Key message
Summary of rice grain size.
Abstract
Rice is one of the most important crops in the world. Increasing rice yield has been an urgent need to support the rapid growth of global population. The size of grains is one of major components determining rice yield; thus, grain size has been an essential target during rice breeding. Understanding the genetic and molecular mechanisms of grain size control can provide new strategies for yield improvement in rice. In general, the final size of rice grains is coordinately controlled by cell proliferation and cell expansion in the spikelet hull, which sets the storage capacity of the grain and limits grain filling. Recent studies have identified several quantitative trait loci and a number of genes as key grain size regulators. These regulators are involved in G protein signaling, the mitogen-activated protein kinase signaling pathway, the ubiquitin–proteasome pathway, phytohormone signalings, or transcriptional regulation. In this review, we summarize current knowledge on grain size control in rice and discuss the genetic and molecular mechanisms of these grain size regulators.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Rice is one of the most important cereal crops in the world and is also the primary food source for about half of the world’s population. Given the rapid increase in global population, improving grain yield has been an urgent need in rice breeding. Rice grain yield is mainly determined by three components: number of panicles per plant, number of grains per panicle, and grain weight. Grain weight is positively associated with grain size. Therefore, grain size is an important agronomical trait for yield improvement in rice.
The rice grain has a typical structure of cereal grains (Fig. 1). The embryo and the endosperm are enclosed by a thin seed coat and covered by the spikelet hull (the husk). The endosperm storing starches and other nutritious compounds occupy the bulk of the mature seed and are the major consumable parts for food. The spikelet hull consists of the palea and the lemma. It not only provides a protective coat, but also forms the container of filling. As the spikelet hull sets the storage capacity of the grain and limits grain growth, it plays a predominant role in determining grain size.
The development of spikelet hull is coordinately regulated by cell proliferation and cell expansion. During early developmental stages, cells in the spikelet hull undergo extensive division to increase cell number. Subsequently, cell division slows down gradually and cell expansion initiates to increase cell size. The final cell number and cell size in different dimensions of the spikelet hull determine grain length, grain width, and grain thickness, therefore influencing the grain size and shape (Fig. 2). Recent studies have identified several quantitative trait loci (QTLs) and a number of genes as key grain size regulators. These regulators have been involved in multiple signaling pathways, including G protein signaling, the mitogen-activated protein kinase (MAPK) signaling pathway, the ubiquitin–proteasome pathway, phytohormone signalings, and transcriptional regulatory factors (Fig. 2, Table 1). In this review, we summarize these findings and discuss the genetic and molecular mechanisms of these regulators in grain size control.
Grain size control in rice. The grain size of rice is regulated by cell proliferation and cell expansion in the spikelet hull. Components of the ubiquitin–proteasome (UPS) pathway, G-protein signaling, MAPK signaling, phytohormones and transcriptional regulatory factors are involved in grain size control. The positive regulators and negative regulators are shown in green and red, respectively. Regulators controlling cell proliferation are placed to the left of the green dotted line; regulators controlling cell expansion are placed to the right of the yellow dotted line; regulators controlling both processes are placed between the green and the yellow dotted lines. *, insufficient evidence or inconsistent results of their roles in cell proliferation/cell expansion process. Bars, 1mm
Control of grain size by G protein signaling
G protein signaling has been involved in a variety of growth and developmental processes in plants and animals. The heterotrimeric G protein complex consists of three subunits: Gα, Gβ, and Gγ. They function with membrane-bound G protein-coupled receptors (GPCRs) to mediate signal transduction to downstream effectors (Hamm 1998). Recent studies suggest that G protein signaling plays a role in grain size control.
GRAIN SIZE3 (GS3) is the first molecularly characterized QTL for grain size. It is the major QTL that contributes to grain-length differences between indica varieties and japonica varieties. The GS3 locus was identified by map-base cloning using near-isogenic lines from cross between Minghui 63 (large grain) and Chuan 7 (small grain) (Fan et al. 2006). GS3 encodes a transmembrane protein with four putative domains: a plant-specific organ size regulation (OSR) domain in the N-terminus, a transmembrane domain, a tumor necrosis factor receptor/nerve growth factor receptor (TNFR/NGFR) family cysteine-rich domain, and a von Willebrand factor type C (VWFC) in the C terminus (Fan et al. 2006; Mao et al. 2010). Intriguingly, the four domains in GS3 protein function differentially in grain size regulation (Mao et al. 2010). The OSR domain is both necessary and sufficient to limit grain size, whereas the C-terminal TNFR/NGFR and VWFC domains have an inhibitory effect on the OSR function. A nonsense mutation carried by the Minghui 63 allele (gs3C165A) causes loss of function of the OSR domain, resulting in long grains. A 1-bp deletion carried by Chuan 7 allele (gs3del357) results in deletion of most of the C-terminal cysteine-rich region, leading to super short grains. By contrast, Zhenshan 97 harboring the wild-type GS3 allele produces medium grains. Sequence analysis of 82 accessions revealed that the gs3C165A allele carried by Minghui 63 is associated with the long-grain varieties widely cultivated in the world, while the gs3del357 related to super short grains is extremely rare, suggesting that the GS3 has been selected during rice breeding.
GS3 shares some homology with DENSE AND ERECT PANICLE1 (DEP1), which is encoded by the QTL locus DEP1/qPE9-1. DEP1/qPE9-1 is characterized as a major rice grain yield QTL for panicle architecture (Huang et al. 2009; Zhou et al. 2009). DEP1 contains the N-terminal region (ORS domain), the putative transmembrane domain, and the C-terminal 4-disulfide-core domain. A gain-of-function mutation at the DEP1 locus results in truncation of the ORS domain, which is very similar to the gain-of-function mutation in the GS3 allele in Chuan 7. This mutant DEP1 allele enhances meristematic activity, resulting in dense and erect panicles, increased grain number per panicle and increased grain yield (Huang et al. 2009).
The N-terminal domain (ORS domain) of GS3 and DEP1 shares significant sequence similarity with the N-terminal domain of Arabidopsis heterotrimeric G protein γ-subunits (Gγ) AGG1 and AGG2, and atypical Gγ AGG3, and therefore was considered as γ-like domains (Chakravorty et al. 2011; Huang et al. 2009; Li et al. 2012b; Mao et al. 2010). In Arabidopsis, AGG3 interacts with Gβ (AGB1), and the role of AGG3 in seed growth is dependent on Gα and Gβ, indicating that these three G proteins function in a same genetic pathway to control seed growth (Li et al. 2012a, b). Loss of function of Gα (RGA1) or suppression of Gβ (RGB1) decreases grain size in rice (Ashikari et al. 1999; Fujisawa et al. 1999; Utsunomiya et al. 2011), suggesting that growth of rice grain is also regulated by Gα and Gβ. The N-terminal domains of GS3 and DEP1 contain several conserved residues critical for binding of Gβ subunit, although it is still unclear whether GS3 and DEP1 act with RGA and RGB1 to regulate rice grain size. Intriguingly, rice GS3 and DEP1 play negative roles in grain size, while AGG3 positively regulates seed growth in Arabidopsis (Li et al. 2012a). It is unclear whether they function with different cofactors or act on different downstream components to control seed development. Further studies would be expected to elucidate why Gγ proteins have different effects on seed growth in Arabidopsis and rice.
Control of grain size by the MAPK signaling pathway
The mitogen-activated protein kinase (MAPK) cascades are evolutionary conserved signaling modules in eukaryotes and play critical roles in transducing developmental and defense signals in plants (Meng and Zhang 2013; Xu and Zhang 2015). Recent studies found that the MAPK cascades are also involved in grain size control. Loss of function of SMALL GRAIN 1 (SMG1)/MITOGEN-ACTIVATED PROTEIN KINASE KINASE4 (OsMKK4) results in small grains due to decreased cell number in spikelet hulls, suggesting that OsMKK4 promotes grain growth (Duan et al. 2014). Similarly, OsMAPK6 also acts positively in grain size control (Liu et al. 2015b). The mutation in OsMAPK6 restricts cell proliferation in spikelet hulls, leading to small grains. OsMKK4 interacts with OsMAPK6 and phosphorylates OsMAPK6, suggesting that OsMKK4 and OsMAPK6 act as a module to control grain size (Kishi-Kaboshi et al. 2010; Liu et al. 2015b). Interestingly, both OsMKK4 and OsMAPK6 affect brassinosteroid (BR) responses and the expression of BR-related genes, suggesting a possible link between BR signaling and the MAPK pathways (Duan et al. 2014; Liu et al. 2015b). It would be worthwhile to identify the upstream MAPKKK and the downstream components of OsMKK4-OsMAPK6 in grain size control.
