Abstract
Bacterial blight (BB) caused by Xanthomonas oryzae pv. oryzae (Xoo) is the most devastating bacterial disease of rice (Oryza sativa L.), a staple food crop that feeds half of the world’s population. In management of this disease, the most economical and effective approach is cultivating resistant varieties. Due to rapid change of pathogenicity in the pathogen, it is necessary to identify and characterize more host resistance genes for breeding new resistant varieties. We have previously identified the BB resistance (R) gene Xa23 that confers the broadest resistance to Xoo strains isolated from different rice-growing regions and preliminarily mapped the gene within a 1.7 cm region on the long arm of rice chromosome 11. Here, we report fine genetic mapping and in silico analysis of putative candidate genes of Xa23. Based on F2 mapping populations derived from crosses between Xa23-containing rice line CBB23 and susceptible varieties JG30 or IR24, six new STS markers Lj36, Lj46, Lj138, Lj74, A83B4, and Lj13 were developed. Linkage analysis revealed that the new markers were co-segregated with or closely linked to the Xa23 locus. Consequently, the Xa23 gene was mapped within a 0.4 cm region between markers Lj138 and A83B4, in which the co-segregating marker Lj74 was identified. The corresponding physical distance between Lj138 and A83B4 on Nipponbare genome is 49.8 kb. Six Xa23 candidate genes have been annotated, including four candidate genes encoding hypothetical proteins and the other two encoding a putative ADP-ribosylation factor protein and a putative PPR protein. These results will facilitate marker-assisted selection of Xa23 in rice breeding and molecular cloning of this valuable R gene.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial blight (BB), caused by Xanthomonas oryzae pv oryzae (Xoo), is one of the most serious and destructive diseases of rice (Oryza sativa L.). This disease has been attracting many researchers, because it is not only important for rice production worldwide, but also a model system to investigate plant–bacteria interactions (Niño-Liu et al. 2006). As to the management of BB in rice production, adoption of host resistance is practically proven the most economical, effective, and eco-friendly approach (Suh et al. 2013). Therefore, scientists have been putting great efforts to identify BB resistance (R) genes from both cultivated rice varieties and wild rice species (Brar and Khush 1997; Kumar et al. 2012). We have previously identified a broad-spectrum BB resistance gene Xa23 from the wild rice Oryza rufipogon (Zhang et al. 1998). Since the Xa23 locus confers complete dominant and high resistance to virtually all Xoo races tested at all growth stages of rice (Zhang et al. 2002; Wang et al. 2013), it has been widely adopted in rice breeding programs (Zhou et al. 2009; Huang et al. 2012). In the present study, we conducted fine genetic mapping of Xa23 to facilitate marker-assisted selection and molecular cloning of this valuable R gene.
Thus far, about 38 BB resistance genes (designated in a series from Xa1 to Xa38) have been identified in rice (Lin et al. 1996; Khush and Angeles 1999; Chen et al. 2002, 2011; Blair et al. 2003; Lee et al. 2003a; Gu et al. 2004; Tan et al. 2004; Cheema et al. 2008; Korinsak et al. 2009; Wang et al. 2009; Zheng et al. 2009; Guo et al. 2010; Miao et al. 2010; Bhasin et al. 2012). Among the identified BB resistance genes, 11 (xa5, xa8, xa13, xa15, xa19, xa20, xa24, xa26b, xa28, xa32, and xa34) are recessive in inheritance (Chen et al. 2011) and the others are dominant in nature. Most of the identified BB resistance genes have been mapped to rice chromosomes, and eight of them have been molecularly cloned, including dominant genes Xa1 (Yoshimura et al. 1998), Xa3 or Xa26 (Sun et al. 2004; Xiang et al. 2006), Xa10 (Tian et al. 2014), Xa21 (Song et al. 1995), Xa27 (Gu et al. 2005) and recessive genes xa5 (Iyer and McCouch 2004; Jiang et al. 2006), xa13 (Chu et al. 2006) and xa25 (Liu et al. 2011).
Although many BB resistance genes have been identified, most of them hold limited application value due to weak resistance, narrow resistance spectrum or recessive inheritance nature. Consequently, only a few of them, such as Xa3, Xa4, Xa7, and Xa21, have been practically used in rice production. In this regard, more effective BB resistance genes should be identified and characterized.
