Abstract
A colorimetric assay based on polydiacetylenes (PDA) nano-liposomes is reported for facile and sensitive detection of alkaline phosphatase (ALP) activity. The critical basis of this method is that the interaction of pyridoxal phosphate (PLP) with nitrogenous group functionalized PDA nano-liposomes induces distinct blue-to-red color changes of PDA nano-liposomes. In the presence of ALP, as a nature substrate, PLP is enzymatically hydrolyzed to form pyridoxal, which cannot interact with PDA nano-liposomes. As a result, the concentration of PLP is reduced and the color change of PDA nano-liposomes is retarded, which is associated with ALP level. Under optimal conditions, the proposed method showed good linear relationship with ALP activity in the range 10–200 U/L with a limit of detection of 2.8 U/L. The detection process could be vividly observed with the naked eye. Additional attempts by using the method for the evaluation of inhibitor efficiency were also achieved with satisfying results. The method was further challenged with real human serum samples, showing consistent results when compared with a commercial standard assay kit. Such simple and easy-to-use approach may provide a new alternative for clinical and biological detection of ALP.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alkaline phosphatase (ALP) is an important biological enzyme that is widely distributed in variety of organisms, playing a vital role in the phosphorous metabolism by catalyzing the dephosphorylation process on substrates containing monophosphate esters [1]. In humans, ALP is actively involved in various important biological processes such as molecule transportation, intracellular regulation, and bone mineralization [2]. It has been shown that the abnormal of ALP activity is associated with a number of diseases, such as hepatobiliary diseases, bone-related diseases, cancers, and diabetes [3]. As such, ALP is one of the widely used clinical bio-markers for diagnosis and monitoring of various diseases [4]. It is therefore in great demand methods that can facilely, sensitively, and selectively assay ALP activity.
Currently, the most widely used method for detection of ALP activity in clinical practice and biological studies is based on classical chromogenic reaction, of which colorless p-nitrophenyl phosphate (p-NPP) is used as the enzyme substrate and could be enzymatically hydrolyzed to yellow p-nitrophenol (p-NP) by ALP [5]. Although this method can assay ALP activity with the superiority of short reacting time and mature automatable procedure, the instability of p-NPP to light and non-specific hydrolysis make this approach uncertain which may produce false positive results [6]. To avoid the limitation and improve detection, diversified methods have been developed for ALP activity based on colorimetry [6, 7], fluorometry [8, 9], electrochemistry [10, 11], and surface-enhanced Raman scattering (SERS) [12, 13]. Among them, colorimetric assay remains to be one of the most popular manners for ALP detection owing to its simplicity, unique visual output, and low cost [14, 15]. In the overall view, most of the sensing principles are running with the assistance of the products generated from ALP-mediated dephosphorylation. These products, such as ascorbic acid (AA) hydrolyzed from ascorbic acid 2-phosphate (AAP), typically have strong reducibility that could further regulate the colorimetric responses of the designed probes [6]. Quite a lot of elegant methods have been designed for assaying ALP based on this strategy by using different nanomaterial-based sensing components [16,17,18,19]. However, due to the complicacy of biological samples, some reductive molecules, such as cysteine, glutathione, and dopamine, may coexist in real samples. These substances could disturb the AA-based detection process, thereby producing certain effects on the selectivity and reliability of these methods for ALP activity in clinical practices [6, 20]. Accordingly, the exploration and development of novel colorimetric methods with new sensing principles and strategies for ALP activity still remain attractive and challenging.
As a representative class of conjugated polymers, polydiacetylenes (PDAs) remain as one of the attractive smart materials for sensing applications owing to their unique optical and structural features [21]. The optical features stem from the special alternating ene-yne conjugated polymer chains of PDAs, which are formed through UV light–induced 1,4-addition reaction of the well-aligned diacetylene monomers. The initially formed conjugated backbone of PDAs is typically in a metastable state with the color of PDAs blue. Under proper external stimulates, the conformation of the conjugated backbone could change into a more stable state, thus inducing a clear color change of PDAs from blue to red [22]. Additionally, due to the amphipathic nature of most diacetylene monomers, PDAs can be easily self-assembled into films, nanowires, micelles, and nano-liposomes, facilitating their usage in aqueous mediums [23]. These unique properties make PDAs particularly useful in exploitation of colorimetric chemo- and bio-sensors [24, 25]. Despite many PDA sensing system developed for given chemical agents [26], ions [27], nucleic acids [28], proteins [29], and viruses [30], few reports concern on the development of PDA-based colorimetric assays for enzyme activity, possibly due to the lack of effective recognition strategies.
