Abstract
The mammalian ovary is a dynamic organ. The coordination of follicle recruitment, selection, and ovulation and the timely development and regression of the corpus luteum are essential for a functional ovary and fertility. Deregulation of any of these processes results in ovarian dysfunction and potential infertility. MicroRNA (miRNA) are short noncoding RNA that regulate developmental processes and time-sensitive functions. The expression of miRNA in the ovary varies with cell type, function, and stage of the estrous cycle. miRNA are involved in the formation of primordial follicles, follicular recruitment and selection, follicular atresia, oocyte-cumulus cell interaction, granulosal cell function, and luteinization. miRNA are differentially expressed in luteal cells at the various stages of the estrous cycle and during maternal recognition of pregnancy, suggesting a role in luteal development, maintenance, and regression. An understanding of the patterns of expression and functions of miRNA in the ovary will lead to novel therapeutics to treat ovarian dysfunction and improve fertility and, potentially, to the development of better contraceptives.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
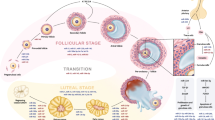
It can be argued that the mammalian ovary is the most dynamic organ in the adult female with continual activation and development of its follicles and corpora lutea (CL). Ovulation involves tissue remodeling and damage of surface epithelial cells to allow for the removal of the oocyte. Involvement of immune cells in this event has led to our understanding that ovulation is a type of inflammatory response (Espey 1980). Immediately following the release of the oocyte, resolution of the inflammation has to occur, which is coincident with the onset of the differentiation of the follicular steroidogenic cells into luteal cells. During luteinization, gene expression is altered to promote considerably greater rates of steroidogenesis coinciding with the remarkable proliferation of endothelial cells. As the corpus luteum (CL) reaches maturity, both the differentiation of steroidogenic cells and the proliferation of endothelial cells essentially cease. Little is known about the mechanisms associated with this transition from luteinization to mature luteal function. Luteolysis marks the end of the estrous cycle or the end of the luteal phase of the menstrual cycle. Proteins associated with steroidogenesis must be rapidly downregulated, proteins associated with tissue remodeling must be rapidly stimulated, and immune cells must be rapidly programmed to facilitate the resolution of the massive tissue degradation. One of the most fascinating features of reproductive biology is that the cell death, immune cell activation, and loss of protein function that occur during luteolysis are prevented when the CL is rescued during maternal recognition of pregnancy (Pate and Landis Keyes 2001; Spencer and Bazer 2004). Although the mechanisms associated with conceptus signaling to the CL during maternal recognition of pregnancy vary with species, this represents yet another important transition, wherein the tissue basically faces a “life-or-death” scenario in all mammals. In addition to their role in follicular and oocyte development, microRNA (miRNA) might be involved in these “life or death” scenarios.
miRNA are short (~22 nucleotides; nt) RNA sequences that regulate gene expression posttranscriptionally (Bartel 2004; Czech and Hannon 2010). They were first discovered to control time-dependent developmental events in Caenorhabditis elegans (Lee et al. 1993; Wightman et al. 1993) and have since been identified in various eukaryotic species (Pasquinelli et al. 2000; Lagos-Quintana et al. 2001; Lau et al. 2001). miRNA can be coded for by separate genes or embedded in intronic or exonic regions of protein coding genes (Bartel 2004) and are transcribed either as single genes or a polycistronic cluster to form stem loop secondary structures (for a review, see Du and Zamore 2005). Drosha ribonuclease type III (DROSHA) and DiGeorge syndrome critical region gene 8 (DGCR8) cleave the stem loop into a pre-miRNA, which is exported into the cytoplasm to be further spliced by Dicer 1, ribonuclease type III (DICER1), resulting in a duplex RNA. One strand of the duplex RNA (mature miRNA) partners with the RNA-induced silencing complex (RISC) and regulates target gene expression. The second (passenger) strand is degraded (Du and Zamore 2005), although it may also be selected, but the mechanism by which a strand is selected to become a mature miRNA is not completely understood (Okamura et al. 2008). Alternatively, miRNA can also be generated by DROSHA/DGCR8-independent or DICER1-independent pathways (Yang and Lai 2011; Ha and Kim 2014). In mammalian species, mature miRNA bind by base-pair complementarity to a seed sequence (7–8 nt) in the 3′ or 5′ untranslated region (UTR) of the messenger RNA (mRNA) to inhibit its translation, either by inducing mRNA degradation or by blocking its translation (Bartel 2004; Lewis et al. 2005; Lytle et al. 2007; Orom et al. 2008). However, some evidence exists that miRNA may also enhance gene expression (Orom et al. 2008; Place et al. 2008). Functional studies of miRNA can be very challenging. For instance, the expression of miRNA can be cell- or tissue-specific and may be under hormonal or temporal control (Reinhart et al. 2000; Sood et al. 2006; Fiedler et al. 2008). Hence, the study of the function of an miRNA in one tissue or a developmental stage might not translate well into other tissues or developmental stages. The function of miRNA is often redundant (Fischer et al. 2015). A single miRNA can target several genes, and one gene can be targeted by several miRNA (Brennecke et al. 2005; Lewis et al. 2005; Lim et al. 2005), adding further challenges to functional studies.
Profiling studies of miRNA in ovarian tissues have confirmed the expression of miRNA in the ovaries of various species, including mice (Ro et al. 2007; Mishima et al. 2008; Ahn et al. 2010), humans (Liang et al. 2007), cows (Hossain et al. 2009; Tripurani et al. 2010; Huang et al. 2011; Miles et al. 2012), goats (Ling et al. 2014), sheep (Di et al. 2014), and pigs (Li et al. 2011). Development of knockout (KO) animal models for Dicer1, an important ribonuclease in miRNA biogenesis, allowed the dissection of the role of miRNA in ovarian function. Dicer1 KO mice were embryonic lethal, suggesting an essential role for DICER1, and subsequently for miRNA, in embryonic development (Bernstein et al. 2003). Mutant mice with hypomorphic Dicer1 alleles (Dicer d/d) were able to survive but were infertile because of the failure of luteal formation and impaired angiogenesis (Otsuka et al. 2008). Conditional knockout (cKO) of Dicer1 from follicular granulosal cells resulted in a number of ovarian functional defects including abnormal oocyte maturation, disrupted follicular development and ovulation, increased follicular atresia, and infertility (Hong et al. 2008; Nagaraja et al. 2008; Gonzalez and Behringer 2009). A detailed comparison of the ovarian defects observed in the Dicer d/d and Dicer1 cKO mice has revealed the complexity of the role of miRNA in ovarian function and the need to study the role of specific miRNA in oocyte maturation, follicular development, and luteal function (Luense et al. 2009). To date, more than 30 miRNA KO animal models exist (for a review, see Vidigal and Ventura 2015). Eleven of these miRNA mouse models are KO or cKO of miRNA that have been found to be important in ovarian function (for a review, see Vidigal and Ventura 2015). However, not all KO models show altered reproductive phenotypes. For instance, mice with either miR-200b or miR-429 deletion exhibit no reproductive defects, but when both miRNA are deleted, the resulting mice are infertile because of disruption of luteinizing hormone synthesis in the pituitary and subsequent failure to ovulate (Hasuwa et al. 2013). Therefore, the deletion of a single miRNA might not produce a phenotypic effect that becomes apparent when additional miRNA are modified.
