Abstract
Pathways regulating threonine, methionine and isoleucine metabolism are very efficiently interconnected in plants. As both threonine and methionine serve as substrates for isoleucine synthesis, their synthesis and catabolism under different developmental and environmental conditions also influence isoleucine availability. Together, methionine gamma-lyase and threonine deaminase maintain the isoleucine equilibrium in plants under varied substrate availabilities. Isoleucine and the two other branched-chain amino acids (BCAAs) (leucine and valine) share four common enzymes in their biosynthesis pathways and thus are coordinately regulated. Induction of free amino acids as osmolytes in response to abiotic stress is thought to play a role in plant stress tolerance. In particular, the accumulation of BCAAs is induced many-fold during osmotic stress. However, unlike in the case of proline, not much research has been focused on understanding the function of the response involving BCAAs. This review describes pathways influencing branched-chain amino acid metabolism and what is known about the biological significance of their accumulation under abiotic stress. A bioinformatics approach to understanding the transcriptional regulation of the genes involved in amino acid metabolism under abiotic stress is also presented.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Unlike plants, animals cannot synthesize all amino acids of primary metabolism by themselves and must obtain the so-called essential amino acids in their diets. For instance, in many monocot grain crops, quantities of two essential amino acids, threonine and methionine, are less abundant than required for human diets and so have to be supplemented from other sources such as legumes, meat, or fish. Poultry, swine, and other non-ruminant domestic animals have specific requirements for each of the essential amino acids, which are often fulfilled by supplementing grains with commercially synthesized amino acids. Such enriched diets, e.g., maize and soybean meal-based diets supplemented with lysine for mammals or methionine for birds, also reduce animal nitrogen excretion by providing an improved balance of the essential amino acids (Newell-McGloughlin 2008).
Targeted manipulation of biosynthetic pathways, plant breeding, and mutant selection (Mertz et al. 1964; Munck et al. 1970; Singh and Axtell 1973; Galili 1995; Imsande 2001) have been used to enhance the levels of either free amino acids or storage proteins, with variable success rates. Unfortunately, because several essential amino acids are synthesized via common substrates and competing enzymes, metabolic engineering attempts to enhance one or more amino acids directly affect metabolism of non-targeted amino acids. Due to the fact that branched-chain amino acids (BCAAs) are essential in human nutrition and agriculture (Singh 1999; Binder et al. 2007), plant-based biosynthesis of isoleucine, leucine, and valine has been the subject of extensive research. Among the three BCAAs, isoleucine belongs to aspartate-derived pathway, whereas valine and leucine are derived from pyruvate. Pathways involving BCAAs in plants are well-investigated because they provide precursors for a number of plant secondary metabolites (Lea and Ireland 1999) and some of the pathway enzymes are herbicide targets (Wittenbach and Abell 1999). For instance, several classes of commercially successful herbicides inhibit the enzyme acetolactate synthase, which is required for the biosynthesis of all three BCAAs (Saari et al. 1994; Singh and Shaner 1995). Recently, yet another important aspect of BCAA metabolism was explored when Prell et al. (2009) confirmed that the supply of BCAAs by the plant is essential for the development of bacteroids and ultimately for symbiotic nitrogen fixation in peas, and possibly legumes generally.
In this review, we discuss the pathways involved in the synthesis of BCAAs derived from threonine and methionine, as well as the significance of their accumulation under abiotic stress. We also propose a bioinformatic analysis scheme based on publicly available microarray data from experiments conducted with Arabidopsis thaliana (Arabidopsis) to elucidate the transcriptional regulation of metabolic genes involved in the abiotic stress-induced amino acid biosynthesis.
Threonine and methionine synthesis and catabolism maintain isoleucine homeostasis in plants
Isoleucine is synthesized from threonine and methionine, which are derived from aspartate via enzymes localized to the plastids. The aspartate-derived amino acid pathway is active during seed development (Galili 1995), and the timing of expression of the threonine biosynthesis genes occurs relatively late during seed development, similar to the timing of storage protein gene expression (Karchi et al. 1993). This allows molecular manipulation of various biosynthetic enzymes to enhance accumulation of precursors such as threonine and/or methionine, and effectively all of the BCAAs. Homoserine kinase (EC 2.7.1.39), which catalyzes the formation of O-phosphohomoserine from homoserine, leads to the formation of either threonine or methionine. Experiments with pea and radish suggest that this enzyme activity is subject to regulation by isoleucine and valine (Thoen et al. 1978; Baum et al. 1983). However, other research with cloned Arabidopsis homoserine kinase did not show such allosteric regulation (Lee and Leustek 1999). Biosynthesis of threonine or methionine in plants is determined by competitive affinities of threonine synthase (EC 4.2.3.1) and cystathionine γ-synthase (EC 2.5.1.48) for a common pool of O-phosphohomoserine. Under conditions of excess methionine, threonine synthase activity is up-regulated by methionine-derived S-adenosylmethionine to help reestablish threonine and methionine equilibrium. Compared to that of cystathionine γ-synthase, the Km of threonine synthase for O-phosphohomoserine drops by 250–500-fold in the presence of S-adenosylmethionine, favoring carbon flow into threonine biosynthesis (Curien et al. 1996; Laber et al. 1996, 1999; Amir et al. 2002). This allosteric regulation mechanism has been confirmed in vitro by independent studies using the crystal structure and enzyme activities of threonine synthase, with and without S-adenosylmethionine (Thomazeau et al. 2001; Curien et al. 2003; Mas-Droux et al. 2006). These studies allowed transgenic approaches for manipulation of threonine synthase activity to enhance methionine levels in plants. Both Arabidopsis threonine synthase point mutations (Bartlem et al. 2000) and antisense constructs in potatoes (Zeh et al. 2001) reduce threonine synthase activity and elevate methionine accumulation.
Mathematical modeling of the aspartate-derived amino acid pathway suggests that threonine concentrations are quite sensitive to perturbation (Curien et al. 2009). Therefore, regulated threonine catabolism may play a central role in the aspartate-derived amino acid pathway. Unlike threonine synthesis, threonine catabolism in plants is initiated by two competing pathways (Fig. 1), one through threonine deaminase/dehydratase (EC 4.3.1.19) and other through threonine aldolase (EC 4.1.2.5). Threonine deaminase, which is localized to the plastids, produces 2-oxobutanoate, an intermediate required for isoleucine synthesis. However, threonine aldolase converts threonine to glycine and acetaldehyde. As neither of the two Arabidopsis threonine aldolases has a predicted plastid-targeting signal peptide, this reaction likely occurs outside of the plastids. Mutations in one of the two Arabidopsis threonine aldolases greatly increase seed threonine (Jander et al. 2004) and cysteine (Lu et al. 2008; Ajjawi et al. 2009) content, whereas knockout mutations in the other are lethal (Joshi et al. 2006). Rescue of this lethal effect by over-expression of feedback-insensitive threonine deaminase is accompanied by excess isoleucine accumulation, suggesting competing activities of these two enzymes. Although most of the glycine required by plants is produced during photorespiration or as a branch off the glycolysis pathway, survival of an Arabidopsis double mutant lacking both threonine aldolases (Joshi et al. 2006) likely underestimates the metabolic need of threonine-derived glycine. The reduced glycine content in the seedlings of a threonine aldolase mutant and the observation that less glycine is available for cell metabolism under high photorespiratory rates (Bourguignon et al. 1998) suggest differential tissue-specific and developmental needs for threonine-derived glycine in plants. Tissue-specific expression patterns of threonine aldolases in Arabidopsis (Joshi et al. 2006) suggest a broader role for these enzymes in maintaining glycine equilibrium or detoxifying excess threonine. Another as yet untested hypothesis is that threonine aldolases could be critical for degrading excess, possibly toxic amounts of threonine, for instance under the conditions where elevated isoleucine feedback inhibits threonine deaminase.
Threonine deaminase in plants is feedback-inhibited by all three BCAAs. Repeat structures in the threonine deaminase enzyme provide separate binding sites for each BCAA (Wessel et al. 2000). Although inhibition of threonine deaminase is herbicidal, deleterious effects can be counteracted by supplementation with 2-ketobutyrate or isoleucine (Szamosi et al. 1994; Mourad and King 1995). Garcia and Mourad (2004) isolated Arabidopsis feedback-insensitive threonine deaminase mutants by ethylmethanesulfonate and site-directed mutagenesis. These mutants accumulated high free isoleucine in the whole plant. However, increased accumulation of isoleucine in the seeds was only observed when threonine aldolase mutant plants were complemented by a feedback-insensitive threonine deaminase mutant allele (Joshi et al. 2006), suggesting that the seed-specific threonine deaminase activity is dependent on substrate availability.
