAA-box are marked; mapped transcription initiation sites are underlined. Tobacco promoter sequences are displayed according to Hajdukiewicz et al. (1997), Sriraman et al. (1998a), Vera and Sugiura (1995), and Meng et al. (1988); spinach promoter sequences according to Iratni et al. (1997)
In case of the rpoB operon encoding three subunits of the plastid-encoded plastid RNA polymerase (PEP), PCR analysis revealed in +Tw assays a prominent fragment of 328 bp (Fig. 4b). This places a transcription start site at position −300. In contrast to the tobacco type-Ia NtPrpoB-345 promoter (Liere and Maliga 1999), CATA and ATCGAA sequence motifs immediately upstream of the initiation site suggest a classification of PrpoB-300 as type Ib NEP promoter (Fig. 5b).
Correspondingly, characterization of rps15 transcript 5′-ends revealed a sole initiation site of 139 bases upstream of the translation initiation site, observable only in RNAs of white leaf tissue (Fig. 4c; +Tw). Directly upstream of the transcription initiation site, a core TCTA and a further upstream-located ACAGAA sequence motif suggest a categorization of Prps15-139 into the group of type-Ib NEP promoters (Fig. 5b).
In case of atpI, we found transcription initiation sites at positions −229 and −225, detectable only in assays with RNA from green leaves (Fig. 4d; +Tg). The directly upstream located −35/−10 promoter elements assigned PatpI-229 to the group of PEP promoters (Fig. 5a).
Discussion
Remarkably few details are known about plastidial promoters in Arabidopsis thaliana. Here, we report on the identification and analyses of sequences at transcription initiation sites of selected plastidial genes. To discriminate between NEP and PEP promoters we compared RNAs from chlorophyll-deficient Arabidopsis plants grown on spectinomycin (Fig. 1) and untreated material (Zubko and Day 1998). Transcription initiation sites were mapped using 5′-RACE combined with enzymatic treatment of RNAs to discriminate between primary and secondary 5′-ends (Bensing et al. 1996; Kühn et al. 2005). By mapping known NEP and PEP promoters (Fig. 2) we were able to prove that we could distinguish between the two promoter types. Interestingly, residual amounts of some PEP-derived transcripts (e.g. psbA, Fig. 2b) were detectable in RNAs from spectinomycin-treated plants. This was also reported for spectinomycin-treated albino Brassica napus plants, which, however, showed no detectable amounts of D1 protein (Zubko and Day 1998). Similarly, RNA gel analysis revealed strongly reduced but still detectable amounts of 16S rRNA, which are likely to be generated by the Pc-promoter Prrn16-139 active in white tissues (Fig. 2d).
In this study, we added transcription initiation sites for atpB, atpI, rps4, rps15, rpoB, accD, and ycf1 to the portfolio of plastidial promoters in Arabidopsis (Figs. 3, 4). We scanned the upstream regions of genes for 5′-ends of primary transcripts using a primer walking approach with several reverse primers in intervals of 100-300 bp (Fig. S1 and S2). Our analyses determined PEP promoters (Fig. 5; PatpB-520, PatpI-229, Prps4-123, Pycf1-34) as well as NEP promoters (Fig. 5; PaccD-252, PaccD-172, PatpB-318, PrpoB-300, Prps4-151, Prps15-139, Pycf1-39, Pycf1-104). Although our scan covered a region of about up to position −800, previous data on putative Arabidopsis rpoB (−563; Inada et al. 1997) and accD (−89; Hanaoka et al. 2005) transcription initiation sites derived by primer extension analyses could not be confirmed (see Fig. S1 for data on rpoB; data for accD not shown). However, this does not rule out the possibility of even further upstream located promoters, as was shown for the tobacco ycf2 gene (NtaPycf2-1577; Hajdukiewicz et al. 1997).
