Abstract
Effective treatment of infectious diseases depends on the ability to rapidly identify the infecting bacteria and the use of sensitive antibiotics. The currently used identification assays usually take more than 72 h to perform and have a low sensitivity. Herein, we present a microbead-based microfluidic platform that is highly sensitive and rapid for bacterial detection and antibiotic sensitivity testing. The platform includes four units, one of which is used for bacterial identification and the other three are used for susceptibility testing. Our results showed that Escherichia coli O157 at a cell density range of 101–105 CFU/μL could be detected within 30 min. Additionally, the effects of three antibiotics on E. coli O157 were evaluated within 4–8 h. Overall, this integrated microbead-based microdevice provides a sensitive, rapid, reliable, and highly effective platform for the identification of bacteria, as well as antibiotic sensitivity testing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Infectious diseases are a leading cause of death [1, 2]. Over 95 % of these deaths are caused by the lack of proper diagnosis and treatment [3]. Conventional methods of diagnosis are time-consuming and cumbersome. In addition, these methods are not feasible for synchronous or high-throughput screening. Therefore, it is critical to develop a tool for shortening the time necessary to identify infectious pathogens and an effective antibiotic regimen for treatment.
Microfluidics is an attractive platform for rapid and multiple analysis [4–12]. Immunomicrobeads coated with a specific antibody for the target pathogen provide a simple and direct method to capture and enrich the target pathogen. This capture/enrichment method (referred to as immunocapture) increases the effective concentration of the bacterium and allows target bacteria to accumulate on the surface of microbeads. This technique has been commonly used for pathogenic separation and detection in microfluidic devices [6, 7]. Similar techniques, such as magnetic immunomicrobeads, have also been used for rapid functional analysis and susceptibility screening [8, 9]. However, these methods lack the capacity for high-throughput screening and the implementation of multiplexed biological assays. Here, we introduce an immunomicrobead-based microfluidic device that can be used for both rapid bacterial identification and antibiotic sensitivity testing. This technology offers at least two advantages: (1) two assays (bacterial identification and drug sensitivity testing) can be performed with a single microfluidic platform and (2) each assay can be performed using a low-density bacterial sample. This technology will reduce the time needed and increase the efficiency to diagnose bacterial infections, eventually enabling patient-specific antibiotic regimens.
Materials and methods
Microfluidic chip fabrication
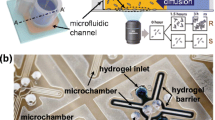
The schematic representation of the microfluidic chip is shown in Fig. 1A. The microfluidic chip consists of four units sharing a common outlet. One unit was utilized for the identification of bacteria in a sample (unit a) and the other three units (units b, c, and d) were used to test the susceptibility of bacteria to three different antibiotics in parallel. Figure 1B shows the details of unit a, which includes a sample inlet (1.5-mm diameter) and a capture chamber. The capture chamber (2 mm × 4 mm × 100 μm) is enclosed by many cuboid micropillars (black dotted line). The distance between neighboring micropillars is 60 μm and it was designed to hold a single layer of glass microbeads (average diameter = 70 μm; Mianyang Guangyao New Material Co., Mianyang, Sichuan, China). Figure 1C shows the details of unit b, which consists of a linear concentration gradient generator (CGG), five downstream parallel mixture chambers, and the corresponding monitoring chamber. The unit has two inlets (1.5-mm diameter), used for medium, drug, or bacterial solution perfusion, and the corresponding cascade microchannels (10 mm × 200 μm × 100 μm). The CGG utilized diffusive mixing to generate a mixture of the two inlets at the cascade microchannels and could generate gradient concentrations of 0:1:2:3:4 theoretically based on the formula in a previous research [11]. Unit b also has two clips (clip 1 and clip 2) that serve as valves to control the liquid stream in the upper and lower microchannels. The workflow of units a and b/c/d is illustrated in Figs. 1D, E. With the entire device, bacteria could be identified, and then the susceptibility of bacteria to three antibiotics could be tested.
