Abstract
The quality of fiber is significant in the upland cotton industry. As complex quantitative traits, fiber quality traits are worth studying at a genetic level. To investigate the genetic architecture of fiber quality traits, we conducted an association analysis using a multi-parent advanced generation inter-cross (MAGIC) population developed from eight parents and comprised of 960 lines. The reliable phenotypic data for six major fiber traits of the MAGIC population were collected from five environments in three locations. Phenotypic analysis showed that the MAGIC lines have a wider variation amplitude and coefficient than the founders. A total of 284 polymorphic SSR markers among eight parents screened from a high-density genetic map were used to genotype the MAGIC population. The MAGIC population showed abundant genetic variation and fast linkage disequilibrium (LD) decay (0.76 cM, r2 > 0.1), which revealed the advantages of high efficiency and power in QTL exploration. Association mapping via a mixed linear model identified 52 significant loci associated with six fiber quality traits; 14 of them were mapped in reported QTL regions with fiber-related or other agronomic traits. Nine markers demonstrated the pleiotropism that controls more than two fiber traits. Furthermore, two SSR markers, BNL1231 and BNL3452, were authenticated as hotspots that were mapped with multi-traits. In addition, we provided candidate regions and screened six candidate genes for identified loci according to the LD decay distance. Our results provide valuable QTL for further genetic mapping and will facilitate marker-based breeding for fiber quality in cotton.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cotton (Gossypium spp.) is an important crop that provides most natural fiber for industrial textiles. Gossypium hirsutum L., also called upland cotton, is the most widely cultivated cotton species and accounts for more than 90% of worldwide production because of its high yield, wide adaptability, and acceptable fiber quality (Zhang et al. 2008; Page et al. 2013). Cotton fibers are single-celled trichomes that provide a unique experimental system to study cell development processes, such as cell growth, biosynthesis, and response mechanisms for outside signals (Qin and Zhu 2011; Haigler et al. 2012; Walford et al. 2012; Guo et al. 2016). Cotton fiber has been involved in pioneering research concerning the genetic basis of its yield and quality through forward or reverse genetics (Haigler et al. 2012; Said et al. 2015).
Recently, molecular selection strategies based on quantitative trait loci (QTL) mapping are efficient tools for breeders to combine economically important traits to create a superior cultivar (Xu et al. 2012; Smykal et al. 2016). Identifying the QTL and dissecting the genetic basis of agronomy traits are critical steps and prerequisites for marker-assisted breeding in crops. Linkage mapping and linkage disequilibrium (LD) mapping are the mainstream methods for dissecting the genetic architecture of complex traits in crops (Mackay and Powell 2007). Traditionally, bi-parental populations, such as F2, recombinant inbred lines (RILs) and chromosome segment substitution lines (CSSLs), are utilized to excavate the QTLs or genes and to evaluate the biological effects for plant phenotypes (Xu et al. 2012; Said et al. 2015). For qualitative traits or main effect loci, linkage mapping is an efficient way to determine the target genes (Han et al. 2015; Ma et al. 2016; Liu et al. 2016a). However, the resolution of quantitative trait mapping in bi-parental populations is usually low (Mackay and Powell 2007). Association mapping offers a high resolution with either prior information on candidate genes or a genome scan with very high marker coverage (Zhao et al. 2011; Li et al. 2013b). Compared to linkage mapping, association analysis is more efficient and powerful for the dissection of complex traits (Cavanagh et al. 2008; Atwell et al. 2010). In addition, association analyses have been successfully employed in many crops (Zhao et al. 2011; Li et al. 2013b; Xu et al. 2016; Sun et al. 2017).
In most cases, researchers conduct an association analysis based on a natural population from a diverse collection of inbred lines or germplasms that may constitute a group with population structure related to geographical origin or reproductive isolation. The population structure will disrupt the association mapping and give false-positive results. The multi-parent advanced generation inter-cross (MAGIC) population has richer genetic diversity and recombination rate without population structure, which provides power and resolution for gene targeting (Cavanagh et al. 2008; Huang et al. 2015; Pascual et al. 2016). The MAGIC population is suitable for genetic map developing and QTL mapping (Huang et al. 2011, 2015; Bandillo et al. 2013). Recently, MAGIC populations have been created for genetic analysis in crops, such as rice (Bandillo et al. 2013), wheat (Huang et al. 2012; Delhaize et al. 2015), maize (Dell’Acqua et al. 2015), chickpea (Agarwal et al. 2015), and cotton (Islam et al. 2016).
Fiber quality traits are quantitative traits with a complex genetic basis (Fang et al. 2017). Several fiber characteristics, such as fiber length (FL), fiber strength (FS), micronaire value (MV), fiber elongation (FE), fiber uniformity (FU), and short fiber content (SFC) are the main determining factors for fiber quality. Dissection of QTLs for these fiber traits will help to enhance the genetic architecture of fiber quality. Recently, several studies were focused on the genetic bases of fiber quality traits using simple sequence repeat (SSR) or single-nucleotide polymorphism (SNP) markers (Said et al. 2015; Nie et al. 2016; Sun et al. 2017; Huang et al. 2017). In this study, an upland cotton MAGIC population containing 960 lines was developed by a convergent cross based on 8 parents. An association analysis was performed based on phenotypic data from five environments, and genotypic data were generated from 284 polymorphic SSRs. This study aimed to take advantage of the MAGIC population to explore the genetic architecture of 6 fiber traits and to provide molecular genetic basis for fiber improvement in breeding.
Materials and methods
Development of the upland cotton MAGIC population
The upland cotton MAGIC population was developed with eight parents. The parents of MAGIC (PMs) were diverse accessions with abundant phenotypic characteristics for high yield and fiber quality (GY2, GY4, GY5, and GY6), pest resistance (KC9, CQ2, and CQ13) and Fusarium wilt and Verticillium wilt resistance (KB10) (Table S1). They were widely distributed in the Yangtze River valley and Yellow River valley and were provided by CCRI (Cotton Research Institute of Chinese Academy of Agricultural Sciences), HAAFS (Hebei Academy of Agriculture and Forestry Sciences), ACYU (Agricultural College of Yangtze University), HSMS (Hubei Province Seed Management Station), and TCCSF (Taicang City Cotton Seed Farm in Jiangsu province) (Table S1).