In our previous investigations, the Xa23 locus has been transferred into a susceptible indica rice variety JG30, resulting in near-isogenic line CBB23 (Zhang et al. 2002). We have previously mapped the Xa23 locus within a 1.7 cm region on the long arm of rice chromosome 11, between molecular markers CP02662 (Wang et al. 2005) and 69B (Wang et al. 2006). We here report genetic fine mapping and in silico analysis of putative candidate genes of Xa23.
Materials and methods
Plant materials
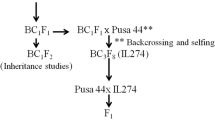
JG30 is an indica rice variety highly susceptible to all Xoo strains tested. CBB23 is a near-isogenic line of Xa23 in genetic background of JG30 (Zhang et al. 2002). IR24 is a Xoo strain PXO99 susceptible indica rice variety from International Rice Research Institute. CBB23 was crossed as male to JG30 and IR24, respectively. The F1 plants were self-pollinated to generate F2 populations. Rice materials used for Xa23 mapping were listed in Table 1. Additional 36 rice varieties/lines used for the polymorphism assays at Xa23 locus were indicated in Fig. 3. All rice plants were grown in field or greenhouse at 28–35 °C in day light.
Bacterial inoculation and plant resistance assessment
The Philippine race 6 (PXO99) of Xoo was used to evaluate disease phenotypes of rice plants. Xoo cells were cultured in PPS medium [ferv-filtering juice of 300 g potato, 5 g peptone, 15 g sucrose, 2 g Na2HPO4·12H2O, and 0.5 g Ca(NO3)2·4H2O] at 28 °C for 48 h. Bacterial suspensions (OD600 = 1.0) with sterile distilled water were inoculated by the leaf-clipping method (Kauffman et al. 1973) in leaves of rice plants at booting stage. For each plant, 3–5 fully expanded leaves were inoculated. Disease symptom was scored 2 weeks post inoculation. The disease symptom was scored by lesion area ratio against the whole leaf through visual assessment (Wang et al. 2005). Plants with lesion areas equal to or less than 15 % were classified as resistant (R) and those with lesion areas larger than 15 % were classified as susceptible (S) plants.
Extraction of genomic DNA and PCR procedure
Genomic DNA of 39 rice varieties/lines and individual plants of the F2 populations were isolated from leaves following the description by McCouch et al. (1988). PCR amplification was carried out with 20 µL reaction volume containing 2 µL of 10× PCR buffer, 1.2 µL of dNTPs (10 mmol L−1), 0.3 µL of each primer (10 µmol L−1), 50 ng of DNA template, 1.0 U of Taq DNA polymerase. The reactions were heated to 94 °C for 3 min followed by 35 cycles of amplification at 94 °C for 30 s, 55–60 °C (depending on primers) for 30 s, 72 °C for 30–60 s, and a final extension at 72 °C for 7 min. Amplified products were separated by electrophoresis in agarose gels with ethidium bromide and photographed under ultraviolet light using the gel documentation system, or separated in 8–10 % denaturing polyacrylamide gel electrophoresis and observed by silver staining.
Development of new molecular markers
Based on Nipponbare reference sequences on NCBI, almost uniformly distributed sequences of the corresponding bacterial artificial chromosome (BAC) or P1-derived artificial chromosome (PAC) clones (http://www.tigr.org) between the markers 69B (Wang et al. 2006) and CP02662 (Wang et al. 2005) were selected for BLAST with all available rice genome sequences to find non-homologous or less homologous regions. Based on this analysis, 107 pairs of STS (sequence-tagged sites) marker primers were designed within this region. New primers were designed to generate PCR fragments ranging 100–1,000 bp in size.
Genetic and physical mapping of Xa23
The genetic map of Xa23 was constructed according to genetic distances of the molecular markers linked to Xa23 locus. Linkage analysis of the polymorphic markers was performed with MAPMAKER/EXP 3.0 (Lincoln et al. 1993). Recombination frequencies were converted to cM using Kosambi function (Kosambi 1944). The physical map was constructed by bioinformatically (http://blast.ncbi.nlm.nih.gov/) locating the linked makers on related BAC and PAC clones of Nipponbare.
Identification of putative candidate genes of Xa23
Based on the targeted region of Xa23 locus, genomic sequence of Nipponbare between the markers Lj138 and A83B4 was downloaded from NCBI (http://www.ncbi.nlm.nih.gov) and analyzed using the online software FGENESH (http://linux1.softberry.com/). Candidate genes in the region of interest were identified by BLAST-Putility (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The Xa23 candidates were finally determined by comparing the FGENESH predictions with those annotated by MSU Rice Genome Annotation Project (http://rice.plantbiology.msu.edu/).