Methionine metabolism in plants is discussed in more detail in another review in this issue (Amir 2010). However, it is particularly relevant to mention methionine catabolism here due to its involvement in isoleucine synthesis. S-Adenosylmethionine synthase (EC 2.5.1.6) directs about 80% of the metabolic flux of methionine to S-adenosylmethionine, which is used to methylate nucleic acids, proteins, lipids, and numerous other plant metabolites. Like other enzymes involved in methionine catabolism, methionine γ-lyase (EC 4.4.1.11) is also not localized to plastids and produces 2-ketobutyrate, methanethiol, and ammonia from methionine in cytoplasm (Fig. 1). This enzyme has been studied extensively in microbes and protozoa (Inoue et al. 1995; Faleev et al. 1996; Hori et al. 1996; Dias and Weimer 1998; McKie et al. 1998; Tokoro et al. 2003; Manukhov et al. 2005). Independent knockout mutations in Arabidopsis methionine γ-lyase increase S-methylmethionine and methionine accumulation in leaves under sulfate-limiting growth conditions (Goyer et al. 2007), as well as free methionine in flowers and seeds under normal growth conditions (Joshi and Jander 2009). Methanethiol produced by methionine γ-lyase is emitted in plants in response to the excess sulfur or methionine accumulation (Schmidt et al. 1985; Boerjan et al. 1994; Saini et al. 1995) and is also incorporated into cysteine (Rebeille et al. 2006; Goyer et al. 2007).
Metabolic profiling experiments carried out using Arabidopsis cells supplemented with labeled [13C]methionine identified labeled S-adenosylmethionine, S-methylmethionine, and isoleucine (Rebeille et al. 2006), suggesting an alternate route of threonine-independent isoleucine synthesis (Fig. 1). Methionine γ-lyase activity was verified in vivo in wild-type Arabidopsis flowers and siliques when labeled isoleucine was recovered from cut stems of plants fed with labeled methionine (Joshi and Jander 2009). Spraying with formasulfuran dramatically increases plant methionine content (Trenkamp et al. 2009) and spraying with sulfonylurea abolishes isoleucine biosynthesis (Rebeille et al. 2006). Inhibition of the same enzyme, acetolactate synthase, by both of these herbicides suggests a direct role for methionine in isoleucine biosynthesis. Additional indirect evidence that supports threonine-independent isoleucine synthesis comes from research showing increased isoleucine synthesis under the conditions that enhance methionine accumulation. For example, increased isoleucine accumulation is observed in potatoes expressing antisense threonine synthase (Zeh et al. 2001), potatoes over-expressing cystathionine γ-synthase (Dancs et al. 2008), and tobacco expressing methionine-insensitive cystathionine γ-synthase genes from Arabidopsis (Hacham et al. 2008).
Transcription of methionine γ-lyase in plants is strongly regulated by exogenous methionine (Rebeille et al. 2006), low sulfate (Goyer et al. 2007), osmotic stress (Less and Galili 2008; Joshi and Jander 2009), and availability of threonine and/or isoleucine (Joshi and Jander 2009). Cornah et al. (2004) found almost 50-fold induction in methionine γ-lyase, along with 8-fold induction in threonine aldolase expression, in Arabidopsis seedlings defective in isocitrate lyase, an enzyme that is otherwise required in the glyoxylate cycle. It is possible that the increased need of glycine in the absence of glyoxylate synthesis is fulfilled by induction of threonine aldolase, leaving less substrate for threonine-dependent isoleucine synthesis via threonine deaminase. Increased methionine γ-lyase transcription thus can fulfill the isoleucine needs caused by limited threonine deaminase activity. Such complementary functional roles of these two Arabidopsis enzymes are confirmed by two observations (a) that feedback-insensitive mutations in threonine deaminase down-regulate methionine γ-lyase transcription and (b) that the isoleucine deficit in a threonine deaminase knock-down mutant is rescued by methionine γ-lyase overproduction (Joshi and Jander 2009).
Biosynthesis of BCAAs in plants
Several detailed reviews that focus on BCAA metabolism in plants have been published recently (Singh and Shaner 1995; Singh 1999; Binder et al. 2007; Jander and Joshi 2009). Biosynthesis of BCAAs occurs exclusively in plastids (Schulze-Siebert et al. 1984) and many of the genes involved in their metabolism are found to carry plastid-targeting signal peptides. However, the catabolism of BCAA, which requires β-oxidation, might occur in mitochondria, peroxisomes, or both (Graham and Eastmond 2002). One unique feature of the BCAA metabolism is that the four biosynthetic enzymes are common to all three amino acids, even though the substrates are different (Fig. 2). Coordinated regulation of BCAAs in plants has been shown by identification of pathway quantitative trait loci (QTL) in pericarp metabolites using a set of tomato introgression lines (Schauer and Fernie 2006), in an ethylmethanesulfonate-induced Arabidopsis mutant screen for increased seed amino acids (Jander et al. 2004), and in Arabidopsis crosses segregating for a mutation in BCAA catabolism (Gu et al. 2009).
The first step in all BCAA syntheses is catalyzed by acetohydroxyacid synthase/acetaolactate synthase (EC 4.1.3.18) (Chipman et al. 1998; Duggleby and Pang 2000). This enzyme catalyzes condensation of two pyruvate molecules to acetolactate, as well as the synthesis of 2-acetohydroxybutyrate from pyruvate and 2-ketobutyrate (Singh and Shaner 1995). Acetolactate synthase activity is feedback-inhibited by leucine and valine (Singh et al. 1988; Durner and Boger 1990; Southan and Copeland 1996; Lee and Duggleby 2001), as well as with greater efficiency by synergistic combinations of BCAAs (Durner and Boger 1988). Such synergism in enzyme inhibition is not observed in the corresponding bacterial and yeast enzymes. Acetolactate synthase consists of large catalytic subunits and small regulatory subunits (Tan et al. 2005). Five commonly occurring mutations that render tolerance to acetolactate synthase-inhibiting herbicides have been found in the large subunit (Tranel and Wright 2002; Jander et al. 2003; Tan et al. 2005). More than 100 weed species have been observed to show resistance to acetolactate synthase inhibitors due to one or more of these mutations (Heap 2009). The herbicides inhibiting acetolactate synthase have no resemblance to the natural substrates and are not competitive inhibitors, suggesting that they bind at a site that is distinct from the enzyme’s active site (Chang and Duggleby 1998), either at the entry site for the substrate or the substrate access channel (Ott et al. 1996; Pang et al. 2003). Resistance mutations reduce binding to the acetolactate synthase enzyme core by some or all inhibitor classes without affecting enzyme activity (Tranel and Wright 2002). The known herbicide target sites in acetolactate synthase are also distinct from those that provide feedback inhibition by BCAAs, and mutant enzymes that are no longer subject to feedback regulation are nevertheless sensitive to herbicides (Wu et al. 1994).
Ketol-acid reductoisomerase (EC 1.1.1.86) and dihydroxy-acid dehydratase (EC 4.2.1.9), which catalyze the next two steps in the shared BCAA biosynthesis pathway (Fig. 2), remain largely uncharacterized in plants (Binder et al. 2007). Although Arabidopsis candidate genes have been identified based on sequence similarities to microbial enzymes, their enzymatic activities and regulatory mechanisms have not been verified. However, spinach (Dumas et al. 1992) and rice (Leung and Guddat 2009) ketol-acid reductoisomerase enzymes have been crystallized and offer potential targets for designing herbicide resistance.
The final transamination step in the synthesis of BCAAs, as well as the first step in the degradative pathways, is catalyzed by BCAA transferases (EC 2.6.1.42) (Binder et al. 2007). A small gene family of six transcribed members in Arabidopsis has been identified by complementation of yeast knockout mutations (Diebold et al. 2002). These are targeted to different sub-cellular locations in the plant, one in the mitochondria, two in the cytosol, and three in the plastids (Diebold et al. 2002). Based on its chloroplast localization, BCAT2 was predicted to be involved in biosynthesis rather than in degradation (Schuster and Binder 2005). This was confirmed recently through drought stress effects on amino acid accumulation in an abscisic acid (ABA)-defective Arabidopsis mutant: Urano et al. (2009) reported that the dehydration-inducible accumulation of BCAAs was correlated with induced expression of BCAT2 gene, which is regulated by endogenous ABA. Nicotiana benthamiana BCAT enzyme has been shown to be involved in the regulation of endogenous hormones by its effect on KNOTTED-like homeobox (KNOX) genes (Gao et al. 2009).
As Binder et al. (2007) have published a thorough review describing various enzymatic steps involved in the catabolism of individual BCAAs in plants, we have opted to leave out the details of this biochemical pathway. Even though much progress has been made in understanding these catabolic routes, the importance of BCAA degradation in plants is still unclear. Excessive accumulation of branched-chain α-keto acids is known to be cytotoxic and can induce apoptosis in mammals (Chuang and Shih 1995; Eden and Benvenisty 1998), but no such observations have been reported in plants. Many-fold higher accumulation of BCAAs during transient osmotic stress followed by rapid depletion during the recovery process clearly implies tight regulation of the BCAA catabolism by environmental perturbation. Additionally, the breakdown products of BCAAs, which include acetyl-CoA, propionyl-CoA, and acetoacetate, are potential energy sources for plants. It has been suggested that, to avoid excessive accumulation, BCAAs promote their own degradation during seed germination, senescence, or under sugar starvation, and thereby provide alternative carbon sources for plants during stress conditions (Taylor et al. 2004). Recently, it was demonstrated that branched-chain and aromatic amino acid catabolism leads to the synthesis of unique aroma volatiles in melon, Cucumis melo (Gonda et al. 2010).