The convergent crossing followed the development of an eight-way cross or funnel crossing (Cavanagh et al. 2008; Huang et al. 2015), and the MAGIC population was obtained by inter-crossing eight parents with three generations and successive selfing (Fig. 1). In the first stage, multiple parents (G0) were inter-mated to generate four bi-parental crosses (two-way): GY5 × KB10 (AB), GY2 × KC9 (CD), CQ13 × GY6 (EF), and CQ2 × GY4 (GH) at the Agricultural College of Yangtze University, Jinzhou, Hubei province in the summer of 2004. Then, four F1 plants (G1) were inter-crossed to derive four-way crosses (G2), including AB × CD (ABCD), and EF × GH (EFGH) at Sanya in Hainan province in the winter of 2004. The four-way crosses formed a female parent group (ABCD) containing 200 individuals and a male parent group (EFGH) containing 200 individuals for subsequent crossing. The last stage involved individual inter-crossing between female and male parent groups that formed 200 inter-mated hybridizations to derive the eight-way crosses (G3) EFGH × ABCD (ABCDEFGH) in 2005. Only 168 of 200 mated generations were successfully multiplied.
To create homozygous individuals, the eight-way cross generations (G3) were self-pollinated with multiple generations. In the first self-crossing, the 168 G3 heterozygotes were self-crossed and generated ten plants from each line in 2006. Approximately 1680 filial lines were obtained after expanding propagation. However, some of the lines were lost in the process of selfing through single seed descent. Finally, a total of 960 lines were preserved after selfing for more than 5 generations from 2007 to 2011. The 8 parents (PMs) and 960 MAGIC lines (MLs) formed the population panel for this study.
Fiber quality phenotyping
The fiber quality traits were measured under multiple environments. The phenotypic data of eight PMs and 960 MLs were collected from five natural environments at Huanggang, Hubei province (HG), Zhoukou, Henan province (ZK), and Jingzhou, Hubei province (JZ). The five natural environments were defined as E1 (HG, 2013), E2 (HG, 2014), E3 (ZK, 2014), E4 (ZK, 2015), and E5 (JZ, 2015). The 968 lines were grown during the planting season at each location. The field plots followed a completely randomized block design. Each plot contained one row of MLs and multi-rows of PMs. The row length and spacing were set as 5 and 1 m, respectively. Each row included 10 plants with 40 cm between individuals. Fiber samples were collected from 20 normally opened bolls from the middle fruit branches of plants in each row. The fiber quality traits, including the fiber upper half mean length (FUHML, mm), fiber strength (FS, cN/tex), micronaire value (MV), fiber elongation (FE, %), fiber uniformity (FU, %), and short fiber content (SFC, %), were measured by HVI1000 Automatic Fiber Determination System in Institute of Cotton Research, Shihezi Academy of Agricultural Sciences. FE data in 2014 Huanggang (E2) and Zhoukou (E3) were missing and only obtained from three environments. The remaining five traits were obtained from all five environments.
Phenotypic data analysis
The analysis of variance (ANOVA), correlation, and repeatability analysis for fiber quality traits were conducted using IBM SPSS Statistics 21 (SPSS, Chicago, IL, USA). Best linear unbiased predictions (BLUP) were used to estimate phenotypic traits across multiple environments based on a linear model. The phenotypic data from BLUP and the broad-sense heritability (h2) were calculated with the R software (http://www.r-project.org). The statistical analysis of the mean, standard error (SE), and Kendall’s tau-b correlation coefficient was calculated using IBM SPSS Statistics 21 (SPSS, Chicago, IL, USA). The ANOVA was carried out to evaluate the effects of genotype (G), environment (E), and the interactions between genotype and environment (G × E).
SSR genotyping
Five young fresh leaves were picked from five plants of each line. The genomic DNA of 8 PMs and 960 MLs was extracted using the modified CTAB method (Paterson et al. 1993). A 10-µL reaction volume for the SSR reaction system was performed for PCR amplification (Lin et al. 2005). The Fragment Analyzer™ 96-Capillary Automated CE System and gel electrophoresis (6% denaturing polyacrylamide gel for SSR and 8% non-denatured polyacrylamide gel electrophoresis for SSCP-SSR, silver staining developing after electrophoresis) were used to visualize the PCR products. A total of 3871 SSR primers were selected for detecting polymorphisms from the interspecific (Gossypium hirsutum × Gossypium barbadense) genetic map, which contained 5152 loci and spanned 4696.03 cM (Li et al. 2016b). After screening the SSR primers by eight PMs, the polymorphic markers were used to genotype the 960 lines (Table 2). The 5 cM range nearly covered by SSR markers were set as effectively covered fragments. The adjacent markers with gaps ≤ 10 cM were treated as the same covered fragments.
Population structure, kinship and LD decay analysis
The genotypic data of polymorphic markers were used to calculate the genetic eigen value of the MAGIC population. Population structure and kinship could cause false positives or deviations in association mapping. Principal component analysis (PCA) and cluster analysis were used to confirm a reasonable population structure. PCA was conducted by the Powermarker software 3.25 (Liu and Muse 2005) and Ntsys 2.1 (Adams and Rohlf 2000). The relative kinship coefficients (K) were estimated by SPAGeDi version 1.4b (Hardy and Vekemans 2002). The PCA matrix for phenotypic variations was calculated by IBM SPSS Statistics 21, whereas the contribution rate of the K matrix was calculated with TASSEL 3.0 (Bradbury et al. 2007). The linkage disequilibrium (D′, r2 and p value) was calculated with TASSEL 3.0. The parameter r2 was used to graphically represent the LD curves with the R software.
Marker–trait association
Marker–trait association analyses for the six fiber quality traits based on the interspecific genetic map (Li et al. 2016b) were conducted using a general linear model (GLM) and mixed linear model (MLM) with the TASSEL 3.0 software package. The naive GLM model ignores the characteristic vector from the population structure. The MLM-incorporated kinship (K matrix) was used as the concomitant variant to eliminate random effects. The threshold for the significance of associations between SSR markers and traits was set as p < 0.01 (− log10(p) > 2), and a Bonferroni threshold was set, so that p < 0.00352 (p = 1/n, − log10(1/284) = 2.45), which has been widely adopted in the previous studies (Wang et al. 2012; Liu et al. 2015b, 2016b). The sequences of significant associated markers were searched from CottonGen Database (http://www.cottongen.org) and assigned a genome location (NAU-genome database of TM-1, Zhang et al. 2015) using a BLAST (Altschul et al. 1994) search with E ≤ 1e−10.
Results
Upland cotton MAGIC population development and phenotypic characteristics of fiber quality
Eight upland cotton accessions with good characteristics in different traits were selected as the founders (PMs) to develop the MAGIC population (Table S1). An eight-way inter-cross was designed to aggregate the alleles from eight PMs. Overall, 960 stable inbred MAGIC lines (MLs) were obtained after selfing for more than six generations from eight-way lines (Fig. 1). Eight PMs and 960 MLs were used for phenotyping, genotyping, and the association analysis.