Schematic of the integrated microfluidic device for bacterial identification and antibiotic sensitivity testing. A Diagram (left) and a picture (right) of the microfluidic device. B Enlarged diagram of unit a. C Enlarged diagram of unit b. D Workflow of unit a. ① Bacteria were injected into the unit. ② The bacteria mixed with the antibody (AB)-coated microbeads to form microbeads–bacteria complexes. ③ A mixture of monoclonal rabbit anti-Escherichia coli antibody (1:100 dilution, Abcam) and FITC-conjugated goat anti-rabbit polyclonal antibody (1:200 dilution, Santa Cruz Biotechnology) was injected into the unit, and the fluorescence intensity was measured using a fluorescence microscope. E) Workflow of units b, c, or d. ① Bacteria were injected into the mixture chamber. ② Antibiotics [norfloxacin (12.8 μg/mL), ceftriaxone (8 μg/mL), or ampicillin (80 μg/mL)] were then injected into the microchannels to generate a series of gradient concentrations of the antibiotic. ③ SYBR green I, a fluorescence dye that only stains live cells, was added to the mixture chambers. ④ The mixture of the bacteria, antibiotic, and SYBR green dye was allowed to mix with AB-coated microbeads to form microbeads–bacteria complexes. ⑤ The fluorescence of SYBR green was captured by a fluorescence microscope and the fluorescence intensity was measured
The microfluidic device was fabricated with polydimethylsiloxane (PDMS, Sylgard 184, Dow Corning, Midland, MI, USA), and the assembly was placed on a glass slide to create an irreversible seal. The microfluidic device was sterilized overnight with UV light. Nonspecific interactions between the chip surface and cells or antibiotics were minimized by coating the walls of the fluid layer and the glass coverslip with 1 % bovine serum albumin.
Validation of the microfluidic chip
To validate the feasibility of the microfluidic platform for biological studies, Escherichia coli O157 at the same initial concentration was cultured in 96-well plates and in the microfluidic chip. The growth states of the bacterial cells were monitored by recording the optical density at a wavelength of 600 nm every 2 h during a period of 16 h. In addition, rhodamine 123 (Rh-123, Sigma Aldrich, St. Louis, MO, USA) was used to validate the performance of the CGG. Culture medium with or without 50 mM Rh-123 was introduced into a chip unit (units b, c, or d) from the drug and medium inlets; a series of solutions containing different concentrations of Rh-123 were formed in the winding microchannels before entering the corresponding downstream mixture chambers. The junctions between the CGG and the mixture chambers were imaged with an inverted fluorescence microscope (Leica, DMI4000), and the fluorescence intensities of the Rh-123 were quantified and compared with the theoretical data.
Bacterial cell culture
E. coli O157 (ATCC 35150, ATCC, Rockville, MD, USA) was used as the target bacteria. E. coli (ATCC 25922) and Staphylococcus aureus (ATCC6538) were used as control strains to evaluate the specificity for pathogenic identification. Bacterial strains were routinely cultivated in brain heart infusion (BHI) broth (Remel, Lenexa, KS, USA), and bacterial concentrations were determined by the standard plating count method (PCM).
Identification of E. coli O157 on the microfluidic chip
Immobilization of antibodies on microbeads was conducted as described previously [13]. To identify whether the microbeads have the ability to capture E. coli O157, 1 μL (105 CFU) of bacterial solution was incubated with the coated microbeads in the capture chamber for 5 min to form a bacteria–microbead complex through immunoreactions. The microbeads were observed by a scanning electron microscope (SEM) (Quanta™ 650 FEG, USA).
The capture efficiency of chip unit a for different concentrations of E. coli O157 was tested using the PCM. Briefly, the glass microbeads were divided into experimental groups (coated with antibody) and control groups (not coated). First, 5 mg of the glass microbeads suspended in 100 μL of phosphate-buffered saline were injected into the chamber via a syringe pump (Baoding Longer Precision Pump Co., Ltd., China) to form a microbead monolayer in the capture chamber. Next, 1 μL of an E. coli O157 suspension with concentrations ranging from 101 CFU/μL to 106 CFU/μL was injected into the unit slowly. The capture efficiency of the microbeads was calculated according to the following formula:
Image processing and analysis
Images were taken using an inverted fluorescence microscope (Leica, DMI4000) with a 10 × 0.3 NA objective equipped with a CCD camera (Retiga 2000R, QImaging). FITC or SYBR green was imaged at an excitation wavelength of 490–495 nm and an emission wavelength of 525–530 nm. Fluorescence intensity was measured using Image-Pro® Plus 6.0 software (Media Cybernetics, Silver Spring, MD, USA). The fluorescence signal from the unoccupied immunomicrobeads was considered as background and subtracted from the signal.
