Abstract
Main conclusion
Plants develop both short-term and transgenerational memory of drought stress through epigenetic regulation of transcription for a better response to subsequent exposure.
Abstract
Recurrent spells of droughts are more common than a single drought, with intermittent moist recovery intervals. While the detrimental effects of the first drought on plant structure and physiology are unavoidable, if survived, plants can memorize the first drought to present a more robust response to the following droughts. This includes a partial stomatal opening in the watered recovery interval, higher levels of osmoprotectants and ABA, and attenuation of photosynthesis in the subsequent exposure. Short-term drought memory is regulated by ABA and other phytohormone signaling with transcriptional memory behavior in various genes. High levels of methylated histones are deposited at the drought-tolerance genes. During the recovery interval, the RNA polymerase is stalled to be activated by a pause-breaking factor in the subsequent drought. Drought leads to DNA demethylation near drought-response genes, with genetic control of the process. Progenies of the drought-exposed plants can better adapt to drought owing to the inheritance of particular methylation patterns. However, a prolonged watered recovery interval leads to loss of drought memory, mediated by certain demethylases and chromatin accessibility factors. Small RNAs act as critical regulators of drought memory by altering transcript levels of drought-responsive target genes. Further studies in the future will throw more light on the genetic control of drought memory and the interplay of genetic and epigenetic factors in its inheritance. Plants from extreme environments can give queues to understanding robust memory responses at the ecosystem level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
By the year 2050, the world population will reach near 10 billion. On the other hand, agricultural production may not be sufficient to feed the growing billions due to global warming and increased frequencies of drought and desertification (Salinger 2005). Drought will complicate matters further by reducing the nutrient availability and altering the patterns of pest and disease infestations of plants. It is estimated that India and the USA currently experience a 15–20% yield loss in agriculture due to drought, projected to increase in the future (Leng and Hall 2019). Severe drought usually causes damage to a plant, preventing reproduction, causing senescence, and ultimately leading to death. However, a short-term moderate drought may permit plant recovery upon stress withdrawal. Sessile plants have evolved several adaptive strategies to survive droughts, viz., drought avoidance with deeper roots, succulent stems, stomatal closure at high water potentials, high tissue elasticity, and crassulacean acid metabolism (CAM) mode of photosynthesis, drought endurance by leaf shedding, and drought tolerance by stomatal closure at low water potentials, low hydraulic conductance of xylem, leaf size reduction and accumulation of osmoprotectants. In addition, some ephemerals like Arabidopsis bypass stress altogether by temporal control of their life cycles (Ward 2016).
But the catch is that single stress events are rare. Plants practically experience frequent spells of drought, often with intermittent recovery intervals. In such a fluctuating stress scenario, the survival of plants critically depends on the ability to “remember” or “learn” and “recall” the past events of drought, altering its physiology to present a more rapid and robust response to the subsequent drought. This adaptive process is designated by priming, memory, and acclimation. It is perplexing that plants have “memory,” usually considered a monopoly of higher animals having a centralized “brain” and complex neuronal networks. However, this assumption is entirely wrong, as all living beings, including plants, are equipped with memorization capacity, the basis of adaptive evolution (Galviz et al. 2020).
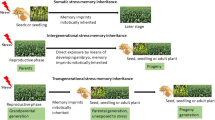
Figure 1 describes the various types of drought memory in plants. Plants remember exposure to drought for several days to weeks. This is due to transcriptomic and metabolomic alterations within the somatic cells and is effective within the same generation. Accumulation of specific dormant signaling molecules like transcription factors (TFs) activated upon the subsequent stress exposure has been found for short-term memory of drought. These molecular events are reflected in physiological events like partial stomatal closure and slowing down photosynthesis as preparatory strategies for the next drought. However, the ultimate level of control of various genes for drought memory lies in the epigenetic regulation of the chromatin, which leads to both short-term memories and the passage of memory to future generations through germ cells developing during the drought exposure. In this review, we look into our current understanding of the molecular mechanisms of plant memory to drought—both in the short term and across generations. We explore gaps in our current knowledge of plant drought memory and promising areas of future explorations and applications for crop improvement.