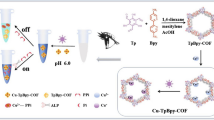
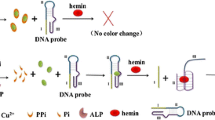
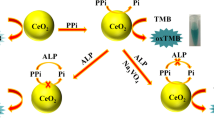
To this end, we provide herein a new sensing strategy for colorimetric assaying of ALP activity with high selectivity and sensitivity based on the specific response of functional PDA nano-liposomes for pyridoxal phosphate (PLP). As illustrated in Scheme 1, two kinds of functional nitrogenous groups of ethylenediamine and imidazolium are incorporated into self-assembly PDA liposome system. The as-prepared PDA nano-liposomes display rapid and distinct blue-to-red color changes in response to PLP due to the strong electrostatic interaction between the positively charged nitrogenous groups of PDA nano-liposomes and the negatively charged phosphate groups of PLP. As a natural substrate of ALP, PLP could be enzymatically hydrolyzed by ALP to generate pyridoxal (PL), which cannot interact with PDA nano-liposomes. It means that the color transition of PDA nano-liposomes for PLP can be impeded and regulated by ALP activity. Thus, a simple PDA-liposome based assay can be established for ALP detection. In addition, when the inhibitor of ALP is introduced into this system, the hydrolysis of ALP for PLP will be inhibited, thereby restoring the color changes of PDA nano-liposomes. Based on these processes, our proposed approach is capable of monitoring of ALP activity and evaluating of inhibitor efficiency in a facile and effective manner, especially avoiding the interferences of reductive species in complex biological samples.
Experimental
Reagents and apparatus
Alkaline phosphatase (ALP) was purchased from Sigma-Aldrich (Saint Louis, MO, USA). 1-(3-Aminopropyl)imidazole, iodomethane, and pyridoxal phosphate (PLP) were purchased from Aladdin (Shanghai, China). N-hydroxysuccinimide (NHS) and 1-(3-dimethylaminopropyl)-3-ethylcarbodiimide hydrochloride (EDC·HCl) were purchased from TCI (Shanghai, China). 2-[4-(2-Hydroxyethyl)piperazin-1-yl]ethanesulfonic acid (HEPES) was purchased from Solarbio (Beijing, China). p-NPP-based assay kit for ALP was obtained from Beyotime Biotechnology (Shanghai, China). Other reagents and solvents were supplied by local commercial suppliers. All reagents were of analytical grade and used as received. All the solutions were prepared with ultrapure water (resistivity ≥ 18.2 MΩ·cm) obtained from a Thermo Scientific water purification system (Waltham, MA, USA).
All the absorption spectra were obtained with a Hitachi U3310 spectrometer (Tokyo, Japan). The chemical structures of the synthesized compounds were characterized by 1H and 13C nuclear magnetic resonance (NMR) spectra using a Bruker Avance 400 MHz/100 MHz spectrometer. Scanning electron microscopy (SEM) was performed on a FEI Verios G4 electron microscope (Hillsboro, OR, USA). Raman spectra were recorded on a Finder One Raman microscope (Zolix, Beijing, China). The Malvern Zetasizer Nano-ZS (Malvern Instruments Co., UK) equipped with a He–Ne laser was used to analyze the size distribution and zeta potential.
Functional PDA liposome preparation
The functionalized diacetylene monomers PCDA-EA and PCDA-AID were synthesized based on previous reported methods [26, 27]. The detailed synthetic routes and NMR spectra can be found in the “Supplementary information.”
Nitrogenous ligand functionalized PDA nano-liposomes were prepared based on a probe sonication method according to the previous studies [29]. Typically, a mixture of PCDA-EA and PCDA-AID with a molar ratio of 1:1 was dissolved in chloroform (1 mL) in a round-bottom flask. Solvent was thoroughly removed by evaporation under vacuum, and then ultrapure water (10 mL) was added to yield a lipid concentration of 1 mM. The resulting suspension was probe sonicated (Scientz92-II, Scientz, Ningbo, China) for 20 min at 80 °C to afford a translucent solution. This solution was allowed to store at 4 °C for 12 h. After being brought to room temperature, the solution was exposed to a hand-held 254-nm UV lamp (1 mW cm−2) for 2 min. The obtained PDA liposome solution with deep blue color was either used freshly as prepared or stored at 4 °C until use.
Colorimetric response of PDA nano-liposomes for PLP
To investigate the colorimetric responses of PDA nano-liposomes for PLP and other anions, various anions (1 mM) were injected into HEPES buffer (10 mM, pH = 7.4) containing PDA nano-liposomes (100 μM), respectively. The resulting mixtures were equilibrated at room temperature for 20 min. Their UV–Vis absorption spectra were then recorded. For concentration-dependent responses of PDA nano-liposomes towards PLP, different concentrations of PLP were added into buffer solution (HEPES, 10 mM, pH = 7.4) containing PDA nano-liposomes (100 μM), respectively. Their absorption spectra were acquired after equilibrium at room temperature for 20 min. To quantify the color change of PDA nano-liposomes, colorimetric response value (CR%) was calculated by the following formula: CR = [P0 – P1]/P0 × 100%, where P = I640/[I640 + I540]. I640 and I540 are the absorption of PDA nano-liposomes at 640 nm and 540 nm, respectively. P0 and P1 are the values before and after the color change.
