Abstract
Spatially explicit capture–recapture (SECR) models are gaining popularity for estimating densities of mammalian carnivores. They use spatially explicit encounter histories of individual animals to estimate a detection probability function described by two parameters: magnitude (g 0), and spatial scale (σ). Carnivores exhibit heterogeneous detection probabilities and home range sizes, and exist at low densities, so g 0 and σ likely vary, but field surveys often yield inadequate data to detect and model the variation. We sampled American black bears (Ursus americanus) on 43 study areas in ON, Canada, 2006–2009. We detected 713 animals 1810 times; however, study area-specific samples were sometimes small (6–34 individuals detected 13–93 times). We compared AIC c values from SECR models fit to the complete data set to evaluate support for various forms of variation in g 0 and σ, and to identify a parsimonious model for aggregating data among study areas to estimate detection parameters more precisely. Models that aggregated data within broad habitat classes and years were supported over those with study area-specific g 0 and σ (ΔAIC c ≥ 30), and precision was enhanced. Several other forms of variation in g 0 and σ, including individual heterogeneity, were also supported and affected density estimates. If study design cannot eliminate detection heterogeneity, it should ensure that samples are sufficient to detect and model it. Where this is not feasible, combing sparse data across multiple surveys could allow for improved inference.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Reliable information about population size is fundamental to the study of ecology and is necessary for effective conservation and management of wildlife populations. Bears and other large and medium-sized mammalian carnivores are notoriously difficult to enumerate because they range widely, occur at low densities, exhibit heterogeneous probabilities of detection and movement rates, and are often secretive or elusive (Garshelis 1992; Karanth 1995; Boulanger et al. 2004; MacKenzie et al. 2005; Kéry et al. 2011; Boitani and Powell 2012). Nevertheless, many carnivore populations are managed intensively because they pose conservation concerns, come into conflict with humans, or are subject to sport harvest. The potential to identify individuals from photographic or genetic detections has enabled researchers to collect sufficient capture–recapture (C–R) data to estimate population size from data collected over days or weeks, whereas multiple years of live-capture data were required previously (Karanth 1995; Woods et al. 1999). Estimates of population density are often preferred over estimates of population size because the former is independent of scale and comparable across studies. However, where occupied habitat extends beyond the study area, population size cannot be reliably converted to density because the area sampled is unknown (Dice 1938; White et al. 1982; Efford 2004). This is particularly problematic in studies of wide-ranging species including most mammalian carnivores (White et al. 1982; Garshelis 1992; Boulanger et al. 2002; Foster and Harmsen 2012).
Spatially explicit capture–recapture (SECR) is a recently-developed modeling framework for estimating animal density directly from spatially explicit C–R data collected on geographically open study areas, without estimating the area sampled (Efford 2004; Borchers and Efford 2008; Gardner et al. 2009). Density is estimated as the intensity of a spatial point process where the points are the unobserved central locations of home ranges, or “activity centers” (Efford 2004; Borchers and Efford 2008; Gardner et al. 2009). The models can accommodate different types of traps including live-traps, camera traps, and passive DNA sampling devices (Efford et al. 2009). They assume demographic closure, independence of captures or detections, and that individuals occupy home ranges, the central locations of which are independent and fixed during sampling (Efford 2004; Gardner et al. 2009). Probability of detection is modeled as a decreasing function of the distance between traps and the unobserved activity centers (Efford 2004; Gardner et al. 2009). The simplest model has two parameters: the magnitude (g 0) and spatial scale (σ) of a half-normal spatial detection probability function; g 0 may be thought of as the probability of detection where a trap is placed at an individual’s activity center (i.e., at distance zero), and σ and the shape of the detection probability function describe how detection probability declines with increasing distance between home range center locations and traps (Efford 2004). By treating spatially variable exposure to traps explicitly, SECR models account for a major source of individual heterogeneity in the detection process (Garshelis 1992; Boulanger et al. 2004; Efford 2004; Royle et al. 2009). Furthermore, both g 0 and σ may be described as functions of covariates.
Several researchers recently concluded that SECR estimates of carnivore density were preferable to those obtained by dividing estimates of population size by estimates of the area sampled (Gardner et al. 2009; Royle et al. 2009, 2011; Obbard et al. 2010; Sollmann et al. 2011; Gerber et al. 2012; Noss et al. 2012). However, due to both the above-described characteristics of carnivores, and the fact that many factors and constraints influence study design, sample sizes and detection probabilities obtained from surveys of mammalian carnivores are frequently insufficient to detect and model variation in detection probabilities (McKelvey and Pearson 2001; Boulanger et al. 2002; Proctor et al. 2010; Doherty et al. 2012; Foster and Harmsen 2012; Gervasi et al. 2012; Sollmann et al. 2012). Consequently, most SECR studies published to date presented results from only one or a few simple models, often fit to sparse data (Gardner et al. 2009, 2010; Royle et al. 2009, 2011; Kéry et al. 2011; Kalle et al. 2011; O’Brien and Kinnaird 2011; Sollmann et al. 2011; Gray and Prum 2012; Noss et al. 2012). Therefore, the sensitivity of SECR density estimates to variation in the parameters of the detection probability function has not been adequately assessed using empirical data. This is problematic because large carnivores frequently exhibit heterogeneous detection probabilities, including among individuals, beyond what can be explained by variable exposure to traps (Noyce et al. 2001; Boulanger et al. 2004; Ebert et al. 2010; Gardner et al. 2010; Obbard et al. 2010; Sollmann et al. 2011). Furthermore, densities estimated from sparse data may not be sufficiently precise to inform management (Ebert et al. 2010; Foster and Harmsen 2012).