Drought memory in plants. Severe drought may kill a plant, but a mild drought or a drought for a short duration can trigger short-term memory, usually established by transcriptional training or metabolic reprogramming, leading to survival under a subsequent drought. Sometimes, the memory may be reset or erased upon a prolonged watered recovery interval, leading to drought sensitivity in the subsequent exposure. The memory of stressed plants is carried over to the next generation, termed transgenerational memory, which is usually attributed to epigenetic changes like DNA methylations. The persistence of memory in two or more generations is intergenerational memory
Transcriptional memory of drought
To understand how plants remember drought in the short term within a single generation, most researchers followed a simple experimental setup, wherein watered plants (W) were given a first drought treatment (D1) followed by a watered recovery interval (R1). Subsequently, more droughts and recovery intervals followed (i.e., D2, R2, D3, etc.). Under these conditions, researchers carried out transcriptome analyses to identify the expression patterns of different genes, revealing four distinct types of memory (Ding et al. 2013). Those genes that exhibited progressively higher expression levels under subsequent droughts were designated as [+/+]. Genes belonging to abscisic acid (ABA)-dependent signaling, TFs, late embryogenesis abundant (LEA) genes, lipid transfer proteins, chaperones, ion transporters, and membrane protectants belonged to this category. A higher expression of these genes in D2 signifies their higher protective functions in the subsequent drought. Genes for ribosome assembly and protein synthesis, and the chlorophyll-binding light-harvesting protein complex of photosystem II (PSII), were progressively downregulated in a [−/−] fashion, signifying attenuation of chloroplast functions. The third class of memory genes included other chloroplast membrane proteins and those belonging to the electron transport chain, which, although downregulated in D1, had increased levels in D2, exhibiting a [−/+] pattern. This signifies the restoration of photosynthetic functions under subsequent drought. The fourth category of transcriptional memory behavior was of the [+/−] type, including genes of phytohormone signaling, ion transporters, and aquaporins, whose expression gets induced in D1 but reduces in D2, possibly functioning to readjust ion homeostasis under re-iterated drought (Ding et al. 2013).
Alteration of photosynthesis under recurrent droughts
The exposure of the plant to the first drought causes an increase in the drought stress hormone ABA leading to stomatal closure, reduced carbon dioxide intake, decreased photosynthesis capacity, and photoinhibition of PS. As a result, accumulating energy in the chloroplast leads to reactive oxygen species (ROS) production, causing cellular damage and further degradation of photosynthetic pigments. To counter this, plants exposed to drought reduce the light-harvesting complex of PSII to minimize the energy accumulation and resultant ROS in the chloroplast (Fleta-Soriano and Munné-Bosch 2016; Godwin and Farrona 2020). Also, as an immediate measure to reduce photosystem damage, a drop in photosynthesis is accompanied by increased heat dissipation and chlorophyll fluorescence. In sugar beet, photosynthesis was drastically reduced in all three cycles of recurrent drought, displaying unexpected increase in chlorophyll fluorescence (Leufen et al. 2016). In potato, the genes encoding chlorophyll a/b binding protein of PSII and proteins involved in chlorophyll biosynthesis had reduced expression in D1. However, the level of these PSII genes increased in D2 through a memory response in both potato and rice (Chen et al. 2020; Auler et al. 2021). In another study in rice, several pigment biosynthesis genes and those encoding photosystem I reaction center proteins were suppressed in D1 but maintained at constant levels in D2. In addition, genes of chloroplast retrograde signaling also displayed memory behavior, suggesting their possible role in drought acclimation (Li et al. 2019).
However, Dipteryx alata, endemic to the Brazilian savanna and adapted to a recurrent drought environment, memorized and acclimated to drought differently, without compromising photosynthesis. Plants exposed to three drought cycles (D3) maintained higher carbon fixation rates than only D1-exposed plants by keeping the stomata open and maintaining higher stomatal conductivity and leaf hydraulic conductance. However, chlorophyll did not degrade in any of the drought cycles. Maintained photosynthesis could fuel cellular defense and respiratory processes in D3 plants, comparable to control levels. In contrast, the D1 plants had higher visible ROS-induced damage due to the stomatal closure, impairment of photosynthesis, and higher respiration rates than control or D3 plants, shunning metabolites from other processes to the Krebs cycle(Alves et al. 2020). Despite having a higher stomatal conductance leading to more water loss, D3 D. alata plants survived drought by increasing the accumulation of solutes for osmotic adjustment(Martorell et al. 2015).
Role of osmolytes under recurrent droughts
Plants can survive droughts for a more extended period by synthesizing and accumulating osmolytes in the cell. These are small-sized, water-soluble neutral molecules, helpful in maintaining positive turgor pressure of the cell, stabilizing the proteins and membranes, and protecting the photosynthesis machinery from ROS (Yancey 2005; Szabados and Savouré 2010; Meena et al. 2019). These include small non-reducing sugars and other amine compounds (Lahuta et al. 2022). Soluble sugars increased along with recurrent drought stresses (Woodruff and Meinzer 2011; Sala et al. 2012; