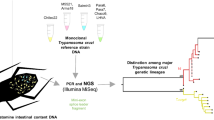
Abstract
Trypanosoma cruzi and Trypanosoma rangeli are kinetoplastid parasites which are able to infect humans in Central and South America. Misdiagnosis between these trypanosomes can be avoided by targeting barcoding sequences or genes of each organism. This work aims to analyze the feasibility of using species-specific markers for identification of intraspecific polymorphisms and as target for diagnostic methods by PCR. Accordingly, primers which are able to specifically detect T. cruzi or T. rangeli genomic DNA were characterized. The use of intergenic regions, generally divergent in the trypanosomatids, and the serine carboxypeptidase gene were successful. Using T. rangeli genomic sequences for the identification of group-specific polymorphisms and a polymorphic AT(n) dinucleotide repeat permitted the classification of the strains into two groups, which are entirely coincident with T. rangeli main lineages, KP1 (+) and KP1 (−), previously determined by kinetoplast DNA (kDNA) characterization. The sequences analyzed totalize 622 bp (382 bp represent a hypothetical protein sequence, and 240 bp represent an anonymous sequence), and of these, 581 (93.3 %) are conserved sites and 41 bp (6.7 %) are polymorphic, with 9 transitions (21.9 %), 2 transversions (4.9 %), and 30 (73.2 %) insertion/deletion events. Taken together, the species-specific markers analyzed may be useful for the development of new strategies for the accurate diagnosis of infections. Furthermore, the identification of T. rangeli polymorphisms has a direct impact in the understanding of the population structure of this parasite.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Trypanosoma cruzi and Trypanosoma rangeli are closely related protozoan parasites which are able to infect humans. Their geographical distribution overlaps in Central and South America sometimes resulting in the occurrence of mixed infections both in vertebrate and invertebrate hosts (Vallejo et al. 2009). T. cruzi is the etiological agent of Chagas’ disease, and T. rangeli is considered harmless for the vertebrate host. However, T. rangeli antigens induce a humoral immune response in the host leading to the production of antibodies that cross-react with T. cruzi antigens and may interfere with the diagnosis of Chagas’ disease (Guhl et al. 1985).
In Brazil, three human cases of trypanosomiasis rangeli were reported in the Amazon region (Coura et al. 1996), and two cases of coinfection with T. cruzi were described (De Sousa et al. 2008). This fact contrasts with the high prevalence of T. rangeli or T. rangeli-like parasites described and isolated from wild mammals and triatomines (Dias et al. 2010).
The characterization of T. rangeli kinetoplast DNA (kDNA) allowed the determination of two main T. rangeli lineages named KP1 (+) and KP1 (−) (Vallejo et al. 2002), which were later confirmed by analysis of other chromosomal regions (Vallejo et al. 2003; Cabrine-Santos et al. 2011). Interestingly, T. rangeli KP1 (−) strains were associated with vectors of the pallescens group, and the T. rangeli KP1 (+) strains were associated with the prolixus group, suggesting the coevolution between these subpopulations and the two species of the genus Rhodnius (Urrea et al. 2005).
DNA-based detection methods can be useful tools for determining intraspecific polymorphisms (Brandão et al. 2008; Jaeger and Brandão 2011) and an alternative for the differential diagnosis of infections. Microsatellites (MS) are repetitive sequences of mostly one to six nucleotides which have a widespread occurrence particularly in multicellular organisms (Devera et al. 2003). In T. cruzi, the amplification of microsatellite polymorphic loci was proven to be an important tool for direct typing T. cruzi in biological samples (Macedo et al. 2001). Single-nucleotide polymorphisms (SNPs) consist mostly of base substitutions in single nucleotide positions and are useful markers in population genetics and evolutionary studies (Landegren et al. 1998). In protozoa, SNPs present in the coding regions of genes may alter the primary structure of a protein involved in drug metabolism and are targets for pharmacogenetic analyses (Evans and Relling 1999).
In view of the importance of the development of new techniques for the correct diagnosis of Chagas’ disease, this study focuses on the specific molecular identification of T. cruzi and T. rangeli based on independent genomic DNA markers and on the study of sources of genetic variability in different isolates of T. rangeli.
Materials and methods
Parasite strains
In this work, we used eight T. rangeli strains and four T. cruzi strains. T. rangeli KP1 (+) strains (P07, P02, P18, P19, and Cas4) were isolated from blood culture and xenodiagnosis from opossums (Didelphis albiventris) captured in Minas Gerais, Brazil (Ramirez et al. 2002). T. rangeli KP1 (−) strains (SO18, SO29, and Gal60) were provided by Dr. Jaime Moreno, Departamento de Biologia, Faculdad de Ciencias Exactas y Naturales, Universidade de Antioquia, Colombia. T. cruzi strains Alv, AQ2 (Cabrine-Santos et al. 2011), Y (Silva and Nussenzweig 1953), and JG (Andrade et al. 1999) were previously characterized. Further information about T. cruzi and T. rangeli strains used in this work are presented in Table 1. DNA samples of Leishmania major Friedlin, Leishmania infantum JPCM5, and Leishmania braziliensis M2904 were provided by Dr. Angela Kaysel Cruz, Faculdade de Medicina de Ribeirão Preto, Universidade de São Paulo, Brasil.