The data of five fiber quality traits, including FUHML, FS, MV, FU, and SFC, were collected from E1 (HG, 2013), E2 (HG, 2014), E3 (ZK, 2014), E4 (ZK, 2015), and E5 (JZ, 2015), while the FE data were only collected from E1, E4, and E5. These traits showed rich variations in different environments (Table S2). The coefficients of variation (CV) for six traits ranged from 1.17% (FU-E2) to 20.13% (SFC-E3) in MLs and ranged from 0.72% (FU-E3) to 16.58% (FE-E4) in PMs (Table S2). The six traits, including FUHML, FS, FE, FU, MV, and SFC, varied from 25.11 to 31.13 mm, 25.85 to 39.18 cN/tex, 2.86 to 9.66%, 82.46 to 87.27%, 4.36 to 5.67, and 7.01 to 13.16% in PMs, respectively. The variations of the six traits were wider in MLs than in PMs and varied from 22.34 to 33.36 mm, 12.04 to 43.72 cN/tex, 1.80 to 10.62%, 78.44 to 88.29%, 2.77 to 6.74, and 4.20 to 23.80%, respectively (Table S2).
The six traits were also merged with BLUP to eliminate the environmental impact. The results also showed that phenotypic variations in MLs were more abundant than variations in PMs. The CVs of six traits in BLUP were 2.12, 3.87, 0.65, 0.10, 3.48, and 1.93% in PMs, whereas they were 2.56, 4.03, 10.60, 0.26, 3.87, and 3.26 in MLs. The broad-sense heritability (h2) of six traits ranged from 40 to 81% in MLs. Only the FU and SFC had relatively low h2, which were 42 and 40%, respectively (Table 1). The ANOVA showed that the genotype (G) and environment (E) had significant effects (p < 0.001) on fiber quality traits (Table S3). The results suggested that both genotype and environment affected phenotypic variation, and genotype played a stable role in most traits. Correlation analysis showed that FUHML and FS were significantly correlated with other traits (p < 0.01) (Table S4). Significant positive correlations were found among six paired traits, FUHML-FS, FUHML-FU, FS-FU, FS-MV, FE-MV, and FU-MV, and there were significant negative correlations in the remaining pairs, except for FE-FU and FE-SFC.
Genotyping and genetic diversity
A total of 644 (17.15%) polymorphic markers screened from 3871 SSR markers by 8 PMs were used to genotype the 960 MLs. Among them, only 284 markers showed clear electrophoresis bands and considerable polymorphism, and the minor allele frequency (MAF) was > 0.05. The 284 polymorphic SSR markers were distributed across 26 chromosomes from 3 (Chr04) to 16 (Chr05 and Chr13) (Table 2). The effective fragment covered by SSR markers accounted for approximately 42.98% of the whole genome and ranged from 19.67% (Chr04) to 65.03% (Chr16). The mean values of genetic diversity for the PMs and MLs were 0.415 and 0.465, respectively (Table 2). In addition, the average polymorphism information content (PIC) values of PMs and MLs were 0.346 and 0.390, respectively. These results suggested that the genetic diversity of MLs based on SSR markers was higher than the diversity of PMs.
Population kinship, genetic structure and linkage disequilibrium
The relative kinship coefficients were calculated as the pairwise relatedness between MLs. The results showed that the kinship values after standardized treatment were low with an average value of 0.057 in MLs, but the values were higher with an average value of 0.490 in PMs. In addition, 52.3% of pairwise kinship coefficients among MLs were 0, and approximately 97.0% of pairwise coefficients were less than 0.3 (Fig. S1). The kinship matrix (K matrix) was used to correct random error in the marker–trait association. The K-matrix could explain 24.81, 20.60, 9.21, 11.41 12.73, and 15.28% of the phenotypic variation for FUHML, FS, FE, FU, MV, and SFC, respectively (Table S5).
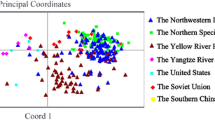
PCA was used to reflect the genetic divergence for MLs and PMs. The top three principal components (PCs), PC1, PC2, and PC3, accounted for 28.76, 13.06, and 7.99% of the genetic variations, respectively. The PC1–PC2 plots showed a dispersive distribution without obvious clusters (Fig. S2). In addition, the PC1–PC2 plots of MLs were scattered more widely than the PMs. It was suggested that the MLs obtained more diversity without population structure. In addition, for the PCA matrix, the top three PCs explained 14.2, 16.8, 0.2, 9.9, 7.7, and 9.8% of the phenotypic variations for FUHML, FS, FE, FU, MV, and SFC, respectively. It was obviously smaller than the K matrix.
The squared Pearson correlation coefficients (r2) were measured by 284 SSR markers to describe the LD relationship between markers. The LD showed a very low level in MLs (Fig. 2). A total of 40,186 possible pair combinations were obtained with an average of r2 = 0.029. The distribution of r2 values was concentrated in the < 0.1 interval that accounted for 93.6% of all pairwise coefficients and 49.1% of the coefficients were < 0.01. In addition, 27.02, 33.70, and 43.49% of the coefficients (r2) were revealed to be significant at p < 0.0001, p < 0.001 and p < 0.01, respectively. The LD decay rate was measured by an r2 plot based on the corresponding genetic distance across the entire genome. The average distance of pair combinations was ~ 0.60 cM with the r2 plot dropping to 0.2, and it was ~ 0.76 cM with the r2 plot dropping to 0.1 (Fig. 2c). The 0.76 cM approximately equaled 400 kb in physical genome map.
Association mapping QTLs for fiber quality traits
The MAGIC population was calculated with a low population structure. The naive general linear model (GLM) and mixed linear model with K matrix (MLM (K)) were used to execute the marker–trait associations based on the phenotypic data of six fiber traits. The Q–Q plot of naive GLM deviated materially from the predicted line, which suggested that there was a higher rate of false positives in GLM. Compared to the GLM, the MLM model could correct the error from the random effects (Fig. S3). For this study, the MLM (K) was chosen as the optimal model.
Two thresholds, p < 0.01 (− log10(p) = 2) and a Bonferroni threshold of p < 0.00352 (1/284, − log10(p) = 2.45) were set as the significance levels. When using the traits data from the BLUP results and MLM (K) model, a total of 52 marker–trait associated loci were significantly associated with 6 traits at p < 0.01 (Table 3), while 31 of the 52 loci were identified with p < 0.00352 (Table S6). The FUHML was associated with the maximum loci of 18, while the other 5 traits, including FS, FE, FU, MV, and SFC, identified 12, 2, 4, 7, and 9 loci, respectively (p < 0.01, Table S6). The allele frequency of the significantly associated markers was dependable as the MAF > 7.15% (Table 3). The percentage of phenotypic variation explained by SSR markers (R2) ranged from 0.71 to 5.47% (Table S6). Furthermore, among the 52 loci, 47 loci shared a significant association with the results using a naive GLM model (Table S7). In addition, 134 significant (p < 0.01) marker–trait association loci were found using trait data from individual environments (E1–E5) (Table S8). A total of 35 significant loci associated with the BLUP data were identified in individual environments. For example, MON_SHIN0376b, which was associated with MV based on BLUP data, were detected in E3, E4, and E5.