Detection of ALP activity
For assaying ALP activity, 40 μL of PLP solution (10 mM) and 30 μL of ALP with varying concentrations ranging from 0.01 to 2 U/mL were sequentially added into 830 μL of HEPES buffer (10 mM, pH = 7.4). The resulting samples were incubated at 37 °C for 30 min, 60 min, or 90 min before 100 μL of PDA liposome solution (1 mM) was respectively added into these samples. After equilibrating at room temperature for 20 min, the UV–Vis absorption spectrum of each of the mixtures was recorded. The limit of detection (LOD) was estimated using the 3σ/m criterion, where σ is the standard deviation of the blank or standard deviation of the intercept and m is the slope of the calibration plot [31].
Inhibitor efficiency evaluation
In order to investigate the inhibition effect of EDTA toward ALP, different concentrations of EDTA and ALP (0.3 U/mL) were mixed in HEPES buffer (10 mM, pH = 7.4) at 37 °C for 20 min. Afterwards, PLP (400 μM) was added into the above solutions and resulting samples were incubated at 37 °C for 60 min. The inhibition efficiency (IE) was calculated using a method described previously [31].
Detection of ALP activity in human serum samples
For ALP detection in serum samples, a standard addition test was first performed in HEPES buffer (10 mM, pH = 7.4) containing 1% fetal bovine serum. Spiked samples with different ALP concentrations (0.01, 0.05, 0.10, 0.15, 0.20 U/mL) were prepared and assayed with this approach. To further verify the potential application of the method for ALP in real samples, five human serum samples obtained from volunteers in our lab were divided into two parts. One was analyzed with a standard p-NPP based colorimetric assay, and the other was tested by our proposed method. Specifically, 20 µL of each of the serum samples was added into 160 µL of HEPES buffer (10 mM, pH = 7.4) containing 400 µM of PLP in a 96-well plate. The resulting mixtures were incubated at 37 °C for 60 min. Subsequently, 20 µL of PDA nano-liposomes (final concentration of 100 µM) was added into each sample. After equilibrium at room temperature for 20 min, the absorption at 640 nm and 540 nm was recorded using a microplate reader (Synergy HT, BioTek, USA). CR values were then calculated using abovementioned formula and the ALP levels in unknown serum samples were then estimated using the standard curve method. All tests were repeated three times. All date are shown as mean ± standard deviation (SD).
Results and discussion
Colorimetric responses of PDA nano-liposomes for PLP
It is reported in previous studies that PDA nano-liposomes bearing quaternary ammonium and amino groups show clear color changes towards ATP and pyrophosphate (PPi) due to the strong electrostatic interaction between quaternary ammonium and phosphate groups [27, 32]. This finding inspired us to synthesize two kinds of diacetylene monomers, PCDA-EA and PCDA-AID, in which 10,12-pentacosadiynoic acids (PCDA) were respectively modified with ethanediamine and imidazolium (Scheme 1). PDA nano-liposomes were then prepared by incorporating these two monomers into PDA supermolecules via self-assembly and photo-induced polymerization. We speculated that the functional cationic head groups could strongly interact with phosphates which may ultimately lead to the color changes of PDA nano-liposomes. To verify this hypothesis, colorimetric responses of the as-prepared PDA nano-liposomes for several phosphates and other anions were first investigated. As shown in Fig. 1a, PDA nano-liposomes displayed noticeable color changes from blue to violet or red in the presence of PPi or pyridoxal phosphate (PLP) that could be observed with the naked eye. While other anions including H2PO4− and HPO42− almost could not induce any color changes. From absorption spectra, it was observed that the addition of PLP caused the most obvious spectral changes of PDA nano-liposomes, in which the absorption band at 640 nm distinctly decreased accompanied by the occurrence and dramatic increase of a new absorption band at 540 nm (Fig. 1b). Further quantitative analysis by CR value showed that PLP induced colorimetric response was as high as 67.4%. A much lower CR values of 30.8% was observed for PPi, while less than 10% of CR values were obtained for other anions (Fig. 1c). These results strongly confirm our conjecture, revealing that our nitrogenous ligand-functionalized PDA nano-liposomes could selectively respond to PLP and PPi (particularly PLP) with colorimetric signals.
