Abstract
A new series of pyrazolyl–chalcone derivatives was prepared in moderate yields via Claisen–Schmidt condensation reaction of 4-acetylpyrazole derivatives with the corresponding aldehydes. The newly synthesized compounds have been fully characterized by 1H NMR, 13C NMR, IR, mass spectrometry, and elemental analysis. The in vitro antimicrobial and anti-cancer activities of the novel compounds were evaluated. Depending on the structure of the molecule, different types of compounds have varying effects on microbial growth effectiveness. 3-(2,4-Dimethoxyphenyl)-1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]prop-2-en-1-one gave the highest antibacterial activity (20 mm) against B. mycoides, whereas 3-(4-chlorophenyl)-1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]prop-2-en-1-one and 1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]-3-(p-tolyl)prop-2-en-1-one had equivalent antibacterial activity (17 mm) against E. coli. The 4-chlorophenyl derivative exhibited the most potent antifungal activity against C. albicans (17 mm). The anti-cancer activity of the prepared compounds was tested against four human cancer cell lines namely A549 (lung carcinoma), MCF7 (human caucasian breast adenocarcinoma), HePG2 (human hepatocellular carcinoma cell line), and BJ1 (normal skin fibroblast). 1-[3,5-Dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]-3-(p-tolyl)prop-2-en-1-one emerged as the most promising compound with IC50 = 44.3 µg/cm3 against A549 and IC50 = 57.9 µg/cm3 against HePG2. Its gene expression, DNA damage values, and DNA fragmentation percentages have been discussed. The expression values of ISL1 and MALL, ASNS and ACLY genes were decreased significantly in treated lung and liver cell lines respectively and positive control compared with negative samples. The expression levels of ISL1 and MALL genes were downregulated in positive control lung cell lines much lower than those in the p-tolyl substituted derivative. The expression levels of ASNS and ACLY genes were downregulated similar to those in positive control liver cell lines. The DNA damage values and DNA fragmentation percentages were increased significantly (P < 0.01) in the treated lung and liver sample compared with the negative control.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
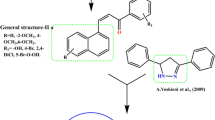
Chalcones are prominent secondary metabolites that exhibit a variety of biological activities that include anti-inflammatory [1,2,3], antiviral [4], antiplatelet [5], antibacterial [6], antimalarial [7], analgesic [8], antioxidant [1, 9], and anti-cancer agents [9,10,11,12]. The existence of active α,β-unsaturated ketone groups in chalcone derivatives is believed to be responsible for their biological activity. Furthermore, nitrogen-containing heterocyclic compounds such as pyrazoles with two nitrogen atoms in their five-membered rings, show considerable pharmaceutical activities [13,14,15,16,17,18,19,20,21,22]. Various drugs have been produced from pyrazole derivatives [23,24,25,26,27]. Besides, the utility of pyrazoles in organic light-emitting diodes applications, liquid crystals, and semiconductors has been intensively studied [28,29,30,31,32]. Based on these facts and in continuation to our research interest directed toward the preparation of bioactive heterocycles [11, 33,34,35], we motivated to synthesize the pyrazolyl–chalcones.
Results and discussion
Chemistry
The starting 1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]ethan-1-one (5) was obtained with good yields following literature procedure [36, 37]. The first step involves the bromination of 1-ethylidene-2-(4-nitrophenyl)hydrazine (1) that affords N-(4-nitrophenyl)acetohydrazonoyl bromide (2). Subsequent reaction of 2 with acetylacetone 4 in the presence of sodium ethoxide as a catalyst leads to the formation of 5. It is proposed that N-(4-nitrophenyl)acetohydrazonoyl bromide (2) in presence of ethoxide converted in to nitrilimine intermediate 3 that adds through 1,3-dipolar cycloaddition to acetylacetone 4 to give the final product (Scheme 1). Claisen–Schmidt condensation of compound 5 with equimolar amounts of arylaldehydes 6 in ethanol in the presence of sodium hydroxide solution resulted in the formation of the pyrazolyl–chalcones 7a–7f (Schemes 2 and 3). The structures of the formed products were confirmed based on inspection of their spectral data.



1H NMR of chalcone 7e indicated the presence of four singlet signals at δ = 2.56, 2.63, 3.87, and 3.91 ppm corresponding to two methyl groups and two methoxy groups. Also, 1H NMR spectrum of compound 7e displayed the two vinyl protons as doublet at 7.33 and 7.83 ppm with coupling constant J = 15.6 Hz (characteristic for the trans configuration of the two olefinic protons). The two doublets at 7.67 and 8.39 ppm with J = 9 Hz are assigned to aromatic protons. The structure of 7e was also confirmed based on 13C NMR that featured 20 signals corresponding to 20 different carbon atoms.