Control of grain size by the ubiquitin–proteasome pathway
Modification of target proteins by ubiquitin chains is an important regulatory process in eukaryotes. Ubiquitination requires the sequential action of three enzymes: the ubiquitin-activating enzyme (E1), the ubiquitin-conjugating enzyme (E2), and the ubiquitin ligase (E3). The diverse length and linkage of the ubiquitin chains have different effects on the target protein. Generally, linkage via K48 leads to protein degradation by the 26S proteasome. The ubiquitin chain can be removed from substrate proteins by the deubiquitinating enzymes (DUBs). Recent studies showed that the ubiquitin–proteasome pathway plays important roles in seed size control.
GW2
The major QTL for grain width (GW2) was identified by map-based cloning using the progeny of a cross between japonica variety WY3 (large grain) and indica variety Fengaizhan-1 (FAZ1; small grain) (Song et al. 2007). GW2 encodes a RING-type E3 ubiquitin ligase. The WY3 allele harbors a 1-bp deletion in the coding region of GW2, leading to a premature stop codon. This loss-of-function mutation enhances cell proliferation in the spikelet hulls and accelerates grain filling, resulting in increased grain width, weight, and yield. Further analysis revealed that another wide grain variety Oochikara carries the same GW2 allele as WY3. Importantly, the GW2 allele in WY3 could increase grain size and yield with little effect on appearance and no reduction in cooking or eating quality and thus could be a useful target in rice breeding.
GW2 shares significant sequence similarity with Arabidopsis E3 ubiquitin ligase DA2, which interacts with the ubiquitin-receptor DA1 to control seed size (Li et al. 2008; Xia et al. 2013). Recent studies show that DA2 can ubiquitylate DA1 to active its peptidase activity, and the activated DA1 then cleaves downstream substrates to control seed and organ growth (Dong et al. 2017). These studies provide some clues to the molecular mechanisms by which GW2 controls grain growth. It would be challenging but interesting to identify the substrates and downstream components of GW2 in grain size control.
OsOTUB1/WTG1
Loss of function of the deubiquitinating enzyme OsOTUB1/WIDE AND THICK GRAIN 1 (WTG1) increases grain width, grain thickness, and grain number per panicle (Huang et al. 2017). OsOTUB1 controls grain size and shape mainly by influencing cell expansion. Overexpression of OsOTUB1 results in narrow, thin, and long grains. Later, OsOTUB1 is also identified as a major QTL determining the ‘new plant type’ (NPT) architecture, which is characterized by larger panicles, stronger culms, and fewer sterile tillers (Wang et al. 2017). OsOTUB1 interacts with OsSPL14/IPA1 that is an important regulator of ideal plant architecture. This interaction limits K63-linked ubiquitination (K63Ub) of OsSPL14 and in turn promotes K48Ub-dependent proteasome degradation of OsSPL14. Downregulation of OsOTUB1 causes accumulation of OsSPL14 and results in the NPT architecture. It will be interesting to investigate whether OsOTUB1 regulates grain size through SPL transcription factors.
Control of grain size by phytohormones
Brassinosteroids
Brassinosteroids (BRs) are a class of polyhydroxysteroid plant hormones that are essential for the proper regulation of multiple physiological processes during plant growth and development (Clouse 2011). The role of BR on grain size control has been shown by a number of studies. The BR-deficient mutants dwarf11 and dwarf2 produce small and short grains, suggesting that BR promotes grain growth (Fang et al. 2016; Hong et al. 2005; Tanabe et al. 2005; Wu et al. 2016). Consistently, some regulators of BR homeostasis have effects on grain size. For example, enhanced expression of SLENDER GRAIN (SLG) causes elevated BR contents, leading to long and narrow grains (Feng et al. 2016), whereas loss of function of XIAO results in typical BR-related phenotypes and reduced grain length (Jiang et al. 2012) .
BRs are perceived by the membrane-localized receptor kinase BRASSINOSTEROID-INSENSITIVE1 (BRI1) and its partner BRI1-ASSOCIATED RECEPTOR KINASE (BAK1). BR signal then initiates a cascade of cellular events, leading to inactivation of BRASSINOSTEROID-INSENSITIVE 2 (BIN2) and activation of the two transcription factors BRASSINAZOLE-RESISTANT1 (BZR1) and BZR2 for transcription of downstream genes (Clouse 2011). In rice, loss of function of OsBRI1 or OsBAK1 results in BR-insensitive phenotypes and small grains (Morinaka et al. 2006; Yuan et al. 2017), whereas overexpression of OsBZR1 increases grain length, grain width and grain weight (Zhu et al. 2015). The downstream components of BR signaling also affect grain size. The rice counterpart of BIN2, GSK2, negatively regulates grain size (Tong et al. 2012). GSK2 can phosphorylate the GRAS family protein DWARF AND LOW-TILLERING (DLT/OsGRAS-32/D62/GS6), a positive regulator that mediates several BR responses in rice (Tong et al. 2012). One of the studies showed that d62 has short and wide grains (Li et al. 2010b), while another study showed that the grain width of gs6 was increased but the grain length was not significantly different compared with the wild-type 93-11 (Sun et al. 2013).
Several regulators of BR signaling were also involved in grain size control. A pair of bHLH proteins POSITIVE REGULATOR OF GRAIN LENGTH 1 (PGL1) and ANTAGONIST OF PGL1 (APG) antagonistically regulate rice grain length and weight by controlling cell elongation in lemma/palea through heterodimerization (Heang and Sassa 2012a). APG negatively regulates grain length, while its function is inhibited by PGL1. Overexpression of PGL1 or suppression of APG results in increased BL sensitivity and long grains, suggesting that PGL1 and APG1 control grain length probably by mediating BR signaling. Similar to PGL1, the atypical bHLH protein PGL2/BRASSINOSTEROID UPREGULATED 1-LIKE1 (OsBUL1) promotes grain length by suppressing the function of APG (Heang and Sassa 2012b). PGL2/OsBUL1 can also form a transcriptional activator complex with another basic helix-loop-helix (bHLH) transcriptional activator OsBUL1 COMPLEX1 (OsBC1) and KxDL motif-containing protein LO9-177 to regulate BR response and grain size (Jang et al. 2017; Jang and Li 2017). The closet homolog of PGL2/OsBUL1, BRASSINOSTEROID UPREGULATED1 (BU1), is also a positively regulator of BR response and grain size (Heang and Sassa 2012b; Tanaka et al. 2009). However, no interaction was detected between BU1 and APG, indicating that BU1 might control grain length independently of APG (Heang and Sassa 2012b). In addition, SHORT GRAIN1 (SG1), a protein with unknown function, acts as a negative regulator of BR response and grain size (Nakagawa et al. 2012). Overexpression of SG1 in rice causes brassinosteroid (BR)-deficient phenotype and short grains, while downregulation of SG1 and SG1-LIKE PROTEIN1 results in long grains.
The QTL for grain size GRAIN SIZE 5 (GS5) encodes a putative serine carboxypeptidase which functions as a positive regulator of grain size (Li et al. 2011b). GS5 was identified by using a double haploid (DH) population (92 lines) derived from a cross between Zhenshan 97 (wide grains) and H94 (slender grains). Higher expression of GS5 increases grain width and grain yield by accelerating cell division and cell expansion in the spikelet hull. Sequence analysis of 51 rice accessions from a wide geographic range revealed that polymorphisms in the promoter region of GS5 are likely correlated with grain width, indicating that natural variation in GS5 contributes to grain size diversity in rice. Furthermore, a recent study found that GS5 regulates grain size by preventing OsBAK1-7 endocytosis and enhancing BR signaling, suggesting a possible link between GS5 and BR signaling in grain size control (Xu et al. 2015a).
