Abstract
In view of the need for easily accessible biomarkers, we evaluated in ADHD children the epigenetic status of the 5′-untranslated region (UTR) in the SLC6A3 gene, coding for human dopamine transporter (DAT). We analysed buccal swabs and sera from 30 children who met DSM-IV-TR criteria for ADHD, assigned to treatment according to severity. Methylation levels at six-selected CpG sites (among which, a CGGCGGCGG and a CGCG motif), alone or in combination with serum titers in auto-antibodies against dopamine transporter (DAT aAbs), were analysed for correlation with CGAS scores (by clinicians) and Conners’ scales (by parents), collected at recruitment and after 6 weeks. In addition, we characterized the DAT genotype, i.e., the variable number tandem repeat (VNTR) polymorphisms at the 3′-UTR of the gene. DAT methylation levels were greatly reduced in ADHD patients compared to control, healthy children. Within patients carrying at least one DAT 9 allele (DAT 9/x), methylation at positions CpG2 and/or CpG6 correlated with recovery, as evident from delta-CGAS scores as well as delta Conners’ scales (‘inattentive’ and ‘hyperactive’ subscales). Moreover, hypermethylation at CpG1 position denoted severity, specifically for those patients carrying a DAT 10/10 genotype. Intriguingly, high serum DAT-aAbs titers appeared to corroborate indications from high CpG1 versus high CpG2/CpG6 levels, likewise denoting severity versus recovery in DAT 10/10 versus 9/x patients, respectively. These profiles suggest that DAT 5′UTR epigenetics plus serum aAbs can serve as suitable biomarkers, to confirm ADHD diagnosis and/or to predict the efficacy of treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Attention deficit/hyperactivity disorder (ADHD) has been internationally recognized as a serious neuro-developmental alteration [18, 20, 68]. The prevalent ADHD symptoms include problems in maintaining attention, excessive motor activity, and impulsivity, which often lead to poor academic performance and impaired social interactions. These symptoms develop quite early in up to 5% of children [67], and often persist into adolescence and adulthood [8]. Other conditions, frequently comorbid with ADHD, include: externalizing disorders, such as oppositional defiant disorder and conduct disorder; compulsive conditions, such as obsessive–compulsive disorder and Tourette’s syndrome [69, 70]; and addictive disorders, such as pathological gambling as well as substance abuse & drug dependence problems [43, 44].
The developmental psychopathology theoretical framework [14, 21] underlined the role of family characteristics in buffering, or even aggravating, ADHD symptomatology. Moreover, among the etiopathogenetic factors for the onset of ADHD, the quality of parent–infant interactions has been proposed to affect the actual phenomenological expression, resulting in a more or less severe constellation of symptoms. As such, in the present work, we took in due account the maternal judgement about their ADHD children, both at recruitment and after 6 weeks of therapy. We are explicitly assuming that Conners’ scales compiled by mothers could reflect a combined index, reflecting the quality of interaction between maternal skills and individual ADHD temperament. As such, severe ADHD at recruitment may well reflect a contribution by poor coping ability of the mother; conversely, recovery after 6 weeks may incorporate a relief perceived by the mother, in turn facilitating the recovery itself. However, this notion shall not be seen as a caveat but rather as a more complete account of the dynamic expression of ADHD in real-life situations.
Although the aetiology of ADHD is multi-factorial and still unclear, this syndrome is viewed by some as a motivational dysfunction, due to an altered cross-talk between fronto-striatal circuits [12, 3, 75]. Some evidence exists of imbalanced prefrontal and/or striatal levels of neurotransmitters, especially dopamine [61, 71]. Part of preclinical ADHD research has hence focused on the dopamine transporter (DAT), because modification of expression and/or function of its gene may well lead to specific ADHD symptoms [6, 7, 48]. Unsurprisingly, knockout DAT mice have been extensively used to reproduce behavioral symptoms of ADHD (see [33]). In the same line, knockout DAT rats have been recently generated (Leo and colleagues, manuscript submitted; Cinque and colleagues, manuscript in preparation), although their behavior has still to be fully phenotyped.
Expression of DAT may be determined by the genetic VNTR polymorphism of the 3′ untranslated region (UTR) of its gene SLC6A3, and it can be finely tuned by epigenetic mechanisms. The latter are mitotically heritable, but reversible, changes in transcription and/or translation of a gene without modification of genomic DNA sequence [40, 65, 66]. Accumulated evidence supports the pivotal role of epigenetics in neuronal development, differentiation and communication, as well as in synaptic plasticity [51]. In the last decade, a role for epigenetics in psychiatric diseases has been recognized [77]. Among epigenetic mechanisms, DNA methylation is the best characterized, and has been consistently implicated in the development of mental disorders [1, 26, 35]. DNA methylation has been the focus of most recent studies concerning addictive psychiatric disorders (see [41, 46]). For instance, selective changes in DNA methylation of BDNF promoter have been observed in peripheral blood mononuclear cells (PBMCs) of subjects with bipolar disorder type II [23] and major depression [22, 27] as well as in schizophrenic patients [24].
Recently, we proposed that an excessive production of DAT protein, possibly accompanied by altered turnover/degradation, could make DAT or its fragments spill into the blood and generate an auto-immune reaction [34]. Indeed, the presence of detectable auto-antibodies against DAT (DAT aAbs) was confirmed in serum samples from ADHD patients [34]. Notably, observed behavioral changes were nicely segregated between the genotypes. On the one hand, for carriers of two 10-repeat (termed 10/10) VNTR alleles, elevated DAT-aAbs titers were likely associated with most severe ADHD symptoms. On the other hand, for patients carrying at least one 9-repeat allele (termed DAT 9/x), the DAT aAbs were rather predicting efficacy of therapy: elevated titers at recruitment were found in subjects showing a considerable behavioral amelioration after 6 weeks of methylphenidate [34]. These data left unanswered the question on how could DAT aAbs serve two apparently opposite functions in either genotype.
Today, diagnoses of ADHD still rely on structured interviews and/or psychometric scales, and there is the need of more objective, possibly biological markers for this condition (see [32, 78]). Purpose of the present study was (1) to ascertain whether DNA methylation at DAT gene 5′-UTR could serve as a biomarker for ADHD and (2) to correlate possible changes in methylation at specific CpG sites with previous data on DAT-aAbs titers [34], as well as with clinical scores of severity (i.e., ADHD symptoms observed at recruitment) and of recovery (i.e., delta-score changes, as observed in ADHD symptoms after 6 weeks of therapy). Our aim was to provide new possible directions to the search of biomarkers, helping clinicians with ADHD diagnosis.
