Abstract
Key message
During abiotic stress low abundant amino acids are not synthesized but they accumulate due to increased protein turnover under conditions inducing carbohydrate starvation (dehydration, salt stress, darkness) and are degraded.
Abstract
Metabolic adaptation is crucial for abiotic stress resistance in plants, and accumulation of specific amino acids as well as secondary metabolites derived from amino acid metabolism has been implicated in increased tolerance to adverse environmental conditions. The role of proline, which is synthesized during Arabidopsis stress response to act as a compatible osmolyte, has been well established. However, conclusions drawn about potential functions of other amino acids such as leucine, valine, and isoleucine are not entirely consistent. This study reevaluates published datasets with a special emphasis on changes in the free amino acid pool and transcriptional regulation of the associated anabolic and catabolic pathways. In order to gain a comprehensive overview about the general direction of amino acid metabolism under abiotic stress conditions a complete map of all currently known enzymatic steps involved in amino acid synthesis and degradation was assembled including also the initial steps leading to the synthesis of secondary metabolites. Microarray datasets and amino acid profiles of Arabidopsis plants exposed to dehydration, high salinity, extended darkness, cold, and heat were systematically analyzed to identify trends in fluxes of amino acid metabolism. Some high abundant amino acids such as proline, arginine, asparagine, glutamine, and GABA are synthesized during abiotic stress to act as compatible osmolytes, precursors for secondary metabolites, or storage forms of organic nitrogen. In contrast, most of the low abundant amino acids are not synthesized but they accumulate due to increased protein turnover under conditions inducing carbohydrate starvation (dehydration, salt stress, extended darkness) and are degraded.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amino acids are involved in several physiological processes in plants apart from constituting proteins. Most importantly, they act as precursors for a diverse set of secondary metabolites and are the transport and storage form for organic nitrogen within the plant. Gln synthesis is the only way to assimilate inorganic nitrogen into organic molecules and thus prerequisite for the production of all other nitrogen containing metabolites (Bernard and Habash 2009). Also, amino acids are tightly linked to carbohydrate metabolism, they are synthesized mainly from intermediates of the calvin cycle and degraded to either TCA cycle intermediates or their precursors (Pratelli and Pilot 2014; Hildebrandt et al. 2015; Supp. Fig. S1). Plants are able to synthesize all 20 proteinogenic amino acids de novo from inorganic carbon, nitrogen, and sulfur compounds (Fig. 1a). The set of secondary metabolites including non-protein amino acids that can be produced via modification of these 20 amino acids is extremely diverse and serves critical functions in plant metabolism such as signaling, defense, structure, interaction with other organisms, and protection from various abiotic stresses (D’Auria and Gershenzon 2005). For example, N-hydroxy-pipecolic acid required for long distance signaling during the establishment of systemic acquired resistance is produced from Lys; Met is a precursor for the synthesis of the hormone ethylene, glucosinolates, as well as the methyl group donor S-adenosylmethionine; polyamines are derived from Arg; the aromatic amino acids can be converted to numerous secondary metabolites such as isoquinoline and indole alkaloids, phenylpropanoids, glucosinolates, and auxin (Supp. Fig. S1; Wittstock and Halkier 2002; Alcázar et al. 2006; Amir 2010; Tzin and Galili 2010; Chen et al. 2018; Hartmann et al. 2018).
Interactions between free and protein-bound amino acid pools. a Schematic presentation of reactions relevant for pool sizes of free and protein bound amino acids. b Fraction of each individual amino acid [%] in the free amino acid pool of non-stressed plants (black bars) or included in proteins (white bars). The mean free amino acid content of non-stressed Arabidopsis plants was calculated from published datasets (Lugan et al. 2010; Watanabe et al. 2013; Krüßel et al. 2014). Absolute contents of free amino acids are shown on the right axis. The protein bound amino acid fractions are based on the theoretical Arabidopsis proteome extracted from the TAIR database (Hildebrandt et al. 2015). To illustrate the effect of protein degradation on the free amino acid pool I calculated the fold change ratio for each amino acid in the mean free pool of non-stressed plants (grey bars) assuming that degradation of the theoretical proteome took place leading to a 10% increase in the most abundant free amino acid, Gln (the mathematical formulas are described in the “Methods” section)
Interestingly, steady state levels of the individual amino acids are very different indicating specific functions in plant metabolism (Fig. 1b, black bars). Some amino acids such as Gln and Glu, Ser, Ala, Pro, Asn and Asp, are present in millimolar concentrations. They are connected to carbohydrate metabolism via very short pathways (Supp. Fig. S1), non-toxic and function e.g. as stores for organic nitrogen (Gln and Asn) or as compatible osmolytes (Pro). In contrast, the low abundant amino acids including Lys, the branched-chain amino acids Leu, Val, and Ile, the aromatic amino acids Phe, Tyr, and Trp, as well as the sulfur amino acids Met and Cys are normally present in low micromolar concentrations. Most of them are energetically costly and their synthesis as well as degradation requires multiple reaction steps (Hildebrandt et al. 2015; Supp. Fig. S1). Another reason for tightly controlling the concentration of individual amino acids might be a high reactivity leading to toxic effects, which has been shown for Cys (Jacob et al. 2003; Park and Imlay 2003). Thus, free amino acid homeostasis has to be regulated by controlling metabolic flux through the relevant pathways, which at the level of amino acid synthesis is mainly achieved by allosteric feedback inhibition (Less and Galili 2008).
However, since the major fraction of amino acids is present in proteins proteolysis as well as protein synthesis massively affect free amino acid pool sizes (Fig. 1). In order to estimate the impact of protein degradation on free amino acid levels correctly, it is important to keep in mind that the average content of the 20 amino acids within proteins varies by less than a factor of 10 with Leu (9.2%) being the most abundant and Trp (1.2%) the least abundant amino acid (Fig. 1b, white bars; Hildebrandt et al. 2015). The effect of protein breakdown on the free amino acid pool can be nicely illustrated by calculating the fold-change ratio for each amino acid on the basis of its mean content in non-stressed Arabidopsis plants (Fig. 1b, grey bars). Degradation of the average proteome leading to a moderate 10% increase in the most abundant free amino acid (Gln) would massively change the relative contents of the low abundant amino acids producing for example a 35-fold increase (3500%) in free Leu.
Proteolysis is increased in response to several abiotic stress conditions to provide amino acids as substrates for ATP production and to remobilize reduced nitrogen and sulfur (Araújo et al. 2011). Large changes in free amino acid levels have been detected in stress treated plants with particularly high fold increases of specific amino acids (Obata and Fernie 2012). The role of Pro as a compatible osmolyte during drought and salt stress has been well established. The genes involved in Pro synthesis are strongly induced during osmotic stress leading to increased production rates and a strong accumulation of Pro to high millimolar levels (Verslues and Sharma 2010). More recent data demonstrated that Pro catabolism is also relevant for increased drought resistance and can act as a buffer of the cellular redox status (Bhaskara et al. 2015; Shinde et al. 2016). But what about the other 19 amino acids? Do any of them have particular functions during abiotic stress response, and is their accumulation caused by increased synthesis rates or rather a consequence of protein degradation? The branched-chain and aromatic amino acids as well as Lys show particularly high fold increases under different stress conditions, and based on bioinformatic studies a strong induction of their synthesis has been postulated (Urano et al. 2009; Joshi et al. 2010; Obata and Fernie 2012). In the case of branched-chain amino acids, the line of arguments leading to this conclusion was based on high fold changes and induced expression of BCAT2, one of six isoforms of branched-chain amino acid transaminase present in Arabidopsis (Urano et al. 2009). However, BCATs not only catalyze the last step of branched-chain amino acid synthesis but also the first step in branched-chain amino acid degradation and it turned out that the particular isoform selected as a marker of increased synthesis is mainly involved in the degradation pathway (Angelovici et al. 2013). A recent study indeed provided experimental evidence showing that branched-chain amino acids increased during drought stress as a consequence of protein degradation (Huang and Jander 2017).