Previous reviews have summarized the role of miRNA in the development of the mammalian reproductive system (Zhao and Rajkovic 2008; Donadeu et al. 2012; Hossain et al. 2012; Nothnick 2012), in ovarian function (Toloubeydokhti et al. 2008; Carletti and Christenson 2009; Christenson 2010; Baley and Li 2012; Donadeu et al. 2012; Imbar and Eisenberg 2014; McGinnis et al. 2015; Li et al. 2015), and in female reproductive diseases and pathologies (Baley and Li 2012; Imbar and Eisenberg 2014; McGinnis et al. 2015; Li et al. 2015). In order not to reiterate topics that have been reviewed previously, this review will primarily focus on the latest findings concerning the expression and role of miRNA in regulating folliculogenesis, oocyte-cumulus cell interactions, and the development and function of the CL.
miRNA regulation of cumulus-oocyte communication and oocyte maturation
Cellular communication between the oocyte and the surrounding somatic cells is essential for appropriate folliculogenesis, oocyte maturation, and ovulation of a healthy oocyte (Matzuk et al. 2002; Hawkins and Matzuk 2010). cKO of Dicer1 in murine granulosal cells increases the number of primordial follicles and results in abnormal recruitment and increased follicular atresia, suggesting an involvement of miRNA in these processes (Lei et al. 2010). The oocytes isolated from Zp3-Dicer1 cKO mice, in which Dicer1 is specifically deleted in the oocyte, are arrested in meiosis because of defects in meiotic spindle formation (Murchison et al. 2007). A recent study by Yuan et al. (2014) has shown that cKO of Dicer1, but not Drosha, in oocytes of murine fetal ovary results in premature ovarian failure and infertility in the adult ovary. Interestingly, Flemr et al. (2013) have discovered a mouse oocyte-specific DICER1 isoform that lacks the N-terminal DExD helicase domain and which has greater cleavage activity than the full-length DICER1. While DICER1 is more efficient in processing pre-miRNA into mature miRNA, the oocyte-specific DICER1 isoform is more efficient in processing long double-stranded RNA into endogenous small interfering RNA (endo-siRNA) in oocytes (Flemr et al. 2013). Therefore, the primary active small RNA in the mouse oocyte is probably endo-siRNA, not miRNA. Recently, a study by Stein et al. (2015) has supported this result by showing the necessity of endo-siRNA for meiotic progression in mouse oocytes. The expression of miRNA varies during the maturation of both human and bovine oocytes (Tesfaye et al. 2009; Tripurani et al. 2010; Abd El Naby et al. 2011; Assou et al. 2013). To date, the efficiency of DICER1 in producing miRNA vs. endo-siRNA has only been described for mouse oocytes.
In bovine oocytes, Tesfaye et al. (2009) identified 59 miRNA that were differentially expressed in immature vs. mature bovine oocytes; 31 miRNA were more predominant in immature bovine oocytes, and 28 were more predominant in mature bovine oocytes. Abd El Naby et al. (2011) profiled miRNA expression in both immature and mature oocytes compared with their surrounding cumulus cells by using polymerase chain reaction (PCR) arrays. In immature oocytes, 39 miRNA were upregulated, and in mature oocytes, 45 miRNA were upregulated, compared with miRNA expression in the surrounding cumulus cells, respectively. Thirty-three miRNA were upregulated in the oocyte compared with cumulus cells, regardless of oocyte maturation status. Of these 33, six miRNA declined in expression during in vitro oocyte maturation. The expression of miRNA in cumulus cells was influenced by the presence of the oocyte, indicating that cumulus-oocyte interactions affected miRNA expression (Abd El Naby et al. 2011). Assou et al. (2013) used small RNAseq and microarrays to compare miRNA and mRNA expression in human metaphase II (MII) oocytes with that in cumulus cells from patients undergoing in vitro fertilization. The number of identified miRNA (32) was greater in cumulus cells than in MII oocytes, in which only three known miRNA were present. All of these miRNA were cell-specific, with no expression being noted in the opposite cell type. Let-7b, let-7c, and miR-21 were the most abundant miRNA in cumulus cells, with 51, 31, and 28 reads, respectively. In MII oocytes, the most abundant miRNA were miR-184 and miR-10a, with 1988 and 555 reads, respectively. In silico analysis of the oocyte-specific miRNA resulted in predicted targets that were associated with the regulation of transcription and cell cycle, whereas predicted targets of miRNA in the cumulus cells were associated with the extracellular matrix and apoptosis. The microarray analysis revealed over 10,000 differentially expressed genes in cumulus cells vs. oocytes, and 224 of those were included in the list of predicted targets of the expressed miRNA. Based on these findings, gene function during cumulus-oocyte maturation seems likely to be regulated by these miRNA. However, importantly, this study was performed on oocytes that failed to fertilize, and hence, the pattern of expression of these miRNA might depict a pathologic, rather than a normal condition. Overall, the differential and cell-specific expression of miRNA in granulosal cells and oocytes and the temporal dependence of expression support the studies with Dicer1 conditional KO models, leading to the conclusion that miRNA are important mediators of granulosal-oocyte communication and oocyte development.
miRNA in follicular development
Recent studies investigating the role of specific miRNA in fetal and neonatal mouse ovarian tissue have revealed three miRNA (mir-143, miR-145, and miR-376a) that are important in the formation and maintenance of the primordial follicle pool (Fig. 1). Transfection of cultured newborn mouse ovaries with miR-376a increases the number of primordial follicles and reduces oocyte apoptosis by targeting the expression of Pcna, which promotes the apoptosis of oocytes in fetal and neonatal mouse ovaries (B. Xu et al. 2011; H. Zhang et al. 2014). Overexpression of miR-376a in fetal mouse ovaries 18.5 days postcoitum (dpc) decreases the expression of proapoptotic genes (Bax, Tnf, and Tnfr-2) and increases the expression of antiapoptotic (Bcl2) and oocyte survival (Pard6a, Lhx8) genes, although the proteins have not been quantified, and so whether these proteins increase or decrease remains unclear (H. Zhang et al. 2014). The expression of miR-143 increases in fetal mouse primordial follicles from 15.5 dpc to 4 days postpartum. The expression of miR-143 has been observed in pregranulosal cells but not in oocytes. Functional studies of miR-143 in cultured fetal mouse ovaries have demonstrated that miR-143 inhibits primordial follicle formation by reducing the proliferation of pregranulosal cells and decreasing the expression of cell-cycle-related genes (J. Zhang et al. 2013). MiR-145 may also have a role in the maintenance of the primordial follicle pool and the regulation of the rate of follicular activation (Yang et al. 2013). Inhibition of miR-145 in neonatal mouse ovaries decreases the expression of the zona pellucida (ZP) genes, Zp1, Zp2, and Zp3, increases the expression of TGFBR2, and leads to the activation of the TGFB signaling pathway, which is important in primordial and primary follicle activation, and many other ovarian functions (Yang et al. 2013). Figure 1 summarizes the miRNA with known functions during follicular development in mouse whole ovarian tissues. The mechanisms by which many other miRNA regulate follicular development in mice or other species remain to be investigated.
MicroRNA (miRNA) regulate early folliculogenesis. The depicted miRNA are those that have been confirmed as regulators of the formation and maintenance of primordial and primary follicles in fetal and neonatal mouse ovaries (B. Xu et al. 2011; Yang et al. 2013; J. Zhang et al. 2013; H. Zhang et al. 2014). Dashed lines represent the function of miRNA on the early steps of folliculogenesis. Dashed lines have been used, instead of solid lines, to indicate that our knowledge of the role of miRNA in these steps is incomplete, and that other miRNA may also be involved
In attempts to understand the role of miRNA in follicular growth and the selection of the dominant follicle, miRNA expression was analyzed in follicles during development. Based on microarray analysis, a total of 523 miRNA were expressed in large vs. small and large healthy vs. large atretic bovine follicles (Sontakke et al. 2014). The expression patterns of five of these miRNA (miR-144, miR-202 [exclusively expressed in gonads], miR-451, miR-652, and miR-873) were confirmed by quantitative PCR. Of these, three (miR-144, miR-202, and miR-873) were expressed at greater concentrations in large healthy follicles compared with large atretic follicles and were suggested to be involved in the regulation of mural cell function in the dominant follicle. miR-873 was considered to be the strongest candidate of those three for playing a role in the selection of a dominant follicle (Sontakke et al. 2014). In a separate study, bovine subordinate and dominant follicles were found to contain 244 common miRNA on both days 3 and 7 of the estrous cycle (Salilew-Wondim et al. 2014). Of these, the let-7 family, miR-10b, miR-26a, miR-27b, and miR-99b were highly expressed regardless of the stage of the estrous cycle (Salilew-Wondim et al. 2014). As the estrous cycle progressed from day 4 to day 7, few changes in miRNA expression were observed in subordinate follicles, whereas greater changes in miRNA expression were noted in the dominant follicles (Salilew-Wondim et al. 2014). In the horse, miR-132, miR-212 miR-21, miR-145, miR-224, and miR-378 were differentially expressed in dominant vs. subordinate or dominant vs. luteinized follicles and were suggested to be involved in follicular selection and ovulation (Schauer et al. 2013). These exciting results present evidence that miRNA might be involved in the selection of the dominant follicle, the mechanism of which has remained largely elusive.
Functional effects of miRNA in granulosal cells
Granulosal cell proliferation and function are essential in follicular development, maturation, and atresia. Functions of individual miRNA can be readily evaluated by using cultures of primary granulosal cells, and these can be performed with cells from a variety of species. Unsurprisingly, the ovarian cell type most studied with regard to regulation by miRNA is the granulosal cell. Several studies have shown miRNA to be involved in the proliferation, survival, function, and/or death of granulosal cells (Table 1). Several miRNA have been found to regulate the proliferation of primary or immortalized cultured granulosal cells. For example, miR-181a was revealed to inhibit granulosal cell proliferation by decreasing proliferating cell nuclear antigen (PCNA) accumulation and targeting AcvrIIa (Sirotkin et al. 2010; Q. Zhang et al. 2013). Overexpression of miR-320 in murine granulosal cells, both in vivo and in vitro, suppressed cell proliferation (Yin et al. 2014). The expression of miR-320 and the suppression of granulosal cell proliferation were further enhanced by miR-383 (Yin et al. 2014). Overexpression of miR-93, which is highly expressed in human polycystic ovaries, increased cell proliferation in immortalized human granulosal tumor cells (KGN) by targeting a cyclin-dependent kinase inhibitor (CDKN1A; Jiang et al. 2015). High concentrations of insulin, mimicking hyperinsulinemia in polycystic ovarian syndrome (PCOS), also induced the expression of miR-93, increased KGN proliferation, and downregulated the expression of CDKN1A (Jiang et al. 2015). However, the pattern of expression of miR-93 in the granulosal cells of healthy ovaries has not been described, and so whether the role of miR-93 in granulosal cell proliferation is general or limited to cells in a PCOS-like environment remains unclear. Whereas the majority of these studies focus on determining the function of one miRNA during granulosal cell proliferation, more comprehensive studies are needed to improve our understanding of the way that interactions of the various miRNA, at different follicular stages, can modulate the proliferation of granulosal cells.
One of the hallmarks of follicular atresia is apoptosis of the granulosal cells. Several miRNA regulate apoptosis in cultured primary granulosal cells of various species (Table 1). Overexpression of miR-23a in human granulosal cells increases apoptosis by targeting XIAP and CASP3, resulting in CASP3 cleavage, which is a marker of apoptosis (Yang et al. 2012). miR-26b induces apoptosis in porcine granulosal cells by targeting SMAD4, both directly and indirectly through USP9X, which regulates the ubiquitination of SMAD4 (Liu et al. 2014a; Shen et al. 2014). miR-34a induces apoptosis of porcine granulosal cells by targeting INHBB expression (Tu et al. 2014). miR-21 may play a role in protection against apoptosis, because inhibition of miR-21 induces apoptosis in both mouse granulosal cells and mouse ovaries and decreases the ovulation rate (Carletti et al. 2010). miR-92a also inhibits apoptosis in porcine granulosal cells by targeting SMAD7 (Liu et al. 2014b).
In addition to supporting the growth and development of the oocyte, granulosal and thecal cells must produce estrogen to support uterine functions, regulate gonadotropin release, and elicit reproductive behaviors. miRNA that serve as regulators of steroidogenesis are summarized in Table 1. Examples of inhibitors of steroidogenesis are miR-378, which inhibits aromatase expression and estradiol production and suppresses the expression of PGR in cultured porcine granulosal cells (S. Xu et al. 2011; Toms et al. 2015), and miR-34a and miR-320, which inhibit estradiol release from human granulosal cells and murine ovaries, respectively (Sirotkin et al. 2009; Yin et al. 2014). In contrast, positive regulators of estradiol production in mouse granulosal cells are miR-383, which increases estradiol but does not affect progesterone release, and miR-132, which promotes estradiol synthesis via the translational repression of NURR1, a negative regulator of CYP19A1 (Yin et al. 2012, 2014; Wu et al. 2015). Another positive regulator of steroidogenesis is miR-320, which stimulates testosterone and progesterone in murine ovaries (Yin et al. 2014).
When considering the biology of the growing or atretic follicle, one might expect genes for proliferation and apoptosis to be inversely regulated by miRNA. However, some individual miRNA have the same effect on both proliferation- and apoptosis-related genes. These miRNA may be fine tuners of their target pathways, or their actions may depend on the presence of other miRNA. For example, the Let-7 family (Let-7b/c/d/g) miRNA decrease proteins associated with proliferation and apoptosis (Sirotkin et al. 2010). Support for a role of the Let-7 family in follicular function has been presented by Cao et al. (2015), who have shown that the expression of let-7a/b/c/i decreases, whereas the expression of let-7 g increases, in atretic porcine follicles. These observations have been followed by functional studies of the role of the Let-7 family in apoptosis. Cultured porcine granulosal cells have been cotransfected with Let-7a/b/c/i mimics or with Let-7 g mimic and assayed for apoptosis. As predicted by the expression data in tissues, overexpression of Let-7a/b/c/i reduces, whereas Let-7 g increases, the proportion of apoptotic cells compared with the negative control transfected group. Together, these studies provide strong evidence for a role of the Let-7 family of miRNA in granulosal cell proliferation, survival, apoptosis, and function. Clearly, miRNA serve to regulate the important functions of granulosal cells that will impact follicular development, but elucidation of the way that these miRNA work together to regulate translation of their many target mRNA will require functional studies of miRNA families and proteomic analyses to determine the physiological effects of the multiple miRNA that are present in these cells.