The major QTL for grain length (qGL3/qGL3.1) was identified by three independent studies (Hu et al. 2012; Qi et al. 2012; Zhang et al. 2012). qGL3/GL3.1 encodes a Ser/Thr phosphatase with Kelch-like repeat domain (OsPPKL1). OsPPKL1 controls cell division in the spikelet by directly dephosphorylating Cyclin-T1;3. A single nucleotide transition from C to A (c. + 1092C → A) causes an aspartate to glutamate change (Asp364Glu) in a conserved AVLDT motif of the second Kelch domain in OsPPKL1, resulting in weaker dephosphorylation activity. The qgl3 allele increases grain length and grain yield without affecting grain quality. Sequencing analysis of the qGL3 locus using 94 rice germplasms showed that only one variety (DT108) carries the (c. + 1092C → A) transition, suggesting that qgl3 is a rare allele. Furthermore, the qgl3 allele could significantly increase grain yield in various rice varieties. Therefore, it could be used in breeding elite rice varieties (Zhang et al. 2012).
There are two OsPPKL1 homologs in rice, OsPPKL2 and OsPPKL3 (Zhang et al. 2012). Interestingly, OsPPKL1 and OsPPKL3 limit grain length, while OsPPKL2 promotes grain growth. OsPPKL2 belongs to a subgroup with Arabidopsis homologs AtBSU1 and AtBSL1, two serine–threonine protein phosphatases that function in brassinosteroid signaling pathway to promote cell elongation and cell division. It would be worthwhile to investigate whether OsPPKLs influence grain length through brassinosteroid-mediated signaling.
The SEED WIDTH ON CHROMOSOME 5 (GW5/qSW5) is a major QTL that determines grain-width differences between indica and japonica varieties (Shomura et al. 2008; Weng et al. 2008). GW5/qSW5 was identified by two independent studies using different recombinant inbred lines (RILs) generated from crosses between Asominori/Nipponbare (wide grains) and IR24/Kasalath (slender grains) (Shomura et al. 2008; Weng et al. 2008). Sequencing results revealed that GW5/qSW5 is associated with a 1212-bp deletion. Transformation of a 11.2-kbp Kasalath fragment covering the deletion region resulted in thin rice grains in Nipponbare background, suggesting that qSW5 is in this region. One of the predicted ORFs (GenBank: Kasalath qSW5 gene, AB433345) in the 11.2-kbp region, which encodes an unknown protein, was proposed to be the qSW5 gene (Shomura et al. 2008), while another study proposed that an ORF (Gene bank: IR24, GW5, DQ991205) encoding an ubiquitin-interacting protein is the GW5 gene (Weng et al. 2008). Thus, GW5 has been proposed to be involved in the proteasome pathway (Li and Li 2014, 2016). However, functional complementation tests using these individual ORFs were lacking. Recent studies revealed that the transformation of Nipponbare with another ORF (LOC_Os05g09520) in the 11.2-kbp Kasalath fragment resulted in thin grains, suggesting that this ORF encodes GW5 (Liu et al. 2017). Meanwhile, this ORF was identified as a major QTL for grain size (GSE5) through genome-wide association analysis (Duan et al. 2017). GSE5/GW5 encodes a plasma membrane-associated protein with IQ calmodulin-binding motifs (Duan et al. 2017; Liu et al. 2017). GSE5/GW5 physically associates with rice calmodulin (OsCAM1), suggesting that calcium signaling may play a role in grain size control. However, how GSE5/GW5 mediates calcium signaling to regulate grain size remains unknown.
Further analysis showed that natural variation in the promoter region of GSE5 contributes to grain size diversity in cultivated rice (Duan et al. 2017; Liu et al. 2017). GSE5 has three major haplotypes in cultivated rice: most japonica varieties have a 1212-bp deletion (DEL2) in the promote region of GSE5; most narrow-grain indica varieties have no deletion; and most wide-grain indica varieties contain a 950-bp deletion (DEL1) and a 367-bp insertion (IN1) in the promoter region of GSE5 and a nucleotide change (G/A) in the first exon of GSE5. DEL1 in indica varieties and DEL2 in japonica varieties associate with decreased expression of GSE5, resulting in wide grains. The DEL1 and DEL2 deletions likely originated from different wild rice accessions during rice domestication and are widely utilized by rice breeders. Knockout of GSE5/GW5 using CRISPR/Cas9 technology in japonica and indica varieties significantly increased grain width and weight, suggesting that the GSE5/GW5 locus may be used to improve rice yield (Duan et al. 2017; Liu et al. 2017). In addition, GSE5/GW5 can repress the kinase activity of GSK3/SHAGGY-like kinase (GSK2), a component of brassinosteriod (BR) signaling pathway, suggesting that GSE5/GW5 may regulate grain width by modulating BR signaling (Liu et al. 2017). Interestingly, GSE5/GW5 controls grain size by regulating cell proliferation in spikelet hulls, while GSK2 affects grain size predominantly by influencing cell expansion in spikelet hulls.
Auxin
Although auxin has been demonstrated to play important roles in many aspects of plant growth and development, its role in grain size control remains elusive. So far, only a few pieces of evidence show a connection between auxin and grain size control. The major QTL for thousand-grain weight (TGW6) was identified by positional cloning using backcrossed inbred lines produced from Nipponbare (heavy grains) and Kasalath (light grains) (Ishimaru et al. 2013). TGW6 encodes IAA-glucose hydrolase, which regulates the transition from the syncytial to the cellular phase during early endosperm development by regulating IAA supply. A 1-bp deletion in the Kasalath allele causes loss of function of TGW6 and results in increased grain weight and grain yield, suggesting that TGW6 negatively regulates grain growth. The Kasalath TGW6 allele can also increase the accumulation of carbohydrates before heading and consequently improve yield without change in grain quality. Analysis of different wild rice lines (Oryza rufipogon) and 69 rice varieties showed that the Kasalath TGW6 allele has probably not been selected in rice breeding and thus could be used in rice yield improvement. Notably, TGW6 influences grain length and grain weight with no effect on husk size, indicating a distinct regulation mechanism from other grain size regulators.
BIG GRAIN1 (BG1), a novel membrane-localized protein, was identified as a positive regulator of grain size (Liu et al. 2015a). Activation of BG1 increases grain size and grain weight due to increased cell proliferation and cell expansion in spikelet hulls, whereas suppression of BG1 results in small grains. BG1 affects auxin response and transport, suggesting that it may control grain growth through auxin signaling. Nonetheless, future studies need to elucidate the function of BG1 in auxin transport/signaling.
Loss of function of SMALL ORGAN SIZE1 (SMOS1), an unusual APETALA2 (AP2)-type transcription factor, results in small grains and organs (Aya et al. 2014). SMOS1 promotes cell expansion and microtubule orientation. The promoter region of SMOS1 gene contains auxin response elements (AuxRE), and the expression of SMOS1 was induced by exogenous auxin treatment, suggesting that SMOS1 acts as an auxin-dependent regulator for cell expansion. SMOS1 directly regulates the expression of PHOSPHATEINDUCED PROTEIN 1 (OsPHI-1) that is involved in cell expansion. In addition, a recently report showed that SMOS1 forms a complex with DLT to integrate auxin and brassinosteroid signaling in rice (Hirano et al. 2017). However, the genetic relationship between SMOS1 and DLT in grain size control is still unclear.