Unlike synthesis, degradation of BCAAs has been suggested to occur outside plastids, either in mitochondria (Anderson et al. 1998) or peroxisomes (Gerbling and Gerhardt 1989). This also explains the need of differential sub-cellular localization of BCAA aminotransferases and, in particular, the Arabidopsis mitochondrial BCAA aminotransferase has been suggested to initiate degradation and perform a central regulatory role in the BCAA turnover (Diebold et al. 2002; Schuster and Binder 2005). Knockout of Arabidopsis isovaleryl-CoA dehydrogenase (At3g45300; E.C. 1.3.99.10), the third enzyme in the leucine catabolic pathway, dramatically increases the accumulation of all three BCAAs in seeds and, by as yet unknown regulatory mechanisms, also increases accumulation of homomethionine and several other amino acids (Gu et al. 2009).
Significance of BCAA accumulation under abiotic stresses
The elevated accumulation of metabolites in response to abiotic stress is an important aspect of plant amino acid metabolism. Plants are subjected to various abiotic stresses, e.g., drought, flooding, heat, and salt, throughout their lives. Osmotic stress, in particular due to drought, salinity and flooding, is considered the most serious problem that limits agricultural crop productivity (Ceccarelli and Grando 1996). As a general response to such stress, all plants, even halophytes, accumulate amino acids, betaines, sugars, organic acids, and other osmolytes in the cytoplasm (Delauney and Verma 1993; Parida and Das 2005). Although this accumulation of osmolytes is ubiquitous in nature, their concentration is dependent on the particular micro-environment, cell or tissue types, developmental status, and the nature and longevity of the particular abiotic stress. A detailed review describing function and regulation of osmolytes in different biological systems has been published recently by Burg and Ferraris (2008).
Amino acid metabolism may play an important role in plant stress tolerance through the accumulation of compatible osmolytes (Campalans et al. 1999), by intracellular pH regulation, and by detoxification of reactive oxygen species, xenobiotics, and heavy metals (Nuccio et al. 1999; Alia et al. 2001). During drought stress, protein residues may be altered by chemical processes and some proteins are degraded by proteases after being irreversibly damaged by the effects of drought stress. In many plant species, most amino acids have been observed to show a characteristic linear increase with the induction of drought stress, followed by a reduction in concentration upon rehydration (Delauney and Verma 1993; Rhodes and Hanson 1993; Good and Zaplachinski 1994). In addition to de novo synthesis, these elevated amino acid pools could result from reduced protein synthesis (Good and Zaplachinski 1994) or general protein breakdown during drought stress. It has been suggested that proteases mobilize amino acids from proteins for the synthesis of compatible osmolytes (Campalans et al. 1999). Less and Galili (2008) studied the expression of all annotated Arabidopsis proteases in response to drought and other abiotic stress, and concluded that increased protease production cannot account for the observed increase in free amino acid accumulation. Nevertheless, one cannot rule out the possibility of up-regulated protein targeting to already existing protease complexes, e.g., by the activity of E3 ubiquitin ligases, in response to osmotic stress.
ABA plays an essential role in plant reactions to osmotic stress, influencing stomatal closure, salt sequestration, growth modifications, production of osmolytes, and other responses. Reduced BCAA accumulation in an ABA-defective Arabidopsis mutant (aba2) and failure to enhance BCAA accumulation by exogenous ABA treatment (Nambara et al. 1998) suggested a role for endogenous ABA in this response. Urano et al. (2009) showed that dehydration stress induces endogenous ABA accumulation, which could regulate global metabolic networks and change the levels of amino acids. Among amino acids that are induced by drought stress, proline has been studied the most extensively (Stewart and Hanson 1980; Delauney and Verma 1993; Rhodes et al. 1994). The accumulation of proline has been observed not only in plants but also in eubacteria, marine invertebrates, protozoa, and algae (McCue and Hanson 1990; Delauney and Verma 1993). A review of plant proline metabolism is included in this issue (Lehmann et al. 2010), and hence a more detailed review of proline accumulation under stress is excluded here. Nevertheless, the inability to enhance drought tolerance with proline overproduction, as well as associated toxic effects, has made the practical application of proline as an osmoprotective metabolite in plants difficult to achieve (Hare and Cress 1997; Nanjo et al. 1999; Hellmann et al. 2000; Srikrishnan et al. 2002; Nanjo et al. 2003; Deuschle et al. 2004).
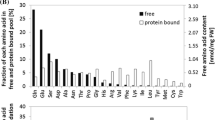
Although proline has been studied most frequently, it is certainly not the only plant amino acid that is overproduced in response to osmotic stress. For instance, proline contributed only 10% of the total free amino acids induced as a response to drought and plant age in flatpea (Lathyrus sylvestris L.), less than the valine, isoleucine, leucine, phenylalanine, and methionine increases (Shen et al. 1989). The relative increase of isoleucine (90-fold) and leucine (150-fold) in drought-stressed Arabidopsis foliage is greater than the 80-fold foliar proline increase (Nambara et al. 1998). Similar results were observed in Arabidopsis leaves under our growth conditions (Fig. 3a). BCAAs in drought-stressed Arabidopsis flowers and siliques also show high fold-increases (Fig. 3b, c). The absolute abundance of free BCAAs in these two tissues under drought stress is about half that of proline. The observed amino acid increases are not limited to Arabidopsis; the fold-change increases in BCAAs of drought-stressed tomato leaves are also higher than that of proline (Fig. 3d). Expression of BCAA aminotransferase, which functions in the last step of the biosynthesis of BCAAs, is induced in response to dehydration stress (Urano et al. 2009). Increases in valine and leucine concentrations upon drought stress could also result from the corresponding increase in the substrate quantities, as was observed in case of pyruvate, which was doubled after 4 days of drought stress (Good and Zaplachinski 1994).
Amino acid accumulation in drought-stressed plant tissues. Fold-change over respective fresh tissue due to dehydration is shown for selected amino acids. Mean ± SD of n = 4–6. Drought experiments and amino acid analyses were carried out as described in Joshi and Jander (2009)
It has been proposed that accumulation of free BCAAs may serve as a substrate for the synthesis of stress-induced proteins and that BCAAs may act as signaling molecules to regulate gene expression (Nambara et al. 1998). Application of BCAAs alters gene expression patterns in plants (De Veylder et al. 1997), yeast (Didion et al. 1996) and other biological systems (Yoshizawa 2004). A set of genes encoding proteins that are rich in BCAAs is induced in response to low temperature or osmotic stress in barley, wheat, and strawberry (Behl et al. 1991; Goddard et al. 1993). Because of their unsubstituted aliphatic side chains with branched alkyl groups, the BCAAs are the most hydrophobic among the standard 20 protein amino acids. BCAAs are often located in the core of the proteins and play a crucial role in determining the structures of globular proteins and interaction of the transmembrane domains of membrane proteins with phospholipid bilayers.
Threonine deaminase and methionine γ-lyase maintain the isoleucine needs of plants under stress
Threonine deaminase and methionine γ-lyase transcription, as well as isoleucine accumulation, are induced in response to osmotic stress (Nambara et al. 1998; Less and Galili 2008; Jander and Joshi 2009; Joshi and Jander 2009). Overlapping but dissimilar expression patterns of Arabidopsis methionine γ-lyase and threonine deaminase in the reproductive tissue (Rebeille et al. 2006) and during vegetative growth (Joshi and Jander 2009) (Supplemental Fig. 2), respectively, suggest different functions in fulfilling isoleucine needs during the course of plant development and in response to stress.
Although reduced threonine deaminase causes severe growth defects in Nicotiana species (Colau et al. 1987; Kang et al. 2006) and Arabidopsis (Joshi and Jander 2009), no such defects are observed in the absence of methionine γ-lyase (Goyer et al. 2007; Joshi and Jander 2009). This suggests that threonine deaminase plays a predominant role in isoleucine synthesis. However, under drought conditions and in reproductive tissue, methionine γ-lyase might play significant role as an alternate route to fulfill increased demand for isoleucine (Less and Galili 2008; Joshi and Jander 2009). Even though the synthesis of BCAAs occurs in all plant parts, based on the transcript abundance of related genes, enzyme activities and carbon flux Singh and Shaner (1995) concluded that this synthesis predominantly takes place in the young tissues in each plant part. Due to species-specific, physiological and environmental differences, it is difficult to monitor quantitative distribution of the tissue-specific synthesis or transport of the BCAAs during the growth of plants. Nevertheless, it is apparent that the reproductive tissues such as flowers, siliques and mature seeds in Arabidopsis accumulate significantly higher levels of BCAAs than rosette leaves (Supplemental Fig. 1), either as a result of tissue-specific biosynthesis or due to transport from vegetative tissue. Transcript abundance of methionine γ-lyase was found to be 5–10 times higher in stems and siliques than in leaves of Arabidopsis (Goyer et al. 2007). In tomato, it has been shown that threonine deaminase transcript accumulation is at least 500 times higher in flowers than in leaves or roots (Samach et al. 1991, 1995). Flowers of potato (Hildmann et al. 1992) and Nicotiana attenuata (Hermsmeier et al. 2001) have also been shown to express threonine deaminase constitutively, implying an enhanced requirement for isoleucine synthesis during reproduction.