One approach to improving estimator performance where data from individual surveys are inadequate to estimate detection probabilities, but data from multiple surveys are available, is to combine the data and estimate those probabilities from all surveys simultaneously (Boulanger et al. 2002; MacKenzie et al. 2005; White 2005). This provides more power to detect and model variation among individuals and sampling occasions. Rather than assuming common detection probabilities across surveys, support for models representing different levels of aggregation may be compared using model selection criteria. One goal of model selection becomes the identification of a model of variation among surveys that describes the data reasonably well, but allows some aggregation to improve precision (Boulanger et al. 2002; MacKenzie et al. 2005; White 2005; Conn et al. 2006). With more aggregation, precision is enhanced at the expense of possible bias (White 2005).
We sampled female American black bears (Ursus americanus) on 43 study areas across ON, Canada (2006–2009), with the aim of providing local density estimates for management purposes. The main objective of the current study was to evaluate support for various forms of variation in the spatial detection probability function, and assess the effect of modeling them on density estimates. In particular, we sought a parsimonious model of variation among study areas that would allow us to estimate local densities more precisely by aggregating the data. We compared AIC c values among candidate SECR models that allowed g 0 and σ to vary among study areas, individuals, sampling occasions, and in response to initial detection, to parameter-reduced models, including models which combined data across all study areas, or subsets of study areas in similar habitat, to estimate detection parameters. We expect these results to be of interest to those attempting to enumerate large carnivores because reliable estimates are required to inform management, but obtaining sample sizes necessary to detect and model heterogeneity in the detection function and yet estimate density with reasonable precision from individual C–R surveys of large carnivores is challenging, and not always feasible.
Methods
Field sampling
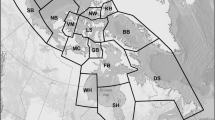
We conducted standardized noninvasive capture–recapture surveys of black bears on 43 study areas in the Boreal and Great Lakes-St. Lawrence (GLSL) forest regions (Rowe 1972) of ON, Canada (Fig. 1) during 2006–2009. Different study areas were sampled each year. During spring and early summer, black bears typically occupy stable home ranges and focus their activities within a smaller home range “core area”; however, they frequently make long-distance excursions from their breeding range when energy rich foods such as berries and nuts become available, typically beginning in mid-July (Rogers 1987; Powell et al. 1997; Schenk et al. 1998; Noyce and Garshelis 2011). We therefore sampled from late May through June (the latest any samples were collected was 06 July) in order to meet the SECR assumptions of demographic closure and static home range center locations (Efford 2004). Most study areas had been established previously for population monitoring by bait-station index lines in habitat representative of the respective management unit (McLaren et al. 2005). On each study area, we snagged bear hair at 20–25 barbed wire hair corrals (Woods et al. 1999), except on three study areas where 15, 17, and 18 corrals were used. Corrals were spaced approximately 2 km apart in curvilinear arrays along secondary roads, 20–200 m from the road itself. We baited corrals with 3 partially-opened tins of sardines in oil suspended from a board nailed 2.5 m high on a tree >2 m from any point along the wire. We collected samples and re-baited corrals one week later, and repeated this for a total of 5 sampling occasions on 36 study areas; bears were sampled on only 4 occasions on 4 study areas, and on six occasions on 3 other study areas. We air dried hair samples in paper envelopes and stored them at room temperature until DNA extraction.
Map of the greater study area in ON, Canada. The Great Lakes are shaded dark grey. Trap locations are marked with black dots; at this scale traps arrays appear as thick black lines surrounded by gray shading which depicts 10 km buffers around all traps within each study area. Rowe’s (1972) Forest region boundary between Boreal and Great Lakes-St. Lawrence Forests is depicted as a 20 km-thick light grey line. Thinner, darker grey lines show Ecoprovince boundaries between the Mid-Boreal Shield in the north, and each of the Lake of the Woods and Southern Boreal Shield to the south (Marshall et al. 1998)
DNA analysis
We did not attempt to extract DNA from samples with fewer than 5 guard hairs with visible roots; >90 % of samples were processed using 10–15 hairs to minimize technical artifacts from low template DNA. When the number of samples exceeded what we could analyze with available resources, we excluded samples collected from adjacent barbs at the same trap on the same occasion. Individuals were identified from their microsatellite genotypes at 10–15 polymorphic loci (Paetkau and Strobeck 1994; Paetkau et al. 1995; Taberlet et al. 1997; Kitahara et al. 2000), and sex was determined from amplification of a region of the Amelogenin gene (Ennis and Gallagher 1994). For more detailed DNA extraction, amplification, and profiling methods and criteria for ascribing samples to individuals see Obbard et al. (2010) and Pelletier et al. (2012).
Data analysis
We excluded males from statistical analyses because we were concerned that our study areas might be too small to estimate densities of male bears reliably as the size of their home ranges could approach or exceed that of our study areas (Alt et al. 1980; Koehler and Pierce 2003; Efford 2011; Marques et al. 2011). We assumed that the height of the barbed wire strand (~50 cm) would exclude cubs and yearlings from the sample (Woods et al. 1999).
We generated study area-specific integration meshes (see Borchers and Efford 2008, Efford et al. 2009) that extended 10 km around all traps on each study area and excluded points that would have fallen in lakes. Mesh points were spaced 0.8–1.0 km apart. We verified that the extent and resolution of our integration meshes were sufficient to avoid bias by fitting SECR models to data from two study areas with >25 recaptures and movement rate estimates near the minimum and maximum across study areas, while varying the extent of the mask and the spacing of mask points. Extents ranged from 5–15 km in increments of 2.5 km; the spacing of points ranged from 0.4–1.2 km in increments of 0.2 km. Density estimates and their coefficients of variation were insensitive to increases in the extent of the mesh beyond 10 km, or with reductions in point spacing below 1.0 km.
We analyzed data from all study areas simultaneously. In cases where study areas were <20 km apart, we verified that no individuals were detected on >1 study area. Study areas were modeled as groups or “sessions” in a multi-session analysis that allowed data to be pooled across study areas to estimate g 0 and σ (Efford et al. 2009). We assumed the total number of individuals was binomially-distributed on each study area. We estimated g 0 and σ by maximizing the conditional likelihood for proximity detectors, and estimated density as a derived parameter using a Horvitz–Thompson-like estimator (see Borchers and Efford 2008; Efford et al. 2009). We used the half-normal form of the detection function, which we suspected would reasonably approximate the above-described home range characteristics of female black bears.