The aim of the present study is to gain a comprehensive overview about amino acid contents and the pathways modifying the free amino acid pool in order to better understand the direction of amino acid metabolism under abiotic stress conditions. A data mining approach was used based on published Arabidopsis microarray data as well as amino acid profiles. However, in order to avoid the pitfalls of previous studies not only fold changes but also the absolute amino acid contents in the tissue were considered. In addition, expression analysis of selected enzymatic steps of a pathway or even single isoforms was replaced by a complete pathway map to systematically evaluate all the available information about potential fluxes in amino acid metabolism during drought, salt, extended darkness, cold, and heat stress.
Results
Selection of metabolomics and transcriptomics datasets
The most frequently studied abiotic stress conditions are dehydration, high salinity, extended darkness, cold, and heat, so that suitable microarray datasets for data mining in Arabidopsis are available. However, growth conditions as well as the severity of the stress affect the metabolic reaction of the plant. Thus, for each of the stress conditions three microarray datasets and one amino acid profile were selected carefully choosing those studies that were based on experimental conditions closely reflecting the naturally occurring stress situation (Supp. Table S1). As far as possible, experiments using plants grown on soil were selected to avoid secondary effects caused by specific properties of an artificial growth system such as the composition of the medium. Stress treatments were started during the vegetative growth stage. Gradual dehydration was achieved by not watering the plants for several days. Plant were cultivated without light to study extended darkness and transferred to 4–8 °C for 24–48 h to apply cold stress or to 37–40 °C for 1–2 h for heat stress treatment. The effect of high salinity on Arabidopsis metabolism is mostly studied in artificial culture systems that allow better control of the NaCl concentration. Therefore, the datasets selected for salt stress are based on experiments using hydroponic culture or artificial soil. The original datasets used have been included in the supplemental material (Supp. Tables S2–S4).
Behaviour of free amino acid pools during different abiotic stress conditions
Amino acid profiles can be analyzed by different approaches and mostly either an HPLC method based on separation of specifically labeled amino acids or a GC/MS approach commontly used for large scale metabolomics is applied. Amino acid contents are either presented as absolute contents in nmol per fresh or dry weight or as relative peak areas of treatment versus control. In order to estimate the pool sizes of individual amino acids and the potential effect of amino acid supply by proteolysis correctly as far as possible datasets reporting absolute values were selected (salt stress and extended darkness, Fig. 2, black and grey bars). For drought, cold, and heat stress only fold changes were available so that in these cases the mean amino acid contents of non-stressed plants (Fig. 1b) were used as a basis for calculating the absolute contents of each amino acid after the stress treatment (Fig. 2, grey bars). The mathematical operations applied are described in the “Methods” section.
Detected versus expected changes in the free amino acid pool after abiotic stress treatments. The amino acid content [nmol/mg FW] of stress treated (grey bars) versus control (black bars) plants was directly taken from the literature for salt stress (Lugan et al. 2010), and extended darkness (Krüßel et al. 2014). For drought (Pires et al. 2016), cold, and heat (Kaplan et al. 2004) stress only fold changes were reported so that the mean amino acid contents of non-stressed plants (see Fig. 1b) were used as a basis for calculating the absolute contents of each amino acid after the stress treatment. White bars represent the expected changes in individual amino acid contents assuming that the entire increase in the free amino acid pool (calculated as the sum of all detected changes) was caused exclusively by protein degradation (the mathematical formulas are described in the “Methods” section). The difference between the detected and the expected change in the content of individual amino acids can be used to estimate whether net synthesis or degradation of the respective amino acid occurs during the stress condition analyzed. In the case of cold stress Gln was not included in the calculation of the free amino acid pool potentially produced by protein degradation since it is most likely the product of inorganic nitrogen assimilation. Alternative scenarios assuming de novo synthesis of Pro during salt stress and Asn during extended darkness can be found in Supp. Fig. S3
To provide a first idea whether the detected increase in individual amino acid contents could be a consequence of protein degradation or would require massive de novo synthesis I calculated the theoretical content of each amino acid assuming that the net increase in the free amino acid pool was caused exclusively by degradation of proteins with an amino acid composition representative for the Arabidopsis proteome (Fig. 1b). Since the amino acid composition of Rubisco, a high abundant protein that is preferentially degraded during stress (Izumi et al. 2010), is very similar to the representative proteome (Supp. Fig. S2), this approach will provide a good estimation of the in vivo situation. The net increase in the free amino acid pool during each stress condition was calculated by subtracting the free amino acid content of control plants from the content in the stressed plants. This increase was then allocated to the individual amino acids according to their relative content in the representative proteome to determine the amount of each amino acid that would be expected to be produced by protein degradation. This approach can of course only be a raw estimation since the total free amino acid pool size also depends on amino acid synthesis and degradation as well as on the synthesis of secondary metabolites or even transport to the roots or seeds (Fig. 1a). However, large differences between expected (Fig. 2, white bars) and detected (Fig. 2, grey bars) amino acid contents provide a sound basis for postulating increased flux of the particular amino acid towards either synthesis or degradation during the stress condition studied.
The amino acid profiles clearly indicate that during salt stress Pro accumulates due to increased synthesis rates, which is in good agreement with previous studies. In addition, massive synthesis of Asn can be postulated to take place during extended darkness and of Gln during cold stress, most likely to provide storage and transport forms for reduced nitrogen. Since massive de novo synthesis would overestimate the amount of amino acids released from protein degradation, an alternative scenario was calculated for cold stress, extended darkness, and salt stress excluding the amino acid with the largest absolute increase as well as its precursors. The synthesis of Asn, Gln, and Pro during the different stress conditions will most likely incorporate both, amino groups generated by transamination of other amino acids as well as free ammonium and therefore the in vivo situation represents a mixture of both theoretical scenarios. In the case of cold stress, high rates of nitrogen assimilation can be postulated (see below) and therefore Gln was generally excluded during the calculation of the theoretical protein derived amino acid pool. For salt stress and extended darkness the situation is less clear and both possible scenarios are presented (Fig. 2; Supp. Fig. S3). The different versions consistently indicate that during drought, darkness, and cold stress plants contained much lower concentrations of Lys, sulfur containing and branched-chain amino acids than expected based on the assumption of protein turnover. Thus, increased catabolism rather than synthesis can be postulated for these low abundant amino acids. Interestingly, free amino acid levels were hardly affected by the heat shock treatment.