Antimicrobial activity
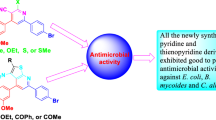
The antimicrobial activity of the synthesized compounds against Gram-negative and Gram-positive bacteria, as well as non-filamentous fungus, was investigated in this work. The antibacterial activity of the tested compounds was evaluated in millimeters by measuring the growth inhibition zone around the holes that contain the individual compounds. Table 1 and Fig. 1 exhibit the antimicrobial effectiveness of various synthesized compounds. The results show that different types of preparations have variable considerable impacts on microbial growth efficacy, which is dependent on the individual synthesized compound. Based on the diameter of the inhibition zone, as shown in Table 1, compound 7e exhibited the greatest antibacterial activity against B. mycoides (20 mm) as a Gram-positive bacteria, while compounds 7b and 7c exhibited equal antibacterial activity (17 mm) against Gram-negative bacteria (E. coli). Furthermore, compound 7b had the greatest antifungal action against C. albicans (17 mm). On the other hand, the lowest antimicrobial activity (14 mm) was observed in the case of compounds 7e and 7f against C. albicans. It is worth noting that all tested compounds shown significant antimicrobial activity against the tested representative microbial strains (Table 1).
Anti-cancer activity
Primary screening
The synthesized compounds 7a–7f were screened against four human cancer cell lines, namely A549 (lung carcinoma), MCF7 (human Caucasian breast adenocarcinoma), HePG2 (human hepatocellular carcinoma cell line), and BJ1 (normal skin fibroblast) at 100 µg/cm3. The results showed that compound 7c possesses anti-cancer activities of 100% and 84% on A549 and HePG2 cell lines, respectively. Most of the compounds showed low activity against A549, MCF7, and HePG2 cell lines as shown in Table 2. So this promising compound 7c was subjected to secondary screening to calculate the IC50 and its selectivity index.
Secondary screening
Concerning IC50, as depicted in Table 3, compound 7c was found as the most promising compound for A549 (lung carcinoma) and HePG2 (human hepatocellular carcinoma cell line) with IC50 = 44.3 and 57.9 µg/cm3, respectively.
Gene expression analysis lung and liver cancer-related genes
Gene expression analysis in lung cancer cell line was performed using lung cancer-related genes, namely ISL1 protein (ISL1) and MAL-like protein (MALL) genes. The results revealed that the expression levels of ISL1 and MALL genes were increased significantly (P < 0.01) in lung cancer cell lines (negative control) compared with treated cell lines (Fig. 2A and B). In contrast, the expression values of ISL1 and MALL genes were decreased significantly (P < 0.05) in treated lung cell lines (7c) and lung cell line treated with 25 nM of doxorubicin (positive control) compared with negative samples. Furthermore, the expression levels of ISL1 and MALL genes were downregulated in doxorubicin-treated lung cell lines much lower than those in 7c.
Alterations in the gene expression level of A ISL1 and B MALL genes in lung cancer cell line; C ASNS and D ACLY genes in liver cancer cell line due to the treatment with the target compound 7c. Tissues with unlike superscript letters were significantly different (P < 0.05). Mean values with different superscript letters (a, b, c, and d) were significantly different (P < 0.05). Negative control, untreated and positive control, doxorubicin
Expression analysis of hepatic cancer-related genes, namely asparagine synthetase (glutamine-hydrolyzing) (ASNS) and ATP citrate lyase (ACLY) genes was performed in liver cancer cell lines. The results revealed that the expression levels of ASNS and ACLY genes were increased significantly (P < 0.01) in liver cancer cell line (negative control) compared with treated cell lines (Fig. 2C and D). In contrast, the expression values of ASNS and ACLY genes were decreased significantly (P < 0.05) in treated (7c) and doxorubicin-treated cell line (positive control) compared with negative control. Furthermore, the expression levels of ASNS and ACLY genes were downregulated in 7c similar to those in doxorubicin-treated cell lines.
DNA damage in lung and liver cell lines using the comet assay
The DNA damage in lung cancer cell lines was determined using comet assay as shown in Table 4. The results showed that untreated cancer cell lines (negative control) samples compared with treated cell lines. However, the DNA damage values were increased significantly (P < 0.01) in treated samples (7c) and doxorubicin-treated cell lines (positive control). Additionally, the highest values of DNA damage were observed in 7c much more than those in doxorubicin-treated cell lines.
Moreover, the comet assay results for the treated liver cancer cell lines are presented in Table 5. The untreated cancer cell lines (negative control) showed significant decrease (P < 0.05) in DNA damage values compared with treated cell lines. In contrary, the DNA damage values were increased significantly (P < 0.01) in treated liver cells (7c) and doxorubicin-treated cell lines (positive control). Furthermore, the highest values of DNA damage were observed in 7c treated cell lines much more than those in positive control cell line.