Materials and methods
Recruitment of patients
Participants included 30 children (5 females aged 6–12 years and 25 males aged 6–14 years), with a formal diagnosis of ADHD formulated by the Child Psychiatry Unit of Tor Vergata University (from April 2010 to March 2012). Fifteen children with typical development were used as healthy controls. All subjects had a full Scale IQ over 84, as assessed by the Wechsler Intelligence Scale—III edition. They were evaluated by child neuro-psychiatrists (G. Giana; M. Troianiello; M.C. Porfirio) who determined the diagnosis of ADHD, according to DSM-IV-TR criteria; a medical work-up excluded any auto-immune disorder. Exclusions were made in case of psychiatric comorbidity (conduct disorder, obsessive–compulsive disorder, bipolar disorder, depression, and psychosis), as assessed by a specific Schedule (K-SADS/PL).
Around two-thirds of children with milder symptoms, judged as not in need of pharmacological treatment, underwent cognitive-behavioral therapy and/or periodic follow-up; one-third of children, with a significant impairment of their adaptive functioning in different areas of life, were assigned to pharmacological treatment with methylphenidate (MPH). In the present paper, as we deal with samples collected at recruitment, these two categories based on the therapeutic intervention decided after enrollment were not taken into consideration.
This study was approved by the Ethical Committee of ISS (Prot. CE-ISS 09/270 of 15 July 2009). Informed consent procedures included searching for consent from the child (using age-adequate approaches) and illustrating to parents the standard consent form; the parents gave their signature (i.e., written informed consent) for the child to participate in this study. We confirm that all potential participants who decided not to participate in the study were not disadvantaged in any way by not participating. In addition, we declare that collected biological materials were used solely to the purpose of previous [34] and of this study. The rules set by the Code of Ethics of the World Medical Association (Declaration of Helsinki), which has been printed in the British Medical Journal (18 July 1964), were fully respected.
Clinical assessment
Each patient was evaluated by trained child neuro-psychiatrists at our unit, according to the DSM-IV and ICD-10 criteria for ADHD. Information was gathered from the clinical interviews and questionnaires with the parents, and from direct observations of the patients.
The Children’s Global Assessment Scale (CGAS) was used by clinicians to measure the overall severity of social and psychiatric functioning: for children aged 4–16 years, CGAS scores range between 1 and 100, with higher scores indicating better functioning. ADHD symptoms were also determined using Conners’ Parent Rating Scale; each item was scored from 0 (not true at all) to 3 (Very much true), giving information about ADHD subtypes (inattentive, hyperactive–impulsive, and combined type).
Parents completed SNAP-IV that elicits DSM-IV-TR criteria for ADHD on a four–point scale of frequency. The semi-structured Schedule for Affective Disorders and Schizophrenia—Present and Lifetime version (K-SADS/PL) and the Child Behavior Checklist/4–18 (CBCL) were used separately, to elicit parents’ and patients’ reports of signs and symptoms that might indicate possible co-morbidities.
Biochemical and genetic assessment
In these children, we were able to collect blood samplings (time T0: basal withdrawal, at recruitment); in addition, we were able to collect buccal swabs using a Catch-all sample collection Swab (Epicentre). As already reported [34], DNA isolated from buccal swabs allowed to determine the DAT genotype, specifically the DAT VNTR polymorphism, well known to be present in the 3′ untranslated region (3′-UTR), rs28363170. Notably, 15 out of 30 patients were homozygous for the 10-repeat allele (DAT 10/10), while 15 of them were carrying at least one 9-repeat allele (DAT 9/x, i.e., 9/9 or 9/10). In addition, the same DNAs were used to measure the methylation status of six CpG sites, selected among those present in the 5′ untranslated region (5′-UTR) of the same gene.
Analysis of DNA methylation
Genomic DNA was prepared from buccal swab samples using the BuccalAmp™ DNA Extraction Kit, following the manufacturer instructions (Epicentre, USA). Briefly, after collecting buccal cells, the swab end was placed into a tube containing QuickExtract DNA extraction solution and rotated a minimum of five times. The tube was vortex mixed for 10 s and incubated at 65 °C for 1 min. After vortex mix for 15 s, the tube was transferred to 98 °C and incubated for 2 min. After vortex mix for 15 s, the DNA was stored until further processing at –20 °C. The yield of DNA is usually between 2 and 14 ng/μl.
The 3′-UTR repeated sequence of the DAT gene was amplified by the polymerase chain reaction (PCR) as it has been described previously [34]. DNAs from the buccal swabs were further processed for assessing amount of methylation in the DAT 5′-UTR sequence (notably, not the transcription promoter region). Amount of methylation was determined in six specific CpG residues (termed M1, M2, M3, M5, M6, and M7; see Fig. 1). Notably, M1–M3 represent a CGGCGGCGG motif, while M5/M6 represents a CGCG motif. The following primers (5′–3′) were used to amplify the gene for DAT: Fwd = AGCTACCATGCCCTATGTGG; Rev = ATCAGCACTCCAAACCCAAC.
Sequence of the 5′-UTR in the DAT gene, with localization of six-chosen CpG residues. Our experimental work discovered three out of six residues which are relevant and useful, for the purpose of ADHD severity (diagnosis) and for prediction or verification of recovery after response to therapy (prognosis)
Bisulfite-treated DNA was amplified by PyroMark PCR Kit (Qiagen, Hilden, Germany) in accordance with the manufacturer’s protocol. PCR conditions were as follows: 95 °C for 15 min, followed by 45 cycles of 94 °C for 30 s, 56 °C for 30 s, 72 °C for 30 s, and, finally, 72 °C for 10 min. PCR products were verified by agarose electrophoresis. Pyrosequencing methylation analysis was conducted using the PyroMark Q24 (Qiagen, Hilden, Germany). The level of methylation was analysed using the PyroMark Q24 Software (Qiagen, Hilden, Germany), which calculates the methylation percentage [mC/(mC + C)] for each CpG site, allowing quantitative comparisons (mC is methylated cytosine and C is unmethylated cytosine).
ELISA methods
As described previously [34], the DAT-EIA-kit (patent details, see below; holder: ISS 100%) was used for the DAT-aAbs detection. Briefly, it is a microtiter immuno-plate for ELISA-based determination of natural antibodies (nAbs) to peptide fragments of human dopamine transporter (hDAT) in the serum. Synthetic peptides corresponding to a fragment of DAT serve as the antigen, and have been proven to detect DAT aAbs in the serum of mice as well [2]. This was a kit designed and custom synthesized by one of authors (O.G.), by selecting on the DAT protein an antigenic sequence (namely, a 19-aminoacid most immuno-reactive portion, residing on the best exposed-portion of the wider extra-cellular loop; U.S. Patent and Trademark Office: EFS ID 13464574; Application No: 61681638; Granstrem et al. 10-AUG-2012 provisional turned into Full Patent No. PN810701WO; Int.l Application No. PCT/EP2013/066845; Publication number WO/2014/023852, 10-AUGUST-2013).