Although both methionine and threonine serve as precursors for isoleucine biosynthesis in plants in response to osmotic stress, alternative plant metabolic pathways cannot be ruled out. Precursors for threonine and methionine-independent isoleucine biosynthesis in microorganisms include glutamate, 2-methylbutyrate, propionate, homolanthionine, and citramalate (Phillips et al. 1972; Kisumi et al. 1977; Monticello et al. 1984; Hochuli et al. 1999; Xu et al. 2004; Kromer et al. 2006; Risso et al. 2008; Wu et al. 2009). Metabolite profiling experiments have shown the presence of citramalate in Arabidopsis (Fiehn et al. 2000; Roessner-Tunali et al. 2003), and it has, therefore, been proposed that citramalate may also serve as a precursor for isoleucine biosynthesis in plants (de Kraker et al. 2007) (Fig. 1). However, there is as yet no direct evidence to support the hypothesis that this pathway functions in response to osmotic stress or under other physiological conditions.
Transcriptional regulation of amino acid synthesis during osmotic stress: a case study
Although extensive allosteric regulation of amino acid biosynthesis has been identified, very little is known about transcriptional regulation of this process. In the case of proline accumulation, a promoter region of proline dehydrogenase (ProDH) was shown to carry a 9-bp sequence (ACTCATCCT) that is a hypo-osmolarity- or proline-responsive element required for the efficient expression of ProDH in response to hypo-osmolarity (Satoh et al. 2002). Another cis element (ACTCAT), a typical binding site of basic leucine zipper (bZIP) transcription factors was identified for hypo-osmolarity-mediated activation of ProDH (Weltmeier et al. 2006).
Recently, a metabolic network analysis done by Urano et al. (2009) revealed that dehydration-increased amino acids contribute more significantly to the dehydration stress responses when they have a global correlation with one another. We extended such research using a bioinformatics approach for identifying regulatory programs of amino acid genes in response to different abiotic stresses. Three microarray gene expression datasets, drought, osmotic, and salt, were downloaded from the Nottingham Arabidopsis Stock Centre (http://arabidopsis.org.uk/) (Craigon et al. 2004) and used for expression correlation and regulatory motif analysis. All data were generated using the Affymetrix ATH1 genome array, which contains ~22,800 probe sets. The CEL files were processed and normalized at the probe level using the GC content-based Robust Multi-array Algorithm (GCRMA) (Wu et al. 2004), and the expression datasets were processed by taking the log ratio between the expression level of treatments and that of the corresponding normal tissues.
Correlation analysis between transcription factors and amino acid metabolism genes
A total of 2,620 transcription factors were obtained from the recent collection of four representative Arabidopsis transcription factor databases: RARTF (http://rarge.gsc.riken.jp/rartf/), AGRIS (http://arabidopsis.med.ohio-state.edu/), DATF (http://datf.cbi.pku.edu.cn/), and PlnTFDB (http://plntfdb.bio.uni-potsdam.de/v3.0/) (Mitsuda and Ohme-Takagi 2009). In order to observe the expression relationships of transcription factors and amino acid genes, we computed the Pearson’s correlation coefficient between the expression profiles of all transcription factors and those of an amino acid biosynthetic gene set. Then, the correlation coefficient matrix of only highly correlated transcription factors and amino acids was clustered using an agglomerative hierarchical clustering algorithm in R. The genes involved in threonine, lysine and methionine metabolism were grouped separately from those involved in BCAA synthesis (Fig. 4). Each group of identified amino acid metabolism-related genes shows either positive or negative correlation with mostly independent stress (drought, osmotic, salt)-induced transcription factors. Positive correlation suggests either direct activation of the amino acid metabolism related genes by the specific transcription factor or through recruitment of other transcription factors. For example, methionine γ-lyase shows strong positive correlation with 12 and 9 independent transcription factors under drought and osmotic stresses, respectively. Interestingly, the same set of transcription factors is co-regulated with S-adenosylmethionine synthase I (At1g02500). Although both of these competitive enzymes are known to be induced under abiotic stress, they compete for a common methionine pool. It is possible that, with the increased need of isoleucine due to stress, S-adenosylmethionine (SAM) produced by this enzyme can activate threonine synthase to increase substrate availability for threonine deaminase. Similarly, a set of four genes involved in the BCAA metabolism, At1g10070 (branched-chain amino acid transferase 2, BCAT2), At1g10060 (branched-chain amino acid transferase 1, BCAT1), At4g11640 (serine racemase, catalyzing dehydration of serine to pyruvate), and At1g18500 (isopropylmalate synthase IPMS1, involved in leucine biosynthesis) are co-regulated with same set of transcription factors under salt stress.
Heatmaps of amino acid-related genes and highly correlated transcription factors identified based on gene expression profiles. A subset of amino acid genes in two groups (lysine, methionine and threonine biosynthesis; isoleucine, valine and leucine biosynthesis; shown on horizontal axis) is highly correlated with specific transcription factors (shown on vertical axis). In the figure, transcription factors and amino acid metabolism-related genes are clustered hierarchically, as indicated by the dendrograms at the top and left of each heatmap respectively
Identification of regulatory motifs associated with amino acid metabolism genes
Here, we present an approach for identifying regulatory motifs associated with amino acid gene expression based on the gene expression coherence and promoter analysis of these genes (Fig. 5). First, expression profiles of these genes were clustered by the hierarchical clustering in order to group co-regulated amino acid genes. Then, the over-represented motifs were screened in the upstream sequences of genes in each cluster using the Athena promoter database (O’Connor et al. 2005). We identified conserved motifs from the amino acid metabolism-related genes that were significantly over-represented in drought and osmotic stress treatments (P value < 0.001). The genes containing the motif-binding sites are also significantly correlated with the corresponding transcription factors in the gene expression profile of a specific biological process. Microarray analysis confirmed that, among 121 rehydration-inducible genes, 48% harbor the ACTCAT motif in their promoters (Oono et al. 2003). ACTCAT is a cis-acting element involved in rehydration-, proline- and low osmolarity-inducible gene expression, and genes containing the ACTCAT cis-acting element in their promoter regions were regarded as responding to low osmolarity as well as to proline treatment (Satoh et al. 2002). The ACTCAT sequence is similar to the yeast GCN4 motif [ATGA(C/G)TCAT], which is recognized by ATB2 subgroup bZIP transcription factors (Satoh et al. 2004). We carried out similar analysis of promoters of other genes involved in amino acid metabolism (Supplemental Table 1). Many of the amino acid transporters carry this conserved motif, suggesting their need and active involvement during the osmotic stress recovery processes. Furthermore, the presence of this conserved motif in both Arabidopsis threonine aldolases (At1g08630 and At3g04520) indicates reduced need of isoleucine synthesis from threonine upon rehydration.
Identification of regulatory motifs associated with the amino acid gene expression. Significant regulatory motifs were identified in upstream sequences of co-regulated amino acid genes. Expression of the corresponding transcription factors is highly correlated with their putative targets under a specific biological process. [Genes in cluster 2 (drought) and 4 (osmotic) are listed in Supplemental Table 2]
From the motif analysis of the osmotic and drought stress experiments, we identified two clusters having conserved motifs recognized by two independent transcription factors. Selected genes from both the clusters are shown in Fig. 5 and the remaining genes in each cluster are listed separately in Supplemental Table 2. The osmotic cluster C4 (Fig. 5) has a conserved motif (TACACTTTTGG), which is a binding site for the Dof zinc finger protein OBP4 (Singh et al. 2002). Two of the genes carrying this motif are involved in methionine biosynthesis and catabolism, and are thus most likely required to enhance isoleucine accumulation. Serine O-acetyltransferase (AT1G55920), which is involved in sulfur assimilation and methionine biosynthesis, and methionine γ-lyase (At1g64660) are known to be highly up-regulated upon drought stress (Less and Galili 2008). The other three genes carrying this motif are known to be up-regulated upon salt-stress (tyrosine transaminase, At5g53970, Roosens et al. 1998), induced under stress (proline dehydrogenase, At5g38710, Hollander-Czytko et al. 2005), or are involved in serine and homoserine biosynthetic process (AT1G72190, which is a part of phosphorylation pathway that is up-regulated upon high salinity, flood and cold, Ho and Saito 2001). Motif analysis of data from drought stress experiments identified a cluster of genes (drought cluster C2 in Fig. 5; Supplemental Table 2) carrying a conserved motif which is a binding site for MADS box transcription factor AGL15 protein (Heck et al. 1995). Among the genes that carry this motif, At5g14780 (formate dehydrogenase) and At1G64660 (methionine γ-lyase) were highly correlated with AGL15 expression. Two other genes from this cluster with APS kinase activities (At2g14750 and At4g39940) carry a binding site for the abiotic stress-induced TEIL (AP2/EREBP family) transcription factor, but were negatively correlated with its expression. These two genes are involved in sulfur activation for the production of sulfonated compounds such as glucosinolates. The negative correlation with drought stressed-induced TEIL transcription factors most likely implies that they down-regulate genes involved in sulfonation reactions, thereby making the substrate adenosine 5′-phosphosulfate available for sulfate reduction (assimilation) by APS reductase (EC 1.8.4.9) for the synthesis of methionine.
