Abstract
Upland cotton (Gossypium hirsutum L.) is the most important fiber crop, and its lint-yield improvement is impeded due to its narrow genetic base and the lack of understanding of the genetic basis of yield. Backcross inbred lines (BILs) or near-isogenic lines (NILs) in the same genetic background differing in lint yield, developed through advanced backcrossing, provide an important genomic resource to study the molecular genetic basis of lint yield. In the present study, a high-yield (HY) group and a low-yield (LY) group each with three BILs were selected from a BIL population between G. hirsutum and G. barbadense. Using a microarray-based comparative transcriptome analysis on developing fibers at 10 days post-anthesis (DPA) between the two groups, 1486 differentially expressed genes (DEGs) were identified. A total of 212 DEGs were further mapped in the regions of 24 yield QTL and 11 yield trait QTL hotspots as reported previously, and 81 DEGs mapped with the 7 lint-yield QTL identified in the BIL population from which the two sets of BILs were selected. Gene Ontology annotations and Blast-Mapping-Annotation-KEGG analysis via Blast2GO revealed that more DEGs were associated with catalytic activity and binding, followed by transporters, nucleic acid binding transcription factors, structural molecules and molecular transducer activities. Six DEGs were chosen for a quantitative RT-PCR assay, and the results were consistent with the microarray analysis. The development of DEGs-based markers revealed that 7 single strand conformation polymorphism-based single nucleotide polymorphic (SSCP-SNP) markers were associated with yield traits, and 3 markers with lint yield. In the present study, we identified a number of yield and yield component QTL-co-localizing DEGs and developed several DEG-based SSCP-SNP markers for the traits, thereby providing a set of candidate genes for molecular breeding and genetic manipulation of lint yield in cotton.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cotton is an important economic crop worldwide and represents the most important natural fiber in the textile industry. China is the leading producer of cotton, particularly in the North China Plain, the Yangtze River Valley, the Yellow River Valley, and the Xinjiang Uygur Autonomous (Northwest) Region. In recent years, the arable land per capita has rapidly decreased, reflecting the increasing human population and decreasing the amount of arable land due to urbanization. Therefore, enhancing fiber yield through breeding has become a major goal for most cotton improvement programs.
The genus Gossypium comprises 46 diploid (2n = 2x = 26) and 5 allotetraploid (2n = 4x = 52) species (Wendel and Albert 1992). The allotetraploid cotton species, including two commercially important cultivated species, G. hirsutum (Upland cotton) and G. barbadense (Sea-Island, or Pima, or Egyptian cotton), originated from a natural hybridization of an extant A and an extant D genome followed by chromosome doubling 1–2 million years ago (Wendel et al. 2009). Upland cotton produces more than 95 % of the world cotton fibers due to its high lint yield and wide adaptations, while G. barbadense is the only cultivated tetraploid relative of G. hirsutum and is valued for its premium fiber quality attributes. Combining the yield from G. hirsutum and the fiber quality from G. barbadense, has been an ongoing goal of cotton breeders and geneticists, with limited success (Zhang et al. 2014).
Cotton fibers are single-celled trichomes that terminally differentiate from single cells located in the epidermis of the ovule. Although all epidermal cells are potential fibers, only approximately 30 % of these cells actually differentiate into fibers (Basra and Malik 1984; Tiwari and Wilkins 1995). Morphologically, the initiation and elongation of each fiber cell is associated with the spherical expansion and protrusion of an epidermal cell above the ovular surface during anthesis. Accordingly, increasing fiber initials and, therefore, lint percentage (i.e., percent of lint weight in seedcotton weight) will contribute to high lint yield. Therefore, understanding the mechanisms that determine which epidermal cells differentiate into fibers is essential for increasing fiber productivity through genetic engineering and molecular breeding. The identification and characterization of quantitative trait loci (QTL) for yield and yield component traits in cotton have been a research focus for more than a decade. In cotton, QTL for agronomic traits, including yield and yield components, have been identified using F2 (Guo et al. 2007), F2/F2:3 (Zhang et al. 2005; Ulloa et al. 2005; He et al. 2005; Yu et al. 2013b), recombinant inbred lines (RILs) (Shen et al. 2007; Wang et al. 2007; Wu et al. 2009; Zhang et al. 2008; Lacape et al. 2010; Yu et al. 2012), or backcross inbred lines (BILs) (Yu et al. 2012, 2013a). Yu et al. (2013a) used a BIL population in G. hirsutum × G. barbadense and identified 39 QTL affecting yield and yield component traits, including lint yield, seedcotton yield, lint percentage, and boll weight. Said et al. (2013, 2015a, b) recently performed the most comprehensive meta-analyses of QTL reported to date. RNA-Seq has been recently used to study the transcriptomes of the two cotton species, i.e., G. barbadense (Pima cotton) and G. hirsutum (Upland including Acala cotton), during boll development (Pang et al. 2012; Lacape et al. 2012; Fang et al. 2014; Paterson et al. 2012). However, it is unknown whether these genes are yield-related or species-specific, and genetic and physical map-based cloning of lint yield-related genes has not been reported. In fact, no genome-wide analyses of differentially expressed genes (DEGs) have been performed to identify genes associated with lint yield and its component traits using an appropriate genetic population in cotton.
In recent years, microarray analysis and RNA-Seq have been employed to identify DEGs in cotton (e.g., Hinchliffe et al. 2010; Curtiss et al. 2011; Gilbert et al. 2013). Compared to a deep RNA-Seq, although the number of unigenes represented in the Affymetrix GeneChips was limited, the number of DEGs identified was similar to that between two NILs differing in fiber length from a deep genome-wide RNA-Seq (Jinfa Zhang, unpublished). Therefore, it is likely that each unigene in the GeneChips may represent several homologous genes when it was designed before the Upland cotton genome TM-1 was recently sequenced (Li et al. 2015; Zhang et al. 2015). Furthermore, the predicted number of genes in the tetraploid Upland cotton is known only until its genome was fully sequenced in 2015. However, there have been no significant reports to date on the use of the microarray array or RNA-Seq technology to identify lint-yield-related transcripts in cotton.
In the present study, RNA from developing fibers at 10 DPA (days post-anthesis) was extracted from both a high-yielding (HY) and a low-yielding (LY) groups each with three BILs and compared for a transcriptome analysis. Lint yield in cotton is the product of boll number, boll weight and lint percentage. Because developing bolls after 10 DPA usually does not abscise, boll number, as one of the important yield components, is determined at this time. Furthermore, 10 DPA fibers are the fastest-growing fibers after fiber initiation, so lint fiber numbers and part of the fiber length are also determined for another yield component-lint percentage. Of course, boll size at 10 DPA also contributes to the final boll weight at maturity. Therefore, we chose 10 DPA fibers to identify differentially expressed genes (DEGs) that may be involved in cotton yield determination during this important fiber developmental stage. However, it should be pointed out that many genes associated with fiber initiation at −3 to 1 DPA (the stage for lint fiber determination) and secondary wall synthesis after 15 DPA (another important stage for fiber weight determination) may be undetected using 10 DPA fibers.
The objective of the present study was to identify DEGs that are associated with lint yield by employing a microarray analysis using Affymetrix GeneChips and to obtain insight into the underlying molecular genetic differences of fiber development in cotton between two groups of backcross inbred lines (BILs) differing in lint yield.