a Colorimetric responses of PDA nano-liposomes (100 µM) in the presence of various anions (1 mM). b UV–Vis absorption spectra and c CR (%) values of PDA nano-liposomes (100 μM) in response to various anions (1 mM). d UV–Vis absorption spectra and e CR (%) values of PDA nano-liposomes (100 μM) versus concentration of PLP. Inset is a photograph of PDA nano-liposomes after addition of PLP. The relevant species in (c) are: 1 F–, 2 Cl–, 3 Br–, 4 I–, 5 CO32–, 6 HCO3–, 7 SO42–, 8 S2O32–, 9 SO32–, 10 H2PO4–, 11 HPO42–, 12 NO3–, 13 Citric Acid (CA), 14 AcO–, 15 EDTA, 16 PPi, 17 PLP
The concentration-dependent response of PDA nano-liposomes for PLP was subsequently studied to determine the minimum amount of PLP required to induce clear blue-to-red color transition. Before that, detection parameters of pH and the concentration of PDA nano-liposomes were first optimized. As shown in Fig. S2, PDA nano-liposomes showed strong colorimetric responses for PLP in acidic condition. When the pH was high than 8.0, slight color change was observed with the CR value of ~ 20%. Considering acid medium may decrease the activity of ALP [1, 3], a slight basic pH of 7.4 was chosen for further studies. In addition, when the concentration of PDA nano-liposomes was above 30 µM, similar colorimetric responses were obtained (Fig. S2c and 2d). We then fixed the amount of PDA nano-liposome to be 100 µM as the color change of PDA nano-liposomes could be easily identified at this concentration. As the amount of PLP increases, clear color changes from blue to violet and finally to red were vividly observed with the naked eye. At least 400 µM of PLP was needed to induce the distinct color transition (Fig. 1e, inset). This observation could be further confirmed by absorption spectra and CR values (Fig. 1d and e). The absorption shifts from 640 to 540 nm were rapidly occurred when the concentration of PLP in the range of 10 – 400 µM with a maximal CR value of ~ 68.1%. Slow spectral changes with slight increase in CR values were observed when the amount of PLP was over 400 µM. Moreover, the addition of PLP resulted in a rapid response of PDA nano-liposomes whose CR values reached to plateau within 20 min (Fig. S3). For clear visual detection, the concentration of PLP was thus fixed at 400 µM in the follow-up studies.
Quantitative assay of ALP activity
As we described in Scheme 1, ALP could readily catalyze the hydrolysis of PLP to form pyridoxal (PL) and orthophosphate (Pi), thereby removing the effect of PLP to PDA nano-liposomes. Reverse colorimetric response that relies on ALP activity should be observed when ALP is introduced into the sensing system. To evaluate the feasibility of this designed strategy, absorption spectra of PDA nano-liposomes at different conditions were recorded. As shown in Fig. S4, in the presence of PLP, clear band shift from 640 to 540 nm was observed (curve b). However, when PLP was first incubated with ALP and then interacted with PDA nano-liposomes, similar absorption spectrum (curve c) as that of pure PDA nano-liposomes (curve a) was obtained. In addition, there was no noticeable absorption change of PDA nano-liposomes when ALP was added (curve d), revealing the negligible effect of ALP on PDA nano-liposomes. These observations thus indicate our approach is highly feasible for the detection of ALP activity.
Further studies were then performed to investigate the analytical performance of this assay for ALP activity. To achieve optimal detection, the effects of pH and temperature on ALP activity were first optimized. It was found that the highest enzyme activity was obtained in HEPES buffer with pH of 7.4 at 37 °C (Fig. S5) in our sensing system. In the presence of ALP, PLP is enzymatically hydrolyzed to produce PL, which then removes the colorimetric response of PLP to PDA nano-liposomes. Therefore, the optical changes of PDA nano-liposomes are highly dependent on the incubation time of PLP with ALP. We set the incubation time of PLP with ALP at 30, 60, and 90 min. The colorimetric responses of PDA nano-liposomes at these three points in time were respectively recorded. As shown in Figs. 2a, c, and S4, with the increasing of ALP activity, the absorption at 640 nm increased accompanied by the decrease of the absorption at 540 nm at all three time points. Quantified analysis by [1-CR] values showed that the colorimetric response of PDA nano-liposomes had good correlation with ALP activity. Good linear relationships were obtained with the ALP concentration in the range of 0.10–0.40 U/mL (R2 = 0.9943), 0.01–0.30 U/mL (R2 = 0.9821), and 0.01–0.20 U/mL (R2 = 0.9243) at the incubation time of 30, 60, and 90 min, respectively (Figs. 2b, d and S6). The limit of detection (LOD) for ALP (based on 3σ/m criterion) was determined as 0.0092 U/mL, 0.0042 U/mL, and 0.0028 U/mL for the incubation time at 30, 60, and 90 min, respectively. The LOD of the proposed method is comparable with some other colorimetric and fluorometric methods (Table 1). More importantly, as the ALP activity in healthy adults is about 0.04–0.19 U/mL, this method holds great potential to assay ALP activity with high accuracy in real samples. Apart from its high sensitivity, this method can also realize the detection through the naked eye by vivid color change from red to blue (Fig. 2e), which may be particularly useful for rapid and on-site analysis of ALP activity.
a and c UV–Vis absorption spectra of the PDA liposome–based sensing system versus different activities of ALP at the incubation time of 30 min and 60 min, respectively. b and d Plot of [1-CR] (%) values of the PDA liposome–based sensing system versus various activities of ALP at the incubation time of 30 and 60 min, respectively. Insets are linear curves of [1-CR] (%) values versus ALP activities. e Visual detection of ALP activity using the proposed method
Selectivity studies were further performed by testing the colorimetric responses of PDA nano-liposomes towards other proteins, metal ions as well as amino acids. As shown in Fig. 3, all these interferents almost had no influence on PDA nano-liposomes with the CR values below 6% (Fig. 3a and b). When they were incubated with PLP instead of ALP, obvious absorption changes with CR values around 60% were recorded, revealing that they had no effects on the colorimetric response of PDA nano-liposomes for PLP (Fig. 3c and d). These results thus indicate that our approach is highly selective for ALP.