Cytokinin
Recent findings suggest that cytokinin plays important roles in controlling grain number and grain size. A major QTL for grain number, Gn1a, encodes cytokinin oxidase/dehydrogenase (OsCKX2) that modulates cytokinin accumulation by catalyzing the degradation of active CKs (Ashikari et al. 2005). Reduced expression of OsCKX2 increases grain number, with no effects on grain size. Thus, Gn1 could be used for increasing grain yield. The expression of OsCKX2 is regulated by LARGE PANICLE (LP) and DROUGHT AND SALT TOLERANCE (DST) (Li et al. 2011a, 2013). LARGER PANICLE (LP) is a Kelch repeat-containing F-box protein localized in endoplasmic reticulum (ER). The lp mutants show increased grain number, grain size and grain yield. OsCKX2 was downregulated in the lp mutants, implying that LP might regulate grain number and grain size by modulating cytokinin level (Li et al. 2011a). DST is a zinc finger transcription factor. It directly regulates OsCKX2 expression in the reproductive meristem, thereby influencing the number of the reproductive organs through modulating CK accumulation (Li et al. 2013). The semidominant DSTreg1 allele affects DST-directed regulation of OsCKX2 expression, leading to elevated CK levels in the inflorescence meristem and increased grain number, grain weight and grain yield (Li et al. 2013).
STRESS_tolerance and GRAIN_LENGTH (OsSGL), an abiotic stress-induced gene, has been involved in stress tolerance and grain length (Wang et al. 2016). OsSGL encodes a putative DUF1645 family protein with unknown function. Overexpression of OsSGL not only enhances drought tolerance but also increases grain length, grain weight and grain number per panicle, resulting in a significant increase in yield. OsSGL promotes grain growth by increasing longitudinal cell number and cell size in the lemma/palea. Transcriptome analysis showed that elevated expression of OsSGL alters the expression of several genes related to CK signaling process, suggesting that OsSGL may regulate grain length and stress response through modulating CK signal transduction.
An-2 is a QTL locus for awn length. The An-2 gene encodes a Lonely Guy (LOG) homologous enzyme that catalyzes the last step of cytokinin synthesis. An-2 promotes awn elongation by enhancing cell division, but decreases grain production by reducing grain weight and grain number. Genetic variation analysis shows that the cultivar allele of An-2 shows significantly reduced nucleotide diversity compared with wild rice, indicating that this locus was selected for reduced awn length and increased grain yield during rice domestication (Gu et al. 2015).
Control of grain size by transcriptional regulatory factors
GLW7
The major QTL for grain length and weight (GLW7) was identified by an approach integrating genome-wide association testing with functional analysis on grain size in a population of 381 japonica varieties (Si et al. 2016). GLW7 encodes the plant-specific transcription factor SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 13 (OsSPL13). OsSPL13 positively regulates grain length and yield by promoting cell expansion in the grain hull. Higher expression of OsSPL13 is associated with large grains in tropical japonica rice due to difference in the 5′ UTR sequence of OsSPL13 which affects transcription and translation. This large-grain allele of GLW7 in tropical japonica rice was introgressed from indica varieties under artificial selection. Furthermore, GLW7 directly associates with the promoter region of SMALL AND ROUND SEED 5 (SRS5) and promotes its expression. SRS5 encodes alpha-tubulin protein (Segami et al. 2012). A semidominant mutation of SRS5 leads to short grains with reduced cell length (Segami et al. 2012), while plants overexpressing SRS5 form long grains (Segami et al. 2017). However, the genetic relationship between GLW7 and SRS5 remains unclear.
GW8
The QTL for grain width (GW8) was identified by analysis of segment substitution lines (SSSLs) from a cross between Basmati385 (slender grain) and HJX74 (wide grain) (Wang et al. 2012). GW8 encodes OsSPL16, which increases grain width and yield by promoting cell division and grain filling (Wang et al. 2012). In Basmati rice, a 10-bp deletion in the deletion in the OsSPL16 promoter region results in reduced transcription and leads to slender grains and better quality of appearance. Haplotype diversity of the OsSPL16 sequence suggests that the Basmati haplotype was selected due to its association with better grain quality, whereas in elite indica varieties the HJX74 haplotype was selected for higher grain productivity.
The major QTL for rice grain length (GL7)/GRAIN WIDTH 7 (GW7)/SLENDER GRAIN ON CHROMOSOME 7 (SLG7) was identified by three independent studies (Wang et al. 2015a, b; Zhou et al. 2015). GL7/GW7/SLG7 encodes a TON1 RECRUITING MOTIF (TRM)-containing protein homologous to Arabidopsis LONGIFOLIA proteins involved in microtubule regulation. In long-grain varieties, elevated expression of GL7 leads to slender grains (Wang et al. 2015a, b), and this beneficial allele of GL7 has been selected in rice breeding (Wang et al. 2015b). One of the studies showed that tandem duplication of a 17.1-kb segment at the GL7 locus leads to upregulation of GL7, resulting in increased grain length and improvement in grain appearance quality (Wang et al. 2015b). By contrast, another study reported that the mutation in the promoter region of GL7 causes its high expression (Wang et al. 2015a). The expression of GL7/GW7/SLG7 was regulated by the SPL transcription factor OsSPL16, which is encoded by the QTL for grain width (GW8) (Wang et al. 2015a). It was shown that OsSPL16/GW8 binds to the promoter of GL7/GW7/SLG7 and represses its transcription to regulate cell proliferation in the spikelet hull (Wang et al. 2015a). Controversially, two studies demonstrated that GL7/GW7/SLG7 regulates grain length through cell expansion. It enhances cell elongation in the grain-length direction and restricts cell expansion in the grain-width direction (Wang et al. 2015b; Zhou et al. 2015). Further studies need to clarify whether GL7/GW7/SLG7 controls grain size through cell proliferation or cell expansion.
GS2/GL2/GLW2/PT2
GRAIN SIZE 2 (GS2)/GRAIN-LENGTH-ASSOCIATED (GL2)/GRAIN LENGTH AND WIDTH 2 (GLW2)/PANICLE TRAITS 2 (PT2), a major QTL for grain length, grain width and weight, was identified independently by five research groups using different F2 populations (Che et al. 2015; Duan et al. 2015; Hu et al. 2015; Li et al. 2016; Sun et al. 2016). GS2 encodes a plant-specific transcription factor GROWTH-REGULATING FACTOR 4 (OsGRF4), which regulates grain size through predominantly increasing cell expansion and slightly promoting cell proliferation in the spikelet hull. The expression of OsGRF4 is regulated by miRNA396. In large-grain varieties, a 2-bp substitution mutation (TC → AA, GS2AA) in miR396 targeting site of OsGRF4 perturbs OsmiR396-directed regulation, causing elevated expression of OsGRF4, and resulting in large and heavy grains and increased grain yield. Sequence analysis of various cultivars revealed that the GS2AA allele is a rare allele and has not been selected by breeders, suggesting that the GS2AA allele could be used to increase grain size and yield (Duan et al. 2015; Hu et al. 2015; Sun et al. 2016).
OsGRF4 interacts with the transcription coactivators GRF-INTERACTING FACTOR 1/2/3 (OsGIF1/2/3) (Duan et al. 2015; Li et al. 2016). Overexpression of OsGIF1 increases grain size and weight in rice (Che et al. 2015; Duan et al. 2015; He et al. 2017; Li et al. 2016). Thus, the OsmiR396–OsGRF4–OsGIFs regulatory module plays important roles in grain size control. In addition, OsGRF4 interacts with GSK2 which functions in BR signaling (Che et al. 2015). GSK2 can repress the transcription activation activity of OsGRF4 and suppress the function of OsGRF4 in grain size control, suggesting that OsGRF4 may function with BR signaling to regulate grain size.
GW6a
The QTL for grain weight (GW6a) was detected using a set of backcrossed inbred lines derived from a cross of Kasalath (light grains) with Nipponbare (heavy grains) (Song et al. 2015). GW6a encodes a new-type GNAT-like protein with intrinsic histone acetyltransferase activity (OsglHAT1). OsglHAT1 is localized in nucleus and functions presumably via regulation of gene expression. Elevated OsglHAT1 expression increases grain weight and grain yield by enhancing cell proliferation in spikelet hulls and accelerating grain filling. GW6a is the first QTL for yield component traits that encodes a chromatin modifier. Importantly, GW6a has not been selected during rice domestication and modern breeding, indicating that it could be exploited in rice yield improvement.