Materials and methods
Plant materials
An interspecific backcross inbred line (BIL) population of 146 lines developed at New Mexico State University was used for the current study. The BIL population was developed from a cross between the high-yield (HY) parent (HYP) Upland cotton SureGrow 747 (SG 747) and the low-yield (LY) parent (LYP) Egyptian cotton (G. barbadense) Giza 75 through two generations of backcrossing using SG 747 as the recurrent parent followed by four generations of selfing. The 146 BILs with a designation of NMGA and their two parents were planted in five environments in three locations: Anyang, Henan province in 2006, 2007 and 2008; Wangjing, Anhui province in 2007; and Aksu, Xinjiang Uyghur Autonomous Region in 2007. The three locations represent the major cotton production regions with three different cultivation systems in China-Yellow River valley (Henan province), Yangtze River valley (Anhui province) and Northwest (Xinjiang Uygur Autonomous Region). The 148 entries were arranged in a randomized complete block design with two replications and single row plots in each environment. The growth conditions, experimental design, and trait including lint-yield evaluation were previously described (Yu et al. 2013a). To represent the BIL population, 17 BILs were selected from the BIL population based on lint yield, boll size, lint percentage and fiber quality traits and analyzed using least significant difference (LSD) tests for significant differences with the SAS software (SAS Institute Inc., USA).
For the current study on lint yield, 6 BILs were then chosen from the 17 BILs based on differences in lint yield and similarities in fiber quality traits. Three BILs (i.e., NMGA-026, NMGA-017, and NMGA-144, designated HY1, HY2, and HY3, respectively) were high-yielding (HY) with an average lint yield of 60.46 kg ha−1, fiber length of 29.32 mm, strength of 28.54 cN tex−1 and micronaire of 4.50; and three other BILs (i.e., NMGA-066, NMGA-095 and NMGA-023, designated LY1, LY2, and LY3, respectively) were low-yielding (LY) with 32.57 kg ha−1, 29.17 mm, 28.76 cN tex−1 and 4.55 for above traits, respectively. The two parents, i.e., SG 747 and Giza 75, differed significantly in lint yield (41.50 vs. 25.53 kg ha−1), fiber length (29.44 vs. 34.02 mm), strength (27.61 vs. 40.23 cN tex−1) and micronaire (5.55 vs. 4.55). The 6 lines, together with their two parents (HYP and LYP), were planted in the Experimental Farm at the Institute of Cotton Research (ICR), Chinese Academy of Agricultural Sciences (CAAS), Anyang, Henan Province, China (E 114°48′, N 36°06′) in the summer of 2009. The field plots were arranged using a randomized complete block design with three replications. Flowers at the day of flowering (i.e., 0 DPA) in each plot were tagged, and cotton boll samples were harvested at 5, 10, 15, 20, and 25 days post anthesis (DPA) before 10:00 a.m. and immediately placed on ice. To minimize experimental errors, all samples within each developmental stage were collected in the same day. Fibers on the ovules of 5–10 developing bolls per replication were dissected, frozen in liquid nitrogen, and stored at −80 °C until further processing.
RNA isolation from cotton fibers
Tissue samples for total RNA isolation were prepared from 1 g of cotton fiber harvested from ovules of each genotype at 5, 10, 15, 20, and 25 DPA on a replication basis. Total RNA was extracted using a Sigma Spectrum™ Plant Total RNA kit (Sigma–Aldrich, St. Louis, MO, USA) with an on-column DNaseI digestion according to the manufacturer’s instructions. An agarose gel electrophoresis was performed and the results showed high RNA integrity, as both 28S and 18S rRNAs in the total RNA were predominantly present (data not shown). The RNA quantity and quality were then determined based on the 260/280 nm ratio and 260/230 nm ratio using a DU®800 spectrophotometer (Beckman Coulter, Brea, CA, USA).
Microarray hybridization and data analysis
For the microarray experiments using the GeneChip® Cotton Genome Array (Affymetrix, Santa Clara, CA, USA), 2 mg of purified total RNA was extracted from 10 DPA fibers from the three biological replications of each HY and LY genotype. The GeneChip® Cotton Genome Array is a single GeneChip brand array comprised of 239,777 probe sets representing 21,854 cotton transcripts, and is based on EST sequences obtained from four cotton species: G. hirsutum, G. raimondii, G. arboreum, and G. barbadense (http://www.affymetrix.com/catalog/131430/AFFY/Cotton+Genome+Array#1_1). The extracted RNA was used for cRNA synthesis and hybridization to the Cotton GeneChip Array based on a standard protocol (Rodriguez-Uribe et al. 2014). The default Affymetrix method MAS 5.0, RMA was used for a primary normalization. The results were submitted to NCBI as a GEO record under the accession number GSE81134 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE81134).
A pairwise comparison between the HY and LY fibers was conducted to identify differentially expressed genes (DEGs). Using the Affymetrix GeneChip Operating System v1.4 (GCOS; Affymetrix), the relative mean signal, detection calls, signal-log ratios and change calls were independently calculated using four different algorithms for each probe set. Fold changes in expression (e.g., >twofold), the Microarray Suite (MAS) 5.0 probe set detection p values, and/or p values (e.g., Bonferroni-corrected p-value threshold <0.05) from the standard statistical tests (e.g., independent t test, Welch’s t test, Wilcoxon’s rank sum test, ANOVA, etc.) were used to identify genes with significant changes in expression. The DEGs were further screened using the Partek software (http://www.partek.com). The Basic Local Alignment Search Tool (BLAST) was used to identify a significant homology between a gene and known gene products. The BLAST search results were subsequently sorted based on gene function to identify DEGs between the HY and LY groups. The GO annotations for the DEGs were obtained using the Blast2GO program (https://www.blast2go.com/). For a pathway enrichment analysis, the DEGs were mapped to the terms in the KEGG (Kyoto Encyclopedia of Genes and Genomes) database using KOBAS 2.0 (KEGG Orthology-Based Annotation System) (Mao et al. 2005). KEGG pathways with corrected p values ≤0.05 were considered statistically significant.
RT-PCR and quantitative reverse transcription PCR
To perform an RT-PCR analysis, the first-strand cDNA was synthesized from 5 µg of total RNA from each tissue sample using the Superscript First-Strand Synthesis System (Invitrogen, Carlsbad, CA, USA). The cDNA fragments were then PCR amplified using gene-specific primers designed from selected DEGs and commercially synthesized (Invitrogen, USA) as listed in Supplementary Data 1 online. The PCR amplification was performed in a PTC-200 thermocycler (Scientific Support, USA). Gel images were then acquired with a gel-scanning imager (Gel Doc-2000, Bio-Rad, USA) and ethidium bromide staining. Based on RT-PCR results, four DEGs were selected for a further real-time quantitative RT-PCR (qRT-PCR) analysis. The qRT-PCR reactions were performed using iTaq™ SYBR Green Super Mix (Bio-Rad Laboratories, USA) via a Bio-Rad CFX96 real-time PCR detection system. The relative quantification (RQ) of the two targets in the treated samples was calculated by the formula \( RQ = E^{{(Ct,{\text{ctrl}} - Ct,{\text{sample}})}} \) , where E is the experimental efficiency of the qPCRs (Bennett et al. 2015). The 18S ribosomal RNA (rRNA) was used as the internal control. Significant differences in gene expression were determined using the t test.
Chromosome location of DEGs in yield trait QTL and QTL hotspots
To locate DEGs with yield and yield trait QTL as anchored by simple sequence repeat (SSR) markers, we collected the marker names in the map of [“Guazuncho2” (G. hirsutum) × “VH8-4602” (G. barbadense)] as published (Said et al. 2013, 2015a). The marker sequence data were downloaded from the CottonGen database (https://www.cottongen.org/data/download/ marker) (Yu et al. 2014). Then, the chromosome locations of markers and DEGs were identified using the BLAST program with the marker sequences and DEG sequences as the query and the genome sequence (Zhang et al. 2015) as the subject. The positions of yield trait QTL hotspot regions as reported by Said et al. (2013, 2015a, b), and the 24 QTL loci for yield as reported by Yu et al. (2013a) on chromosomes were identified using anchoring markers’ locations in the map. The candidate genes in the regions of QTL hotspots or loci were then identified and used for further studies.
