Abstract
We describe three new Henneguya spp. (Myxobolidae) found parasitizing two species of cichlid fish from the Amazon basin, Brazil: H. tucunarei n. sp. from gill filaments of Cichla monoculus and H. tapajoensis n. sp. from gill filaments of Cichla pinima, both from the Tapajós River, Pará State and H. jariensis n. sp. in the fins of Cichla monoculus from the Jari River, Amapá State. We based descriptions on myxospore morphology and small subunit ribosomal DNA sequences, and used a phylogenetic analysis to compare the new Henneguya species with known relatives. Spores of the three species had similar morphology and morphometrics, but differed molecularly 5–7.5%, and were no more than 94% similar to any other sequence in GenBank. Together with having different hosts, these data supported the diagnosis of the parasites as distinct, novel species. Maximum likelihood and Bayesian analyses showed that H. tucunarei n. sp., H. tapajoensis n. sp., and H. jariensis n. sp. plus Henneguya paraensis (which parasitizes Cichla temensis) formed a well-supported sub-clade of Henneguya parasites of cichlids from the Amazon basin, in a lineage sister to those in characiforms hosts. Our analysis was consistent with previous studies that suggest that aquatic environment and vertebrate host group are the strongest correlates with phylogenetic signals in the Myxobolidae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Myxozoans are a species-rich group of endoparasitic Cnidaria that infect a wide range of vertebrate and invertebrate hosts worldwide (Lom and Dyková 2006; Fiala et al. 2015). They are major parasites in both wild and farmed fish, with more than 2400 nominal species (Zhang 2011), distributed in 63 genera, and classified mainly by spore morphology (Fiala et al. 2015). Although most species do not cause disease, some are highly pathogenic and give rise to serious ecological and economic impacts on their host fish populations (Kent et al. 2001; Feist and Longshaw 2006; Feist 2008).
Knowledge about the diversity of myxozoan parasites in South America is limited, especially in the Amazon basin, as this region has the world’s richest and most diversified ichthyofauna (Reis et al. 2016), all of which are potential hosts of myxozoans.
The genus Henneguya Thélohan, 1892, encompasses > 200 species and are most commonly found in fresh water fishes (Eiras 2002; Eiras and Adriano 2012). Fifty-five Henneguya species have been described in Brazil, of which 16 were reported from Amazon basin fishes (Eiras 2002; Eiras and Adriano 2012; Rocha et al. 2014; Videira et al. 2015; Mathews et al. 2016; Velasco et al. 2016). Only three species have been reported from cichlids in the Amazon: H. amazonica Rocha, Matos, and Azevedo 1992, from gill lamellae of Crenicichla lepidota; H. aequidens Videira et al. 2015, from gill filaments of Aequidens plagiozonatus; and H. paraensis Velasco et al. 2016, from gill filaments of Cichla temensis. Given there are at least a hundred additional cichlid species known from the Amazon basin (Froese and Pauly 2017), we hypothesized that there are many undescribed Henneguya species present in this system.
Accordingly, in this study, we describe three new freshwater Henneguya species parasitizing Cichla monoculus Agassiz 1831, and Cichla pinima Kullander and Ferreira 2006, from different areas in the Amazon basin, Brazil. Cichla monoculus and C. pinima are popularly known as “tucunaré” in Portuguese or “peacock bass” in English. These fish are endemic to the Amazon basin and do not perform reproductive migration (Kullander and Ferreira 2006). They are important as subsistence, recreational, and commercial food resource (Willis et al. 2007; Freitas and Campos 2014). We used morphological analysis in combination with ssrDNA sequence data to describe the new parasites, and showed their phylogenetic position within the large Myxobolus/Henneguya clade (Family Myxobolidae).
Material and methods
Sampling and morphological analysis
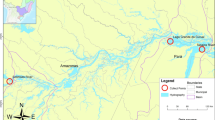
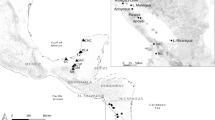
Fish (N = 57) were collected using seine net and fishing hook from two localities (Fig. 1): C. monoculus (N = 29; average length 33.3 cm, range 26–51 cm) and C. pinima (N = 22; average length 32 cm, range 27.5–52 cm), were collected from the Tapajós River, municipality of Santarém, Pará State, in October 2014 and March 2015 (GPS coordinates given in Results); C. monoculus (N = 6; average length 28 cm, range 24–30 cm) were collected from the Jari River, Municipality of Vitória do Jari, Amapá State, in October 2015. The sampling and access to genetic heritage was authorized by the Brazilian Ministry of the Environment (SISBIO n° 44268-4 and SisGen A33CB83).
Fish were killed by overdose of benzocaine solution (70 mg L−1), measured, and necropsied. The research methodology was approved by the Ethics Committee on Animal Use, University of Campinas (CEUA/UNICAMP n° 3846-1). Under a stereomicroscope, the organs were inspected for the presence of myxozoan cysts. Individual cysts were excised and fixed in 10% neutral-buffered formalin for myxospore measurements (Lom and Arthur 1989) and 100% ethanol for ssrDNA sequencing. Myxospores were photographed using differential interference contrast and measured using a Carl Zeiss Axio Imager A2 light microscope equipped with an Axio Cam, and AxioVision AxioVs 40V4.8.2 software, following the guidelines of Lom and Arthur (1989), except that we use the term “polar tubule” instead of “polar filament” (Ben-David et al. 2016). Type myxospores were air-dried onto glass slides, stained with Giemsa, and deposited in the Museum of Zoology “Adão José Cardoso” University of Campinas, São Paulo State, Brazil.