Statistical analysis
The following variables, already used for publication [34], were presently re-evaluated:
-
1.
scores obtained in CGAS and in subscales of Conners’ scales, both at recruitment and after 6 weeks of therapy; delta scores (value after 6 weeks of therapy minus value at recruitment); and their possible correlation with:
-
2.
semi-quantitative DAT-aAbs titers, obtained through the DAT-EIA-kit assay, in recruited ADHD children.
In detail, we fully investigated the potential correlation between amount of methylation in the six specific CpG residues (see Fig. 1) and the scores obtained in the clinical questionnaires. We also verified the potential correlation between said amounts of methylation and the DAT-aAbs titers found in the serum. These correlations were run for each genotype separately (i.e., DAT 10/10 on one side and DAT 9/x on the other hand; n = 15 each) by the Pearson’s R value (threshold for significance with n = 15 is 0.4973 at p < 0.05 and 0.4259 at 0.10 < p < 0.05).
Results
Gross methylation decrease in ADHD
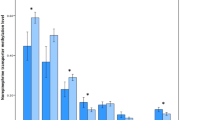
A very first and impressive result (see Fig. 2) shows reduced (nearly halved) levels of methylation, in all six-selected positions, for all of the ADHD patients (see red line, ADHD) compared to healthy controls (see black line, CT). This is a new and unexpected piece of data, since no obvious link is known in the literature, for DAT gene, between a biomarker of epigenetic control (namely, amount of methylation in general) and ADHD.
Statistical correlation between DAT aAbs and clinical scores
We found a correlation between DAT-aAbs titer and CGAS score of −0.186 for 9/x patients and −0.395 for 10/10 patients, confirming a weak tendency towards a link between DAT-aAbs titer and ADHD severity, but only within 10/10 patients. These data extend previous findings [34] that, in patients showing the worst severity of ADHD profile, a very high titer could be found in 10/10 carriers only.
We also found a correlation between DAT-aAbs titer and delta-CGAS (i.e., score after 6 weeks minus score at enrolment) of +0.5216 for 9/x patients and of +0.2689 for 10/10 patients. These data confirm a strong and significant link between DAT-aAbs titer and likelihood of recovery, but only for 9/x patients. Consistently, previous findings [34] showed that, in 9/x patients only, a high titer at enrolment was correlated with likelihood of recovery after 6 weeks of therapy.
Therefore, two apparently contradictory pictures segregated along genotypes. It could be asked how can it be that the elevated DAT-aAbs titers are possibly serving as an index of severity (in 10/10 patients), while seemingly are an index of recovery in 9/x patients. At this point, we can only speculate that these titers might mirror specific (but yet unknown) molecular features of DAT that in turn depend on the presence or not of the 9-repeat allele.
Statistical correlation between DAT aAbs and methylation
When DAT-aAbs titers were subjected to formal correlations with methylation data, it was clearly evident that the titer correlated with different CpG sites depending on genotype. Specifically, DAT-aAbs titers correlated with methylation of CpG site at position M1 for patients with DAT 10/10 genotype, and conversely with methylation of CpG sites at positions M2 and M6 for patients with a 9/x genotype.
Therefore, we could hypothesize that methylation at different sites could well serve a biomarker role similar to DAT aAbs. Specifically, the elevated methylation at CpG M1, alone or jointly with DAT-aAbs titers, may possibly serve as an index of severity (in 10/10 patients), while it serves as an index of recovery in 9/x patients when it occurs at CpG M2 or M6, alone or together with titers (see Table 1).
Statistical correlation between clinical scores and methylation
For DAT 10/10 patients, we confirmed indeed that hypermethylation at CpG M1, as linked to the titer, would have a role in predicting severity. We found indeed that only CpG at position M1 correlated negatively (Pearson’s R = −0.480) with CGAS value, and slightly with some of the Conners’ subscales (see Table 2). Therefore, hypermethylation at CpG M1—much alike high DAT-aAbs titers—denotes a low CGAS score, indexing severity of ADHD symptoms.
For DAT 9/x patients, correlations between the six-selected CpG methylation sites and scores from clinical scales (at enrolment) were never significant (data not shown). Therefore, methylation does not appear to serve as a diagnostic index for this genotype.
For DAT 9/x patients, we confirmed indeed that hypermethylation at CpG M2 and/or M6, as linked to the titer, would have a role in predicting recovery. We found indeed that the CpG at positions M2 and M6 correlated positively with delta-CGAS value (Pearson’s R = +0.537 and +0.648, respectively), as well as slightly negatively with some of the Conners’ subscales (see Table 3). In particular, elevated M2 and/or M6 methylation is associated with positive and larger delta values, suggestive of the amelioration of CGAS scores after a 6-week therapy. In addition, both CpG M2 and (to a lesser extent) M6 were linked to greatly negative delta values, suggestive of amelioration in the scores for the ‘inattentive’ subscale as well as to less ‘hyperactivity’ after a 6-week therapy. Overall, hypermethylation at CpG M2 and/or M6, much alike high DAT-aAbs titers, denotes (already at enrolment) the likelihood of a quite quick recovery.
For DAT 10/10 patients, correlations between the six-selected CpG methylation sites and delta scores from clinical scales were never significant (data not shown). Therefore, methylation does not appear to serve a prognostic index for this genotype.
Discussion
The present study provides new insights into possible strategies for diagnosing ADHD and/or for predicting (and/or monitoring) treatment efficacy. In particular, we provide preliminary evidence that particular CpG sites are hypomethylated, in the 5′-UTR (transcribed but untranslated region) of the SLC6A3 gene for human DAT. Our data report for the first time an unexpected link between methylation of specific CpG sites and titers of circulating DAT aAbs, which were previously shown to be detectable and linked to ADHD [34]. However, this was not true for all patients but occurred only in association with their genotype, namely, the VNTR polymorphism at 3′-UTR. Specifically, we already demonstrated that the amounts of DAT aAbs, which are detectable in the bloodstream, do correlate with ADHD symptoms’ severity in the case of a 10/10 genotype and/or with likelihood to respond positively after treatment in the case of patients carrying at least one 9-repeat allele [34]. In the present study, we were able to extend this observation, further discovering that the amounts of methylation are also correlated with DAT-aAbs titers, depending on genotype, and hence with ADHD symptoms’ severity or with likelihood to respond positively to treatment.