Results
Resistance patterns of rice plants
Plants of two F2 populations derived from the crosses of JG30/CBB23 and IR24/CBB23 were inoculated individually with Xoo strain PXO99 at booting stage. Since disease phenotypes of resistant and susceptible F2 plants were clearly distinguishable, like their R and S parents (Fig. 1a), the lesion areas on the plant leaves were scored by visual assessment (Wang et al. 2005). For the F2 population derived from JG30/CBB23, distribution of F2 plants based on lesion area was bimodal, with an apparent valley at lesion area approximately 16–25 % (Fig. 1b). Inoculation assessment revealed that 1,930 plants of the F2 population were resistant and 632 plants were susceptible; the R:S ratio fits 3:1 well (χ 2 = 0.13, P = 0.72) (Table 1). Likewise, the R:S ratio of the F2 population derived from IR24/CBB23 fits 3:1 perfectly (χ 2 = 0.03, P = 0.86) (Table 1). These results indicated that the BB resistance in CBB23 was controlled by a single dominant gene.
Reaction patterns of rice plants to Xanthomonas oryzae pv. oryzae strain PXO99. a Leaves of parents JG30, CBB23, IR24 and the F1 plants of cross JG30/CBB23 were presented to show the lesion patterns: S susceptible, R resistant. Pictures were taken 14-day post inoculation. b Distribution, based on the ratio of lesion area against the whole leaf area, of 2,562 F2 plants derived from the cross JG30/CBB23. Lesion area (%) was scored 14-day post inoculation
High-resolution genetic and physical mapping of Xa23
In our previous investigations, the Xa23 locus was located within a 1.7 cM region on long arm of rice chromosome 11 between molecular markers 69B (Wang et al. 2006) and CP02662 (Wang et al. 2005). The RFLP marker 69B was developed from the PAC clone 69B15 from indica rice Guangluai 4 (Wang et al. 2006). We then sequenced the ends of 69B and located it, by BLASTN (http://blast.ncbi.nlm.nih.gov/Blast.cgi), at nucleotide position 23846538-23848957 on chromosome 11 of japonica rice Nipponbare (NCBI Reference Sequence: NC_008404). CP02662 is an EST marker located at nucleotide position 24176326-24176528 on Nipponbare chromosome 11 (NC_008404) (Wang et al. 2005). We accordingly speculated that Xa23 locus resides within a region corresponding to the 330-kb region (23846538-24176528 of NC_008404) of Nipponbare. To narrow down the region harboring the Xa23 gene, 107 pairs of PCR primers were designed across the 330-kb sequences by searching the non- or low-homologous sequences. The surveyed results showed that six pairs of the designed primers (Table 2) revealed reliable polymorphisms between JG30 and CBB23 (Fig. 2a). The PCR products amplified from JG30 and CBB23 were sequenced to confirm the polymorphisms.
Genetic and physical mapping of Xa23 gene. a Polymorphisms between JG30 (J) and CBB23 (C) were revealed by STS markers Lj36, Lj46, Lj138, A83B4, Lj13, and Lj74. Molecular genotypes of some susceptible F2 plants revealed by Lj74 were also shown. b Genetic map of Xa23 locus. Xa23 was mapped between the markers Lj138 and A83B4 on chromosome 11 (11S). The six new markers identified in this study, including the co-segregating maker Lj74, were shown in bold. c Physical map of Xa23 locus. Xa23 was located in a region corresponding to a 49.8 kb interval in the PAC clone P0480H08 of Nipponbare
The six newly developed markers Lj36, Lj46, Lj138, Lj74, A83B4, and Lj13 (Table 2) were then used to survey the 632 susceptible and 167 resistant (randomly chosen) F2 individuals derived from cross of JG30/CBB23. Results showed that Lj36, Lj46, Lj138, Lj74, A83B4, and Lj13 revealed 4, 3, 1, 0, 4, and 14 recombinant susceptible individuals, respectively (Table 3). Thus, the Xa23 gene was defined to a 0.4 cM region between markers Lj138 and A83B4, in which the co-segregating marker Lj74 was identified (Table 3; Fig. 2b). To confirm this fine-mapping results, we used markers Lj46, Lj74 and A83B4 to survey all resistant and susceptible F2 individuals derived from IR24/CBB23 (Table 1), and similar genetic map was obtained (data not shown).