The 52 marker–trait associated loci were from 40 SSR markers and distributed on 22 chromosomes, except for Chr03, Chr10, Chr17, and Chr22 (Fig. 3). Based on the marker sequences and the NAU-genome database for TM-1, 37 of the 40 identified SSR markers were located on a physical map using BLAST methods (Table S9). The other three markers were below the threshold or located in uncertain positions. There were nine markers associated with multiple traits (Table S10). For example, MON_CGR5525 was associated with three traits, including FUHML, FU, and SFC. Based on studies in recent years, 14 markers were found to be mapped with the same traits or related traits (Fig. 3, Table S11). For example, BNL1231, which was associated with FS, FE, and SFC, was also located in the loci controlling FUHML, FS, MV, and some yield or plant type traits in other studies.
Distribution of the marker–trait loci identified by association analysis on the published genetic map. Genetic distances in centimorgans (cM) are indicated on the left side of the linkage map. The marker name and associated fiber quality traits are indicated on the right side. Asterisk indicates that the markers were the overlapped marker in this study with other studies, as listed in Table S11
The candidate regions for marker–trait loci were set around the LD decay distance as 400 kb (r2 < 0.1). The numbers of genes ranged from 4 to 107 with an average of 53 in the candidate regions (Table S9). There were three regions covering only few genes that were less than ten. They were TMB0029b, MON_DPL0906, and MON_DPL0375c, which were significantly associated with the same trait FUHML, linked 2, 3, and 4 genes in the confidence genome intervals (Tables S9 and S12). In addition, there were 20 genes with the functions mainly affected fiber development via resource references, located in neighboring site of the significant SSR loci by BLAST (Table S13). Six genes, Gh_A01G1843 (GhPIP1-2), Gh_A01G1915 (GhACT1), Gh_A09G1461 (GhACT1), Gh_A13G0162 (GhLIM1), Gh_D01G0018 (GhPIP2-4), and Gh_D01G1810 (GhCER6) were located in the candidate regions with a short distance < 400 kb. In addition, the other 14 genes were near the marker–trait loci within < 3 Mb. Several reported genes were found multi-copy loci to be related to different loci; for example, GhACT1 was linked with four loci, HAU4552b, MON_CGR5707, MON_DPL0042, and NAU7153.
Discussion
In this study, an association analysis was performed to explore the genetic basis of fiber quality traits in an upland cotton MAGIC population. Association mapping is a powerful and popular tool for studying the genetic basis of complex traits in plants (Yu et al. 2008; Ingvarsson and Street 2011; Yang et al. 2012). The efficiency of association analysis is determined by the diversity of a population, the experimental population size, and reasonable statistical methods (Yan et al. 2009; Bush and Moore 2012). We developed a MAGIC population with sufficient diversity and individuals. The MAGIC population takes advantage of populations that are known to have high efficiency, high precision, and low false-positive rate (Mackay and Powell 2007; Cavanagh et al. 2008; Huang et al. 2015).
The wider range of phenotypic and genotypic variations generated in MAGIC recombination lines is beneficial for carrying out genetic research of complex traits (Cavanagh et al. 2008; Huang et al. 2015). To maintain relatively high levels of diversity in the MAGIC population, eight parents were selected from different areas with various characteristics and convergent crosses were performed to obtain sufficient recombination (Fig. 1, Table S1). Multiple parents could carry more alleles than a bi-parental program. Although the eight parents had relatively low polymorphism at the genetic level, they created more abundant phenotypic and genotypic variations in the MAGIC lines. The screened SSR markers used for genotyping demonstrated that the MAGIC population had higher genetic diversity compared to the PMs (Table 2) as well as higher diversity than the population used in the association analysis for cotton (Zhao et al. 2014; Liu et al. 2015a). In addition, the MAGIC population had relatively abundant phenotypic variations compared to the PMs. The multiple parents inter-cross is a breeding method that could gather favorable alleles and create excellent new varieties through recombination (Cavanagh et al. 2008; Huang et al. 2015). In this study, more MLs had better properties in terms of fiber quality with fiber lengths > 30 mm and fiber strengths > 30 cN/tex. The selection of germplasm from the cotton MAGIC population could potentially improve fiber traits. There are 960 MLs in the MAGIC population, which is larger than the natural population used in association analysis of upland cotton conducted in the previous studies (Islam et al. 2016; Nie et al. 2016; Sun et al. 2017), which ensures a high efficiency in QTL mapping.
The LD decay is an important factor that determines the mapping resolution of an association analysis. The LD level is affected by recombination and some factors that generate change of allele frequency distribution, such as mutations, gene drift, selection, and migration. Recombination is the major factor that decides the speed of LD decay. On the other hand, the LD can reflect the historical condition of recombination in most inbred plant populations. For example, Zea mays, which is an allogamy crop, was confirmed to have short attenuation distances of < 100 kb (Gore et al. 2009; Dinesh et al. 2016). The average distance of LD decay in upland cotton was more than 3 cM (~ 1.5 Mb) in the previous studies using SSR markers (Abdurakhmonov et al. 2008; Fang et al. 2013; Li et al. 2016a). The results limit the depth exploration of QTL mapping and gene discovery, such as map-based cloning. However, the increase in the recombination rate through artificial crossing could efficiently break the genetic linkage and speed up LD decay. In our study, the distance of the LD decay of the MAGIC population was 0.76 cM (r2 = 0.1), which was estimated with 284 SSR markers. It is a relatively low LD level that represents suitable precision for QTL mapping.