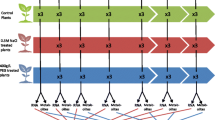
Transcriptional regulation of amino acid metabolism during abiotic stress conditions. Log2-fold changes in the expression of all currently known enzymes involved in amino acid synthesis and degradation pathways as well as of the enzymes catalyzing committed steps leading to the synthesis of amino acid derived secondary metabolites (shown in italics) were extracted from selected microarray datasets (Supp. Table S1). The colored squares in the figure represent the sum of all expression data available for the respective branch of the pathway in the three datasets analyzed for each abiotic stress condition (drought, salt, extended darkness, cold, heat). Arrows leading to amino acids represent the synthesis pathways, arrows pointing away from the amino acid show catabolic reactions, and metabolites that can be interconverted by a single set of enzymes are connected by arrows pointing in both directions. Pathways and metabolites marked by a question mark are presently unknown. The complete enzyme list including the microarray results as well as the position of each enzyme within the pathway scheme is shown in the supplemental material (Supp. Table S3; Supp. Fig. S1). 2-OB 2-oxobutyrate, 2-OG 2-oxoglutarate, 3-MOB 3-methyl-2-oxobutanoate, 3-MOP 3-methyl-2-oxopentanoate, 3-PG 3-phosphoglycerate, 4-MOP 4-methyl-2-oxopentanoate, GABA γ-aminobutyric acid, Glx glyoxylate, IAA indole-3-acetic acid, OA oxaloacetic acid, Pyr pyruvate, SAM S-adenosyl methionine
Transcriptional regulation of amino acid metabolism during abiotic stress
In order to present a clear but also comprehensive overview about the transcriptional regulation of amino acid metabolism during abiotic stress conditions I produced a scheme containing all currently known enzymes involved in amino acid synthesis and degradation pathways in Arabidopsis (Supp. Fig. S1; Supp. Table S3). In general, arrows leading to amino acids represent the synthesis pathways whereas arrows pointing away from the amino acid show catabolic reactions. However, several amino acids such as Thr, Gly, and Ser can be interconverted via reversible reactions and those are indicated by arrows pointing in both directions. Also, it has to be considered that some amino acids serve as precursors for the synthesis of others, e.g. Lys, Thr, Met, and Ile are all derived from Asp. The enzymes catalyzing committed steps leading to the synthesis of amino acid derived secondary metabolites (shown in italics) were also included in the scheme so that their effect on amino acid pool sizes can be estimated. However, a detailed evaluation of secondary metabolism during stress is not within the scope of this study.
Log2-fold changes in the expression of all enzymes involved in amino acid metabolism were extracted from the selected datasets (Supp. Table S3). All expression data available for the individual branches of the pathways were summed up for each stress condition analyzed (drought, salt, extended darkness, cold, heat). The results were converted to a color gradient to provide a visual impression of the direction of metabolism for each of the 20 proteinogenic amino acids (Fig. 3). Since several pathways are closely interconnected it is in some cases not possible to clearly distinguish between anabolic and catabolic reactions and thus essential to consider the complete scheme in order to understand and correctly interpret the different datasets. For example, the strong decrease in free Gln and Glu in salt stressed plants (Fig. 2) makes perfect sense considering that the synthesis pathways for Pro and GABA, which are both derived from Glu, are highly induced (Fig. 3d). The clear induction of Gln synthesis during cold stress (Fig. 3d) and Asn synthesis during extended darkness (Fig. 3a) are also very consistent with the amino acid profiles.
Looking at the color distribution it is obvious that the synthesis pathways for most low abundant amino acids including Lys, Met, Cys, Val, Leu, Ile, His, and Phe are generally down-regulated during abiotic stress. Production of polyamines from Arg seems to be slightly induced during drought, salt and cold stress (Fig. 3d). In contrast, the synthesis of glucosinolates as an additional class of secondary metabolites is strongly down-regulated on a transcriptional level by all abiotic stress conditions analyzed. The presently known degradation pathways of the low abundant amino acids are consistently induced during dehydration, salt stress, and darkness.
Combined information from transcriptome and metabolome analysis reveals direction of amino acid metabolism during abiotic stress
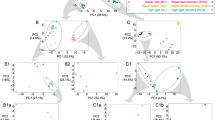
To make a systematic comparison and correct interpretation of the results from transcriptome and metabolome datasets more convenient, the most relevant data points regarding the direction of amino acid metabolism were compiled into one diagram (Fig. 4a). For each amino acid, induction of degradation or synthesis pathways is indicated on the left side (“Transcriptomics”) and can be directly compared to the results of the metabolite analysis shown on the right side (“Metabolomics”). For the metabolite data, the difference between amino acid contents detected in stressed plants and the theoretical profile resulting from proteolysis was calculated. Transcriptomics and metabolomics datasets are consistent if bars point into the same direction with bars to the left indicating increased catabolism and bars to the right indicating increased synthesis of the respective amino acid.
Combined information from transcriptome and metabolome studies reveal direction of amino acid metabolism during abiotic stress conditions. a Transcriptomics: the mean relative expression level (log2-ratio stress/control) of enzymes involved in either the degradation pathway (left side) or the synthesis pathway (right side) of the respective amino acid was calculated for each abiotic stress condition (drought, salt, extended darkness, cold, heat). Only increased mean expression levels (log2 > 0) are shown. Metabolomics: for each amino acid the theoretical content produced by protein degradation (see Fig. 2, white bars) was subtracted from the actual contend measured in stressed Arabidopsis leaves (see Fig. 2, grey bars). b In order to identify general trends in the expression levels of genes involved in protein degradation versus synthesis the sum of log2-fold changes in the expression of all genes included in bin 29.5. (“protein degradation”) or bin 29.2. (“protein synthesis”) in the MapMan annotation file (version Ath_AGI_LOCUS_TAIR10_Aug2012) was calculated in the three studies analyzed for each stress conditions, respectively. In addition, the sum of changes in the expression levels of autophagy-related proteins is shown
The induction of Gln synthesis during cold stress and Asn synthesis during extended darkness already described in the previous section is nicely visible also in this representation (Fig. 4a, blue and black bars). In addition, the catabolic pathways for Lys, the branched-chain amino acids and the sulfur containing amino acids are clearly induced by drought, salt, and darkness, which is very consistent with the amino acid profiles (Fig. 4a, grey, white, and black bars). However, interestingly during cold stress the microarray datasets indicate a strong down-regulation of these pathways (Fig. 4a, blue bars and Fig. 3) although the amino acid contents are very low. Looking for a possible explanation for this obvious discrepancy I analyzed general trends in the expression levels of genes involved in protein synthesis and degradation as well as in autophagy, a process associated with bulk nutrient remobilization (Fig. 4b; Supp. Table S4). Indeed, the microarray datasets indicate an induction of autophagy and net protein degradation during drought stress, extended darkness, and less pronounced also during salt stress (Fig. 4b, grey, white, and black bars) and thus confirm the conclusions drawn so far. In contrast, cold stress clearly induces net protein synthesis (Fig. 4b, blue bars), which nicely resolves the apparent conflict described above and also is in line with the postulated high rates of nitrogen assimilation via the glutamine synthetase–glutamate synthase pathway (Fig. 2). During heat stress there seems to be a balance between protein synthesis and degradation (Fig. 4b, red bars), resulting in rather constant steady state levels of the individual free amino acids and (Fig. 2) and little changes in amino acid metabolism compared to control conditions (Fig. 4a, red bars).