DNA extraction, amplification, and sequencing
Cysts preserved in 100% ethanol were mechanically ruptured on a microscopic slide under a cover slip, and the content washed into a 2-mL tube using ATL Buffer from the DNeasy® Blood & Tissue Kit (animal tissue protocol) (QIAGEN Inc., California, USA). Total DNA was extracted following the manufacturer’s instructions.
Partial small-subunit ribosomal DNA (ssrDNA) was amplified using a two-round PCR. Initial amplification was performed using primers ERIB1 (ACCTGGTTGATCCTGCCAG; Barta et al. 1997) and ERIB10 (CTTCCGCAGGTTCACCTACGG; Barta et al. 1997), followed by a semi-nested second round using primer set ERIB1 and ACT1R (AATTTCACCTCTCGCTGCCA; Hallett and Diamant 2001), and either Myxigen4f (GTGCCTTGAATAAATCAGAG; Diamant et al. 2004) or a new primer that we designed MyxidF4 (GTTAAAACGCTCGTAGTTGG) with ERIB10. The new primer was designed to align with a conserved region from an alignment of myxozoan sequences, downloaded from GenBank.
PCR was performed in 20 μl reaction volumes that comprised of the following: 1 μl DNA (10–50 ng/μl), 1.25 U GoTaq Flexi polymerase (Promega, San Luis Obispo, California, USA), 0.2 μl mM each dNTP, 0.50 μl each primer (10 pmol), 4 μl 5× GoTaq Flexi clear buffer, 2.4 μl MgCl2 (1.5 mM), 0.50 μl BSA, 0.4 μl Rediload dye (Invitrogen, Carlsbad, California, USA), and ultrapure water. PCR cycling was performed on a PTC-200 Thermocycler (MJ Research Inc., Watertown, Massachusetts, USA) using a block preheated to 95 °C, and cycle conditions of: 95 °C for 2 min, followed by 35 cycles of 94 °C for 30 s, 58 °C for 30 s, and 72 °C for 90 s (with first-round primers or 60 s when using second-round primers), following by a terminal extension at 72 °C for 5 min. Negative and positive controls were included in each PCR run. Amplicons were electrophoresed in a 1.5% agarose gel in Tris-EDTA buffer (0.045 M Tris, 0.001 M EDTA, pH 8.0) stained with SYBRsafe (Invitrogen By Life Technologies, Maryland, USA) alongside a 1 kb Plus DNA Ladder (Invitrogen By Life Technologies, Maryland, USA) at 90 V for 40 min, and then analyzed on a Compact Digimage System transilluminator (MajorScience, California, USA). Amplicons were purified using the QIAquick PCR Purification Kit (QIAGEN Inc., California, USA) according to the manufacturer’s instructions. Amplicons were sequenced in both directions using a BigDye 102 Terminator v3.1 Cycle Sequencing kit (Applied Biosystems, California, USA) in an ABI 3730 DNA 103 Analyzer (Applied Biosystems, California, USA) at Life Sciences Core Facility (LaCTAD) from the University of Campinas, Brazil, for H. tucunarei n. sp., and at the Center for Genome Research and Biocomputing sequencing facility at Oregon State University, USA, for H. tapajoensis n. sp. and H. jariensis n. sp.
Sequencing assembly and phylogenetic analysis
Sequences were imported into BioEdit 7.1.3.0 (Hall 2011) and aligned manually to produce consensus sequences. Chromatograms were checked to access the quality of the sequences and presence of any polymorphisms. A nucleotide BLASTn (Altschul et al. 1997) search was conducted to confirm amplification of myxozoan DNA.
Phylogenetic analyses were performed on an alignment of 34 ssrRNA sequences from the most closely related freshwater myxozoans, as determined by the BLASTn search, and included all Myxobolus or Henneguya sequences from South America available from the NCBI database (accession numbers are indicated in the phylogenetic tree). Sequences were aligned with ClustalW (Thompson et al. 1994) using default parameters. Gaps were treated as missing data. Phylogenetic trees were calculated using maximum likelihood (ML) and Bayesian (BI) analysis. The more distant, “marine lineage” myxozoans Ceratomyxa auerbachi (EU616734) and Ceratomyxa diamanti (FJ204246) were used as outgroup taxa. A second ML analysis including myxobolid parasites of African cichlids was performed using the same parameters described above (supplementary material—Fig. S1).
The optimum evolutionary model for the dataset was obtained by the Akaike Information Criterion using jModelTest 0.1.1 (Posada 2008). We performed ML analysis using PhyML version 3.0 (Guindon and Gascuel 2003) implemented via the web server (http://www.atgc-montpellier.fr/phyml/) (Guindon et al. 2010). Bootstrap support was calculated using 1000 replicates. Bayesian phylogenetic relationships were inferred using Mr. Bayes v.3.0 (Ronquist and Huelsenbeck 2003), with posterior probabilities estimated from 1,000,000 generations via two independent runs of four simultaneous Markov Chain Monte Carlo algorithms with every 100th tree saved with the burn-in set to 10% (100,000 generations). We tested results with Tracer v1.4.1 (Rambaut and Drummond 2007) (http://beast.bio.ed.ac.uk/Tracer), and visualized trees using Figtree 1.3.1 (Rambaut 2008) and edited them for publication using Adobe Photoshop (Adobe Systems Inc., California, USA). We performed a genetic distance analysis of Henneguya parasites from Cichla using the p-distance model matrix in MEGA 7.0 (Kumar et al. 2016), with default parameters; gaps and missing data were deleted.