We defined a set of candidate models of variation in g 0 and σ based on previously published information about the probabilities of detection, home ranges, and movements of black bears (Garshelis and Pelton 1980; Rogers 1987; Powell et al. 1997; Noyce et al. 2001; Koehler and Pierce 2003; Mowat et al. 2005; Obbard et al. 2010). We used the small-sample corrected version of Akaike’s Information Criterion (AIC c ; Hurvitch and Tsai 1989) to identify the most parsimonious models in the candidate set. We assessed support for general and trap-specific responses to prior detection affecting g 0, (hereafter denoted b and bk, respectively) and differences in g 0 among individuals (h), years, and study areas. For σ, we considered differences among individuals, sampling occasions (t), years, study areas, and 4 different patterns of variation among habitat types. Individual heterogeneity was modeled using two-point finite mixture distributions (Pledger 2000; Borchers and Efford 2008). Three of the habitat type covariates used Rowe’s (1972) Forest Regions to ascribe study areas to different habitat types; the fourth used Ecoprovince boundaries (“ECOP”; Marshall et al. 1998; see Fig. 1). Habitat productivity for black bears is superior in eastern GLSL than Boreal Forests (Rowe 1972; Kolenosky 1990; Obbard and Howe 2008). We hypothesized that σ would be smaller in GLSL than Boreal Forest Regions, and might also vary between eastern and western GLSL Forests, and that bears in the Lake of the Woods and Southern Boreal Shield Ecoprovinces might have lower σ than those in the Mid-Boreal Shield. The simplest habitat covariate (FR2) had two levels, and discriminated only between Boreal and GLSL Forests. FR3 further separated eastern from western GLSL forests. FR3I combined eastern and western GLSL forests, but included a separate “intermediate” level for study areas that fell within 10 km of the Forest Region boundary. We fit a total of 76 models. The most constrained models in the candidate set were the null model and models with each form of variation in isolation. The most general model had 173 parameters and allowed for study area-specific g 0 and σ, and study area-specific differences among individuals (equivalent to fitting a model with h affecting g 0 and σ to each study area-specific data set). The most general additive model included differences in g 0 among individuals, after initial detection, and among study areas, and differences in σ among sampling occasions, individuals and study areas. Model fitting was prohibitively time-consuming on a stand-alone desktop computer. To reduce the total number of models, we initially emphasized the simplest habitat covariate, and later crossed other habitat covariates with the best-supported models of variation among individuals, years, sampling occasions, and in response to initial detection. We used the facilities of the high performance computing network “SHARCNET” (http://www.sharcnet.ca) to fit many models simultaneously. Integration meshes were generated using program DENSITY (version 4.4.5.1; Efford et al. 2004; Efford 2010); all other analyses were performed in the R programming environment version 2.15 (R Development Core Team 2012) using the “secr” package version 2.4.0 (Efford 2012).
Results
Field sampling and DNA analysis
The number of hair samples collected on each study area ranged from 104 to 860, from which we obtained 38 to 352 genetic detections (including multiple detections of the same individual at the same trap and occasion; Table 1). The number of unique females detected on each study area ranged from 6 to 34, and the total number of detections of females included in spatially explicit encounter histories (i.e., excluding multiple detections of the same individual at the same trap and occasion) ranged from 13 to 93 (Table 1). The mean of maximum distances moved among traps ranged from 881 to 5459 m, and the mean distance between successive detection locations ranged from 352 to 2677 m (Table 1).
Data analysis
Models of detection heterogeneity that minimized AIC c included a trap-specific response to initial detection and differences among individuals and years affecting g 0, and differences in σ among individuals, years, sampling occasions, and habitat types (Table 2). Model selection uncertainty was limited to whether σ differed among years, and which habitat type covariate best approximated variation in σ among study areas (Table 2). Models with study area specific g 0 and σ were not supported (Table 2). The top-ranked model that excluded h in either parameter (ΔAIC c = 108) had the same structure otherwise as the top-ranked model. The most general model, with study area-specific g 0, σ, and effects of h, had ΔAIC c = 409. Variance calculation failed when this model was fit, and parameter estimates for some study areas were not identifiable or were at a boundary. The null model ranked last (ΔAIC c = 439).
Densities estimated from the top 8 AIC c -ranked models (Σw i = 1.00) were similar in magnitude (mean \( \hat{D} \) across study areas estimated from 1st–8th ranked models, respectively = 11.9, 12.2, 11.2, 11.0, 9.8, 9.4, 12.1, and 11.3), precision (mean CV across study areas = 0.27 from all 4 models that included differences in σ among years, and = 0.26 from models that did not), and the pattern of variation among study areas (Spearman’s rank correlation coefficients from pairwise comparisons of \( \hat{D} \) ranged from 0.903 to 0.986). Correlations among estimates from different models were strongest, ranging from 0.976 to 0.986, for the top 4 AIC c -ranked models (Σw i = 0.94). \( \hat{D} \) from supported models ranged from 3 to 30 females aged >1 year/100 km2 and was generally higher in more productive habitat, but variable in eastern GLSL forests (Fig. 2).