The tobacco atpB gene is transcribed from at least three NEP (NtaPatpB-255, −502/−488, −611) and two PEP promoters (NtaPatpB-289, −329; Hajdukiewicz et al. 1997). However, similar to maize (NEP ZmaPatpB-601 and PEP ZmaPatpB-298; Silhavy and Maliga 1998) only one PEP (PatpB-520) and one NEP promoter (PatpB-318) are driving this gene in Arabidopsis. Although sequences of tobacco NtaPatpB-488 and NtaatpB-611 PEP promoters are conserved within the upstream region of the Arabidopsis atpB gene, they are not utilized. This is reminiscent to the diverse promoter usage observed for clpP and rrn16. Although the promoter regions of both genes are well conserved, different types of RNA polymerases and cis-elements are used in distinct plants (Baeza et al. 1991; Sriraman et al. 1998a; reviewed in Liere and Börner 2007).
Interestingly, ycf1 is transcribed from a highly conserved NEP promoter in tobacco and Arabidopsis (Fig. 5b; Pycf1-39, Ntaycf1-41). However, apart from an additional NEP promoter (Pycf1-104) active in both green and white leaves, a PEP promoter (Pycf1-34) located at the NEP promoter position of Pycf1-39 takes over transcription in green leaves. Besides the rrn16 Pc and PEP promoters in Arabidopsis, this is the second report of a defined DNA sequence serving as a promoter for both, NEP and PEP.
However, upstream sequences of some genes, such as accD, atpI, rps4, and rpoB, are not highly conserved. Nevertheless, the accD and rpoB genes of both tobacco and Arabidopsis are transcribed by NEP. Although one NEP promoter precedes the tobacco accD gene (NtaPaccD-129; Hajdukiewicz et al. 1997), two NEP promoters were found to transcribe the Arabidopsis accD gene (PaccD-172, −252). However, rpoB seems to be transcribed from a sole NEP promoter in both plants (PrpoB-300, NtaPrpoB-345; Liere and Maliga 1999). Similarly, as formerly predicted from studies in barley (Hess et al. 1993) and Arabidopsis mutants (Nagashima et al. 2004), the Arabidopsis rps15 is transcribed by NEP from Prps15-139. The spinach rps4 gene was shown to be transcribed by PEP (Tahar et al. 1986) which seems also to be the case in Arabidopsis (Prps4-123). However, an additional NEP promoter (Prps4-151) was also detected in white, spectinomycin-treated Arabidopsis leaves. Conversely, atpI in tobacco is transcribed by both NEP and PEP (NEP NtaPatpI-207, PEP NtaPatpI-130; Miyagi et al. 1998), whereas transcription of the Arabidopsis atpI gene is driven by a sole PEP promoter (PatpI-229). The diversity of individual promoter usage in different plants therefore suggests species-specific solutions in controlling gene expression in plastids.
Although genes exist that are transcribed from a single promoter, transcription of plastidial genes and operons by multiple promoters seems to be a rather common feature (Fig. 6). Furthermore, a number of genes are reported to be co-transcribed with other genes within an operon and to additionally possess an individual promoter upstream of their coding region (e.g. trnG and psbA; reviewed in Liere and Börner 2007). The role of most multiple promoters upstream of plastidial genes and operons is not fully understood; however, some are well characterized. The blue-light-responsive promoter (BRLP) of psbD–psbC is thought to differentially maintain the ability to re-synthesize and replace damaged D2 and CP43 photosystem components in mature chloroplasts (Christopher and Mullet 1994). It has been shown that two promoters are responsible for differential transcription of the petE operon in maize. While monocistronic petE transcripts accumulate under light conditions, usage of a further upstream located promoter results in polycistronic transcripts covering the petL–petE–psaJ gene cluster in the dark (Haley and Bogorad 1990).