Single nucleotide polymorphism (SNP) identification and confirmation using single strand conformation polymorphism (SSCP) analysis
Sequence alignment was conducted using BLAST. Potential SNPs were identified via visual inspection based on a comparative alignment between the 106 DEG sequences obtained from the Affymetrix GeneChip Cotton Genome Array and the 183,035 sequences with SNPs obtained from CottonGen (http://www.cottongen.org/data/download/marker) (E value ≤ 1×10−20). For SNP confirmation, primers annealing immediately upstream or downstream of an SNP site were designed as the forward or reverse primers, respectively (Supplementary Data 2). The primers were designed using the Primer Premier 6.0 software (http://www.premierbiosoft.com) based on the following parameters: 18–26 nucleotides (nt) (20 nt optimum) in primer length, annealing at 55 °C, 35–60 % for GC content, and 100–300 bp for PCR product size. Briefly, each 20 µL PCR reaction contained 10 ng of DNA, 10 µL of TaKaRa Premix Ex Tag, and 0.4 µL of each 20 µM primer. A touchdown PCR was performed in a Bio-Rad C1000 Thermal Cycler with the following conditions: denaturation at 94 °C for 3 min, followed by 29 cycles at 94 °C for 30 s, touchdown primer annealing beginning at 56 °C for 30 s and extension at 72 °C for 60 s, and a final extension at 72 °C for 10 min. The PCR products of SNP markers were resolved using the polyacrylamide gel electrophoresis according to the SSCP method of Lu et al. (2009). Polymorphic markers were scored as 0 for the absence and 1 for the presence of an SSCP fragment in the 17 BILs (from which the 6 BILs were selected for the current study) and were used for a correlation analysis with lint yield, yield traits and fiber quality using SAS (SAS Institute Inc., USA).
Results
Microarray and data analysis
Using the Affymetrix GeneChip Cotton Genome Array with multiple probes for each gene, a comparison between the HY and LY BIL groups further minimized complications due to background differences resulting from the use of only two NILs. The six BILs were selected based on lint yield, boll size, and lint percentage to form a high-yield (HY) group (including three BILs designated HY1, HY2, and HY3) and a low-lint yield (LY) group (with three BILs designated LY1, LY2, and LY3). The two groups of BILs had significant differences in seedcotton yield, lint yield, lint percent, and boll weight, but no significant differences in fiber length and elongation (see "Materials and methods"). Although genetically, the BILs showed 87.5 % recovery of the recurrent parent used in developing the BIL population from which the six BILs were selected, our simple sequence repeat (SSR) and SSCP-SNP analysis of 2349 markers showed that the genetic similarity between the HY group and the LY group was as high as 98.5 % (unpublished), indicating a highly similar background between the lines used in the study. Therefore, although two backcrosses were used in the development of the BIL population, the BILs selected in this study may be considered NILs.
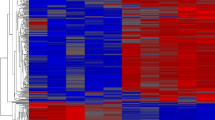
A total of 1486 genes were differentially expressed (DE) between the two groups (≥twofold change; Bonferroni-corrected P value threshold <0.05) (Supplementary Data 3). Compared to the LY group, 278 (19 % of 1486) of the DEGs were up-regulated, and 1208 genes (81 % of 1486) were down-regulated in the HY group. A cluster analysis of the gene expression data from the 1486 DE genes (DEGs) separated all the three HY lines from two of the three LY BILs (Fig. 1). Specifically, HY2 and HY3 had the highest similarity in gene expression patterns, followed by the high similarity of gene expression between LY1 and LY2. The result demonstrated the reliability of the selection of BILs and grouping method used in the present study, providing a great confidence in the obtained results. However, one of the three LY lines, i.e., LY3 was grouped together with the HY group, indicating that high levels of similarity in phenotypes (here lint yield, yield traits and fiber quality) may not always be reflected by high levels of similarity in gene expressions. The DEGs identified between the two BIL groups may indicate a common gene regulation mechanism for fiber development-related (more specifically, yield-related) genes in cotton, but these genes need to be further identified through a co-localization analysis with lint yield QTL.
To identify the developmental processes that differed between the two groups of BILs, the 1486 DEGs were categorized into 38 GO terms based on biological process, cellular component and molecular function, respectively (Fig. 2). Under the cellular component category, DEGs were further categorized as cell (521 genes, 35.06 % of the 1486 DEGs) and organelle (373 genes, 25.10 %). For the biological processes, metabolic process (649 genes, 43.67 %) and cellular process (544 genes, 36.61 %) represented a majority of the DEGs identified. Under the molecular function category, binding (525 genes, 35.33 %) and catalytic activity (476 genes, 32.03 %) were the two most abundant subcategories followed by transporter activity, nucleic acid binding, transcription factor activity, structural molecule activity, and molecular transducer activity. Interestingly, binding, catalytic, and transporter activity were associated with hormone binding, hydrolase activity, transferase activity, and carbohydrate transporter activity.
A gene expression enrichment analysis was further performed on the 278 genes upregulated in the HY group and 1,208 genes upregulated in the LY group. The 278 upregulated DEGs in the HY group were categorized into 37 GO terms based on the biological process, cellular component, and molecular function, respectively (Fig. 3). For the biological processes, macromolecule metabolic process (29 genes, 10.43 % of the upregulated 278 DEGs), and organic substance biosynthetic process (22 genes, 7.91 %) had the highest numbers of the DEGs identified among the GO terms. Under the molecular function category, ion binding (33 genes, 11.87 %) and transferase activity (22 genes, 7.91 %) were the two most abundant subcategories, followed by hydrolase activity, organic cyclic compound binding, oxidoreductase activity, transmembrane transporter activity, and small molecule binding. Under the cellular component category, the DEGs were further categorized as intracellular (54 genes, 19.42 %) and intracellular organelle (43 genes, 15.47 %). The 1208 upregulated DEGs in the LY group were also categorized into 39 GO terms based on the biological process, cellular component, and molecular function (Fig. 4). For the biological processes, organic substance metabolic process (443 genes, 36.67 % of the 1208 DEGs) and cellular metabolic process (407 genes, 33.39 %) represented a majority of the DEGs identified. Under the molecular function category, organic cyclic compound binding (346 genes, 28.64 %) and heterocyclic compound binding (346 genes, 28.64 %) were the two most abundant subcategories, followed by ion binding, small molecule binding, hydrolase activity, transferase activity, carbohydrate derivative binding, oxidoreductase activity, and protein binding. Under the cellular component category, DEGs were further categorized as intracellular (433 genes, 35.84 %) and intracellular part (427 genes, 35.35 %).
The 1486 DEGs identified from the microarray analysis were further analyzed using KOBAS (i.e., KEGG Orthology-Based Annotation System) to identify the metabolic pathways in which they function. KOBAS mapped 467 of the 1486 DEGs to 102 KEGG pathways. The pathways with the highest unigene representation were starch and sucrose metabolism (32, 6.85 %), phenylpropanoid biosynthesis (24, 5.14 %), and phenylalanine metabolism (23, 4.93 %). As expected (Gilbert et al. 2013), starch and sucrose metabolism was the most abundant pathway in fiber development at 10 DPA (Table 1).
