Abstract
To investigate whether Trypanosoma cruzi populations found in chagasic cardiopathic and non-cardiopathic patients are genetically differentiated, three molecular microsatellite markers were analysed. This analysis was also applied to compare T. cruzi samples from peripheral blood or dejections of Triatoma infestans fed on the blood of the same patients. In order to obtain the first objective, analyses of predominant T. cruzi genotypes were conducted using three approaches: a locus-by-locus analysis; a Fisher method across three loci; and analysis of molecular variance by Genepop and Arlequin programs. Only with one locus and on the blood samples was a significant differentiation detected among non-cardiopathic and cardiopathic groups, which was not confirmed by the other two methods. On the contrary, with the three approaches, it was found that T. cruzi clones present in the blood of patients are genetically differentiated from those detected in dejections of T. infestans fed on the same patients. Our results showed that the most frequent lineage both in blood as well as in triatomine dejection samples was TcI. No significant difference in T. cruzi lineage distribution was observed among chagasic cardiopathic and non-cardiopathic patients. The majority of the samples (50–60%) had only one T. cruzi clone (uniclonal) either in blood or dejection samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
It is estimated that 15–16 million people are infected with Trypanosoma cruzi in Latin America and 75–90 million are exposed to infection (Rodrigues Coura and Pinto Dias 2009). T. cruzi is a diploid organism, and diverse studies have demonstrated that sexual reproduction is absent or very rare and that the reproduction of this parasite is eminently clonal (Tibayrenc and Ayala 1988). Despite their high degree of genetic variability, T. cruzi isolates can be classified into two major phylogenetic lineages and six discrete typing units (DTU), which, according to the last agreement of several researchers, will be named from now on as: TcI (DTU I), TcII (DTU IIb), TcIII (DTU IIc), TcIV (DTU IIa), TcV (DTU IId) and TcVI (DTU IIe; Brisse et al. 2000; Zingales et al. 2009). There is also consensus that even within the same six current lineages, there is greater genetic variability which may be related to still unknown aspects of the biology of T. cruzi (Llewelyn et al. 2009a, b; Zingales et al. 2009).
PCR, PCR hybridization, RAPD and microsatellite markers have demonstrated that isolated stocks from vectors or wild mammals are more complex than those obtained from patients in the acute phase of Chagas disease and than those obtained from patients with chronic disease (Oliveira et al. 1998). Clonal selection may also occur during the isolation of the parasite to be maintained “in vitro” (Deane et al. 1984; Solari et al. 2001). As a consequence of such clonal selection, the parasites obtained from culture for molecular characterization can be different from those present in the infected host (Macedo et al. 2002). An analysis of the distribution of clones of T. cruzi in humans and vectors (Triatoma infestans) of the same area found significant differences, attributed to particular characteristics of the human host and vector (Breniere et al. 1998; Llewelyn et al. 2009a).
The importance of the possible role of multiclonal infections has not been sufficiently weighed. Genetic analyses in T. cruzi have verified that a human host (Breniere et al. 1992) and a given insect vector frequently are infected by two clones of the parasite, phylogenetically distant (Rozas et al. 2007). It has been postulated that multiple infections by genetically diverse clones constitutes an important evolutionary strategy for many pathogens (Tibayrenc et al. 1986; Tibayrenc and Ayala 1991).
The technique of microsatellites described by Macedo et al. (2001) has high sensitivity, proven in biological samples, since it allows detection of the equivalent to the middle of DNA genome of a parasite. In addition, the technique is highly reproducible and allows discrimination between several clones in a single test.
Although the interruption of vectorial transmission in Chile was obtained some years ago, there are still thousands of people infected with the parasite in Chile, many of whom have developed important cardiological affections. Recent reports show that in patients with decompensated heart failure and similar clinical conditions, those with Chagas disease had worse prognoses (Pereira Silva et al. 2008).
To determine if there are genetic differences between the T. cruzi populations of non-cardiopathic and cardiopathic persons and to investigate whether human and insect hosts select T. cruzi populations, genotypic multilocus analyses of different types of samples were conducted using the Genepop and Arlequin programmes (Excoffier et al. 1992; Raymond and Rousset 1995a, b).
Materials and methods
Patients
The study included two chronic chagasic patient groups composed of 42 non-cardiopathic and 32 cardiopathic individuals, both groups with positive conventional serology (ELISA and indirect immunofluorescence, IF) and circulating T. cruzi (PCR-positive). All patients were from the IV, V and Metropolitan Regions of Chile, endemic areas of the disease, and were treated with allopurinol (8.5 mg kg−1 day−1 for 60 days) in 1992. The ad hoc protocols and regulations of the Ethics Committee of the Faculty of Medicine, University of Chile, were followed throughout the course of this investigation (Apt et al. 1998).
