Abstract
Bombyx mori Nucleopolyhedrovirus (BmNPV), which is a member of the Baculoviridae family, is a significant pathogen of the silkworm. The infection of BmNPV is often lethal and causes about 20% loss of cocoon in the silk industry annually. To explore the effects of different gene inhibition strategies on the replication cycle of baculovirus, we constructed the mutant virus to infect BmN cells directly and further identified ie0, ie1, and gp64 as the essential viral genes of BmNPV. To elucidate the significance of the inhibition effect of different interference strategies, we characterized and constructed the recombinant BmNPV that carried a single or multigene-interfering cassette. The results showed that the inhibition effect of dsie1 on target gene expression, virus titer, and silkworm mortality was significantly better than that of dsie0 and dsgp64. It also showed that the dsie1 interference produced fewer progeny virions and was less lethal, which indicates that ie1 played a more critical role in the BmNPV replication cycle. Furthermore, the inhibitory effect of the virus titer and mortality indicated that the multigene co-interference constructed by the baculovirus expression system was significantly better than the interference of any single-gene (p < 0.05). In summary, the strategy of multigene synergy can achieve the function of continuous interference and provide a new platform for the breeding of silkworm disease resistant. In addition, this strategy improves the various traits of the silkworm.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bombyx mori nucleopolyhedrovirus (BmNPV) is the primary pathogen affecting silkworm cocoon production in Asia, and its infection causes about 20% of silkworm cocoon loss annually (Jiang and Xia 2014; Chen et al. 2017). Compared with other viruses, BmNPV is very stable because it is encapsulated by strong polyhedrin proteins that prevent the penetration against low pH and drug from reaching viral particles. As a result of the deepening research of the BmNPV, the viral genome sequence has been thoroughly uncovered (Xia et al. 2004; Cheng et al. 2008; Choi and Guarino 1995; Guarino and Summers 1986; Kovacs et al. 1992). Baculoviruses virions are present as two types, budded virions (BV) and occluded virions (ODV). They are similar in their nucleocapsid structure, but different in the origin and composition of their envelopes and their roles in the virus life cycle. BV virions spreads the infection throughout the insect and ODV virions in the form of inclusion bodies that spreads the infection between the insects (Jiang et al. 2012; Gomi et al. 1997, 1999; Lu and Iatrou 1997; Subbaiah et al. 2013). The ie0 gene, which encoded the IE-0 protein, is mainly distributed in the cytoplasm and has a significant regulatory effect on the RNA transcription of vp39, lef9 and P10 genes of the virus (de Jong et al. 2011; Kuzio et al. 1984; Rankin et al. 1986). This regulatory effect indicates that the gene has a vital activation function for viral gene transcription and viral infection (Senkevich et al. 1994). The ie1 gene is one of the earliest discovered genes of baculovirus, and it is an early gene transactivator and advanced expression factor (Jiang et al. 2013). The encoded IE-1 protein, which is mainly distributed in the DNA replication center of BmNPV in the nucleus, is an essential protein for baculovirus replication (Dai et al. 2004), and its function involves transcriptional activation of DNA replication, DNA damage, and apoptosis regulation (Lu and Iatrou 1997; Nagai et al. 2011; Nagamine et al. 2011). BmNPV envelope glycoprotein GP64 is a type I integral membrane protein, which is presented on the surface of infected cells and baculoviruses (Kadlec et al. 2008; Oomens et al. 1995). GP64, the most abundant proteins found associated with the process of BmNPV entry by endocytosis, is a fatty acid-acylated glycoprotein and a low-pH activated envelope fusion protein, and it is necessary for the process of the virus invading host and budding (Blissard and Wenz 1992; Li et al. 2011; Rahman and Gopinathan 2003).
The baculovirus multigene expression system (BMES) is the most widely used eukaryotic expression system, and it has the advantages of high expression level and excellent processing mechanism for post-translational modifications (Carbonell et al. 1985; Mena and Kamen 2011). Having transposable site cre-loxp and mini-Tn7, BMES can be used to accomplish the co-expression of multiple complex proteins so that it has been widely used in the field of gene delivery and biopesticides (Qi et al. 2016; Sun et al. 2010; Yao et al. 2012a, b; Zheng et al. 2018). RNA interference (RNAi) is a mechanism that explicitly silences the expression of a target gene. This process mainly depends on the endogenous or exogenous double-stranded RNA (dsRNA) mediated specific degradation of mRNA in the cell, resulting in the inhibition of the corresponding target gene expression and functional deletion (Fire et al. 1998; Hannon 2002). The occurrence of RNAi can be divided into three phase (1) the dsRNA is uniformly cut by RNase III (Dicer) into small-interfering RNA (siRNA) with a size of 21–25 nt in the cell (Bernstein et al. 2001; Zamore et al. 2000); (2) the RNA-induced silencing complex (RISC) is composed of siRNA and endonucleases, where the nucleic acid part plays a targeted role, and the protein part acts to degrade mRNA during the effect phase; (3) the targeted siRNA combined with the mRNA to form dsRNA again under the action of RNA-dependent RNA polymerase (RdRP) (Dougherty and Parks 1995). The newly formed dsRNA is further cut into siRNA by Dicer and enters the next cycle to achieve the purpose of amplification of interference effects (McIntyre and Fanning 2006; Saleh et al. 2009). Several reports demonstrated that the RNAi had been used to inhibit the activity of corresponding homologous genes (Elbashir et al. 2001a, b; Fire et al. 1998; Hammond et al. 2001). Currently, the RNAi has been widely used in eukaryote such as plants, fungi, insects, metazoans and mammals (Hammond et al. 2001; Isobe et al. 2004; Valdes et al. 2003), and it also has a broad application in viral infection (Isobe et al. 2004; Valdes et al. 2003).