The markers were then landed on the reference sequences of Nipponbare by bioinformatical analysis (Fig. 2c). Based on the pairwise BLAST analysis, the BACs and PACs were aligned as a contig map covering the Xa23 locus. The markers Lj46, Lj138, Lj74, and A83B4 were landed on the same PAC clone P0480H08. The physical interval between Lj138 and A83B4 is about 49.8 kb (Fig. 2c).
Xa23 candidate genes
Recently, the quality of Nipponbare reference genome sequences has been improved (Sakai et al. 2013; Kawahara et al. 2013). Within the region flanked by the newly developed markers Lj138 and A83B4 (from 22182291 to 22232136) of the updated Os-Nipponbare-Reference-IRGSP-1.0 (http://rapdb.dna.affrc.go.jp/), nine genes have been annotated (http://rice.plantbiology.msu.edu/), including LOC_Os11g37650 that encodes the DWARF27 protein required for biosynthesis of strigolactones and thereby regulating rice tiller bud outgrowth (Lin et al. 2009) and two transposon protein-encoding genes (LOC_Os11g37590 and LOC_Os11g37600). The remaining six annotated genes are candidates of Xa23 (Table 4). They encode four hypothetical proteins (LOC_Os11g37580, LOC_Os11g37610, LOC_Os11g37620 and LOC_Os11g37630), a putative ADP-ribosylation factor protein (LOC_Os11g37640) and a putative vegetative storage protein (LOC_Os11g37660).
Discussion
The Xa23 locus in CBB23 is originally from wild rice O. rufipogon (Zhang et al. 2002). Studies in the past decade repetitively verified that the single Xa23 locus confers high resistance to more than 30 representative Xoo strains from China, Philippines, Japan, Korea, and Bangladesh. In fact, no naturally occurring Xoo isolate that can overcome the Xa23-mediated resistance has been identified so far (Wang et al. 2013). Furthermore, Xa23 locus confers dominant BB resistance at all growth stages, an important feature for hybrid rice breeding (Zhang et al. 2002). Thus, CBB23 has been widely adopted in rice breeding programs in China (Zhou et al. 2009; Huang et al. 2012). In this study, we identified six new STS markers co-segregated with or closely linked to the Xa23 locus and mapped the Xa23 gene within a 0.4 cM region, corresponding to a 49.8 kb physical distance on Nipponbare genome, in which 6 Xa23 candidate genes have been annotated. The high-resolution genetic map of Xa23 locus and the co-segregating or closely linked markers will certainly facilitate both marker-assisted selection and molecular cloning of Xa23.
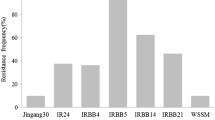
Among the six STS markers developed in this study, Lj36 and Lj13 are dominant markers, generating amplicons in the Xa23-donor parent CBB23 but not in the recipient parent JG30 (Fig. 2a). Lj46, Lj138, and A83B4 are co-dominant markers, but polyacrylamide gel electrophoresis must be adopted to differentiate the CBB23- and JG30-amplicons due to their very small (3–5 bp) differences in size (Table 2; Fig. 2a). Comparatively, Lj74 should be the most effective marker for selection of Xa23, because it is a co-segregating and co-dominant marker, generating CBB23- and JG30-amplicons with 114 bp difference in size (Table 2), clearly differentiated in agarose gel (Fig. 2a). We used Lj74 to survey the polymorphisms of 36 additional rice varieties/lines, including 24 lines harboring different BB resistance genes. The results showed that the specific 983-bp band can be amplified only from Xa23-containing varieties/lines (Fig. 3), indicating that Lj74 would be very useful in rice breeding for the marker-assisted selection of Xa23.