Discussion
Environmental conditions such as aridity, extreme temperatures (heat, cold, freezing), and high salt concentrations are major abiotic stress factors limiting plant growth and agronomical yield. Several reviews and metastudies have been published addressing different aspects of amino acid metabolism and their relevance for abiotic stress resistance (Less and Galili 2008; Joshi et al. 2010; Araújo et al. 2011; Obata and Fernie 2012; Krasensky and Jonak 2012). However, conclusions drawn about the metabolism of low abundant amino acids were not entirely consistent and this study reevaluates published datasets with a special emphasis on changes in the free amino acid pool and associated pathways. Special care was taken to select experiments conducted under physiological conditions so that the results reflect general trends in Arabidopsis stress response. Of course, not every stressful condition can be covered by this approach, and it has to be kept in mind that factors such as the severity of the stress or the light regime can also affect amino acid levels. The main findings are summarized in Fig. 5. In order to understand the general direction of protein and amino acid metabolism it is important to keep in mind that the different abiotic stress conditions have specific but also overlapping physiological and metabolic effects (Bartels and Sunkar 2005; Zhu 2016). For example, several abiotic stresses such as dehydration, high salinity, and freezing ultimately result in desiccation of the cell and osmotic imbalance. In order to prevent a loss of turgor and to protect proteins from denaturation plants accumulate compatible osmolytes which can be sugars but also amino acids or proteins. In reaction to dehydration plants also close their stomata to prevent transpirational water loss, which however interferes with photosynthesis due to a decline in intracellular CO2 levels resulting in reduced photosynthesis rates and increased oxidative stress (Mahajan and Tuteja 2005).
Summary of changes in the free amino acid pool during abiotic stress conditions. The intensity of the arrows indicates the flux through the respective pathway. a During drought and salt stress free amino acids are mainly produced by protein degradation. Low abundant amino acids are degraded and Pro is synthesized as compatible osmolyte. Also, amino acids are used as precursors for the synthesis of secondary metabolites. b Extended darkness leads to protein degradation for the production of amino acids as an alternative substrate for mitochondrial respiration. Asn is used for N-storage. c Cold stress leads to net protein synthesis, N-fixation into Gln, and the synthesis of GABA, polyamines and indole alkaloids from amino acids. d During heat stress protein degradation and synthesis are approximately balanced so that only minor changes in the free amino acid pool occur
Differential roles of protein degradation and synthesis during abiotic stress conditions
According to the results of the present study autophagy and protein degradation are highly relevant for the free amino acid pool during dehydration, salt stress, and extended darkness (Fig. 5a, b). This finding agrees very well with experimental data demonstrating a sharp decrease of the total protein content and increased protease activity in plants exposed to osmotic stress, drought, or extended darkness (Araújo et al. 2010; Pires et al. 2016; Huang and Jander 2017). Autophagy, which involves degradation of complete subcellular compartments, is induced in response to a lack of carbohydrate supply to remobilize lipids and proteins (Araújo et al. 2011). Increased proteolysis is essential to remove proteins that have been damaged during the stressful condition and it also provides amino acids as alternative respiratory substrates when photosynthesis rates are decreased due to stomatal closure, which occurs during drought and to a lesser extent also salt stress (Araújo et al. 2011). Extended darkness obviously also interferes with photosynthesis and can thus be used to study the effects of carbohydrates starvation on amino acid metabolism separated from osmotic stress or other consequences of abiotic stress.
In contrast, protein synthesis seems to be particularly relevant during extreme temperature stress (Fig. 5c, d). A well-established defense reaction against protein aggregation is the increased production of heat-shock proteins, which are chaperones preserving or reestablishing the native structure of proteins. In addition, antioxidant enzymes are highly induced by different stress conditions (Mahajan and Tuteja 2005; Hasanuzzaman et al. 2013). Interestingly, plants exposed to low temperatures also synthesize large amounts of non-enzymatic proteins to increase their protein content and decrease the freezing point thus preventing ice formation (Guy 1990). The high rates of incorporation into proteins can be expected to keep the contents of the low abundant amino acids low during cold stress, which is reflected by the amino acid profile (Fig. 2).
Some high abundant amino acids are synthesized to act as osmolytes or nitrogen stores
Accumulation of free amino acids in plants exposed to abiotic stress has been repeatedly reported (Joshi et al. 2010; Obata and Fernie 2012; Krasensky and Jonak 2012). This increase can result from stress-induced protein breakdown. However, plants might also actively synthesize particular amino acids with specific beneficial functions during stress response, and it is important to identify these amino acids and their functions in order to fully unravel the principles of increased stress resistance. Plants cope with osmotic stress by decreasing their intracellular osmotic potential via accumulation of compatible solutes, i.e. sugars and also amino acid derived metabolites such as Pro, and the non-protein amino acid γ-aminobutyric acid (GABA) (Krasensky and Jonak 2012). Pro synthesis and the GABA shunt are important for stress tolerance and both metabolites are considered to act as ROS scavenger (Bouché and Fromm 2004; Szabados and Savouré 2010). The results of the present analysis are in good agreement with already established knowledge showing that Pro synthesis is induced during drought, salt, and cold stress but not during heat (Fig. 5; Lv et al. 2011; Verslues and Sharma 2010). However, it has to be noted that a more prominent role of Pro with up to 100-fold accumulation has been reported for Arabidopsis plants exposed to low water potential using either detached leaves or PEG treatment as an experimental system (Urano et al. 2009; Verslues and Sharma 2010). The effect of gradual dehydration on plants grown in soil analyzed here was probably less severe and a strong response seems to occur rather late, shortly before the plant dies. Since dehydration as well as high salinity both lead to osmotic stress, a partial overlap in stress responses can be expected but there are also a number of specific effects. This meta-analysis suggests a much stronger increase in protein degradation during drought compared to salt stress probably due to an increased demand for amino acids as substrates for ATP production reminiscent of extended darkness conditions. In contrast, high salinity clearly led to increased Pro synthesis, whereas during gradual dehydration Gln accumulated, which might act as a precursor for rapid Pro synthesis during later stages of more severe drought stress. The observed decrease in Pro content and induction of proline dehydrogenase after extended darkness are well in line with the literature and can be explained by oxidation of Pro as an alternative respiratory substrate (Ábrahám et al. 2003). However, while the microarray datasets analyzed in the frame of the present study also indicate enhanced Pro synthesis, which might serve as a non-toxic storage form for the amino groups liberated during the oxidation of other amino acids, an inhibitory effect of darkness on the induction of Δ-1-pyrroline-5-carboxylate synthetase has also been reported (Ábrahám et al. 2003). This discrepancy might be due to the different growth conditions used since sucrose supply from the medium will delay the carbohydrate starvation response.
The present dataset in addition indicates an induction of Gln and Asn synthesis during stress (Fig. 5b, c). Gln synthetase is required for reassimilating the nitrogen liberated during amino acid catabolism and it is also involved in primary nitrogen assimilation in situations of increased protein synthesis such as cold acclimation of the plant (Bernard and Habash 2009). Asn can be used for nitrogen transport and storage within the plant (Lam et al. 2003). The high level of consistency with previous results demonstrates that the approach used in this study is suitable to identify general directions of amino acid metabolism during abiotic stress.
Lys, branched-chain and sulfur amino acids accumulate due to proteolysis and are catabolized
Combined analysis of transcriptome and metabolome datasets revealed a clear trend for the metabolism of specific low abundant amino acids during abiotic stress. Branched-chain amino acids, Lys and sulfur amino acids are not synthesized but their degradation pathways are strongly induced during dehydration, salt stress and extended darkness (Fig. 5a, b). In contrast to some high abundant amino acids (Pro, GABA, Gln, Asn), accumulation of the normally low abundant amino acids is a consequence of increased protein turnover during abiotic stress. This conclusion has already been confirmed experimentally for the branched-chain amino acids Leu, Val and Ile, which are degraded by partially overlapping pathways (Hildebrandt et al. 2015; Huang and Jander 2017). Drought-induced accumulation of branched-chain amino acids was decreased in the presence of a protease inhibitor but not affected by inhibition of their synthesis pathway. ABA signaling seems to be involved since a mutant line deficient in stress-induced ABA biosynthesis also had lower branched-chain amino acid contents than the wild type after different osmotic stress treatments (Huang and Jander 2017).