There is extensive literature with discordant results on the involvement of 3′-UTR VNTR polymorphism of DAT in ADHD [17, 31, 79] as far as severity is concerned but also for likelihood to respond after treatment [47, 64]. These studies indicate that the 3′ VNTR might not be the functional site itself, but it could act as a tagging marker for an alternative functional site that contributes to the ADHD phenotype [79]. It is, however, quite new to find a role for the 5′-UTR in general and for specific CpG methylation sites in particular, especially as it is clearly a function of VNTR at 3′-UTR; therefore, we deal with factors residing thousands of base pairs apart. It is also quite new to find a link between 5′-UTR methylation, a factor residing on DNA well before it is transcribed, and DAT aAbs, which may well be due to a sort of auto-immune response against a self protein, and anyway implies the translation and cell-surface expression of DAT protein to act as an (auto)-antigen. Such a link is not obvious, since it is not easy to figure out what happened to transcribed mRNA and then to translation into DAT protein (where these steps enhanced or lowered?).
Our data leave also unanswered the question about what levels of promoter methylation as well as DAT protein are expressed in different brain regions of ADHD patients as well as in lymphocytes. With regard to this aspect, there are no consistent data in the literature about VNTR polymorphism and density of DAT protein in the brain and in lymphocytes of ADHD subjects [13, 39, 55, 58]. It could be proposed that DAT aAbs in the bloodstream may somehow mirror the quantity of DAT present on lymphocytes’ surface, though this link should be demonstrated. Even more difficult issue is to demonstrate, in humans, that peripheral markers may somewhat mirror central levels of the same markers. Use of animal models is therefore warranted as they permit to compare central and peripheral parameters related to a given behavioral phenotype (see [82]). Overall, our data obtained on epithelial cells support a correlation between DAT-aAbs levels and amount of methylation, in few particular 5′-UTR CpG sites.
A working hypothesis
Consistently with our data, it has been already observed that DNA methylation downstream of the transcription starting site (TSS) more tightly correlates with repression of gene transcription than methylation upstream of the TSS, i.e., in the promoter region [9]. DNA methylation at 5′-UTR should imply less transcription of DNA into mRNA, yet the various CpG sites are related to high DAT-aAbs titers. If indeed titers mirror DAT protein levels in the bloodstream, these may indicate a large extent of translation, at least in the periphery. It is, therefore, unclear how less transcription may result in more translation! It is tempting to speculate that either the 9- or the 10-repeat VNTR (alleles of polymorphism at 3′-UTR) makes it more likely that a few mRNAs generate a lot of DAT protein, for instance, by enhancing mRNA stability. As an alternative explanation, we shall posit that one (or more) of the CpG (when methylated) may act as a binding site for a putative molecule that stimulates transcription (instead of repressing it). Normally, in such a case, we would have more mRNAs (though with a normal stability), thus justifying the 5′-UTR (rather than the 3′-UTR) as promoting expression of DAT protein and hence of DAT aAbs. In this context, it has been shown that methylation at specific sites of a genomic sequence can either reduce [29, 30, 45] or enhance [52] transcription factors attachment to DNA regulatory regions.
One allele only was associated with likelihood of recovery, and it appeared to be the 9-repeat one [34]; in our hands, the 10-repeat allele confers resistance to treatment instead. It is know from our previous paper [34] that 9/x patients are showing decreased DAT-aAbs titers combined with a positive outcome of treatment. This notion implies that either the transcription or the translation should perhaps be modulated; namely, we assume that either transcription of mRNAs or their translation may be somewhat reduced or blocked after few weeks of therapeutic, environmental stimulation. We speculate, therefore, of any between either possibility: first, a CpG-favoured (5′-UTR-based) mRNA, whose transcription may be blocked, or second, an over-stable (3′-UTR-based) mRNA which translation can be turned off, by means of appropriate therapeutic interventions. In the case of methylphenidate, where the drug acts by increasing extra-cellular dopamine, a feedback can be proposed, so that the production of further DAT protein is dampened. However, by means of environmental stimulation, also the cognitive and behavioral therapies may turn out to produce quite easily the same effect.
To explain how genotypes interact with mRNAs, it would be enough to elucidate which mechanism, between the 3′-UTR and the 5′-UTR-based ones (just described above), is likely prone to environmental modulation. If environment can switch off translation, when a 3′-UTR VNTR allele confers excessive stability to its mRNAs, the 9-repeat allele is candidate for this role and the 10-repeat allele is then linked to 5′-UTR CpG-based inflexible transcription. If environment can switch off transcription, when a 5′-UTR CpG allele confers allowance to produce its mRNAs, the 9-repeat allele is candidate for this role and the 10-repeat allele is linked to 3′-UTR VNTR-based inflexible translation. By deduction, the second possibility appears much more likely, since an eventual switch off of transcription on one allele (via the 5′-UTR) would leave unaltered translation of over-stable mRNA by the other allele; conversely, the eventual switch off of translation would act on mRNAs from both alleles.
Others already suggested that DNA methylation can be influenced by cis-acting DNA sequence variation located on the same chromosome [5, 28, 57, 59, 81] and our results might thus be seen of relevance in the attempt to integrate genetic variants and DNA methylation. A possible concern might be the distance between the CpG sites under study and the VNTR. These CpG sites are located ≈1000 of bp away from the VNTR; however, it has been demonstrated that regulation in cis arrangement can actually occur at great distances [5].
It is tempting to speculate that 10-repeat VNTR allele confers a great stability to mRNAs, enhancing the likelihood that few mRNAs give still rise to a lot of protein, which production in turn may be out of control. Indeed, as this is an hypothesis based on environmental effects on transcription through the 5′-UTR, there is no way for environment to act on over-stable mRNA produced by 10-repeat 3′-UTR. We shall posit, conversely, that M2 and M6 are sites for binding of a molecule that allows transcription. Normally, when these sites are methylated, many more of normally stable mRNAs are translated; however, it is relatively easy to get them blocked, as environment is able to remove the molecule allowing mRNAs to be translated (after few weeks of therapeutic stimuli).
Specifically, as far as CpG M1 (1,444,716; chromosome 5) methylation is concerned, these levels are closely associated with DAT-aAbs titers in 10/10 patients, as both are indicative of ADHD severity in these patients. To discuss the role of M1, we underline that expansion of a CGG repeat, in the 5′-UTR of the FMR1 gene, is a genetic anomaly that, when accompanied by epigenetic modifications (mainly DNA methylation), results in the inactivation of the FMR1 gene and X-fragile symptoms (see [26, 74]). The importance of DNA 5′-UTR methylation is confirmed by the rare males who are unaffected by X-fragile, since they carry unmethylated full mutations, hence not repressing FMR1 transcription. By applying a similar reasoning to our data, we suppose DAT levels to be very high within the DAT 10/10 patients; as such, hypermethylation at M1 as well as high DAT-aAbs may be two attempts of trying to reduce DAT transcription or function, respectively. This interpretation implies that translation is elevated, as the 3′-UTR confers over-stability to 10-repeat mRNAs.