Densities of female black bears aged >1 year (per 100 km2) on 43 study areas in ON, Canada, 2006–2009, estimated from the top AIC c -ranked spatially explicit capture–recapture model, within different habitat types (see Fig. 1). Study area-specific estimates were grouped into habitat types four ways (see Fig. 1): (1) Boreal and Great Lakes-St. Lawrence (GLSL) forests (a), (2) Boreal forests and GLSL forests in eastern and western ON (b), Boreal Forests, GLSL Forests, and study areas within 10 km of the Forest region boundary (c), and Southern Boreal Shield, Mid-Boreal Shield, and Lake of the Woods Ecoprovinces (d)
Densities estimated from high-ranking models were similar because these models all had a similar structure. However, \( \hat{D} \) varied when different forms of variation in g 0 and σ were modeled (Table 3). Individual heterogeneity increased \( \hat{D} \) and reduced its precision, but the effect size was smaller where other forms of variation were also included in the estimating model (Table 3). Forest region and year effects altered the pattern of variation in \( \hat{D} \) among study areas. For example, lower σ in GLSL Forests than in Boreal Forests was associated with higher \( \hat{D} \) in the former (Tables 3, 4). Differences in g 0 among years reduced \( \hat{D} \) on study areas sampled in all years other than 2006, and differences in σ among years increased \( \hat{D} \) in 2006 relative to estimates from models without year effects (Table 3). Models with study area effects yielded density estimates which differed considerably from estimates from supported models in some cases (Table 3), and were relatively imprecise (mean CV across study areas from the highest-ranking model with study area effects on either parameter = 0.39). Effects of initial detection on g 0, and variation in σ among sampling occasions had little effect on \( \hat{D} \) here (Table 3).
Parameter estimates from high-ranking models showed that g 0 increased in response to initial detection at the same trap (Table 4). Animals traveled farther to encounter traps in Boreal than GLSL forests; bears on study areas within 10 km of the forest region boundary had intermediate σ (Table 4). Parameter estimates from the highest-ranked models with ECOP and FR3 covariates indicated that σ was lower in the Lake of the Woods Ecoprovince than in the Southern Boreal Shield, and in Western than Eastern GLSL Forests (β SBS = 0.089, SE 0.061, β MBS = 0.335, SE 0.086; β GLSL E = −0.069, SE 0.063, β GLSL W = −0.187, SE 0.078). Bears also traveled farther on successive sampling occasions during the first 5 weeks of sampling (the estimate of σ on occasion 6 was based on data from only 3 study areas and was imprecisely estimated; Table 4). Models with differences in σ among years indicated that bears did not travel as far to encounter traps in 2006 as in other years (Table 4) Year effects indicated and that g 0 was highest in 2008 and lowest in 2006 (Table 3). Year effects were imprecise except in the case of lower g 0 in 2006 (Table 4).
Approximately 13 % of individuals were assigned to the 2nd mixture of individuals with higher g 0 and σ than other individuals (Table 4). Point estimates of g 0 from the top AIC c -ranked model for the first mixture of individuals were 0.29 in 2008 and 0.10 in 2006, but increased to 0.52 and 0.23 respectively after initial detection (Table 4). Estimates of the same parameters for the second mixture of individuals were 0.64 in 2008 and 0.34 in 2006, and 0.82 and 0.56 after initial detection. Point estimates of σ on occasion 5 in 2008 for the first mixture of individuals were 1395 m in GLSL forests, 1876 m in Boreal forests, and 1539 m on study areas within 10 km of the Forest Region boundary; concurrent estimates for the second mixture were 2040, 2743, and 2250 m (Table 3).
Discussion
Capture–recapture surveys of mammalian carnivores frequently yield samples that provide insufficient information to select among candidate models or to estimate abundance precisely from models that allow for important forms of variation (Norris and Pollock 1995; Boulanger et al. 2002, 2004; Link 2004; Royle et al. 2009; Ebert et al. 2010; Kéry et al. 2011; Marucco et al. 2011; O’Brien and Kinnaird 2011; Sollmann et al. 2011; Doherty et al. 2012; Foster and Harmsen 2012; Gray and Prum 2012). Small samples are a consequence of both the biological characteristics of carnivores, and the fact that study designs and their ability to provide moderate to large samples are frequently constrained by factors such as habitat fragmentation, small population size, limited knowledge of the population under study, the need to place traps in specific habitats or along known travel corridors to maximize detection probabilities, conflicting research priorities, or limited funding (Boitani and Powell 2012). Sharing information across surveys increases power to detect heterogeneity and improves precision at the possible expense of bias (White 2005; Anderson 2008). Our results demonstrate empirically that by combining sparse data from multiple standardized surveys, researchers may be able to detect and model forms of variation in SECR model parameters that are likely to be present in C–R data from carnivores, and yet estimate densities with reasonable precision.
Snagging hairs of American black bears on 5 occasions on our curvilinear arrays of 15–25 traps yielded samples that were insufficient to model heterogeneity or estimate density precisely enough to inform management on some study areas. Furthermore, we sometimes obtained few repeat detections of the same individuals at different traps, particularly nonadjacent traps, which may have rendered our estimates of σ prone to sampling error. The use of habitat and year covariates to describe variation in detection among study areas, and model selection using AIC c , allowed us to identify a model that allowed for variation among study areas but also improved the precision of \( \hat{D} \) by aggregating data to estimate some of the parameters of the detection function. Of course, there remains the potential for bias introduced by pooling the data across study areas. Different models of variation in detection probabilities among study areas, or more general combinations of the covariates we considered (e.g., including interactions) might have been supported had we considered them.