Culture conditions and genomic DNA purification
Epimastigote forms of T. cruzi and T. rangeli were cultured at 28 °C in liver infusion tryptose (LIT) medium supplemented with 3 % (v/v) human urine (Ferreira et al. 2007), and the cell density was determined by counting in a hemocytometer.
Parasite genomic DNA was isolated by alkaline lysis of 40 mL of epimastigote culture in the exponential phase of growth containing approximately 2.5 × 107 parasites/mL (Vallejo et al. 1999). DNA samples were quantified at 260 nm using the BioPhotometer (Eppendorf).
Primer design
Primers used in PCR (Table 2) were synthesized by Invitrogen (CA, USA) and Integrated DNA Technologies (IDT, IA, USA). PCR was carried out using primers designed from three classes of sequences: (i) the conserved HSP70 chaperone gene with 97 % identity between T. rangeli and T. cruzi (TrypHSP70-F/TrypHSP70-R) used as a control, (ii) species-specific T. cruzi DNA, and (iii) species-specific T. rangeli DNA.
Species-specific T. cruzi primers (TcIR-F/TcIR-R and TcSCP-F/TcSCP-R) were designed from the intergenic region located between a hypothetical protein gene (GenBank accession number XM_801619.1) and an ubiquitin gene (XM_801620.1) and a T. cruzi serine carboxypeptidase, respectively (El-Sayed et al. 2005).
A hypothetical protein and two anonymous sequences potentially species-specific for T. rangeli (Ferreira et al. 2010) were used for the design of primer pairs Tr1F/Tr2R, Tr3F/Tr4R, and Tr5F/Tr6R, respectively. Tr1F-B/Tr2R-B, T6R-B primers were designed to obtain longer amplicons and consequently more quality on nucleotide sequence.
PCR conditions
Telomeric PCR (Chiurillo et al. 2003) was used to confirm the species of T. cruzi and T. rangeli. The reaction was conducted in a final volume of 30 μL containing 12 mM Tris–HCl (pH 9.0), 2.5 mM MgCl2, 0.24 mM of dNTPs, 1.25 units of Taq DNA polymerase recombinant (Invitrogen), 30 mM KCl, 0.4 μM of primers, and 1 ng of genomic DNA. PCR conditions were: 94 °C for 4 min and 30 cycles of 55 °C for 30 s, 72 °C for 40 s, and 94 °C for 1 min, followed by 55 °C for 1 min and 72 °C for 3 min.
PCR with primers designed from conserved and specific sequences of T. cruzi, T. rangeli, and Leishmania spp. were conducted in a final volume of 30 μL containing 10 mM Tris–HCl (pH 9.0), 2.5 mM MgCl2, 0.24 mM of dNTPs, 1.25 units of Taq DNA polymerase recombinant (Invitrogen), 30 pmol of each primer pair, and 1 ng of DNA. The amplification program consisted of an initial denaturation at 95 °C for 4 min, followed by 30 cycles of denaturation at 95 °C for 40 s, annealing at different temperatures according to the pair of primers used (Table 2) for 20 s, and extension at 72 °C for 15 s followed by final extension at 72 °C for 10 min in a MyCycler Thermal Cycler (Bio-Rad, CA, USA). All PCR reactions were optimized aiming at increasing specificity of the technique (Table 2).
The amplification products were visualized by 1.5 % (w/v) TAE agarose gel electrophoresis stained with 0.5 μg/mL of ethidium bromide solution and photographed through the specific filter in VersaDoc Imaging System (Bio-Rad).
Cloning of T. rangeli microsatellite fragments
Fragments containing the microsatellites of T. rangeli Cas4 were cloned into the pCR2.1-TOPO vector, forming the plasmid pCR-TrMS-Cas4, and the recombinant vector was transformed into Escherichia coli according to manufacturer instructions (Invitrogen). The transformed cells were cultured on LB agar containing ampicillin (100 μg/mL) overnight at 37 °C. Colonies were picked, consecutively numbered, and inoculated into LB/ampicillin medium and cultured at 180 rpm overnight at 37 °C for the extraction of plasmid DNA.
DNA sequencing
The amplification products of P07, P02, Cas4, SO18, SO19, and Gal60 strains of T. rangeli were excised from the agarose gel and purified using the Silica Bead DNA Gel Extraction Kit (Fermentas Life Sciences, Canada) and subjected to DNA sequencing in triplicate. Additionally, both strands of the pCR-TrMS-Cas4 were sequenced. A plasmid isolated from a previously described genome survey sequence library of T. rangeli P07 strain (pUC19-TrMS-P07, GenBank accession number FI569284) was purified by the alkaline lysis method and sequenced as described (Ferreira et al. 2010). The reaction was carried out in the MyCycler Thermal Cycler under the following conditions: 96 °C for 1 min, followed by 40 cycles at 96 °C for 15 s, 50 °C for 15 s, and 60 °C for 4 min. The nucleotide sequences were analyzed in an ABI PRISM 3130xl Genetic Analyzer sequencer (Applied Biosystems) using the Kit Big Dye v3.1 (Applied Biosystems) and the appropriate forward and reverse primers. Only bases with quality value (QV) above 20 (<1 % error in reading of bases) were considered for analysis (QV = −10 log10(Pe), where Pe is the probability of error).