Conversely, CpG M2 or M6 (1,444,713 or 1,444,685 on chromosome 5) are closely associated with DAT-aAbs titers in DAT 9/x patients, as both are indicative of likelihood to respond positively after treatment in these patients. We may suppose DAT levels to be high as well, therefore promoting ADHD, but to possess room for being then diminished. Such interpretation implies that an elevated translation can still be reduced, as the stability of 3′-UTR 9-repeats mRNAs may be modulated. In addition, the CGCG motif (M6) is a putative target for members of a novel family of calmodulin-binding transcription activators (CAMTAs), reported in various species: CAMTAs regulate transcription through binding to a specific CGCG element [73] and have intriguingly been recognized as integrators of stress responses [60]. As for the present data, the M6 methylation may modulate CAMTAs binding after few weeks of therapy indeed, and hence somewhat reduce or block the excessive mRNA transcription, at least on carriers of that DAT 9-repeat allele (in one copy at least). In support of this notion, we already reported [34] that recovery of ADHD symptoms was also yielding to a decrease of DAT aAbs, which in turn may mirror a reduced expression of DAT protein, at least on lymphocytes.
Remarks on the role of DAT 5′-UTR and/or DAT aAbs
Little attention was given so far to the 5′ untranslated region of SLC6A3 despite potential consequences on gene expression of sequence variation in this region [38, 53]. One work identified many single nucleotide polymorphisms (SNPs), covering exon 1 and intron 1 until the start of exon 2 (+2106). However, the role of the CpG methylation within these sites is less well understood.
In this line, a striking link appears to exist to psycho-immunological interactions [36, 42]. Recent literature provides a clear indication that anti-neuronal antibodies may target a wide range of CNS proteins, including neuro-receptors [25, 37, 49, 50, 80, 83]. Nearly nothing is known for DAT aAbs; yet, in the periphery (namely, in the bloodstream), dopamine can mediate communication between immune cells, and the cross-talk between the immune and nervous systems [10, 72]. Dopamine can probably be important for suppressing T-regulatory cells (Treg) which are involved in auto-immunity [19, 54]. Altered activity of Treg (due to abnormal dopamine) could lead to uncontrolled function of effector T cells and to auto-immunity [62, 63]. Thus, DAT over-expression outside the brain, as in lymphocytes of ADHD children, could lead to dysregulation of neuro-immune systems [4]. The issue of a role for neuro-immune components in ADHD deserves, however, further investigation.
The interactions between genetics, epigenetics, (auto)-immunity on one side, and social as well as familiar environment on the other hands, are at the basis of the concept of multifinality: according to the Developmental Psychopathology framework, a specific risk factor can produce a multiplicity of outcomes depending on other causes or developmental contexts [15, 16, 76]. Thus, the effect of methylation or of a polymorphism, associated with a maladaptive developmental outcome, may well vary depending on other environmental factors such as, e.g., the quality of parent–infant interactions or parental caregiving capacity [11, 56]. It is tempting to propose that methylation levels in correlation with maternal Conners’ scores of the present study may provide an index not only for children behavior in itself but more in general for a deviant mother–infant dynamics along the ADHD dimensions.
The importance of such environmental factors and of the proposed biomarkers (genotype, methylation, and aAbs for DAT as well other candidate genes) suggests the need, in future studies, to evaluate them in both parents and children. Moreover, a problematic issue to be addressed would be whether biomarkers found in our clinical sample have a certain degree of stability of if they vary over time during development. Molecular epigenetics as well as auto-antibodies, in fact, may well operate over quick and subtle or slow and persistent processes. Very importantly, it will be crucial to understand whether changes in these biomarkers are indicative of discrete, adverse environmental events (e.g., early traumatic experiences), or more complex, persistent factors (e.g., parental quality of caregiving, neglect, or maltreatment).
Conclusion
Unfortunately, to date, clinical diagnosis of ADHD is solely based on structured interviews or on questionnaires. Nevertheless, reliability of these criteria for ADHD diagnostics remains a matter of debate. This study provides a hint towards a new and potentially very useful biomarker for diagnosis and/or prognosis. Our clinical experimentation demonstrates that determination of DNA methylation levels at specific CpG residues within the 5′-UTR region of the human DAT gene, alone or in combination with quantitation in blood of auto-antibodies (aAbs) against specific peptide fragments of DAT, is a robust and reliable marker of ADHD.
Our present data suggest that CpG methylation and/or DAT aAbs can be used to diagnose ADHD at least in the patients who are homozygous for the 10-repeat VNTR alleles. Instead, in carriers of at least one 9-repeat allele, our available biomarkers can only predict the efficacy of therapeutic approaches, consisting of cognitive and behavioral interventions with or without psycho-stimulant drugs. The main limitation of this study is the relative small sample size. Another limitation may be represented by tools used to classify ADHD symptoms. First, CGAS scores are an estimator of global functioning and thus provide an indirect assay of ADHD severity, and second, Conners’ scores by teachers are lacking, which would add an insight into children functioning in a school context. Actually, in our previous paper [34], we report Conners’ scales for both teachers and fathers, along with their correlations to aAbs titers. Presently, we decided to limit our investigations to the two scores for which we had a first measure at recruitment and a second measure 6 weeks after, namely, CGAS and mothers’ Conners (as we could run formal correlations with delta-score changes). Further studies in larger series of patient—together with their parents and perhaps also first-degree relatives—and with an extensive psychometric panel are warranted to monitor the effectiveness of the proposed biomarkers, by ascertaining whether or not they show a consistent change during the progress of symptoms’ recovery upon various ADHD therapeutic treatments.