The titres of the patients studied by IF fluctuated between 1/60 (minimum positive) and 1/1,280 (maximum obtained by us). Patients had antibody levels determined by ELISA at an optical density (OD) of 490 nm ranging between 1.596 and 2.026. The cutoff value was 0.2 DO, using epimastigotes as antigens, as usual (Zulantay et al. 2004).
Cardiology study
Each patient was given a 12-lead electrocardiogram (EKG) examination twice a year during the 13-year follow-up. Therefore, each patient had had at least 20 EKGs at the time of this study. The cardiac evaluation, in addition to the EKG, included a chest radiograph and, in some cases, an Eco-Doppler. The double-blind protocol recommended by the World Health Organization was followed. The cardiologist analyzing the EKG profiles was unaware of the patient’s infection status or any previous EKG results (Apt et al. 2003).
Each patient was classified as chronic chagasic cardiomyopathy (CCC) only if the different analysis mentioned above (conventional serology, PCR, cardiologist evaluation, EKG profiles and chest radiographs) remained consistent with the diagnosis of CCC for a period of 10 years.
The xenodiagnosis test and triatomine samples
The xenodiagnosis test uses two cylindrical wooden boxes, each containing seven uninfected third-instar T. infestans nymphs, maintained in our laboratory since 1950 (Schenone 1999). The insects were allowed to feed for 20–30 min on the arm of each patient. Microscopic examination of insect dejections was performed 30, 60 and 90 days after feeding triatomines with peripheral blood from infected individuals. The sensitivity of this test is about 80% (Zulantay et al. 2007).
The faecal samples of all the triatomines obtained after 30, 60 and 90 days from each patient were pooled and used for PCR analysis. The material was placed in Diamond medium, incubated at 98°C for 15 min and centrifuged at 4,000 rpm for 3 min. The supernatant was loaded onto a P6 column to remove low-molecular-weight DNA polymerase inhibitors, and the wash through was collected and frozen at −20°C until use.
Detection of T. cruzi DNA in peripheral blood and triatomine dejection samples
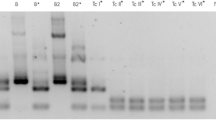
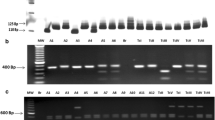
PCR was performed in triplicate using DNA extracted from peripheral blood samples (PCR-B) or triatomine dejections (PCR-D) following procedures previously described (Solari et al. 2001). Briefly, 800 μL of blood sample was used for nucleic acid extraction and the DNA suspended in 50 μL of sterile water. The reaction mix contained 10 pmol of forward primer 121 (5′-AAA TAA TGT ACG GG(T/G) GAG ATG CAT GA-3′), 10 pmol of reverse primer 122 (5′-GGT TCG ATT GGG GTT GGT GTA ATA TA-3′), 5 μL of amplification buffer (1.5 mM MgCl2, 50 mM KCl, 10 mM Tris–HCl pH 8.3), 0.2 mM of each dNTP, 1.5 U of Taq DNA polymerase (Promega) and 5 μL of template in a final volume of 50 μL. The amplification procedure included an initial denaturation at 94°C for 4 min followed by 35 cycles (94°C for 30 s, 64°C for 30 s and 72°C for 30 s). A final incubation at 72°C for 10 min was included to ensure full elongation of the amplified fragments. A Techne TC-412 thermocycler and a DNA clean chamber (AirClean 600 PCR workstation) were used to prevent contamination. DNAs prepared from a confirmed infected individual and from a non-infected individual were included as positive and negative controls, respectively. A similar PCR protocol was used to process the triatomine samples. Amplification products were separated by electrophoresis in 2% agarose gels in 0.5× TBE buffer (44.5 mM Tris–HCl, pH 8.0, 44.5 mM boric acid and 1.1 mM EDTA) as described (Venegas et al. 2009a).
Analysis of microsatellites
Three microsatellite loci were analysed (SCLE10, SCLE11 and MCLE01) with a modification of the technique described by Oliveira et al. (1998) using a second amplification with an aliquot of 1 μL from the first PCR and the same amplification conditions. The alleles were detected using primers stained with fluorescence (Oliveira et al. 1998). The amplification products were sent to the Roy J. Carver Biotechnology Center–University of Illinois, USA, for analysis by capillary electrophoresis and fluorescence detection with an automatic sequencer and an appropriate software programme. The number of base pairs of each allele was determined using cloned alleles from each marker as controls. In this analysis, the minimal detectable peak height was set to 80 arbitrary fluorescence units (FU).