Results
We observed plasmodia that contained myxospores with morphological characteristics of the genus Henneguya in 3.4% (1/29) C. monoculus and 45% (10/22) C. pinima caught in the Tapajós River and in 17% (1/6) C. monoculus collected in the Jari River.
Taxonomic summary and morphological data
Phylum: Cnidaria Verrill, 1865
Unranked Sub-phylum Myxozoa Grassé (1970)
Class: Myxosporea Bütschli, 1881
Order: Bivalvulida Shulman, 1959
Family: Myxobolidae Thélohan, 1892
Genera: Henneguya Thélohan, 189
Henneguya tucunarei n. sp. (Figs. 2a, 3a)
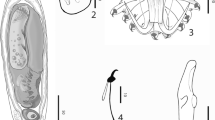
Nomarski differential interference contrast images of wet-mount myxospores of three novel Henneguya species. A: Henneguya tucunarei n. sp. from gill filaments of Cichla monoculus; B: Henneguya tapajoensis n. sp. from gill of Cichla pinima; C: Henneguya jariensis n. sp. from fins of Cichla monoculus from Jari River. Bars = 10 μm
Type host: Cichla monoculus Agassiz 1831, Cichliformes, Cichlidae
Prevalence: 1 of 29 (3.4%)
Type locality: Tapajós River, near the city of Santarém, Pará State, Brazil Coordinates: 02° 20′ 03″ S 54° 52′ 33″ W
Site of infection: Gill filaments
Type material: A glass slide with stained spores (syntype) was deposited in the collection of the Museum of Zoology “Adão José Cardoso” of University of Campinas (UNICAMP), São Paulo, Brazil (accession number ZUEC MYX71). The ssrDNA sequence was deposited in GenBank with accession number KY751401.
Etymology: The specific name refers to the Portuguese common name of the host, “tucunaré.”
Description: Whitish, oval-shaped plasmodia, diameter ≤ 300 μm, contained mature and immature myxospores. Mature myxospores were ellipsoid in valvular view and biconvex in sutural view, average total spore length of 43.8 μm (range 36.1–49.6 μm, N = 14), average body length 14 μm (range 12.1–15.7 μm, N = 29), average width 6.1 μm (range 4.9–7.8 μm, N = 26). Each valve cell had a caudal projection, average length 28.1 μm (range 19.6–35.6 μm, N = 12). Two polar capsules, pyriform, at the anterior pole of the spore, equal in size, average length 3.4 μm (range 2.5–4.6 μm, N = 23), average width 2 μm (range 1.3–2.6 μm, N = 22), with 3–4 turns of the polar tubule (Table 1).
Henneguya tapajoensis n. sp. (Figs. 2b, 3b).
Type host: Cichla pinima Kullander and Ferreira 2006, Cichliformes, Cichlidae
Prevalence: 10 of 22 (45.4%)
Type locality: Tapajós River, near the city of Santarém, Pará State, Brazil Coordinates: 02° 20′ 03″ S 54° 52′ 33″ W
Site of infection: Gill filaments
Type material: A glass slide with stained spores (syntype) was deposited in the collection of the Museum of Zoology “Adão José Cardoso” of the University of Campinas (UNICAMP), São Paulo, Brazil, under accession number ZUEC MYX 72). The ssrDNA sequence was deposited in GenBank with accession number KY751402.
Etymology: The specific name refers to the river (Tapajós) where the parasite was found.
Description: Several small and slightly elongate plasmodia (diameter ≤200 μm) occurred within a single gill filament. Mature myxospores, average total length 54.6 μm (range 47.2–62.2 μm, N = 23), with an ellipsoidal spore body, average length 16.4 μm (range 14.5–19.1 μm, N = 30), average width 7 μm (range 5.7–9.3 μm, N = 24), and average thickness 5 μm (range 4.9–5.0 μm, N = 3). Valve cells extended to equal-sized caudal processes, average length 39 μm (range 31.7–46.5 μm, N = 22). Two polar capsules equal in size, pyriform, average length 4.2 μm (range 2.9–5 μm, N = 20), average width 2.1 μm (range 1.5–2.8 μm, N = 20) with 4–5 turns of the polar tubule.
Henneguya jariensis n. sp. (Figs. 2c, 3c).
Type host: Cichla monoculus Agassiz 1831, Cichliformes, Cichlidae
Prevalence: 1 of 6 (17%)
Type locality: Jari River, municipality of Vitória do Jari, (District of Jarilândia), Amapá State, Brazil.
Coordinates: 01° 07′ 29″ S 51° 59′ 42″ W
Site of infection: Soft tissue of fins.
Type material: A glass slide with stained spores (syntype) was deposited in the collection of the Museum of Zoology “Adão José Cardoso” of the University of Campinas (UNICAMP), São Paulo, Brazil (accession number ZUEC MYX73). The ssrDNA sequence was deposited in GenBank with accession number KY751403.
Etymology: The specific name is based on the river (Jari) where the parasite was found.
Description: Small, spherical, whitish plasmodia (diameter ≤ 500 μm) in soft tissue of the fins of C. monoculus. Mature myxospores had thin, smooth walls with symmetric valves, average total length 46.7 μm (range 43.9–49.2 μm, N = 15), average spore body length 13.4 μm (range 11.9–14.6 μm, N = 21), average width 6.5 μm (range 5.5–7.3 μm, N = 20). Valve cells extended to equal length caudal processes, average length 33.2 μm (range 30.2–37 μm, N = 16). The polar capsules, elongate ovoid, equal sized, average length 4 μm (range 3.4–4.3 μm, N = 21), average width 2 (range 1.7–2.4 μm, N = 21) containing polar tubules with 4 turns.