References
Abdolmaleky HM, Thiagalingam S, Wilcox M (2005) Genetics and epigenetics in major psychiatric disorders: dilemmas, achievements, applications, and future scope. Am J Pharmacogenom 5:149–160
Adriani W, Koot S, Columba-Cabezas S, Romano E, Travaglini D, van den Bos R, Granstrem O, Ali SF, Laviola G (2012) Immunization with DAT fragments is associated with long-term striatal impairment, hyper-activity and reduced cognitive flexibility in mice. Behav Brain Funct 8:54
Albayrak Ö, Friedel S, Schimmelmann BG, Hinney A, Hebebrand J (2008) Genetic aspects in attention-deficit/hyperactivity disorder. J Neural Transm 115(2):305–315
Auci DL, Fikrig S, Rodriquez J (1997) Methylphenidate and the immune system. J Am Acad Child Adolesc Psychiatry 36:1015–1016
Bell JT, Pai AA, Pickrell JK, Gaffney DJ, Pique-Regi R, Degner JF et al (2011) DNA methylation patterns associate with genetic and gene expression variation in HapMap cell lines. Genome Biol 12:R10
Bannon MJ (2005) The dopamine transporter: role in neurotoxicity and human disease. Toxicol Appl Pharmacol 204:355–360
Berridge KC (2007) The debate over dopamine’s role in reward: the case for incentive salience. Psychopharmacology 191:391–431
Biederman J, Monuteaux MC, Mick E, Spencer T, Wilens TE, Silva JM, Snyder LE, Faraone SV (2006) Young adult outcome of attention deficit hyperactivity disorder: a controlled 10-year follow-up study. Psychol Med 36:167–179
Brenet F, Moh M, Funk P, Feierstein E, Viale AJ, Socci ND, Scandura JM (2011) DNA methylation of the first exon is tightly linked to transcriptional silencing. PLoS ONE 6:e14524
Buttarelli FR, Fanciulli A, Pellicano C, Pontieri FE (2011) The dopaminergic system in peripheral blood lymphocytes: from physiology to pharmacology and potential applications to neuropsychiatric disorders. Curr Neuropharmacol 9:278–288
Caspi A, McClay J, Moffitt TE, Mill J, Martin J, Craig IW et al (2002) Role of genotype in the cycle of violence in maltreated children. Science 297(5582):851–854
Chambers RA, Potenza MN (2003) Neurodevelopment, impulsivity, and adolescent gambling. J Gambl Stud 19:53–84
Cheon KA, Ryu YH, Kim JW, Cho DY (2003) The homozygosity for 10-repeat allele at dopamine transporter gene and dopamine transporter density in Korean children with attention deficit hyperactivity disorder: relating to treatment response to methylphenidate. Eur Neuropsychopharmacol 15:95–101
Cicchetti D, Dawson G (2002) Editorial: multiple levels of analysis. Dev Psychopathol 14(03):417–420
Cicchetti D, Rogosch FA (1996) Equifinality and multifinality in developmental psychopathology. Dev Psychopathol 8:597–600
Cicchetti D, Rogosch FA (1999) Psychopathology as risk for adolescent substance use disorders: a developmental psychopathology perspective. J Clin Child Psychol 28(3):355–365
Cook EH, Stein MA, Krasowski MD, Cox NJ, Olkon DM, Kieffer JE et al (1995) Association of attention-deficit disorder and the dopamine transporter gene. Am J Hum Genet 56:993–998
Cortese S (2012) The neurobiology and genetics of attention-deficit/hyperactivity disorder (ADHD): what every clinician should know. Eur J Paediatr Neurol 16:422–433
Cosentino M, Fietta AM, Ferrari M, Rasini E, Bombelli R, Carcano E, Saporiti F, Meloni F, Marino F, Lecchini S (2007) Human CD4+CD25+ regulatory T cells selectively express tyrosine hydroxylase and contain endogenous catecholamines subserving an autocrine/paracrine inhibitory functional loop. Blood 109(2):632–642
Curatolo P, Paloscia C, D’Agati E, Moavero R, Pasini A (2009) The neurobiology of attention deficit/hyperactivity disorder. Eur J Paediatr Neurol 13:299–304
Curtis WJ, Cicchetti D (2007) Emotion and resilience: a multilevel investigation of hemispheric electroencephalogram asymmetry and emotion regulation in maltreated and non-maltreated children. Dev Psychopathol 19(03):811–840
D’Addario C, Dell’Osso B, Galimberti D, Palazzo MC, Benatti B, Di Francesco A, Scarpini E, Altamura AC, Maccarrone M (2013) Epigenetic modulation of BDNF gene in patients with major depressive disorder. Biol Psychiatry 73:e6–e7
D’Addario C, Dell’Osso B, Palazzo MC, Benatti B, Lietti L, Cattaneo E, Galimberti D, Fenoglio C, Cortini F, Scarpini E, Arosio B, Di Francesco A, Di Benedetto M, Romualdi P, Candeletti S, Mari D, Bergamaschini L, Bresolin N, Maccarrone M, Altamura AC (2012) Selective DNA methylation of BDNF promoter in bipolar disorder: differences among patients with BDI and BDII. Neuropsychopharmacology 37:1647–1655
D’Addario C, Micale V, Di Bartolomeo M, Stark T, Pucci M, Sulcova A, Palazzo M, Babinska Z, Cremaschi L, Drago F, Altamura AC, Maccarrone M, Dell’Osso B (2017) A preliminary study of endocannabinoid system regulation in psychosis: Distinct alterations of CNR1 promoter DNA methylation in patients with schizophrenia. Schizophr Res, in Press (EPub Jan 17)
Dalmau J, Lancaster E, Martinez-Hernandez E, Rosenfeld MR, Balice-Gordon R (2011) Clinical experience and laboratory investigations in patients with anti-NMDAR encephalitis. Lancet Neurol 10:63–74
Das S, Kubota T, Song M, Daniel R, Berry-Kravis EM, Prior TW, Popovich B, Rosser L, Arinami T, Ledbetter DH (1997) Methylation analysis of the fragile X syndrome by PCR. Genet Test 1:151–155
Dell’Osso B, D’Addario C, Palazzo MC, Benatti B, Camuri G, Galimberti D, Fenoglio C, Scarpini E, Di Francesco A, Maccarrone M, Altamura AC (2014) Epigenetic modulation of BDNF gene: differences in DNA methylation between unipolar and bipolar patients. J Affect Disord 166:330–333
Docherty SJ, Davis OS, Haworth CM, Plomin R, D’Souza U, Mill J (2012) A genetic association study of DNA methylation levels in the DRD4 gene region finds associations with nearby SNPs. Behav Brain Funct 8:31
Doerfler W (1983) DNA methylation and gene activity. Annu Rev Biochem 52:93–124
Egger G, Liang G, Aparicio A, Jones PA (2004) Epigenetics in human disease and prospects for epigenetic therapy. Nature 429:457–463