In addition to regulating genes in the cells of origin, recent studies have demonstrated the presence of extracellular miRNA in follicular fluid (Donadeu and Schauer 2013; Sang et al. 2013; Sohel et al. 2013; Santonocito et al. 2014; Da Silveira et al. 2015). miRNA occur in exosomal and nonexosomal fractions of the follicular fluid (Sohel et al. 2013; Santonocito et al. 2014), and their expression varies depending on the stage of follicular development (Sohel et al. 2013). Exosomal miRNA is probably taken up by follicular cells by endocytosis and results in the increased concentration of miRNA and modulation of targeted mRNA (Sohel et al. 2013; Da Silveira et al. 2015).
Much less work has been carried out to elucidate the regulation of miRNA expression in ovarian cells. Ovarian functions are under the control of the pituitary gonadotropins, follicle-stimulating hormone, luteinizing hormone, and in the case of rodents, prolactin. The gonadotropins stimulate steroidogenesis in granulosal, thecal, and luteal cells by the activation of enzymatic activity, the stimulation of cholesterol uptake, and the mobilization and maintenance of key structural and functional proteins. Thus, if miRNA can serve as modifiers of steroidogenesis, the expression of those miRNA must be under the control of the gonadotropins. In addition, transcription factors, cytokines, and other factors can control miRNA in preantral granulosal cells (Table 2).
miRNA in the corpus luteum
The role of miRNA in the CL was first evident when female Dicer d/d hypomorphic mice were found to be infertile because of impaired angiogenesis that resulted in a failure of luteal development (Otsuka et al. 2008). Expression of TIMP1 and luteal angiogenesis were partially restored by injection of miR-17-5p and let-7b (Otsuka et al. 2008). However, using a different hypomorphic Dicer1 model, Yang et al. (2005) observed impaired embryonic angiogenesis, rather than a defect in luteal development. Otsuka et al. (2008) suggested that angiogenesis in different tissues may have different sensitivities to concentrations of DICER1, because of the involvement of different miRNA in embryonic vs. luteal development. Moreover, the deletion of various elements of Dicer1 possibly results in diverse dysfunctional proteins with dissimilar phenotypic results. In both cases, homozygous embryos died, although in the latter model, one homozygous male survived (Otsuka et al. 2007), for reasons that the authors could not explain, and was used to generate the mouse line used in the study by Otsuka et al. (2008).
To date, few functional studies have been conducted on miRNA in the CL, but more is known about miRNA expression in the CL from profiling studies. In the ovine CL, the most abundantly expressed miRNA are Let-7a, Let-7b, miR-16b, miR-21, and mir-125b (McBride et al. 2012). The let-7 family and miR-21 are also among the most abundant miRNA in the bovine CL, together with miR-140, miR-199a-3p, and miR-320 (Maalouf et al. 2014). Notably, the let-7 family range from 0.2 to 4 million reads in the bovine CL (Maalouf et al. 2014), a far greater number than the 31–51 reads that have been observed in human cumulus cells by Assou et al. (2013). The greater expression of the let-7 family in the CL compared with the follicle suggests an important function in fully differentiated steroidogenic cells.
Comparisons of miRNA expression in early and midcycle CL (McBride et al. 2012; S.W. Maalouf et al., in preparation), in functional and regressing CL (Ma et al. 2011), and in CL of cyclic and pregnant cows (Maalouf et al. 2014) are paving the way toward an understanding of the numbers and types of miRNA that are involved in the regulation of the functional transitions that occur during the lifespan of the CL. McBride et al. (2012) have reported that nine miRNA decrease and eight miRNA increase during the follicular-luteal transition in the sheep (Fig. 2). One of the upregulated miRNA is miR-21, which is abundantly expressed in mature CL (Maalouf et al. 2014) and occurs in higher amounts in CL collected on day 10 compared with day 4 of the estrous cycle (S.W. Maalouf et al., in review), suggesting that miR-21 is involved in luteinization. In murine granulosal cells, miR-21 expression inhibits granulosal cell apoptosis (Fiedler et al. 2008; Carletti et al. 2010), indicating a role in survival, whereas expression in the late CL and corpus albicans (McBride et al. 2012) implies a role in apoptosis and regression. Because miRNA function may be time-, tissue-, and species-specific, a determination of whether the role of miR-21 in the regressing CL is opposite to that in murine granulosal cells will be of interest.
Stage-specific expression of miRNA in the corpus luteum (CL). Only those miRNA that have been found to be differentially expressed by stage in miRSeq or microarray analyses are depicted. Upregulated miRNA are listed in blue, and downregulated miRNA are shown in orange during the following steps: luteinization (McBride et al. 2012; Kitahara et al. 2013; Iwamune et al. 2014), maturation (McBride et al. 2012; Dai et al. 2014; S.W. Maalouf et al., in review), regression (Ma et al. 2011; McBride et al. 2012), and rescue (MRP maternal recognition of pregnancy; Maalouf et al. 2014)
Dai et al. (2014) have reported a significant increase in the miR-126 concentration in midcycle compared with early cycle bovine CL. This concentration is maintained in the late cycle CL but declines in the regressed CL, although the time after onset of luteolysis has not been determined in their study. In situ hybridization assays have shown miR-126 expression to be localized in luteal steroidogenic cells. Using bioinformatic analysis, Dai et al. (2014) have identified Talin2 (TLN2), a cytoskeletal protein involved in integrin signaling (Petrich 2009), as a predicted target of miR-126. Dual luciferase assays in NIH3T3 fibroblastic cells have shown that miR-126 binds to the wild-type, but not to a mutated sequence in the 3′UTR of TLN2 mRNA and reduces luciferase activity, indicating that TLN2 is a direct target for miR-126. Furthermore, the mRNA expression of TLN2 is greatest, whereas that of the TLN2 protein is least during midcycle CL. The authors (Dai et al. 2014) conclude that TLN2 expression is inversely correlated with that of mir-126-3p in the bovine CL and may play a role in regulating luteal function. We have also investigated miR-126 in the bovine CL, and similar to the results reported by Dai et al. (2014), miR-126 expression is upregulated on day 10 compared with day 4 CL. However, in our experiments, miR-126 expression is also present in luteal endothelial cells (S.W. Maalouf and J.L. Pate, unpublished observations).
Once fully differentiated, the structural features of the CL are maintained, and steroidogenesis continues at a high rate. Another important “switch” in luteal structure/function occurs at the end of the estrous cycle, when the CL is induced to undergo rapid luteolysis. To determine which miRNA impact the switch from luteal maintenance to regression, Ma et al. (2011) identified, on a microarray, seven miRNA that were greater in amount in midcycle bovine CL compared with regressed CL, and six miRNA that were greater in amount in regressed CL (Fig. 2). The functional state of these CL was estimated by visual evaluation of the ovaries at the time of collection. miR-378 exhibited the greatest fold change and was suggested to be important in luteal maintenance. The concentration of miR-378 was greater in mid- and late-cycle CL compared with regressed CL. Interferon gamma receptor 1 (IFNGR1), a predicted target of miR-378, was reduced in late-cycle CL and elevated in regressed CL. The negative correlation of miR-378 and IFNGR1 protein in late-cycle and regressed CL was interpreted as evidence for a regulatory role of miR-378 in the regulation of IFNG-related signaling pathways. IFNG is present in the CL and induces apoptosis in cultured luteal cells (Fairchild and Pate 1989; Petroff et al. 1999; Cannon and Pate 2006) and is suggested to mediate the effects of immune cells in regressing CL (Walusimbi and Pate 2013). Given the potential importance of IFNG signaling in luteal regression, functional studies to determine whether the receptor is a target of miR-378 are certainly warranted.