Molecular and phylogenetic analysis
We obtained ssrDNA sequences from the three novel Henneguya species: 1478 bp from H. tucunarei n. sp. (GenBank accession number KY751401), 1932 bp from H. tapajoensis n. sp. (KY751402), and 1951 bp from H. jariensis n. sp. (KY751403). The sequences were not more than 94% similar to any other sequence in GenBank, and were all most similar to Henneguya paraensis (from Cichla temensis; Velasco et al. 2016), with similarities of 94, 92, and 94% respectively. A similarity matrix of the ssrDNA gene sequences comparing only Henneguya sequences from Cichla spp. showed that the smallest genetic distance was 5% between H. tucunarei and H. paraensis, and the largest was 7.5% between H. tapajoensis and H. paraensis (Table 2).
We identified GTR + G as the optimum evolutionary model based on jModelTest. ML and BI phylogenetic trees showed two main clades, A and B (Fig. 4). Clade A had several sub-clades formed by parasites of Characiformes, Siluriformes, Cichliformes, and Perciformes hosts, from different continents. Clade B was formed by Myxobolus spp. from Characiformes, and included Myxobolus cordeiroi Adriano, Arana, Alves, Ceccarelli, Henrique-Silva and Maia 2009, and Thelohanellus marginatus Rocha et al. 2014, which are parasites of Siluriformes, and all them are parasites of exclusively South American fish. We observed that the three novel species, H. tucunarei n. sp., H. tapajoensis n. sp., and H. jariensis n. sp., grouped with H. paraensis in a well-supported sub-clade in both methods; this sub-clade represented a lineage of Henneguya species parasites of Cichla spp. that was sister to a lineage formed by Henneguya/Myxobolus parasites of South American characiforms.
Consensus ML phylogenetic tree using ssrDNA sequences of selected Myxobolus and Henneguya species. GenBank accession numbers given in parenthesis. Nodal supports are indicated for ML with a bootstrap of 1000 replicates, and BI with posterior probabilities, respectively. Values for weakly supported nodes (< 50) are not shown. Scale is 0.25
Taxonomic affinities
The three new Henneguya species were compared with congeners from the Amazon basin, and from freshwater fish from other regions, based on morphological and molecular data when available (Eiras 2002; Eiras and Adriano 2012; Carriero et al. 2013; Moreira et al. 2014a, b; Naldoni et al. 2014; Rocha et al. 2014; Videira et al. 2015; Mathews et al. 2016; Velasco et al. 2016).
-
H. tucunarei n. sp.: considering only Amazonian species, H. tucunarei myxospores resemble morphologically those of H. paraensis, which also parasites a fish of the genus Cichla. Morphometrically, H. tucunarei myxospores are distinct from H. paraenis, with longer polar capsules and greater spore body width, shorter spore body, and more turns of the polar tubules (Table 1). ssrDNA sequences were 5% different (Table 2). In comparison with the two other novel species described herein, H. tucunarei n. sp. is similar morphologically, but differs morphometrically from H. tapajoensis n. sp. by having shorter spore total length and caudal appendages, and fewer polar tubule coils (Table 1); and a genetic difference of 6.9% (Table 2). Compared with H. jariensis n. sp., H. tucunarei n. sp. has only subtle morphometric differences: slighter shorter spore total length and appendages (Table 1), but is distinct molecularly, with 5.3% difference in ssrDNA. In comparison with Henneguya spp. from different geographic regions, H. tucunarei n. sp. resembles morphologically the spores of Henneguya salmonicola Ward 1919, a parasite of salmonid fishes in North America, but has a longer spore body (14.1 versus 12.4) and is narrower (6.1 versus 7.9 μm), and molecularly differs by at least 25% in ssrDNA. The ssrDNA sequence of H. tucunarei n. sp. was at maximum 94% similar to any myxosporean available in GenBank.
-
H. tapajoensis n. sp.: in comparison with other Henneguya described from the Amazon, H. tapajoensis n. sp. resembled Henneguya amazonica Rocha, Matos, and Azevedo 1992, which also parasitizes gills of cichlid fish. However, the length of spore body and caudal appendences of H. tapajoensis n. sp. were shorter than H. amazonica, and its polar capsules were larger and wider (Table 2). In comparison with the other novel species described herein, H. tapajoensis n. sp. is larger in all dimensions (Table 2) and is 7.1% different genetically (Table 1). Regarding Henneguya species from other geographical areas, H. tapajoensis n. sp. resembles Henneguya acuta Bond 1939, a parasite of the esocid Esox masquinongy in North America. However, myxospores and polar capsules of H. acuta are longer, with more polar tubule coils that H. tapajoensis. There are no molecular data for H. acuta to inform the comparison. The ssrDNA sequence of H. tapajoensis n. sp. had a maximum similarity of 92% with other myxosporean available in GenBank.