Among the six candidates of Xa23, LOC_Os11g37580 is a hypothetical protein with no functional domain predicted based on its amino acid sequences. The gene is highly conserved in rice, poplar, Brachypodium, maize and sorghum, but its biological function is unclear. LOC_Os11g37610 is also a hypothetical protein with a transmembrane region. Orthologous genes exist in rice but not in other organisms. LOC_Os11g37620 is another hypothetical protein with three transmembrane regions. No orthologous gene has been found. LOC_Os11g37630 is a hypothetical protein with high identity to uncharacterized leaf senescence protein-like proteins of rice, conserved in poplar, Arabidopsis, Brachypodium, maize, grapevine and sorghum. LOC_Os11g37630 contains a transmembrane region and two PMR5 domains. Plant proteins with PMR5 have a C-rich sugar binding domain followed by the PC-Esterase (acyl esterase) domain. Plant proteins with PMR5 may play important roles in host–pathogen interactions, regulation of transpiration and stress resistance (Xin et al. 2007; Anantharaman and Aravind 2010). LOC_Os11g37640 is an ADP-ribosylation factor (ARF)-like protein with a signal peptide at N-terminus. ARF proteins are conserved in various organisms. In plants, ARF proteins have been reported to play roles in controlling cell cycle during seed development, intracellular signaling, and membrane trafficking (Matheson et al. 2007; Cevher-Keskin 2013), associated with endocytosis in plant cells (Naramoto et al. 2010) and replication of red clover necrotic mosaic virus, a plant RNA virus (Hyodo et al. 2013). The relationship between ARF proteins and plant disease resistance has been established (Lee et al. 2003b; Lee and Sano 2007; Böhlenius et al. 2010; Nielsen et al. 2012). Rice ARF protein has been demonstrated to be involved in fungal disease response (Lee et al. 2003b). LOC_Os11g37660 is a putative vegetative storage protein, containing nine tandem pentatricopeptide repeats (PPRs). PPR proteins are eukaryote-specific RNA-binding proteins, involved in multiple aspects of RNA metabolism of organellar genes (Nakamura et al. 2012). Recent investigation has shown that the PPR protein PPR8522 is necessary for maize embryogenesis and vegetative development (Sosso et al. 2012). PPR proteins usually act in a gene-specific manner (Nakamura et al. 2012; Härtel et al. 2013). So far, no one has reported that a PPR protein is involved in plant disease resistance.
We have cloned the cognate avr-gene of Xa23 from Xoo strain PXO99 (GenBank: GU732172.1). The avrXa23 encodes a member of transcription activator-like (TAL) effectors (Wang et al. 2013). It has been demonstrated that Xa23-dependent BB resistance was resulted from a classical gene-for-gene interaction between CBB23 and Xoo strains (Wang et al. 2013). Thus, we speculate that Xa23 should be a member of the so-called executor type R genes whose expressions are activated by TAL effectors. Among the cloned dominant BB resistance genes, only Xa10 (Tian et al. 2014) and Xa27 (Gu et al. 2005) belong to the executor type R genes. Notably, recent work revealed that the conserved domains of Xa10 are highly homologous with the hypothetical protein (ABA94457) deduced from LOC_Os11g37620, even if their nucleotide sequences are largely different (Tian et al. 2014). Therefore, LOC_Os11g37620 is most likely the candidate of Xa23. However, LOC_Os11g37620 presents in Nipponbare that lacks the Xa23-mediated BB resistance. Thus, we speculate that the Xa23 might encode a protein different from ABA94457. Accomplishment of molecular cloning of Xa23 would confirm this speculation.
References
Anantharaman V, Aravind L (2010) Novel eukaryotic enzymes modifying cell-surface biopolymers. Biology Direct 5:1. doi:10.1186/1745-6150-5-1
Bhasin H, Bhatia D, Raghuvanshi S, Lore JS, Sahi GK, Kaur B, Vikal Y, Singh K (2012) New PCR-based sequence-tagged site marker for bacterial blight resistance gene Xa38 of rice. Mol Breed 30:607–611