The RNAi has been widely used in the anti-virus strategy of the silkworm. However, it has not had a stable and robust anti-BmNPV protective effect on the virus yet. Improving the effect and stability of RNAi is an urgent problem to be solved. In this study, single-gene and multigene interference cassettes were driven by the cytoplasmic actin 3 (A3) promoter of the silkworm, and the recombinant baculovirus that carries different interfering cassettes were constructed successfully by BMES. Consequently, the interference effect of the multigene combination will provide a theoretical basis strategy for the breeding of the silkworm resistant to BmNPV and improve the various characteristic of the silkworm.
Materials and methods
Bacterial strains, plasmids, viral Bacmid, reagents and larvae
Escherichia coli DH10B, BW23474, and TOP10 were used for the propagation of BmBacmid, R6kγ origin derived plasmids and other general plasmids, respectively. E. coli SW106 BmBacmid containing BmBacmid, pHelper, and pGB2Ωinv was constructed previously (Yao et al. 2010, 2012). Plasmids pFBDM-A3 and pUCDM-A3 were provided by Lunguang Yao (Nanyang Normal University, China), which contained a constitutive A3 promoter upstream of the multiple clone sites (MCS), as well as the modified BmBacmid with gentamycin or chloramphenicol resistance gene by mini-Tn7 or cre-loxp transposition (Sun et al. 2009; Yao et al. 2010, 2012a, b). The genes of ie0, ie1, and gp64 were amplified from genomic DNA of the BmNPV and constructed the recombinant donor vectors. The enhanced green fluorescent protein (egfp) and mCherry fluorescent protein gene were used as reporter genes. Pfu Taq, restriction enzymes, and T4 DNA ligase were purchased from NEB (New England Biolabs, England), while dl-α-ε Diaminopimelic acid (DAP) was bought from Sigma (cat.D1377, USA). Luria Bertani (LB) medium (10 g of tryptone, 10 g of NaCl and 5 g of yeast extract in 1 L of broth, pH 7.5) was used for cloning and growing the plasmids. Bombyx mori (BmN) cells were maintained at 27 °C in Grace’s cell culture medium (Grace’s medium) supplemented with 10% FBS (Gibco). Silkworm variety is 9·Fu × 7·Xiang.
Construction of donor vectors
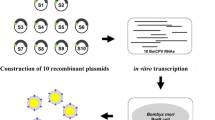
Donor vectors (Fig. 1) were used to construct recombinant baculoviruses expressing different RNAi cassettes under the control of the A3 promoter. In brief, the donor vector pUCDM-A3-dsie0-IG (Fig. 1a) was used as the backbone in ie0-triggered gene silencing. The formal interference gene of ie0 (amplified with primer dsie0-F and dsie0-R, Table 1) was cloned into plasmid pUCDM-A3 between XmaI and XhoI under the control of the A3 promoter. The anti-form interference gene of ie0 (also amplified with primer dsie0-F and dsie0-R, Table 1) was cloned between XhoI and NdeI to the downstream of the formal ie0. The interference genes by the formal and anti-form ie0 were constituted the RNAi cassette dsie0. The ires-dependent enhanced green fluorescent reporter gene expression cassette (IG, amplified with primer ig-F and ig-R, Table 1) was ligated to the downstream of the dsie0 by NdeI and SphI. For the donor vector pFBDM-A3-dsie1-IM (Fig. 1b), primers dsie1-F and dsie1-R (Table 1) were used to amplify the dsie1 by BmNPV genome, the formal gene ie1 was ligated into the BamHI/SacI and the anti-form gene ie1 was ligated into the SacI/XbaI, respectively. The ires-dependent mCherry fluorescent reporter gene expression cassette (IM, amplified with primer im-F and im-R, Table 1) was ligated to the downstream of the dsie1 by XbaI and PstI. The RNAi cassette dsgp64 was amplified with primer dsgp64-F and dsgp64-R (Table 1) and cloned into the pFBDM-A3 via XmaI/XhoI (formal gene gp64) and XhoI/NdeI (anti-form gene gp64) to the downstream of the promoter A3. The IM was ligated to the downstream of the dsgp64 by NdeI and SphI to form donor vector pFBDM-A3-dsgp64-IY (Fig. 1c). The donor vector pFBDM-A3-dsie1-IM was digested with SpeI and XbaI to release DNA A3-dsie1, and the fragment was cloned into the pFBDM-A3-dsie1-IM via SpeI and XbaI to construct donor vector pFBDM-A3-dsie1-A3-dsgp64-IM (Fig. 1d).