DNA fragmentation in lung and liver cancer cell lines
The rate of DNA fragmentation determined in lung cancer cell lines is summarized in Table 6 and Fig. 3A. The results found that untreated cancer cell lines (negative control) samples of lung cancer cell lines exhibited a significant decrease (P < 0.01) in DNA fragmentation rates compared with those in treated samples (7c) and doxorubicin-treated cell lines (positive control). However, the DNA fragmentation values were increased significantly (P < 0.01) in treated sample compared with untreated cancer cell lines (negative control).
Furthermore, the effect of the treatments on the percentage of DNA fragmentation in the liver cancer cell lines was investigated (Table 7 and Fig. 3B). The results found that untreated cancer cell lines (negative control) showed significant decrease (P < 0.01) in DNA fragmentation rates compared with those in treated samples (7c) and doxorubicin-treated cell lines (positive control). However, the DNA fragmentation values were increased significantly (P < 0.01) in treated cell compared with negative control. Moreover, the highest values of DNA fragmentation value were observed in 7c much more than those in positive liver cancer cell lines.
Conclusion
In conclusion, using the Claisen–Schmidt condensation reaction, pyrazolyl–chalcone derivatives were prepared by condensation of 5-acetylpyrazole derivatives with the corresponding aldehydes. The antimicrobial activity of various synthesized compounds against Gram-negative and Gram-positive bacteria, as well as non-filamentous fungus, was demonstrated. Depending on the specific structure, different compounds have varying significant effects on microbial growth efficacy. The antibacterial activity of the studied compounds was greater than those of the positive controls employed, suggesting that these compounds might be used to limit microbial spread in future. The expression levels of ISL1 and MALL genes were downregulated in positive control lung cell lines much lower than those in 7c. But, the expression levels of ASNS and ACLY genes were downregulated in 7c similar to those in positive control liver cell lines. The DNA damage values and DNA fragmentation percentages were increased significantly (P < 0.01) in the treated lung and liver (7c) sample compared with the negative control.
Experimental
Melting points were measured with a Stuart melting point apparatus. The IR spectra were recorded using a FT-IR Bruker Vector 22 spectrophotometer as KBr pellets. The 1H and 13C NMR spectra were recorded in CDCl3 as solvent on Varian Gemini NMR spectrometer at 300 MHz using TMS as internal standard. Chemical shifts are reported as δ values in ppm. Mass spectra were recorded with a Shimadzu GCMS–QP–1000 EX mass spectrometer in EI (70 eV) model. The elemental analyses were performed at the Microanalytical Center, Cairo University. 1-Ethylidene-2-(4-nitrophenyl)hydrazine (1) [36], N-(4-nitrophenyl)acetohydrazonoyl bromide (2) [48], and 1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]ethan-1-one (5) [36] were prepared using the reported procedures.
Synthesis of 1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]prop-2-en-1-one derivatives 7a–7f
To a stirred solution of 1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]ethan-1-one (5, 1.0 mmol, 259 mg) and the appropriate aldehyde 6 (1.0 mmol, 106 mg for 6a, 140 mg for 6b, 120 mg for 6c, 136 mg for 6d, 166 mg for 6e, 196 mg for 6f) in 30 cm3 ethanol, 5 cm3 sodium hydroxide solution (20%) was added and reaction mixture was stirred for 6 h at room temperature and left overnight. The resulting solid product that precipitated was filtered, washed with water and crystallized from a suitable solvent to give the corresponding 1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]prop-2-en-1-one derivatives 7a–7f. The purity of the synthesized compounds was tested by TLC that indicated the presence of one compound in each case.
1-[3,5-Dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]-3-phenylprop-2-en-1-one (7a, C20H17N3O3)
Yellow crystals (crystallized from acetonitrile); m.p.: 176–178 °C; yield: 64%; IR: \(\overline{\nu }\) = 1654 (C=O) cm−1; 1H NMR (300 MHz, CDCl3): δ = 2.58 (s, 3H, CH3), 2.64 (s, 3H, CH3), 7.22 (d, 1H, vinyl-H, J = 15.3 Hz), 7.42–7.74 (m, 8H, Ar–H and vinyl-H), 8.37 (d, 2H, Ar–H, J = 8.7 Hz) ppm; 13C NMR (75 MHz, CDCl3): δ = 13.3, 14.8, 122.1, 124.7, 125.2, 125.4, 128.2, 128.9, 130.5, 134.6, 143.4, 143.5, 143.6, 146.6, 150.7, 187.0 ppm; MS (EI, 70 eV): m/z (%) = 347 (M+, 100), 270 (51.24), 198 (25.03), 103 (28.73), 77 (48.54).