Blair MW, Garris AJ, Iyer AS, Chapman B, Kresovich S, McCouch SR (2003) High resolution genetic mapping and candidate gene identification at the xa5 locus for bacterial blight resistance in rice (Oryza sativa L.). Theor Appl Genet 107:62–73
Böhlenius H, Mørch SM, Godfrey D, Nielsen ME, Thordal-Christensen H (2010) The multivesicular body-localized GTPase ARFA1b/1c is important for callose deposition and ROR2 syntaxin-dependent preinvasive basal defense in barley. Plant Cell 22:3831–3844
Brar DS, Khush GS (1997) Alien introgression in rice. Plant Mol Biol 35:35–47
Cevher-Keskin B (2013) ARF1 and SAR1 GTPases in endomembrane trafficking in plants. Int J Mol Sci 14:18181–18199
Cheema KK, Grewal NK, Vikal Y, Sharma R, Lore JS, Das A, Bhatia D, Mahajan R, Gupta V, Bharaj TS, Singh K (2008) A novel bacterial blight resistance gene from Oryza nivara mapped to 38 kb region on chromosome 4 and transferred to Oryza sativa L. Genet Res 90:397–407
Chen H, Wang S, Zhang Q (2002) A new gene for bacterial blight resistance in rice located on chromosome 12 identified from Minghui 63, and elite restorer line. Phytopathology 92:750–754
Chen S, Liu X, Zeng L, Ouyang D, Yang J, Zhu X (2011) Genetic analysis and molecular mapping of a novel recessive gene xa34(t) for resistance against Xanthomonas oryzae pv. oryzae. Theor Appl Genet 122:1331–1338
Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen JL, Zhang Q, Wang S (2006) Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev 20:1250–1255
Fan YL, Chen XW, Wang CL, Zhu LH, Zhang Q, Zhao KJ (2006) Mapping the rice bacterial blight resistance gene Xa23 with RFLP markers and converting RFLP to STS Marker. Acta Agron Sin 32:931–935
Gu K, Tian D, Yang F, Wu L, Sreekala C, Wang D, Wang GL, Yin Z (2004) High-resolution genetic mapping of Xa27(t), a new bacterial blight resistance gene in rice, Oryza sativa L. Theor Appl Genet 108:800–807
Gu K, Yang B, Tian D, Wu L, Wang D, Sreekala C, Yang F, Chu Z, Wang GL, White FF, Yin Z (2005) R gene expression induced by a type-III effector triggers disease resistance in rice. Nature 435:1122–1125
Guo SB, Zhang DP, Lin XH (2010) Identification and mapping of a novel bacterial blight resistance gene Xa35(t) originated from Oryza minuta. Sci Agric Sin 43:2611–2618
Härtel B, Zehrmann A, Verbitskiy D, van der Merwe JA, Brennicke A, Takenaka M (2013) MEF10 is required for RNA editing at nad2-842 in mitochondria of Arabidopsis thaliana and interacts with MORF8. Plant Mol Biol 81:337–346
Huang B, Xu JY, Hou MS, Ali J, Mou TM (2012) Introgression of bacterial blight resistance genes Xa7, Xa21, Xa22 and Xa23 into hybrid rice restorer lines by molecular marker-assisted selection. Euphytica 187:449–459
Hyodo K, Mine A, Taniguchi T, Kaido M, Mise K, Taniguchi H, Okuno T (2013) ADP ribosylation factor 1 plays an essential role in the replication of a plant RNA virus. J Virol 87:163–176
Iyer AS, McCouch SR (2004) The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol Plant-Microbe Interact 17:1348–1354
Jiang GH, Xia ZH, Zhou YL, Wan J, Li DY, Chen RS, Zhai WX, Zhu LH (2006) Testifying the rice bacterial blight resistance gene xa5 by genetic complementation and further analyzing xa5 (Xa5) in comparison with its homolog TFIIAg1. Mol Genet Genomics 275:354–366
Kauffman HE, Reddy APK, Hsieh SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis Report 57:537–541
Kawahara Y, Bastide M, Hamilton JP, Kanamori H, McCombie WR, Ouyang S, Schwartz DC, Tanaka T, Wu J, Zhou S, Childs KL, Davidson RM, Lin H, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee SS, Kim J, Numa H, Itoh T, Buell CR, Matsumoto T (2013) Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6:4
Khush GS, Angeles ER (1999) A new gene for resistance to race 6 of bacterial blight in rice, Oryza sativa L. Rice Genet Newsl 16:92–93