Conclusion and future prospects
New evidence suggests that threonine and methionine together regulate isoleucine homeostasis under certain physiological and growth conditions. Despite complex regulation of the enzymes responsible for their synthesis, these three amino acids are maintained in equilibrium. In view of crop improvement, this makes it complicated to manipulate the synthesis of aspartate-derived amino acids, as many of the enzymes regulating their metabolism are in shared pathways or compete for common substrates. However, knocking out the catabolic pathway enzymes may offer opportunities to enhance particular substrates, e.g., threonine by knocking out threonine aldolases (Joshi et al. 2006) or methionine by knocking out methionine γ-lyase (Joshi and Jander 2009) without affecting plant fitness.
In spite of the large relative increase in BCAA accumulation in response to abiotic stress, not much research has been directed toward their role as protective osmolytes. Isoleucine is produced from threonine and methionine, with threonine deaminase and methionine γ-lyase as the committing enzymes, respectively. Expression patterns and tissue specificities of these two enzymes are non-overlapping, implying that they have different functions based on plant isoleucine requirements. Methionine, threonine, and isoleucine biosynthesis is regulated in a network that is linked to physiological and growth requirements. We have used abiotic stress-induced production of BCAAs as a method for identifying transcription factors that may be involved in regulating drought-induced amino acid biosynthesis. Future experiments can be designed with selected transcription factors that are strongly co-regulated with BCAA metabolism. It will be possible to analyze stress-induced amino acid content in the mutant Arabidopsis lines where expression of selected transcription factors is knocked out or reduced. If any of the identified transcription factor mutations results in altered BCAA biosynthesis, subsequent over-expression of particular transcription factors in plants might elevate BCAA biosynthesis and enhance abiotic stress tolerance in plants. The same genes could also be used to increase the nutritive value of crop plants by up-regulating amino acid biosynthesis in a more targeted manner. For instance, expression of particular transcription factors from the patatin promoter in potatoes (Jefferson et al. 1990; Liu et al. 1990) or the seed-specific phaseolin promoter (Karchi et al. 1993) in grain crops might allow up-regulation of BCAA accumulation in a tissue-specific manner.
Some studies suggest that additional as yet unknown regulatory mechanisms can contribute to the control of amino acid accumulation and more general plant responses to osmotic stress. For instance, Arabidopsis protein kinases KIN10 (At3g01090) and KIN11 (At3g29160) have been shown to control convergent reprogramming of transcription in response to stress conditions (Baena-Gonzalez et al. 2007). Transgenic KIN10 over-expression that enhanced starvation tolerance also caused almost sixfold increase in methionine γ-lyase expression (Baena-Gonzalez and Sheen 2008). It will be interesting to determine whether this regulation of methionine γ-lyase is a direct effect of protein kinase or involves less direct regulation of gene expression. Future research will determine how the established mechanisms of allosteric regulation, the transcription factors described in this article, and other, as yet unidentified regulatory mechanisms are integrated to control the osmotic stress-induced accumulation of amino acids in plants. A better understanding of this process will provide new opportunities for regulating plant amino acid biosynthesis pathways in a targeted manner.
References
Ajjawi I, Lu Y, Savage LJ, Bell SM, Last RL (2009) Large scale reverse genetics in Arabidopsis: case studies from the Chloroplast 2010 Project. Plant Physiol. Epub ahead of print. doi:10.1104/pp.109.148494
Alia, Mohanty P, Matysik J (2001) Effect of proline on the production of singlet oxygen. Amino Acids 21:195–200
Amir R (2010) Current understanding of the factors regulating methionine content in vegetative tissues of higher plants. Amino Acids (in press)
Amir R, Hacham Y, Galili G (2002) Cystathionine gamma-synthase and threonine synthase operate in concert to regulate carbon flow towards methionine in plants. Trends Plant Sci 7:153–156
Anderson MD, Che P, Song JP, Nikolau BJ, Wurtele ES (1998) 3-Methylcrotonyl coenzyme A carboxylase is a component of the mitochondrial leucine catabolic pathway in plants. Plant Physiol 118:1127–1138
Baena-Gonzalez E, Sheen J (2008) Convergent energy and stress signaling. Trends Plant Sci 13:474–482
Baena-Gonzalez E, Rolland F, Thevelein JM, Sheen J (2007) A central integrator of transcription networks in plant stress and energy signalling. Nature 448:938–942
Bartlem D, Lambein I, Okamoto T, Itaya A, Uda Y, Kijima F, Tamaki Y, Nambara E, Naito S (2000) Mutation in the threonine synthase gene results in an over-accumulation of soluble methionine in Arabidopsis. Plant Physiol 123:101–110
Baum HJ, Madison JT, Thompson JF (1983) Feedback inhibition of homoserine kinase from radish leaves. Phytochemistry 22:2409–2412
Behl RK, Moawad AM, Achtnich W (1991) Amino acid and protein profile changes in a spring wheat mutant under prolonged heat stress. Ann Biol 7:63–68
Binder S, Knill T, Schuster J (2007) Branched-chain amino acid metabolism in higher plants. Physiol Plant 129:68–78
Boerjan W, Bauw G, Vanmontagu M, Inze D (1994) Distinct phenotypes generated by overexpression and suppression of S-adenosyl-l-methionine synthetase reveal developmental patterns of gene silencing in tobacco. Plant Cell 6:1401–1414
Bourguignon J, Rebéillé F, Douce R (1998) Serine and glycine metabolism in higher plants. In: Singh BK (ed) Plant amino acids. Biochemistry & biotechnology. Marcel Dekker Inc., New York, pp 111–146
Burg MB, Ferraris JD (2008) Intracellular organic osmolytes: function and regulation. J Biol Chem 283:7309–7313
Campalans A, Messeguer R, Goday A, Pages M (1999) Plant responses to drought, from ABA signal transduction events to the action of the induced proteins. Plant Physiol Biochem 37:327–340
Ceccarelli S, Grando S (1996) Drought as a challenge for the plant breeder. Plant Growth Regul 20:149–155
Chang AK, Duggleby RG (1998) Herbicide-resistant forms of Arabidopsis thaliana acetohydroxyacid synthase: characterization of the catalytic properties and sensitivity to inhibitors of four defined mutants. Biochem J 333:765–777
Chipman D, Barak ZA, Schloss JV (1998) Biosynthesis of 2-aceto-2-hydroxy acids: acetolactate synthases and acetohydroxyacid synthases. Biochim Biophys Acta 1385:401–419
Chuang DT, Shih VE (1995) Disorders of branched-chain amino acid and keto acid metabolism. In: Scriver CR, Beaudet AL, Sly WS, Valle D (eds) The metabolic basis of inherited disease, vol 1, 7th edn. McGraw-Hill, New York, pp 1239–1277
Colau D, Negrutiu I, Van Montagu M, Hernalsteens JP (1987) Complementation of a threonine dehydratase-deficient Nicotiana plumbaginifolia mutant after Agrobacterium tumefaciens-mediated transfer of the Saccharomyces cerevisiae ILV1 gene. Mol Cell Biol 7:2552–2557
Cornah JE, Germain V, Ward JL, Beale MH, Smith SM (2004) Lipid utilization, gluconeogenesis, and seedling growth in Arabidopsis mutants lacking the glyoxylate cycle enzyme malate synthase. J Biol Chem 279:42916–42923
Craigon DJ, James N, Okyere J, Higgins J, Jotham J, May S (2004) NASCArrays: a repository for microarray data generated by NASC’s transcriptomics service. Nucl Acids Res 32:D575–D577
Curien G, Dumas R, Ravanel S, Douce R (1996) Characterization of an Arabidopsis thaliana cDNA encoding an S-adenosylmethionine-sensitive threonine synthase. Threonine synthase from higher plants. FEBS Lett 390:85–90