Corroboration of the microarray data
To validate the results obtained from the microarray analysis, a RT-PCR analysis of eight genes was first performed on developing fibers at 15, 20, and 25 DPA from the six BILs and their parents. The eight genes included ribosomal protein L16 (designated GhRPL16), kinesin-related 11 (designated GhKIF11), filament-like plant protein 4 (designated GhFPP4), a gene of unknown function (designated Unknown), alpha-expansin 4 (designated GhEXPA4), ABC transporter G family member 10-like (designated GhABC-10), bacterial-induced peroxidase (designated GhPOX), and glucose-1-phosphate adenylyltransferase large subunit 3 (designated GhAPL3). Once different expression levels were found among the eight genotypes, 4 of the 8 genes were chosen for a further quantitative RT-PCR (qRT-PCR) analysis on developing fibers at 10 DPA. These four genes included two up-regulated and two down-regulated DEGs in the HY group identified by the microarray analysis (Fig. 5). The selected genes were GhEXPA4 (GhiAffx.26845.1.S1_at), GhRPL16 (Ghi.10647.1.S1_s_at), GhAPL3 (Gra.1083.1.A1_at), and Unknown (GhiAffx.32232.1.S1_at). Based on the results from the six BILs and the two parents, the coefficients of correlation between microarray and qRT-PCR were positive for two genes and significantly positive for another gene, indicating a relative congruence between the microarray and qRT-PCR results. The microarray analysis of the alpha-expansin 4 gene showed a reduction in gene expression in LY compared to HY, and similar results were obtained from the qRT-PCR analysis (Fig. 5). One of the four probe sets targeting the Unknown gene (GhiAffx.32232.1.S1_at) exhibited a reduction in expression in the LY group as compared with the HY group, which was consistent with the qRT-PCR analysis (Fig. 5). Two of the four probe sets targeting the genes encoding for ribosomal protein GhPRL16 (Ghi.10647.1.S1_s_at) and glucose-1-phosphate adenylyltransferase large subunit 3 (Gra.1083.1.A1_at) exhibited increased expression in the LY group compared to the HY group, congruent with the results of qRT-PCR analysis (Fig. 5). The results indicated that the differential expression of the four genes between the HY and LY groups was consistent between both the qRT-PCR and microarray analyses.
Quantitative RT-PCR analysis of GhEXPA4, an unknown gene, GhRPL16 and GhAPL3, and its comparison with the results from a microarray analysis in developing fibers at 10 days post-anthesis (DPA). LYP, low-yielding parent Giza 75, LY1, NMGA-066, LY2, NMGA-095, LY3, NMGA-023, HY1, NMGA026, HY2, NMGA-17, HY3, NMGA-144, HYP, high-yielding (HY) parent SG747
Two other genes, i.e., the gene coding for bacterial-induced peroxidase (GhPOX) (Ghi.3212.1.A1_at) and the gene coding for kinesin-related 11 (GhK1F11) (Ghi.10183.1.S1_at), were finally chosen for a dynamic qRT-PCR analysis of fibers at various fiber development stages (i.e., 5, 10, 15, 20, and 25 DPA). The expression of GhPOX was the highest in fibers at 10 DPA than in the other fiber developmental stages, and the expression of GhPOX was up-regulated in the LY group than in the HY group in 5-15 DPA, especially 10 DPA fibers (Fig. 6). However, for the GhKIF11 gene, its expression at the mRNA level was the highest at the 5 DPA fibers and then linearly decreased until 20–25 DPA at a negligent level. Furthermore, its expression in fibers at 5 and 10 DPA was higher in the LY lines than in the HY lines. These results were again consistent with the microarray analysis for the two genes in fibers at 10 DPA.
Candidate DEGs in lint-yield QTL regions identified in the BIL population
The completion of the G. hirsutum (AD) genome sequencing (Li et al. 2015; Zhang et al. 2015) has made it possible to determine the chromosome distribution of the DEGs identified in this study. Based on published data for the cotton genome, out of the 1486 DEGs, 536 were mapped onto chromosome locations from At_chr1 to At_chr13 of the At subgenome, 698 to Dt_chr1 to Dt_chr13 of the Dt subgenome, and the remaining DEGs (i.e., 125 genes) were on unmapped scaffolds. Interestingly, 115 genes were homologous to genes on both the At subgenome and the Dt subgenome at the sequence identify of 79.89–99.45 %.
Co-localization of quantitative trait loci (QTL) with the identified 1486 DEGs will facilitate the identification of candidate genes for lint yield (LY) and its component traits (Supplementary Data 4). We mapped 212 DEGs with the 24 yield and yield trait QTL (Yu et al. 2013a), and 12 yield trait QTL hotspots per a previous meta-analysis (Said et al. 2013) (Supplementary Data 5). These 212 genes were distributed in A01 (4 DEGs), A03 (2), A05 (1), A07 (26), A09 (6), A10 (4), A11 (2), A12 (1), A13 (8), D02 (38), D03 (8), D04 (9), D06 (20), D07 (9), D11 (8), D12 (31), and D13 (25). Of the 212 DEGs, 156 were categorized into 47 GO terms based on the biological process, cellular component, and molecular function (Fig. 7). Under the cellular component category, the DEGs were further categorized as cell part (89 genes, 42.58 % of 212 DEGs) and membrane-bounded organelle (67 genes, 32.06 %). For the biological process, organic substance metabolic process (55 gene, 26.32 %), and primary metabolic process (55 gene, 26.32 %) had more DEGs identified. Under the molecular function category, ion binding (51 genes, 24.40 %), heterocyclic compound binding (30 genes, 14.35 %), and organic cyclic compound binding (30 genes, 14.35 %) were the three most abundant subcategories, followed by hydrolase activity, oxidoreductase activity, transferase activity, lyase activity, and isomerase activity.
Since the six BILs were selected from a G. hirsutum × G. barbadense backcross inbred line (BIL) population analysis (Yu et al. 2013a), we focused on the 11 lint-yield QTL identified in the BILs to identify lint-yield-related DEGs. Of the 11 yield QTL, 7 were found to co-localize with 81 DEGs, as shown in Supplementary Data 6. The 7 QTL for lint yield were qLY-06A-c1-1 on At_chr1, qLY-06A-c7-1 on At_chr7, qLY-07A-c9-1 on At_chr9, qLY-07A-c13-1 on At_chr13, qLY-07X-c14-1 on Dt_chr2, and qLY-07X-c18-1 and qLY-06A-c18-1 in different regions on Dt_chr13.
The At subgenome carried more LY QTL with DEGs. The qLY-06A-c1-1 on At-chr1 had three co-localized DEGs encoding for a blue copper protein, a lysine histidine transporter-like 8, and a dva-1 polyprotein. The qLY-06A-c7-1 on At_chr07 had 2 co-localized DEGs encoding for an RNA-binding-protein 38 and an NAD-binding rossmann-fold superfamily protein. The qLY-07A-c9-1 locus had one co-localized DEG encoding for a mitotic checkpoint serine threonine-protein kinase isoform 2. The qLY-07A-c13-1 on At_chr13 carried a histone gene.
On the Dt subgenome, the qLY-07X-c14-1 locus had 30 co-localized DEGs encoding for a transcription factor 21, a calcium-dependent lipid-binding (domain) family protein isoform 3, protein far-red elongated hypocotyl 3, beta-glucosidase family protein, among others. The qLY-07X-c18-1 locus had 21 co-localized DEGs encoding for a DNA replication licensing factor mcm 4, branched-chain-amino-acid aminotransferase-like protein 2, protein root initiation defective 3-like, microtubule-associated protein 65-1 isoform 1, among others. The qLY-06A-c18-1 locus had one co-localized DEG encoding for a calmodulin-binding-protein isoform 1.