The RNAi cassette on recombinant donor vector. a The map of pUCDM-A3-dsie0-IG shows an RNAi cassette including promoter A3, interference gene ie0 (dsie0), and ires-dependent egfp (IG) expression cassette. b The map of pFBDM-A3-dsie1-IM shows an RNAi cassette including promoter A3, interference gene ie1 (dsie1), and ires-dependent mCherry (IM) expression cassette. c The map of pFBDM-A3-dsgp64-IM shows an RNAi cassette including interference gene gp64 (dsgp64) and IM expression cassette are driven by A3. d Two RNAi cassettes on recombinant donor vector pFBDM-A3-dsie1-A3-dsgp64-IM, which contains the interference gene fragment of dsgp64 and dsie1, and the fluorescent reporter gene expression cassette IM was ligated to the downstream of the dsgp64
Introduction of RNAi cassette into BmBacmid
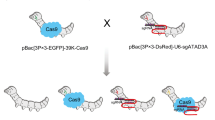
The RNAi cassettes A3-dsie1-IM (Fig. 2b), A3-dsgp64-IM (Fig. 2c), and A3-dsie0-IG (Fig. 2d), were introduced into the E. coli SW106 BmBacmid (Fig. 2a) by mini-Tn7 and cre-loxp transposition as described in a previous study, respectively (Sun et al. 2009; Yao et al. 2012a, b, 2007). The positive clones grown on Cm (Gm)/Kan/Tet/Spe/DAP were screened by white–blue plaque selection and PCR screening (Zheng et al. 2018), to generate three kinds of recombinant BmBacmids, including BmBacmid-dsie1-IM (Fig. 2f), BmBacmid-dsgp64-IM (Fig. 2g), BmBacmid-dsie0-IG (Fig. 2h). The A3-dsgp64-IM-A3-dsie1 (Fig. 2e) was introduced into asd-deletion type of E. coli SW106, which contained BmBacmid-A3-dsie0-IG by mini-Tn7 transposition, to construct the recombinant BmBacmid-dsie0-dsie1-dsgp64-IG-IM (Fig. 2i).
Construction of RNAi cassette-bearing recombinant BmBacmid and baculoviruses. aE. coli BmMultiBac/Δasd Sw106/PGB2ΩInv. b–e RNAi and fluorescent reporter gene expression cassettes were driven by promoter A3 (bdsie1 + IM. cdsgp64 + IM. ddsie0 + IG. edsie1 + dsgp64 + IM). f–i recombinant BmBacmid (fdsie1 + IM. gdsgp64 + IM. hdsie0 + IG. Idsie0 + dsie1 + dsgp64 + IG + IM). j–n observation of insect BmN cells infected by recombinant BmBacmid, Bar 100 µm (j BmBacmid-dsie1-IM, attendance at 512 nm. k BmBacmid-dsgp64-IM, attendance at 512 nm. l BmBacmid-dsieo-IG, attendance at 488 nm. m BmBacmid-dsie0-dsie1-dsgp64-IG-IM, attendance at 488 nm. n BmBacmid-dsie0-dsie1-dsgp64-IG-IM, attendance at 512 nm). o merge by m + n. p–s purified recombinant baculoviruses were observed by TEM, Bar 0.2 µm (p BmNPV-dsie1-IM. q BmNPV-dsgp64-IM. r BmNPV-dsie0-IG. s BmNPV-dsie0-dsie1-dsgp64-IG-IM)
Production of recombinant BmNPV
The E. coli SW106 containing recombinant BmBacmid with the different RNAi cassettes (dsie0, dsie1, dsgp64, and dsie0 + dsie1 + dsgp64) were cultured until the OD600 = 0.5–1 (attendance at 600 nm). These bacteria were collected by centrifuged (3000g) and resuspended in serum-free insect medium. The bacterial suspension was adjusted to different densities (105–108 cells/mL) with serum-free Grace’s medium (Yao et al. 2012a, b). BmN cells were cultured overnight in a 12-well plate until the cell density was approximately 70–80%. The supernatant was discarded, and different concentrations of bacteria were added to the corresponding wells. After cultured at 28 °C for 4–5 h, the bacteria in each well were washed out by serum-free Grace’s medium, then 500 µL of fresh Grace’s medium (with 10% FBS and 0.075% of penicillin) was added and incubated for 4–5 days post infected (d.p.i.). BmN cells were infected successfully by examining the fluorescence in the corresponding well by fluorescence microscope (Eclipse Ti-S, Nikon, Japan). The supernatant was collected and infected again with BmN cells. When the fluorescence appeared again after 3–4 d.p.i, the recombinant baculovirus was successfully constructed and distributed in the cell supernatant.
Titer determination of BmNPV
The early-exponential-phase BmN cells were diluted to 106 cells/mL with Sf-900 II serum-free medium (SFM), and then 100 µL working volume was placed into the different microwell in a 96-well plate. Serial baculoviral dilutions were also prepared and diluted in serial 10-fold dilutions to 10− 8 by SFM. The cell culture medium was removed and 100 µL of each virus dilutions was added (12-well per dilution) to the cell monolayer and then incubated at 28 °C. Following 4 h incubation, the supernatant in each well was replaced with 100 µL fresh insect medium. Plates were checked daily once for 4–5 days until the fluorescence was observed to reach maximum. The baculovirus titers were expressed as the 50% tissue culture infective dose (TCID50) according to the standard method of Reed and Muench (Rattanarojpong et al. 2016; Wang et al. 2018).
Baculovirus purification and transmission electron microscope
The supernatant of silkworm-derived BmN cells infected with recombinant baculovirus (MOI = 1) was collected at 4 d.p.i. Baculovirus in the supernatant was purified by two rounds of sucrose gradient ultracentrifugation according to standard methods (O’Reilly 1997).
Purified baculoviruses were adsorbed onto glow discharge-activated carbon coated grids for 2 min. Moreover, the sample-coated grids were washed three times with distilled water, following a negative staining with 1% uranyl acetate for 45 s. Images were acquired using the FEI Talos F200S transmission electron microscope (TEM).
Quantitative PCR
Total RNA was extracted from infected BmN cells by Trizol Reagent (Invitrogen Inc., Carlsbad, CA, USA) according to the manufacturer’s protocol. The extracted RNA was then reverse transcribed by the PrimeScript™ RT reagent Kit with gDNA Eraser (Perfect Real Time., Takara, Japan). The cDNA was used as a template for quantitative polymerase chain reaction (Q-PCR), which was achieved by SYBR Premix Ex Taq™ (Tli RNase H Plus., Takara, Japan) to assay samples with specific primers (Table 2). The results were subsequently analyzed by Bio-Rad CFX Manager.
Statistical analysis
All values presented as mean ± standard deviations (SD). GraphPad Prism 7 software was used for data analysis. One-way ANOVA followed by Tukey’s post hoc test was used to determine the significant difference. p < 0.05 was considered statistically significant (Rattanarojpong et al. 2016).
Results
Generation of RNAi cassette-bearing recombinant baculoviruses
The recombinant BmBacmids containing interference cassettes were successfully constructed and directly transfected into cultured BmN cells by the invasion protein. Without diaminopimelic acid (DAP) in medium, the cell wall of DAP auxotrophic E. coli cannot be synthesized so that the bacteria will disrupt and release recombinant BmBacmid. The released BmBacmids will generate infective recombinant baculovirus particles in insect cells. 4 d.p.i later, the infected BmN cells turned green (attendance at 488 nm) or red (attendance at 512 nm), caused by the fluorescent protein eGFP or mCherry, and the figures indicated that the target gene on the recombinant BmBacmid was expressed and produced recombinant baculovirus as expected (Fig. 2j, k, m–o). The culture supernatant was collected and centrifuged at 80,000g when the fluorescence reached the maximum (5 d.p.i), and the mature baculoviral particles were successfully observed in the pellet. In this study, the purified recombinant BmBacmids were respectively transfected into BmN cells to produce four kinds of recombinant BmNPV named with BmNPV-dsie1-IM (Fig. 2p), BmNPV-dsgp64-IM (Fig. 2q), BmNPV-dsie0-IG (Fig. 2r) and BmNPV-dsie0-dsie1-dsgp64-IG-IM (Fig. 2s).
Inhibition of gene expression by RNAi cassette
To determine the inhibition by different target genes interferences, we used four recombinant baculoviruses, which carrying interfering cassettes on those mentioned above to infect normal BmN cells (MOI = 0.1). The recombinant baculovirus BmNPV-IM carrying mCherry fluorescent protein as control and the same treatment was performed. After 12 h, 24 h, 36 h, 48 h and 60 h post infection (h.p.i), the supernatant and pellet were separated by centrifuging at 4 °C 1500g. The supernatant was collected for subsequent titer determination, and the centrifuge pellets were extracted by Trizol Reagent and reverse transcription by Oligo dT primers (Perfect Real Time., Takara, Japan). The Q-PCR assay results indicated that RNAi cassettes of four different interference modes showed inhibition on the expression levels of the target genes ie0, ie1, gp64 and baculovirus capsid protein vp39 (the gene BmRP49 as the reference. Table 2) (Fig. 3).
Because of the gene interference in the infected BmN cells by BmNPV-dsie1-IM, the expression level of the ie1 was down-regulated by 23.75 fold (12 h.p.i), 30.77 fold (24 h.p.i), 19.03 fold (36 h.p.i), 9.38 fold (48 h.p.i), and 3.58 fold (60 h.p.i), respectively, compared with control (p < 0.05). Furthermore, the expression levels of ie0 and gp64 genes showed a certain decrease in the corresponding period when the maximum reduction rate at 24 h.p.i. Due to its interfering with the dsie0, the expression of ie0 in the infected BmN cells was reduced by 25.71 fold (12 h.p.i), 16.99 fold (24 h.p.i), 7.58 fold (36 h.p.i), 2.25 fold (48 h.p.i), and 1.81 fold (60 h.p.i), respectively, compared with control (p < 0.05). Similarly, the expression level of ie1 and gp64 in the corresponding period also reduced by dsie0 and appeared in the largest reduction at 24 h.p.i. Interference with the envelope protein gp64 gene resulted in down-regulation of gp64 gene expression by 3.22 fold (12 h.p.i), 8.09 fold (24 h.p.i) 3.26 fold (36 h.p.i), 3.05 fold (48 h.p.i), and 2.23 fold (60 h.p.i), respectively, compared with control (p < 0.05). The expression levels of ie1 and gp64 genes also decreased and reached the largest in 24–36 h.p.i (Fig. 3).
When the baculovirus multigene expression system was used to co-interfere with dsie0, dsie1, and dsgp64, the expression of the target genes showed significant down-regulation, compared with the single-interfere. The expression level of ie0 was reduced by 89.21 fold (12 h.p.i), 14.35 fold (24 h.p.i), 4.33 fold (36 h.p.i), 2.44 fold (48 h.p.i), and 2.55 fold (60 h.p.i) (p < 0.05), respectively. The ie1 gene expression level was reduced by 15.72 fold (12 h.p.i), 8.81 fold (24 h.p.i), fourfold (36 h.p.i), 5.47 fold (48 h.p.i), and 3.49 fold (60 h.p.i) (p < 0.05), respectively. The gp64 gene expression level was reduced by 3.88 fold (12 h.p.i), 6.28 fold (24 h.p.i), 1.73 fold (36 h.p.i), 2.18 fold (48 h.p.i), and 2.03 fold (60 h.p.i) (p < 0.05), respectively. In addition, all of the interference cassettes could decrease the expression of vp39, and the effect of multigene co-interference was the most significant within 12–36 h post infected, compared with the control (p < 0.05) (Fig. 3).
To evaluate the interference differences among ie0, ie1, and gp64 genes more clearly and get better interference, we also carried out the study on the expression level of the capsid protein VP39 under different interference strategies. The analysis of Q-PCR results showed that interference strategy of dsie1 could reduce the expression of vp39 by 53.94% and 44.96% (p < 0.05), compared with dsie0 and dsgp64 at 60 h.p.i, respectively. Compared with the dsie0, the dsgp64 decreased the expression of vp39 by 47.68% (p < 0.05) in the early stage of virus infection (12 h.p.i), indicating that the inhibition of early interference on dsgp64 was better. In addition, the vp39 expression of the dsie0 was reduced by 16.03–68.77% in the terminal stage of infection (24–60 h.p.i, compared with dsgp64), which indicates that the inhibition of dsie0 is better than dsgp64 in the middle and later period. It is likely that the gp64 gene is driven by extreme early active promoter p64 (Zheng et al. 2018), which has an early activity about 2–3 times than A3 promoter. Interestingly, the multigene co-interference strategy reduced vp39 gene expression by 29.47–68.40% (p < 0.05), compared with any single-gene interference in the early stage of infection (Fig. 3).
Inhibition of baculovirus proliferation by RNAi cassette
The collected recombinant baculovirus supernatant was directly infected with normal BmN cells referring to the TCID50 method by 96-well plate. The inhibition of virus titers by different interference cassettes at different infecting periods were decreased significantly, compared with the control (Fig. 4). Concretely, when it interfered with the dsie0, the virus titer was decreased by 2.20 fold (12 h.p.i), 4.16 fold (24 h.p.i) 3.48 fold (36 h.p.i), 2.48 fold (48 h.p.i), and 1.99 fold (60 h.p.i), respectively (p < 0.05). The interference results of the dsie1 indicated that the titer of virus was decreased by 3.12 fold (12 h.p.i), 8.05 fold (24 h.p.i), 5.05 fold (36 h.p.i), 3.06 fold (48 h.p.i) and 2.16 fold (60 h.p.i), respectively (p < 0.05). The viral titers that interfere with dsgp64 were decreased by 1.71 fold (12 h.p.i), 2.58 fold (24 h.p.i), 2.96 fold (36 h.p.i), 2.24 fold (48 h.p.i), and 1.77 fold (60 h.p.i), respectively (p < 0.05). When the dsie0 + dsie1 + dsgp64 were carried by the BmNPV-dsie0-dsie1-dsgp64-IG-IM, the reduction of the virus titer was significant, which was reduced by 3.38 fold (12 h.p.i), 6.19 fold (24 h.p.i), 4.66 fold (36 h.p.i), threefold (48 h.p.i), and 2.16 fold (60 h.p.i), respectively (p < 0.05). Moreover, the inhibitory effect of recombinant virus titers reached a maximum at 24 h.p.i in both single- and multi-gene.
Protection of silkworm BmNPV mortality by interference cassette
To evaluate the protective effect of the dsie0, dsie1, dsgp64, and dsie0 + dsie1 + dsie1 on the mortality of BmNPV, a total of 1350 silkworms were injected to challenge with baculoviruses. Five of recombinant baculoviruses containing BmNPV-IM (as control), BmNPV-dsie0-IG, BmNPV-dsie1-IM, BmNPV-dsgp64-IM, and BmNPV-dsie0-dsie1-dsgp64-IG-IM were injected into the silkworm with the concentration of 106/mL, 107/mL and 108/mL, respectively. The silkworm death caused by BmNPV infection was counted daily (Fig. 5a) and dissected (Fig. 5b, c, d).
Anatomy of silkworm and mortality statistics. a Infected larvae showed fluorescence when observed with gel imaging system (Bio-Rad Gel Doc XR+). The hemolymph (b), sericterium (c) and fat body (d) from silkworm larvae infected with recombinant baculoviruses BmNPV-dsie0-dsie1-dsgp64-IG-IM was observed by Eclipse Ti-S, Nikon. The tissue images were taken at the bright, green (488 nm) and red (580 nm). Bar 100 µm. e–g Cumulative mortality of B. mori by different concentrations of BmNPV-infection. e Baculoviral injection of 106/mL, f Baculoviral injection of 107/mL, g Baculoviral injection of 108/mL. (*p < 0.05)
The results indicated that when the injected concentration of baculovirus was 108/mL, any single-gene interference strategy has non-specific protection on the mortality of the silkworm against BmNPV infection at 8 d.p.i, except in the co-interference of multigene (3.33%), compared with the control (Fig. 5e). However, when baculovirus concentration of 107/mL were injected into B. mori, the mortality rate decreased to varying degrees (Fig. 5f). Concrete, the cumulative mortality rates of silkworm were reduced by 5.56% (dsie0), 17.78% (dsie1), and 12.22% (dsgp64), respectively. The results of the cumulative mortality by BmNPV-dsie0-dsie1-dsgp64-IG-IM showed that multigene co-interference could reduce mortality by 20% at day 8 post-infection, compared with control (p < 0.05). The results of 106 / mL BmNPV-injected indicated that the cumulative mortality rate of the silkworm, which interfered with dsie0, dsie1, and dsgp64, could be reduced by 23.46%, 46.91%, and 19.75%, respectively, compared with the control at 8 d.p.i (p < 0.05) (Fig. 5g). Interestingly, the lowest mortality rate for BmNPV infection in silkworm was the multigene co-interference, which reached the protection rate of 55.56% at day 8 post-infection, compared with the control (p < 0.05).
Discussion
With analyzing the results of virus titer by TCID50, we know that the virus titer of the dsie1 model was decreased by 76% and 61% (p < 0.05), respectively, compared with dsie0 and dsgp64 within 12–36 h.p.i. The multigene co-interference strategy constructed by MultiBac inhibited the virus titer with 84%, significantly (p < 0.05). The effect of gene expression level was amplified gradually as the virus increases, resulting in a better inhibition of the same interference strategy on individuals. (Liu et al. 2015). Similarly, the statistical results of silkworm mortality also illustrate this point.
Since the ie1 was an early expression and regulatory gene of BmNPV replication, its transcript product (mRNA) can also encode the protein IE0 by cleavage, indicating that the ie1 has multiple functions of feedback regulation and activation of other promoters (Valdes et al. 2003). The role of the ie1 was interfered and degraded at an early stage, which not only decreased the expression levels of IE1 and IE0, but also regulated the expression. Furthermore, the infection cycle of baculovirus was a cascade process, which showed that the inhibitory on the proliferation of BmNPV was more intense (Sijen et al. 2001). In addition, when the injection concentration is too high, the B. mori had shown the symptoms of BmNPV infected and death before the interference mechanism has taken works, even as the interference cassettes constructed on MultiBac were identical to the baculovirus copies. Consequently, the MultiBac-based co-interference strategy improved by 1.18–2.81 times, compared with the single-gene (p < 0.05), and the inhibitory effect of RNAi on low viral-copy is more pronounced.
Because of the specific expression sequence of RNAi, the RNAi technology constructed by donor vector can generate siRNA in the BmN cells, instantaneously or stably, thus achieving the longtime interference or inhibition (Jiang et al. 2013; Brummelkamp et al. 2002; Chi et al. 2003; Hammond et al. 2001). The study showed that the recombinant baculovirus, which contained the interfering cassettes, would be noticeable interference effect in the early stage of infection and weakened gradually in the later stage of the replication of the virus. Since the impact of poly-gene co-interference is better than that of the single, it indicates that a recombinant baculovirus introduces fewer target interference elements, which may render the deficiency of dsRNA. Additionally, RNAi has stringent sequence specificity, and the viruses were likely to mutate and gain an escape mechanism against RNAi (Elbashir et al. 2001a, b; Leonard and Schaffer 2005; Robalino et al. 2007). According to the previous MultiBac system, the multi-gene co-interference system was successfully constructed and implemented as expected.
BMES system has the advantages of high expression level and large capacity of foreign genes. The multiple interference elements can be directly introduced into MultiBac by combining the polyclonal sites of donor vector (Yao et al. 2012a, b). The strategy can effectively avoid the cumbersome operation of the lipofection technology and give interference genes the ability to work continuously in the process of virus proliferation. The multigene co-interference system can make use of the flexibility of multiple cloning sites and the interfering gene insertions, providing an experimental platform for further RNAi mechanism.
References
Bernstein E, Caudy AA, Hammond SM, Hannon GJ (2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409(6818):363–366
Blissard GW, Wenz JR (1992) Baculovirus gp64 envelope glycoprotein is sufficient to mediate pH-dependent membrane fusion. J Virol 66(11):6829–6835
Brummelkamp TR, Bernards R, Agami R (2002) A system for stable expression of short interfering RNAs in mammalian cells. Science 296(5567):550–553
Carbonell LF, Klowden MJ, Miller LK (1985) Baculovirus-mediated expression of bacterial genes in dipteran and mammalian cells. J Virol 56(1):153–160
Chen S, Hou C, Bi H, Wang Y, Xu J, Li M, James AA, Huang Y, Tan A (2017) Transgenic clustered regularly interspaced short palindromic repeat/Cas9-mediated viral gene targeting for antiviral therapy of Bombyx mori nucleopolyhedrovirus. J Virol 91(8):e02465–e02416. (https://doi.org/10.1128/JVI.02465-16)
Cheng D, Xia Q, Duan J, Wei L, Huang C, Li Z, Wang G, Xiang Z (2008) Nuclear receptors in Bombyx mori: insights into genomic structure and developmental expression. Insect Biochem Mol Biol 38(12):1130–1137
Chi JT, Chang HY, Wang NN, Chang DS, Dunphy N, Brown PO (2003) Genomewide view of gene silencing by small interfering RNAs. Proc Natl Acad Sci USA 100(11):6343–6346
Choi J, Guarino LA (1995) The baculovirus transactivator IE1 binds to viral enhancer elements in the absence of insect cell factors. J Virol 69(7):4548–4551
Dai X, Willis LG, Huijskens I, Palli SR, Theilmann DA (2004) The acidic activation domains of the baculovirus transactivators IE1 and IE0 are functional for transcriptional activation in both insect and mammalian cells. J Gen Virol 85(Pt 3):573–582
de Jong J, Theilmann DA, Arif BM, Krell PJ (2011) Immediate-early protein ME53 forms foci and colocalizes with GP64 and the major capsid protein VP39 at the cell membranes of Autographa californica multiple nucleopolyhedrovirus-infected cells. J Virol 85(19):9696–9707
Dougherty WG, Parks TD (1995) Transgenes and gene suppression: telling us something new? Curr Opin Cell Biol 7(3):399–405
Elbashir SM, Harborth J, Lendeckel W, Yalcin A, Weber K, Tuschl T (2001a) Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411(6836):494–498
Elbashir SM, Martinez J, Patkaniowska A, Lendeckel W, Tuschl T (2001b) Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. Embo J 20(23):6877–6888
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391(6669):806–811
Gomi S, Zhou CE, Yih W, Majima K, Maeda S (1997) Deletion analysis of four of eighteen late gene expression factor gene homologues of the baculovirus, BmNPV. Virology 230(1):35–47
Gomi S, Majima K, Maeda S (1999) Sequence analysis of the genome of Bombyx mori nucleopolyhedrovirus. J Gen Virol 80(Pt 5):1323–1337
Guarino LA, Summers MD (1986) Functional mapping of a trans-activating gene required for expression of a baculovirus delayed-early gene. J Virol 57(2):563–571
Hammond SM, Caudy AA, Hannon GJ (2001) Post-transcriptional gene silencing by double-stranded RNA. Nat Rev Genet 2(2):110–119
Hannon GJ (2002) RNA interference. Nature 418(6894):244–251
Isobe R, Kojima K, Matsuyama T, Quan GX, Kanda T, Tamura T, Sahara K, Asano SI, Bando H (2004) Use of RNAi technology to confer enhanced resistance to BmNPV on transgenic silkworms. Arch Virol 149(10):1931–1940
Jiang L, Xia Q (2014) The progress and future of enhancing antiviral capacity by transgenic technology in the silkworm Bombyx mori. Insect Biochem Mol Biol 48:1–7
Jiang L, Wang G, Cheng T, Yang Q, Jin S, Lu G, Wu F, Xiao Y, Xu H, Xia Q (2012) Resistance to Bombyx mori nucleopolyhedrovirus via overexpression of an endogenous antiviral gene in transgenic silkworms. Arch Virol 157(7):1323–1328
Jiang L, Zhao P, Wang G, Cheng T, Yang Q, Jin S, Lin P, Xiao Y, Sun Q, Xia Q (2013) Comparison of factors that may affect the inhibitory efficacy of transgenic RNAi targeting of baculoviral genes in silkworm, Bombyx mori. Antiviral Res 97(3):255–263
Kadlec J, Loureiro S, Abrescia NG, Stuart DI, Jones IM (2008) The postfusion structure of baculovirus gp64 supports a unified view of viral fusion machines. Nat Struct Mol Biol 15(10):1024–1030
Kovacs GR, Choi J, Guarino LA, Summers MD (1992) Functional dissection of the Autographa californica nuclear polyhedrosis virus immediate-early 1 transcriptional regulatory protein. J Virol 66(12):7429–7437
Kuzio J, Rohel DZ, Curry CJ, Krebs A, Carstens EB, Faulkner P (1984) Nucleotide sequence of the p10 polypeptide gene of Autographa californica nuclear polyhedrosis virus. Virology 139(2):414–418
Leonard JN, Schaffer DV (2005) Computational design of antiviral RNA interference strategies that resist human immunodeficiency virus escape. J Virol 79(3):1645–1654
Li G, Tang Q, Chen H, Yao Q, Ning D, Chen K (2011) Display of Bombyx mori nucleopolyhedrovirus GP64 on the Bacillus subtilis spore coat. Curr Microbiol 62(5):1368–1373
Liu Y, Zhang L, Zhang Y, Liu D, Du E, Yang Z (2015) Functional analysis of RNAi suppressor P19 on improving baculovirus yield and transgene expression in Sf9 cells. Biotechnol Lett 37(11):2159–2166
Lu M, Iatrou K (1997) Characterization of a domain of the genome of BmNPV containing a functional gene for a small capsid protein and harboring deletions eliminating three open reading frames that are present in AcNPV. Gene 185(1):69–75
McIntyre GJ, Fanning GC (2006) Design and cloning strategies for constructing shRNA expression vectors. Bmc Biotechnol 6(1):1–8
Mena JA, Kamen AA (2011) Insect cell technology is a versatile and robust vaccine manufacturing platform. Expert Rev Vaccines 10(7):1063–1081
Nagai S, Alves CA, Kobayashi M, Ikeda M (2011) Comparative transient expression assay analysis of hycu-hr6- and IE1-dependent regulation of baculovirus gp64 early promoters in three insect cell lines. Virus Res 155(1):83–90
Nagamine T, Abe A, Suzuki T, Dohmae N, Matsumoto S (2011) Co-expression of four baculovirus proteins, IE1, LEF3, P143, and PP31, elicits a cellular chromatin-containing reticulate structure in the nuclei of uninfected cells. Virology 417(1):188–195
O’Reilly DR (1997) Use of baculovirus expression vectors. Methods Mol Biol 62:235–246
Oomens AG, Monsma SA, Blissard GW (1995) The baculovirus GP64 envelope fusion protein: synthesis, oligomerization, and processing. Virology 209(2):592–603
Qi Q, Yao L, Liang Z, Yan D, Li Z, Huang Y, Sun J (2016) Production of human type II collagen using an efficient baculovirus-silkworm multigene expression system. Mol Genet Genomics 291(6):2189–2198
Rahman MM, Gopinathan KP (2003) Characterization of the gene encoding the envelope fusion glycoprotein GP64 from Bombyx mori nucleopolyhedrovirus. Virus Res 94(1):45–57
Rankin C, Ladin BF, Weaver RF (1986) Physical mapping of temporally regulated, overlapping transcripts in the region of the 10K protein gene in Autographa californica nuclear polyhedrosis virus. J Virol 57(1):18–27
Rattanarojpong T, Khankaew S, Khunrae P, Vanichviriyakit R, Poomputsa K (2016) Recombinant baculovirus mediates dsRNA specific to rr2 delivery and its protective efficacy against WSSV infection. J Biotechnol 229:44–52