Korinsak S, Sriprakhon S, Sirithanya P, Jairin J, Korinsak S, Vanavichit A, Toojinda T (2009) Identification of microsatellite markers (SSR) linked to a new bacterial blight resistance gene xa33(t) in rice cultivar ‘Ba7’. Maejo Intern J Sci and Technol 3:235–247
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Kumar PN, Sujatha K, Laha GS, Rao KS, Mishra B, Viraktamath BC, Hari Y, Reddy CS, Balachandran SM, Ram T, Madhav MS, Rani NS, Neeraja CN, Reddy GA, Shaik H, Sundaram RM (2012) Identification and fine-mapping of Xa33, a novel gene for resistance to Xanthomonas oryzae pv. oryzae. Phytopathology 102:222–228
Lee MH, Sano H (2007) Attenuation of the hypersensitive response by an ATPase associated with various cellular activities (AAA) protein through suppression of a small GTPase, ADP ribosylation factor, in tobacco plants. Plant J 51:127–139
Lee KS, Rasabandith S, Angeles ER, Khush GS (2003a) Inheritance of resistance to bacterial blight in 21 cultivars of rice. Phytopathology 93:147–152
Lee WY, Hong JK, Kim CY, Chun HJ, Park HC, Kim JC, Yun DJ, Chung WS, Lee SH, Lee SY, Cho MJ, Lim CO (2003b) Over-expressed rice ADP-ribosylation factor 1 (RARF1) induces pathogenesis-related genes and pathogen resistance in tobacco plants. Physiol Plant 119:573–581
Lin XH, Zhang DP, Xie YF, Gao HP, Zhang Q (1996) Identifying and mapping a new gene for bacterial blight resistance in rice based on RFLP markers. Phytopathology 86:1156–1159
Lin H, Wang R, Qian Q, Yan M, Meng X, Fu Z, Yan C, Jiang B, Su Z, Li J, Wang Y (2009) DWARF27, an iron-containing protein required for the biosynthesis of strigolactones, regulates rice tiller bud outgrowth. Plant Cell 21:1512–1525
Lincoln S, Daly M, Lander E (1993) Constructing genetic maps with Mapmaker/Exp 3.0, vol 3. Whitehead Institute Technical Report, Cambridge
Liu Q, Yuan M, Zhou Y, Li X, Xiao J, Wang S (2011) A paralog of the MtN3/saliva family recessively confers race-specific resistance to Xanthomonas oryzae in rice. Plant Cell Environ 34:1958–1969
Matheson LA, Hanton SL, Rossi M, Latijnhouwers M, Stefano G, Renna L, Brandizzi F (2007) Multiple roles of ADP-ribosylation factor 1 in plant cells include spatially regulated recruitment of coatomer and elements of the Golgi matrix. Plant Physiol 143:1615–1627
McCouch SR, Kochert Q, Yu ZH, Wang ZY, Khush GS, Coffman WR, Tanksley SD (1988) Molecular mapping of rice chromosome. Theor Appl Genet 26:815–829
Miao LL, Wang CL, Zheng CK, Che JY, Gao Y, Wen YC, Li GQ, Zhao KJ (2010) Molecular mapping of a new gene for resistance to rice bacterial blight. Sci Agric Sin 43:3051–3058
Nakamura T, Yagi Y, Kobayashi K (2012) Mechanistic insight into pentatricopeptide repeat proteins as sequence-specific RNA-binding proteins for organellar RNAs in plants. Plant Cell Physiol 53:1171–1179
Naramoto S, Kleine-Vehn J, Robert S, Fujimoto M, Dainobu T, Paciorek T, Ueda T, Nakano A, Van Montagu MC, Fukuda H, Friml J (2010) ADP-ribosylation factor machinery mediates endocytosis in plant cells. Proc Natl Acad Sci USA 107:21890–21895
Nielsen ME, Feechan A, Böhlenius H, Ueda T, Thordal-Christensen H (2012) Arabidopsis ARF-GTP exchange factor, GNOM, mediates transport required for innate immunity and focal accumulation of syntaxin PEN1. Proc Natl Acad Sci USA 109:11443–11448
Niño-Liu DO, Ronald PC, Bogdanove AJ (2006) Xanthomonas oryzae pathovars: model pathogens of a model crop. Mol Plant Pathol 7:303–324
Sakai H, Lee SS, Tanaka T, Numa H, Kim J, Kawahara Y, Wakimoto H, Yang CC, Iwamoto M, Abe T, Yamada Y, Muto A, Inokuchi H, Ikemura T, Matsumoto T, Sasaki T, Itoh T (2013) Rice annotation project database (RAP-DB): an integrative and interactive database for rice genomics. Plant Cell Physiol 54:e6. doi:10.1093/pcp/pcs183
Song WY, Wang GL, Chen LL, Kim HS, Pi LY, Holsten T, Gardner J, Wang B, Zhai WX, Zhu LH, Fauquet C, Ronald P (1995) A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 270:1804–1806
Sosso D, Canut M, Gendrot G, Dedieu A, Chambrier P, Barkan A, Consonni G, Rogowsky PM (2012) PPR8522 encodes a chloroplast-targeted pentatricopeptide repeat protein necessary for maize embryogenesis and vegetative development. J Exp Bot 63:5843–5857