SECR represents an improvement over boundary strip methods for estimating carnivore density, especially where a conservation concerns exists because the latter are prone to overestimation (Gardner et al. 2009; Obbard et al. 2010; Sollmann et al. 2011; Gerber et al. 2012; Noss et al. 2012). However, considerable variation in \( \hat{D} \) estimated from different models fit to the same data in our study demonstrates the potential to obtain biased density estimates by fitting an inappropriate SECR model. In particular, individual heterogeneity poses one of the greatest challenges to researchers attempting to estimate animal abundance from C–R data (Davis et al. 2003; Link 2004; Lukacs and Burnham 2005; Ebert et al. 2010; Marucco et al. 2011). SECR models represent an improvement in this regard because they treat heterogeneity associated with variable exposure to traps explicitly (Efford 2004; Royle et al. 2009). In our study, individual heterogeneity was still unambiguously supported even though we used a spatial detection model and limited the data set to females. Innate differences in cautious behavior (DeBruyn 1999; Noyce et al. 2001) or differences among bears of different ages or social classes (Miller et al. 1997; Woods et al. 1999; Noyce et al. 2001; Boulanger et al. 2006) could have caused g 0 to vary among individuals independently of spatial effects. Home range sizes of female black bears also vary with age and social status (Alt et al. 1980; Rogers 1987; Wooding and Hardisky 1994; Costello 2008). Most previous SECR analyses of carnivore data did not attempt to model individual heterogeneity, except, in some cases, for differences between sexes (Gardner et al. 2009, 2010; Royle et al. 2009, 2011; Kalle et al. 2011; Kéry et al. 2011; Sollmann et al. 2011, 2012; Gray and Prum 2012; Noss et al. 2012). However, carnivores other than black bears also typically exhibit individual heterogeneity in detection probabilities and movement rates beyond what can be explained by spatial effects and sex, including differences among age and social classes (Boulanger et al. 2002, 2004, 2006; Cubaynes et al. 2010; Ebert et al. 2010; Marucco et al. 2011; Sollmann et al. 2012). O’Brien and Kinnaird (2011) attributed lack of support for individual heterogeneity in SECR data from four species of African carnivores to small sample size rather than a lack of differences among individuals, and Sollmann et al. (2012), in an SECR analysis of black bear data, noted that there was much residual variation in sex-specific SECR model parameters that may have been attributable to individual heterogeneity.
As age and social status cannot be inferred from genetic samples, we must rely on statistical approaches to correct for the associated variation. This requires larger samples to achieve similar precision (Pledger 2000; Dorazio and Royle 2003) and raises concerns about the sensitivity of density estimates to how individual heterogeneity is modeled (Dorazio and Royle 2003; Link 2004). These problems should not lead us to abandon models that correct for heterogeneity, because if it is present in the data, such models will generally yield more accurate estimates with better confidence interval coverage (Pledger 2000, 2005; Dorazio and Royle 2003; Boulanger et al. 2004; Link 2004; Cubaynes et al. 2010; Proctor et al. 2010). If a reduced-parameter model is selected a priori or because the power to detect variation is low, associated density estimates are not only potentially inaccurate, but also overstate precision (White et al. 1982; Boulanger et al. 2004). Proctor et al. (2010) and Ebert et al. (2010) recommended considering models that allow for individual heterogeneity for estimation purposes even when tests or model selection criteria do not detect it, because it is likely present in noninvasive data sets but power to detect it is often low. Statisticians might question the validity of such an approach, and in any case the parameters of SECR models that allow for individual heterogeneity may be inestimable when data are sparse (O’Brien and Kinnaird 2011; Sollmann et al. 2012; this study). Among the forms of variation we considered, individual heterogeneity had the greatest effect on \( \hat{D} \) when modeled. Furthermore, variation among individuals was apparently overestimated, leading to even higher \( \hat{D} \), when other forms of variation present in the data were not modeled explicitly. We agree with those who argued that if individual heterogeneity cannot be eliminated by study design or explained using covariates, then it is key to obtain sample sizes and detection probabilities sufficient to detect and model it (Boulanger et al. 2002; Lukacs and Burnham 2005; Marucco et al. 2011). More generally, we recommend treating variation in SECR model parameters as thoroughly as has become standard practice in conventional C–R studies. Where only sparse data from a single survey are available, SECR models that ignore individual heterogeneity could still be used to generate conservative estimates for management purposes because correcting for individual heterogeneity increases \( \hat{D} \).
Although estimating abundance is the main goal of most C–R studies, identifying models of variation in g 0 and σ that minimize AIC c allows researchers to evaluate support for competing hypotheses about animal behavior (Kéry et al. 2011; Sollmann et al. 2011). For instance, comparisons of home range sizes of a species in different parts of its range or at different densities are of interest, but are often confounded by small samples of instrumented animals and differences in home range estimation methods among studies (Powell et al. 1997). In the case of American black bears, abundant prior research allows us to verify that our model selection results are consistent with independent information. For example, positive effects of initial detection were common where trap sites were baited (Boersen et al. 2003; Dreher et al. 2007; Immell and Anthony 2008; Gardner et al. 2010). The presence and magnitude of behavioral responses to detection at hair corrals may be related to the reliability and quantity of the food reward. Our baits (3 tins of sardines) provided a small food reward, were sometimes consumed by non-target species, and otherwise were likely consumed by the first bear to visit the trap; nevertheless, g 0 approximately doubled after initial detection. Responses to initial detection were not supported where the same bait was used but sampling occasions were separated by week-long intervals with no bait present (Obbard et al. 2010), or in two studies where a lure that provided little or no food reward was used (Belant et al. 2005; Sollmann et al. 2012). In one study where baits provided a larger food reward, the positive response was even stronger than that observed here (Wegan 2008; Gardner et al. 2010). A positive response to initial detection affecting g 0 had very slight effects on \( \hat{D} \) here, but other studies suggest that failure to model such responses could cause underestimation (Borchers and Efford 2008; Gardner et al. 2010). We did not test for effects of initial detection on σ because we did not expect the small food reward to cause female black bears to deviate from their normal spring and early summer movement patterns, which are strongly influenced by social factors (Rogers 1987; Costello 2008; Castle 2010). However, the possibility that bears would move outside their normal home range in search of, or following the scent of, a baited trap after encountering one within it may warrant further investigation.