GL4
The quantitative trait locus for grain length (GL4) in African rice was detected using F2 population derived from a cross between introgression line GIL25 (long grains) and a cultivar of African cultivated rice IRGC102305 (short grains) (Wu et al. 2017). GL4 encodes a Myb-like protein sharing highly identity with SH4/SHA1, its orthologue in Asian wild rice. SH4 was previously reported to control seed shattering, and the Ossh4 allele resulting with non-shattering has been selected during the domestication of Asian cultivated rice O. sativa. GL4 regulates grain length in African cultivated rice (O. glaberrima) by promoting longitudinal cell elongation in the glumes. Like SH4/SHA1 in Asian wild rice, GL4 also controls seed shattering. A single nucleotide substitution (C760T) in the IRGC102305 allele causes a premature stop codon, leading to small grains and loss of grain shattering during African rice domestication. By contrast, GIL25 harboring the wild-type GL4 allele has increased grain length and grain yield. Further studies showed that GL4/SH4 is a key domestication gene with pleiotropic effects, which controls both grain size and shattering in rice. During crop domestication, increasing seed size and reducing seed shattering were two main selection targets. Ossh4 and Ogsh4 were selected in parallel during the domestication of Asian and African rice, resulting in loss of grain shattering. The Ossh4 mutation does not change the seed size in Asian cultivated rice, while Ogsh4 leads to small seeds in African cultivated rice. Replacing the Ogsh4 allele with the Ossh4 allele would enhance the grain yield of O. glaberrima.
Other factors in grain size control
P450 family proteins
The CYP78A subfamily P450 monooxygenase GIANT EMBRYO (GE)/BIG GRAIN2 (BG2)/GRAIN LENGTH 3.2 (GL3.2) is critical for coordinating rice embryo and endosperm development. The GE/BG2 gene was identified by three independent studies (Nagasawa et al. 2013; Xu et al. 2015b; Yang et al. 2013). It is expressed predominantly in the scutellar epithelium, the interface region between embryo and endosperm, and coordinates the development of the embryo and endosperm (Nagasawa et al. 2013; Yang et al. 2013). Loss of function of GE leads to large embryos and small endosperm, whereas GE overexpression causes small embryos and enlarged endosperm, suggesting that GE is crucial for the coordinated development of the embryo and the endosperm.
Another putative cytochrome P450, CYP704A3, is also responsible for grain length (Tang et al. 2016). The expression of CYP704A3 was regulated by miRNA. A SNP at the miRNA binding site in the 3′-UTR region of CYP704A3 is associated with rice grain size. Downregulation of CYP704A3 via RNAi increases grain length.
Cytochrome P450s play important roles in a variety of biosynthetic pathways. Recently, several cytochrome P450s have been involved in seed size control. In Arabidopsis, KLUH (KLU)/CYP78A5 promotes seed and organ growth in a non-cell-autonomous manner (Adamski et al. 2009; Anastasiou et al. 2007; Eriksson et al. 2010; Wang et al. 2008). It was proposed that KLU can generate a mobile growth signal that is distinct from the classic phytohormones (Anastasiou et al. 2007). The expression of KLU is regulated by the NGATHA-like B3 domain transcriptional repressor (NGAL2)/SUPPRESSOR OF DA1 (SOD7) and its homolog NGAL3/DEVELOPMENT- RELATED PCG TARGET IN THE APEX 4 (DPA4). NGAL2/SOD7 directly binds to the promoter of KLUH (KLU)/CYP78A5 and represses the transcription of KLU to regulate seed growth. Two homologs of KLU, EOD3/CYP78A6 and CYP78A9, control seed size in Arabidopsis by promoting both cell proliferation and cell expansion in maternal integuments.
GAD1
GRAIN NUMBER, GRAIN LENGTH AND AWN DEVELOPMENT1 (GAD1)/REGULATOR OF AWN ELONGATION 2 (RAE2) was identified by two independent studies (Bessho-Uehara et al. 2016; Jin et al. 2016). GAD1/RAE2 encodes a small secretary signal peptide belonging to the EPIDERMAL PATTERNING FACTOR-LIKE family. It promotes grain elongation and awn development by enhancing cell division at the apices of glumes. Loss of function of GAD1 results in increased number of grains per panicle, short grains, and awnless phenotype. The GAD1/RAE2 precursor is specifically cleaved by its requisite processing enzymes, SUBTILISIN-LIKE PROTEASE 1 (SLP1), in the rice spikelet (Bessho-Uehara et al. 2016). EPF/EPFL family members have been shown to regulate multiple biological processes in plants (Murphy et al. 2012). In Arabidopsis, the EPFL family peptides bind to the membrane-bond ERECTA family receptors, which transduce the signals through MAPK cascade to regulate stomata development (Bergmann and Sack 2007; Lampard et al. 2009; Lee et al. 2012, 2015; Pillitteri and Dong 2013). It would be interesting to identify the receptor for GAD1 and investigate whether the OsMKK4-OsMPK6 module acts in a same pathway with GAD1 to regulate grain size in the future.
FUWA
The fuwa mutant shows compact plant architecture with wide, thick and short grains (Chen et al. 2015). FUWA encodes an NHL domain-containing protein that is evolutionary conserved. Downregulation of FUWA results in erect panicles and increased grain size in both indica and japonica rice, suggesting a potential approach to improve agronomic traits. Several cyclins and cyclin-dependent kinases were upregulated in the fuwa mutant, suggesting that FUWA controls grain growth by regulating cell-cycle progression. The detailed mechanisms of FUWA in grain size control remains to be further investigated.
OsKinesin-13A
SMALL AND ROUND SEED 3 (SRS3)/SMALL AND ROUND GRAINS (SAR1)/OsKINESIN-13A is an active microtubule depolymerase, which mainly distributes on vesicles derived from the Golgi apparatus and is destined for the cell surface (Deng et al. 2015). Loss of function of OsKinesin-13A leads to short grains due to decreased cell elongation in the glumes (Deng et al. 2015; Kitagawa et al. 2010). The srs3 mutant shows defective orientation of cellulose microfibrils and microtubule turnover, suggesting that OsKinesin-13A may control cell elongation and grain length through affecting cellulose microfibril orientation and vesicle transport (Deng et al. 2015).
DEP2/SRS1
DENSE AND ERECT PANICLE 2 (DEP2)/SMALL AND ROUND SEED 1 (SRS1)/ERECT PANICLE2-1 (EP2-1) is involved in the control of both panicle architecture and grain size (Abe et al. 2010; Li et al. 2010a; Zhu et al. 2010). The dep2 mutant has dense and erect panicles as well as small and round grains. DEP2 encodes an endoplasmic reticulum-localized protein without any known functional domain. The reduced grain length of srs1-1 is due to the reduction in both cell length and cell number in the longitudinal direction, and the elongation of the cells in the lateral direction of the lemma (Abe et al. 2010). Interestingly, although the dep2 mutant has a compact plant architecture, the grain production is comparable to that of the wild type, indicating that this allele has important implications for rice breeding.
OsAGSW1
A chloroplast-localized ABC1 protein kinase, OsAGSW1 (ABC1-like kinase related to grain size and weight), is involved in the regulation of grain size and weight. OsAGSW1 promotes grain growth by regulating the number of external parenchyma cells. Overexpression of OsAGSW1 increases the number of external parenchyma cells in the spikelet hull, leading to increased grain size, grain weight, grain filling rate and 1000-grain weight. However, the molecular mechanisms by which OsAGSW1 controls grain growth are still unclear.