Mapping more DEGs with previously reported yield trait QTL hotspots on the G. hirsutum genome
Since the BIL population, including the six lines used in this study only mapped 7 QTL for lint yield, explaining 6.88–16.77 % of the phenotypic variation, we also used the 20 yield trait QTL hotspots reported from a meta-analysis (Said et al. 2013) to identify more lint-yield-related genes. A total of 70 DEGs were found to co-localize with 12 yield trait QTL hotspots for boll weight (BW), lint percent (LP), seedcotton yield (SCY), lint yield (LY), and boll number (BN), while other yield trait QTL hotspots carried no DEGs (Fig. 8). The 12 yield trait QTL hotspots with DEGs were mapped onto 11 chromosomes, including 4 At and 7 Dt chromosomes, i.e., At_chr3, At_chr9, At_chr12, At_chr13, Dt_chr2, Dt_chr3, Dt_chr4, Dt_chr6, Dt_chr7, Dt_chr12, and Dt_chr13 (Fig. 8).
Of these 12 yield trait QTL hotspots, 6 were mapped to 3 pairs of homoelogous chromosomes (i.e., At_chr3 vs. Dt_chr3, At_chr12 vs. Dt_chr12, and At_chr13 vs. Dt_chr13). The c3_BN/LP_mQTL_Hotspots(5) was identified at 1.76–2.82 cM, and carried 5 QTL on At_chr3, and contained 2 DEGs, encoding for an isoform 3 and a pectin lyase-like superfamily protein isoform 1. The c17_LP/BW/_mQTL_Hotspots(4) with 4 QTL residing at 22.25–39.58 cM on Dt_chr3 contained 8 DEGs, including genes encoding for proteins, such as ubiquitin carboxyl-terminal hydrolase isozyme l5, RNA polymerase II transcriptional coactivator kelp, luminal-binding-protein 5, activating signal cointegrator 1, DNAJ heat shock N-terminal domain-containing, RNA pseudouridine synthase 7 isoform × 2, copper transporter 1-like, and scarecrow-like protein 9. Since the two hotspots were not located in the same regions of the two homeologous chromosomes (At_chr3 vs. Dt_chr3), no common DEGs were identified.
Similarly, no common DEGs were identified for QTL on other two pairs of homeologous chromosomes. The c12_LP/YLD/LP_mQTL_Hotspots(5) was identified at 76.44–77.19 cM on At_chr12 and carried 5 QTL, and it contained only 1 DEG encoding for a serine threonine-protein kinase. As a comparison, the c26_BW/LY/LP/YLD_mQTL_Hotspots(4) was identified at 50.67–59.04 on Dt_chr12 and carried 4 QTL, and it contained 16 DEGs, including genes encoding for an acid beta-fructofuranosidase-like, protein cornichon homolog 4-like, protein iq-domain 1, among others. Another QTL hotspot on At_chr13 named c13_LY/BW/SCY_mQTL_Hotspots(3) was identified at 79.04-79.57 cM and carried 3 QTL, containing 5 DEGs coding for alpha-glucan isozyme, histone h4, histone h2b, probable serine threonine-protein kinase, and serine threonine-protein phosphatase 6 regulatory subunit 3. Only one DEG was co-localized with the c18_LP/BW_mQTL_Hotspots(4) carrying 4 QTL at 59.98–60.35 cM on Dt_chr13, which encodes for a rubisco subunit binding-protein alpha subunit.
In addition to the above three At-subgenome chromosomes, the c9_LP/SCY/LY/BN_mQTL_Hotspots(10) with 10 QTL ranged from 1.12 to 6.98 cM on At_chr9 and contained 5 genes, including genes encoding for a calcium-activated outward-rectifying potassium isoform 1, alpha- glucan phosphorylase l chloroplastic amyloplastic for 2 genes, chaperone protein chloroplastic, and e3 ubiquitin-protein ligase ring1-like.
Four additional Dt subgenome chromosomes carried yield trait QTL hotspots. A yield trait QTL hotspot, i.e., c14_BW/LP/LY_mQTL_Hotspots(9) carrying 9 QTL, was located at 0–5.13 cM on Dt_chr2, and it contained 8 DEGs, encoding for proteins, such as calreticulin 3 isoform 1, selenium-binding-protein 2-like, cytochrome p450, chalcone synthase, dihydroxy-acid chloroplastic, glutamine synthetase leaf chloroplastic, non-specific lipid-transfer protein at2g13820-like, and transcription factor 21. The c22_LP/BW/SCY_mQTL_Hotspots(12) with 12 QTL ranged from 46.37 to 50.57 cM in Dt_chr4, and contained 6 DEGs encoding for a magnesium transporter mrs2-4-like, organ-specific protein s2-like, a squalene monooxygenase, a polygalacturonase at1g48100, an eukaryotic translation initiation factor 5b, and an unknown gene. The c25_BW_mQTL_Hotspots(13) ranged from 0.69 to 2.82 cM with 13 QTL on Dt_chr6, and it contained 6 DEGs, including genes encoding for a c3hc4 transcription factor, an allene oxide synthase, an ent-kaurene chloroplastic, a hypothetical protein F383_19939, a probable isoaspartyl peptidase l-asparaginase 2, and a proteasome subunit alpha type-3. The c16_LY/YLD/LP/SCY/LY_mQTL_Hotspots(6) with 6 QTL ranged from 18.34 to 29.32 cM on Dt_chr7, and contained 9 co-localized DEGs coding for a ubiquitin-conjugating enzyme19 isoform 1, a sulfite reductase, a hydrophobic lti6a, a transcription factor myb86-like protein, a uncharacterized calcium-binding-protein at1g02270-like isoform x1, an auxin response factor 18, an aspartic proteinase nepenthesin-1, an o-acyltransferase wsd1-like, and a golgi snap receptor complex member 1–2.
Single nucleotide polymorphic (SNP) locus identification and association with agronomic traits
To illustrate the utility of the DEGs identified in the present study, a sequence alignment was conducted via BLAST with 106 gene sequences from DEGs against 183,035 sequences in a reference database downloaded from the CottonGen database (http://www.cottongen.org/data/download/marker). The results showed that 56 gene sequences did not contain SNPs, whereas 45 gene sequences contained 209 putative SNPs (E value ≤ 1 × 10−20). An example of the results of the PCR amplification using the SSCP method for genomic DNAs from the six NILs used in this study is shown in Fig. 9. Among the 45 primer pairs, 24 pairs (53 %) yielded polymorphic SSCP markers, revealing a total of 81 SSCP-SNP loci (Supplementary Data 7). One gene (gb|DW225231.1) was identified to encode for a protein kinase and it carried 6 SNP loci which was located in c25_BW_mQTL_Hotspots(13). Another gene (gb|L04497.1) was identified to encode for myb-like transcription factor 1 and contained 1 SNP locus which was located in the same region for QTL qLY-07A-c13-1 and qSCY-07A-c13-1. A further correlation analysis between SSCP markers and 10 agronomic traits in the 17 BILs revealed that 26 markers were significantly (at P < 0.05 for 22 markers) or highly significantly (at P < 0.01 for 4 markers) associated with one or more agronomic traits, including 7 markers for yield component traits, and 3 makers for lint yield, 17 markers for fiber quality, and 1 marker for cottonseed oil content (Supplementary Data 8). Interestingly, repulsion linkage phases were found for several polymorphic SSCP markers amplified using the same primer pairs designed for 3 genes, resulting in opposite correlations. This may reflect the nature of co-dominance of these markers amplified by the same primer pairs designed from the same genes.