That σ increased during successive sampling occasions is consistent with telemetry studies that demonstrated increasing movement rates and home range sizes of black bears before and during the breeding season in early summer (Alt et al. 1980; Garshelis and Pelton 1980; Rogers 1987; Castle 2010). Black bears have also been shown to have smaller home ranges in higher quality habitat (Powell et al. 1997; Jones and Pelton 2003; Koehler and Pierce 2003). GLSL Forests provide more productive bear habitat than Boreal Forests (Rowe 1972; Kolenosky 1990; Obbard and Howe 2008) so higher σ in Boreal than GLSL forests was also expected. Possible explanations for the supported differences in g 0 and σ among years include differences in proportion of females accompanied by cubs if reproduction was synchronous, differences in food abundance during sampling or in the previous foraging season, differences in the timing of the onset of spring and associated bear behaviors, and effects of weather conditions on the rate of DNA degradation between the time samples were deposited and collected.
The patterns of variation in female bear density and home range size across ON described by our estimates are consistent with prior information indicating that both are related to habitat quality—the former directly and the latter inversely (Lindzey et al. 1986; Garshelis 1994; Powell et al. 1997; Koehler and Pierce 2003), though we note that our density estimates are partly a function of the way spatial variation in σ was modeled. Reproductive rates of black bears are generally higher in higher quality habitat (Garshelis 1994) but may also increase where densities are reduced by anthropogenic mortality (Czetwertynski et al. 2007; Obbard and Howe 2008); whether these latter increases are a consequence of increased home range size due to reduced competition for space or patchily-distributed food resources is not well-understood. That some of the lowest densities occurred in more southerly portions of the greater study area in eastern GLSL forests likely reflects greater habitat fragmentation and increased anthropogenic mortality near the southern limit of contiguous forests where human population density is higher and agricultural and urban development are more common. If black bears increase their home range size where densities are reduced, anthropogenic effects on bear density in the southeast may also explain why σ was larger on average in eastern than western GLSL forests.
References
Alt GL, Matula GJ Jr, Alt FW, Lindzey JS (1980) Dynamics of home range and movements of adult black bears in northeastern Pennsylvania. Int Conf Bear Res Manage 4:131–136
Anderson DR (2008) Model based inference in the life sciences: a primer on evidence. Springer, New York
Belant JL, Van Stappen JF, Paetkau D (2005) American black bear population size and genetic diversity at Apostle Islands National Lakeshore. Ursus 16:85–92
Boersen MR, Clark JD, King TL (2003) Estimating black bear population density and genetic diversity at Tensas River, Louisiana using microsatellite DNA markers. Wildl Soc Bull 31:197–207
Boitani N, Powell RA (eds) (2012) Carnivore ecology and conservation: a handbook of techniques. Oxford University Press, Oxford
Borchers DL, Efford MG (2008) Spatially explicit maximum likelihood methods for capture–recapture studies. Biometrics 64:377–385
Boulanger J, White GC, McLellan BN, Woods J, Proctor M, Himmer S (2002) A meta- analysis of grizzly bear DNA mark-recapture projects in British Columbia, Canada. Ursus 13:137–152
Boulanger J, Stenhouse G, Munro R (2004) Sources of heterogeneity bias when DNA mark-recapture sampling methods are applied to grizzly bear (Ursus arctos) populations. J Mammal 85:618–624
Boulanger J, Proctor M, Himmer S, Stenhouse G, Paetkau D (2006) An empirical test of DNA mark-recapture sampling strategies for grizzly bears. Ursus 17:149–158
Castle JH (2010) Mate searching as a cost of reproduction for American black bears (Ursus americanus) in the Great Lakes St. Lawrence forest region of Ontario, Canada. Thesis, Trent University, Peterborough, Canada
Conn PB, Arthur AD, Bailey LL, Singleton GR (2006) Estimating the abundance of mouse populations of known size: promises and pitfalls of new methods. Ecol Appl 16:829–837
Costello CM (2008) The spatial ecology and mating system of black bears (Ursus americanus) in New Mexico. Ph.D. dissertation, Montana State University, Bozeman, USA
Cubaynes S, Pradel R, Choquet R, Duchamp C, Gaillard J-M, Lebreton J-D, Marboutin E, Miquel C, Reboulet A-M, Poillot C, Taberlet P, Giminez O (2010) Importance of accounting for detection heterogeneity when estimating abundance: the case of French wolves. Conserv Biol 24:621–626
Czetwertynski SM, Boyce MS, Schmiegelow FK (2007) Effects of hunting on demographic parameters of American black bears. Ursus 18:1–18
Davis SA, Akison LK, Farroway LN, Singleton GR, Leslie KE (2003) Abundance estimators and truth: accounting for individual heterogeneity in wild house mice. J Wildl Manage 67:634–645
DeBruyn TD (1999) Walking with bears. Lyons Press, Washington DC
Dice LR (1938) Some census methods for mammals. J Wildl Manage 2:119–130
Doherty PF, White GC, Burnham KP (2012) Comparison of model building and selection strategies. J Ornithol 152(Supplement 2):317–323
Dorazio RM, Royle AJ (2003) Mixture models for estimating the size of a closed population when capture rates vary among individuals. Biometrics 59:351–364
Dreher BP, Winterstein SR, Scribner KT, Lukacs PM, Etter DR, Rosa GJM, Lopez VA, Libants S, Filcek KB (2007) Noninvasive estimation of black bear abundance incorporating genotyping errors and harvested bear. J Wildl Manage 71:2684–2693
Ebert C, Knauer F, Storch I, Hohmann U (2010) Individual heterogeneity as a pitfall in population estimates based on non-invasive genetic sampling—review and recommendations. Wildl Biol 16:225–240