Discussion and perspective
In the past decades, there have been great research progresses on grain size control in rice. A number of grain size regulators related to several signaling pathways were identified (Fig. 2, Table 1). However, our understanding of the mechanisms of grain size control is just beginning and full of gaps. The molecular roles of some regulators in grain size control are still unclear or controversial. The genetic relationships between different regulators and the molecular interactions between different signaling pathways are largely unknown. In previous studies, the mutant alleles used by independent research groups were usually in different genetic backgrounds, which might lead to inconsistent conclusions. Besides, the near-isogenic lines used for genetic analyses might contain other mutations. These problems can now be resolved by newly emerging genome-editing technologies, including CRISPR/Cas9, which allow researchers to knock out candidate genes and analyze their genetic interactions in the same genetic background. In addition, system biology and new biotechnologies will facilitate the studies on grain size control. For instants, the genome-wide association study and modern omics analysis could help to identify novel grain size regulators. Future challenges are to elucidate the molecular mechanisms of identified regulators in grain growth control, identify novel regulators to fill up the gaps in each signaling pathway, and build up genetic frameworks regulating grain size.
Recent studies showed that some of the seed size regulators have conserved functions between rice and other plant species. For example, several components in BR signaling and G protein signaling influence seed size in both rice and Arabidopsis (Li and Li 2015, 2016); the ubiquitin ligase GW2 and its homologs in Arabidopsis, wheat and maize have conserved functions in seed size control (Bednarek et al. 2012; Li et al. 2010; Song et al. 2007; Xia et al. 2013). These pieces of evidences indicate that different plant species share similar mechanisms to control seed and organ growth. Therefore, studies on rice grain size could help us to understand the mechanisms of seed size control in other crops.
Author contribution statement
NL and YL conceived and wrote the manuscript. RX and DP helped to prepare and revise the manuscript.
References
Abe Y, Mieda K, Ando T, Kono I, Yano M, Kitano H, Iwasaki Y (2010) The SMALL AND ROUND SEED1 (SRS1/DEP2) gene is involved in the regulation of seed size in rice. Genes Genet Syst 85:327–339
Adamski NM, Anastasiou E, Eriksson S, O’Neill CM, Lenhard M (2009) Local maternal control of seed size by KLUH/CYP78A5-dependent growth signaling. Proc Natl Acad Sci USA 106:20115–20120. https://doi.org/10.1073/pnas.0907024106
Anastasiou E, Kenz S, Gerstung M, MacLean D, Timmer J, Fleck C, Lenhard M (2007) Control of plant organ size by KLUH/CYP78A5-dependent intercellular signaling. Dev Cell 13:843–856. https://doi.org/10.1016/j.devcel.2007.10.001
Ashikari M, Wu J, Yano M, Sasaki T, Yoshimura A (1999) Rice gibberellin-insensitive dwarf mutant gene Dwarf 1 encodes the alpha-subunit of GTP-binding protein. Proc Natl Acad Sci USA 96:10284–10289
Ashikari M et al (2005) Cytokinin oxidase regulates rice grain production. Science 309:741–745. https://doi.org/10.1126/science.1113373
Aya K, Hobo T, Sato-Izawa K, Ueguchi-Tanaka M, Kitano H, Matsuoka M (2014) A novel AP2-type transcription factor, SMALL ORGAN SIZE1, controls organ size downstream of an auxin signaling pathway. Plant Cell Physiol 55:897–912. https://doi.org/10.1093/pcp/pcu023
Bednarek J et al (2012) Down-regulation of the TaGW2 gene by RNA interference results in decreased grain size and weight in wheat. J Exp Bot 63:5945–5955. https://doi.org/10.1093/jxb/ers249
Bergmann DC, Sack FD (2007) Stomatal development. Annu Rev Plant Biol 58:163–181. https://doi.org/10.1146/annurev.arplant.58.032806.104023
Bessho-Uehara K et al (2016) Loss of function at RAE2, a previously unidentified EPFL, is required for awnlessness in cultivated Asian rice. Proc Natl Acad Sci USA 113:8969–8974. https://doi.org/10.1073/pnas.1604849113
Chakravorty D et al (2011) An atypical heterotrimeric G-protein gamma-subunit is involved in guard cell K(+)-channel regulation and morphological development in Arabidopsis thaliana. Plant J Cell Mol Biol 67:840–851. https://doi.org/10.1111/j.1365-313x.2011.04638.x
Che R et al (2015) Control of grain size and rice yield by GL2-mediated brassinosteroid responses. Nat Plants 2:15195. https://doi.org/10.1038/nplants.2015.195
Chen J et al (2015) An evolutionarily conserved gene, FUWA, plays a role in determining panicle architecture, grain shape and grain weight in rice. Plant J Cell Mol Biol 83:427–438. https://doi.org/10.1111/tpj.12895
Clouse SD (2011) Brassinosteroids. Arabidopsis Book 9:e0151. https://doi.org/10.1199/tab.0151
Deng ZY et al (2015) OsKinesin-13A is an active microtubule depolymerase involved in glume length regulation via affecting cell elongation. Sci Rep 5:9457. https://doi.org/10.1038/srep09457
Dong H et al (2017) Ubiquitylation activates a peptidase that promotes cleavage and destabilization of its activating E3 ligases and diverse growth regulatory proteins to limit cell proliferation in Arabidopsis. Genes Dev 31:197–208. https://doi.org/10.1101/gad.292235.116
Duan P et al (2014) SMALL GRAIN 1, which encodes a mitogen-activated protein kinase kinase 4, influences grain size in rice. Plant J 77:547–557. https://doi.org/10.1111/tpj.12405
Duan P et al (2015) Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nat Plants 2:15203. https://doi.org/10.1038/nplants.2015.203
Duan P et al (2017) Natural variation in the promoter of GSE5 contributes to grain size diversity in rice. Mol Plant 10:685–694. https://doi.org/10.1016/j.molp.2017.03.009
Eriksson S, Stransfeld L, Adamski NM, Breuninger H, Lenhard M (2010) KLUH/CYP78A5-dependent growth signaling coordinates floral organ growth in Arabidopsis. Curr Biol 20:527–532. https://doi.org/10.1016/j.cub.2010.01.039
Fan C et al (2006) GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor Appl Genet 112:1164–1171. https://doi.org/10.1007/s00122-006-0218-1
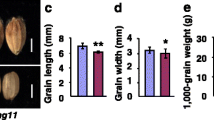
Fang N et al (2016) SMALL GRAIN 11 controls grain size, grain number and grain yield in rice. Rice (NY) 9:64. https://doi.org/10.1186/s12284-016-0136-z
Feng Z et al (2016) SLG controls grain size and leaf angle by modulating brassinosteroid homeostasis in rice. J Exp Bot 67:4241–4253. https://doi.org/10.1093/jxb/erw204
Fujisawa Y et al (1999) Suppression of the heterotrimeric G protein causes abnormal morphology, including dwarfism, in rice. Proc Natl Acad Sci USA 96:7575–7580
Gu B et al (2015) An-2 encodes a cytokinin synthesis enzyme that regulates awn length and grain production in rice. Mol Plant 8:1635–1650. https://doi.org/10.1016/j.molp.2015.08.001
Hamm HE (1998) The many faces of G protein signaling. J Biol Chem 273:669–672
He Z et al (2017) OsGIF1 positively regulates the sizes of stems, leaves, and grains in rice. Front Plant Sci 8:1730. https://doi.org/10.3389/fpls.2017.01730
Heang D, Sassa H (2012a) Antagonistic actions of HLH/bHLH proteins are involved in grain length and weight in rice. PLoS ONE 7:e31325. https://doi.org/10.1371/journal.pone.0031325