In silico analysis with consensus generated by alignment of forward and reverse sequences was performed using the BLAST program (Altschul et al. 1990) and the ClustalW program (Thompson et al. 1994) to achieve the multiple alignments.
Phylogenetic analysis
For the phylogenetic analysis, we first conducted a statistical selection of best-fit models of nucleotide substitution for both type of data, i.e., six nucleotide polymorphisms and five microsatellite sequences, using the jModelTest2 software, that provides estimates of model selection uncertainty, parameter importance, and model-averaged parameter estimates, including model-averaged tree topologies (Darriba et al. 2012). The phenograms were calculated using the molecular evolutionary genetics analysis (MEGA5) integrated tool (Tamura et al. 2011) and visualized using the FigTree graphical viewer (tree.bio.ed.ac.uk/software/figtree).
Results
Primer design and amplification of genomic markers
The detection of size-specific telomeric sequences by PCR allowed the confirmation of the T. rangeli and T. cruzi species. As expected, it were present amplified products of 170 bp in P07, P18, P19, SO18, and SO29 T. rangeli strains and of 100 bp products in AQ2, ALV, Y, and JG T. cruzi strains (Fig. 1).
Characterization of parasite species by telomeric PCR. The amplified products were observed in 2.0 % agarose gel stained with ethidium bromide. Tc189Fw2/Tc189Rv3 and TrF3/TrR8 primers were used in PCR assays that amplify duplex telomeric and subtelomeric sequences (Chiurillo et al. 2003) of Trypanosoma cruzi and Trypanosoma rangeli, respectively. T. rangeli strains (1: P07, 2: P18, 3: P19, 4: Cas4, 5: SO18, and 6: SO29); T. cruzi stains (7: AQ2, 8: ALV, 9: Y, and 10: JG); 11: negative control (NC). MM molecular marker 100 bp (Invitrogen Life Technologies, Carlsbad, CA), Bands expected 170 bp for T. rangeli and 100 bp for T. cruzi
Primer pair TrypHSP70-F/TrypHSP70-R amplified a 241-bp fragment in both T. rangeli and T. cruzi DNA samples (Fig. 2a), as a control for DNA quality and integrity in the reactions. T. cruzi DNA-specific amplification was achieved with primer pairs IGTc-F/IGTc-R and TcSCP-F/TcSCP-R, which specifically amplified products of 295 and 621 bp in T. cruzi samples, respectively (Fig. 2b, c).
Specificity of the primers designed for Trypanosoma cruzi and Trypanosoma rangeli. Amplification products separated by electrophoresis and observed in 1.5 % agarose gel stained with ethidium bromide. Primers designed from the T. cruzi and T. rangeli chaperone HSP70 conserved sequence: a TrypHSP70-F/TrypHSP70-R. Specific T. cruzi primers: b ubiquitin intergenic region (TcIR-F/TcIR-R) and c serine carboxypeptidase (TcSCP-F/TcSCP-R). Specific T. rangeli primers: d Tr1F/Tr2R, e Tr3F/Tr4R, and f Tr5F/Tr6R. T. rangeli strains (1: P07, 2: P18, 3: P19, 4: Cas4, 5: SO18, and 6: SO29); T. cruzi strains (7: AQ2, 8: ALV, 9: Y, and 10: JG); Leishmania spp. strains (11: L. major, 12: L. infantum, 13: L. braziliensis); 14: negative control (NC). MM Molecular Marker 100 bp (Invitrogen Life Technologies, Carlsbad, CA)
In the T. rangeli-specific amplification reactions, PCR with Tr1F/Tr2R primers amplified a fragment of 189 bp only in T. rangeli strains, confirming the specificity of the PCR (Fig. 2d). Tr3F/Tr4R primers generated fragments of 260 bp specifically in T. rangeli (Fig. 2e). PCR with primers Tr5F/Tr6R generated amplicons in both KP1 (+) and KP1 (−) strains. The expected 280 bp fragment was obtained only for T. rangeli KP1 (+) strains, and a smaller fragment was obtained from KP1 (−) strains (Fig. 2f). For all primers used, no amplification of Leishmania spp. DNA was detected.
Group-specific polymorphisms in T. rangeli
In order to investigate further polymorphisms among KP1 (+) and KP1 (−) T. rangeli strains, amplicons generated from the Tr1F-B/Tr2R-B and Tr5F/Tr6R-B primer pairs were sequenced. Both sequences totalize 622 bp (382 bp represent a hypothetical protein sequence, and 240 bp represent an anonymous sequence), and of these, 581 (93.3 %) are conserved sites and 41 bp (6.7 %) are polymorphic, with 9 transitions (21.9 %), 2 transversions (4.9 %), and 30 (73.2 %) insertion/deletion (indel) events.