Electrophoretic analysis of single strand conformation polymorphism (SSCP)-based single nucleotide polymorphic (SNP) markers developed for the genes encoding for MYB (a), RDL (b), and MADS-3 (c). 1 LY1, 2 NMGA-060, 3 HY3, 4 NMGA-101, 5 HY1, 6 NMGA-062, 7 NMGA-092, 8 NMGA-043, 9 NMGA-140, 10 NMGA-089, 11 LY3, 12 LY2, 13 NMGA-070, 14 NMGA-051, 15 NMGA-105, 16 NMGA-012, and 17 HY2
Discussion
The present study performed a microarray-based comparative transcriptome analysis on developing fibers at 10 days post-anthesis (DPA) between the two groups (high yield—HY vs. low yield—LY) of BILs. A total of 1486 differentially expressed genes (DEGs) were identified, providing the first glimpse of DEGs into the relationship to genetic variation of fiber yield in cotton. Several fiber development-related genes that exhibited altered expression profiles in HY group and LY group were validated through qRT-PCR. Specifically, gene expression levels of GhKIF11 and GhPOX genes were relatively higher at 15 and 20 DPA in the LY group than in the HY group. We further mapped 212 DEGs with the 24 yield QTL and 11 yield trait QTL hotspots as reported by others, and 81 DEGs with the 7 lint-yield QTL mapped in the BIL population from which the two groups of BILs were chosen for this study. Molecular markers from the DEGs closely associated with the lint-yield-related QTL were developed. The results will provide a good foundation for fine mapping and map-based cloning of lint-yield genes and facilitate targeted gene transfer from G. barbadense to Upland cotton to further improve cotton yield.
Mapped DEGs with the yield QTL and yield trait QTL hotspots as reported by others, and the SSCP markers associated with important agronomic traits from the association analysis provide a good foundation for map-based cloning and MAS of lint-yield QTL. Thus, this study took one of the first comprehensive genomic approaches to genotype and profile the interspecific Pima × Upland backcross inbred lines during fiber development to associate DNA markers and gene expression levels with yield traits (Michaelson et al. 2009; Pang et al. 2012). In cotton, several microarray platforms have been developed and used to study the expression of genes associated with fiber development (Arpat et al. 2004; Shi et al. 2006; Wu et al. 2005, 2007; Lee et al. 2007), fiber traits or cotton evolution via comparisons of genotypes with differing fiber qualities (Hinchliffe et al. 2005, 2010; Wu et al. 2008; Li et al. 2010), or normal genotypes vs. mutants (Lee et al. 2006; Shi et al. 2006; Yang et al. 2006; Bolton et al. 2009; Gilbert et al. 2013). However, a comprehensive comparison between the results obtained in the present study and previously published results is difficult to obtain due to the use of different genetic systems. To our knowledge, this fiber yield DEG study is the first to use genotyping and expression profiling to examine BILs from an interspecific Egyptian cotton × Upland cotton BIL population during fiber development to associate DNA markers and gene expression levels with yield traits.
Lint yield is the product of boll number, boll weight and lint percentage. Boll weight is the sum of lint weight (lint fiber number × fiber length × fiber fineness) and seed weight, and lint percentage is the ratio of lint weight by boll weight. Fiber fineness can be measured by the diameter of the mature fiber, but it is usually determined by micronaire (i.e., the fiber weight per-unit fiber length). Therefore, it should be recognized that the number of mature fibers developed from fiber initials is also a contributing factor to lint yield. To identify DEGs for the understanding of its genetic and genomic basis, it will require a similar comparative transcriptome study among BILs or NILs differing in fiber initials using ovules at the fiber initiation stage, i.e., −3 to 1 DPA. Furthermore, because fiber fineness (usually measured based on micronaire), as an important contributing factor to fiber weight, is majorly determined during secondary wall biosynthesis stage after 15 DPA, many genes associated with fiber fineness are unlikely detected using 10 DPA fibers. Therefore, the current study is not intended to detect most DEGs that are associated with lint yield and its component traits due to the use of 10 DPA fibers that did not cover the entire spectrum of fiber initiation and development and the genetic variation in yield limited by the BILs used in this study.
Since the microarray analysis was performed in 2009–2010 prior to the publication of the tetraploid cotton genome sequences (Li et al. 2015; Zhang et al. 2015), two DEGs genes, i.e., GhKIF11 and GhPOD, were chosen for a dynamic analysis of gene expression during fiber development. GhKIF11 expression levels were relatively higher at 15 and 20 DPA in the LY group compared to the HY group, but it was not located in any of the QTL regions for lint yield and yield traits. Previous studies have identified more than a dozen of kinesins that are abundantly expressed during fiber development in Upland cotton. Kong et al. (2015) reported a member of the kinesin-4 subfamily named GhKINESIN-4A that shares the highest homology to the FRA1 (fragile fiber 1) protein previously reported in Arabidopsis thaliana. FRA1 or KINESIN-4A has been implicated in the deposition and orientation of cellulose microfibrils during secondary cell wall synthesis in fiber cells in A. thaliana (Zhou et al. 2007). A kinesin-like calmodulin-binding-protein involved in long-chain fatty acid biosynthesis showed greater than 50-fold accumulation in 10 DPA fiber cells compared with 0 DPA ovules (Ji et al. 2003). Preuss et al. (2003, 2004) proposed that MT-based motor kinesins actively participate in fiber growth and identified several kinesin-encoding genes that are abundantly expressed in developing cotton fibers. Tiwari and Wilkins (1995) reported diffuse cell expansion of fiber development for more than 10 days prior to the initiation of secondary cell wall deposition. Although these results demonstrate that the GhKIF11 gene is closely associated with fiber cell wall biosynthesis, the differential expression of GhKIF11 between the LY and HY groups was not one of the causal genes underlying lint-yield QTL in our study. Its expression may be directly or indirectly regulated by one of the lint-yield QTL, and therefore, it is indirectly related to lint yield.
The GhPOX (encoding for peroxidase) expression levels in the LY group were also relatively higher at 10 and 15 DPA, compared with the HY group. The transcript level of GhPOX increased >5.0-fold at 10 DPA (i.e., the period of fast fiber cell elongation) in the LY group. Most plant POX isozymes catalyze oxidoreductions between H2O2 and variable reductants and have been implicated in many physiological processes, such as the polymerization of cell wall compounds, defense against pathogen attacks, salt tolerance, oxidative stress, and the metabolism of phytohormones and alkaloids (Hiraga et al. 2001; Gabaldón et al. 2005; Passardi et al. 2005; Costa et al. 2008). A total of 73 AtPOX and 138 OsPOX genes were identified within the A. thaliana and Oryza sativa genomes, respectively (Tognolli et al. 2002; Welinder et al. 2002; Duroux and Welinder 2003; Passardi et al. 2004). A microarray analysis of G. hirsutum gene expression revealed that the Class III peroxidase genes were up-regulated in fast elongating fiber cells (Shi et al. 2006). Another microarray analysis indicated that GhPOX1 plays an important role during fiber cell elongation through the production of reactive oxygen species (Mei et al. 2009). Similar to GhKIF11, this gene was also not located in any of the lint-yield QTL regions, and therefore, is unlikely one of the candidate genes for lint-yield QTL.
However, through a comparative genome-wide gene expression study between two groups of BILs contrasting in lint yield and co-localization of DEGs with lint yield and yield trait QTL, 1486 DEGs were narrowed to 212 genes in the QTL regions, providing an important list of genes for further studies. An in silico analysis of the QTL-co-localized DEGs based on the sequenced TM-1 genome allowed identification of putative single nucleotide polymorphisms (SNPs) for the development of SSCP-SNP markers for an association analysis with the lint yield and yield traits, resulting in the identification of several yield and yield trait-associated DEG-based SNP markers. Although more studies are needed, this positional (physical map) candidate gene approach will assist in the fast and targeted identification of candidate genes for lint yield and other traits to facilitate molecular breeding and map-based cloning of QTL in cotton.