Several studies using mutant lines have demonstrated the functional significance of amino acid degradation for stress resistance. The degradation pathways for the branched-chain amino acids are rather complex and not completely known. The initial steps, transamination followed by two oxidation steps, are catalyzed by a common set of enzymes for Leu, Val, and Ile (Hildebrandt et al. 2015). Isovaleryl-CoA dehydrogenase, catalyzing the second oxidation step, transfers electrons to the electron-transfer flavoprotein (ETF)/ETF:ubiquinone oxidoreductase (ETFQO) complex, which is bound to the inner mitochondrial membrane and donates electrons to the respiratory chain (Däschner et al. 2001; Ishizaki et al. 2005; Araújo et al. 2010). Loss-of-function mutants for IVDH and ETFQO, as well as for d-2-hydroxyglutarate dehydrogenase (D2HGDH), an enzyme involved in Lys catabolism that also transfers electrons to the ETF/ETFQO complex, are less tolerant to drought stress and extended darkness than the wild type (Ishizaki et al. 2005, 2006; Araújo et al. 2010; Pires et al. 2016). These lines accumulate the amino acids that are degraded by the respective pathways (BCAAs, Lys) but also show additional changes in the free amino acid pool during stress, indicating close interactions of the amino acid metabolic networks.
There are a number of possible explanations for the importance of these specific amino acid degradation pathways for stress tolerance. Due to their complex structure, oxidation of branched-chain amino acids and Lys produces high amounts of ATP, which might be essential for survival during stress conditions provoking carbohydrate starvation (Hildebrandt et al. 2015). In addition, the amino acids or their derivatives could have specific functions, e.g. as signaling molecules, so that high concentrations would be toxic. Leu has been suggested to be a metabolic signal, and Lys is the precursor for N-hydroxy-pipecolic acid, a transmitter during the establishment of systemic acquired resistance (Hannah et al. 2010; Chen et al. 2018; Hartmann et al. 2018). However, the datasets analyzed here do not indicate any particular role of N-hydroxy-pipecolic acid in abiotic stress response.
Met seems to be degraded to methane thiol and 2-oxobutyrate by methionine-γ-lyase (MGL) rather than converted to glucosinolates, ethylene, or the methyl group donor S-adenosyl methionine (Fig. 3). Since 2-oxobutyrate is a precursor of Ile synthesis, induction of MGL could also be interpreted as an indicator for increased Ile synthesis during stress. However, Thr deamination, which is consistently down-regulated during abiotic stress, has been shown to be the main route of 2-oxobutyrate production for Ile synthesis (Joshi and Jander 2009; Less and Galili 2008). Also, expression of the enzymes catalyzing subsequent steps in Ile production is decreased so that it is reasonable to consider MGL as a Met catabolic enzyme during abiotic stress.
Cysteine can be catabolized via several different routes that either produce H2S or oxidize the thiol group to thiosulfate or sulfate (Fig. 3; Hildebrandt et al. 2015). During drought, salt stress, and extended darkness the mitochondrial cysteine oxidation pathway catalyzing complete sulfur oxidation via the sulfur dioxygenase ETHE1 is strongly induced, and also breakdown of the cysteine dimer cystine by cystine lyase is upregulated (Supp. Fig. S1; Supp. Table S3 reactions D15–D17 and D41). In contrast, cysteine desulfhydrases (reaction D19) are repressed by dehydration as well as cold stress and slightly induced by extended darkness in the microarray datasets analyzed here. The cytosolic cysteine desulfhydrase DES1 generates H2S as a signaling molecule, which has been shown to positively regulate stomatal closure but also to act as a repressor of autophagy (Gotor et al. 2013; Scuffi et al. 2014). Since both of these processes are relevant for abiotic stress resistance the regulation of cysteine metabolism might be particularly relevant. Production of reduced sulfur for the synthesis of iron–sulfur clusters by cysteine desulfurases (reaction D18) is at least at a transcriptional level not affected by abiotic stress.
Aromatic amino acids are precursors for the synthesis of secondary metabolites
The aromatic amino acids Tyr, Phe, and Trp are precursors for a diverse set of secondary metabolites with different functions in stress resistance (Tzin and Galili 2010). The microarray datasets indicate increased production of individual secondary metabolites during dehydration, salt and cold stress (Fig. 5a, c). However, a detailed analysis of plant secondary metabolism in the different abiotic stress conditions is clearly without the scope of this study. Only the first committed steps of the pathways were included in order to estimate the effect of secondary metabolite production on the free amino acid pool sizes. In general, the amount of aromatic amino acids produced by protein degradation seem to be sufficient for secondary metabolite production since the synthesis pathways of the aromatic amino acids are not affected or even repressed by abiotic stress except for Trp synthesis, which is induced by cold stress (Fig. 5c). Interestingly, Tyr degradation is strongly up-regulated during drought, salt stress and extended darkness showing a similar pattern as the other low abundant amino acids discussed in the previous section. In the case of Trp, Phe and also His it is difficult to draw a final conclusion about the direction of metabolism since the catabolic pathways are currently unknown. After unraveling these pathways it might be possible to combine all low abundant amino acids into a common scheme.
Regulation of amino acid metabolism during abiotic stress
Enzyme activities are regulated by a combination of several different mechanisms on the level of transcription, translation, post-translational modification and protein stability. It has been shown, that changes in protein levels do not always reflect the change in transcript levels. Therefore, transcriptome analysis alone cannot represent the complete picture. However, it is unlikely that plants will apply different regulatory strategies with largely contrasting effects to individual enzymes or pathways, so that clear trends in transcriptional changes still provide valuable clues. Ideally, hypotheses developed from the analysis of microarray datasets should be tested for consistency with other available datasets, and in this case the amino acid profiles were used. Transcriptional changes in response to abiotic stress were in general slightly stronger for the catabolic pathways than for the biosynthetic genes, which has also been noticed in a previous bioinformatics approach (Less and Galili 2008). Pro metabolism is an exception since abiotic stress leads to a strong induction of the biosynthetic enzymes. It is well established that the biosynthesis of most amino acids in plants is regulated by allosteric inhibition of the commited step by the endproduct of the respective pathway (Galili 1995; Radwanski and Last 1995; Joshi et al. 2010; Tzin and Galili 2010; Hong et al. 2000). Allosteric feedback inhibition is suitable for sensing amino acid levels and increasing the flux through the synthesis pathways when amino acids are removed by protein synthesis for example during active growth or cold stress. A local accumulation of amino acids during stress could be achieved by up-regulation of amino acid export from the site of synthesis (e.g. plastids) that would release feedback inhibition of biosynthetic enzymes leading to enhanced synthesis rates. However, the datasets analyzed here clearly show a down-regulation of several amino acid synthesis pathway on a transcriptional level making this scenario rather unlikely.