a UV–vis absorption spectra and b CR (%) values of PDA nano-liposomes in the presence of various species. c UV–vis absorption spectra and d CR (%) values of PDA nano-liposomes in response to PLP solutions in which they are pre-incubated with various species. The relevant species in (b) and (d) are 1 GOX, 2 GSH, 3 THR, 4 TRY, 5 BSA, 6 K+, 7 Na+, 8 Mg2+, 9 Fe3+, 10 Zn2+, 11 Phe, 12 Ala, 13 Gly, 14 Met, 15 Lys, 16 Leu, 17 Pro, 18 Ser, 19 Asn, 20 His, 21 PLP for (b) and ALP for (d)
Mechanism of detection for PLP and ALP
We attributed the excellent selectivity of this method for ALP to the two key features of the detection system: (i) strong and specific interactions of PDA nano-liposomes with PLP, which could eventually induce the distinct color changes, and (ii) the enzymatic hydrolysis of PLP by ALP, which could remove the effect of PLP on PDA nano-liposomes. To verify the detection mechanism described above, the electrostatic interactions of PDA nano-liposomes at different conditions were first investigated through zeta potential measurement. As shown in Fig. S7, blank PDA nano-liposomes displayed a zeta potential of 29.27 mV. It is worth noting that the zeta potential of PDA nano-liposomes could keep constant around 30 mV along with the storage time, revealing the high stability of PDA nano-liposomes (Fig. S8). In the presence of PLP, the value sharply decreased to 5.71 mV, clearly indicating an electrostatic interaction between PLP and PDA nano-liposomes. However, when PLP was hydrolyzed under the catalysis of ALP, the zeta potential of PDA nano-liposomes slightly decreased from 29.27 to 23.83 mV, similar to the value when ALP was added into PDA nano-liposomes. The decreased zeta potential can be ascribed to the weak electrostatic interactions between PDA nano-liposomes and ALP, as negative charged nature of most proteins in physiological condition. In addition, other analytes, such as GSH, K+, and Phe, had little effects on zeta potentials of PDA nano-liposomes. It was worth noting that the addition of H2PO4− and THR could also induce noticeable decreases of zeta potential to 20.13 mV and 15.0 mV, respectively. These phenomena suggest that PDA nano-liposomes used in this study could also produce electrostatic interactions with other negative charged substances. Nevertheless, the optical properties of PDA nano-liposomes were not altered, as confirmed in Fig. 3a.
To gain insight into structural origins of the observed zeta potential changes, the size distributions and morphologies of PDA nano-liposomes in different conditions were characterized with DLS and SEM. As shown in Fig. 4 and S9, the as-prepared PDA nano-liposomes possess a spherical structure (Fig. 4b) with the size around 61.95 nm (Fig. 4a). The size of PDA nano-liposomes dramatically increased to 1112.25 nm with massive aggregations observed on SEM images in the presence of PLP (Fig. 4c). However, with the introduction of ALP, the size of PDA nano-liposomes slightly increased to 130.89 nm, and no obvious aggregation was observed (Fig. S10). These changes were quite similar to that of PDA nano-liposomes containing ALP alone. Interestingly, other substances including H2PO4−, GSH, K+, Phe, and THR all caused slight increases in size distribution around 90 nm (Fig. S10). PDA nano-liposomes were all well-dispersed without any aggregation, although some of the analytes (such as H2PO4− and THR) could also produce certain degree of electrostatic interaction with PDA nano-liposomes (Fig. S11).