Arabidopsis genes with multiple promoters. Schematic synopsis which shows the multiple PEP and NEP promoters of Arabidopsis genes. Boxes represent genes, while open arrowheads denote PEP promoters, filled black arrowheads type-I NEP promoters, filled light gray arrowhead the PclpP-58 type-II NEP promoter, and filled dark gray arrowhead the Prrn16-139 Pc promoter. The promoters are named based on their position in respect to the translation initiation site (+1)
In spite of the observed diversity between plastidial genes of Arabidopsis and other species of higher plants, our data support also the existence of common themes in promoter usage that have been deduced mainly from studies on transcription in tobacco plastids. Mixed NEP and PEP promoters typically are found upstream of housekeeping genes which need to be transcribed during full plastidial development (Maliga 1998). Consequently, both promoter types are believed to differentially express their cognate gene during plant development (reviewed in Liere and Maliga 2001). NEP promoters seem to be generally recognized in youngest and non-green tissues early in plant development, while PEP takes over in maturating photosynthetically active chloroplasts (Bisanz-Seyer et al. 1989; Baumgartner et al. 1993; Hajdukiewicz et al. 1997; Kapoor et al. 1997; Emanuel et al. 2004). Large spurious transcripts initiated by NEP cover the entire plastome in tobacco Δrpo mutants lacking PEP, suggesting that besides selective promoter utilization, post-transcriptional processes also determine the transcript pattern of plastids (Krause et al. 2000; Legen et al. 2002). Data derived from analyses of the developmental gradient in maize leaves suggest that, as plastids mature, the stability of transcripts generated by NEP declines, although the transcriptional activity by NEP increases (Cahoon et al. 2004). However, exclusively NEP-transcribed regions encoding housekeeping functions, such as the rpoB operon and rps15 gene, suggest that NEP is important for proper gene expression and regulation also in mature chloroplasts. Indeed, NEP and PEP are active throughout leaf development in Arabidopsis, although PEP seems to play a major role in mature leaves (Demarsy et al. 2006; Zoschke et al. 2007). Interestingly, exclusively PEP-transcribed genes code for proteins with a role in photosynthesis. As the major active polymerase in mature chloroplasts, present data point to PEP as a prominent target for regulation signals including redox control, not yet determined for NEP (for review see Forsberg et al. 2001; Liere and Maliga 2001; Pfannschmidt and Liere 2005). Since plants that turn to a parasitic lifestyle lose photosynthetic genes as well as PEP promoters (Wolfe et al. 1992a, b; Krause et al. 2003; Berg et al. 2004), transcription and regulation of gene expression by PEP might be connected to photosynthesis. The knowledge of plastidial promoters in Arabidopsis will help to define the role of both the NEP and PEP RNA polymerases in plant development in future experiments.
References
Baeza L, Bertrand A, Mache R, Lerbs-Mache S (1991) Characterization of a protein binding sequence in the promoter region of the 16S rRNA gene of the spinach chloroplast genome. Nucleic Acids Res 19:3577–3581
Baumgartner BJ, Rapp JC, Mullet JE (1993) Plastid genes encoding the transcription/translation apparatus are differentially transcribed early in barley (Hordeum vulgare) chloroplast development: evidence for selective stabilization of psbA mRNA. Plant Physiol 101:781–791
Bensing BA, Meyer BJ, Dunny GM (1996) Sensitive detection of bacterial transcription initiation sites and differentiation from RNA processing sites in the pheromone-induced plasmid transfer system of Enterococcus faecalis. Proc Natl Acad Sci USA 93:7794–7799