Prevention of luteolysis is a critical event in the establishment of pregnancy. We have hypothesized that maternal recognition of pregnancy includes changes in luteal miRNA that target pathways important for luteal survival or regression. The profiling of miRNA expression in the bovine CL on day 17 of the estrous cycle or pregnancy by using miRSeq resulted in 12 differentially expressed miRNA (Maalouf et al. 2014). Four were in greater amounts, and eight were in lower amounts in the CL of early pregnancy compared with those of the estrous cycle (Fig. 2). Predicted target analysis of the differentially expressed miRNA have revealed genes that are involved in apoptosis and regulation of the immune response, processes that must be targeted in the CL to prevent luteal regression (Atli et al. 2012; Poole and Pate 2012; Walusimbi and Pate 2013). MiR-122, one of the upregulated miRNA, is a key regulator of cholesterol and fatty acid metabolism in mouse liver (Esau et al. 2006), an inhibitor of breast cancer tumorigenesis by targeting IGF1R, and a potential regulator of the PI3K/AKT/mTOR signaling pathway (Wang et al. 2012). Another upregulated miRNA, MiR-302b, is an inhibitor of apoptosis in tumor cells (Chen et al. 2014; Ge et al. 2014). Retinoic acid induces the expression of miR-302b in human malignant glioma cells via the RAR alpha pathway (Chen et al. 2014). Therefore, during maternal recognition of pregnancy in the cow, changes occur in luteal miRNA that are consistent with a role in facilitation of luteal survival.
Maalouf et al. (2014) also reported the expression of 46 novel miRNA on day 17 bovine CL. Of these, 21 had been reported in other species but had not been previously reported for the cow, whereas 25 had not been previously reported in any species. Three of the novel miRNA were differentially expressed in CL of cyclic compared with pregnant animals (Fig. 2). Mir-10170 was downregulated in the CL of pregnancy, and miR-10178 was present in the CL of the cycle but absent during pregnancy. Conversely, miR-10163 was only present in the CL of pregnant animals. Target analysis of the differentially expressed novel miRNA also revealed apoptosis and immune response signaling as significant pathways (Maalouf et al. 2014). The role of these miRNA in luteal function and survival is the subject of current investigations underway in our laboratory. Figure 2 depicts the miRNA that are currently known to be expressed during specific stages of luteal development, maintenance, rescue, or regression.
Concluding remarks and future directions
miRNA can now be added to the list of endocrine, autocrine, and paracrine mediators that regulate ovarian function. Advancements in genetic tools and high-throughput sequencing technologies have allowed rapid advancement in the profiling of the expression of miRNA in the ovary from several species. Profiling studies have demonstrated that miRNA are expressed in all types of ovarian cells, and functional studies indicate roles in follicular and luteal development. The complex nature of miRNA target interaction, regulation, and function, however, pose challenges for functional studies. Whereas some miRNA appear single-handedly to regulate specific signaling pathways, most miRNA act in clusters and are fine tuners of cellular functions. In both cases, the function of miRNA is affected by the hormonal and cellular milieux. Hence, focusing on only one miRNA in functional studies becomes very challenging and may not result in significant measurable biological changes. A better approach will be to assess the function of a cluster of miRNA that are coexpressed and more likely to affect the functions of other miRNA. Therefore, an understanding of the mechanism(s) of action of miRNA in ovarian function requires global comprehension of the network of miRNA-target interactions within the milieux of hormones and growth factors that dominate ovarian function. Hence, profiling studies are important not only in order to draw a spatial and temporal map of miRNA expression in the ovary, but also to provide clues with regard to the function or regulation of miRNA.
In addition, much of our understanding about miRNA function is derived from mouse models. Although mouse models have been highly instrumental in revealing the role of miRNA in ovarian follicular development, species differences in miRNA expression and function may pose an obstacle to the application of miRNA diagnostics in medical and/or veterinary practices. In addition, because of the spatial and temporal regulation of miRNA, the identification of miRNA targets and functions may not translate well from diseased to healthy tissues with different tissue microenvironments. Novel genetic tools such as CRISPR and other technologies hold a promising future for attaining a deeper understanding of miRNA function in the ovary from other species.
Finally, the advancement in our knowledge of miRNA expression and function should help to fill in the gaps in the miRNA network analysis programs available and to improve their accuracy to predict bona fide targets and miRNA-target interactions. A better appreciation of miRNA function and miRNA signature in the ovary could have tremendous impacts on reproductive health by aiding the rapid diagnosis of reproductive disorders, the enhancement of embryo quality and survival during assisted reproductive technologies, and the design of novel contraceptive methods.
References
Abd El Naby WS, Hagos TH, Hossain MM, Salilew-Wondim D, Gad AY, Rings F, Cinar MU, Tholen E, Looft C, Schellander K, Hoelker M, Tesfaye D (2011) Expression analysis of regulatory microRNAs in bovine cumulus oocyte complex and preimplantation embryos. Zygote 21:31–51
Ahn HW, Morin RD, Zhao H, Harris RA, Coarfa C, Chen ZJ, Milosavljevic A, Marra MA, Rajkovic A (2010) MicroRNA transcriptome in the newborn mouse ovaries determined by massive parallel sequencing. Mol Hum Reprod 16:463–471
Assou S, Al-edani T, Haouzi D, Philippe N, Lecellier C-H, Piquemal D, Commes T, Ait-Ahmed O, Dechaud H, Hamamah S (2013) MicroRNAs: new candidates for the regulation of the human cumulus-oocyte complex. Hum Reprod 28:3038–3049
Atli MO, Bender RW, Mehta V, Bastos MR, Luo W, Vezina CM, Wiltbank MC (2012) Patterns of gene expression in the bovine corpus luteum following repeated intrauterine infusions of low doses of prostaglandin F2alpha. Biol Reprod 86:130
Baley J, Li J (2012) MicroRNAs and ovarian function. J Ovarian Res 5:8
Bartel DP (2004) MicrorNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bernstein E, Kim SY, Carmell MA, Murchison EP, Alcorn H, Li MZ, Mills AA, Elledge SJ, Anderson KV, Hannon GJ (2003) Dicer is essential for mouse development. Nat Genet 35:215–217