Genotyping of predominant T. cruzi clone in each DNA sample
The determination of the predominant T. cruzi clone genotype in samples containing multiple alleles per locus was conducted taking into account the distance and the height in FU of the highest two peaks, the major and second peaks, directly in the electropherogram. When the height ratio between the second and the major peak was <0.33, it was concluded that the predominant clone genotype in this sample has only a single allele and therefore is homozygous. By contrast, if the height ratio between the second and the major peaks was ≥0.33, it was analysed further locus by locus. Thus, if the peaks were obtained by the SCLE10 marker and are separated by a maximum distance not superior to 32 bp, it is concluded that the genotype of predominant clone is formed by two alleles and therefore is heterozygous. But if the distance, at which two peaks are located, is >32 bp, they are considered different clones and we must find a third peak that should locate within 32 bp from the major peak. If the height ratio between the third and the major peaks was ≥0.33, it was concluded that the predominant SCLE10 genotype is heterozygous and formed by the major and the third peaks. On the contrary, if the height ratio between the third and the major peaks was <0.33, finally, it was concluded that the predominant SCLE10 genotype in this sample has only a single allele and therefore is homozygous. Similar analysis was conducted, using the SCLE11 and MCLE01 markers, with the exception that the maximum distance of separation between the major and the second peaks were 16 and 20 bp, respectively. This approach was applied with corresponding SCLE10, SCLE11 and MCLE01 locus in each T. cruzi sample (Electronic supplementary materials (ESM) Table S1). These criteria were used based on the height ratio and distances between the two alleles found in different T. cruzi heterozygous clones analysed previously by us (Venegas et al. 2009b).
The minimal number of T. cruzi clones in each triatomine sample was estimated based on the total number of alleles found in each sample divided by two. An odd number of alleles was considered as another clone, i.e. three alleles must be from two clones.
Classification of T. cruzi DNA samples
From each patient was obtained a peripheral blood sample and was subjected to xenodiagnosis. From peripheral blood and dejection of xenodiagnosis triatomines was extracted a DNA sample, respectively. T. cruzi DNA samples were classified into four groups according to their origins: N-B (Blood from 42 non-cardiopathic individuals), C-B (blood from 32 cardiopathic individuals, N-X (dejections of xenodiagnosis of the same 42 non-cardiopaths) and C-X (dejections of xenodiagnosis of the same 32 cardiopaths; ESM Table S1).
Analysis of genetic differentiation
This was conducted by three approaches based on genotype frequencies locus by locus using the SCLE10, SCLE11 and MCLE01 loci; multilocus analysis by the Fisher method across these three loci; and analysis of molecular variance (AMOVA). Genotype differentiation by the first and the second approaches was studied with the Genepop programme using the Markov algorithm and the Fisher method (Raymond and Rousset 1995a, b). The AMOVA studies were performed with the Arlequin software using the genotype data determined from the predominant T. cruzi clone in each sample, with unknown gametic phase, inferring the haplotypes from the distance matrix using the sum of squared size difference (RST) as molecular distance and a significant P value of genetic differences among population samples of 0.05 (Weir and Cockerham 1984; Excoffier et al. 1992).
Lineage determination of predominant T. cruzi clones
We used the Geneclass programme with the Bayesian method described by Rannala and Mountain (1997) and by Piry et al. (2004), with a cutoff score of 70%. As reference populations of each known T. cruzi lineage, the genotypic data of SCLE10, SCLE11 and MCLE01 markers from several T. cruzi clones published in the literature was used (De Freitas et al. 2006; Venegas et al. 2009b). According to this criterion, 16.7%, 28.1%, 14.3% and 21.9% of samples from the N-B, C-B, N-X and C-X T. cruzi DNA groups, respectively, were not taken into account to assign the corresponding lineages (ESM Table S1).
Results
Study of T. cruzi populations in chronic chagasic patients
In order to determine whether the T. cruzi populations found in non-cardiopathic and cardiopathic chagasic patients are genetically differentiated, 42 or 32 patients from each respective group were studied using the described SCLE10, SCLE11 and MCLE01 microsatellite markers (Oliveira et al. 1998). To achieve this goal, the genotype of predominant clone in each T. cruzi DNA sample was determined (“Materials and methods” and ESM Table S1). Other interesting information that can be obtained directly from ESM Table S1 is the efficiency of each marker and the multiclonality of each T. cruzi sample. The least efficient marker either in blood or triatomine dejection samples was SCLE10, with the lowest detection percentage in blood samples (Table S2). SCLE11 and MCLE01 showed efficiencies which ranged from 76.2% to 93.3%.
Multiclonality estimation
The minimal number of T. cruzi clones estimated in each blood or dejection sample is shown in ESM Table S1 and Fig. S1. As can be seen, the majority of samples (50–60%) are composed of only one T. cruzi clone (uniclonal) either in blood or dejection samples (Fig. S1). A minimum of two or three T. cruzi clones were found in 10% and 25% of samples, respectively. There were a minimum of four T. cruzi clones in only 2.2% of the blood samples from the non-cardiopathic chagasic group (N-B). No statistically significant differences were found between sample groups.