Curien G, Ravanel S, Dumas R (2003) A kinetic model of the branch-point between the methionine and threonine biosynthesis pathways in Arabidopsis thaliana. Eur J Biochem 270:4615–4627
Curien G, Bastlen O, Robert-Genthon M, Cornish-Bowden A, Cardenas ML, Dumas R (2009) Understanding the regulation of aspartate metabolism using a model based on measured kinetic parameters. Mol Sys Biol 5:271. doi:10.1038/msb.2009.29
Dancs G, Kondrak M, Banfalvi Z (2008) The effects of enhanced methionine synthesis on amino acid and anthocyanin content of potato tubers. BMC Plant Biol 8:65. doi:10.1186/1471-2229-8-65
de Kraker JW, Luck K, Textor S, Tokuhisa JG, Gershenzon J (2007) Two Arabidopsis genes (IPMS1 and IPMS2) encode isopropylmalate synthase, the branchpoint step in the biosynthesis of leucine. Plant Physiol 143:970–986
De Veylder L, Segers G, Glab N, Van Montagu M, Inze D (1997) Identification of proteins interacting with the Arabidopsis Cdc2aAt protein. J Exp Bot 48:2113–2114
Delauney AJ, Verma DPS (1993) Proline biosynthesis and osmoregulation in plants. Plant J 4:215–223
Deuschle K, Funck D, Forlani G, Stransky H, Biehl A, Leister D, van der Graaff E, Kunzee R, Frommer WB (2004) The role of delta(1)-pyrroline-5-carboxylate dehydrogenase in proline degradation. Plant Cell 16:3413–3425
Dias B, Weimer B (1998) Conversion of methionine to thiols by Lactococci, Lactobacilli, and Brevibacteria. App Environ Microbiol 64:3320–3326
Didion T, Grauslund M, KiellandBrandt MC, Andersen HA (1996) Amino acids induce expression of BAP2, a branched-chain amino acid permease gene in Saccharomyces cerevisiae. J Bact 178:2025–2029
Diebold R, Schuster J, Daschner K, Binder S (2002) The branched-chain amino acid transaminase gene family in Arabidopsis encodes plastid and mitochondrial proteins. Plant Physiol 129:540–550
Duggleby RG, Pang SS (2000) Acetohydroxyacid synthase. J Biochem Mol Biol 33:1–36
Dumas R, Job D, Ortholand JY, Emeric G, Greiner A, Douce R (1992) Isolation and kinetic-properties of acetohydroxy acid isomeroreductase from spinach (Spinacia oleracea) chloroplasts overexpressed in Escherichia coli. Biochem J 288:865–874
Durner J, Boger P (1988) Acetolactate synthase from barley (Hordeum vulgare L.)—purification and partial characterization. Zeitschrift Fur Naturforschung 43:850–856
Durner J, Boger P (1990) Oligomeric forms of plant acetolactate synthase depend on flavin adenine-dinucleotide. Plant Physiol 93:1027–1031
Eden A, Benvenisty N (1998) Characterization of a branched-chain amino-acid aminotransferase from Schizosaccharomyces pombe. Yeast 14:189–194
Faleev NG, Troitskaya MV, Paskonova EA, Saporovskaya MB, Belikov VM (1996) l-Methionine-gamma-lyase in Citrobacter intermedius cells: stereochemical requirements with respect to the thiol structure. Enzyme Microb Technol 19:590–593
Fiehn O, Kopka J, Dormann P, Altmann T, Trethewey RN, Willmitzer L (2000) Metabolite profiling for plant functional genomics. Nat Biotechnol 18:1157–1161
Galili G (1995) Regulation of lysine and threonine synthesis. Plant Cell 7:899–906
Gao F, Wang CZ, Wei CH, Li Y (2009) A branched-chain aminotransferase may regulate hormone levels by affecting KNOX genes in plants. Planta 230:611–623
Garcia EL, Mourad GS (2004) A site-directed mutagenesis interrogation of the carboxy-terminal end of Arabidopsis thaliana threonine dehydratase/deaminase reveals a synergistic interaction between two effector-binding sites and contributes to the development of a novel selectable marker. Plant Mol Biol 55:121–134
Gerbling H, Gerhardt B (1989) Peroxisomal degradation of branched-chain 2-oxo acids. Plant Physiol 91:1387–1392
Goddard NJ, Dunn MA, Zhang L, White AJ, Jack PL, Hughes MA (1993) Molecular analysis and spatial expression pattern of a low-temperature-specific barley gene, BLT101. Plant Mol Biol 23:871–879
Gonda I, Bar E, Portnoy V, Lev S, Burger J, Schaffer AA, Tadmor Y, Gepstein S, Giovannoni JJ, Katzir N, Lewinsohn E (2010) Branched-chain and aromatic amino acid catabolism into aroma volatiles in Cucumis melo L. fruit. J Exp Bot. Epub ahead of print
Good AG, Zaplachinski ST (1994) The effects of drought stress on free amino-acid accumulation and protein-synthesis in Brassica napus. Physiol Plant 90:9–14
Goyer A, Collakova E, Shachar-Hill Y, Hanson AD (2007) Functional characterization of a methionine gamma-lyase in Arabidopsis and its implication in an alternative to the reverse trans-sulfuration pathway. Plant Cell Phys 48:232–242
Graham IA, Eastmond PJ (2002) Pathways of straight and branched chain fatty acid catabolism in higher plants. Progr Lipid Res 41:156–181
Gu L, Daniel Jones A, Last RL (2009) Broad connections in the Arabidopsis seed metabolic network revealed by metabolite profiling of an amino acid catabolism mutant. Plant J. Epub ahead of print
Hacham Y, Matityahu I, Schuster G, Amir R (2008) Overexpression of mutated forms of aspartate kinase and cystathionine γ-synthase in tobacco leaves resulted in the high accumulation of methionine and threonine. Plant J 54(2):260–271
Hare PD, Cress WA (1997) Metabolic implications of stress-induced proline accumulation in plants. Plant Growth Regul 21:79–102
Heap I (2009) International survey of herbicide resistant weeds. www.weedscience.com. Retrieved 12 Dec 2009 (online)
Heck GR, Perry SE, Nichols KW, Fernandez DE (1995) AGL15, a MADS domain protein expressed in developing embryos. Plant Cell 7:1271–1282
Hellmann H, Funck D, Rentsch D, Frommer WB (2000) Hypersensitivity of an Arabidopsis sugar signaling mutant toward exogenous proline application. Plant Physiol 123:779–789
Hermsmeier D, Schittko U, Baldwin IT (2001) Molecular interactions between the specialist herbivore Manduca sexta (Lepidoptera, Sphingidae) and its natural host Nicotiana attenuata. I. Large-scale changes in the accumulation of growth- and defense-related plant mRNAs. Plant Physiol 125:683–700
Hildmann T, Ebneth M, Pena-Cortes H, Sanchez-Serrano JJ, Willmitzer L, Prat S (1992) General roles of abscisic and jasmonic acids in gene activation as a result of mechanical wounding. Plant Cell 4:1157–1170
Ho CL, Saito K (2001) Molecular biology of the plastidic phosphorylated serine biosynthetic pathway in Arabidopsis thaliana. Amino Acids 20:243–259
Hochuli M, Patzelt H, Oesterhelt D, Wuthrich K, Szyperski T (1999) Amino acid biosynthesis in the halophilic archaeon Haloarcula hispanica. J Bact 181:3226–3237
Hollander-Czytko H, Grabowski J, Sandorf I, Weckermann K, Weiler EW (2005) Tocopherol content and activities of tyrosine aminotransferase and cystine lyase in Arabidopsis under stress conditions. J Plant Phys 162:767–770
Hori H, Takabayashi K, Orvis L, Carson DA, Nobori T (1996) Gene cloning and characterization of Pseudomonas putida l-methionine-alpha-deamino-gamma-mercaptomethane-lyase. Cancer Res 56:2116–2122
Imsande J (2001) Selection of soybean mutants with increased concentrations of seed methionine and cysteine. Crop Sci 41:510–515
Inoue H, Inagaki K, Sugimoto M, Esaki N, Soda K, Tanaka H (1995) Structural-analysis of the l-methionine gamma-lyase gene from Pseudomonas putida. J Biochem 117:1120–1125
Jander G, Joshi V (2009) Aspartate-derived amino acid biosynthesis in Arabidopsis thaliana. In: Last RL (ed) The Arabidopsis book. The American Society of Plant Biologists, Rockville, MD, pp 1–15. doi:10.1199/tab.0121
Jander G, Baerson SR, Hudak JA, Gonzalez KA, Gruys KJ, Last RL (2003) Ethylmethanesulfonate saturation mutagenesis in Arabidopsis to determine frequency of herbicide resistance. Plant Physiol 131:139–146
Jander G, Norris SR, Joshi V, Fraga M, Rugg A, Yu S, Li L, Last RL (2004) Application of a high-throughput HPLC-MS/MS assay to Arabidopsis mutant screening; evidence that threonine aldolase plays a role in seed nutritional quality. Plant J 39:465–475