Statistical analyses
To evaluate the reliability of the experimental results, each experiment was repeated at least three times. Data were presented as means ± standard deviation. A paired Student’s t-test was used to compare the study groups. Significance was defined at p ≤ 0.05. All statistical analyses were performed using SPSS 12.0 software (SPSS Inc.).
Results
Validation of the microfluidic chip
To validate the feasibility of the microfluidic platform for biological studies, we monitored the growth of E. coli O157 cultured in the microfluidic chip and compared it with that grown in 96-well plates. As shown in Fig. 2A, the growth curve for E. coli O157 cells in the chip was in close agreement with that in the 96-well plate for a period of 16 h. The statistical analysis showed that there were no significant differences between the two curves (p > 0.05).
Validation of the microfluidic chip. A The growth curves of Escherichia coli O157 cultured in the microfluidic chip and a 96-well plate over a period of 16 h. B Performance validation of the concentration gradient generator (CGG) in the microfluidic device. a Fluorescence intensities of rhodamine 123 (Rh-123) in the mixing channels connecting the CGG and the mixture chambers [1–5 represent mixing channel 1 (close to the medium inlet) to channel 5 (close to the drug inlet), respectively]. b Quantified comparison of the actual and theoretical data for the fluorescence intensity of different concentrations of Rh-123 solution generated by the CGG
To validate the CGG in the microfluidic device, the experimental gradient values were compared with the theoretical values, using Rh-123 as an indicator. There was good agreement between the experimental and theoretical data (correlation coefficient = 0.9953) (Fig. 2B), indicating the feasibility of the CGG.
Identification of E. coli O157 on the microfluidic device
An SEM was used to examine whether the antibody-coated microbeads have the capability of capturing bacteria. As shown in Fig. 3, a large amount of E. coli O157 was present on the surface of a microbead selected randomly, indicating that the microbeads had a high capacity for specific capture.
The capture efficiencies of E. coli O157 cells in the experimental groups and the control groups were examined. As shown in Fig. 4, for the experimental groups, the capture rate was 85–92 %, with the bacterial concentrations varying from 4 × 101 CFU/μL to 4 × 105 CFU/μL. The capture rate was only 61.41 % when the bacterial concentration reached 4 × 106 CFU/μL. This finding was probably because the amount of bacteria in the sample was much larger than the number of antibody sites on the glass microbeads. For the control groups, the capture rates were all below 20 %, which was ascribed to nonspecific adsorptions. These results suggest that the antibody-coated microbeads provide ideal capture efficiency when the bacterial sample concentrations range from 4 × 101 CFU/μL to 4 × 105 CFU/μL.
We further validated the capture efficiency and specificity by fluorescence imaging of the FITC-labeled secondary antibody. Images were taken, and the fluorescence intensity of the bacteria at various concentrations was measured. As shown in Fig. 5A, the bacterial concentration was directly proportional to the fluorescence intensity. Furthermore, as shown in Fig. 5B, there was a correlation between the bacterial concentration and the immunofluorescence measurement. These data demonstrate that our platform can be used for the quantitative determination of bacterial concentrations ranging from 101 to 105 CFU/μL, with high sensitivity.
Capture efficiency and specificity of the microfluidic device. A Fluorescence images of the capture chamber with different concentrations of E. coli O157 suspensions. a–e The representative concentrations of E. coli O157 samples were 4 × 105 CFU/μL, 4 × 104 CFU/μL, 4 × 103 CFU/μL, 4 × 102 CFU/μL, and 4 × 101 CFU/μL, respectively. f Control group: sample with no bacteria. B Correlation between Ig (CFU/μL) and fluorescence mean values obtained from antibody-modified microbeads in the capture chamber (with respect to the control group). C Fluorescence images of three strains. a E. coli O157, b E. coli (ATCC 25922), and c S. aureus. D Quantitative measurement of C
The bacterial specificity was also assayed. As controls, E. coli (ATCC 25922) and S. aureus (ATCC 6538) diluted to the same concentration as E. coli O157 (105 CFU/μL) were tested. As shown in Fig. 5C, bright fluorescence was observed with E. coli O157, but almost no fluorescence was seen from the two control strains, indicating that our platform could specifically recognize E. coli O157.