-
H. jariensis n. sp.: compared to other Amazonian species, morphologically, H. jariensis resembles Henneguya astyanax Vital, Corral, Matos, and Azevedo 2003, but is morphometrically slightly shorter in spore body and polar capsules. Whereas H. astyanax parasitizes the gill filaments of a characid fish, H. jariensis n. sp. infects fins of a cichlid. The new species resembles the spores of Henneguya garavelli Martins and Onaka 2006, a parasite of Cyphocharax nagelii in Brazil. However, myxospores of H. jariensis n. sp. are wider and have smaller polar capsules with more turns of the polar tubule. There are no molecular data on H. astyanax and H. garavelli to perform the genetic comparisons. The ssrDNA sequence of H. jariensis n. sp. had a maximum similarity of 94% with other myxosporeans available in GenBank.
Discussion
Cichlids are a diverse, ecologically and economically important fish group in the Amazon basin (Batista and Petrere 2003; Dos Santos et al. 2012). Characterization of their parasite fauna is central to better informing management decisions and risk assessments of their populations, particularly as their use in aquaculture expands. To date, only two myxosporean parasites were known to infect Cichla species: H. paraensis, a parasite of the gills of C. temensis (Velasco et al. 2016) and Ceratomyxa brasiliensis Zatti et al. 2017, from gallbladder of C. monoculus (Zatti et al. 2017). Herein, we describe three additional Henneguya species, from Amazonian Cichla spp.: Henneguya tucunarei n. sp. and H. jariensis n. sp. from C. monoculus, and H. tapajoensis n. sp. from C. pinima.
The three novel Henneguya species were typical of the genus in both myxospore morphology and in the histozoic nature of cyst development in the fish host. As there is morphological similarity between the many members of family Myxobolidae (primarily Myxobolus and Henneguya species), which have small, simple myxospores, it is essential that species descriptions include additional characters, such as host identity, tissue tropism, and genetic sequence data (Lom and Arthur 1989; Molnár et al. 2006). Herein, we combined multiple characters to describe three new Henneguya species from closely related Amazonian cichlid fish. These descriptions form an important part of documenting myxozoan diversity in this region and reinforce the importance of molecular analysis to better understand the taxonomy of the Myxozoa. The new species were morphometrically and genetically distinct from each other, and from previously described Henneguya species (Table 1).
Recently, Abdel-Gaber et al. (2017) re-described five species of myxozoan from Myxobolidae (genus Myxobolus and Triangula), infecting the Nile tilapia Oreochromis niloticus (Perciformes: Cichlidae) collected from River Nile, Africa. The descriptions were based on morphometry and molecular sequencing of the ssrDNA. We attempted to use those sequence data in our phylogenetic analysis to test the hypothesis that myxobolids from Africa and South America are closely related based on their fish hosts. Unfortunately, attempts to perform phylogenetic analyses that included those species resulted in a tree where the African species appeared in a very long branch (see supplementary material—Fig. S1), which suggest that either there is no close genetic relationship between South American and African myxobolid parasites of cichlid fish or there is some inconsistency in those African sequences.
Within Henneguya from other regions, including South American species, our phylogenetic analysis gave insights into the evolutionary context of the three novel species, to reveal correlations with host and geographic locality. The phylogenetic tree (Fig. 4) had two primary branches: the larger clade, A, was comprised of multiple sub-clades of Henneguya/Myxobolus species grouping by vertebrate host fish family from different continents, and the smaller clade B, consisted of exclusively South American fish belonging to four families of the order Characiformes. Within clade A, the three novel Henneguya species, H. tucunarei n. sp., H. tapajoensis n. sp., and H. jariensis n. sp., with H. paraensis, formed a well-supported group of parasites of Amazonian freshwater Cicha spp., which was closely related to Henneguya and Myxobolus parasites of South American characiforms (families Anostomidae and Serrasalmidae). This clustering of parasite species with Cichla spp. is in concordance with previous studies that show vertebrate host affinity is an important phylogenetic signal within the Myxobolidae (Fiala 2006; Ferguson et al. 2008; Carriero et al. 2013; Rocha et al. 2014). Fine-scale structure within the group of Cichla parasites is resolved by tissue tropism: either fin (H. jariensis n. sp.) or gill (H. tucunarei n. sp. and H. tapajoensis n. sp.).
Although species of the genus Cichla are reported to be sedentary (Hoeinghaus et al. 2003), their distribution within the Amazon basin can be relatively restricted, in the case of both Cichla ocellaris Bloch and Schneider 1801, and Cichla intermedia Machado-Allison 1971, to wide-ranging C. monoculus, found throughout the basin (Kullander and Ferreira 2006; Willis et al. 2007). C. pinima distribution is restricted to the lower Tapajós River, lower Xingu River, and lower Tocantins River (Kullander and Ferreira 2006). We hypothesize that host behavior drives both parasite endemism and radiation within the basin. We found myxosporeans in hosts with both limited (C. pinima) and wide-ranging (C. monoculus) behavior, with the highest prevalence of infection (45%) in C. pinima, the host with the narrowest distribution; and the corollary—two different species of Henneguya at low infection prevalences (only single, infected fish) in the widest ranging host, C. monoculus, from localities ~500 km apart. Our results show that in the lower Tapajós region, where C. pinima and C. monoculus are sympatric, the prevalence of H. tapajoensis n. sp. was 45% in the first fish host, but unrecorded in the second. This fine-scale host specificity is a known characteristic of Henneguya species (Molnár 1998; Molnár et al. 1998). However, further study and molecular characterization of the myxobolid fauna that infects Cichla spp. will improve our understanding of the species diversity, host specificity, and evolutionary history of myxozoan parasites within the Amazon basin.