3-(4-Chlorophenyl)-1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]prop-2-en-1-one (7b, C20H16ClN3O3)
Yellow crystals (crystallized from DMF); m.p.: 192–194 °C; yield: 62%; IR: \(\overline{\nu }\) = 1655 (C=O) cm−1; 1H NMR (300 MHz, CDCl3): δ = 2.57 (s, 3H, CH3), 2.64 (s, 3H, CH3), 7.18 (d, 1H, vinyl-H, J = 15.6 Hz), 7.26 (d, 2H, Ar–H, J = 9.3 Hz), 7.39 (d, 2H, Ar–H, J = 8.7 Hz), 7.53 (d, 2H, Ar–H, J = 8.7 Hz), 7.63, 7.69 (d, 1H, vinyl-H, J = 15.9 Hz), 8.40 (d, 2H, Ar–H, J = 8.7 Hz) ppm; 13C NMR (75 MHz, CDCl3): δ = 13.3, 14.7, 122.3, 124.7, 125.4, 125.7, 129.2, 129.4, 133.0, 136.4, 142.2, 143.3, 144.1, 146.8, 151.5, 186.9 ppm; MS (EI, 70 eV): m/z = 381 (M+).
1-[3,5-Dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]-3-(p-tolyl)prop-2-en-1-one (7c, C21H19N3O3)
Yellow crystals (crystallized from ethanol); m.p.: 156–158 °C; yield: 62%; IR: \(\overline{\nu }\) = 1654 (C=O) cm−1; 1H NMR (300 MHz, CDCl3): δ = 2.41 (s, 3H, CH3), 2.57 (s, 3H, CH3), 2.63 (s, 3H, CH3), 7.16 (d, 1H, vinyl-H, J = 15.9 Hz), 7.25 (d, 2H, Ar–H, J = 7.8 Hz), 7.53 (d, 2H, Ar–H, J = 7.8 Hz), 7.66, 7.72 (d, 1H, vinyl-H, J = 15.9 Hz), 7.68 (d, 2H, Ar–H, J = 9 Hz), 8.36 (d, 2H, Ar–H, J = 9 Hz) ppm; 13C NMR (75 MHz, CDCl3): δ = 13.2, 14.7, 21.4, 122.2, 124.4, 124.6, 125.2, 128.2, 129.6, 131.8, 141.0, 143.4, 143.5, 143.6, 146.5, 150.6, 187.1 ppm; MS (EI, 70 eV): m/z (%) = 361 (M+, 100), 346 (24.26), 270 (28.81), 198 (15.59), 115 (19.45).
1-[3,5-Dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]-3-(4-methoxyphenyl)prop-2-en-1-one (7d, C21H19N3O4)
Yellow crystals (crystallized from acetonitrile); m.p.: 174–176 °C; yield: 64%; IR: \(\overline{\nu }\) = 1651 (C=O) cm−1; 1H NMR (300 MHz, CDCl3): δ = 2.57 (s, 3H, CH3), 2.63 (s, 3H, CH3), 3.87 (s, 3H, OCH3), 6.94 (d, 2H, Ar–H, J = 9 Hz), 7.14 (d, 1H, vinyl-H, J = 15.6 Hz), 7.59 (d, 2H, Ar–H, J = 8.7 Hz), 7.65 (d, 2H, Ar–H, J = 8.7 Hz), 7.61, 7.71 (d, 1H, vinyl-H, J = 15.6 Hz), 8.37 (d, 2H, Ar–H, J = 9 Hz) ppm; 13C NMR (75 MHz, CDCl3): δ = 13.2, 14.6, 55.3, 114.4, 115.9, 123.1, 124.7, 125.4, 127.2, 130.1, 143.4, 143.7, 145.2, 146.7, 151.4, 161.6, 187.4 ppm; MS (EI, 70 eV): m/z = 377 (M+).
3-(2,4-Dimethoxyphenyl)-1-[3,5-dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]prop-2-en-1-one (7e, C22H21N3O5)
Yellow crystals (crystallized from acetonitrile); m.p.: 172–174 °C; yield: 60%; IR: \(\overline{\nu }\) = 1643 (C=O) cm−1; 1H NMR (300 MHz, CDCl3): δ = 2.56 (s, 3H, CH3), 2.63 (s, 3H, CH3), 3.87 (s, 3H, OCH3), 3.91 (s, 3H, OCH3), 6.49–6.57 (m, 2H, Ar–H), 7.33 (d, 1H, vinyl-H, J = 15.6 Hz), 7.50 (d, 1H, Ar–H, J = 8.4 Hz), 7.67 (d, 2H, Ar–H, J = 9 Hz), 7.83 (d, 1H, vinyl-H, J = 15.6 Hz), 8.39 (d, 2H, Ar–H, J = 9 Hz) ppm; 13C NMR (75 MHz, CDCl3): δ = 13.2, 14.6, 55.4, 55.4, 98.3, 105.3, 116.7, 122.4, 124.0, 124.6, 125.1, 131.3, 139.5, 143.1, 143.7, 146.4, 150.6, 160.2, 162.9, 187.8 ppm; MS (EI, 70 eV): m/z (%) = 407 (M+, 9.22), 104 (100), 90 (32.30), 79 (93.63), 63 (20.24).