Association mapping based on a MAGIC population is a powerful and reliable tool for detecting variation in genomic regions. There were three statistical models, including GLM (naive, Q, and PCA), MLM (K, Q + K, and PCA + K), and AD test, that were used in association analyses (Yang et al. 2014; Sun et al. 2016; Xu et al. 2016). In general, the population structure is the major factor that increases the number of false positives in marker–trait association (Yu et al. 2006). The convergent cross could eliminate or attenuate the effect of population structure. Given the noninterference from structure for a MAGIC population and the preponderance of MLM (Mackay et al. 2014; Sallam and Martsch 2015), we chose the MLM (K) model for the current study. Although the low density of SSR markers resulting from low polymorphism is inconvenient for delicate loci and gene mining, 52 significant loci were identified for six fiber quality traits (Table 2; Fig. 3), which provides preliminary and systematic studies for the genetic basis of fiber quality traits. In addition, MLM (K) is a reliable method for complex trait association analysis, and most of the loci were simultaneously identified by phenotypic data after BLUP as well as traits in a single environment (Table 3). The marker–trait loci identified by MLM (K) were also frequently shared with the GLM model. Furthermore, compared to the previous studies of QTL mapping in cotton using SSR markers published in an online database (http://www2.cottonqtldb.org:8081), 14 markers coincided with the previous results. Nine of them, including BNL1231, BNL3089, BNL3452, DPL0354, Gh277, HAU1129, HAU2026, NAU3419, and NAU6468, were reported to be located in QTLs for fiber traits. DPL0345 and NAU3419 were stable loci associated with fiber length that were also detected by two previous researchers (Cao et al. 2014; Jamshed et al. 2016). The SSR markers BNL1231 and BNL3452 were two hotspots mapped with multi-traits, including fiber quality, yield, and resistance traits (Table S11) (Said et al. 2015). In addition, MON_CGR5525, Gh185, BNL1231, HAU1321, MON_SHIN0376a, TMB1838, MON_DC20017, MON_DPL0441, and NBRI_HQ526817 were the new loci that control more than two fiber traits (Table S10). These loci express pleiotropism and create multiple effects in fiber development.
Along with the development of sequencing technology and genomics, association mapping has been used for candidate gene mining (Ma et al. 2016; Andres et al. 2017). The identified significant SSR markers have been anchored on physical locations that provided candidate regions in confidence interval based on LD decay distance. The genes located in these regions could be further verified by fine mapping or using reverse genetics methods. The candidate genes could also be screened by genome annotation (Sun et al. 2016). For example, the annotation gene for Gh_A06G0913 in Arabidopsis thaliana, AT5G25170, has been reported to code PPPDE putative thiol peptidase family protein which is related to cell development (Kim et al. 2013), while Gh_A06G0913 is located in MON_DPL0375c-FUHML locus (Table S12). In addition, the genes with known functions can also provide reference for candidate genes. GhPIP1-2, GhPIP2-4, GhACT1, and GhCER6, located in candidate regions of HAU4552b (SFC), MON_CGR5707 (MV), MON_CGR5352b (FE), HAU1058 (FUHML), and MON_DPL0542 (FUHML), were involved in fiber elongation or development proved by gene expression or transgenic technology (Luo et al. 2003; Li et al. 2005, 2013a; Qin et al. 2007; Liu et al. 2008). Besides, numbers of genes near the candidate markers but out of confidence regions should also be taken attention as linkage or cluster, such as GhXTH, GhFBP7, GhGA20ox2, GhKNL1, GhHOX3-A, GhPRF1_D, GhWLIM1a, GhSusA1, GhF3H, GhHOX3-D, and GhCaM7 were also important during fiber growth (Lee et al. 2010; Xiao et al. 2010; Zhang et al. 2011; Argiriou et al. 2012; Jiang et al. 2012; Han et al. 2013; Tan et al. 2013; Gong et al. 2014; Shan et al. 2014; Tang et al. 2014). The candidate loci and genes found in our study are a valuable resource for cotton genetic analyses as well as cotton improvement.
In conclusion, a cotton MAGIC population containing 960 lines was developed from 8 parents, and the population showed abundant phenotypic and genetic variation as well as a low LD level. The association mapping based on 284 SSR markers provides useful information for understanding the genetic basis of fiber traits and for genetic improvement of cotton fiber quality.
References
Abdurakhmonov IY, Kohel RJ, Yu JZ, Pepper AE, Abdullaev AA, Kushanov FN, Salakhutdinov IB, Buriev ZT, Saha S, Scheffler BE, Jenkins JN, Abdukarimov A (2008) Molecular diversity and association mapping of fiber quality traits in exotic G. hirsutum L. germplasm. Genomics 92:478–487
Adams DC, Rohlf FJ (2000) Ecological character displacement in Plethodon: biomechanical differences found from a geometric morphometric study. Proc Natl Acad Sci USA 97(8):4106
Agarwal G, Sabbavarapu MM, Singh VK, Thudi M, Sheelamary S, Gaur PM, Varshney RK (2015) Identification of a non-redundant set of 202 in silico SSR markers and applicability of a select set in chickpea (Cicer arietinum L.). Euphytica 205:381–394
Altschul SF, Boguski MS, Gish W, Wootton JC (1994) Issues in searching molecular sequence databases. Nat Genet 6(2):119–129
Andres RJ, Coneva V, Frank MH, Tuttle JR, Samayoa LF, Han SW, Kaur B, Zhu L, Fang H, Bowman DT, Rojas-Pierce M, Haigler CH, Jones DC, Holland JB, Chitwood DH, Kuraparthy V (2017) Modifications to a LATE MERISTEM IDENTITY1 gene are responsible for the major leaf shapes of Upland cotton (Gossypium hirsutum L.). Proc Natl Acad Sci USA 114:57–66