Berg S, Krause K, Krupinska K (2004) The rbcL genes of two Cuscuta species, C. gronovii and C. subinclusa, are transcribed by the nuclear-encoded plastid RNA polymerase (NEP). Planta 219:541–546
Binder S, Brennicke A (2003) Gene expression in plant mitochondria: transcriptional and post-transcriptional control. Philos Trans R Soc Lond B Biol Sci 358:181–189
Bisanz-Seyer C, Li Y-F, Seyer P, Mache R (1989) The components of the plastid ribosome are not accumulated synchronously during the early development of spinach plants. Plant Mol Biol 12:201–211
Cahoon AB, Harris FM, Stern DB (2004) Analysis of developing maize plastids reveals two mRNA stability classes correlating with RNA polymerase type. EMBO Rep 5:801–806
Chang C-C, Sheen J, Bligny M, Niwa Y, Lerbs-Mache S, Stern DB (1999) Functional analysis of two maize cDNAs encoding T7-like RNA polymerases. Plant Cell 11:911–926
Christopher DA, Mullet JE (1994) Separate photosensory pathways co-regulate blue light/ultraviolet-A-activated psbD-psbC transcription and light-induced D2 and CP43 degradation in barley (Hordeum vulgare) chloroplasts. Plant Physiol 104:1119–1129
Demarsy E, Courtois F, Azevedo J, Buhot L, Lerbs-Mache S (2006) Building up of the plastid transcriptional machinery during germination and early plant development. Plant Physiol 142:993–1003
Emanuel C, Weihe A, Graner A, Hess WR, Börner T (2004) Chloroplast development affects expression of phage-type RNA polymerases in barley leaves. Plant J 38:460–472
Fey V, Wagner R, Brautigam K, Wirtz M, Hell R, Dietzmann A, Leister D, Oelmuller R, Pfannschmidt T (2005) Retrograde plastid redox signals in the expression of nuclear genes for chloroplast proteins of Arabidopsis thaliana. J Biol Chem 280:5318–5328
Forsberg J, Rosenquist M, Fraysse L, Allen JF (2001) Redox signalling in chloroplasts and mitochondria: genomic and biochemical evidence for two-component regulatory systems in bioenergetic organelles. Biochem Soc Trans 29:403–407
Gray MW (1993) Origin and evolution of organelle genomes. Curr Opin Genet Dev 3:884–890
Gruissem W, Tonkyn JC (1993) Control mechanisms of plastid gene expression. Crit Rev Plant Sci 12:19–55
Gruissem W, Greenberg BM, Zurawski G, Hallick RB (1986) Chloroplast gene expression and promoter identification in chloroplast extracts. Methods Enzymol 118:253–270
Hajdukiewicz PTJ, Allison LA, Maliga P (1997) The two RNA polymerases encoded by the nuclear and the plastid compartments transcribe distinct groups of genes in tobacco plastids. EMBO J 16:4041–4048
Haley J, Bogorad L (1990) Alternative promoters are used for genes within maize chloroplast polycistronic transcription units. Plant Cell 2:323–333
Hanaoka M, Kanamaru K, Takahashi H, Tanaka K (2003) Molecular genetic analysis of chloroplast gene promoters dependent on SIG2, a nucleus-encoded sigma factor for the plastid-encoded RNA polymerase, in Arabidopsis thaliana. Nucleic Acids Res 31:7090–7098
Hanaoka M, Kanamaru K, Fujiwara M, Takahashi H, Tanaka K (2005) Glutamyl-tRNA mediates a switch in RNA polymerase use during chloroplast biogenesis. EMBO Rep 6:545–550
Hess WR, Börner T (1999) Organellar RNA polymerases of higher plants. Int Rev Cytol 190:1–59
Hess WR, Prombona A, Fieder B, Subramanian AR, Börner T (1993) Chloroplast rps15 and the rpoB/C1/C2 gene cluster are strongly transcribed in ribosome-deficient plastids: evidence for a functioning non-chloroplast-encoded RNA polymerase. EMBO J 12:563–571
Hoffer PH, Christopher DA (1997) Structure and blue-light-responsive transcription of a chloroplast psbD promoter from Arabidopsis thaliana. Plant Physiol 115:213–222
Inada H, Seki M, Morikawa H, Nishimura M, Iba K (1997) Existence of three regulatory regions each containing a highly conserved motif in the promoter of plastid-encoded RNA polymerase gene (rpoB). Plant J 11:883–890