Allele frequency distribution of microsatellite markers in four T. cruzi DNA sample groups
Figure 1 shows the allele frequencies of the four T. cruzi DNA sample groups, N-B, C-B, N-X and C-X, described before. Short alleles ranging from 85 to 141 bp were detected only in the xenodiagnosis samples with the SCLE10 marker (Fig. 1a). A similar situation occurred with the MCLE01 marker; the short alleles between 92 and 120 bp were only detected in the xenodiagnosis samples (Fig. 1c). With the SCLE11 marker, short alleles (between 95 and 103 bp) were detected only in the blood samples (Fig. 1b).
Allele frequencies of microsatellite markers detected in predominant T. cruzi clones found in four sample groups: peripheral blood from a group of 42 non-cardiopathic (N-B) and 32 cardiopathic (C-B) chagasic patients. The non-cardiopathic and cardiopathic groups were studied by xenodiagnosis. The T. infestans dejections from the corresponding xenodiagnosis of 42 non-cardiopathic (N-X) and 32 cardiopathic (C-X) chagasic patients were analysed. a–c Analysis with the microsatellite markers: SCLE10, SCLE11 and MCLE01, respectively
Analysis of population differentiation of four T. cruzi sample groups based on SCLE10, SCLE11 and MCLE01 genotypes of predominant clones
In order to determine if genetic differentiation exists between non-cardiopathic and cardiopathic predominant T. cruzi populations and to investigate whether the human and triatomine hosts influence these T. cruzi populations, genotype analyses were performed on the N-B, C-B, N-X and C-X T. cruzi sample groups based on three approaches: locus by locus using the exact G test, the Fisher method across the three loci and AMOVA. The first and the second ones were performed using the Genepop programme (Raymond and Rousset 1995a, b) and the third one using the Arlequin programme (Schneider et al. 2000).
Significant genotype differentiation was found comparing non-cardiopathic and cardiopathic groups only using SCLE11 marker in the blood samples (pair N-B/C-B); the result was not confirmed by the Fisher method across three loci (Table 1).
Significant genotype differentiation was found comparing blood with xenodiagnosis T. cruzi samples both using the locus-by-locus approach and the Fisher method (pairs N-B/N-X or C-B/C-X); in both cases, the P values were lower than 0.05 (Table 1).
To confirm or refute the above results, the third approach, AMOVA, was performed. The first structure tested considered two groups: one composed of the N-B and C-B T. cruzi samples and a second composed of the N-X and C-X T. cruzi samples. As can be seen in Table 2, among these two groups, there is a high percentage of variation, 54.3%, whilst the variation among genotypes within each group was very low (−0.16%). As displayed below Table 2, the corresponding variant component of 54.3% was significant.
The AMOVA considering only two T. cruzi samples, N-B and C-B or N-X and C-X, to further investigate the genetic differentiation among non-cardiopathic and cardiopathic T. cruzi genotypes was performed, but no significant structure was detected (data not shown).
Study of lineages of T. cruzi sample groups
Lineage distribution of predominant T. cruzi clones was studied in the four N-B, C-B, N-X and C-X T. cruzi sample groups. The Geneclass software and genotypic data from described T. cruzi clones were used to determine T. cruzi lineages as described in “Materials and methods”.
The lineage distribution of predominant clones in the four N-B, C-B, N-X and C-X T. cruzi sample groups is shown in Table 3 and ESM Fig. S2. As can be observed, the most frequent lineage in all groups was TcI, 50–60% of samples. No clear differences can be observed between non-cardiopathic and cardiopathic T. cruzi sample groups, but there was a slightly higher but not significant percentage of TcI in non-cardiopathic group either in blood (N-B and C-B) or xenodiagnosis (N-X and C-X) T. cruzi sample groups. Interestingly, the TcIII lineage was more frequently detected in xenodiagnosis groups (N-X or C-X); this was statistically significant (Table 3 and ESM Fig. S2). Finally, the hybrid lineage (TcV or TcVI) only was found in one of the non-cardiopathic blood (N-B) samples.
Discussion
Two main contrasting hypotheses have been proposed to explain the pathogenesis of Chagas disease: one based on the host immune response and the other on the genetic characteristics of the parasite. The first, the autoimmune hypothesis, proposes that the humoral and/or cellular immune response against the parasite produces an inflammatory effect on the tissues, destroying them and provoking a functional failure which in the heart is expressed as myocarditis and loss of conductivity, which, in some cases, might led to death (Acosta and Santos-Buch 1985; Higuchi et al. 2003). These autoimmune hypotheses suggest that the inflammatory effect may remain even without the presence of the parasite (Higuchi et al. 2003).
The other hypothesis states that the genetic characteristics of the T. cruzi clones infecting the patient determine the fate of Chagas disease pathogenesis and their clinical forms (Zingales et al. 1998; Tibayrenc and Ayala 1999; Coura et al. 2002). This latter hypothesis has been suggested by the dissimilar prevalence of Chagas manifestations which have been found in different regions of Latin America, in parallel with the high genetic heterogeneity of the T. cruzi taxon in which a differential distribution of the main lineages associated either with the different transmission cycles and/or mammalian reservoirs is also found (Miles et al. 1981; Zingales et al. 1998, 1999; Brisse et al. 2000; Coura et al. 2002; Buscaglia and Di Noia 2003).