Analysis of multiple sequence alignments of these sequences allowed the identification of group-specific polymorphisms (GSP) in T. rangeli. Eight transitions and two transversions enabled the discrimination between T. rangeli KP1 (+) and KP1 (−) genotypes. Thus, the first pair of primers (ALPTr1F-B/ALPTr2R-B) enabled detection of the following polymorphisms in KP1 (−) strains compared to KP1 (+) strains: [KP1 (+) > KP1 (−)]:128 T > C, 160A > C, 196A > G, 223G > A and 338C > T (Fig. 3a). A detail of an electropherogram depicting an example of the GSP at position 128, with a nucleotide T in the T. rangeli KP1 (+) strains (P07, P02, and Cas4) and one substitution (128 T > C) in T. rangeli KP1 (−) strains (SO18, S029, and Gal60) is shown in Online Resource 1a and b. By using the Jukes and Cantor (JC) substitution model suggested by jModelTest2 for phylogenetic analysis, it is possible to determine a pattern of two groups (group 1: P02, P07, and Cas4 and group 2: SO29, Gal60, and SO18), which are in agreement with the parasite genetic groups [KP1 (+) and KP1 (−), respectively] previously described (Online Resource 1c). For the second pair of primers (ALPTr5F/ALPTr6R-B), the following polymorphisms were observed: 25 T > C, 26C > T, 82C > T, 90 T > C, and 100C > T (Fig. 3b).
Multiple sequence alignment from different strains of T. rangeli amplified by PCR using the following primers: a Tr1F-B/Tr2R-B and b Tr5F/Tr6R-B. The comparison among nucleotide sequences of strains was performed using the ClustalW program. Only representative sequences of KP1 (+) (P07 and Cas4) and KP1 (−) (SO18) are shown. Group-specific polymorphisms are indicated by boxes and dinucleotide repeats are underlined. Only polymorphic regions of the amplicons are shown
At position 214, it is possible to observe a pyrimidine-to-pyrimidine transition (Y) in the T. rangeli P07 strain. The other T. rangeli strains have only the nucleotide T in this locus (Online Resource 2a). The identification of this polymorphism has been made possible by the fact that the P07 strain sequences were generated from both, amplified DNA (with sequencing of the two alleles) and recombinant DNA (with the sequencing of a single allele). The detail of the electropherogram generated from recombinant DNA of the pUC19-TrSNP-P07 of T. rangeli GSS (GenBank FI569298) shows the presence of C nucleotide (Online Resource 2b), and the amplified genomic DNA shows the presence of two nucleotides (C and T) in the position 214 (Online Resource 2c).
Microsatellites in T. rangeli
Sequences obtained with primers Tr5F/Tr6R-B generated PCR products with distinct sizes due to the variability in the number of AT(n) in T. rangeli strains. Dinucleotide repeats (DRs) constitute one type of microsatellites. Figure 3b shows the multiple alignments of the sequences obtained from these primers. Sequencing of the pUC19-TrMS-P07 revealed the occurrence of 20 DRs (Online Resource 3a). On the other hand, the determination of the number of DR in the Cas4 amplicon was initially hampered due to the presence superimposed nucleotides after the 15th DR. The analysis of the chromatogram obtained from Cas4 strain suggested that amplification of the MS locus in the Cas4 strain generated two fragments with different sizes. In order to confirm this hypothesis, amplicons of the Cas4 MS locus (pCR-TrMS-Cas4) were cloned in a plasmid vector. Sequencing of recombinant clones confirmed the presence of a fragment of 16 DRs (pCR-TrMS-Cas4a) and another with just 15 DRs (pCR-TrMS-Cas4b) in the T. rangeli Cas4 strain (Online Resource 3a). In all KP1 (−) strains, only five DRs were observed, generating an amplified product of smaller size in SO18, SO29, and Gal60 T. rangeli strains (Online Resource 3b). As suggested by jModelTest2, we have used Kimura substitution model (K80) for phylogenetic analysis of the AT(n) DRs studied. As observed in Online Resource 3c, microsatellite analysis also allowed the formation of two groups entirely coincident with the clustering observed for GSP. Other microsatellites such as AG(3), CTA(2), GTTT(2), and GTTT(2) were also observed; however, these microsatellites showed no polymorphisms among the strains studied.
Discussion
In this study, we have characterized a set of primers which are able to specifically detect T. cruzi or T. rangeli DNA in culture samples. Several techniques have been used to distinguish T. rangeli and T. cruzi, using different PCR targets (Vallejo et al. 1999; Guhl et al. 2002). Among these, the most commonly used are PCRs with primers that recognize repetitive sequences and that are highly sensitive, since each cell of the parasite can carry from tens to thousands copies of a particular target. However, the use of PCR-based amplification of highly polymorphic regions could be a disadvantage in epidemiological studies by the lack of parameters for the comparison of sequences.
According to El-Sayed et al. (2005), fewer than 5 % of the protein domains identified in the trypanosomatid genomes studied so far are unique to a single species. Nevertheless, identification and diagnosis should be improved by targeting species-specific, lineage-specific sequences, or genes encoding proteins with putative roles in host–parasite relationships. In this context, we have explored the divergence in the intergenic regions of trypanosomatids by designing primers to specifically amplify T. cruzi sequences (the intergenic region between a hypothetical protein gene and an ubiquitin gene). Another genomic target used in this work was the serine carboxypeptidase protein, which in T. cruzi is involved in the process of the parasite entry in a wide variety of mammalian cells (Caler et al. 1998), which is not found in Trypanosoma brucei or in L. major (El-Sayed et al. 2005). We have demonstrated that this target also does not generate amplicons in T. rangeli and could be used in differential diagnosis between trypanosomes. The amplification obtained reached the expected objectives with the desired amplification of specific fragments in samples of T. cruzi.