1-[3,5-Dimethyl-1-(4-nitrophenyl)-1H-pyrazol-4-yl]-3-(3,4,5-trimethoxyphenyl)prop-2-en-1-one (7f, C23H23N3O6)
Yellow crystals (crystallized from acetonitrile); m.p.: 112–114 °C; yield: 61%; IR: \(\overline{\nu }\) = 1647 (C=O) cm−1; 1H NMR (300 MHz, CDCl3): δ = 2.55 (s, 3H, CH3), 2.62 (s, 3H, CH3), 3.91 (s, 3H, OCH3), 3.92 (s, 6H, 2OCH3), 6.83 (s, 2H, Ar–H), 7.13 (d, 1H, vinyl-H, J = 15.6 Hz), 7.62 (d, 1H, vinyl-H, J = 15.6 Hz), 7.70 (d, 2H, Ar–H, J = 9 Hz), 8.39 (d, 2H, Ar–H, J = 9 Hz) ppm; 13C NMR (75 MHz, CDCl3): δ = 13.3, 14.4, 55.9, 60.8, 105.4, 121.4, 124.6, 124.9, 125.1, 125.2, 140.2, 142.9, 143.5, 143.9, 146.5, 153.3, 155.9, 187.7 ppm; MS (EI, 70 eV): m/z (%) = 437 (M+, 100), 407 (10.03), 198 (27.53), 181 (44.70), 117 (21.23), 76 (32.02).
Antimicrobial evaluation
Bacillus mycoides, Escherichia coli, and Candida albicans were employed as Gram-positive and -negative bacterial strains, respectively, as well as a non-filamentous fungus strain, to evaluate the antimicrobial efficacy of synthesized compounds. B. mycoides, E. coli, and C. albicans were grown and kept on modified nutrient agar medium slants. The modified medium contained (in g/dm3): 0.5 glucose, 0.25 NaCl, 3.0 peptone, 1.5 yeast extract, 1.5 meat extract, and 20.0 agar. Before autoclaving, the pH was adjusted to 7.0. The generated microbial cultures were maintained on agar slants at 4 °C after they had fully matured. Seeded microbial strains were cultivated on nutrient agar medium (70,148 Nutrient agar, Fluka, Spain) with the following components (g/dm3) for antimicrobial testing at 37 °C using the agar diffusion technique: meat extract (1.0 g), yeast extract (2.0 g), peptone (5.0 g), sodium chloride (5.0 g), and agar (15.0 g). The pH of the nutrient agar medium was adjusted to 7.0 after suspending 28 g of ready medium in 1.0 L of distilled water. The medium was then autoclaved for 15 min at 121 °C and 1.5 atmospheres to sterilize it. [38] All of the chemicals utilized were of high purity and analytical grade.
Agar diffusion technique for antibacterial activity determination
The microbial cultures were inoculated in the nutritional agar medium (70,148 Nutrient agar, Fluka) using 100 mm3 of re-suspended overnight culture at 37 °C (1 × 107 CFU/100 mm3) to assess the effectiveness of the synthesized compounds as antimicrobial agents. After injecting the necessary microbial inoculums into the liquefied media at about 45 °C, the nutrient agar medium was poured onto Petri plates. After the plates had solidified at room temperature, holes of 12 mm were made using a sterile cork borer, and then 100 mm3 of dissolved compounds (10 mg/cm3) in DMSO were applied to the holes prepared in Petri plates. Culture test plates were incubated at 37 °C overnight. The AATCC Test Method was used to measure the generated inhibitory zones [39]. Tobramycin (10 mcg) and gentamicin (10 mcg) were used as positive controls [40].
Anti-cancer evaluation
Cell viability was assessed by the mitochondrial-dependent reduction of yellow MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) to purple formazan (Mosmann,1983).
All the following procedures were done in a sterile area using a Laminar flow cabinet biosafety class II level (Baker, SG403INT, Sanford, ME, USA). Cells were suspended in DMEM-F12 medium (for A549, MCF7 and HePG2) besides one normal cell line (BJ1), 1% antibiotic–antimycotic mixture (10,000 U/cm3 potassium penicillin, 10,000 µg/cm3 streptomycin sulfate, and 25 µg/cm3 amphotericin B) and 1% L-glutamine at 37 °C under 5% CO2.