The first aim of the present work was to investigate the distribution of multiclonality in non-cardiopathic and cardiopathic chagasic patients and in blood and xenodiagnosis samples. Stocks of T. cruzi are obtained normally by cultivating parasitic isolates of triatomine vectors or host mammals. The patients in endemic areas may be infected by multiple contacts with different triatomines, and these may also have fed on different infected individuals. This promiscuity causes the formation of multiclonal populations in host vectors, leading to the isolation of multiclonal stocks that grow in culture. In fact, many molecular studies based on zymodemes, esquizodemes, PCR and RAPD have revealed that the stocks of T. cruzi are multiclonal (Macedo et al. 1992; Torres et al. 2004; Rozas et al. 2007). Our results showed no significant differences among frequency of multiclonality between non-cardiopathic and cardiopathic chagasic groups either in peripheral blood or xenodiagnosis samples. One possible explanation for this result is that the spectrum of clones of T. cruzi present in xenodiagnosis samples was previously reduced in the human host, as suggested by the literature (Oliveira et al. 1998), minimizing the quantitative differences between the T. cruzi populations present in both hosts. The high percentage of uniclonal samples (composed only of one clone) observed in the present work, either in blood or dejection samples (50–60%), is in accordance with this idea.
The second objective was to determine whether the T. cruzi populations found in the non-cardiopathic group were genetically different from those found in the cardiopathic group. Three approaches were used: a locus-by-locus analysis, a Fisher method across three loci and the method of AMOVA. The first and the second approaches were conducted using the Genepop programme (Raymond and Rousset 1995a, b). The third approach, AMOVA, was performed using the Arlequin software (Schneider et al. 2000). In all approaches, the SCLE10, SCLE11 and MCLE01 markers described were used (Oliveira et al. 1998). In addition, we only considered the predominant T. cruzi clone present in a sample because it is possible to determine the genotype of more than one locus simultaneously, as was done in the study of other parasites using microsatellite markers (Anderson et al. 1999). Only in the first approach and using the SCLE11 locus was a significant differentiation found among non-cardiopathic and cardiopathic groups analysing blood samples (pair N-B/C-B), which was not confirmed either by the Fisher or the AMOVA methods. The most likely explanation might be due to experimental error produced by single-locus detection. However, we cannot exclude that small genetic differentiation exist between T. cruzi clones present in non-cardiophatic and cardiophatic chagasic patients, as we have found early using another method (Venegas et al. 2009a). Further works must be performed to resolve this important question.
The third objective of the present work was to confirm previous results which show that the kind of host, vertebrate or invertebrate, exerts marked differential selection on T. cruzi clones, which has been observed using schizodemes (Morel et al. 1980), microsatellite markers (Oliveira et al. 1998) and kDNA probes (Coronado et al. 2006: Venegas et al. 2009a). To do this, analyses were performed to compare T. cruzi populations from peripheral blood or dejections of T. infestans fed on the blood of the same patients (xenodiagnosis). With the three approaches mentioned before, locus by locus, Fisher across three loci and AMOVA, a clear significant genetic differentiation was detected between T. cruzi groups. It is important to mention that all these results could not due to the low efficiency found with SCLE10 in blood samples because either with SCLE11 or MCLE01 (which showed similar efficiencies), a significant genetic differentiation among blood and xenodiagnosis samples was also found. These results confirm the mentioned literature evidence which indicates that immune response of the mammalian host against the infecting parasite tends to reduce the spectrum of circulating T. cruzi clones, changing the relative abundance of different lineages. In contrast, the intestine of insect vector offers a more permissive environment, allowing the amplification both minor and predominant T. cruzi clones (Oliveira et al. 1998; Coronado et al. 2006; Venegas et al. 2009a).
The fourth objective of our work was to determine the lineage distribution, TcI-TcVI according to the recommended nomenclature (Zingales et al. 2009), of T. cruzi predominant clones found in non-cardiopathic and cardiopathic chagasic patients in their respective blood and xenodiagnosis samples. The results were only partially concordant with previous reports (Coronado et al. 2006; Venegas et al. 2009a). In the present study, the most prevalent lineage was TcI, both in peripheral blood and xenodiagnosis samples, with percentages between 50% and 60%. Similarly, Coronado et al. (2006) found that TcI lineage was the most prevalent in humans, but in xenodiagnosis was TcV (DTU IId; Coronado et al. 2006); this last lineage was not detected in the present xenodiagnosis samples.