The use of these genomic approaches for designing of the specific primers for T. rangeli from a Brazilian KP1 (+) strain was successful. Primers Tr1F/Tr2R and Tr2R/Tr3F allowed the conduction of the PCR with specific amplification of DNA from T. rangeli. Additionally, primers Tr5F/Tr6R, besides the specific amplification of T. rangeli, allowed the observation of a differential amplification pattern in T. rangeli samples. We have not observed amplification of undesired DNA fragments of Leishmania spp.
Sequencing of the amplicons from the PCR reaction using Tr1F-B/Tr2R-B and Tr5F/Tr6R-B primers allowed identification of the GSPs. These polymorphisms are entirely coincident with T. rangeli main lineages, KP1 (+) and KP1 (−), previously determined by kDNA classification (Vallejo et al. 2002) and for other molecular studies that show a bipolarity of T. rangeli genotype (Grisard et al. 1999; Cabrine-Santos et al. 2011; Cuervo et al. 2013). The GSPs detected in this study (ten events) are mainly transitions and occurred approximately one GSP every 56 nucleotides. Compared to other sequence polymorphisms, the frequency of the T. rangeli GSPs is approximately ten times greater than the frequency of SNPs detected in other protozoa. In Plasmodium falciparum, the frequency of SNPs was determined from the sequence of chromosome 3 and was estimated in 1/917 bp (Mu et al. 2002). In Plasmodium vivax, the study of a 100-kb region revealed a frequency of 1/523 bp, mainly due to the large number of intergenic regions analyzed (Feng et al. 2003). In other trypanosomatids, such as T. brucei and T. cruzi, the frequency of SNPs is estimated in 1/453 and 1/611 bp, respectively (Jackson et al. 2010, TcSNP v2 online resource).
The presence of polymorphism in T. rangeli P07 strain was possible by comparing the sequence of a recombinant clone with the sequence of an amplicon. The recombinant clone showed the presence of a cytosine in position 214, since during the cloning process, a single allele is represented in the plasmid (Ferreira et al. 2010). In contrast, the sequence of corresponding region of the amplicon showed the presence of both cytosine and thymine in position 214 (C > T transition). This observation can be explained by the fact that, after the initial denaturation, both alleles of the target were amplified by PCR. The analysis of the locus containing the AT(n) DR with primers Tr5F/Tr6F-B Cas4 strain also revealed the presence of a polymorphism between alleles of the Cas4 T. rangeli strain. To our knowledge, these polymorphisms have not been described for this parasite and may have arisen from sexual crosses between T. rangeli strains, a process already described for other trypanosomes (Gaunt et al. 2003; Jenni et al. 1986).
Furthermore, sequencing of the PCR products generated from primers Tr5F/Tr6R enabled the determination of the presence of polymorphic DRs which divide the species in two groups related to the previous kDNA classification, thus justifying the different sizes of amplicons observed in agarose gels. These results increase the studies of the genetic structure of T. rangeli showing a polymorphism in a species-specific genomic target in strains that comprise a single group according to kDNA classification (Vallejo et al. 2002). In our analysis, the microsatellite identified consists of AT(n) repetitions, which are considered more stable than others microsatellites, in particular, CG-rich trinucleotides repeats and CA DRs (Kuhl and Caskey 1993).
In trypanosomatids, genes are organized into polycistronic transcription units separated by intergenic regions with frequent DRs in T. cruzi (Andersson et al. 1998). Repetitive sequences were considered initially as useless genomic elements accumulated as junk (Ohno 1972). Actually, DRs have been proposed to be involved in regulation of gene expression, DNA metabolic processes, gene evolution (Li et al. 2002), and as target sequences for specific protein recognition (Epplen et al. 1996). The anonymous sequence that served as target for identifying microsatellite in T. rangeli may be an intergenic region. In T. cruzi, there is evidence that intergenic regions have specific regulatory sequences responsible for operating at different levels, transcriptional and posttranslational (Venegas et al. 2010; Duhagon et al. 2011).
Indel events and microsatellite variation in mini-exon gene and internal transcribed spacer (ITS) sequences of rDNA (Grisard et al. 1999; Beltrame-Botelho et al. 2005) have been described as sources of genetic variation in T. rangeli. In T. cruzi, the analysis of indel events in the spliced leader intergenic region (SL-IR) allowed the identification of subgroups within the Tc I group (Herrera et al. 2007; Cura et al. 2010). In T. rangeli, polymorphisms in the SL-IR and in the small subunit rDNA (SSU rDNA) (Maia da Silva et al. 2004) and cathepsin L-like gene (Ortiz et al. 2009) suggest five subpopulations denoted T. rangeli A-E. In T. cruzi, the genetic distance among strains can be demonstrated by phylogenetic analysis based on microsatellites (Oliveira et al. 1998). The T. rangeli analysis of subgroups provides a better understanding of the epidemiology and interaction between the parasite and the vertebrate and invertebrate hosts (Guhl et al. 2002).