Faraone SV, Perlis RH, Doyle AE, Smoller JW, Goralnick JJ, Holmgren MA et al (2005) Molecular genetics of attention deficit/hyperactivity disorder. Biol Psychiatry 57:1313–1323
Faraone SV, Bonvicini C, Scassellati C (2014) Biomarkers in the diagnosis of ADHD—promising directions. Curr Psychiatry Rep 16:497
Gainetdinov RR, Caron MG (2003) Monoamine transporters: from genes to behavior. Annu Rev Pharmacol Toxicol 43:261–284
Giana G, Romano E, Porfirio MC, D’Ambrosio R, Giovinazzo S, Troianiello M, Barlocci E, Travaglini D, Granstrem O, Pascale E, Tarani L, Curatolo P, Laviola G, Adriani W (2015) Detection of auto-antibodies to DAT in the serum: interactions with DAT genotype and psycho-stimulant therapy for ADHD. J Neuroimmunol 278:212–222
Grayson DR, Chen Y, Costa E, Dong E, Guidotti A, Kundakovic M, Sharma RP (2006) The human reelin gene: transcription factors (+), repressors (−) and the methylation switch (±) in schizophrenia. Pharmacol Ther 111:272–286
Graus F, Saiz A, Dalmau J (2010) Antibodies and neuronal autoimmune disorders of the CNS. J Neurol 257:509–517
Graus F, Saiz A, Lai M, Bruna J, López F, Sabater L, Blanco Y, Rey MJ, Ribalta T, Dalmau J (2008) Neuronal surface antigen antibodies in limbic encephalitis: clinical-immunologic associations. Neurology 71:930–936
Greenwood TA, Kelsoe JR (2003) Promoter and intronic variants affect the transcriptional regulation of the human dopamine transporter gene. Genomics 82:511–520
Heinz A, Goldman D, James DW, Palmour R, Hommer D, Gorey JG et al (2000) Genotype influences in vivo dopamine transporter availability in human striatum. Neuropsychopharmacology 22:133–139
Henikoff S, Matzke MA (1997) Exploring and explaining epigenetic effects. Trends Genet 13:293–295
Hillemacher T, Frieling H, Buchholz V, Hussein R, Bleich S, Meyer C, John U, Bischof A, Rumpf HJ (2015) Alterations in DNA-methylation of the dopamine-receptor 2 gene are associated with abstinence and health care utilization in individuals with a lifetime history of pathologic gambling. Prog Neuropsychopharmacol Biol Psychiatry 63:30–34
Hoekstra PJ, Minderaa RB (2005) Tic disorders and obsessive-compulsive disorder: is autoimmunity involved? Int Rev Psychiatry 17:497–502
Hollander E, Buchalter AJ, DeCaria CM (2000) Pathological gambling. Psychiatr Clin North Am 23:629–642
Hollander E, Sood E, Pallanti S, Baldini-Rossi N, Baker B (2005) Pharmacological treatments of pathological gambling. J Gambl Stud 21:99–110
Holliday R (1987) DNA methylation and epigenetic defects in carcinogenesis. Mutat Res 181:215–217
Jirtle RL, Skinner MK (2007) Environmental epigenomics and disease susceptibility. Nat Rev Genet 8:253–262
Joober R, Grizenko N, Sengupta S, Amor LB, Schmitz N, Schwartz G, Karama S, Lageix P, Fathalli F, Torkaman-Zehi A, Ter Stepanian M (2007) Dopamine transporter 3′-UTR VNTR genotype and ADHD: a pharmaco-behavioural genetic study with methylphenidate. Neuropsychopharmacology 32:1370–1376
Jucaite A, Fernell E, Halldin C, Forssberg H, Farde L (2005) Reduced midbrain dopamine transporter binding in male adolescents with attention-deficit/hyperactivity disorder: association between striatal dopamine markers and motor hyperactivity. Biol Psychiatry 57:229–238
Lai M, Hughes EG, Peng X, Zhou L, Gleichman AJ, Shu H, Matà S, Kremens D, Vitaliani R, Geschwind MD, Bataller L, Kalb RG, Davis R, Graus F, Lynch DR, Balica-Gordon R, Dalmau J (2009) AMPA receptor antibodies in limbic encephalitis alter synaptic receptor location. Ann Neurol 65:424–434
Lancaster E, Lai M, Peng X, Hunghes E, Constantinescu R, Raizer J, Friedman D, Skeen MB, Grisold W, Kimura A, Ohta K, Iizuka T, Guzman M, Graus F, Moss SJ, Balice-Gordon R, Dalmau J (2010) Antibodies to the GABA(B) receptor in limbic encephalitis with seizures: case series and characterisation of the antigen. Lancet Neurol 9:67–76
Levenson JM, Sweatt JD (2006) Epigenetic mechanisms: a common theme in vertebrate and invertebrate memory formation. Cell Mol Life Sci 63:1009–1016
Lopez-Serra L, Ballestar E, Fraga MF, Alaminos M, Setein F, Esteller M (2006) A profile of methyle CpG binding domain protein occupancy of hypermethylated promoter CpG islands in tumor suppressor genes in human cancer. Cancer Res 66:8342–8346
Kelada SN, Costa-Mallen P, Checkoway H, Carlson CS, Weller TS, Swanson PD, Franklin GM, Longstreth WT Jr, Afsharinejad Z, Costa LG (2005) Dopamine transporter (SLC6A3) 5′ region haplotypes significantly affect transcriptional activity in vitro but are not associated with Parkinson’s disease. Pharmacogenet Genom 15:659–668
Kipnis J, Cardon M, Avidan H, Lewitus GM, Mordechay S, Rolls A, Shani Y, Schwartz M (2004) Dopamine, through the extracellular signal-regulated kinase pathway, downregulates CD4 + CD25 + regulatory T-cell activity: implications for neurodegeneration. J Neurosci 24:6133–6143
Martinez D, Gelernter J, Abi-Dargham A, van Dyck CH, Kegeles L, Innis RB et al (2001) The variable number of tandem repeats polymorphism of the dopamine transporter gene is not associated with significant change in dopamine transporter phenotype in humans. Neuropsychopharmacology 24:553–560
Martins C, Gaffan EA (2000) Effects of early maternal depression on patterns of infant-mother attachment: a meta-analytic investigation. J Child Psychol Psychiatry 41(6):737–746
Milani L, Lundmark A, Nordlund J, Kiialainen A, Flaegstad T, Jonmundsson G, Kanerva J, Schmiegelow K, Gunderson KL, Lönnerholm G, Syvänen AC (2009) Allele-specific gene expression patterns in primary leukemic cells reveal regulation of gene expression by CpG site methylation. Genome Res 19:1–11
Mill J, Asherson P, Browes C, D’Souza U, Craig I (2002) Expression of the dopamine transporter gene is regulated by the 3′-UTR VNTR: evidence from brain and lymphocytes using quantitative RT-PCR. Am J Med Genet 114:975–979
Mill J, Tang T, Kaminsky Z, Khare T, Yazdanpanah S, Bouchard L et al (2008) Epigenomic profiling reveals DNA-methylation changes associated with major psychosis. Am J Hum Genet 82:696–711