In the previous study of our group, which used cloned highly variable regions of minicircles (HVRm) as probes, the most frequent lineages found in blood samples were TcII and hybrid lineages TcV and/or TcVI (Venegas et al. 2009a; Zingales et al. 2009). However, in the xenodiagnosis samples, similar as was detected in the present work, TcI also was prevalent together with the TcII lineage. These differences with the previous reports could only partly be explained due to experimental error in the assignation of the lineages, particularly considering that the cutoff and confidence score was not very high (70%) to avoid the loss of too many samples. Other possible explanations could be due to different kinds of molecular markers used in the three studies and the fact that the two earlier studies did not distinguish among predominant clones present in multiclonal T. cruzi samples (Coronado et al. 2006; Venegas et al. 2009a). Finally, in the present work, a greater frequency of the TcIII (DTU IIc) lineage was found in xenodiagnosis than in blood sample groups, which was not studied earlier in similar works (Coronado et al. 2006; Venegas et al. 2009a). Interestingly, the importance of this lineage in vertebrates and invertebrates hosts is only recently being evaluated (Llewelyn et al. 2009b).
Concluding statements
In general, the results do not support the hypothesis that the genetic characteristics of the T. cruzi populations are the determining factors that cause or induce the development of chronic chagasic cardiomyopathy. On the other hand, the results are in agreement with the idea that the type of host (vertebrate or invertebrate) plays a crucial role in the selection of T. cruzi populations. It will be very relevant to confirm these results with a larger number of loci as well as patients from other regions of our country to determine if it is a general phenomenon of the chronic chagasic cardiomyopathy in Chile or just a local phenomenon of patients from some regions.
References
Acosta AM, Santos-Buch CA (1985) Autoimmune myocarditis induced by Trypanosoma cruzi. Lab Invest 71:1255–1261
Anderson TJC, Su X-Z, Bockarie M, Lagog M, Day KP (1999) Twelve microsatellite markers for characterization of Plasmodium falciparum from finger-prick blood samples. Parasitology 119:113–125
Apt W, Aguilera X, Arribada A, Perez C, Miranda C, Sanchez G, Zulantay I, Cortes P, Rodriguez J, Juri D (1998) Treatment of chronic Chagas’ disease with itraconazole and allopurinol. Am J Trop Med Hyg 59:133–138
Apt W, Arribada A, Zulantay I, Sánchez G, Vargas SL, Rodríguez J (2003) Itraconazole or allopurinol in the treatment of chronic American trypanosomiasis: the regression and prevention of electrocardiographic abnormalities during 9 years of follow-up. Ann Trop Med Parasitol 97:23–29
Breniere SF, Bosseno MF, Revollo S, Rivera MT, Tibayrenc M (1992) Direct identification of Trypanosoma cruzi natural clones in vectors and host blood by PCR technique. Am J Trop Med Hyg 46:335–341
Breniere SF, Bosseno MF, Telleria J, Bastrenta B, Yaksic N, Noireau F, Alcazar JL, Barnabé C, Wincker P, Tibayrenc M (1998) Different behavior of two Trypanosoma cruzi major clones: transmission and circulation in young Bolivian patients. Exp Parasitol 89:285–295
Brisse S, Barnabé C, Tibayrenc M (2000) Identification of six Trypanosoma cruzi phylogenetic lineages by random amplified polymorphic DNA and multilocus enzyme electrophoresis. Int J Parasitol 30:35–44
Buscaglia CA, Di Noia JM (2003) Trypanosoma cruzi clonal diversity and the epidemiology of Chagas’ disease. Microbes Infect 5:419–427
Coronado X, Zulantay I, Albrecht H, Rozas M, Apt W, Ortiz S, Rodríguez J, Sánchez G, Solari A (2006) Variation in Trypanosoma cruzi clonal composition detected in blood patients and xenodiagnosis triatomines: implications in the molecular epidemiology of Chile. Am J Trop Med Hyg 74:1008–1012
Coura J, Junqueira A, Fernandes O, Valente S, Miles M (2002) Emerging Chagas’ disease in Amazonian Brazil. Trends Parasitol 18:171–176
Deane MP, Soussa MA, Preira NM, Goncalves AM, Momen H, Morel CM (1984) Trypanosoma cruzi: inoculations schedules and reisolation methods select individual strains from doubly infected mice, as demonstrated by schizodeme and zymodeme analysis. J Protozool 231:2765–280
De Freitas JM, Augusto-Pinto L, Pimental JR, Bastos-Rodrigues L, Vanessa F, Gonçalves VF, Teixeira SMR, Chiari E, Junqueira ACV, Fernández O, Macedo AM, Machado CR, Pena SDJ (2006) Ancestral genomes, sex, and the population structure of Trypanosoma cruzi. PLoS Pathog 2:0226–0235