The results obtained by the use of new genomic DNA markers confirm the specificity of PCR for discriminating T. cruzi and T. rangeli strains and for identifying the presence of polymorphisms in parasite strains. Moreover, the use of the Tr5F/Tr6R-B primers even opened the perspective of identifying different genotypes of T. rangeli. These primers could be used in further analysis to test the identification of T. rangeli and T. cruzi in humans and other biological fluids.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. doi:10.1016/S0022-2836(05)80360-2
Andersson B, Aslund L, Tammi M, Tran AN, Hoheisel JD, Pettersson U (1998) Complete sequence of a 93.4-kb contig from chromosome 3 of Trypanosoma cruzi containing a strand-switch region. Genome Res 8(8):809–816
Andrade LO, Machado CR, Chiari E, Pena SD, Macedo AM (1999) Differential tissue distribution of diverse clones of Trypanosoma cruzi in infected mice. Mol Biochem Parasitol 100(2):163–172
Beltrame-Botelho IT, Gaspar-Silva D, Steindel M, Dávila AM, Grisard EC (2005) Internal transcribed spacers (ITS) of Trypanosoma rangeli ribosomal DNA (rDNA): a useful marker for inter-specific differentiation. Infect Genet Evol 5(1):17–28. doi:10.1016/j.meegid.2004.05.007
Brandão A, Samudio F, Fernandes O, Calzada JE, Sousa OE (2008) Genotyping of Panamanian Trypanosoma cruzi stocks using the calmodulin 3’UTR polymorphisms. Parasitol Res 102(3):523–526. doi:10.1007/s00436-007-0775-5
Cabrine-Santos M, Ramírez LE, Lages-Silva E, de Souza BF, Pedrosa AL (2011) Sequencing and analysis of chromosomal extremities of Trypanosoma rangeli in comparison with Trypanosoma cruzi lineages. Parasitol Res 108(2):459–466. doi:10.1007/s00436-010-2087-4
Caler EV, Vaena de Avalos S, Haynes PA, Andrews NW, Burleigh BA (1998) Oligopeptidase B-dependent signaling mediates host cell invasion by Trypanosoma cruzi. EMBO J 17(17):4975–4986. doi:10.1093/emboj/17.17.4975
Chiurillo MA, Crisante G, Rojas A, Peralta A, Dias M, Guevara P et al (2003) Detection of Trypanosoma cruzi and Trypanosoma rangeli infection by duplex PCR assay based on telomeric sequences. Clin Diagn Lab Immunol 10(5):775–779
Coura JR, Fernandes O, Arboleda M, Barrett TV, Carrara N, Degrave W et al (1996) Human infection by Trypanosoma rangeli in the Brazilian Amazon. Trans R Soc Trop Med Hyg 90(3):278–279
Cuervo C, Thomas MC, López MC, Puerta CJ (2013) Sequence polymorphism in the Trypanosoma rangeli HSP70 coding genes allows typing of the parasite KP1(+) and KP1(−) groups. Exp Parasitol 133(4):447–453. doi:10.1016/j.exppara.2013.01.001
Cura CI, Mejía-Jaramillo AM, Duffy T, Burgos JM, Rodriguero M, Cardinal MV et al (2010) Trypanosoma cruzi I genotypes in different geographical regions and transmission cycles based on a microsatellite motif of the intergenic spacer of spliced-leader genes. Int J Parasitol 40(14):1599–1607. doi:10.1016/j.ijpara.2010.06.006
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModel Test2: more models, new heuristics and parallel computing. Nat Methods 9(8):772. doi:10.1038/nmeth.2109
De Sousa MA, da Silva Fonseca T, Dos Santos BN, Dos Santos Pereira SM, Carvalhal C, Hasslocher Moreno AM (2008) Trypanosoma rangeli Tejera, 1920, in chronic Chagas’ disease patients under ambulatory care at the Evandro Chagas Clinical Research Institute (IPEC-Fiocruz, Brazil). Parasitol Res 103(3):697–703. doi:10.1007/s00436-008-1033-1
Devera R, Fernandes O, Coura JR (2003) Should Trypanosoma cruzi be called “cruzi” complex? A review of the parasite diversity and the potential of selecting population after in vitro culturing and mice infection. Mem Inst Oswaldo Cruz 98(1):1–12
Dias FB, Quartier M, Romaña CA, Diotaiuti L, Harry M (2010) Tamandua tetradactyla Linnaeus, 1758 (Myrmecophagidae) and Rhodnius robustus Larrousse, 1927 (Triatominae) infection focus by Trypanosoma rangeli Tejera, 1920 (Trypanosomatidae) in Attalea phalerata Mart. ex Spreng (Arecaceae) palm tree in the Brazilian Amazon. Infect Genet Evol 10(8):1278–1281. doi:10.1016/j.meegid.2010.06.020
Duhagon MA, Smircich P, Forteza D, Naya H, Williams N, Garat B (2011) Comparative genomic analysis of dinucleotide repeats in Tritryps. Gene 487(1):29–37. doi:10.1016/j.gene.2011.07.022
El-Sayed NM, Myler PJ, Blandin G, Berriman M, Crabtree J, Aggarwal G et al (2005) Comparative genomics of trypanosomatid parasitic protozoa. Science 309(5733):404–409. doi:10.1126/science.1112181
Epplen JT, Kyas A, Mäueler W (1996) Genomic simple repetitive DNAs are targets for differential binding of nuclear proteins. FEBS Lett 389(1):92–95
Evans WE, Relling MV (1999) Pharmacogenomics: translating functional genomics into rational therapeutics. Science 286(5439):487–491