References
Abdel-Gaber R, Abdel-Ghaffar F, Maher S, El-Mallah AM, Al Quraishy S, Mehlhorn H (2017) Morphological re-description and phylogenetic relationship of five myxosporean species of the family Myxobolidae infecting Nile tilapia. Dis Aquat Org 124(3):201–214. https://doi.org/10.3354/dao03118
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLASTn and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3302. https://doi.org/10.1093/nar/25.17.3389
Azevedo C, Matos E (1995) Henneguya adherens n. Sp. (Myxozoa, Myxosporea), parasite of the Amazonian fish, Acestrorhynchus falcatus. J Eukaryot Microbiol 42(5):515–518
Azevedo C, Matos E (1996) Henneguya malabarica sp. nov. (Myxozoa, Myxobolidae) in the Amazonian fish Hoplias malabaricus. Parasitol Res 82(3):222–224
Azevedo C, Matos E (2002) Fine structure of the myxosporean, Henneguya curimata n. Sp., parasite of the Amazonian fish, Curimata inormata (Teleostei, Curimatidae). J Eukaryot Microbiol 49(3):197–200
Azevedo C, Corral L, Matos E (1997) Light and ultrastructural data on Henneguya testicularis n. Sp. (Myxozoa, Myxobolidae), a parasite from the testis of the Amazonian fish Moenkhausia oligolepis. Syst Parasitol 37(2):111–114
Azevedo C, Casal G, Matos P, Matos E (2008) A new species of Myxozoa, Henneguya rondoni n. Sp. (Myxozoa), from the peripheral nervous system of the Amazonian fish, Gymnorhamphichthys rondoni (Teleostei). J Eukaryot Microbiol 55(3):229–234. https://doi.org/10.1111/j.1550-7408.2008.00317.x
Azevedo C, Casal G, Matos P, Alves A, Matos E (2011) Henneguya torpedo sp. nov. (Myxozoa), a parasite from the nervous system of the Amazonian teleost Brachyhypopomus pinnicaudatus (Hypopomidae). Dis Aquat Org 93(3):235–342. https://doi.org/10.3354/dao02292
Barta JR, Martin DS, Liberator PA, Dashkevicz M, Anderson JW, Feighner SD, Elbrecht A, Perkins-Barrow A, Jenkins MC, Danforth HD, Ruff MD, Profous-Juchelka H (1997) Phylogenetic relationships among eight Eimeria species infecting domestic fowl inferred using complete small subunit ribosomal DNA sequences. J Parasitol 83(2):262–271. https://doi.org/10.2307/3284453
Batista VS, Petrere M (2003) Characterization of the commercial fish production landed at Manaus, Amazonas State, Brazil. Acta Amaz 33(1):53–66. https://doi.org/10.1590/1809-4392200331066
Ben-David J, Atkinson SD, Pollak Y, Yossifon G, Shavit U, Bartholomew JL, Lotan T (2016) Myxozoan polar tubules display structural and functional variation. Parasit Vectors 9(1):549. https://doi.org/10.1186/s13071-016-1819-4
Carriero MM, Adriano EA, Silva MRM, Ceccarelli PS, Maia AAM (2013) Molecular phylogeny of the Myxobolus and Henneguya genera with several new South American species. PLoS One 8(9):e73713. https://doi.org/10.1371/journal.pone.0073713
Casal G, Matos E, Azevedo C (2003) Light and electron microscopic study of the myxosporean, Henneguya friderici n. Sp. from the Amazonian teleostean fish, Leporinus friderici. Parasitology 126:313–319
Dos Santos CHS, De Sousa CFS, Paula-Silva MN, Val AL, Almeida-Val VMF (2012) Genetic diversity in Cichla monoculus (Spix & Agassiz, 1931) populations: implications for management and conservation. Am J Environ Sci 8:35–41
Diamant A, Whipps CM, Kent ML (2004) A new species of Sphaeromyxa (Myxosporea: Sphaeromyxina: Sphaeromyxidae) in devil firefish, Pterois miles (Scorpaenidae), from the northern Red Sea: morphology, ultrastructure, and phylogeny. J Parasitol 90(6):1434–1442. https://doi.org/10.1645/GE-336R
Eiras J (2002) Synopsis of the species of the genus Henneguya Thélohan, 1892 (Myxozoa: Myxosporea: Myxobolidae). Syst Parasitol 52(1):43–54. https://doi.org/10.1023/A:1015016312195
Eiras J, Adriano EA (2012) A checklist of new species of Henneguya Thélohan, 1892 (Myxozoa: Myxosporea, Myxobolidae) described between 2002 and 2012. Syst Parasitol 83(2):95–104. https://doi.org/10.1007/s11230-012-9374-7
Eiras JC, Malta JC, Varela A, Pavanelli GC (2004) Henneguya schizodon n. Sp. (Myxozoa, Myxobolidae), a parasite of the Amazonian teleost fish Schizodon fasciatus (Characiformes, Anostomidae). Parasite 11(2):169–173
Feijó MM, Arana S, Ceccarelli PS, Adriano EA (2008) Light and scanning electron microscopy of Henneguya arapaima n. Sp. (Myxozoa: Myxobolidae) and histology of infected sites in pirarucu (Arapaima gigas: Pisces: Arapaimidae) from the Araguaia River, Brazil. Vet Parasitol 157(1–2):59–64. https://doi.org/10.1016/j.vetpar.2008.06.009
Feist SW (2008) Myxozoan diseases. In: Eiras JC, Segner H, Wahli T, Kapoor BG (eds) fish diseases. Science publishers, Enfield, pp 613–682