Excoffier L, Smouse P, Quattro J (1992) Analysis of molecular variance inferred from metric distance among DNA haplotypes. Application to human mitochondrial restriction data. Genetics 131:479–491
Higuchi ML, Benvenuti LA, Reis MM, Metzger M (2003) Pathophysiology of the heart in Chagas’ disease: current status and new developments. Cardiovasc Res 60:96–107
Llewelyn MS, Miles MA, Carrasco HJ, Lewis MD, Yeo M, Segobia M, Vargas J, Torrico F, Diosque P, Valente V, Valente S, Gaunt MW (2009a) Genome-scale multilocus microsatellite typing of Trypanosoma cruzi discrete typing unit reveals phylogeographic structure and specific genotypes linked to human infection. PLoS Pathog 5:1–9
Llewelyn MS, Lewis MD, Acosta N, Yeo M, Carrasco HJ, Vargas J, Torrico F, Miles MA, Gaunt MW (2009b) Trypanosoma cruzi IIc: phylogenetic and phylogeographic insights from sequence and microsatellite analysis and potential impact on emergent Chagas disease. Plos Neglected Tropical Diseases 3:1–10
Macedo AM, Martins SM, Chiari E, Pena SDJ (1992) DNA fingerprinting of Trypanosoma cruzi: a new tool for characterization of strains and clones. Mol Biochem Parasitol 55:147–154
Macedo AM, Pimenta J, Aguiar RS, Melo AIR, Chiari E, Zingales B, Pena SDJ, Oliveira RP (2001) Usefulness of microsatelites typing in population genetic studies of Trypanosoma cruzi. Mem Inst Oswaldo Cruz 96:407–413
Macedo AM, Oliveira RP, Pena SDJ (2002) Chagas disease: role of parasite genetic variation in pathogenesis. Expert Rev Mol Med 4:1–16
Miles M, Cedillos R, Povoa M, de Souza A, Prata A, Macedo V (1981) Do radically dissimilar Trypanosoma cruzi strains (zymodemes) cause Venezuelan and Brazilian forms of Chagas’ disease? Lancet 1:1338–1340
Morel C, Chiari E, Plessmann Camargo E, Mattei DM, Romanha AJ, Simpsont L (1980) Strains and clones of Trypanosoma cruzi can be characterized by pattern of restriction endonuclease products of kinetoplast DNA minicircles. Proc Natl Acad Sci USA 77:6810–6814
Oliveira RP, Broude NE, Macedo AM, Cantor CR, Smith CL, Pena SDJ (1998) Probing the genetic population structure of Trypanosoma cruzi with polymorphic microsatellites. Proc Natl Acad Sci USA 95:3776–3780
Pereira Silva C, Del Carlo CH, de Oliveira Junior MT, Scipioni A, Strunz-Cessaro C, Franchini JA, Pereira AC (2008) Why do patients with chagasic cardiomyopathy have worse outcomes than those with non-chagasic cardiomiopathy. Arq Bras Cardiol 91:358–362
Piry S, Alapetite A, Cornuet J-M, Paetkau D, Baudouin L, Estoup A (2004) GeneClass2: a software for genetic assignment and first-generation migrant detection. J Hered 95:536–539
Rannala B, Mountain JL (1997) Detecting immigration by using multilocus genotypes. Proc Natl Acad Sci USA 94:9197–9201
Rozas M, Botto-Mahan C, Coronado X, Ortiz S, Cattan PE, Solari A (2007) Coexistence of Trypanosoma cruzi genotypes in wild and periodomestic mammals in Chile. Am J Trop Med Hyg 77:647–653
Raymond M, Rousset F (1995a) An exact test for population differentiation. Evolution 49:1283–1286
Raymond M, Rousset F (1995b) GENEPOP version 1.2: population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rodrigues Coura J, Pinto Dias JC (2009) Epidemiology, control and surveillance of Chagas disease 100 years after its discovery. Mem Inst Oswaldo Cruz 104:31–40
Schenone H (1999) Xenodiagnosis. Mem Inst Oswaldo Cruz 94:289–294
Schneider S, Roessli D, Excoffier L (2000) Arlequin: a software for population genetic data. Genetics and Biometry Laboratory, University of Geneva, Switzerland
Solari A, Campillay R, Ortiz S, Wallace A (2001) Identification of Trypanosoma cruzi genotypes circulating in Chilean chagasic patients. Exp Parasitol 97:226–233
Tibayrenc M, Ayala F (1988) Isoenzyme variability in T. cruzi, the agent of Chagas disease: genetical, taxonomical and epidemiological significance. Evolution 42:277–292
Tibayrenc M, Ayala JF (1991) Towards a population genetic of microorganisms: the clonal theory of parasitic protozoa. Parasitol Today 7:228–232
Tibayrenc M, Ayala F (1999) Evolutionary genetics of Trypanosoma and Leishmania. Microbes Infect 1:465–472
Tibayrenc M, Ward P, Moya A, Ayala FJ (1986) Natural populations of Trypanosma cruzi, the agent of Chagas disease, have a complex multiclonal structure. Proc Natl Acad Sci USA 83:115–119
Torres JP, Ortiz S, Muñoz S, Solari A (2004) Trypanosoma cruzi isolates from Chile are heterogeneous and composed of mixed populations when characterized by schizodeme and Southern analyses. Parasitology 128:161–168