Abbreviations
- HY:
-
High-yield
- LY:
-
Low-yield
- HYP:
-
HY parent
- LYP:
-
LY parent
- DEGs:
-
Differentially expressed genes
- NILs:
-
Near isogenic lines
- DPA:
-
Days post-anthesis
- QTL:
-
Quantitative trait loci
References
Arpat A, Waugh M, Sullivan J, Gonzales M, Frisch D et al (2004) Functional genomics of cell elongation in developing cotton fibers. Plant Mol Biol 54:911–929
Basra AS, Malik CP (1984) Development of the cotton fiber. Int Rev Cytol 89:65–113
Bennett J, Hondred D, Register JC (2015) Keeping qRT-PCR rigorous and biologically relevant. Plant Cell Rep 34:1–3
Bolton JJ, Soliman KM, Wilkins TA, Jenkins JN (2009) Aberrant expression of critical genes during secondary cell wall biogenesis in a cotton mutant, Ligon Lintless-1 (Li-1). Comp Funct Genomics 65:9301–9308
Costa MM, Hilliou F, Duarte P, Pereira LG, Almeida I et al (2008) Molecular cloning and characterization of a vacuolar class III peroxidase involved in the metabolism of anticancer alkaloids in Catharanthus roseus. Plant Physiol 146:403–417
Curtiss J, Rodriguez-Uribe L, Stewart JM, Zhang JF (2011) Identification of differentially expressed genes associated with semigamy in Pima cotton (Gossypium barbadense L.) through comparative microarray analysis. BMC Plant Biol 11:49
Duroux L, Welinder KG (2003) The peroxidase gene family in plants: a phylogenetic overview. J Mol Evol 57:397–407
Fang L, Tian R, Li X, Chen J, Wang S et al (2014) Cotton fiber elongation network revealed by expression profiling of longer fiber lines introgressed with different Gossypium barbadense chromosome segments. BMC Genom 15:838–853
Gabaldón C, López-Serrano M, Pedreno MA, Ros Barceló A (2005) Cloning and molecular characterization of the basic peroxidase isoenzyme from Zinnia elegans, an enzyme involve in lignin biosynthesis. Plant Physiol 139:1138–1154
Gilbert MK, Turley RB, Kim HJ, Li P, Thyssen G (2013) Transcript profiling by microarray and marker analysis of the short cotton (Gossypium hirsutum L.) fiber mutant Ligon lintless-1 (Li1). BMC Genom 14:403–420
Guo WZ, Cai CP, Wang CB, Han ZG, Song XL et al (2007) A microsatellite-based, gene-rich linkage map reveals genome structure, function and evolution in Gossypium. Genetics 176:527–541
He D, Lin Z, Zhang X, Nie Y, Guo X et al (2005) Mapping QTLs of traits contributing to yield and analysis of genetic effects in tetraploid cotton. Euphytica 144:141–149
Hinchliffe DJ, Wilkins W, Cantrell RG, Zhang JF (2005) Comparative microarray analysis of genes differentially expressed during fiber development of Upland and Pima cotton. In: Proceedings of Beltwide Cotton Conference, p883–896
Hinchliffe DJ, Meredith WR, Yeater KM, Kim HJ, Woodward AW (2010) Triplett BA: near-isogenic cotton germplasm lines that differ in fiber-bundle strength have temporal differences in fiber gene expression patterns as revealed by comparative high-throughput profiling. Theor Appl Genet 120:1347–1366
Hiraga S, Sasaki K, Ito H, Ohashi Y, Matsui H (2001) A large family of class III plant peroxidases. Plant Cell Physiol 42:462–468
Ji SJ, Lu YC, Feng JX, Wei G, Li J (2003) Isolation and analyses of genes preferentially expressed during early cotton fiber development by subtractive PCR and cDNA array. Nucl Acids Res 31(10):2534–2543
Kong Z, Ioki M, Braybrook S, Li S, Ye ZH (2015) Kinesin-4 functions in vesicular transport on cortical microtubules and regulates cell wall mechanics during cell elongation in plants. Mol Plant 8:1011–1023
Lacape JM, Llewellyn D, Jacobs J, Arioli T, Becker D (2010) Meta-analysis of cotton fiber quality QTLs across diverse environments in a Gossypium hirsutum x G. barbadense RIL population. BMC Plant Biol 10:132–156
Lacape JM, Claverie M, Vidal RO, Carazzolle MF, Guimaraes PGA (2012) Deep sequencing reveals differences in the transcriptional landscapes of fibers from two cultivated species of cotton. PLoS One 7:10–19
Lee JJ, Hassan OS, Gao W, Wei NE, Kohel RJ (2006) Developmental and gene expression analyses of a cotton naked seed mutant. Planta 223:418–432
Lee JJ, Woodward AW, Chen ZJ (2007) Gene expression changes and early events in cotton fibre development. Ann Bot 100:1391–1401
Li LY, Yu JW, Zhai HH, Huang SL, Li XL (2010) Comparative analysis of cotton fiber development related gene expression profiling. Mol Plant Breed 8:488–496
Li XM, Gao WH, Guo HL, Zhang XL, Fang DD (2014) Development of EST-based SNP and InDel markers and their utilization in tetraploid cotton genetic mapping. BMC Genom 15:1046–1047
Li FG, Fan GY, Lu CR, Xiao GH, Zou CS et al (2015) Genome sequence of cultivated Upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat Biotechnol 33:524–529
Lu YZ, Curtiss J, Percy RG, Hughs SE, Yu JW (2009) DNA polymorphisms of genes involved in fiber development in a selected set of cultivated tetraploid cotton. Crop Sci 49:1695–1704
Mao X, Cai T, Olyarchuk JG, Wei L (2005) Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21:3787–3793
Mei WQ, Qin YM, Song WQ, Li J, Zhu YX (2009) Cotton GhPOX1 encoding plant class III peroxidase may be responsible for the high level of reactive oxygen species production that is related to cotton fiber elongation. J Genet Genomics 36:141–150
Michaelson JJ, Loguercio S, Beyer A (2009) Detection and interpretation of expression quantitative trait loci (eQTL). Methods 48:265–276
Pang MX, Percy RG, Stewart JM, Hughs E, Zhang JF (2012) Comparative transcriptome analysis of Pima and Acala cotton during boll development using 454 pyrosequencing technology. Mol Breed 30:1143–1153
Passardi F, Longet D, Penel C, Dunand C (2004) The class III peroxidase multigenic in land plants family in rice and its evolution. Phytochemistry 65:1879–1893
Passardi F, Cosio C, Penel C, Dunand C (2005) Peroxidases have more functions than a Swiss army knife. Plant Cell Rep 24:255–265
Paterson AH, Wendel JF, Gundlach H, Guo H, Jenkins J et al (2012) Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492:423–427
Preuss ML, Delmer DP, Liu B (2003) The cotton kinesin-like calmodulin-binding protein associates with cortical microtubules in cotton fibers. Plant Physiol 132:154–160
Preuss ML, Kovar DR, Lee YR, Staiger CJ, Delmer DP, Liu B (2004) A plant-specific kinesin binds to actin microfilaments and interacts with cortical microtubules in cotton fibers. Plant Physiol 136:3945–3955
Rodriguez-Uribe L, Abdelraheem A, Tiwari R, Sengupta-Gapalan C, Hughs SE, Zhang JF (2014) Identification of drought response genes in a drought tolerant cotton (Gossypium hirsutum L.) under irrigated field conditions and development of candidate gene markers for drought tolerance. Mol Breed 14:1776–1796
Said JI, Lin ZX, Zhang XL, Song MZ, Zhang JF (2013) A comprehensive meta QTL analysis for fiber quality, yield, yield related and morphological traits, drought tolerance, and disease resistance in tetraploid cotton. BMC Genom 14:776–798