Feist SW, Longshaw M (2006) Phylum Myxozoa. In: Woo PTK (ed) Fish diseases and disorders: protozoan and metazoan infections, 2nd edn. CAB International, Oxfordshire, pp 230–296. https://doi.org/10.1079/9780851990156.0230
Ferguson JA, Atkinson SD, Whipps CM, Kent ML (2008) Molecular and morphological analysis of Myxobolus spp. pf salmonids fishes with the description of a new Myxobolus species. J Parasitol 94:322–1334. https://doi.org/10.1645/GE-1606.1
Fiala I (2006) The phylogeny of Myxosporea (Myxozoa) based on small subunit ribosomal RNA gene analysis. Int J Parasitol 36(14):1521–1534. https://doi.org/10.1016/j.ijpara.2006.06.016
Fiala I, Bartošova-Sojková P, Okamura B, Hartikainen H (2015) Adaptive radiation and evolution within the Myxozoa. In: Okamura B, Gruhl A, Bartholomew JL (eds) Myxozoan evolution. Ecology and development. Springer, Cham, pp 69–84. https://doi.org/10.1007/978-3-319-14753-6_4
Freitas CEC, Campos CP (2014) Yield per recruit of the peacock bass Cichla monoculus (Spix and Agassiz, 1831) caught in Lago Grande at Manacapuru (Amazonas - Brazil). Braz J Biol 4:226–230
Froese R, Pauly D (2017) FishBase. World Wide Web electronic publication. http://www.fishbase.org. Accessed November 2017
Guindon S, Gascuel O (2003) A simple, fast and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52(5):696–604. https://doi.org/10.1080/10635150390235520
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59(3):307–321. https://doi.org/10.1093/sysbio/syq010
Hall TA (2011) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Hallett SL, Diamant A (2001) Ultrastructure and small-subunit ribosomal DNA sequence of Henneguya lesteri n. sp. (Myxosporea), a parasite of sand whiting Sillagoanalis (Sillaginidae) from the coast of Queensland, Australia. Dis Aquat Org 46(3):197–212. https://doi.org/10.3354/dao046197
Hoeinghaus DJ, Layman CA, Arrington AD, Winemiller KO (2003) Movement of Cichla species (Cichlidae) in a Venezuelan floodplain river. Neotrop Ichthyol 1(2):121–126. https://doi.org/10.1590/S1679-62252003000200006
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Kullander SO, Ferreira EJG (2006) A review of the South American cichlid genus Cichla, with descriptions of nine new species (Teleostei: Cichlidae). Ichthyol Explor Fresh 17:289–398
Kent ML, Andree KB, Bartholomew JL, El-Matbouli M, Desser SS, Devlin RH, Feist SW, Hedrick RP, Hoffmann RW, Khattra J, Hallett SL, Lester RJ, Longshaw M, Palenzeula O, Siddall ME, Xiao C (2001) Recent advances in our knowledge of the Myxozoa. J Eukaryot Microbiol 48(4):395–413. https://doi.org/10.1111/j.1550-7408.2001.tb00173.x
Lom J, Arthur JR (1989) A guideline for the preparation of species descriptions in Myxosporea. J Fish Dis 12(2):151–156. https://doi.org/10.1111/j.1365-2761.1989.tb00287.x
Lom J, Dyková I (2006) Myxozoan genera: definition and notes on taxonomy, life-cycle terminology and pathogenic species. Folia Parasitol 53(1):1–36
Matos E, Tajdari J, Azevedo C (2005) Ultrastructural studies of Henneguya rhamdia n. Sp. (Myxozoa) a parasite from the Amazon teleost fish, Rhamdia quelen (Pimelodidae). J Eukaryot Microbiol 52(6):532–537
Mathews PD, Maia AA, Adriano EA (2016) Henneguya melini n. sp. (Myxosporea: Myxobolidae), a parasite of Corydoras melini (Teleostei: Siluriformes) in the Amazon region: morphological and ultrastructural aspects. Parasitol Res 115(9):3599–3604. https://doi.org/10.1007/s00436-016-5125-z
Molnár K (1998) Taxonomic problems, seasonality and histopathology of Henneguya creplini (Myxosporea) infection of the pikeperch Stizostedion lucioperca in Lake Balaton. Folia Parasitol 45(4):261–269
Molnár K, Ranzani-Paiva MJ, Eiras JC, Rodrigues EL (1998) Myxobolus macroplasmodialis sp. n. (Myxozoa: Myxosporea), a parasite of the abdominal cavity of the characid teleost, Salminus maxillosus, in Brazil. Acta Protozool 37:241–245
Molnár K, Marton S, Eszterbauer E, Székely C (2006) Comparative morphological and molecular studies on Myxobolus spp. infecting chub from the River Danube, Hungary, and description of M. muellericus sp. n. Dis Aquat Org 73(1):49–61. https://doi.org/10.3354/dao073049
Moreira GSA, Adriano EA, Silva MRM, Ceccarelli OS, Maia AAM (2014a) Morphology and 18S rDNA sequencing identifies Henneguya visibilis n. sp., a parasite of Leporinus obtusidens from Mogi Guaçu River, Brazil. Parasitol Res 113(1):81–90. https://doi.org/10.1007/s00436-013-3629-3
Moreira GSA, Adriano EA, Silva MRM, Ceccarelli PS, Maia AAM (2014b) The morphological and molecular characterization of Henneguya rotunda n. sp., a parasite of the gill arch and fins of Salminus brasiliensis from the Mogi Guaçu River, Brazil. Parasitol Res 113(5):1703–1711. https://doi.org/10.1007/s00436-014-3815-y