Antibiotic susceptibility of E. coli O157
We employed chip units b, c, and d to test the susceptibility of E. coli O157 to norfloxacin, ceftriaxone, and ampicillin. For norfloxacin, the gradient concentrations were 0 (control group), 1.6, 3.2, 4.8, and 6.4 μg/mL; for ceftriaxone, they were 0 (control group), 1, 2, 3, and 4 μg/mL; and for ampicillin, they were 0 (control group), 10, 20, 30, and 40 μg/mL. With the increasing concentration of each antibiotic, an apparent change in the fluorescence signal was observed within 4–8 h (Fig. 6). These results indicated that both norfloxacin and ceftriaxone inhibited bacterial growth effectively, in a statistically significant dose- and time-dependent manner, whereas E. coli O157 was resistant to amipicillin when the concentration was less than 30 μg/mL (Fig. 6).
Antibiotic sensitivity testing. A The fluorescence images of the corresponding monitoring chambers observed at 4 and 8 h after: a norfloxacin, b ceftriaxone, and c ampicillin treatment. B Temporal changes to the fluorescence intensity of the corresponding monitoring chambers with gradient concentrations of norfloxacin (a), ceftriaxone (b), and ampicillin (c) at 4 and 8 h. The error bars represent standard deviations (n = 3)
The three antibiotic sensitivity tests were also determined by an absorbance-based assay performed in 96-well plates. The results were consistent with those obtained on the microfluidic platform. However, the 96-well plate assay takes a much longer time and consumes larger amounts of reagents (data not shown).
Discussion
In this study, we present a rapid and effective microbead-based microfluidic platform that could be used for both bacterial identification and antibiotic sensitivity testing. Using this system, E. coli O157 could be distinguished from 1 μl of sample with a concentration of 101–105 CFU/μL within 30 min. The antibiotic susceptibility to three antibiotics could be tested within 4–8 h. Unlike many microfluidic devices used currently [7, 14–16], another advantage of our system is the integration of four chip units into a multifunctional platform to enable multiple assays. As a result, the whole process from bacterial identification to relational antibiotic susceptibility testing could be carried out on a single platform, which greatly improves the application efficiency.
Sensitivity is a major concern in evaluating a microfluidic platform. Compared with previous detection platforms, we made some improvements to increase the sensitivity. First, the use of microbeads increases the surface area for antibody immobilization. Thus, the immunocapture efficiency was higher (85–92 %) than offline immunomagnetic separation, whose capture rates are 44–86 %, or the planar methods, whose capture rates are 21 % [17, 18]. Second, we modified the depth of the capture chamber, making it larger (100 μm) than the diameter of the beads (70 μm), so the beads could be aligned as a monolayer in the capture chamber. This design maximized the optical accessibility of bacteria to the detection light, and high fluorescence signal strength was achieved. Third, the use of CGG provided the advantage of automatically generating gradient concentrations of antibiotics. It greatly enhanced the efficiency of the antibiotic susceptibility tests and, therefore, enhanced the clinical applicability of this platform.
In summary, we demonstrated a microbead-based platform for rapid bacterial identification and antibiotic susceptibility testing. The presented immunocapture technology marks an essential step towards this versatile platform for the integration of microbead- and cell-based assays. With this platform, the assay time was shortened from a typical 72 h to 8.5 h for the whole process. This integrated and flexible microfluidic platform has the potential to be incorporated as a routine clinical test device used for the early diagnosis and treatment of infectious diseases.