Jefferson R, Goldsbrough A, Bevan M (1990) Transcriptional regulation of a patatin-1 gene in potato. Plant Mol Biol 14:995–1006
Joshi V, Jander G (2009) Arabidopsis methionine gamma-lyase is regulated according to isoleucine biosynthesis needs but plays a subordinate role to threonine deaminase. Plant Physiol 151:367–378
Joshi V, Laubengayer KM, Schauer N, Fernie AR, Jander G (2006) Two Arabidopsis threonine aldolases are nonredundant and compete with threonine deaminase for a common substrate pool. Plant Cell 18:3564–3575
Kang JH, Wang L, Giri A, Baldwin IT (2006) Silencing threonine deaminase and JAR4 in Nicotiana attenuata impairs jasmonic acid-isoleucine-mediated defenses against Manduca sexta. Plant Cell 18:3303–3320
Karchi H, Shaul O, Galili G (1993) Seed-specific expression of a bacterial desensitized aspartate kinase increases the production of seed threonine and methionine in transgenic tobacco. Plant J 3:721–727
Kisumi M, Komatsubara S, Chibata I (1977) Pathway for isoleucine formation from pyruvate by leucine biosynthetic-enzymes in leucine-accumulating isoleucine revertants of Serratia marcescens. J Biochem 82:95–103
Kromer JO, Heinzle E, Schroder H, Wittmann C (2006) Accumulation of homolanthionine and activation of a novel pathway for isoleucine biosynthesis in Corynebacterium glutamicum McbR deletion strains. J Bact 188:609–618
Laber B, Clausen T, Huber R, Messerschmidt A, Egner U, Müller-Fahrnow A, Pohlenz H-D (1996) Cloning, purification, and crystallization of Escherichia coli cystathionine [beta]-lyase. FEBS Lett 379:94–96
Laber B, Maurer W, Hanke C, Grafe S, Ehlert S, Messerschmidt A, Clausen T (1999) Characterization of recombinant Arabidopsis thaliana threonine synthase. Eur J Biochem 263:212–221
Lea PJ, Ireland RJ (1999) Nitrogen metabolism in higher plants. In: Singh BK (ed) Plant amino acids: biochemistry and biotechnology. Marcel Dekker, New York, pp 1–47
Lee YT, Duggleby RG (2001) Identification of the regulatory subunit of Arabidopsis thaliana acetohydroxyacid synthase and reconstitution with its catalytic subunit. Biochemistry 40:6836–6844
Lee M, Leustek T (1999) Identification of the gene encoding homoserine kinase from Arabidopsis thaliana and characterization of the recombinant enzyme derived from the gene. Arch Biochem Biophys 372:135–142
Lehmann S, Funck F, Szabados L, Rentsch D (2010) Proline metabolism and transport in plant development. Amino Acids (in press)
Less H, Galili G (2008) Principal transcriptional programs regulating plant amino acid metabolism in response to abiotic stresses. Plant Physiol 147:316–330
Leung EWW, Guddat LW (2009) Conformational changes in a plant ketol-acid reductoisomerase upon Mg2+ and NADPH binding as revealed by two crystal structures. J Mol Biol 389:167–182
Liu XJ, Prat S, Willmitzer L, Frommer WB (1990) Cis regulatory elements directing tuber-specific and sucrose-inducible expression of a chimeric class I patatin promoter/GUS-gene fusion. Mol Gen Genet 223:401–406
Lu Y, Savage LJ, Ajjawi I, Imre KM, Yoder DW, Benning C, DellaPenna D, Ohlrogge JB, Osteryoung KW, Weber AP, Wilkerson CG, Last RL (2008) New connections across pathways and cellular processes: industrialized mutant screening reveals novel associations between diverse phenotypes in Arabidopsis. Plant Physiol 146:1482–1500
Manukhov IV, Demidkina TV, Zavilgelsky GB (2005) Evolution of a gene encoding l-methionine gamma-lyase in Enterobacteriaceae family genomes. FEBS J 272:478
Mas-Droux C, Biou V, Dumas R (2006) Allosteric threonine synthase—reorganization of the pyridoxal phosphate site upon asymmetric activation through S-adenosylmethionine binding to a novel site. J Biol Chem 281:5188–5196
McCue KF, Hanson AD (1990) Drought and salt tolerance—towards understanding and application. Trends Bact 8:358–362
McKie AE, Edlind T, Walker J, Mottram JC, Coombs GH (1998) The primitive protozoon Trichomonas vaginalis contains two methionine gamma-lyase genes that encode members of the gamma-family of pyridoxal 5′-phosphate-dependent enzymes. J Biol Chem 273:5549–5556
Mertz ET, Nelson OE, Bates LS (1964) Mutant gene that changes protein composition and increases lysine content of maize endosperm. Science 145:279
Mitsuda N, Ohme-Takagi M (2009) Functional analysis of transcription factors in Arabidopsis. Plant Cell Phys 50:1232–1248
Monticello DJ, Hadioetomo RS, Costilow RN (1984) Isoleucine synthesis by clostridium-sporogenes from propionate or alpha-methylbutyrate. J Gen Microbiol 130:309–318
Mourad G, King J (1995) l-O-Methylthreonine-resistant mutant of Arabidopsis defective in isoleucine feedback-regulation. Plant Physiol 107:43–52
Munck L, Karlsson KE, Hag-Berg A (1970) Gene for improved nutritional value in barley seed protein. Science 168:985–987
Nambara E, Kawaide H, Kamiya Y, Naito S (1998) Characterization of an Arabidopsis thaliana mutant that has a defect in ABA accumulation: ABA-dependent and ABA-independent accumulation of free amino acids during dehydration. Plant Cell Phys 39:853–858
Nanjo T, Kobayashi M, Yoshiba Y, Kakubari Y, Yamaguchi-Shinozaki K, Shinozaki K (1999) Antisense suppression of proline degradation improves tolerance to freezing and salinity in Arabidopsis thaliana. FEBS Lett 461:205–210
Nanjo T, Fujita M, Seki M, Kato T, Tabata S, Shinozaki K (2003) Toxicity of free proline revealed in an Arabidopsis T-DNA-tagged mutant deficient in proline dehydrogenase. Plant Cell Phys 44:541–548
Newell-McGloughlin M (2008) Nutritionally improved agricultural crops. Plant Physiol 147:939–953
Nuccio ML, Rhodes D, McNeil SD, Hanson AD (1999) Metabolic engineering of plants for osmotic stress resistance. Curr Opin Plant Biol 2:128–134
O’Connor TR, Dyreson C, Wyrick JJ (2005) Athena: a resource for rapid visualization and systematic analysis of Arabidopsis promoter sequences. Bioinformatics 21:4411–4413
Oono Y, Seki M, Nanjo T, Narusaka M, Fujita M, Satoh R, Satou M, Sakurai T, Isida J, Akiyama K, Maruyama K, Sato S, Yamaguchi-Shinozaki K, Shinozaki K (2003) Monitoring expression profiles of Arabidopsis gene expression during rehydration process after dehydration using ca. 7000 full-length cDNA microarray. Plant Cell Phys 44:S46–S46
Ott KH, Kwagh JG, Stockton GW, Sidorov V, Kakefuda G (1996) Rational molecular design and genetic engineering of herbicide resistant crops by structure modeling and site-directed mutagenesis of acetohydroxyacid synthase. J Mol Biol 263:359–368
Pang SS, Guddat LW, Duggleby RG (2003) Molecular basis of sulfonylurea herbicide inhibition of acetohydroxyacid synthase. J Biol Chem 278:7639–7644
Parida AK, Das AB (2005) Salt tolerance and salinity effects on plants: a review. Ecotox Environ Safety 60:324–349
Phillips AT, Nuss JI, Moosic J, Foshay C (1972) Alternate pathway for isoleucine biosynthesis in Escherichia coli. J Bact 109:714–719
Prell J, White JP, Bourdes A, Bunnewell S, Bongaerts RJ, Poole PS (2009) Legumes regulate Rhizobium bacteroid development and persistence by the supply of branched-chain amino acids. Proc Natl Acad Sci USA 106:12477–12482
Rebeille F, Jabrin S, Bligny R, Loizeau K, Gambonnet B, Van Wilder V, Douce R, Ravanel S (2006) Methionine catabolism in Arabidopsis cells is initiated by a gamma-cleavage process and leads to S-methylcysteine and isoleucine syntheses. Proc Natl Acad Sci USA 103:15687–15692
Rhodes D, Hanson AD (1993) Quaternary ammonium and tertiary sulfonium compounds in higher-plants. Ann Rev Plant Phys Plant Mol Biol 44:357–384
Rhodes D, Samaras Y, D’Urzo MP, Bressan RA, Garcia-Rios MG, Csonka LN (1994) Proline accumulation during drought and salinity. J Exp Bot 45:46
Risso C, Van Dien SJ, Orloff A, Lovley DR, Coppi MV (2008) Elucidation of an alternate isoleucine biosynthesis pathway in Geobacter sulfurreducens. J Bact 190:2266–2274
Roessner-Tunali U, Urbanczyk-Wochniak E, Czechowski T, Kolbe A, Willmitzer L, Fernie AR (2003) De novo amino acid biosynthesis in potato tubers is regulated by sucrose levels. Plant Physiol 133:683–692