Said JI, Knapka JA, Song MZ, Zhang JF (2015a) Cotton QTLdb: a cotton QTL database for QTL analysis, visualization, and comparison between Gossypium hirsutum and G. hirsutum x G. barbadense populations. Mol Genet Genom 290:1615–1625
Said JI, Song MZ, Wang HT, Lin ZX, Zhang XL, Fang DD, Zhang JF (2015b) A comparative meta-analysis of QTL between intraspecific Gossypium hirsutum and interspecific G. hirsutum × G. barbadense populations. Mol Genet Genom 290:1003–1025
Shen X, Guo W, Lu Q, Zhu X, Yuan Y (2007) Genetic mapping of quantitative trait loci for fiber quality and yield trait by RIL approach in Upland cotton. Euphytica 155:371–380
Shi YH, Zhu SW, Mao XZ, Feng JX, Qin YM (2006) Transcriptome profiling, molecular biological, and physiological studies reveal a major role for ethylene in cotton fiber cell elongation. Plant Cell 18:651–664
Tiwari SC, Wilkins TA (1995) Cotton (Gossypium hirsutum) seed trichomes expand via diffuse growing mechanism. Can J Bot 73:746–757
Tognolli M, Penel C, Creppin J, Simon P (2002) Analysis and expression of the large class III peroxidase gene family in Arabidopsis thaliana. Gene 288:129–138
Ulloa M, Saha S, Jenkins JN, Meredith WR, McCarty JC (2005) Chromosomal assignment of RFLP linkage groups harboring important QTLs on an intraspecific cotton (Gossypium hirsutum L.) Joinmap. J Hered 96:132–144
Wang B, Guo W, Zhu X, Wu Y, Huang N (2007) QTL mapping of yield and yield components for elite hybrid derived-RILs in upland cotton. J Genet Genomics 34:35–45
Welinder KG, Justesen AF, Kjærsgård IVH, Jensen RB, Rasmussen SK et al (2002) Structural diversity and transcription of class III peroxidases from Arabidopsis thaliana. Eur J Biochem 269:6063–6081
Wendel J, Albert VA (1992) Phylogenetics of the cotton genus (Gossypium): characteristic weighted parsimony analysis of chloroplast-DNA restriction site data and its systematic and biogeographic implications. Syst Bot 17:115–143
Wendel JF, Brubaker C, Alvarez I, Cronn R, Stewart JM et al (2009) Evolution and natural history of the cotton genus. In: Paterson AP (ed) Genetics and Genomics of Cotton. Springer, New York. pp 3–22
Wu Y, Rozenfeld S, Defferrard A, Ruggiero K, Udall JA et al (2005) Cycloheximide treatment of cotton ovules alters the abundance of specific classes of mRNAs and generates novel ESTs for microarray expression profiling. Mol Genet Genom 274:477–493
Wu Y, Llewellyn DJ, White R, Ruggiero K, Al-Ghazi Y et al (2007) Laser capture microdissection and cDNA microarrays used to generate gene expression profiles of the rapidly expanding fibre initial cells on the surface of cotton ovules. Planta 226:1475–1490
Wu Z, Soliman KM, Bolton JJ, Saha S, Jenkins JN (2008) Identification of differentially expressed genes associated with cotton fiber development in a chromosomal substitution line (CS-B22sh). Funct Integr. Genom 8:165–174
Wu J, Gutierrez OA, Jenkins JN, McCarty JC, Zhu J (2009) Quantitative analysis and QTL mapping for agronomic and fiber traits in an RI population of Upland cotton. Euphytica 165:231–245
Yang SS, Cheung F, Lee JJ, Ha M, Wei EN, Triplett BA, Town CD, Chen ZJ (2006) Accumulation of genome-specific transcripts, transcription factors and phytohormonal regulators during early stages of fiber cell development in allotetraploid cotton. Plant J 47:761–775
Yu JW, Yu SX, Fan SL, Song MZ, Zhai HH et al (2012) Mapping quantitative trait loci for cottonseed oil, protein and gossypol content in a Gossypium hirsutum × Gossypium barbadense backcross inbred line population. Euphytica 187:191–201
Yu JW, Zhang K, Yu SX, Fan SL, Song MZ et al (2013a) Mapping quantitative trait loci for lint yield and fiber quality across environments in a Gossypium hirsutum × Gossypium barbadense backcross inbred line population. Theor Appl Genet 126:275–287
Yu JW, Yu SX, Gore M, Wu M, Zhai HH et al (2013b) Identification of quantitative trait loci across interspecific F2, F2:3 and testcross populations for agronomic and fiber traits in tetraploid cotton. Euphytica 191:375–389
Yu J, Jung S, Cheng CH, Ficklin SP, Lee T et al (2014) CottonGen: a genomics, genetics and breeding database for cotton research. Nucl Acids Res 42:1229–1236
Zhang ZS, Xiao YH, Luo M, Li XB, Luo XY et al (2005) Construction of a genetic linkage map and QTL analysis of fiber-related traits in upland cotton (Gossypium hirsutum L). Euphytica 144:91–99
Zhang YX, Lin ZX, Xia QZ, Zhang MJ, Zhang XL (2008a) Characteristics and analysis of simple sequence repeats in the cotton genome based on a linkage map constructed from a BC1 population between Gossypium hirsutum and G. barbadense. Genome 51:534–546
Zhang JF, Percy RG, McCarty JC Jr (2014) Introgression genetics and breeding between Upland and Pima cotton: a review. Euphytica 198:1–12
Zhang TZ, Hu Y, Jiang WK, Fang L, Guan XY et al (2015) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol 33:531–537
Zhang YX, Lin ZX, Xia QZ, Zhang MJ, Zhang XL (2008b) Characteristics and analysis of simple sequence repeats in the cotton genome based on a linkage map constructed from a BC1 population between Gossypium hirsutum and G. barbadense. Genome 51:534–546
Zhou JL, Qiu J, Ye ZH (2007) Alteration in secondary wall deposition by overexpression of the Fragile Fiber1 kinesin-like protein in Arabidopsis. J Integr Plant Biol 49:1235–1243
Acknowledgments
The research was sponsored by a grant from the National Natural Science Foundation of China (Grant 31301367 and Grant 31301368) and the 863 National High-Tech Research Development Program in China (Grant 2012AA101108). The research was also in part supported by the New Mexico Agricultural Experiment Station.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
This study was funded by the National Natural Science Foundation of China (Grant Nos. 31301367; 31301368) and the 863 National High-Tech Research Development Program in China (Grant No. 2012AA101108).
Conflict of interest
The authors declare that they no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
438_2016_1216_MOESM2_ESM.xlsx
Supplementary Data 2. Primer sequences for 45 genes used for the analysis of single strand conformation polymorphism (SSCP)-based single nucleotide polymorphic (SNP) markers (XLSX 14 kb)
438_2016_1216_MOESM3_ESM.xlsx
Supplementary Data 3. Microarray results for the differentially expressed genes (DEGs) identified in the high- and low-yield groups (XLSX 178 kb)
438_2016_1216_MOESM4_ESM.xlsx
Supplementary Data 4. Mapping of DEGs with the 24 yield QTL and 12 yield trait QTL hotspots on the genomes of G. hirsutum (XLSX 68 kb)
Rights and permissions
About this article
Cite this article
Man, W., Zhang, L., Li, X. et al. A comparative transcriptome analysis of two sets of backcross inbred lines differing in lint-yield derived from a Gossypium hirsutum × Gossypium barbadense population. Mol Genet Genomics 291, 1749–1767 (2016). https://doi.org/10.1007/s00438-016-1216-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-016-1216-x