Heang D, Sassa H (2012b) An atypical bHLH protein encoded by POSITIVE REGULATOR OF GRAIN LENGTH 2 is involved in controlling grain length and weight of rice through interaction with a typical bHLH protein APG. Breed Sci 62:133–141. https://doi.org/10.1270/jsbbs.62.133
Hirano K et al (2017) SMALL ORGAN SIZE 1 and SMALL ORGAN SIZE 2/DWARF AND LOW-TILLERING form a complex to integrate auxin and brassinosteroid signaling in rice. Mol Plant 10:590–604. https://doi.org/10.1016/j.molp.2016.12.013
Hong Z et al (2005) The Rice brassinosteroid-deficient dwarf2 mutant, defective in the rice homolog of Arabidopsis DIMINUTO/DWARF1, is rescued by the endogenously accumulated alternative bioactive brassinosteroid, dolichosterone. Plant Cell 17:2243–2254. https://doi.org/10.1105/tpc.105.030973
Hu Z et al (2012) A Kelch motif-containing serine/threonine protein phosphatase determines the large grain QTL trait in rice. J Integr Plant Biol 54:979–990. https://doi.org/10.1111/jipb.12008
Hu J et al (2015) A rare allele of GS2 enhances grain size and grain yield in rice. Mol Plant 8:1455–1465. https://doi.org/10.1016/j.molp.2015.07.002
Huang X et al (2009) Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet 41:494–497. https://doi.org/10.1038/ng.352
Huang K, Wang D, Duan P, Zhang B, Xu R, Li N, Li Y (2017) WIDE AND THICK GRAIN 1, which encodes an otubain-like protease with deubiquitination activity, influences grain size and shape in rice. Plant J Cell Mol Biol. https://doi.org/10.1111/tpj.13613
Ishimaru K et al (2013) Loss of function of the IAA-glucose hydrolase gene TGW6 enhances rice grain weight and increases yield. Nat Genet 45:707–711. https://doi.org/10.1038/ng.2612
Jang S, Li HY (2017) Oryza sativa BRASSINOSTEROID UPREGULATED1 LIKE1 induces the expression of a gene encoding a small leucine-rich-repeat protein to positively regulate lamina inclination and grain size in rice. Front Plant Sci 8:1253. https://doi.org/10.3389/fpls.2017.01253
Jang S, An G, Li HY (2017) Rice leaf angle and grain size are affected by the OsBUL1 transcriptional activator complex. Plant Physiol 173:688–702. https://doi.org/10.1104/pp.16.01653
Jiang Y, Bao L, Jeong SY, Kim SK, Xu C, Li X, Zhang Q (2012) XIAO is involved in the control of organ size by contributing to the regulation of signaling and homeostasis of brassinosteroids and cell cycling in rice. Plant J Cell Mol Biol 70:398–408. https://doi.org/10.1111/j.1365-313X.2011.04877.x
Jin J et al (2016) GAD1 encodes a secreted peptide that regulates grain number, grain length, and awn development in rice domestication. Plant Cell 28:2453–2463. https://doi.org/10.1105/tpc.16.00379
Kishi-Kaboshi M et al (2010) A rice fungal MAMP-responsive MAPK cascade regulates metabolic flow to antimicrobial metabolite synthesis. Plant J 63:599–612. https://doi.org/10.1111/j.1365-313X.2010.04264.x
Kitagawa K et al (2010) A novel kinesin 13 protein regulating rice seed length. Plant Cell Physiol 51:1315–1329. https://doi.org/10.1093/pcp/pcq092
Lampard GR, Lukowitz W, Ellis BE, Bergmann DC (2009) Novel and expanded roles for MAPK signaling in Arabidopsis stomatal cell fate revealed by cell type-specific manipulations. Plant Cell 21:3506–3517. https://doi.org/10.1105/tpc.109.070110
Lee JS et al (2012) Direct interaction of ligand-receptor pairs specifying stomatal patterning. Genes Dev 26:126–136. https://doi.org/10.1101/gad.179895.111
Lee JS et al (2015) Competitive binding of antagonistic peptides fine-tunes stomatal patterning. Nature 522:439–443. https://doi.org/10.1038/nature14561
Li N, Li Y (2014) Ubiquitin-mediated control of seed size in plants. Front Plant Sci 5:332. https://doi.org/10.3389/fpls.2014.00332
Li N, Li Y (2015) Maternal control of seed size in plants. J Exp Bot 66:1087–1097. https://doi.org/10.1093/jxb/eru549
Li N, Li Y (2016) Signaling pathways of seed size control in plants. Curr Opin Plant Biol 33:23–32. https://doi.org/10.1016/j.pbi.2016.05.008
Li Y, Zheng L, Corke F, Smith C, Bevan MW (2008) Control of final seed and organ size by the DA1 gene family in Arabidopsis thaliana. Genes Dev 22:1331–1336. https://doi.org/10.1101/gad.463608
Li F et al (2010a) Rice DENSE AND ERECT PANICLE 2 is essential for determining panicle outgrowth and elongation. Cell Res 20:838–849. https://doi.org/10.1038/cr.2010.69
Li W, Wu J, Weng S, Zhang Y, Zhang D, Shi C (2010b) Identification and characterization of dwarf 62, a loss-of-function mutation in DLT/OsGRAS-32 affecting gibberellin metabolism in rice. Planta 232:1383–1396. https://doi.org/10.1007/s00425-010-1263-1
Li Q, Li L, Yang X, Warburton M, Bai G (2010c) Relationship, evolutionary fate and function of two maize co-orthologs of rice GW2 associated with kernel size and weight. BMC Plant Biol 10:143
Li M et al (2011a) Mutations in the F-box gene LARGER PANICLE improve the panicle architecture and enhance the grain yield in rice. Plant Biotechnol J 9:1002–1013. https://doi.org/10.1111/j.1467-7652.2011.00610.x
Li Y et al (2011b) Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat Genet 43:1266–1269. https://doi.org/10.1038/ng.977
Li S, Liu W, Zhang X, Liu Y, Li N, Li Y (2012a) Roles of the Arabidopsis G protein gamma subunit AGG3 and its rice homologs GS3 and DEP1 in seed and organ size control. Plant Signal Behav 7:1357–1359. https://doi.org/10.4161/psb.21620
Li S et al (2012b) The plant-specific G protein gamma subunit AGG3 influences organ size and shape in Arabidopsis thaliana. New Phytol 194:690–703. https://doi.org/10.1111/j.1469-8137.2012.04083.x
Li S et al (2013) Rice zinc finger protein DST enhances grain production through controlling Gn1a/OsCKX2 expression. Proc Natl Acad Sci USA 110:3167–3172. https://doi.org/10.1073/pnas.1300359110
Li S et al (2016) The OsmiR396c-OsGRF4-OsGIF1 regulatory module determines grain size and yield in rice. Plant Biotechnol J 14:2134–2146. https://doi.org/10.1111/pbi.12569
Liu L et al (2015a) Activation of Big Grain1 significantly improves grain size by regulating auxin transport in rice. Proc Natl Acad Sci USA 112:11102–11107. https://doi.org/10.1073/pnas.1512748112
Liu S et al (2015b) OsMAPK6, a mitogen-activated protein kinase, influences rice grain size and biomass production. Plant J Cell Mol Biol 84:672–681. https://doi.org/10.1111/tpj.13025
Liu J et al (2017) GW5 acts in the brassinosteroid signalling pathway to regulate grain width and weight in rice. Nat Plants 3:17043. https://doi.org/10.1038/nplants.2017.43
Mao H et al (2010) Linking differential domain functions of the GS3 protein to natural variation of grain size in rice. Proc Natl Acad Sci USA 107:19579–19584. https://doi.org/10.1073/pnas.1014419107
Meng X, Zhang S (2013) MAPK cascades in plant disease resistance signaling. Annu Rev Phytopathol 51:245–266. https://doi.org/10.1146/annurev-phyto-082712-102314
Morinaka Y, Sakamoto T, Inukai Y, Agetsuma M, Kitano H, Ashikari M, Matsuoka M (2006) Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice. Plant Physiol 141:924–931. https://doi.org/10.1104/pp.106.077081
Murphy E, Smith S, De Smet I (2012) Small signaling peptides in Arabidopsis development: how cells communicate over a short distance. Plant Cell 24:3198–3217. https://doi.org/10.1105/tpc.112.099010
Nagasawa N et al (2013) GIANT EMBRYO encodes CYP78A13, required for proper size balance between embryo and endosperm in rice. Plant J Cell Mol Biol 75:592–605. https://doi.org/10.1111/tpj.12223
Nakagawa H et al (2012) Short grain1 decreases organ elongation and brassinosteroid response in rice. Plant Physiol 158:1208–1219. https://doi.org/10.1104/pp.111.187567