Brennecke J, Stark A, Russell RB, Cohen SM (2005) Principles of microRNA-target recognition. PLoS Biol 3:e85
Cannon MJ, Pate JL (2006) Indoleamine 2,3-dioxygenase participates in the interferon-gamma-induced cell death process in cultured bovine luteal cells. Biol Reprod 74:552–559
Cao R, Wu WJ, Zhou XL, Xiao P, Wang Y, Liu HL (2015) Expression and preliminary functional profiling of the let-7 family during porcine ovary follicle atresia. Mol Cells 38:304–311
Carletti MZ, Christenson LK (2009) MicroRNA in the ovary and female reproductive tract. J Anim Sci 87(14 Suppl):E29–E38
Carletti MZ, Fiedler SD, Christenson LK (2010) MicroRNA21 blocks apoptosis in mouse periovulatory granulosa cells. Biol Reprod 83:286–295
Chen PH, Shih CM, Chang WC, Cheng CH, Lin CW, Ho KH, Su PC, Chen KC (2014) MicroRNA-302b-inhibited E2F3 transcription factor is related to all trans retinoic acid-induced glioma cell apoptosis. J Neurochem 131:731–742
Christenson LK (2010) MicroRNA control of ovarian function. Anim Reprod 7:129–133
Czech B, Hannon GJ (2010) Small RNA sorting: matchmaking for Argonautes. Nat Rev Genet 12:19–31
Da Silveira JC, de Andrade GM, Nogueira MF, Meirelles FV, Perecin F (2015) Involvement of miRNAs and cell-secreted vesicles in mammalian ovarian antral follicle development. Reprod Sci (in press)
Dai L, Xu J, Liu S, Ma T, Zhu Y, Xu F, Gao Y, Yuan B, Wang S, Zhang Y, Sun G, Zhang J (2014) Characterization of miR-126-3p and its target Talin2 in the bovine corpus luteum during the oestrus cycle. Reprod Domest Anim 49:913–919
Di R, He J, Song S, Tian D, Liu Q, Liang X, Ma Q, Sun M, Wang J, Zhao W, Cao G, Wang J, Yang Z, Ge Y, Chu M (2014) Characterization and comparative profiling of ovarian microRNAs during ovine anestrus and the breeding season. BMC Genomics 15:899
Donadeu FX, Schauer SN (2013) Differential miRNA expression between equine ovulatory and anovulatory follicles. Domest Anim Endocrinol 45:122–125
Donadeu FX, Schauer SN, Sontakke SD (2012) Involvement of miRNA in ovarian follicular and luteal development. J Endocrinol 215:323–334
Du T, Zamore PD (2005) MicroPrimer: the biogenesis and function of microRNA. Development 132:4645–4652
Esau C, Davis S, Murray SF, Yu XX, Pandey SK, Pear M, Watts L, Booten SL, Graham M, McKay R, Subramaniam A, Propp S, Lollo BA, Freier S, Bennett CF, Bhanot S, Monia BP (2006) miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 3:87–98
Espey LL (1980) Ovulation as an inflammatory reaction: a hypothesis. Biol Reprod 22:73–106
Fairchild DL, Pate JL (1989) Interferon-gamma induction of major histocompatibility complex antigens on cultured bovine luteal cells. Biol Reprod 40:453–457
Fiedler SD, Carletti MZ, Hong X, Christenson LK (2008) Hormonal regulation of microRNA expression in periovulatory mouse mural granulosa cells. Biol Reprod 79:1030–1037
Fischer S, Handrick R, Aschrafi A, Otte K (2015) Unveiling the principle of microRNA-mediated redundancy in cellular pathway regulation. RNA Biol 12:238–247
Flemr M, Malik R, Franke V, Nejepinska J, Sedlacek R, Vlahovicek K, Svoboda P (2013) A retrotransposon-driven dicer isoform directs endogenous small interfering RNA production in mouse oocytes. Cell 155:807–816
Ge T, Yin M, Yang M, Liu T, Lou G (2014) MicroRNA-302b suppresses human epithelial ovarian cancer cell growth by targeting RUNX1. Cell Physiol Biochem 34:2209–2220
Gonzalez G, Behringer RR (2009) Dicer is required for female reproductive tract development and fertility in the mouse. Mol Reprod Dev 76:678–688
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524
Hasuwa H, Ueda J, Ikawa M, Okabe M (2013) miR-200b and miR-429 function in mouse ovulation and are essential for female fertility. Science 341:71–73
Hawkins SM, Matzuk MM (2010) Oocyte-somatic cell communication and microRNA function in the ovary. Ann Endocrinol (Paris) 71:144–148
Hong X, Luense LJ, McGinnis LK, Nothnick WB, Christenson LK (2008) Dicer1 is essential for female fertility and normal development of the female reproductive system. Endocrinology 149:6207–6212
Hossain MM, Ghanem N, Hoelker M, Rings F, Phatsara C, Tholen E, Schellander K, Tesfaye D (2009) Identification and characterization of miRNAs expressed in the bovine ovary. BMC Genomics 10:443
Hossain MM, Sohel MM, Schellander K, Tesfaye D (2012) Chracterization and importance of microRNAs in mammalian gonadal functions. Cell Tissue Res 349:679–690
Huang J, Ju Z, Li Q, Hou Q, Wang C, Li J, Li R, Wang L, Sun T, Hang S, Gao Y, Hou M, Zhong J (2011) Solexa sequencing of novel and differentially expressed microRNAs in testicular and ovarian tissues in Holstein cattle. Int J Biol Sci 7:1016–1026
Imbar T, Eisenberg I (2014) Regulatory role of microRNAs in ovarian function. Fertil Steril 101:1524–1530
Iwamune M, Nakamura K, Kitahara Y, Minegishi T (2014) MicroRNA-376a regulates 78-kilodalton glucose-regulated protein expression in rat granulosa cells. PLoS One 9:e108997
Jiang L, Huang J, Li L, Chen Y, Chen X, Zhao X, Yang D (2015) MicroRNA-93 promotes ovarian granulosa cells proliferation through targeting CDKN1A in polycystic ovarian syndrome. J Clin Endocrinol Metab 100:E729–E738
Kitahara Y, Nakamura K, Kogure K, Minegishi T (2013) Role of microRNA-136-3p on the expression of luteinizing hormone-human chorionic gonadotropin receptor mRNA in rat ovaries. Biol Reprod 89:114
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294:853–858
Lau NC, Lim LP, Weinstein EG, Bartel DP (2001) An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294:858–862
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854
Lei L, Jin S, Gonzalez G, Behringer RR, Woodruff TK (2010) The regulatory role of Dicer in folliculogenesis in mice. Mol Cell Endocrinol 315:63–73
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120:15–20
Li M, Liu W, Wang T, Guan J, Luo Z, Chen H, Wang X, Chen L, Ma J, Mu Z, Jiang AA, Zhu L, Lang Q, Zhou X, Wang J, Zeng W, Li N, Li K, Gao X, Li X (2011) Repertoire of porcine microRNAs in adult ovary and testis by deep sequencing. Int J Biol Sci 7:1045–1055
Li Y, Fang Y, Liu Y, Yang X (2015) MicroRNAs in ovarian function and disorders. J Ovarian Res 8:51–58
Liang M, Yao G, Yin M, Lu M, Tian H, Liu L, Lian J, Huang X, Sun F (2013) Transcriptional cooperation between p53 and NF-kB p65 regulates microRNA-224 transcription in mouse ovarian granulosa cells. Mol Cell Endocrinol 370:119–129
Liang Y, Ridzon D, Wong L, Chen C (2007) Characterization of microRNA expression profiles in normal human tissues. BMC Genomics 8:166