Suh JP, Jeung JU, Noh TH, Cho YC, Park SH, Park HS, Shin MS, Kim CK, Jena KK (2013) Development of breeding lines with three pyramided resistance genes that confer broad-spectrum bacterial blight resistance and their molecular analysis in rice. Rice 6:5. doi:10.1186/1939-8433-6-5
Sun X, Cao Y, Yang Z, Xu C, Li X, Wang S, Zhang Q (2004) Xa26, a gene conferring resistance to Xanthomonas oryzae pv oryzae in rice, encodes an LRR receptor kinase-like protein. Plant J 37:517–527
Tan GX, Ren X, Weng QM, Shi ZY, Zhu LL, He GC (2004) Mapping of a new resistance gene to bacterial blight in rice line introgressed from Oryza officinalis. Acta Genet Sin 31:724–729
Tian D, Wang J, Zeng X, Gu K, Qiu C, Yang X, Zhou Z, Goh M, Luo Y, Murata-Hori M, White FF, Yin Z (2014) The rice TAL effector-dependent resistance protein XA10 triggers cell death and calcium depletion in the endoplasmic reticulum. Plant Cell. http://www.plantcell.org/cgi/doi/10.1105/tpc.113.119255
Wang CL, Qi HX, Pan HJ, Li JB, Fan YL, Zhang Q, Zhao KJ (2005) EST-markers flanking the rice bacterial blight resistance gene Xa23 and their application in marker-assisted selection. Sci Agric Sin 38:1996–2001
Wang CL, Chen LT, Zeng CZ, Zhang QY, Liu PQ, Liu YG, Fan YL, Zhang Q, Zhao KJ (2006) Chromosome walking for fine mapping of Xa23 gene locus by using genomic libraries. Chin J Rice Sci 20:355–360
Wang CT, Wen GS, Lin XH, Liu XQ, Zhang DP (2009) Identification and fine mapping of the new bacterial blight resistance gene, Xa31(t), in rice. Eur J Plant Pathol 123:235–240
Wang CL, Qin TF, Yu H, Zhang XP, Che JY, Gao Y, Zheng CK, Yang B, Zhao KJ (2013) The broad bacterial blight resistance of rice line CBB23 is triggered by a novel TAL effector of Xanthomonas oryzae pv. oryzae. Mol Plant. doi:10.1111/mpp.12092
Xiang Y, Cao Y, Xu C, Li X, Wang S (2006) Xa3, conferring resistance for rice bacterial blight and encoding a receptor kinase-like protein, is the same as Xa26. Theor Appl Genet 113:1347–1355
Xin Z, Mandaokar A, Chen J, Last RL, Browse J (2007) Arabidopsis ESK1 encodes a novel regulator of freezing tolerance. Plant J 49:786–799
Yoshimura S, Yamanouchi U, Katayose Y, Toki S, Wang ZX, Kono I, Kurata N, Yano M, Iwata N, Sasaki T (1998) Expression of Xa1, a bacterial blight-resistance gene in rice, is induced by bacterial inoculation. Proc Natl Acad Sci USA 95:1663–1668
Zhang Q, Lin SC, Zhao BY, Wang CL, Yang WC, Zhou YL, Li DY, Chen CB, Zhu LH (1998) Identifying of a new gene for resistance to bacterial blight from O. rufipogon. Rice Genet Newsl 15:138–142
Zhang Q, Wang CL, Zhao KJ, Yang WC, Qiao F, Zhou YL, Jiang QX, Liu GC (2002) Development of near- isogenic line CBB23 with a new resistance gene to bacterial blight in rice and its application. Chin J Rice Sci 16:206–210
Zheng CK, Wang CL, Yu YJ, Liang YT, Zhao KJ (2009) Identification and molecular mapping of Xa32(t), a novel resistance gene for bacterial blight (Xanthomonas oryzae pv. oryzae) in rice. Acta Agron Sin 35:1173–1180
Zhou YL, Xu JL, Zhou SC, Yu J, Xie XW, Xu MR, Sun Y, Zhu LH, Fu BY, Gao YM, Li ZK (2009) Pyramiding Xa23 and Rxo1 for resistance to two bacterial diseases into an elite indica rice variety using molecular approaches. Mol Breed 23:279–287
Acknowledgments
This work was supported by the National High-Technology Research Program (The “863” Program: Grant No. 2006AA10Z106) of Ministry of Science and Technology of China and National Natural Science Foundation of China (Grant No. 31171812).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann.
C. Wang, Y. Fan and C. Zheng contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wang, C., Fan, Y., Zheng, C. et al. High-resolution genetic mapping of rice bacterial blight resistance gene Xa23 . Mol Genet Genomics 289, 745–753 (2014). https://doi.org/10.1007/s00438-014-0848-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-014-0848-y