Argiriou A, Kalivas A, Michailidis G, Tsaftaris A (2012) Characterization of PROFILIN genes from allotetraploid (Gossypium hirsutum) cotton and its diploid progenitors and expression analysis in cotton genotypes differing in fiber characteristics. Mol Biol Rep 39:3523–3532
Atwell S, Huang YS, Vilhjalmsson BJ, Willems G, Horton M, Li Y, Meng D, Platt A, Tarone AM, Hu TT, Jiang R, Muliyati NW, Zhang X, Amer MA, Baxter I, Brachi B, Chory J, Dean C, Debieu M, de Meaux J, Ecker JR, Faure N, Kniskern JM, Jones JD, Michael T, Nemri A, Roux F, Salt DE, Tang C, Todesco M, Traw MB, Weigel D, Marjoram P, Borevitz JO, Bergelson J, Nordborg M (2010) Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465:627–631
Bandillo N, Raghavan C, Muyco PA, Sevilla MAL, Lobina IT, Dilla-Ermita CJ, Tung CW, McCouch S, Thomson M, Mauleon R, Singh RK, Gregorio G, Redona E, Leung H (2013) Multi-parent advanced generation inter-cross (MAGIC) populations in rice: progress and potential for genetics research and breeding. Rice 6(1):11
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Bush WS, Moore JH (2012) Chap. 11: Genome-wide association studies. PLoS Comput Biol 8:e1002822
Cao ZB, Wang P, Zhu XF, Chen H, Zhang TZ (2014) SSR marker-assisted improvement of fiber qualities in Gossypium hirsutum using G. barbadense introgression lines. Theor Appl Genet 127(3):587–594
Cavanagh C, Morell M, Mackay I, Powell W (2008) From mutations to MAGIC: resources for gene discovery, validation and delivery in crop plants. Curr Opin Plant Biol 11:215–221
Delhaize E, Rathjen TM, Cavanagh CR (2015) The genetics of rhizosheath size in a multiparent mapping population of wheat. J Exp Bot 66:4527–4536
Dell’Acqua M, Gatti DM, Pea G, Cattonaro F, Coppens F, Magris G, Hlaing AL, Aung HH, Nelissen H, Baute J, Frascaroli E, Churchill GA, Inze D, Morgante M, Pe ME (2015) Genetic properties of the MAGIC maize population: a new platform for high definition QTL mapping in Zea mays. Genome Biol 16:167
Dinesh A, Patil A, Zaidi PH, Kuchanur PH, Vinayan MT, Seetharam K (2016) Genetic diversity, linkage disequilibrium and population structure among CIMMYT maize inbred lines, selected for heat tolerance study. Maydica 61(3):M29
Fang DD, Hinze LL, Percy RG, Li P, Deng D, Thyssen G (2013) A microsatellite-based genome-wide analysis of genetic diversity and linkage disequilibrium in Upland cotton (Gossypium hirsutum L.) cultivars from major cotton-growing countries. Euphytica 191:391–401
Fang XM, Liu XY, Wang XQ, Wang WW, Liu DX, Zhang J, Liu DJ, Teng ZH, Tan ZY, Liu F, Zhang FJ, Jiang MC, Jia XL, Zhong JW, Yang JH, Zhang ZS (2017) Fine-mapping qFS07.1 controlling fiber strength in upland cotton (Gossypium hirsutum L.). Theor Appl Genet 130:795–806
Gong SY, Huang GQ, Sun X, Qin LX, Li Y, Zhou L, Li XB (2014) Cotton KNL1, encoding a class II KNOX transcription factor, is involved in regulation of fibre development. J Exp Bot 6(15):4133–4147
Gore MA, Chia JM, Elshire RJ, Sun Q, Ersoz ES, Hurwitz BL, Peiffer JA, McMullen MD, Grills GS, Ross-Ibarra J, Ware DH, Buckler ES (2009) A first-generation haplotype map of maize. Science 326:1115–1117
Guo K, Du XQ, Tu LL, Tang WX, Wang PC, Wang MJ, Liu Z, Zhang XL (2016) Fibre elongation requires normal redox homeostasis modulated by cytosolic ascorbate peroxidase in cotton (Gossypium hirsutum). J Exp Bot 67:3289–3301
Haigler CH, Betancur L, Stiff MR, Tuttle JR (2012) Cotton fiber: a powerful single-cell model for cell wall and cellulose research. Front Plant Sci 3:104
Han LB, Li YB, Wang HY, Wu XM, Li CL, Luo M, Wu SJ, Kong ZS, Pei Y, Jiao GL, Xia GX (2013) The dual functions of WLIM1a in cell elongation and secondary wall formation in developing cotton fibers. Plant Cell 25(11):4421–4438
Han LJ, Chen J, Mace ES, Liu YS, Zhu MJ, Yuyama N, Jordan DR, Cai HW (2015) Fine mapping of qGW1, a major QTL for grain weight in sorghum. Theor Appl Genet 128:1813–1825
Hardy OJ, Vekemans X (2002) SPAGEDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Huang XQ, Paulo MJ, Boer M, Effgen S, Keizer P, Koornneef M, van Eeuwijk FA (2011) Analysis of natural allelic variation in Arabidopsis using a multiparent recombinant inbred line population. Proc Natl Acad Sci USA 108:4488–4493
Huang BE, George AW, Forrest KL, Kilian A, Hayden MJ, Morell MK, Cavanagh CR (2012) A multiparent advanced generation inter-cross population for genetic analysis in wheat. Plant Biotechnol J 10:826–839
Huang BE, Verbyla KL, Verbyla AP, Raghavan C, Singh VK, Gaur P, Leung H, Varshney RK, Cavanagh CR (2015) MAGIC populations in crops: current status and future prospects. Theor Appl Genet 128:999–1017
Huang C, Nie XH, Shen C, You CY, Li W, Zhan WX, Zhang XL, Lin ZX (2017) Population structure and genetic basis of the agronomic traits of upland cotton in China revealed by a genomewide association study using high-density SNPs. Plant Biotechnol J. https://doi.org/10.1111/pbi.12722
Ingvarsson PK, Street NR (2011) Association genetics of complex traits in plants. New Phytol 189:909–922
Islam MS, Thyssen GN, Jenkins JN, Zeng L, Delhom CD, McCarty JC, Deng DD, Hinchliffe DJ, Jones DC, Fang DD (2016) A MAGIC population-based genome-wide association study reveals functional association of GhRBB1_A07 gene with superior fiber quality in cotton. BMC Genom 17:903
Jamshed M, Jia F, Gong J, Palanga KK, Shi Y, Li J, Shang H, Liu A, Chen T, Zhang Z, Cai J, Ge Q, Liu Z, Lu Q, Deng X, Tan Y, Rashid HO, Sarfarz Z, Hassan M, Gong W, Yuan Y (2016) Identification of stable quantitative trait loci (QTLs) for fiber quality traits across multiple environments in Gossypium hirsutum recombinant inbred line population. BMC Genom 17(1):197
Jiang YJ, Guo WZ, Zhu HY, Ruan YL, Zhang TZ (2012) Overexpression of GhSusA1 increases plant biomass and improves cotton fiber yield and quality. Plant Biotechnol J 10:301–312
Kim Y, Jo H, Lim CJ (2013) Deubiquitinating activity of Sdu1, a putative member of the PPPDE peptidase family, in Schizosaccharomyces pombe. Can J Microbiol 59:789–796
Lee J, Burns TH, Light G, Sun Y, Fokar M, Kasukabe Y, Fujisawa K, Maekawa Y, Allen RD (2010) Xyloglucan endotransglycosylase/hydrolase genes in cotton and their role in fiber elongation. Planta 232:1191–1205
Li XB, Fan XP, Wang XL, Cai L, Yang WC (2005) The cotton ACTIN1 gene is functionally expressed in fibers and participates in fiber elongation. Plant Cell 17:859–875
Li DD, Ruan XM, Zhang J, Wu YJ, Wang XL, Li XB (2013a) Cotton plasma membrane intrinsic protein 2 s (PIP2s) selectively interact to regulate their water channel activities and are required for fibre development. New Phytol 199:695–707
Li H, Peng ZY, Yang XH, Wang WD, Fu JJ, Wang JH, Han YJ, Chai YC, Guo TT, Yang N, Liu J, Warburton ML, Cheng YB, Hao XM, Zhang P, Zhao JY, Liu YJ, Wang GY, Li JS, Yan JB (2013b) Genome-wide association study dissects the genetic architecture of oil biosynthesis in maize kernels. Nat Genet 45:43–50
Li CQ, Ai NJ, Zhu YJ, Wang YQ, Chen XD, Li F, Hu QY, Wang QL (2016a) Association mapping and favourable allele exploration for plant architecture traits in upland cotton (Gossypium hirsutum L.) accessions. J Agr Sci 154:567–583
Li XM, Jin X, Wang HT, Zhang XL, Lin ZX (2016b) Structure, evolution, and comparative genomics of tetraploid cotton based on a high-density genetic linkage map. DNA Res 23(3):283–293
Lin ZX, He DH, Zhang XL, Nie YC, Guo XP, Feng CH, Stewart JM (2005) Linkage map construction and mapping QTL for cotton fibre quality using SRAP, SSR and RAPD. Plant Breeding 124:180–187
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Liu DQ, Tu LL, Wang L, Li YJ, Zhu LF, Zhang XL (2008) Characterization and expression of plasma and tonoplast membrane aquaporins in elongating cotton fibers. Plant Cell Rep 27:1385–1394
Liu GC, Mei HX, Wang S, Li XH, Zhu XF, Zhang TZ (2015a) Association mapping of seed oil and protein contents in upland cotton. Euphytica 205:637–645
Liu YX, Wang L, Mao SS, Liu K, Lu YL, Wang JR, Wei YM, Zheng YL (2015b) Genome-wide association study of 29 morphological traits in Aegilops tauschii. Sci Rep 5:15562
Liu HJ, Shi JP, Sun CL, Gong H, Fan XM, Qiu FZ, Huang XH, Feng Q, Zheng XX, Yuan NN, Li CS, Zhang ZY, Deng YT, Wang JC, Pan GT, Han B, Lai JS, Wu YR (2016a) Gene duplication confers enhanced expression of 27-kDa gamma-zein for endosperm modification in quality protein maize. Proc Natl Acad Sci USA 113:4964–4969
Liu S, Fan CC, Li JN, Cai GQ, Yang QY, Wu J, Yi XQ, Zhang CY, Zhou YM (2016b) A genome-wide association study reveals novel elite allelic variations in seed oil content of Brassica napus. Theor Appl Genet 129(6):1203–1215
Luo M, Xiao YH, Hou L, Luo XY, Li DM, Pei Y (2003) Cloning and expression analysis of a LIM-domain protein gene from cotton (Gossypium hirsuturm L.). J Genet Genomics 30(2):175–182
Ma D, Hu Y, Yang CQ, Liu BL, Fang L, Wan Q, Liang WH, Mei GF, Wang LJ, Wang HP, Ding LY, Dong CG, Pan MQ, Chen JD, Wang S, Chen SQ, Cai CP, Zhu XF, Guan XY, Zhou BL, Zhu SJ, Wang JW, Guo WZ, Chen XY, Zhang TZ (2016) Genetic basis for glandular trichome formation in cotton. Nat Commun 7:10456
Mackay I, Powell W (2007) Methods for linkage disequilibrium mapping in crops. Trends Plant Sci 12:57–63
Mackay IJ, Bansept-Basler P, Barber T, Bentley AR, Cockram J, Gosman N, Greenland AJ, Horsnell R, Howells R, O’Sullivan DM, Rose GA, Howell PJ (2014) An eight-parent multiparent advanced generation inter-cross population for winter-sown wheat: creation, properties, and validation. G3-Genes Genom Genet 4:1603–1610
Nie XH, Huang C, You CY, Li W, Zhao WX, Shen C, Zhang BB, Wang HT, Yan ZH, Dai BS, Wang MJ, Zhang XL, Lin ZX (2016) Genome-wide SSR-based association mapping for fiber quality in nation-wide upland cotton inbreed cultivars in China. BMC Genom 17:352
Page JT, Huynh MD, Liechty ZS, Grupp K, Stelly D, Hulse AM, Ashrafi H, Van Deynze A, Wendel JF, Udall JA (2013) Insights into the evolution of cotton diploids and polyploids from whole-genome re-sequencing. G3-Genes Genom Genet 3:1809–1818
Pascual L, Albert E, Sauvage C, Duangjit J, Bouchet JP, Bitton F, Desplat N, Brunel D, Le Paslier MC, Ranc N, Bruguier L, Chauchard B, Verschave P, Causse M (2016) Dissecting quantitative trait variation in the resequencing era: complementarity of bi-parental, multi-parental and association panels. Plant Sci 242:120–130
Paterson AH, Brubaker CL, Wendel JF (1993) A rapid method for extraction of cotton. Plant Mol Biol Rep 11(2):1993–1993
Qin YM, Zhu YX (2011) How cotton fibers elongate: a tale of linear cell-growth mode. Curr Opin Plant Biol 14:106–111
Qin YM, Hu CY, Pang Y, Kastaniotis AJ, Hiltunen JK, Zhu YX (2007) Saturated very-long-chain fatty acids promote cotton fiber and Arabidopsis cell elongation by activating ethylene biosynthesis. Plant Cell 19:3692–3704
Said JI, Song M, Wang H, Lin Z, Zhang X, Fang DD, Zhang J (2015) A comparative meta-analysis of QTL between intraspecific Gossypium hirsutum and interspecific G. hirsutum × G. barbadense populations. Mol Genet Genom 290:1003–1025
Sallam A, Martsch R (2015) Association mapping for frost tolerance using multi-parent advanced generation inter-cross (MAGIC) population in faba bean (Vicia faba L.). Genetica 143:501–514
Smykal P, Varshney RK, Singh VK, Coyne CJ, Domoney C, Kejnovsky E, Warkentin T (2016) From Mendel’s discovery on pea to today’s plant genetics and breeding: commemorating the 150th anniversary of the reading of Mendel’s discovery. Theor Appl Genet 129:2267–2280
Sun CM, Wang BQ, Yan L, Hu KN, Liu S, Zhou YM, Guan CY, Zhang ZQ, Li JN, Zhang JF, Chen S, Wen J, Ma CZ, Tu J, Shen JX, Fu TD, Yi B (2016) Genome-wide association study provides insight into the genetic control of plant height in rapeseed (Brassica napus L.). Front Plant Sci 7:1102
Sun ZW, Wang XF, Liu ZW, Gu QS, Zhang Y, Li ZK, Ke HF, Yang J, Wu JH, Wu LQ, Zhang GY, Zhang CY, Ma ZY (2017) Genome-wide association study discovered genetic variation and candidate genes of fibre quality traits in Gossypium hirsutum L. Plant Biotechnol J 15(8):982–996
Tan JF, Tu LL, Deng FL, Hu HY, Nie YC, Zhang XL (2013) A genetic and metabolic analysis revealed that cotton fiber cell development was retarded by flavonoid naringenin. Plant Physiol 162:86–95
Tang WX, Tu LL, Yang XY, Tan JF, Deng FL, Hao J, Guo K, Lindsey K, Zhang XL (2014) The calcium sensor GhCaM7 promotes cotton fiber elongation by modulating reactive oxygen species (ROS) production. New Phytol 202(2):509–520
Walford SA, Wu Y, Llewellyn DJ, Dennis ES (2012) Epidermal cell differentiation in cotton mediated by the homeodomain leucine zipper gene, GhHD-1. Plant J 71:464–478
Wang M, Yan JB, Zhao JR, Song W, Zhang XB, Xiao YN, Zheng YL (2012) Genome-wide association study (GWAS) of resistance to head smut in maize. Plant Sci 196:125–131
Xiao YH, Li DM, Yin MH, Li XB, Zhang M, Wang YJ, Dong J, Zhao J, Luo M, Luo XY, Hou L, Hu L, Pei Y (2010) Gibberellin 20-oxidase promotes initiation and elongation of cotton fibers by regulating gibberellin synthesis. J Plant Physiol 167:829–837
Xu YB, Lu YL, Xie CX, Gao SB, Wan JM, Prasanna BM (2012) Whole-genome strategies for marker-assisted plant breeding. Mol Breeding 29:833–854
Xu LP, Hu KN, Zhang ZQ, Guan CY, Chen S, Hua W, Li JN, Wen J, Yi B, Shen JX, Ma CZ, Tu JX, Fu TD (2016) Genome-wide association study reveals the genetic architecture of flowering time in rapeseed (Brassica napus L.). DNA Res 23:43–52
Yan JB, Shah T, Warburton ML, Buckler ES, McMullen MD, Crouch J (2009) Genetic characterization and linkage disequilibrium estimation of a global maize collection using SNP markers. PloS ONE 4:e8451
Yang J, Ferreira T, Morris AP, Medland SE, Consortium, Replication DIG, Meta-analysis C, Madden PA, Heath AC, Martin NG, Montgomery GW, Weedon MN, Loos RJ, Frayling TM, McCarthy MI, Hirschhorn JN, Goddard ME, Visscher PM (2012) Genetic Investigation of ANthropometric Traits (GIANT). Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat Genet 44:369–375
Yang N, Lu YL, Yang XH, Huang J, Zhou Y, Ali F, Wen WW, Liu J, Li J, Yan JB (2014) Genome wide association studies using a new nonparametric model reveal the genetic architecture of 17 agronomic traits in an enlarged maize association panel. PLoS Genet 10:e1004573
Yu JM, Pressoir G, Briggs WH, Vroh Bi I, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Yu J, Holland JB, McMullen MD, Buckler ES (2008) Genetic design and statistical power of nested association mapping in maize. Genetics 178:539–551
Zhang HB, Li Y, Wang B, Chee PW (2008) Recent advances in cotton genomics. Int J Plant Genom 2008:742304
Zhang M, Zheng XL, Song SQ, Zeng QW, Hou L, Li DM, Zhao J, Wei Y, Li XB, Luo M, Xiao YH, Luo XY, Zhang JF, Xiang CB, Pei Y (2011) Spatiotemporal manipulation of auxin biosynthesis in cotton ovule epidermal cells enhances fiber yield and quality. Nat Biotechnol 29:453–458
Zhang TZ, Hu Y, Jiang WK, Fang L, Guan XY, Chen JD, Zhang JB, Saski CA, Scheffler BE, Stelly DM, Hulse-Kemp AM, Wan Q, Liu BL, Liu CX, Wang S, Pan MQ, Wang YK, Wang DW, Ye WX, Chang LJ, Zhang WP, Song QX, Kirkbride RC, Chen XY, Dennis E, Llewellyn DJ, Peterson DG, Thaxton P, Jones DC, Wang Q, Xu XY, Zhang H, Wu HT, Zhou L, Mei GF, Chen SQ, Tian Y, Xiang D, Li XH, Ding J, Zuo QY, Tao LN, Liu Y, Li J, Lin Y, Hui YY, Cao ZS, Cai CP, Zhu XF, Jiang Z, Zhou BL, Guo WZ, Li RQ, Chen ZJ (2015) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol 33:531–537
Zhao K, Tung CW, Eizenga GC, Wright MH, Ali ML, Price AH, Norton GJ, Islam MR, Reynolds A, Mezey J, McClung AM, Bustamante CD, McCouch SR (2011) Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2:467
Zhao YL, Wang HM, Chen W, Li YH, Gong HY, Sang XH, Huo FC, Zeng FC (2014) Genetic diversity and population structure of elite cotton (Gossypium hirsutum L.) germplasm revealed by SSR markers. Plant Syst Evol 301:327–336
Acknowledgements
We would like to thank Dingguo Li and Xiaofang Li for developing the MAGIC population as well as thank Wu Li, Chao Shen, Tianwang Wen, Bin Gao, De Zhu, and Muhammad Mahmood Ahmed for their assistance with collection of trait data. This research was supported by the National Natural Science Foundation of China (Grant No. 31371674) and the Special Fund for Agro-scientific Research in the Public Interest (Grant No. 201303008).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
The authors declare that this study complies with the current laws of the country in which the experiments were performed. This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Histogram frequency distribution of pairwise relative kinship coefficients estimated in MLs (TIFF 258 KB)
Fig. S2
Principal component analysis of 8 PMs and 960 Mls. The top two principal components, PC1 and PC2, are illustrated in the bottom panels (TIFF 910 KB)
Fig. S3
Phenotype histogram and the Q–Q plots in association analysis. (a)–(f) The phenotype histogram of fiber upper half mean length (FUHML), fiber strength (FS), fiber elongation (FE), fiber uniformity (FU), micronaire value (MV), and short fiber content (SFC). (g)-(l) The Q–Q plots of GLM and K models for FUHML, FS, FE, FU, MV, and SFC (TIF 1600 KB)
Rights and permissions
About this article
Cite this article
Huang, C., Shen, C., Wen, T. et al. SSR-based association mapping of fiber quality in upland cotton using an eight-way MAGIC population. Mol Genet Genomics 293, 793–805 (2018). https://doi.org/10.1007/s00438-018-1419-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-018-1419-4