Cells were cultured for 10 days, then seeded at concentration of 10 × 103 cells/well in fresh complete growth medium in 96-well microtiter plastic plates at 37 °C for 24 h under 5% CO2 using a water-jacketed carbon dioxide incubator (Sheldon, TC2323, Cornelius, OR, USA). Media were aspirated, fresh medium (without serum) was added and cells were incubated either alone (negative control) or with different concentrations of sample to give a final concentration of (100–50–25–12.5–6.25–3.125–0.78 and 1.56 ug/cm3). After 48 h of incubation, medium was aspirated, 40 mm3 MTT salt (2.5 μg/cm3) was added to each well and incubated for further four hours at 37 °C under 5% CO2. To stop the reaction and dissolving the formed crystals, 200 mm3 of 10% sodium dodecyl sulfate (SDS) in deionized water was added to each well and incubated overnight at 37 °C. A positive control which composed of 100 µg/cm3 was used as a known cytotoxic natural agent who gives 100% lethality under the same conditions [41].
The absorbance was then measured using a microplate multi-well reader (Bio-Rad Laboratories Inc., model 3350, Hercules, California, USA) at 595 nm and a reference wavelength of 620 nm. A statistical significance was tested between samples and negative control (cells with vehicle) using independent t-test by SPSS 11 program. DMSO is the vehicle used for dissolution of plant extracts and its final concentration on the cells was less than 0.2%. The percentage of change in viability was calculated according to the formula:
A probit analysis was carried for IC50 determination using SPSS 11 program. In the present study, the degree of selectivity of the synthetic compounds is expressed as: SI = IC50 of pure compound in a normal cell line/IC50 of the same pure compound in cancer cell line, where IC50 is the concentration required to kill 50% of the cell population.
Gene expression analysis
Quantitative real-time PCR method
RNA isolation and reverse transcription (RT) reaction
RNeasy Mini Kit (Qiagen, Hilden, Germany) supplemented with DNaseI (Qiagen) digestion step was used to isolate total RNA from Lung and liver cancer cell lines according to the manufacturer’s protocol. Isolated total RNA was treated with one unit of RQ1 RNAse-free DNAse (Invitrogen, Germany) to digest DNA residues, re-suspended in DEPC-treated water and quantified photospectrometrically at 260 nm. Purity of total RNA was assessed by the 260/280 nm ratio which was between 1.8 and 2.1. Additionally, integrity was assured with ethidium bromide-stain analysis of 28S and 18S bands by formaldehyde-containing agarose gel electrophoresis. Aliquots were used immediately for reverse transcription (RT), otherwise they were stored at − 80 °C.
Complete Poly(A)+ RNA isolated from lung and liver cell lines was reverse-transcribed into cDNA in a total volume of 20 mm3 using RevertAid™ First-Strand cDNA Synthesis Kit (Fermentas, Germany). An amount of total RNA (5 µg) was used with a master mix. The master mix consisted of 50 mM MgCl2, 10 × RT buffer (50 mM KCl; 10 mM Tris–HCl; pH 8.3), 10 mM of each dNTP, 50 µM oligo-dT primer, 20 IU ribonuclease inhibitor (50 kDa recombinant enzyme to inhibit RNase activity) and 50 IU MuLV reverse transcriptase. The mixture of each sample was centrifuged for 30 s at 1000 g and transferred to the thermocycler. The RT reaction was carried out at 25 °C for 10 min, followed by 1 h at 42 °C, and finished with a denaturation step at 99 °C for 5 min. Afterward, the reaction tubes containing RT preparations were flash-cooled in an ice chamber until being used for cDNA amplification through quantitative real-time polymerase chain reaction (qRT-PCR).
Real-Time PCR (qPCR)
Determination of the lung and liver cell line cDNA copy number was carried out using StepOne™ Real-Time PCR system from applied Biosystems (Thermo Fisher Scientific, Waltham, MA USA). PCR reactions were set up in 25 mm3 reaction mixtures containing 12.5 mm3 1 × SYBR® Premix Ex TaqTM (TaKaRa, Biotech. Co. Ltd.), 0.5 mm3 0.2 μM sense primer, 0.5 mm3 0.2 μM antisense primer, 6.5 mm3 distilled water, and 5 mm3 of cDNA template. The reaction program was allocated to 3 steps. First step was at 95 °C for 3 min. Second step consisted of 40 cycles in which each cycle divided to 3 steps: (a) at 95 °C for 15 s; (b) at 55 °C for 30 s; and (c) at 72 °C for 30 s. The third step consisted of 71 cycles which started at 60 °C and then increased about 0.5 °C every 10 s up to 95 °C. At the end of each sqRT-PCR, a melting curve analysis was performed at 95 °C to check the quality of the used primers. Each experiment included a distilled water control. The sequences of specific primer of the lung (ISL1 and MALL genes [42], and liver (ASNS and ACLY genes [43]) cancer-related genes were designed and listed in Table 8. At the end of each qPCR, a melting curve analysis was performed at 95 °C to check the quality of the used primers. The relative quantification of the target to the reference was determined using the 2−ΔΔCT method [44].