Efford MG (2004) Density estimation in live-trapping studies. Oikos 106:598–610
Efford MG (2010) Program DENSITY (version 4.4.5.1): software for spatially explicit capture–recapture. http://www.otago.ac.nz/density/index.html
Efford MG (2011) Estimation of population density by spatially explicit capture–recapture analysis of data from area searches. Ecology 92:2202–2207
Efford MG (2012) secr: Spatially explicit capture–recapture models. R package version 2.3.2. http://CRAN.R-project.org/package=secr
Efford MG, Dawson DK, Robbins CS (2004) DENSITY: software for analyzing capture–recapture data from passive detector arrays. Anim Biodivers Conserv 27:217–228
Efford MG, Borchers DL, Byrom AE (2009) Density estimation by spatially explicit capture–recapture: likelihood-based methods. In: Thompson DL, Cooch EG, Conroy MJ (eds) Modeling demographic processes in marked populations. Springer, New York, pp 255–269
Ennis S, Gallagher TS (1994) A PCR-based sex-determination assay in cattle based on the bovine amelogenin locus. Anim Genet 25:425–427
Foster RJ, Harmsen BJ (2012) A critique of density estimation from camera-trap data. J Wildl Manage 76:224–236
Gardner B, Royle JA, Wegan MT (2009) Hierarchical models for estimating density from DNA mark-recapture studies. Ecology 90:1106–1115
Gardner B, Royle JA, Wegan MT, Rainbolt R, Curtis P (2010) Estimating black bear density using DNA data from hair snares. J Wildl Manage 74:318–325
Garshelis DL (1992) Mark-recapture density estimation for animals with large home ranges. In: McCullough DR, Barret RH (eds) Wildlife 2001: populations. Elsevier Applied Science, London, pp 1098–1109
Garshelis DL (1994) Density-dependent population regulation of black bears. In: Taylor M (ed) Density-dependent population regulation of black, brown, and polar bears. Int Conf Bear Res Manage Monograph 3. Knoxville, pp 3–14
Garshelis DL, Pelton MR (1980) Activity of black bears in the Great Smoky Mountains National Park. J Mammal 61:8–19
Gerber BD, Karpanty SM, Kelly MJ (2012) Evaluating the potential biases in carnivore capture–recapture studies associated with the use of lure and varying density estimation techniques using photographic-sampling data of the Malagasy civet. Popul Ecol 54:43–54
Gervasi V, Ciucci P, Boulanger J, Randi E, Boitani L (2012) A multiple data source approach to improve abundance estimates of small populations: the brown bear in the Apennines, Italy. Biol Conserv 152:10–20
Gray TNE, Prum S (2012) Leopard density in post-conflict landscape, Cambodia: evidence from spatially explicit capture–recapture. J Wildl Manage 76:163–169
Hurvitch CM, Tsai C-L (1989) Regression and time series model selection in small samples. Biometrika 76:297–307
Immell D, Anthony RG (2008) Estimation of black bear abundance using a discrete DNA sampling device. J Wildl Manage 72:324–330
Jones MD, Pelton MR (2003) Female American black bear use of managed forest and agricultural lands in coastal North Carolina. Ursus 14:188–197
Kalle R, Ramesh T, Qureshi Q, Sankar K (2011) Density of tiger and leopard in a tropical deciduous forest of Mudumalai Tiger Reserve, southern India, as estimated using photographic capture–recapture sampling. Acta Theriol 56:335–342
Karanth KU (1995) Estimating tiger (Panthera tigris) populations from camera trap data using capture–recapture models. Biol Conserv 71:333–338
Kéry M, Gardener B, Stoeckle T, Weber D, Royle AJ (2011) Use of spatial capture–recapture modeling and DNA data to estimate densities of elusive animals. Conserv Biol 25:356–364
Kitahara E, Isagi Y, Ishibashi Y, Saitoh T (2000) Polymorphic microsatellite DNA markers in the Asiatic black bear Ursus thibetanus. Mol Ecol 9:1661–1662
Koehler GM, Pierce DJ (2003) Black bear home range sizes in Washington: climatic, vegetative, and social influences. J Mammal 84:81–91
Kolenosky GB (1990) Reproductive biology of black bears in east-central Ontario. Int Conf Bear Res Manage 8:385–392
Lindzey FG, Barber KR, Peters RD, Meslow EC (1986) Responses of a black bear population to a changing environment. Int Conf Bear Res Manage 6:57–63
Link WA (2004) Nonidentifiability of population size from capture–recapture data with heterogeneous detection probabilities. Biometrics 59:1123–1130
Lukacs PM, Burnham KP (2005) Review of capture–recapture methods applicable to noninvasive genetic sampling. Mol Ecol 14:3909–3919
MacKenzie DI, Nichols JD, Sutton N, Kawanishi K (2005) Improving inferences in population studies of rare species that are detected imperfectly. Ecology 86:1101–1113
Marques TA, Thomas L, Royle AJ (2011) A hierarchical model for spatial capture–recapture data: comment. Ecology 92:526–528
Marshall IB, Wiken EB, Hirvonen H (1998) (Compilers) Terrestrial ecoprovinces of Canada. Ecosystem Sciences Directorate, Environment Canada and Research Branch, Agriculture and Agri-Food Canada, Ottawa/Hull
Marucco F, Boitani L, Pletscher DH, Schwartz MK (2011) Bridging the gaps between non-invasive genetic sampling and population parameter estimation. Eur J Wildl Res 57:1–13
McKelvey KS, Pearson DE (2001) Population estimation with sparse data: the role of estimators versus indices revisited. Can J Zool 79:1754–1765
McLaren, MA, Obbard ME, Pond, BA (2005) Black bear population index network: Results from 2000–2005 and efficiency of the technique. Ontario Ministry of Natural Resources, South-central Science and Information Section, Bracebridge, Ontario, Canada
Miller SD, White GC, Sellers RA, Reynolds HV, Schoen JW, Titus K, Barnes VG Jr, Smith RB, Nelson RR, Ballard WB, Schwartz CC (1997) Brown and black bear density estimation in Alaska using radiotelemetry and replicated mark-resight techniques. Wildl Monogr 133
Mowat G, Heard DC, Seip DR, Poole KG, Stenhouse G, Paetkau DW (2005) Grizzly Ursus arctos and black bear U. americanus densities in the interior mountains of North America. Wildl Biol 11:31–48