Roosens N, Thu TT, Iskandar HM, Jacobs M (1998) Isolation of the ornithine-delta-aminotransferase cDNA and effect of salt stress on its expression in Arabidopsis thaliana. Plant Physiol 117:263–271
Saari L, Coterman J, Thill D (1994) Resistance to acetolactate synthase inhibiting herbicides. In: Powles S, Holtum J (eds) Herbicide resistance in plants: biology and biochemistry. Lewis Publishers, Boca Raton, pp 81–139
Saini HS, Attieh JM, Hanson AD (1995) Biosynthesis of halomethanes and methanethiol by higher-plants via a novel methyltransferase reaction. Plant Cell Environ 18:1027–1033
Samach A, Hareven D, Gutfinger T, Ken-Dror S, Lifschitz E (1991) Biosynthetic threonine deaminase gene of tomato: isolation, structure, and upregulation in floral organs. Proc Natl Acad Sci USA 88:2678–2682
Samach A, Broday L, Hareven D, Lifschitz E (1995) Expression of an amino acid biosynthesis gene in tomato flowers: developmental upregulation and MeJa response are parenchyma-specific and mutually compatible. Plant J 8:391–406
Satoh R, Nakashima K, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2002) ACTCAT, a novel cis-acting element for proline- and hypoosmolarity-responsive expression of the ProDH gene encoding proline dehydrogenase in Arabidopsis. Plant Physiol 130:709–719
Satoh R, Fujita Y, Nakashima K, Shinozaki K, Yamaguchi-Shinozaki KY (2004) A novel subgroup of bZIP proteins functions as transcriptional activators in hypoosmolarity-responsive expression of the ProDH gene in Arabidopsis. Plant Cell Phys 45:309–317
Schauer N, Fernie AR (2006) Plant metabolomics: towards biological function and mechanism. Trends Plant Sci 11:508–516
Schmidt A, Rennenberg H, Wilson LG, Filner P (1985) Formation of methanethiol from methionine by leaf tissue. Phytochemistry 24:1181–1185
Schulze-Siebert D, Heineke D, Schultz G (1984) Biosynthesis of pyruvate derived amino-acids during photosynthetic carbon metabolism in spinach chloroplasts. Plant Physiol 75(Suppl 1):7
Schuster J, Binder S (2005) The mitochondrial branched-chain aminotransferase (AtBCAT-1) is capable to initiate degradation of leucine, isoleucine and valine in almost all tissues in Arabidopsis thaliana. Plant Mol Biol 57:241–254
Shen L, Foster JG, Orcutt DM (1989) Composition and distribution of free amino-acids in flatpea (Lathyrus sylvestris L.) as influenced by water deficit and plant-age. J Exp Bot 40:71–79
Singh BK (1999) Biosynthesis of valine, leucine and isoleucine. In: Singh BK (ed) Plant amino acids: biochemistry and biotechnology. Marcel Dekker, New York, pp 227–247
Singh R, Axtell JD (1973) A mutant gene (hl) in sorghum which improves lysine concentration in the grain. Crop Sci 13:535–539
Singh BK, Shaner DL (1995) Biosynthesis of branched chain amino acids: from test tube to field. Plant Cell 7:935–944
Singh BK, Stidham MA, Shaner DL (1988) Assay of acetohydroxyacid synthase. Anal Biochem 171:173–179
Singh KB, Foley RC, Onate-Sanchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5:430–436
Southan MD, Copeland L (1996) Physical and kinetic properties of acetohydroxyacid synthase from wheat leaves. Physiol Plant 98:824–832
Srikrishnan M, Cotte Bvd, Montagu Mv, Verbruggen N (2002) Altered levels of proline dehydrogenase cause hypersensitivity to proline and its analogs in Arabidopsis. Plant Physiol 128:73–83
Stewart CR, Hanson AD (1980) Proline accumulation as a metabolic response to water stress. In: Kramer PJ, Turner NC (eds) Adaptation of plants to water and high temperature stress. Wiley, New York, pp 173–189
Szamosi IT, Shaner DL, Singh BK (1994) Inhibition of threonine dehydratase is herbicidal. Plant Physiol 106:1257–1260
Tan SY, Evans RR, Dahmer ML, Singh BK, Shaner DL (2005) Imidazolinone-tolerant crops: history, current status and future. Pest Manag Sci 61:246–257
Taylor NL, Heazlewood JL, Day DA, Millar AH (2004) Lipoic acid-dependent oxidative catabolism of alpha-keto acids in mitochondria provides evidence for branched-chain amino acid catabolism in Arabidopsis. Plant Physiol 134:838–848
Thoen A, Rognes S, Aarnes H (1978) Biosynthesis of threonine from homoserine in pea seedling: homoserine kinase. Plant Sci Lett 13:103–112
Thomazeau K, Curien G, Dumas R, Biou V (2001) Crystal structure of threonine synthase from Arabidopsis thaliana. Protein Sci 10:638–648
Tokoro M, Asai T, Kobayashi S, Takeuchi T, Nozaki T (2003) Identification and characterization of two isoenzymes of methionine gamma-lyase from Entamoeba histolytica—a key enzyme of sulfur-amino acid degradation in an anaerobic parasitic protist that lacks forward and reverse trans-sulfuration pathways. J Biol Chem 278:42717–42727
Tranel PJ, Wright TR (2002) Resistance of weeds to ALS-inhibiting herbicides: what have we learned? Weed Sci 50:700–712
Trenkamp S, Eckes P, Busch M, Fernie AR (2009) Temporally resolved GC–MS-based metabolic profiling of herbicide treated plants treated reveals that changes in polar primary metabolites alone can distinguish herbicides of differing mode of action. Metabolomics 5:277–291
Urano K, Maruyama K, Ogata Y, Morishita Y, Takeda M, Sakurai N, Suzuki H, Saito K, Shibata D, Kobayashi M, Yamaguchi-Shinozaki K, Shinozaki K (2009) Characterization of the ABA-regulated global responses to dehydration in Arabidopsis by metabolomics. Plant J 57:1065–1078
Weltmeier F, Ehlert A, Mayer CS, Dietrich K, Wang X, Schutze K, Alonso R, Harter K, Vicente-Carbajosa J, Droge-Laser W (2006) Combinatorial control of Arabidopsis proline dehydrogenase transcription by specific heterodimerisation of bZIP transcription factors. EMBO J 25:3133–3143
Wessel PM, Graciet E, Douce R, Dumas R (2000) Evidence for two distinct effector-binding sites in threonine deaminase by site-directed mutagenesis, kinetic, and binding experiments. Biochemistry 39:15136–15143
Wittenbach V, Abell L (1999) Inhibitors of valine, leucine, and isoleucine biosynthesis. In: Singh BK (ed) Plant amino acids: biochemistry and biotechnology. Marcel Dekker, New York, pp 385–416
Wu K, Mourad G, King J (1994) A valine-resistant mutant of Arabidopsis thaliana displays an acetolactate synthase with altered feedback-control. Planta 192:249–255
Wu ZJ, Irizarry RA, Gentleman R, Martinez-Murillo F, Spencer F (2004) A model-based background adjustment for oligonucleotide expression arrays. J Am Stat Assoc 99:909–917
Wu B, Zhang B, Feng X, Rubens JR, Huang R, Hicks LM, Pakrasi HB, Tang YJ (2009) Alternate isoleucine synthesis pathway in cyanobacterial species. Microbiology. Epub ahead of print. doi:10.1099/mic.0.031799-0
Xu H, Zhang YZ, Guo XK, Ren SX, Staempfli AA, Chiao JS, Jiang WH, Zhao GP (2004) Isoleucine biosynthesis in Leptospira interrogans serotype lai strain 56601 proceeds via a threonine-independent pathway. J Bact 186:5400–5409
Yoshizawa F (2004) Regulation of protein synthesis by branched-chain amino acids in vivo. Biochem Biophys Res Commun 313:417–422
Zeh M, Casazza AP, Kreft O, Roessner U, Bieberich K, Willmitzer L, Hoefgen R, Hesse H (2001) Antisense inhibition of threonine synthase leads to high methionine content in transgenic potato plants. Plant Physiol 127:792–802
Acknowledgments
This work was funded by grants from the National Science Foundation (MCB-0416567), the Binational Agriculture Research and Development Agency (US-3910-06) and the Triad Foundation to GJ, and National Science Foundation grants DBI-0501778 and DBI-0820405 to ZF.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Joshi, V., Joung, JG., Fei, Z. et al. Interdependence of threonine, methionine and isoleucine metabolism in plants: accumulation and transcriptional regulation under abiotic stress. Amino Acids 39, 933–947 (2010). https://doi.org/10.1007/s00726-010-0505-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-010-0505-7