Feng X, Carlton JM, Joy DA, Mu J, Furuya T, Suh BB et al (2003) Single-nucleotide polymorphisms and genome diversity in Plasmodium vivax. Proc Natl Acad Sci U S A 100(14):8502–8507. doi:10.1073/pnas.1232502100
Ferreira KA, Lemos-Júnior PE, Lages-Silva E, Ramírez LE, Pedrosa AL (2007) Human urine stimulates in vitro growth of Trypanosoma cruzi and Trypanosoma rangeli. Parasitol Res 101(5):1383–1388. doi:10.1007/s00436-007-0654-0
Ferreira KA, Ruiz JC, Dias FC, Lages-Silva E, Tosi LR, Ramírez LE et al (2010) Genome survey sequence analysis and identification of homologs of major surface protease (gp63) genes in Trypanosoma rangeli. Vector Borne Zoonotic Dis 10(9):847–853. doi:10.1089/vbz.2009.0128
Gaunt MW, Yeo M, Frame IA, Stothard JR, Carrasco HJ, Taylor MC et al (2003) Mechanism of genetic exchange in American trypanosomes. Nature 421(6926):936–939. doi:10.1038/nature01438
Grisard EC, Campbell DA, Romanha AJ (1999) Mini-exon gene sequence polymorphism among Trypanosoma rangeli strains isolated from distinct geographical regions. Parasitology 118(Pt 4):375–382
Guhl F, Hudson L, Marinkelle CJ, Morgan SJ, Jaramillo C (1985) Antibody response to experimental Trypanosoma rangeli infection and its implications for immunodiagnosis of South American trypanosomiasis. Acta Trop 42(4):311–318
Guhl F, Jaramillo C, Carranza JC, Vallejo GA (2002) Molecular characterization and diagnosis of Trypanosoma cruzi and T. rangeli. Arch Med Res 33(4):362–370
Herrera C, Bargues MD, Fajardo A, Montilla M, Triana O, Vallejo GA et al (2007) Identifying four Trypanosoma cruzi I isolate haplotypes from different geographic regions in Colombia. Infect Genet Evol 7(4):535–539. doi:10.1016/j.meegid.2006.12.003
Jackson AP, Sanders M, Berry A, McQuillan J, Aslett MA, Quail MA et al (2010) The genome sequence of Trypanosoma brucei gambiense, causative agent of chronic human African trypanosomiasis. PLoS Negl Trop Dis 4(4):e658. doi:10.1371/journal.pntd.0000658
Jaeger LH, Brandão A (2011) The composition of upstream open reading frames (uORF) in four genes from Trypanosoma cruzi typical strains. Parasitol Res 109(4):1205–1208. doi:10.1007/s00436-011-2430-4
Jenni L, Marti S, Schweizer J, Betschart B, Le Page RW, Wells JM et al (1986) Hybrid formation between African trypanosomes during cyclical transmission. Nature 322(6075):173–175. doi:10.1038/322173a0
Kuhl DP, Caskey CT (1993) Trinucleotide repeats and genome variation. Curr Opin Genet Dev 3(3):404–407
Landegren U, Nilsson M, Kwok PY (1998) Reading bits of genetic information: methods for single-nucleotide polymorphism analysis. Genome Res 8(8):769–776
Li YC, Korol AB, Fahima T, Beiles A, Nevo E (2002) Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review. Mol Ecol 11(12):2453–2465
Macedo AM, Pimenta JR, Aguiar RS, Melo AI, Chiari E, Zingales B et al (2001) Usefulness of microsatellite typing in population genetic studies of Trypanosoma cruzi. Mem Inst Oswaldo Cruz 96(3):407–413
Maia Da Silva F, Noyes H, Campaner M, Junqueira ACV, Coura JR, Añez N et al (2004) Phylogeny, taxonomy and grouping of Trypanosoma rangeli isolates from man, triatomines and sylvatic mammals from the widespread geographical origin based on SSU and ITS ribosomal sequences. Parasitology 129:459–561. doi:10.1017/S0031182004005931
Marquez DS, Ramirez LE, Moreno J, Pedrosa AL, Lages-Silva E (2007) Trypanosoma rangeli: RAPD-PCR and LSSP-PCR analyses of isolates from southeast Brazil and Colombia and their relation with KP1 minicircles. Exp Parasitol 117(1):35–42. doi:10.1016/j.exppara.2007.03.007
Mu J, Duan J, Makova KD, Joy DA, Huynh CQ, Branch OH et al (2002) Chromosome-wide SNPs reveal an ancient origin for Plasmodium falciparum. Nature 418(6895):323–326. doi:10.1038/nature00836
Ohno S (1972) So much “junk” DNA in our genome. Brookhaven Symp Biol 23:366–370
Oliveira RP, Broude NE, Macedo AM, Cantor CR, Smith CL, Pena SD (1998) Probing the genetic population structure of Trypanosoma cruzi with polymorphic microsatellites. Proc Natl Acad Sci U S A 95(7):3776–3780
Ortiz PA, Maia da Silva F, Cortez AP, Lima L, Campaner M, Pral EM et al (2009) Genes of cathepsin L-like proteases in Trypanosoma rangeli isolates: markers for diagnosis, genotyping and phylogenetic relationships. Acta Trop 112(3):249–259. doi:10.1016/j.actatropica.2009.07.036
Ramirez LE, Lages-Silva E, Alvarenga-Franco F, Matos A, Vargas N, Fernandes O et al (2002) High prevalence of Trypanosoma rangeli and Trypanosoma cruzi in opossums and triatomids in a formerly-endemic area of Chagas disease in Southeast Brazil. Acta Trop 84(3):189–198