Norris JL, Pollock KH (1995) A capture–recapture model with heterogeneity and behavioural response. Environ Ecol Stat 2:303–313
Noss AJ, Gardner B, Maffei L, Cuéllar E, Montaño R, Romero-Muñoz A, Sollmann R, O’Connell AF (2012) Comparison of density estimation methods for mammal populations with camera traps in the Kaa-Iya del Gran Chaco landscape. Anim Conserv 15:527–535
Noyce KV, Garshelis DL (2011) Seasonal migrations of black bears: causes and consequences. Behav Ecol Sociobiol 65:823–835
Noyce KV, Garshelis DL, Coy PL (2001) Differential vulnerability of black bears to trap and camera sampling and resulting biases in mark-recapture estimates. Ursus 12:211–225
Obbard ME, Howe EJ (2008) Demography of black bears in hunted and unhunted areas of the boreal forest of Ontario. J Wildl Manage 72:869–880
Obbard ME, Howe EJ, Kyle CJ (2010) Empirical comparison of density estimators for large carnivores. J Appl Ecol 47:76–84
O’Brien TG, Kinnaird MF (2011) Density estimation of sympatric carnivores using spatially explicit capture–recapture methods and standard trapping grid. Ecol Appl 21:2908–2616
Paetkau D, Strobeck C (1994) Microsatellite analysis of genetic variation in black bear populations. Mol Ecol 3:489–495
Paetkau D, Calvert W, Stirling I, Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354
Pelletier A, Obbard ME, Mills K, Howe EJ, Burrows FG, White BN, Kyle CJ (2012) Delineating genetic groupings in continuously distributed species across largely homogeneous landscapes: a study of American black bears (Ursus americanus) in Ontario, Canada. Can J Zool 90:999–1014
Pledger S (2000) Unified maximum likelihood estimates for closed capture–recapture models using mixtures. Biometrics 56:434–442
Pledger S (2005) The performance of mixture models in heterogeneous closed population capture–recapture. Biometrics 61:868–873
Powell RA, Zimmerman JW, Seaman DE (1997) Ecology and behavior of North American black bears: home ranges, habitat and social organization. Chapman and Hall, London
Proctor M, McLellan B, Boulanger J, Apps C, Stenhouse G, Paetkau D, Mowat G (2010) Ecological investigations of grizzly bears in Canada using DNA from hair, 1995–2005: a review of methods and progress. Ursus 21:169–188
R Development Core Team (2012) R: a language and environment for statistical computing, version 2.15. R Foundation for Statistical Computing, Vienna. ISBN 3-900051-07-0, http://www.R-project.org
Rogers LL (1987) Effects of food supply and kinship on social behavior, movements, and population growth of black bears in northeastern Minnesota. Wildl Monogr 97
Rowe JS (1972) Forest regions of Canada. Canadian Forestry Service Publication No. 1300. Environment Canada, Ottawa
Royle AJ, Karanth U, Gopalaswamy AM, Kumar NS (2009) Bayesian inference in camera trapping studies for a class of spatial capture–recapture models. Ecology 90:3233–3244
Royle AJ, Magoun AJ, Gardner B, Valkenburg P, Lowell RE (2011) Density estimation in a wolverine population using spatial capture–recapture models. J Wildl Manage 75:604–611
Schenk A, Obbard ME, Kovacs KM (1998) Genetic relatedness and home-range overlap among female black bears (Ursus americanus) in northern Ontario, Canada. Can J Zool 76:1511–1519
Sollmann R, Furtado MM, Gardner B, Hofer H, Jácomo ATA, Tôrres NM, Silveira L (2011) Improving density estimates for elusive carnivores: accounting for sex-specific detection and movements using spatial capture–recapture models for jaguars in central Brazil. Biol Conserv 144:1017–1024
Sollmann R, Gardner B, Belant JL (2012) How does spatial study design influence density estimates from capture recapture models? PLoS ONE 7:e34575
Taberlet P, Camarra J–J, Griffin S, Hanotte O, Waits LP, Dubois-Paganon C, Burke T, Bouvet J (1997) Noninvasive genetic tracking of the endangered Pyrenean brown bear population. Mol Ecol 6:869–876
Wegan M (2008) Aversive conditioning, population estimation, and habitat preference of black bears (Ursus americanus) on Fort Drum Military Installation in northern New York. MSc Thesis, Cornell University, Ithaca, USA
White GC (2005) Correcting wildlife counts using detection probabilities. Wildl Res 32:211–216
White GC, Anderson DR, Burnham KP, Otis DL (1982) Capture–recapture and removal methods for sampling closed populations. Los Alamos National Laboratory Rep. LA-8787-NERP, Los Alamos, NM, USA
Wooding JB, Hardisky TS (1994) Home range, habitat use, and mortality of black bears in North-Central Florida. Int Conf Bear Res Manage 9:349–356
Woods JG, Paetkau D, Lewis D, McLellan BN, Proctor M, Strobeck C (1999) Genetic tagging of free-ranging black and brown bears. Wildl Soc Bull 27:616–627
Acknowledgments
We thank the many biologists and technicians with the Ontario Ministry of Natural Resources and the Natural Resources DNA Profiling and Forensics Centre who collected hair samples and conducted genetic analyses. The Applied Research and Development Branch, Ontario Ministry of Natural Resources provided funding. Murray Efford provided helpful advice on the theory and implementation of SECR models on the DENSITY | secr online forum (http://www.phidot.org). Comments from the associate editor and two anonymous reviewers helped us improve upon an earlier version of the manuscript. This work was made possible by the facilities of the Shared Hierarchical Academic Research Computing Network (SHARCNET: http://www.sharcnet.ca) and Compute/Calcul Canada.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Howe, E.J., Obbard, M.E. & Kyle, C.J. Combining data from 43 standardized surveys to estimate densities of female American black bears by spatially explicit capture–recapture. Popul Ecol 55, 595–607 (2013). https://doi.org/10.1007/s10144-013-0389-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10144-013-0389-y