Iratni R, Diederich L, Harrak H, Bligny M, Lerbs-Mache S (1997) Organ-specific transcription of the rrn operon in spinach plastids. J Biol Chem 272:13676–13682
Kapoor S, Sugiura M (1999) Identification of two essential sequence elements in the nonconsensus Type II PatpB-290 plastid promoter by using plastid transcription extracts from cultured tobacco BY-2 cells. Plant Cell 11:1799–1810
Kapoor S, Suzuki JY, Sugiura M (1997) Identification and functional significance of a new class of non-consensus-type plastid promoters. Plant J 11:327–337
Khan MS (2005) Unraveling the complexities of plastid transcription in plants. Trends Biotech 23:535–538
Krause K, Berg S, Krupinska K (2003) Plastid transcription in the holoparasitic plant genus Cuscuta: parallel loss of the rrn16 PEP-promoter and of the rpoA and rpoB genes coding for the plastid-encoded RNA polymerase. Planta 216:815–823
Krause K, Maier RM, Kofer W, Krupinska K, Herrmann RG (2000) Disruption of plastid-encoded RNA polymerase genes in tobacco: expression of only a distinct set of genes is not based on selective transcription of the plastid chromosome. Mol Gen Genet 263:1022–1030
Kühn K, Weihe A, Börner T (2005) Multiple promoters are a common feature of mitochondrial genes in Arabidopsis. Nucleic Acids Res 33:337–346
Legen J, Kemp S, Krause K, Profanter B, Herrmann RG, Maier RM (2002) Comparative analysis of plastid transcription profiles of entire plastid chromosomes from tobacco attributed to wild-type and PEP-deficient transcription machineries. Plant J 31:171–188
Lerbs-Mache S (1993) The 110-kDa polypeptide of spinach plastid DNA-dependent RNA polymerase: single-subunit enzyme or catalytic core of multimeric enzyme complexes? Proc Natl Acad Sci USA 90:5509–5513
Liere K, Börner T (2007) Transcription of plastid genes. In: Grasser KD (ed) Regulation of transcription in plants. Blackwell, Oxford, pp 184–224
Liere K, Kaden D, Maliga P, Börner T (2004) Overexpression of phage-type RNA polymerase RpoTp in tobacco demonstrates its role in chloroplast transcription by recognizing a distinct promoter type. Nucleic Acids Res 32:1159–1165
Liere K, Kestermann M, Müller U, Link G (1995) Identification and characterization of the Arabidopsis thaliana chloroplast DNA region containing the genes psbA, trnH and rps19′. Curr Genet 28:128–130
Liere K, Maliga P (1999) In vitro characterization of the tobacco rpoB promoter reveals a core sequence motif conserved between phage-type plastid and plant mitochondrial promoters. EMBO J 18:249–257
Liere K, Maliga P (2001) Plastid RNA Polymerases. In: Andersson B, Aro E-M (eds) Regulation of photosynthesis. Kluwer, Dordrecht, pp 29–49
Link G (1994) Plastid differentiation: organelle promoters and transcription factors. In: Nover L (ed) Plant promoters and transcription factors—results and problems in cell differentiation. Springer, Berlin, pp 65–85
López-Juez E, Pyke KA (2005) Plastids unleashed: their development and their integration in plant development. Int J Dev Biol 49:557–577
Maliga P (1998) Two plastid polymerases of higher plants: an evolving story. Trends Plant Sci 3:4–6
Meng BY, Tanaka M, Wakasugi T, Ohme M, Shinozaki K, Sugiura M (1988) Cotranscription of the genes encoding two P700 chlorophyll a apoproteins with the gene for ribosomal protein CS14: determination of the transcriptional initiation site by in vitro capping. Curr Genet 14:395–400
Miyagi T, Kapoor S, Sugita M, Sugiura M (1998) Transcript analysis of the tobacco plastid operon rps2/atpI/H/F/A reveals the existence of a non-consensus type II (NCII) promoter upstream of the atpI coding sequence. Mol Gen Genet 257:299–307
Moazed D, Noller HF (1987) Interaction of antibiotics with functional sites in 16S ribosomal RNA. Nature 327:389–394