Pillitteri LJ, Dong J (2013) Stomatal development in Arabidopsis. Arabidopsis Book 11:e0162. https://doi.org/10.1199/tab.0162
Qi P et al (2012) The novel quantitative trait locus GL3.1 controls rice grain size and yield by regulating Cyclin-T1;3. Cell Res 22:1666–1680. https://doi.org/10.1038/cr.2012.151
Segami S, Kono I, Ando T, Yano M, Kitano H, Miura K, Iwasaki Y (2012) Small and round seed 5 gene encodes alpha-tubulin regulating seed cell elongation in rice. Rice (NY) 5:4. https://doi.org/10.1186/1939-8433-5-4
Segami S, Takehara K, Yamamoto T, Kido S, Kondo S, Iwasaki Y, Miura K (2017) Overexpression of SRS5 improves grain size of brassinosteroid-related dwarf mutants in rice (Oryza sativa L.). Breed Sci 67:393–397. https://doi.org/10.1270/jsbbs.16198
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M (2008) Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet 40:1023–1028. https://doi.org/10.1038/ng.169
Si L et al (2016) OsSPL13 controls grain size in cultivated rice. Nat Genet 48:447–456. https://doi.org/10.1038/ng.3518
Song XJ, Huang W, Shi M, Zhu MZ, Lin HX (2007) A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet 39:623–630. https://doi.org/10.1038/ng2014
Song XJ et al (2015) Rare allele of a previously unidentified histone H4 acetyltransferase enhances grain weight, yield, and plant biomass in rice. Proc Natl Acad Sci USA 112:76–81. https://doi.org/10.1073/pnas.1421127112
Sun L et al (2013) GS6, a member of the GRAS gene family, negatively regulates grain size in rice. J Integr Plant Biol 55:938–949. https://doi.org/10.1111/jipb.12062
Sun P et al (2016) OsGRF4 controls grain shape, panicle length and seed shattering in rice. J Integr Plant Biol 58:836–847. https://doi.org/10.1111/jipb.12473
Tanabe S et al (2005) A novel cytochrome P450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant, dwarf11, with reduced seed length. Plant Cell 17:776–790. https://doi.org/10.1105/tpc.104.024950
Tanaka A et al (2009) BRASSINOSTEROID UPREGULATED1, encoding a helix-loop-helix protein, is a novel gene involved in brassinosteroid signaling and controls bending of the lamina joint in rice. Plant Physiol 151:669–680. https://doi.org/10.1104/pp.109.140806
Tang W et al (2016) SNP-based analysis of genetic diversity reveals important alleles associated with seed size in rice. BMC Plant Biol 16:93. https://doi.org/10.1186/s12870-016-0779-3
Tong H et al (2012) DWARF AND LOW-TILLERING acts as a direct downstream target of a GSK3/SHAGGY-like kinase to mediate brassinosteroid responses in rice. Plant Cell 24:2562–2577. https://doi.org/10.1105/tpc.112.097394
Utsunomiya Y et al (2011) Suppression of the rice heterotrimeric G protein beta-subunit gene, RGB1, causes dwarfism and browning of internodes and lamina joint regions. Plant J Cell Mol Biol 67:907–916. https://doi.org/10.1111/j.1365-313X.2011.04643.x
Wang JW, Schwab R, Czech B, Mica E, Weigel D (2008) Dual effects of miR156-targeted SPL genes and CYP78A5/KLUH on plastochron length and organ size in Arabidopsis thaliana. Plant Cell 20:1231–1243. https://doi.org/10.1105/tpc.108.058180
Wang S et al (2012) Control of grain size, shape and quality by OsSPL16 in rice. Nat Genet 44:950–954. https://doi.org/10.1038/ng.2327
Wang S et al (2015a) The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat Genet 47:949–954. https://doi.org/10.1038/ng.3352
Wang Y et al (2015b) Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat Genet 47:944–948. https://doi.org/10.1038/ng.3346
Wang M et al (2016) OsSGL, a novel pleiotropic stress-related gene enhances grain length and yield in rice. Sci Rep 6:38157. https://doi.org/10.1038/srep38157
Wang S et al (2017) Non-canonical regulation of SPL transcription factors by a human OTUB1-like deubiquitinase defines a new plant type rice associated with higher grain yield. Cell Res. https://doi.org/10.1038/cr.2017.98
Weng J et al (2008) Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res 18:1199–1209. https://doi.org/10.1038/cr.2008.307
Wu Y, Fu Y, Zhao S, Gu P, Zhu Z, Sun C, Tan L (2016) CLUSTERED PRIMARY BRANCH 1, a new allele of DWARF11, controls panicle architecture and seed size in rice. Plant Biotechnol J 14:377–386. https://doi.org/10.1111/pbi.12391
Wu W et al (2017) A single-nucleotide polymorphism causes smaller grain size and loss of seed shattering during African rice domestication. Nat Plants 3:17064. https://doi.org/10.1038/nplants.2017.64
Xia T et al (2013) The ubiquitin receptor DA1 interacts with the E3 ubiquitin ligase DA2 to regulate seed and organ size in arabidopsis. Plant Cell 25:3347–3359. https://doi.org/10.1105/tpc.113.115063
Xu J, Zhang S (2015) Mitogen-activated protein kinase cascades in signaling plant growth and development. Trends Plant Sci 20:56–64. https://doi.org/10.1016/j.tplants.2014.10.001
Xu C, Liu Y, Li Y, Xu X, Li X, Xiao J, Zhang Q (2015a) Differential expression of GS5 regulates grain size in rice. J Exp Bot 66:2611–2623. https://doi.org/10.1093/jxb/erv058
Xu F et al (2015b) Variations in CYP78A13 coding region influence grain size and yield in rice. Plant Cell Environ 38:800–811. https://doi.org/10.1111/pce.12452
Yang W et al (2013) Control of rice embryo development, shoot apical meristem maintenance, and grain yield by a novel cytochrome p450. Mol Plant 6:1945–1960. https://doi.org/10.1093/mp/sst107
Yuan H et al (2017) 08SG2/OsBAK1 regulates grain size and number, and functions differently in Indica and Japonica backgrounds in rice. Rice (NY) 10:25. https://doi.org/10.1186/s12284-017-0165-2
Zhang X et al (2012) Rare allele of OsPPKL1 associated with grain length causes extra-large grain and a significant yield increase in rice. Proc Natl Acad Sci USA 109:21534–21539. https://doi.org/10.1073/pnas.1219776110
Zhou Y et al (2009) Deletion in a quantitative trait gene qPE9-1 associated with panicle erectness improves plant architecture during rice domestication. Genetics 183:315–324. https://doi.org/10.1534/genetics.109.102681
Zhou Y et al (2015) Natural variations in SLG7 regulate grain shape in rice. Genetics 201:1591–1599. https://doi.org/10.1534/genetics.115.181115
Zhu K et al (2010) Erect panicle2 encodes a novel protein that regulates panicle erectness in indica rice. Genetics 184:343–350. https://doi.org/10.1534/genetics.109.112045
Zhu X et al (2015) Brassinosteroids promote development of rice pollen grains and seeds by triggering expression of Carbon Starved Anther, a MYB domain protein. Plant J Cell Mol Biol 82:570–581. https://doi.org/10.1111/tpj.12820
Acknowledgements
We apologize to our colleagues whose work is not covered in this review due to limited space. This work was supported by grants from the Ministry of Agriculture of China (Grant 2016ZX08009-003), the Ministry of Science and Technology of China (Grants 2016YFD0100501), the National Natural Science Foundation of China (Grants 91535203; 31425004; 31771340; 31571742; 31400249), and the Strategic Priority Research Program Molecular Mechanism of Plant Growth and Development’ of Chinese Academy of Sciences (Grant XDPB0401).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by L. Lepiniec, H. North, G. Ingram.
A contribution to the special issue ‘Seed Biology’.
Rights and permissions
About this article
Cite this article
Li, N., Xu, R., Duan, P. et al. Control of grain size in rice. Plant Reprod 31, 237–251 (2018). https://doi.org/10.1007/s00497-018-0333-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00497-018-0333-6