References
Boedicker JQ, Li L, Kline TR, Ismagilov RF (2008) Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 8:1265–1272. doi:10.1039/b804911d
Carrigan SD, Scott G, Tabrizian M (2004) Toward resolving the challenges of sepsis diagnosis. Clin Chem 50:1301–1314. doi:10.1373/clinchem.2004.032144
Nguyen HB, Rivers EP, Abrahamian FM, Moran GJ, Abraham E, Trzeciak S, Huang DT, Osborn T, Stevens D, Talan DA (2006) Severe sepsis and septic shock: review of the literature and emergency department management guidelines. Ann Emerg Med 48:28–54. doi:10.1016/j.annemergmed.2006.02.015
Sauer-Budge AF, Mirer P, Chatterjee A, Klapperich CM, Chargin D, Sharon A (2009) Low cost and manufacturable complete microTAS for detecting bacteria. Lab Chip 9:2803–2810. doi:10.1039/B904854E
Yager P, Edwards T, Fu E, Helton K, Nelson K, Tam MR, Weigl BH (2006) Microfluidic diagnostic technologies for global public health. Nature 442:412–418. doi:10.1038/nature05064
Mujika M, Arana S, Castaño E, Tijero M, Vilares R, Ruano-López JM, Cruz A, Sainz L, Berganza J (2009) Magnetoresistive immunosensor for the detection of Escherichia coli O157:H7 including a microfluidic network. Biosens Bioelectron 24:1253–1258. doi:10.1016/j.bios.2008.07.024
Srinivasan B, Varshney M, Tung S, Li Y (2005) A microfluidic filter chip for highly sensitive chemiluminescence detection of E. coli O157:H7. In: Proceedings of the ASME 2005 International Mechanical Engineering Congress and Exposition, Orlando, Florida, USA, November 2005. doi:10.1115/IMECE2005-81881
Kinnunen P, Sinn I, McNaughton BH, Newton DW, Burns MA, Kopelman R (2011) Monitoring the growth and drug susceptibility of individual bacteria using asynchronous magnetic bead rotation sensors. Biosens Bioelectron 26:2751–2755. doi:10.1016/j.bios.2010.10.010
Sinn I, Kinnunen P, Albertson T, McNaughton BH, Newton DW, Burns MA, Kopelman R (2011) Asynchronous magnetic bead rotation (AMBR) biosensor in microfluidic droplets for rapid bacterial growth and susceptibility measurements. Lab Chip 11:2604–2611. doi:10.1039/C0LC00734J
Sia SK, Whitesides GM (2003) Microfluidic devices fabricated in poly(dimethylsiloxane) for biological studies. Electrophoresis 24:3563–3576. doi:10.1002/elps.200305584
Jeon NL, Dertinger SK, Chiu DT, Choi IS, Stroock AD, Whitesides GM (2000) Generation of solution and surface gradients using microfluidic systems. Langmuir 16:8311–8316. doi:10.1021/la000600b
Duffy DC, McDonald JC, Schueller OJ, Whitesides GM (1998) Rapid prototyping of microfluidic systems in poly(dimethylsiloxane). Anal Chem 70:4974–4984. doi:10.1021/ac980656z
Guan X, Zhang HJ, Bi YN, Zhang L, Hao DL (2010) Rapid detection of pathogens using antibody-coated microbeads with bioluminescence in microfluidic chips. Biomed Microdevices 12:683–691. doi:10.1007/s10544-010-9421-6
Jing W, Zhao W, Liu S, Li L, Tsai C-T, Fan X, Wu W, Li J, Yang X, Sui G (2013) Microfluidic device for efficient airborne bacteria capture and enrichment. Anal Chem 85:5255–5262. doi:10.1021/ac400590c
Mohan R, Mukherjee A, Sevgen SE, Sanpitakseree C, Lee J, Schroeder CM, Kenis PJ (2013) A multiplexed microfluidic platform for rapid antibiotic susceptibility testing. Biosens Bioelectron 49:118–125. doi:10.1016/j.bios.2013.04.046
Sun P, Liu Y, Sha J, Zhang Z, Tu Q, Chen P, Wang J (2011) High-throughput microfluidic system for long-term bacterial colony monitoring and antibiotic testing in zero-flow environments. Biosens Bioelectron 26:1993–1999. doi:10.1016/j.bios.2010.08.062
Xia N, Hunt TP, Mayers BT, Alsberg E, Whitesides GM, Westervelt RM, Ingber DE (2006) Combined microfluidic–micromagnetic separation of living cells in continuous flow. Biomed Microdevices 8:299–308. doi:10.1007/s10544-006-0033-0
Yang L, Banada PP, Chatni MR, Lim KS, Bhunia AK, Ladisch M, Bashir R (2006) A multifunctional micro-fluidic system for dielectrophoretic concentration coupled with immuno-capture of low numbers of Listeria monocytogenes. Lab Chip 6:896–905. doi:10.1039/B607061M
Acknowledgments
This study was supported by the Special Fund for Health Scientific Research in the Public Interest from the National Health and Family Planning Commission of China (no. 201202011).
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
J. He and X. Mu contributed equally to the article.
Rights and permissions
About this article
Cite this article
He, J., Mu, X., Guo, Z. et al. A novel microbead-based microfluidic device for rapid bacterial identification and antibiotic susceptibility testing. Eur J Clin Microbiol Infect Dis 33, 2223–2230 (2014). https://doi.org/10.1007/s10096-014-2182-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-014-2182-z