Mollet IG, Malm HA, Wendt A, Orho-Melander M, Eliasson L (2016) Integrator of stress responses calmodulin binding transcription activator 1 (Camta1) regulates miR-212/miR-132 expression and insulin secretion. J Biol Chem 291:18440–18452
Oades RD (1998) Frontal, temporal and lateralized brain function in children with attention-deficit hyperactivity disorder: a psychophysiological and neuropsychological viewpoint on development. Behav Brain Res 94:83–95
Pacheco R, Contreras F, Zouali M (2014) The dopaminergic system in autoimmune diseases. Front Immunol 5:117
Pacheco R, Riquelme E, Kalergis AM (2010) Emerging evidence for the role of neurotransmitters in the modulation of T cell responses to cognate ligands. Cent Nerv Syst Agents Med Chem 10:65–83
Pasini A, Sinibaldi L, Paloscia C, Douzgou S, Pitzianti MB, Romeo E, Curatolo P, Pizzuti A (2013) Neurocognitive effects of methylphenidate on ADHD children with different DAT genotypes: a longitudinal open label trial. Eur J Paediatr Neurol 17:407–414
Petronis A (2003) Epigenetics and bipolar disorder: new opportunities and challenges. Am J Med Genet C Semin Med Genet 123C:65–75
Pidsley R, Mill J (2011) Research Highlights: epigenetic changes to serotonin receptor gene expression in schizophrenia and bipolar disorder. Epigenomics 3:537–538
Polanczyk G, De Lima MS, Horta BL, Biederman J, Rohde LA (2007) The worldwide prevalence of ADHD: a systematic review and metaregression analysis. Am J Psychiatry 164:942–948
Purper-Ouakil D, Ramoz N, Lepagnol-Bestel AM, Gorwood P, Simonneau M (2011) Neurobiology of attention deficit/hyperactivity disorder. Pediatr Res 69:69–76
Rizzo R, Gulisano M, Calì PV, Curatolo P (2010) ADHD and epilepsy in children with Tourette syndrome: a triple comorbidity? Acta Paediatr 99:1894–1896
Rizzo R, Gulisano M, Calì PV, Curatolo P (2013) Tourette Syndrome and comorbid ADHD: current pharmacological treatment options. Eur J Paediatr Neurol 17:421–428
Sagvolden T, Sergeant JA (1998) Attention deficit/hyperactivity disorder—from brain dysfunction to behaviour. Behav Brain Res 94:1–10
Sarkar C, Basu B, Chakroborty D, Dasgupta PS, Basu S (2010) The immunoregulatory role of dopamine: an update. Brain Behav Immun 24:525–528
Shen C, Yang Y, Du L, Wang H (2015) Calmodulin-binding transcription activators and perspectives for applications in biotechnology. Appl Microbiol Biotechnol 99:10379–10385
Tabolacci E, Chiurazzi P (2013) Epigenetics, fragile X syndrome and transcriptional therapy. Am J Med Genet A. 161A:2797–2808
Thapar A, Holmes J, Poulton K, Harrington R (1999) Genetic basis of attention deficit and hyperactivity. Br J Psychiatry 174(2):105–111
Tronick E, Hunter RG (2016) Waddington, dynamic systems, and epigenetics. Front Behav Neurosci 10:107
Tsankova N, Renthal W, Kumar A, Nestler EJ (2007) Epigenetic regulation in psychiatric disorders. Nat Rev Neurosci 8:355–367
Wallis D (2010) The search for biomarkers for attention deficit/hyperactivity disorder. Drug News Perspect 23:438–449
Yang B, Chan RC, Jing J, Li T, Sham P, Chen RY (2007) A meta-analysis of association studies between the 10-repeat allele of a VNTR polymorphism in the 3′-UTR of dopamine transporter gene and attention deficit hyperactivity disorder. Am J Med Genet B Neuropsychiatr Genet 144B:541–550
Zandi MS, Irani SR, Lang B, Waters P, Jones PB, McKenna P, Coles AJ, Vincent A, Lennox BR (2010) Disease-relevant autoantibodies in first episode schizophrenia. J Neurol 258:686–688
Zhang D, Cheng L, Badner JA, Chen C, Chen Q, Luo W et al (2010) Genetic control of individual differences in gene-specific methylation in human brain. Am J Hum Genet 86:411–419
Zoratto F, Romano E, Pascale E, Pucci M, Falconi A, Dell’Osso B, Maccarrone M, Laviola G, D’Addario C, Adriani W (2017) Down-regulation of serotonin and dopamine transporter genes in individual rats expressing a gambling-prone profile: a possible role for epigenetic mechanisms. Neuroscience 340:101–116
Zuliani L, Graus F, Giometto B, Bien C, Vincent A (2012) Central nervous system neuronal surface antibody associated syndrome: review and guidelines for recognition. J Neurol Neurosurg Psychiatry 83:638–645
Acknowledgements
Research performed under the young-investigator “ADHD-sythe” project (“under 40” call 2007) and the EU “NeuroGenMRI” project (ERAnet “PrioMedChild”), both from Italian Ministry of Health (for Italian partner, to W. A.); in part, supported under Grant Agreements ref. 278367 “EMTICS—European Multicentre Tics In Children Studies” and ref. 603016 “MATRICS—Multidisciplinary Approaches to Translational Research In Conduct Syndromes” by European Seventh Framework Programme (FP7) 2007–2013 (for Italian partner, to G. L.). We acknowledge Dr Nadia Francia and Dr Viola Cardia for their precious help with all management and patenting issues. There are only two items for potential conflict of interest to be disclosed: (1) Adriani W, Laviola G, Pascale E, D’Addario C “Metodo Per Determinare Il Deficit Di Attenzione Con Iperattività (Method to determine attention deficit and hyperactivity disorder)”; Patent Application, deposit in ITALY at No. 102016000129938 (22-DECEMBER-2016). (2) Granstrem O, Adriani W, Laviola G, Porfirio MC, Curatolo P “Biomarkers for validation of A.D.H.D. diagnosis and monitoring of therapy efficacy”; Full Patent No. PN810701WO; Int.l Application No. PCT/EP2013/066845; Publication number WO/2014/023852 (10-AUGUST-2013).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Adriani, W., Romano, E., Pucci, M. et al. Potential for diagnosis versus therapy monitoring of attention deficit hyperactivity disorder: a new epigenetic biomarker interacting with both genotype and auto-immunity. Eur Child Adolesc Psychiatry 27, 241–252 (2018). https://doi.org/10.1007/s00787-017-1040-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00787-017-1040-9