Naldoni J, Maia AAM, Silva MRM, Adriano EA (2014) Henneguya cuniculator sp. nov., a parasite of spotted sorubim Pseudoplatystoma corruscans in the São Francisco Basin, Brazil. Dis Aquat Org 107(3):211–221. https://doi.org/10.3354/dao02685
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256. https://doi.org/10.1093/molbev/msn083
Rambaut A (2008) FigTree v1.1.1: tree figure drawing tool, available from http://tree.bio.ed.ac.uk/software/figtree/
Rambaut A, Drummond AJ (2007) Trace v1.4, available from http://beast.bio.ed.ac.uk/Tracer
Reis RE, Albert JS, Dario FD, Mincarone MM, Petry P, Rocha LA (2016) Fish biodiversity and conservation in South America. J Fish Biol 89(1):12–47. https://doi.org/10.1111/jfb.13016
Rocha E, Matos E, Azevedo C (1992) Henneguya amazonica n.Sp. (Myxozoa, Myxobolidae), parasitizing the gills of Crenicichla lepidota Heckel, 1840 (Teleostei, Cichlidae) from Amazon river. Eur J Protistol 28(3):273–278
Rocha S, Casal G, Garcia P, Matos E, Al-Quraishy S, Azevedo C (2014) Ultrastructure and phylogeny of the parasite Henneguya carolina sp. nov. (Myxozoa), from the marine fish Trachinotus carolinus in Brazil. Dis Aquat Org 112(2):139–148. https://doi.org/10.3354/dao02794
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19(12):11572–11574. https://doi.org/10.1093/bioinformatics/btg180
Thompson JD, Higgins DG, Gilson TJ (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucl Acid Res 22(22):4673–4680. https://doi.org/10.1093/nar/22.22.4673
Videira M, Velasco M, Azevedo R, Silva R, Gonçalves Matos P, Matos E (2015) Morphological aspects of Henneguya aequidens n. sp. (Myxozoa: Myxobolidae) in Aequidens plagiozonatus Kullander, 1984 (Teleostei: Cichlidae) in the Amazon region, Brazil. Parasitol Res 114(3):1159–1162. https://doi.org/10.1007/s00436-014-4295-9
Velasco M, Videira M, Nascimento LCS, Matos P, Gonçalves EC, Matos E (2016) Henneguya paraensis n. sp. (Myxozoa; Myxosporea), a new gill parasite of the Amazonian fish Cichla temensis (Teleostei: Cichlidae): morphological and molecular aspects. Parasitol Res 115(5):177–91787. https://doi.org/10.1007/s00436-016-4916-6
Vital P, Corral L, Matos E, Azevedo C (2003) Ultrastructural aspects of the myxosporean Henneguya astyanax n. Sp. (Myxozoa: Myxobolidae), a parasite of the Amazonian teleost Astyanax keithi (Characidae). Dis Aquat Org 53(1):55–60
Willis SC, Nunes MS, Montana CG, Farias IP, Lovejoy NR (2007) Systematics, biogeography, and evolution of the Neotropical peacock bases Cichla (Perciformes: Cichlidae). Mol Phy Evol 44:291–307
Zatti SA, Atkinson SD, Bartholomew JL, Maia AAM, Adriano EA (2017) Amazonian waters harbour an ancient freshwater Ceratomyxa lineage (Cnidaria: Myxosporea). Acta Trop 169:100–106. https://doi.org/10.1016/j.actatropica.2017.02.006
Zhang ZQ (2011) Animal biodiversity: an introduction to higher-level classification and taxonomic richness. Zootaxa 3148:7–12
Acknowledgments
The authors thank Prof. Dr. Lincoln Corrêa (UFOPA), Prof. Dr. Marcos Tavares-Dias, and Marcos Oliveira for assistance with fieldwork; the fishermen of the communities of Jari do Socorro; Santarém, Pará State; and of Jarilândia, Amapá State, for their local knowledge of fish availability and provision of material for study; and Prof. Dr. Regina Maura Bueno Franco (UNICAMP) for providing some of the equipment used in this study.
Funding
Part of this research was conducted while SAZ was a visiting scholar at Oregon State University, USA, which was sponsored by the CAPES Foundation within the Ministry of Education, Brazil (grant n. BEX–6729/2015-00). SAZ was also supported by a Ph.D. scholarship provided by CAPES to UNICAMP. This study was supported by the São Paulo Research Foundation–FAPESP (Procs. n. 2016/22047 to EAA). EAA received a research productivity grant from the Brazilian Fostering Agency CNPq (Proc. n. 301886/2016-4).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Fig. S1
Consensus ML phylogenetic tree using ssrDNA sequences of selected Myxobolus/Henneguya species, under GTR+G model. GenBank accession numbers are given in parenthesis. Nodal supports are indicated for ML with a bootstrap of 1000 replicates. Values for weakly supported nodes (< 50) are not shown. Asterisk corresponds to the African cichlids (Abdel-Gaber et al. 2017). Scale is 0.20. (GIF 1179 kb)
Rights and permissions
About this article
Cite this article
Zatti, S.A., Atkinson, S.D., Maia, A.A.M. et al. Novel Henneguya spp. (Cnidaria: Myxozoa) from cichlid fish in the Amazon basin cluster by geographic origin. Parasitol Res 117, 849–859 (2018). https://doi.org/10.1007/s00436-018-5762-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-018-5762-5