DNA damage using the comet assay
The DNA damage using comet assay was determined using lung and liver cancer cell lines according to Olive et al. [45]. After the trypsin treatment to produce a single cell suspension, approximately 1.5 × 104 cells were embedded in 0.75% low-gelling-temperature agarose and rapidly pipetted onto a pre-coated microscope slide. Samples were lysed for 4 h at 50 °C in 0.5% SDS, 30 mM EDTA, pH 8.0. After rinsing overnight at room temperature in Tris/borate/EDTA buffer, pH 8.0, samples were electrophoresed for 25 min at 0.6 V/cm, then stained with propidium iodide. Slides were viewed using a fluorescence microscope with a CCD camera, and 150 individual comet images were analyzed from each sample for tail moment, DNA content, and percentage DNA in tail. For each sample, about 100 cells were examined to determine the percentage of cells with DNA damage that appear like comets. The non-overlapping cells were randomly selected and were visually assigned a score on an arbitrary scale of 0–3 (i.e., class 0 = no detectable DNA damage and no tail; class 1 = tail with a length less than the diameter of the nucleus; class 2 = tail with length between 1 × and 2 × the nuclear diameter; and class 3 = tail longer than 2 × the diameter of the nucleus) based on perceived comet tail length migration and relative proportion of DNA in the nucleus [46].
DNA fragmentation assay
The DNA fragmentation assay in lung and liver cancer cell lines was performed in concordance with the premises established by Yawata [47] with some modifications. Briefly, after 24 h of exposure of lung and liver cancer cell lines to the tested substances in different Petri dishes (60 × 15 mm, Greiner), the cells were trypsinized, suspended, homogenized in 1 cm3 of medium and centrifuged (10 min at 800 rpm). Low molecular weight genomic DNA was extracted as described in Yawata [47]. Approximately, 1 × 106 cells were plated and treated with the tested substances in various treatments. All the cells (including floating cells) were harvested by trypsinization and washed with Dulbecco`s Phosphate-Buffered Saline. Cells were lysed with the lysis buffer containing 10 mM Tris (pH 7.4), 150 mM NaCl, 5 mM ethylenediaminetetraacetic acid (EDTA), and 0.5% Triton X-100 for 30 min on ice. Lysates were vortexed and cleared by centrifugation at 10,000 g for 20 min. Fragmented DNA in the supernatant was extracted with an equal volume of neutral phenol:chloroform:isoamyl alcohol mixture (25:24:1) and analyzed electrophoretically on 2% agarose gels containing 0.1 μg/cm3 ethidium bromide.
References
Bandgar BP, Gawande SS, Bodade RG, Gawande NM, Khobragade CN (2009) Bioorg Med Chem 17:8168
Bekhit AA, Abdel-Aziem T (2004) Bioorg Med Chem 12:1935
Hsieh H-K, Tsao L-T, Wang J-P, Lin C-N (2000) J Pharm Pharmacol 52:163
Onyilagha JC, Malhotra B, Elder M, French CJ, Towers GHN (1997) Can J Plant Pathol 19:133
Lin C-N, Hsieh H-K, Ko H-H, Hsu M-F, Lin H-C, Chang Y-L, Chung M-I, Kang J-J, Wang J-P, Teng C-M (2001) Drug Dev Res 53:9
Asiri AM, Khan SA (2011) Molecules 16:523
Li R, Kenyon GL, Cohen FE, Chen X, Gong B, Dominguez JN, Davidson E, Kurzban G, Miller RE, Nuzum EO, Rosenthal PJ, McKerrow JH (2002) J Med Chem 38:5031
Heidari MR, Foroumadi A, Amirabadi A, Samzadeh-Kermani A, Azimzadeh BS, Eskandarizadeh A (2009) Ann NY Acad Sci 1171:399
Shenvi S, Kumar K, Hatti KS, Rijesh K, Diwakar L, Reddy GC (2013) Eur J Med Chem 62:435
Sashidhara KV, Kumar A, Kumar M, Sarkar J, Sinha S (2010) Bioorg Med Chem Lett 20:7205
Fathi EM, Sroor FM, Mahrous KF, Mohamed MF, Mahmoud K, Emara M, Elwahy AHM, Abdelhamid IA (2021) Chem Select 6:6202