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, Johnson JM (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433:769–773
Ling YH, Ren CH, Guo XF, Xu LN, Huang YF, Luo JC, Zhang YH, Zhang XR, Zhang ZJ (2014) Identification and characterization of microRNAs in the ovaries of multiple and uniparous goats (Capra hircus) during follicular phase. BMC Genomics 15:339
Liu J, Du X, Zhou J, Pan Z, Liu H, Li Q (2014a) MicroRNA-26b functions as a proapoptotic factor in porcine follicular granulosa cells by targeting Sma- and Mad- related protein 4. Biol Reprod 91:146
Liu J, Yao W, Yao Y, Du X, Zhou J, Ma B, Liu H, Li Q, Pan Z (2014b) MiR-92a inhibits porcine ovarian granulosa cell apoptosis by targeting Smad7 gene. FEBS Lett 588:4497–4503
Luense LJ, Carletti MZ, Christenson LK (2009) Role of dicer in female fertility. Trends Endocrinol Metab 20:265–272
Lytle JR, Yario TA, Steitz JA (2007) Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5′UTR as in the 3′UTR. Proc Natl Acad Sci U S A 104:9667–9672
Ma T, Jiang H, Gao Y, Zhao Y, Dai L, Xiong Q, Xu Y, Zhao Z, Zhang J (2011) Microarray analysis of differentially expressed microRNAs in non-regressed and regressed bovine corpus luteum tissue; microRNA-378 may suppress luteal cell apoptosis by targeting the interferon gamma receptor 1 gene. J Appl Genet 52:481–486
Maalouf SW, Liu W-S, Albert I, Pate JL (2014) Regulating life or death: potential role of microRNA in rescue of the corpus luteum. Mol Cell Endocrinol 398:78–88
Matzuk MM, Burns KH, Viveiros MM, Eppig JJ (2002) Intercellular communication in the mammalian ovary: oocytes carry the conversation. Science 296:2178–2180
McBride D, Carré W, Sontakke SD, Hogg CO, Law A, Donadeu FX, Clinton M (2012) Identification of miRNAs associated with the follicular-luteal transition in the ruminant ovary. Reproduction 144:221–233
McGinnis LK, Luense LJ, Christenson LK (2015) MicroRNA in ovarian biology and disease. Cold Spring Harb Perspect Med 5.pii:a022962
Miles JR, McDaneld TG, Wiedmann RT, Cushman RA, Echternkamp SE, Vallet JL, Smith TP (2012) MicroRNA expression profile in bovine cumulus-oocyte complexes: possible role of let-7 and miR-106a in the development of bovine oocytes. Anim Reprod Sci 130:16–26
Mishima T, Takizawa T, Luo SS, Ishibashi O, Kawahigashi Y, Mizuguchi Y, Ishikawa T, Mori M, Kanda T, Goto T, Takizawa T (2008) MicroRNA (miRNA) cloning analysis reveals sex differences in miRNA expression profiles between adult mouse testis and ovary. Reproduction 136:811–822
Murchison EP, Stein P, Xuan Z, Pan H, Zhang MQ, Schultz RM, Hannon GJ (2007) Critical roles for dicer in the female germline. Genes Dev 21:682–693
Nagaraja AK, Andrewu-Vieyra C, Frnaco HL, Ma L, Chen R, Han DY, Zhu H, Agno JE, Gunaratne PH, DeMayo FJ, Matzuk MM (2008) Deletion of dicer in somatic cells of the female reproductive tract causes sterility. Mol Endocrinol 22:2336–2352
Nothnick WB (2012) The role of micro-RNAs in the female reproductive tract. Reproduction 143:559–576
Okamura K, Phillips MD, Tyler DM, Duan H, Chou YT, Lai EC (2008) The regulatory activity of microRNA* species has substantial influence on microRNA and 3′UTR evolution. Nat Struct Mol Biol 15:354–363
Orom UA, Nielsen FC, Lund AH (2008) MicroRNA-10a binds the 5′UTR of ribosomal protein mRNAs and enhances their translation. Cell 30:460–471
Otsuka M, Jing Q, Georgel P, New L, Chen J, Mois J, Kang YJ, Jiang Z, Du X, Cook R, Das SC, Pattnaik AK, Beutler B, Han J (2007) Hypersusceptibility to vesicular stomatitis virus infection in Dicer-1-deficient mice is due to impaired miR24 and miR93 expression. Immunity 27:123–134
Otsuka M, Zheng M, Hayashi M, Lee J-D, Yoshino O, Lin S, Han J (2008) Impaired microrNA processing causes corpus luteum insufficiency and infertility in mice. J Clin Invest 118:1944–1954
Pan B, Toms D, Shen W, Li J (2015) MicroRNA-378 regulates oocyte maturation via the suppression of aromatase in porcine cumulus cells. Am J Physiol Endocrinol Metab 308:E525–E534
Pasquinelli AE, Reinhart BJ, Slack F, Martindale MQ, Kuroda MI, Maller B, Hayward DC, Ball EE, Degnan B, Muller P, Spring J, Srinivasan A, Frishman M, Finnerty J, Corbo J, Levine M, Leahy P, Davidson E, Ruvkun G (2000) Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408:86–89
Pate JL, Landis Keyes P (2001) Immune cells in the corpus luteum: friends of foes? Reproduction 122:665–676
Petrich BG (2009) Talin-dependent integrin signaling in vivo. Thromb Haemost 101:1020–1024
Petroff MG, Petroff BK, Pate JL (1999) Expression of cytokine messenger ribonucleic acids in the bovine corpus luteum. Endocrinology 140:1018–1021
Place RF, Li LC, Pookot D, Noonan EJ, Dahiya R (2008) MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc Natl Acad Sci U S A 105:1608–1613
Poole DH, Pate JL (2012) Luteal microenvironment directs resident T lymphocyte function in cows. Biol Reprod 86:29
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE, Horvitz HR, Ruvkun G (2000) The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403:901–906
Ro S, Song R, Park C, Zheng H, Sanders KM, Yan W (2007) Cloning and expression profiling of small RNAs expressed in the mouse ovary. RNA 13:2366–2380
Salilew-Wondim D, Ahmad I, Gebremedhn S, Sahadevan S, Hossain MM, Rings F, Hoelker M, Tholen E, Neuhoff C, Looft C, Schellander K, Tesfaye D (2014) The expression pattern of microRNAs in granulosa cells of subordinate and dominant follicles during the early luteal phase of the bovine estrous cycle. PLoS One 9:e106795
Sang Q, Yao Z, Wang H, Feng R, Wang H, Zhao X, Xing Q, Jin L, He L, Wu L, Wang L (2013) Identification of microRNAs in human follicular fluid: characterization of microRNAs that govern steroidogenesis in vitro and are associated with polycystic ovary syndrome in vivo. J Clin Endocrinol Metab 98:3068–3079
Santonocito M, Vento M, Guglielmino MR, Battaglia R, Wahlgren J, Ragusa M, Barbagallo D, Borzi P, Rizzari S, Maugeri M, Scollo P, Tatone C, Valadi H, Purello M, Di Pietro C (2014) Molecular characterization of exosomes and their microRNA cargo in human follicular fluid: bioinformatics analysis reveals that exosomal microRNAs control pathways involved in follicular maturation. Fertil Steril 102:1751–1761
Schauer SN, Sontakke SD, Watson ED, Esteves CL, Donadeu FX (2013) Involvement of miRNAs in equine follicle development. Reproduction 146:273–282
Sen A, Prizant H, Light A, Biswas A, Hayes E, Lee HJ, Barad D, Gleicher N, Hammes SR (2014) Androgens regulate ovarian follicular development by increasing follicle stimulating hormone receptor and microRNA-125b expression. Proc Natl Acad Sci U S A 111:3008–3013