Silva LHP, Nussenzweig CM (1953) Sobre uma cepa de Trypanosoma cruzi altamente virulenta para o camundongo branco. Folia Clínica e Biológica 29:191
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. doi:10.1093/molbev/msr121
TcSNP v2 online resource/database. Available at http://snps.tcruzi.org. Accessed 26 June 2013
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Urrea DA, Carranza JC, Cuba CA, Gurgel-Gonçalves R, Guhl F, Schofield CJ et al (2005) Molecular characterization of Trypanosoma rangeli strains isolated from Rhodnius ecuadoriensis in Peru, R. colombiensis in Colombia and R. pallescens in Panama, supports a co-evolutionary association between parasites and vectors. Infect Genet Evol 5(2):123–129. doi:10.1016/j.meegid.2004.07.005
Vallejo GA, Guhl F, Chiari E, Macedo AM (1999) Species specific detection of Trypanosoma cruzi and Trypanosoma rangeli in vector and mammalian hosts by polymerase chain reaction amplification of kinetoplast minicircle DNA. Acta Trop 72(2):203–212
Vallejo GA, Guhl F, Carranza JC, Lozano LE, Sánchez JL, Jaramillo JC et al (2002) kDNA markers define two major Trypanosoma rangeli lineages in Latin-America. Acta Trop 81(1):77–82
Vallejo GA, Guhl F, Carranza JC, Moreno J, Triana O, Grisard EC (2003) Parity between kinetoplast DNA and mini-exon gene sequences supports either clonal evolution or speciation in Trypanosoma rangeli strains isolated from Rhodnius colombiensis, R. pallescens and R. prolixus in Colombia. Infect Genet Evol 3(1):39–45
Vallejo GA, Guhl F, Schaub GA (2009) Triatominae-Trypanosoma cruzi/T. rangeli: vector-parasite interactions. Acta Trop 110(2–3):137–147. doi:10.1016/j.actatropica.2008.10.001
Venegas J, Miranda S, Coñoepan W, Pîchuantes S, Jercic MI, González C et al (2010) Microsatellite marker analysis shows differentiation among Trypanosoma cruzi populations of peripheral blood and dejections of Triatoma infestans fed on the same chronic chagasic patients: microsatellite marker analysis and T. cruzi. Parasitol Res 107(4):855–863. doi:10.1007/s00436-010-1939-2
Acknowledgments
Authors thank the members of the Laboratório Multiusuário-UFTM for processing DNA sequences and Maria Teresa Santos, MSc, from the International Cooperation Department-UFTM for English review. This work was supported by FAPEMIG (Fundação de Amparo à Pesquisa do Estado de Minas Gerais)—APQ—00769–11.
Conflict of interest
The authors declare that they have no conflict of interest and that the experiments comply with the current laws in Brazil.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
Detail of electropherograms showing the sequence region obtained with Tr1F-B/Tr2R-B primers with presence of a Group-Specific Polymorphisms at position 128. a) nucleotide T in T. rangeli KP1 (+) strains (P07, P02, and Cas4). b) nucleotide C in T. rangeli KP1 (−) strains (SO18, S029, and Gal60). c) Phenogram build using the JC substitution model with sequences where SNPs were found in T. rangeli, showing a pattern of two different groups KP1 (+) and KP1 (−). (PDF 36 kb)
Online Resource 2
Detail of multiple sequence alignment from different strains of T. rangeli and electropherograms at position 214. a) Comparison among amino acid sequences of P07, P02, Cas4, SO18, SO29, and Gal60 strains was performed using the ClustalW program from products amplified by PCR using primers Tr1F-B/Tr2R-B. The areas highlighted in red consist of polymorphic regions. b) Detail of electropherograms showing the nucleotides at position 214 of the recombinant clone sequence of P07 T. rangeli strain (pUC19-TrMS-P07, GenBank Access FI569284) and c) in the same position on the amplicon sequence obtained by PCR. (PDF 12 kb)
Online Resource 3
Detail of electropherograms showing the dinucleotide repeats (DRs) in T. rangeli strains. a) P07 T. rangeli KP1 (+) strain (pUC19-TrMS-P07) with 20 DRs, and Cas4 T. rangeli KP1 (+) strain (pCR-TrMS-Cas4a and pCR-TrMS-Cas4b) with 16 and 15 DRs, respectively. b) SO18, S029, and Gal60 T. rangeli KP1 (−) strains with 05 DRs (SO18, SO29, and Gal60). c) Phenogram build using the K80 substitution model with the microsatellite sequences found in T. rangeli, showing a pattern of two different groups similar to KP1 (+) and KP1 (−) groups. (PDF 75 kb)
Rights and permissions
About this article
Cite this article
Ferreira, K.A.M., Fajardo, E.F., Baptista, R.P. et al. Species-specific markers for the differential diagnosis of Trypanosoma cruzi and Trypanosoma rangeli and polymorphisms detection in Trypanosoma rangeli . Parasitol Res 113, 2199–2207 (2014). https://doi.org/10.1007/s00436-014-3872-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-014-3872-2