a Particle size of PDA nano-liposomes after the addition of different analytes. SEM images of PDA nano-liposomes with different additives: b PDA nano-liposomes only, c PDA nano-liposomes after the addition of PLP. Scale bar: 500 nm. d Raman spectra of PDA nano-liposomes (cyan line), PDA nano-liposomes in response to PLP (magenta line), PDA nano-liposomes in response to ALP pre-treated PLP (orange line) and PDA nano-liposomes in response to H2PO4– (violet line)
It is well known that the color transition of PDA nano-liposomes is due to conformational changes of conjugated backbone [21, 24]. During polymerization, the formation of intramolecular hydrogen bonding or van der Waals interactions between side chains imposes a mechanical strain on the conjugated backbone. Specific and strong interactions with side chains could lead to the interfacial perturbations and thus release the strain on the backbone, thereby inducing the color change of PDA nano-liposomes [29]. In our system, we found that PLP could strongly bind with PDA nano-liposomes through electrostatic interaction and eventually induce the aggregation of PDA nano-liposomes. These intra- and inter-molecular interactions were believed to produce interfacial disruption and subsequently caused the conformational change of the conjugated backbone. While the presence of ALP could directly remove the effects of PLP, which could be proven from the zeta potential, size, and morphological changes of PDA nano-liposomes as described above. Although some other negative charged ions or proteins could also interact with PDA nano-liposomes, the interactions were much weak as no aggregation could be observed. It means that they may not produce sufficient perturbations to trigger the structural change of the conjugated backbone. This inference could be further confirmed by Raman spectra analysis. As shown in Fig. 4d, the pure PDA nano-liposomes showed strong peaks at the frequencies of 1451, 1513, 2083, and 2119 cm−1, in which peaks at 1451 and 1513 cm−1 belong to − C = C − stretching and peaks at 2083 and 2119 cm−1 belong to − C≡C − stretching. In general, the stretching frequencies of the conjugated backbone in blue phase are at lower frequencies (1451 and 2083 cm−1). When exposure of the blue PDA to external stimulus, the stretching could shift to higher frequencies (1513 and 2119 cm−1) due to the relaxation of PDA conjugated backbones [26]. In our cases, the two stretching states were co-existed in blue PDA nano-liposomes, possibly due to the more volatile backbone of PDA nano-liposomes in this metastable state. However, in the presence of PLP, similar frequency shifts were observed as the peaks at 1451 and 2083 cm−1 disappeared (Fig. 4d, magenta line). Of note, when ALP was introduced into the system, similar Raman spectrum as that of blank PDA nano-liposomes was observed (Fig. 4d, orange line). Moreover, other species including H2PO4− (Fig. 4d, violet line), K+, GSH, and THR showed no effects on Raman spectra of PDA nano-liposomes (Fig. S12). These results suggest that conformational alteration of PDA conjugated backbone occurs only when PLP is presented, while this effect can be suppressed when ALP is introduced. Taken together, one can concluded that the strong interactions of PLP with PDA nano-liposomes induced the clear blue-to-red color change and achieved the selective detection for ALP since it can specially and efficiently catalyze the hydrolysis of PLP.
Application for ALP inhibitor screening
Since inhibitor screening of an enzyme is of great importance in drug design, we then extended the applicability of this method in evaluation of ALP inhibitor efficiency. To this end, EDTA, an irreversible and noncompetitive inhibitor of ALP, was tested. From Fig. 5, it can be seen that the absorption band at 640 nm gradually decreased with the increasing of EDTA concentration, along with significant increases in absorption band at 540 nm (Fig. 5a). This observation clearly indicates that ALP activity is inhibited by EDTA, resulting in decreased hydrolysis rate of PLP. By plotting the inhibition efficiency as a function of EDTA concentration, the IC50 was calculated to be 3.63 µM at given ALP concentration of 0.3 U/mL (Fig. 5b). In addition, the inhibitor assay process could also be reflected through color changes (Fig. 5c). Noticeable changes from blue to red could be observed with the increasing of EDTA. These results demonstrate that the colorimetric assay developed in this study could also be used to screen the inhibitor of ALP in a facile and visual manner.
Detection of ALP activity in serum samples
As the high sensitivity and selectivity of this method for ALP activity, we further examined the accuracy of the method for detection of ALP in spiked serum samples and real human serum samples. Standard final concentrations of ALP ranging from 0.01 to 0.20 U/mL were added into HEPES buffer containing 1% fetal bovine serum to prepare spiked serum samples. As shown in Table S1, the level of ALP in unspiked sample was not detected. Based on the standard curve, all the spiked samples were detected with the recoveries of ALP in the range of 97.97–103.01%. The method was further challenged with real human serum samples. Five unrelated human serum samples were analyzed both with a standard p-NPP-based method and our proposed approach. As shown in Fig. 6, the ALP activities detected by this method were well consistent with the values obtained from the p-NPP based method. The results thus indicate that our proposed colorimetric method could be used for analysis of ALP in biological samples with acceptable accuracy, and may provide a simpler alternative for assaying of ALP activity in clinical applications or biological studies.
Conclusions
We reported a new PDA liposome-based colorimetric assay for ALP activity with the assistant of PLP. This approach is on the basis of the pioneering findings that PDA nano-liposomes bearing quaternary ammonium or amino groups could produce optical response to specific phosphates. Two diacetylene monomers modified with ethylenediamine and imidazolium were synthesized and self-assembled into PDA nano-liposomes at a 1:1 ratio. It was found that the as-prepared PDA nano-liposomes show selective and clear chromatic responses to PLP. In the presence of ALP, PLP was hydrolyzed to form PL, which retarded the colorimetric response of PDA nano-liposomes. Because the reduced concentration of PLP is highly rely on the ALP levels, the faded color processes can thus be used to detect the activity of ALP. Under optimal sensing conditions, the proposed method can be facilely used to assay ALP activity with high sensitivity and selectivity. Evaluation of the inhibitor efficiency could also be achieved with this approach. Besides, a satisfying accuracy was obtained when the method was applied for the detection of ALP in human serum samples, suggesting its potential use in real clinic samples. As PDA materials are temperature sensitive, the disadvantage of the assay may come from the differences in ambient temperature which may produce certain errors in the analysis results. Considering the facile preparation, easy-to-use assaying process and visual detection nature of PDA nano-liposomes, we believe that this work may not only provide a new approach for ALP activity but also offer a new avenue for the design and development of PDA nanomaterial-based sensing system towards other enzymes with similar strategies.