Venegas J, Coñoepan W, Pichuantes S, Miranda S, Apt W, Arribada A, Zulantay I, Coronado X, Rodriguez J, Reyes E, Solari A, Sánchez G (2009a) Differential distribution of Trypanosama cruzi clones in human chronic cardiopathic and non-cardiopathic individuals. Acta Trop 109:187–193
Venegas J, Coñoepan W, Sergio Pichuantes S, Miranda S, Jercic MI, Gajardo M, Sánchez G (2009b) Phylogenetic analysis of microsatellite markers further supports the two hybridization events hypothesis as the origin of the Trypanosoma cruzi lineages. Parasitol Res 105:191–199
Weir BS, Cockerham CC (1984) Estimating the F-statistics for the analysis of population structure. Evolution 38:1358–1370
Zingales B, Souto BR, Mangia R, Lisboa C, Campbell D, Coura J, Jansen A, Fernandes O (1998) Molecular epidemiology of American trypanosomiasis in Brazil based on dimorphisms of rRNA and mini-exon gene sequences. Int J Parasitol 28:105–112
Zingales B, Stolf B, Souto R, Fernandes O, Briones M (1999) Epidemiology, biochemistry and evolution of Trypanosoma cruzi lineages based on ribosomal RNA sequences. Mem Inst Oswaldo Cruz 94:159–164
Zingales B, Andrade SG, Briones MRS, Campbell DA, Chiari E, Fernandes O, Guhl F, Lages-Silva E, Macedo AM, Machado CR, Miles MA, Romanha AJ, Sturm NR, Tibayrenc M, Schijman AG (2009) A new consensus for Trypanosoma cruzi intraspecific nomenclature: second revision meeting recommends TcI to TcVI. Mem Inst Oswaldo Cruz 104:1051–1054
Zulantay I, Honores P, Sánchez G, Apt W, Ortiz S, Osuna A, Rojas A, López B, Solari A (2004) Use of polymerase chain reaction and hybridization assay to detect Trypanosoma cruzi in chronic chagasic patients treated with itraconazole and allopurinol. Diagn Microbiol Infect Dis 48:253–257
Zulantay I, Apt W, Gil LC, Rocha C, Mundaca K, Solari A, Sánchez G, Rodríguez C, Martinez G, De Pablos LM, Sandobal L, Rodríguez J, Vilchez S, Osuna A (2007) The PCR based-detection of Trypanosoma cruzi in the faeces of Triatoma infestans fed on patients with chronic American trypanosomiasis gives higher sensitivity and a quieker result than routine xenodiagnosis. Ann Trop Med Parasitol 101:673–679
Acknowledgements
This study was supported by FONDECYT, nos. 1040731 and 1070837. We would like to thank Prof. Jorge Rodriguez at the Escuela de Salud Pública, Facultad de Medicina, Universidad de Chile for help with the statistical analysis of the T. cruzi lineages, shown in Table 3.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary materials
Below is the link to the electronic supplementary material.
Fig. S1
Frequency of multiclonality based on Trypanosoma cruzi microsatellite markers SCLE10, SCLE11 and MCLE01in four sample groups. N-B and C-B: groups of blood samples from 42 non-cardiopathic or 32 cardiopathic chagasic patients, respectively. N-X and C-X: groups of dejection samples from Triatoma infestans fed on the same 42 non-cardiopathic or 32 cardiopathic chagasic patients, respectively (xenodiagnosis, see “Materials and methods”). Numbers on the left represent the minimal number of T. cruzi clones detected in each sample (DOC 24.5 kb)
Fig. S2
Lineage frequency of predominant Trypanosoma cruzi clones found in cardiopathic and non cardiopathic chagasic patients. N-B and C-B: groups of blood samples from 42 non-cardiopathic or 32 cardiopathic chagasic patients, respectively. N-X and C-X: groups of dejection samples from Triatoma infestans fed on the same 42 non-cardiopathic or 32 cardiopathic chagasic patients, respectively (xenodiagnosis, see “Materials and methods”). Lineage nomenclature according to Zingales et al. (2009) (DOC 24.5 kb)
Table S1
Genetic patterns of Trypanosoma cruzi clones found in peripheral blood and xenodiagnosis of chagasic patients determined by microsatellite markers SCLE10, SCLE11 and MCLE01 (DOC 709 kb)
Table S2
Efficiency of T. cruzi microsatellite markers SCLE10, SCL11 and MCLE01 in cardiopathic and non-cardiopathic chagasic individuals (DOC 27.0 kb)
Rights and permissions
About this article
Cite this article
Venegas, J., Miranda, S., Coñoepan, W. et al. Microsatellite marker analysis shows differentiation among Trypanosoma cruzi populations of peripheral blood and dejections of Triatoma infestans fed on the same chronic chagasic patients. Parasitol Res 107, 855–863 (2010). https://doi.org/10.1007/s00436-010-1939-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-010-1939-2