Robalino J, Bartlett TC, Chapman RW, Gross PS, Browdy CL, Warr GW (2007) Double-stranded RNA and antiviral immunity in marine shrimp: inducible host mechanisms and evidence for the evolution of viral counter-responses. Dev Comp Immunol 31(6):539–547
Saleh MC, Tassetto M, van Rij RP, Goic B, Gausson V, Berry B, Jacquier C, Antoniewski C, Andino R (2009) Antiviral immunity in Drosophila requires systemic RNA interference spread. Nature 458(7236):346–350
Senkevich TG, Koonin EV, Buller RM (1994) A poxvirus protein with a RING zinc finger motif is of crucial importance for virulence. Virology 198(1):118–128
Sijen T, Fleenor J, Simmer F, Thijssen KL, Parrish S, Timmons L, Plasterk RH, Fire A (2001) On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 107(4):465–476
Subbaiah EV, Royer C, Kanginakudru S, Satyavathi VV, Babu AS, Sivaprasad V, Chavancy G, Darocha M, Jalabert A, Mauchamp B, Basha I, Couble P, Nagaraju J (2013) Engineering silkworms for resistance to baculovirus through multigene RNA interference. Genetics 193(1):63–75
Sun JC, Zhang EH, Yao LG, Zhang HL, Jin PF (2009) A high efficient method of constructing recombinant Bombyx mori (silkworm) multiple nucleopolyhedrovirus based on zero-background Tn7-mediated transposition in Escherichia coli. Biotechnol Prog 25(2):524–529
Sun JC, Yao LG, Yao N, Xu H, Jin PF, Kan YC (2010) Production of recombinant Bombyx mori nucleopolyhedrovirus in silkworm by intrahaemocoelic injection with invasive diaminopimelate auxotrophic Escherichia coli containing BmNPV-Bacmid. Biotechnol Appl Biochem 57(3):117–125
Valdes VJ, Sampieri A, Sepulveda J, Vaca L (2003) Using double-stranded RNA to prevent in vitro and in vivo viral infections by recombinant baculovirus. J Biol Chem 278(21):19317–19324
Wang Q, Xie H, Zeng W, Wang L, Liu C, Wu J, Wang Y, Li Y, Bergmann SM (2018) Development of indirect immunofluorescence assay for TCID50 measurement of grass carp reovirus genotype II without cytopathic effect onto cells. Microb Pathog 114:68–74
Xia Q, Zhou Z, Lu C, Cheng D, Dai F, Li B, Zhao P, Zha X, Cheng T, Chai C, Pan G, Xu J, Liu C, Lin Y, Qian J, Hou Y, Wu Z, Li G, Pan M, Li C, Shen Y, Lan X, Yuan L, Li T, Xu H, Yang G, Wan Y, Zhu Y, Yu M, Shen W, Wu D, Xiang Z, Yu J, Wang J, Li R, Shi J, Li H, Li G, Su J, Wang X, Li G, Zhang Z, Wu Q, Li J, Zhang Q, Wei N, Xu J, Sun H, Dong L, Liu D, Zhao S, Zhao X, Meng Q, Lan F, Huang X, Li Y, Fang L, Li C, Li D, Sun Y, Zhang Z, Yang Z, Huang Y, Xi Y, Qi Q, He D, Huang H, Zhang X, Wang Z, Li W, Cao Y, Yu Y, Yu H, Li J, Ye J, Chen H, Zhou Y, Liu B, Wang J, Ye J, Ji H, Li S, Ni P, Zhang J, Zhang Y, Zheng H, Mao B, Wang W, Ye C, Li S, Wang J, Wong GK, Yang H (2004) A draft sequence for the genome of the domesticated silkworm (Bombyx mori). Science 306(5703):1937–1940
Yao LG, Liu ZC, Zhang XM, Kan YC, Zhou JJ (2007) A highly efficient method for the generation of a recombinant Bombyx mori nuclear-polyhedrosis-virus Bacmid and large-scale expression of foreign proteins in silkworm (B. mori) larvae. Biotechnol Appl Biochem 48(Pt 1):45–53
Yao LG, Sun JC, Xu H, Kan YC, Zhang X, Yan HC (2010) A novel economic method for high throughput production of recombinant baculovirus by infecting insect cells with Bacmid-containing diminopimelate-auxotrophic Escherichia coli. J Biotechnol 145(1):23–29
Yao LG, Jin PF, Su S, Xu H, He J, Peng L, Sun JC (2012a) Quantitative analysis of three commonly-used insect cell-specific promoters’ activities by transient and baculovirus-mediated expression. Afr J Biotechnol 11(5):1037–1045
Yao LG, Wang S, Su S, Yao N, He J, Peng L, Sun JC (2012b) Construction of a baculovirus-silkworm multigene expression system and its application on producing virus-like particles. Plos One 7(3):e32510. https://doi.org/10.1371/journal.pone.0032510
Zamore PD, Tuschl T, Sharp PA, Bartel DP (2000) RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 101(1):25–33
Zheng H, Wang X, Ren FF, Zou SL, Feng M, Xu LL, Yao LG, Sun JC (2018) Construction of a highly efficient display system for baculovirus and its application on multigene co-display. Mol Genet Genomics 293(5):1265–1277
Acknowledgements
This study was funded by the National Natural Science Foundation of China (Grant Nos. 31872426, 31372373), the Natural Science Foundation of Guangdong Province, China (Grant No. 2016A030311018).
Author information
Authors and Affiliations
Contributions
HZ and JS (J Sun) coordinated the project. HZ and FR performed the research. HZ JS (J Song) and JS (J Sun) wrote the manuscript. QL and JS (J Sun) contributed new methods and improved the manuscript. QL, ZC, and MF performed the data analysis. HZ, JL, and JS (J Sun) interpreted the context of results. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by S. Hohmann.
Rights and permissions
About this article
Cite this article
Zheng, H., Ren, F., Lu, Q. et al. An efficient method for multigene co-interference by recombinant Bombyx mori nucleopolyhedrovirus. Mol Genet Genomics 294, 111–120 (2019). https://doi.org/10.1007/s00438-018-1491-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-018-1491-9