References
Fernandez NJ, Kidney BA (2019) Alkaline phosphatase: beyond the liver. Vet Clin Pathol 36:223–233
Qian Z, Chai L, Tang C, Huang Y, Chen J, Feng H (2015) Carbon quantum dots-based recyclable real-time fluorescence assay for alkaline phosphatase with adenosine triphosphate as substrate. Anal Chem 87:2966–2973
Sharma U, Pal D, Prasad R (2014) Alkaline phosphatase: an overview. Ind J Clin Biochem 29:269–278
Huang S, Yao J, Chu X, Ning G, Zhou Z, Liu Y, Xiao Q (2020) A ratiometric fluorescent assay for evaluation of alkaline phosphatase activity based on ionic liquid–functionalized carbon dots. Microchim Acta 187:271
Bowers GN Jr, McComb RB (1966) A continuous spectrophotometric method for measuring the activity of serum alkaline phosphatase. Clin Chem 12:70–89
Niu X, Ye K, Wang L, Lin Y, Du D (2019) A review on emerging principles and strategies for colorimetric and fluorescent detection of alkaline phosphatase activity. Anal Chim Acta 1086:29–45
Tang Z, Chen H, He H, Ma C (2019) Assays for alkaline phosphatase activity: progress and prospects. TrAC Trends Anal Chem 113:32–43
Han Y, Chen J, Li Z, Chen H, Qiu H (2020) Recent progress and prospects of alkaline phosphatase biosensor based on fluorescence strategy. Biosens Bioelectron 148:111811
Wang K, Wang W, Zhang XY, Jiang AQ, Yang YS, Zhu HL (2021) Fluorescent probes for the detection of alkaline phosphatase in biological systems: recent advances and future prospects. TrAC Trends Anal Chem 136:116189
Pan MC, Lei YM, Chai YQ, Yuan R, Zhuo Y (2020) In situ controllable generation of copper nanoclusters confined in a poly-L-cysteine porous film with enhanced electrochemiluminescence for alkaline phosphatase detection. Anal Chem 92:13581–13587
Mahato K, Purohit B, Kumar A, Chandra P (2020) Clinically comparable impedimetric immunosensor for serum alkaline phosphatase detection based on electrochemically engineered Au-nano-Dendroids and graphene oxide nanocomposite. Biosens Bioelectron 148:111815
Zhang J, He L, Zhang X, Wang J, Yang L, Liu B, Jiang C, Zhang Z (2017) Colorimetric and SERS dual-readout for assaying alkaline phosphatase activity by ascorbic acid induced aggregation of Ag coated Au nanoparticles. Sens Actuators B 253:839–845
Sun D, Xu W, Liang C, Shi W, Xu S (2020) Smart surface-enhanced resonance Raman scattering nanoprobe for monitoring cellular alkaline phosphatase activity during osteogenic differentiation. ACS Sens 5:1758–1767
Hu Q, Zhou B, Dang P, Li L, Kong J, Zhang X (2017) Facile colorimetric assay of alkaline phosphatase activity using Fe(II)-phenanthroline reporter. Anal Chim Acta 950:170–177
Wang C, Gao J, Cao Y, Tan H (2018) Colorimetric logic gate for alkaline phosphatase based on copper (II)-based metal-organic frameworks with peroxidase-like activity. Anal Chim Acta 1004:74–81
Tian F, Zhou J, Ma J, Liu S, Jiao B, He Y (2019) MnO2 nanosheets as oxidase mimics for colorimetric detection of alkaline phosphatase activity. Microchim Acta 186:408
Liu SG, Han L, Li N, Xiao N, Ju YJ, Li NB, Luo HQ (2018) A fluorescence and colorimetric dual-mode assay of alkaline phosphatase activity via destroying oxidase-like CoOOH nanoflakes. J Mater Chem B 6:2843–2850
Hayat A, Bulbul G, Andreescu S (2014) Probing phosphatase activity using redox active nanoparticles: a novel colorimetric approach for the detection of enzyme activity. Biosens Bioelectron 56:334–339
Wu T, Hou W, Ma Z, Liu M, Liu X, Zhang Y, Yao S (2019) Colorimetric determination of ascorbic acid and the activity of alkaline phosphatase based on the inhibition of the peroxidase-like activity of citric acid-capped Prussian Blue nanocubes. Microchim Acta 186:123
Chen H, Zhou Z, Lu Q, Wu C, Liu M, Zhang Y, Yao S (2019) Molecular structure regulation and enzyme cascade signal amplification strategy for upconversion ratiometric luminescent and colorimetric alkaline phosphatase detection. Anal Chim Acta 1051:160–168
Yarimaga O, Jaworski J, Yoon B, Kim JM (2012) Polydiacetylenes: supramolecular smart materials with a structural hierarchy for sensing, imaging and display applications. Chem Commun 48:2469–2485
Fang F, Meng F, Luo L (2020) Recent advances on polydiacetylene-based smart materials for biomedical applications. Mater Chem Front 4:1089–1104