Conclusions
Using a combined analysis of transcriptome and metabolome datasets this study could identify general trends in amino acid metabolism during different abiotic stress conditions. Some high abundant amino acids such as Pro, Arg, Asn, Gln, and GABA are synthesized to act as compatible osmolytes, precursors for secondary metabolites, or nitrogen stores. In contrast, during stress conditions that induce carbohydrate starvation (dehydration, salt, darkness) the low abundant amino acids are not synthesized but they accumulate due to increased protein degradation and are catabolized. An exception might be Trp, which is required for the synthesis of different groups of secondary metabolites, and in some stress situations the supply by proteolysis could be not sufficient so that additional de novo synthesis is induced. By using a more comprehensive map of amino acid synthesis and degradation pathways this approach was able to reconcile apparent conflicts between expression data and biochemical results identified in previous studies based on only selected enzymatic steps (Urano et al. 2009; Joshi et al. 2010; Obata and Fernie 2012). Looking at the complete pathway map makes it easier to identify individual problematic datapoints and correctly interpret the general metabolic trends. It reveals quite clearly that the breakdown of amino acids for the production of carbon, nitrogen, and energy-associated molecules does normally not involve the synthesis of other amino acids but there are alternative less energy consuming routes (Fig. 3; Supp. Fig. S1). For example Asp will most likely be transaminated to oxaloacetate rather than converted to Lys, Met, or Thr for degradation. However, the power of omics approaches for the study of amino acid metabolism will be considerably improved after complete elucidation of the presently unknown (Trp, Phe, His) or only partially known (Leu, Val, Ile, Lys) catabolic pathways. The present analysis should provide a general impression of amino acid metabolism during abiotic stress in Arabidopsis, but it is not intended to be an exhaustive analysis of all possible reactions.
Methods
Data source
A complete map of currently known enzymatic steps involved in amino acid synthesis and degradation including also the initial steps of amino acid derived secondary metabolism (Supp. Fig. S1; Supp. Table S3) was assembled based on information available in databases [BioCyc (Caspi et al. 2016), UniProt (The UniProt Consortium 2017), KEGG (Kanehisa et al. 2017)] and selected publications (Less and Galili 2008; Pratelli and Pilot 2014; Hildebrandt et al. 2015). In addition many reactions were confirmed individually using original research articles to avoid false annotations. Genes involved in protein synthesis and degradation were selected based on the MapMan annotation file version Ath_AGI_LOCUS_TAIR10_Aug2012 (Thimm et al. 2004).
The amino acid composition of the theoretical Arabidopsis proteome has been published before (Hildebrandt et al. 2015). The mean free amino acid contents of non-stressed Arabidopsis plants were calculated from five published datasets (Lugan et al. 2010 (2×), Watanabe et al. 2013 (2×), Krüßel et al. 2014). Amino acid profiles of stressed plants were taken from Kaplan et al. (2004), Lugan et al. (2010), Krüßel et al. (2014), and Pires et al. (2016). The microarray datasets used for this study have been published in Seki et al. (2002), Gong et al. (2005), Kilian et al. (2007), Perera et al. (2008), Usadel et al. (2008), Ludwików et al. (2009), Shedge et al. (2010), Suzuki et al. (2011), Pandey et al. (2013), Kim et al. (2013), and Maeda et al. (2014). Expression levels were either downloaded from a repository (Genevestigator, Hruz et al. 2008) or supplied in the supplemental information of the respective publications. A list of the datasets selected for each stress conditions including a summary of the experimental conditions is provided in Supp. Table S1.
Metabolomics data analysis
Calculation of fold changes in free amino acid contents due to degradation of the theoretical proteome (Fig. 1b)
To calculate the fold change ratio for each amino acid assuming that protein degradation took place leading to a 10% increase of the most abundant free amino acid (Gln) the first step was to calculate the absolute increase in Gln by multiplying the mean free Gln content of non-stressed plants (2.93 nmol/mg FW) by 0.1. The resulting 0.293 nmol/mg FW represent 3.5% of the degraded representative protein, and all other absolute increases could be calculated by rule of proportion using their relative content in proteins. For each amino acid, the absolute increase was then added to the mean free content and the result was divided by the mean free content.
Calculation of free amino acid contents expected due to degradation of the theoretical proteome in stressed plants (Fig. 2)
The amino acid composition of the theoretical Arabidopsis proteome (Fig. 1b) was used as a basis to calculate the content of each amino acid that can be expected to result from degradation of a representative protein. The total increase in free amino acids was calculated as the sum of changes in the contents of all individual amino acids analyzed. Then, for each amino acid the relative content in proteins was multiplied by the total increase in free amino acids divided by the sum of relative contents of all amino acids included in the respective profile and the result was added to the free control content of the respective amino acid.
For drought stress, the published amino acid contents were corrected using the relative water content of the analyzed samples (24% for stressed leaves and 93% for control, Pires et al. 2016):
Gene expression data analysis
Log2-fold changes in the expression level of genes in stress treated versus control plants were extracted from the microarray datasets, and only significant changes (FDR < 0.05) were considered for further analysis. Expression data was sorted into the complete list of proteins involved in amino acid metabolism (Supp. Table S3) as well as in the list of proteins involved in autophagy, protein synthesis and protein degradation (Supp. Table S4). All log2-ratios available for the individual branches of the amino acid metabolic pathways were summed up for each stress condition analyzed (drought, salt, extended darkness, cold, heat) and the results were converted to a color gradient ranging from − 10 to 10 (Fig. 3).