Nagashima A, Hanaoka M, Motohashi R, Seki M, Shinozaki K, Kanamaru K, Takahashi H, Tanaka K (2004) DNA microarray analysis of plastid gene expression in an Arabidopsis mutant deficient in a plastid transcription factor sigma, SIG2. Biosci Biotechnol Biochem 68:694–704
Pfannschmidt T, Liere K (2005) Redox regulation and modification of proteins controlling chloroplast gene expression. Antioxid Redox Signal 7:607–618
Reznikov W, Siegle DA, Cowing DW, Gross CA (1985) The regulation of transcription initiation in bacteria. Annu Rev Genet 19:355–387
Serino G, Maliga P (1998) RNA polymerase subunits encoded by the plastid rpo genes are not shared with the nucleus-encoded plastid enzyme. Plant Physiol 117:1165–1170
Shen Y, Danon A, Christopher DA (2001) RNA binding-proteins interact specifically with the Arabidopsis chloroplast psbA mRNA 5′ untranslated region in a redox-dependent manner. Plant Cell Physiol 42:1071–1078
Shiina T, Tsunoyama Y, Nakahira Y, Khan MS (2005) Plastid RNA polymerases, promoters, and transcription regulators in higher Plants. In: Int Rev Cytol, Academic Press, New York, pp 1–68
Sikdar SR, Serino G, Chaudhuri S, Pal M (1998) Plastid transformation in Arabidopsis thaliana. Plant Cell Rep V 18:20–24
Silhavy D, Maliga P (1998) Mapping of the promoters for the nucleus-encoded plastid RNA polymerase (NEP) in the iojap maize mutant. Curr Genet 33:340–344
Sriraman P, Silhavy D, Maliga P (1998a) The phage-type PclpP-53 plastid promoter comprises sequences downstream of the transcription initiation site. Nucleic Acids Res 26:4874–4879
Sriraman P, Silhavy D, Maliga P (1998b) Transcription from heterologous rRNA operon promoters in chloroplasts reveals requirement for specific activating factors. Plant Physiol 117:1495–1499
Tahar SB, Bottomley W, Whitfeld PR (1986) Characterization of the spinach chloroplast genes for the S4 ribosomal protein, tRNAThr (UGU) and tRNASer (GGA). Plant Mol Biol 7:63–70
Vera A, Sugiura M (1995) Chloroplast rRNA transcription from structurally different tandem promoters: an additional novel-type promoter. Curr Genet 27:280–284
Weihe A (2004) The transcription of plant organelle genomes. In: Daniell H, Chase CD (eds) Molecular biology and biotechnology of plant organelles. Kluwer, Dordrecht, pp 213–237
Weihe A, Börner T (1999) Transcription and the architecture of promoters in chloroplasts. Trends Plant Sci 4:169–170
Wolfe KH, Morden CW, Ems SC, Palmer JD (1992a) Rapid evolution of the plastid translational apparatus in a nonphotosynthetic plant: loss or accelerated sequence evolution of tRNA and ribosomal protein genes. J Mol Evol 35:304–317
Wolfe KH, Morden CW, Palmer JD (1992b) Function and evolution of a minimal plastid genome from a nonphotosynthetic parasitic plant. Proc Natl Acad Sci USA 89:10648–10652
Zoschke R, Liere K, Börner T (2007) From seedling to mature plant: Arabidopsis plastidial genome copy number, RNA accumulation and transcription are differentially regulated during leaf development. Plant J:in press
Zubko MK, Day A (1998) Stable albinism induced without mutagenesis: a model for ribosome free plastid inheritance. Plant J 15:265–271
Acknowledgments
We thankfully acknowledge the participation of Daniela Kaden and Kristina Kühn in the initial phase of our studies on initiation sites of chloroplast transcripts. This work was supported by grants from the Deutsche Forschungsgemeinschaft (SFB429).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Herrmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Swiatecka-Hagenbruch, M., Liere, K. & Börner, T. High diversity of plastidial promoters in Arabidopsis thaliana . Mol Genet Genomics 277, 725–734 (2007). https://doi.org/10.1007/s00438-007-0222-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-007-0222-4