Weston M, Tjandra AD, Chandrawati R (2020) Tuning chromatic response, sensitivity, and specificity of polydiacetylene-based sensors. Polym Chem 11:166–183
Chen X, Zhou G, Peng X, Yoon J (2012) Biosensors and chemosensors based on the optical responses of polydiacetylenes. Chem Soc Rev 41:4610–4630
Kim C, Hong C, Lee K (2021) Structures and strategies for enhanced sensitivity of polydiacetylene (PDA) based biosensor platforms. Biosens Bioelectron 181:113120
Wang Y, Pei H, Jia Y, Liu J, Li Z, Ai K, Lu Z, Lu L (2017) Synergistic tailoring of electrostatic and hydrophobic interactions for rapid and specific recognition of lysophosphatidic acid, an early-stage ovarian cancer biomarker. J Am Chem Soc 139:11616–11621
Kim KM, Oh DJ, Ahn KH (2011) Zinc(II)-dipicolylamine-functionalized polydiacetylene–liposome microarray: a selective and sensitive sensing platform for pyrophosphate ions. Chem Asian J 6:122–127
Zhu Y, Qiu D, Yang G, Wang M, Zhang Q, Wang P, Ming H, Zhang D, Yu Y, Zou G, Badugu R, Lakowicz JR (2016) Selective and sensitive detection of MiRNA-21 based on gold-nanorod functionalized polydiacetylene microtube waveguide. Biosens Bioelectron 85:198–204
Wang DE, Gao X, You S, Chen M, Ren L, Sun W, Yang H, Xu H (2020) Aptamer-functionalized polydiacetylene liposomes act as a fluorescent sensor for sensitive detection of MUC1 and targeted imaging of cancer cells. Sens Actuators B 309:127778
Qian X, Städler B (2020) Polydiacetylene-based biosensors for the detection of viruses and related biomolecules. Adv Funct Mater 30:2004605
Wang DE, Zhang Y, Li T, Tu Q, Wang J (2014) Self-immolative trigger-initiated polydiacetylene probe for β-glucuronidase activity. RSC Adv 4:16820–16823
Jeon H, Lee S, Li Y, Park S, Yoon J (2012) Conjugated polydiacetylenes bearing quaternary ammonium groups as a dual colorimetric and fluorescent sensor for ATP. J Mater Chem 22:3795–3799
Yang D, Guo Z, Tang Y, Miao P (2018) Poly(thymine)-templated selective formation of copper nanoparticles for alkaline phosphatase analysis aided by alkyne−azide cycloaddition “click” reaction. ACS Appl Nano Mater 1:168–174
Jiao H, Chen J, Li W, Wang F, Zhou H, Li Y, Yu C (2014) Nucleic acid-regulated perylene probe-induced gold nanoparticle aggregation: a new strategy for colorimetric sensing of alkaline phosphatase activity and inhibitor screening. ACS Appl Mater Interfaces 6:1979–1985
Zeng HH, Liu F, Peng ZQ, Yu K, Rong LQ, Wang Y, Wu P, Liang RP, Qiu JD (2020) Lanthanide phosphate nanoparticle-based one-step optical discrimination of alkaline phosphatase activity. ACS Appl Nano Mater 3:2336–2345
Cai M, Ding C, Wang F, Ye M, Zhang C, Xian Y (2019) A ratiometric fluorescent assay for the detection and bioimaging of alkaline phosphatase based on near infrared Ag2S quantum dots and calcein. Biosens Bioelectron 137:148–153
Hu Q, He M, Mei Y, Feng W, Jing S, Kong J, Zhang X (2017) Sensitive and selective colorimetric assay of alkaline phosphatase activity with Cu(II)-phenanthroline complex. Talanta 163:146–152
Ma JL, Yin BC, Wu X, Ye BC (2016) Copper-mediated DNA-scaffolded silver nanocluster on−off switch for detection of pyrophosphate and alkaline phosphatase. Anal Chem 88:9219–9225
Li CM, Zhen SJ, Wang J, Li YF, Huang CZ (2013) A gold nanoparticles-based colorimetric assay for alkaline phosphatase detection with tunable dynamic range. Biosens Bioelectron 43:366–371
Deng J, Yu P, Wang Y, Mao L (2015) Real-time ratiometric fluorescent assay for alkaline phosphatase activity with stimulus responsive infinite coordination polymer nanoparticles. Anal Chem 87:3080–3086
Acknowledgements
We would like to thank the Analytical & Testing Center of Northwestern Polytechnical University for SEM tests and analyses. Also, we thank Dr. Zhao Lei from School of Life Science and Technology, Xidian University, for his help in Raman spectral test.
Funding
This study was supported by the National Natural Science Foundation of China (Nos. 32001055 and 81772409), China Postdoctoral Science Foundation (2019M653753).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, DE., You, S., Huo, W. et al. Colorimetric detection of alkaline phosphatase activity based on pyridoxal phosphate–induced chromatic switch of polydiacetylene nano-liposomes. Microchim Acta 189, 70 (2022). https://doi.org/10.1007/s00604-022-05175-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-022-05175-y