References
Ábrahám E, Rigó G, Székely G, Nagy R, Koncz C, Szabados L (2003) Light-dependent induction of proline biosynthesis by abscisic acid and salt stress is inhibited by brassinosteroid in Arabidopsis. Plant Mol Biol 51:363–372
Alcázar R, Marco F, Cuevas JC, Patron M, Ferrando A, Carrasco P, Tiburcio AF, Altabella T (2006) Involvement of polyamines in plant response to abiotic stress. Biotechnol Lett 28:1867–1876. https://doi.org/10.1007/s10529-006-9179-3
Amir R (2010) Current understanding of the factors regulating methionine content in vegetative tissues of higher plants. Amino Acids 39:917–931. https://doi.org/10.1007/s00726-010-0482-x
Angelovici R, Lipka AE, Deason N, Gonzalez-Jorge S, Lin H, Cepela J, Buell R, Gore MA, Dellapenna D (2013) Genome-wide analysis of branched-chain amino acid levels in Arabidopsis seeds. Plant Cell 25:4827–4843. https://doi.org/10.1105/tpc.113.119370
Araújo WL, Ishizaki K, Nunes-Nesi A, Larson TR, Tohge T, Krahnert I, Witt S, Obata T, Schauer N, Graham IA, Leaver CJ, Fernie AR (2010) Identification of the 2-hydroxyglutarate and isovaleryl-CoA dehydrogenases as alternative electron donors linking lysine catabolism to the electron transport chain of Arabidopsis mitochondria. Plant Cell 22:1549–1563. https://doi.org/10.1105/tpc.110.075630
Araújo WL, Tohge T, Ishizaki K, Leaver CJ, Fernie AR (2011) Protein degradation—an alternative respiratory substrate for stressed plants. Trends Plant Sci 16:489–498. https://doi.org/10.1016/j.tplants.2011.05.008
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24:23–58. https://doi.org/10.1080/07352680590910410
Bernard SM, Habash DZ (2009) The importance of cytosolic glutamine synthetase in nitrogen assimilation and recycling. New Phytol 182:608–620. https://doi.org/10.1111/j.1469-8137.2009.02823.x
Bhaskara GB, Yang T, Verslues PE (2015) Dynamic proline metabolism: importance and regulation in water limited environments. Front Plant Sci 6:484. https://doi.org/10.3389/fpls.2015.00484
Bouché N, Fromm H (2004) GABA in plants: just a metabolite? Trends Plant Sci 9:110–115. https://doi.org/10.1016/j.tplants.2004.01.006
Caspi R, Billington R, Ferrer L, Foerster H, Fulcher CA, Keseler IM, Kothari A, Krummenacker M, Latendresse M, Mueller LA, Ong Q, Paley S, Subhraveti P, Weaver DS, Karp PD (2016) The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res 44:D471–D480. https://doi.org/10.1093/nar/gkv1164
Chen Y, Holmes EC, Rajniak J, Kim J, Tang S, Fischer CR, Mudgett MB, Sattely ES (2018) N-hydroxy-pipecolic acid is a mobile metabolite that induces systemic disease resistance in Arabidopsis. Proc Natl Acad Sci USA 115:E4920–E4929. https://doi.org/10.1073/pnas.1805291115
D’Auria JC, Gershenzon J (2005) The secondary metabolism of Arabidopsis thaliana: growing like a weed. Curr Opin Plant Biol 8:308–316. https://doi.org/10.1016/j.pbi.2005.03.012
Däschner K, Couée I, Binder S (2001) The mitochondrial isovaleryl-coenzyme a dehydrogenase of arabidopsis oxidizes intermediates of leucine and valine catabolism. Plant Physiol 126:601–612
Galili G (1995) Regulation of lysine and threonine synthesis. Plant Cell 7:899–906. https://doi.org/10.1105/tpc.7.7.899
Gong Q, Li P, Ma S, Indu Rupassara S, Bohnert HJ (2005) Salinity stress adaptation competence in the extremophile Thellungiella halophila in comparison with its relative Arabidopsis thaliana. Plant J 44:826–839. https://doi.org/10.1111/j.1365-313X.2005.02587.x
Gotor C, García I, Crespo JL, Romero LC (2013) Sulfide as a signaling molecule in autophagy. Autophagy 9:609–611. https://doi.org/10.4161/auto.23460
Guy CL (1990) Cold acclimation and freezing stress tolerance: role of protein metabolism. Annu Rev Plant Physiol Plant Mol Biol 41:187–223. https://doi.org/10.1146/annurev.pp.41.060190.001155
Hannah MA, Caldana C, Steinhauser D, Balbo I, Fernie AR, Willmitzer L (2010) Combined transcript and metabolite profiling of Arabidopsis grown under widely variant growth conditions facilitates the identification of novel metabolite-mediated regulation of gene expression. Plant Physiol 152:2120–2129. https://doi.org/10.1104/pp.109.147306
Hartmann M, Zeier T, Bernsdorff F, Reichel-Deland V, Kim D, Hohmann M, Scholten N, Schuck S, Bräutigam A, Hölzel T, Ganter C, Zeier J (2018) Flavin monooxygenase-generated N-hydroxypipecolic acid is a critical element of plant systemic immunity. Cell 173:456–469.e16. https://doi.org/10.1016/j.cell.2018.02.049
Hasanuzzaman M, Nahar K, Alam MM, Roychowdhury R, Fujita M (2013) Physiological, biochemical, and molecular mechanisms of heat stress tolerance in plants. Int J Mol Sci 14:9643–9684. https://doi.org/10.3390/ijms14059643
Hildebrandt TM, Nunes Nesi A, Araújo WL, Braun H (2015) Amino acid catabolism in plants. Mol Plant 8:1563–1579. https://doi.org/10.1016/j.molp.2015.09.005
Hong Z, Lakkineni K, Zhang Z, Verma DP (2000) Removal of feedback inhibition of delta(1)-pyrroline-5-carboxylate synthetase results in increased proline accumulation and protection of plants from osmotic stress. Plant Physiol 122:1129–1136
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinform 2008:420747. https://doi.org/10.1155/2008/420747
Huang T, Jander G (2017) Abscisic acid-regulated protein degradation causes osmotic stress-induced accumulation of branched-chain amino acids in Arabidopsis thaliana. Planta. https://doi.org/10.1007/s00425-017-2727-3
Ishizaki K, Larson TR, Schauer N, Fernie AR, Graham IA, Leaver CJ (2005) The critical role of Arabidopsis electron-transfer flavoprotein:ubiquinone oxidoreductase during dark-induced starvation. Plant Cell 17:2587–2600. https://doi.org/10.1105/tpc.105.035162
Ishizaki K, Schauer N, Larson TR, Graham IA, Fernie AR, Leaver CJ (2006) The mitochondrial electron transfer flavoprotein complex is essential for survival of Arabidopsis in extended darkness. Plant J 47:751–760. https://doi.org/10.1111/j.1365-313X.2006.02826.x
Izumi M, Wada S, Makino A, Ishida H (2010) The autophagic degradation of chloroplasts via rubisco-containing bodies is specifically linked to leaf carbon status but not nitrogen status in Arabidopsis. Plant Physiol 154:1196–1209. https://doi.org/10.1104/pp.110.158519
Jacob C, Giles GI, Giles NM, Sies H (2003) Sulfur and selenium: the role of oxidation state in protein structure and function. Angew Chem Int Ed Engl 42:4742–4758. https://doi.org/10.1002/anie.200300573
Joshi V, Jander G (2009) Arabidopsis methionine γ-lyase is regulated according to isoleucine biosynthesis needs but plays a subordinate role to threonine deaminase. Plant physiol 151:367–378
Joshi V, Joung J, Fei Z, Jander G (2010) Interdependence of threonine, methionine and isoleucine metabolism in plants: accumulation and transcriptional regulation under abiotic stress. Amino Acids 39:933–947. https://doi.org/10.1007/s00726-010-0505-7
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45:D353–D361. https://doi.org/10.1093/nar/gkw1092
Kaplan F, Kopka J, Haskell DW, Zhao W, Schiller KC, Gatzke N, Sung DY, Guy CL (2004) Exploring the temperature-stress metabolome of Arabidopsis. Plant Physiol 136:4159–4168. https://doi.org/10.1104/pp.104.052142
Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D’Angelo C, Bornberg-Bauer E, Kudla J, Harter K (2007) The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 50:347–363. https://doi.org/10.1111/j.1365-313X.2007.03052.x