Shen G, Lin Y, Yang X, Zhang J, Xu Z, Jia H (2014) MicroRNA-26b inhibits epithelial-mesenchymal transition in hepatocellular carcinoma by targeting USP9X. BMC Cancer 14:393
Sirotkin AV, Ovcharenko D, Grossmann R, Laukova M, Mlyncek M (2009) Identification of microRNAs controlling human ovarian cell steroidogenesis via a genome-scale screen. J Cell Physiol 219:415–420
Sirotkin AV, Laukova M, Ovcharenko D, Brenaut P, Mlyncek M (2010) Identification of microRNAs controlling human ovarian cell proliferation and apoptosis. J Cell Physiol 223:49–56
Sirotkin AV, Alexa R, Kisova G, Harrath AH, Alwasel S, Ovcharenko D, Mlyncek M (2014) MicroRNAs control transcription factor NF-kB (p65) expression in human ovarian cells. Funct Integr Genomics 15:271–275
Sohel MM, Hoelker M, Noferesti SS, Salilew-Wondim D, Tholen E, Looft C, Rings F, Uddin MJ, Spencer TE, Schellander K, Tesfaye D (2013) Exosomal and non-exosomal transport of extra-cellular microRNAs in follicular fluid: implications for bovine oocyte developmental competence. PLoS One 8:e78505
Sontakke SD, Mohammed BT, McNeilly AS, Donadeu FX (2014) Characterization of microRNAs differentially expressed during bovine follicle development. Reproduction 148:271–283
Sood P, Krek A, Zavolan M, Macino G, Rajewsky N (2006) Cell-type-specific signatures of microRNAs on target mRNA expression. Proc Natl Acad Sci U S A 103:2746–2751
Spencer TE, Bazer FW (2004) Conceptus signals for establishment and maintenance of pregnancy. Reprod Biol Endocrinol 2:49
Stein P, Rozhkov NV, Li F, Cardenas FL, Davydenk O, Le V, Gregory BD, Hannon GJ, Schultz RM (2015) Essential role for endogenous siRNAs during meiosis in mouse oocytes. PLoS Genet 11:e1005013
Tesfaye D, Worku D, Rings F, Phatsara C, Tholen E, Schellander K, Hoelker M (2009) Identification and expression profiling of microRNAs during bovine oocyte maturation using heterologous approach. Mol Reprod Dev 76:665–677
Toloubeydokhti T, Bukulmez O, Chegini N (2008) Potential regulatory functions of microRNAs in the ovary. Semin Reprod Med 26:469–478
Toms D, Xu S, Pan B, Wu D, Li J (2015) Progesterone receptor expression in granulosa cells is suppressed by microRNA-378-3p. Mol Cell Endocrinol 399:95–102
Tripurani SK, Xiao C, Salem M, Yao J (2010) Cloning and analysis of fetal ovary microRNAs in cattle. Anim Reprod Sci 120:16–22
Troppmann B, Kossack N, Nordhoff V, Schuring AN, Gromoll J (2014) MicroRNA miR-513a-3p acts as co-regulator of luteinizing hormone/chorionic gonadotropin receptor gene expression in human granulosa cells. Mol Cell Endocrinol 390:65–72
Tu F, Pan ZX, Yao Y, Liu HL, Liu SR, Xie Z, Li QF (2014) MiR-34a targets the inhibin beta B gene, promoting granulosa cell apoptosis in the porcine ovary. Genet Mol Res 13:2504–2512
Vidigal JA, Ventura A (2015) The biological functions of miRNAs: lessons from in vivo studies. Trends Cell Biol 25:137–147
Walusimbi SS, Pate JL (2013) Physiology and endocrinology symposium: role of immune cells in the corpus luteum. J Anim Sci 91:1650–1659
Wang B, Wang H, Yang Z (2012) MiR-122 inhibts cell proliferation and tumorigenesis of breast cancer by targeting IGF1R. PLoS One 7:e47053
Wightman B, Ha I, Ruvkun G (1993) Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75:855–862
Wu S, Sun H, Zhang Q, Jiang Y, Fang T, Cui I, Yan G, Hu Y (2015) MicroRNA-132 promotes estradiol synthesis in ovarian granulosa cells via translational repression of Nurr1. Reprod Biol Endocrinol 13:94
Xu B, Hua J, Zhang Y, Jiang X, Zhang H, Ma T, Zheng W, Sun R, Shen W, Sha J, Cooke HJ, Shi Q (2011) Proliferating cell nuclear antigen (PCNA) regulates primordial follicle assembly by promoting apoptosis of oocytes in fetal and neonatal mouse ovaries. PLoS One 6:e16046
Xu B, Zhang U-W, Tong X-H, Liu Y-S (2015) Characterization of miNRA profile in human cumulus granulosa cells: identification of microRNAs that regulate Notch signaling and are associated with PCOS. Mol Cell Endocrinol 404:26–36
Xu S, Linher-Melville K, Yang BB, Wu D, Li J (2011) Micro-RNA378 (miR-378) regulates ovarian estradiol production by targeting aromatase. Endocrinology 152:3941–3951
Yang JS, Lai EC (2011) Alternative miRNA biogenesis pathways and the interpretation of core miRNA pathway mutants. Mol Cell 43:892–903
Yang S, Wang S, Luo A, Ding T, Lai Z, Shen W, Ma X, Cao C, Shi L, Jiang J, Rong F, Ma L, Tian Y, Du X, Lu Y, Li Y, Wang S (2013) Expression patterns and regulatory functions of microRNAs during the initiation of primordial follicle development in the neonatal mouse ovary. Biol Reprod 89:126
Yang WJ, Yang DD, Na S, Sandusky GE, Zhang Q, Zhao G (2005) Dicer is required for embryonic angiogenesis during mouse development. J Biol Chem 280:9330–9335
Yang X, Zhou Y, Peng S, Wu L, Lin HY, Wang S, Wang H (2012) Differentially expressed plasma microRNAs in premature ovarian failure patients and the potential regulatory function of miR-23a in granulosa cell apoptosis. Reproduction 144:235–244
Yao G, Yin M, Lian J, Tian H, Liu L, Li X, Sun F (2010) MicroRNA-224 is involved in transforming growth factor-beta-mediated mouse granulosa cell proliferation and granulosa cell function by targeting Smad4. Mol Endocrinol 24:540–551
Yin M, Lu M, Yao G, Tian H, Lian J, Liu L, Liang M, Wang Y, Sun F (2012) Transactivation of microRNA-383 by steroidogenic factor-1 promotes estradiol release from mouse ovarian granulosa cells by targeting RBMS1. Mol Endocrinol 26:1129–1143
Yin M, Wang X, Yao G, Lu M, Liang M, Sun Y, Sun F (2014) Transactivation of microRNA-320 by microrNA-383 regulates granulosa cell functions by targeting E2F1 and SF-1 proteins. J Biol Chem 289:18239–18257
Yuan S, Ortogero N, Wu Q, Zheng H, Yan W (2014) Murine follicular development requires oocyte dicer but not drosha. Biol Reprod 91:39
Zhang H, Jiang X, Zhang Y, Xu B, Hua J, Ma T, Zheng W, Sun R, Shen W, Cooke HJ, Hao Q, Qiao J, Shi Q (2014) microRNA 376a regulates follicle assembly by targeting Pcna in fetal and neonatal mouse ovaries. Reproduction 148:43–54
Zhang J, Ji X, Zhou D, Li Y, Lin J, Liu J, Luo H, Cui S (2013) MiR-143 is critical for the formation of primordial follicles in mice. Front Biosci (Landmark Ed) 18:588–597
Zhang Q, Sun H, Jiang Y, Ding L, Wu S, Fang T, Yan G, Hu Y (2013) MicroRNA-181a suppresses mouse granulosa cell proliferation by targeting activin receptor IIA. PLoS One 8:e59667
Zhao H, Rajkovic A (2008) MicroRNAs and mammalian ovarian development. Semin Reprod Med 26:461–468
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maalouf, S.W., Liu, W.S. & Pate, J.L. MicroRNA in ovarian function. Cell Tissue Res 363, 7–18 (2016). https://doi.org/10.1007/s00441-015-2307-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-015-2307-4