Mohamed MF, Sroor FM, Ibrahim NS, Salem GS, El-Sayed HH, Mahmoud MM, Wagdy M-AM, Ahmed AM, Mahmoud A-AT, Ibrahim SS, Ismail MM, Eldin SM, Saleh FM, Hassaneen HM, Abdelhamid IA (2020) Invest New Drugs 39:98
Karrouchi K, Radi S, Ramli Y, Taoufik J, Mabkhot Y, Al-aizari F, Ansar MH (2018) Molecules 23:134
Khan MF, Alam MM, Verma G, Akhtar W, Akhter M, Shaquiquzzaman M (2016) Eur J Med Chem 120:170
Iglesias-Arteaga MA, Pérez-Fernández R, Goya P, Elguero J (2014) ARKIVOC 2014:233
Fustero S, Sánchez-Roselló M, Barrio P, Simón-Fuentes A (2011) Chem Rev 111:6984
Fustero S, Simón-Fuentes A, Sanz-Cervera JF (2009) Org Prep Proced Int 41:253
Gomha SM, Edrees MM, Faty RAM, Muhammad ZA, Mabkhot YN (2017) Chem Cent J 11:37
Altalbawy F (2013) Int J Mol Sci 14:2967
Zagni C, Citarella A, Oussama M, Rescifina A, Maugeri A, Navarra M, Scala A, Piperno A, Micale N (2019) Int J Mol Sci 20:945
Bennani FE, Doudach L, Cherrah Y, Ramli Y, Karrouchi K, Ansar Mh, Faouzi MEA (2020) Bioorg Chem 97:103470
Othman IMM, Alamshany ZM, Tashkandi NY, Gad-Elkareem MAM, Anwar MM, Nossier ES (2021) Bioorg Chem 114:105078
Ansari A, Ali A, Asif M, Shamsuzzaman S (2017) New J Chem 41:16
Surendra Kumar R, Arif IA, Ahamed A, Idhayadhulla A (2016) Saudi J Biol Sci 23:614
Jakob KS, Ganguly S (2016) Int J Pharm Pharm Sci 8:75
Alam R, Wahi D, Singh R, Sinha D, Tandon V, Grover A, Rahisuddin (2016) Bioorg Chem 69:77
Bekhit AA, Hassan AMM, Abd El Eazik HA, El-Miligy MMM, El-Agroudy EJ, Bekhit AE-DA (2015) Eur J Med Chem 94:30
Karcı F, Karcı F, Demirçalı A, Yamaç M (2013) J Mol Liq 187:302
Burschka J, Kessler F, Nazeeruddin MK, Grätzel M (2013) Chem Mater 25:2986
Chou P-T, Chi Y (2007) Chem Eur J 13:380
Kauhanka UM, Kauhanka MM (2006) Liq Cryst 33:121
Wang M, Zhang J, Liu J, Xu C, Ju H (2002) J Lumin 99:79
Tantawy MA, Sroor FM, Mohamed MF, El-Naggar ME, Saleh FM, Hassaneen HM, Abdelhamid IA (2020) Anti-Cancer Agents Med Chem 20:70
Abdelhamid IA, Abdelmoniem AM, Sroor FM, Ramadan MA, Ghozlan SAS (2020) Synlett 31:895
Sroor FM, Basyouni WM, Tohamy WM, Abdelhafez TH, El-awady MK (2019) Tetrahedron 75:130749
Shawali AS, Hassaneen, HM (1976) Indian J Chem 7: 549
Stanovnik B, Svete J (2002) Product class 1: pyrazoles. In: Neier R, Bellus D (eds) Science of synthesis 2: hetarenes, vol 12. Thieme, Stuttgart, New York
Abouelnaga AM, Meaz TM, Othman AM, Ghazy RA, El Nahrawy AM (2020) SILICON 13:623
Mohamed SAA, El-Sakhawy M, Nashy ELSHA, Othman AM (2019) Int J Biol Macromol 136:774
Sroor FM, Othman AM, Tantawy MA, Mahrous KF, El-Naggar ME (2021) Bioorg Chem 112:104953
Thabrew MI, Hughes RD, McFarlane IG (1997) J Pharm Pharmacol 49:1132
Watanabe T, Miura T, Degawa Y, Fujita Y, Inoue M, Kawaguchi M, Furihata C (2010) Cancer Cell Int 10:2
Saur D, Seidler B, Schneider G, Algül H, Beck R, Senekowitsch-Schmidtke R, Schwaiger M, Schmid RM (2005) Gastroenterology 129:1237
Yang Q, Feng MH, Ma X, Li HC, Xie W (2017) Oncol Lett 14:6071
Olive PL, Banáth JP, Durand RE (2012) Radiat Res 178:AV35
Collins A, Dusinska M, Franklin M, Somorovska M, Petrovska H, Duthie S, Fillion L, Panayiotidis M, Raslova K, Vaughan N (1997) Environ Mol Mutagen 30:139
Yawata A, Adachi M, Okuda H, Naishiro Y, Takamura T, Hareyama M, Takayama S, Reed JC, Imai K (1998) Oncogene 16:2681
Hegarty AF, Cashman MP, Scott FL (1972) J Chem Soc Perkin Trans 2:1381
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kamel, M.G., Sroor, F.M., Othman, A.M. et al. Structure-based design of novel pyrazolyl–chalcones as anti-cancer and antimicrobial agents: synthesis and in vitro studies. Monatsh Chem 153, 211–221 (2022). https://doi.org/10.1007/s00706-021-02886-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00706-021-02886-5