Kim Y, Park S, Gilmour SJ, Thomashow MF (2013) Roles of CAMTA transcription factors and salicylic acid in configuring the low-temperature transcriptome and freezing tolerance of Arabidopsis. Plant J 75:364–376. https://doi.org/10.1111/tpj.12205
Krasensky J, Jonak C (2012) Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J Exp Bot 63:1593–1608. https://doi.org/10.1093/jxb/err460
Krüßel L, Junemann J, Wirtz M, Birke H, Thornton JD, Browning LW, Poschet G, Hell R, Balk J, Braun H, Hildebrandt TM (2014) The mitochondrial sulfur dioxygenase ETHYLMALONIC ENCEPHALOPATHY PROTEIN1 is required for amino acid catabolism during carbohydrate starvation and embryo development in Arabidopsis. Plant Physiol 165:92–104. https://doi.org/10.1104/pp.114.239764
Lam H, Wong P, Chan H, Yam K, Chen L, Chow C, Coruzzi GM (2003) Overexpression of the ASN1 gene enhances nitrogen status in seeds of Arabidopsis. Plant Physiol 132:926–935. https://doi.org/10.1104/pp.103.020123
Less H, Galili G (2008) Principal transcriptional programs regulating plant amino acid metabolism in response to abiotic stresses. Plant Physiol 147:316–330. https://doi.org/10.1104/pp.108.115733
Ludwików A, Kierzek D, Gallois P, Zeef L, Sadowski J (2009) Gene expression profiling of ozone-treated Arabidopsis abi1td insertional mutant: protein phosphatase 2C ABI1 modulates biosynthesis ratio of ABA and ethylene. Planta 230:1003–1017. https://doi.org/10.1007/s00425-009-1001-8
Lugan R, Niogret M, Leport L, Guégan J, Larher FR, Savouré A, Kopka J, Bouchereau A (2010) Metabolome and water homeostasis analysis of Thellungiella salsuginea suggests that dehydration tolerance is a key response to osmotic stress in this halophyte. Plant J 64:215–229. https://doi.org/10.1111/j.1365-313X.2010.04323.x
Lv W, Lin B, Zhang M, Hua X (2011) Proline accumulation is inhibitory to Arabidopsis seedlings during heat stress. Plant Physiol 156:1921–1933. https://doi.org/10.1104/pp.111.175810
Maeda H, Song W, Sage T, Dellapenna D (2014) Role of callose synthases in transfer cell wall development in tocopherol deficient Arabidopsis mutants. Front Plant Sci 5:46. https://doi.org/10.3389/fpls.2014.00046
Mahajan S, Tuteja N (2005) Cold, salinity and drought stresses: an overview. Arch Biochem Biophys 444:139–158. https://doi.org/10.1016/j.abb.2005.10.018
Obata T, Fernie AR (2012) The use of metabolomics to dissect plant responses to abiotic stresses. Cell Mol Life Sci 69:3225–3243. https://doi.org/10.1007/s00018-012-1091-5
Pandey N, Ranjan A, Pant P, Tripathi RK, Ateek F, Pandey HP, Patre UV, Sawant SV (2013) CAMTA 1 regulates drought responses in Arabidopsis thaliana. BMC Genomics 14:216. https://doi.org/10.1186/1471-2164-14-216
Park S, Imlay JA (2003) High levels of intracellular cysteine promote oxidative DNA damage by driving the fenton reaction. J Bacteriol 185:1942–1950
Perera IY, Hung C, Moore CD, Stevenson-Paulik J, Boss WF (2008) Transgenic Arabidopsis plants expressing the type 1 inositol 5-phosphatase exhibit increased drought tolerance and altered abscisic acid signaling. Plant Cell 20:2876–2893. https://doi.org/10.1105/tpc.108.061374
Pires MV, Pereira Júnior AA, Medeiros DB, Daloso DM, Pham PA, Barros KA, Engqvist MKM, Florian A, Krahnert I, Maurino VG, Araújo WL, Fernie AR (2016) The influence of alternative pathways of respiration that utilize branched-chain amino acids following water shortage in Arabidopsis. Plant Cell Environ 39:1304–1319. https://doi.org/10.1111/pce.12682
Pratelli R, Pilot G (2014) Regulation of amino acid metabolic enzymes and transporters in plants. J Exp Bot 65:5535–5556. https://doi.org/10.1093/jxb/eru320
Radwanski ER, Last RL (1995) Tryptophan biosynthesis and metabolism: biochemical and molecular genetics. Plant Cell 7:921–934. https://doi.org/10.1105/tpc.7.7.921
Scuffi D, Álvarez C, Laspina N, Gotor C, Lamattina L, García-Mata C (2014) Hydrogen sulfide generated by L-cysteine desulfhydrase acts upstream of nitric oxide to modulate abscisic acid-dependent stomatal closure. Plant Physiol 166:2065–2076. https://doi.org/10.1104/pp.114.245373
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Taji T, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31:279–292
Shedge V, Davila J, Arrieta-Montiel MP, Mohammed S, Mackenzie SA (2010) Extensive rearrangement of the Arabidopsis mitochondrial genome elicits cellular conditions for thermotolerance. Plant Physiol 152:1960–1970. https://doi.org/10.1104/pp.109.152827
Shinde S, Villamor JG, Lin W, Sharma S, Verslues PE (2016) Proline coordination with fatty acid synthesis and redox metabolism of chloroplast and mitochondria. Plant Physiol 172:1074–1088. https://doi.org/10.1104/pp.16.01097
Suzuki N, Sejima H, Tam R, Schlauch K, Mittler R (2011) Identification of the MBF1 heat-response regulon of Arabidopsis thaliana. Plant J 66:844–851. https://doi.org/10.1111/j.1365-313X.2011.04550.x
Szabados L, Savouré A (2010) Proline: a multifunctional amino acid. Trends Plant Sci 15:89–97. https://doi.org/10.1016/j.tplants.2009.11.009
The UniProt Consortium (2017) UniProt: the universal protein knowledgebase. Nucleic Acids Res 45:D158–D169. https://doi.org/10.1093/nar/gkw1099
Thimm O, Bläsing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller LA, Rhee SY, Stitt M (2004) mapman: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939. https://doi.org/10.1111/j.1365-313X.2004.02016.x
Tzin V, Galili G (2010) New insights into the shikimate and aromatic amino acids biosynthesis pathways in plants. Mol Plant 3:956–972. https://doi.org/10.1093/mp/ssq048
Urano K, Maruyama K, Ogata Y, Morishita Y, Takeda M, Sakurai N, Suzuki H, Saito K, Shibata D, Kobayashi M, Yamaguchi-Shinozaki K, Shinozaki K (2009) Characterization of the ABA-regulated global responses to dehydration in Arabidopsis by metabolomics. Plant J 57:1065–1078. https://doi.org/10.1111/j.1365-313X.2008.03748.x
Usadel B, Bläsing OE, Gibon Y, Retzlaff K, Höhne M, Günther M, Stitt M (2008) Global transcript levels respond to small changes of the carbon status during progressive exhaustion of carbohydrates in Arabidopsis rosettes. Plant Physiol 146:1834–1861. https://doi.org/10.1104/pp.107.115592
Verslues PE, Sharma S (2010) Proline metabolism and its implications for plant-environment interaction. Arabidopsis Book 8:e0140. https://doi.org/10.1199/tab.0140
Watanabe M, Balazadeh S, Tohge T, Erban A, Giavalisco P, Kopka J, Mueller-Roeber B, Fernie AR, Hoefgen R (2013) Comprehensive dissection of spatiotemporal metabolic shifts in primary, secondary, and lipid metabolism during developmental senescence in Arabidopsis. Plant Physiol 162:1290–1310. https://doi.org/10.1104/pp.113.217380
Wittstock U, Halkier BA (2002) Glucosinolate research in the Arabidopsis era. Trends Plant Sci 7:263–270
Zhu J (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324. https://doi.org/10.1016/j.cell.2016.08.029
Acknowledgements
I thank Michael Senkler for helping me with the extraction of the expression data and Hans-Peter Braun for critically reading the manuscript.
Author information
Authors and Affiliations
Contributions
TMH performed all tasks necessary to produce this manuscript.
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hildebrandt, T.M. Synthesis versus degradation: directions of amino acid metabolism during Arabidopsis abiotic stress response. Plant Mol Biol 98, 121–135 (2018). https://doi.org/10.1007/s11103-018-0767-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-018-0767-0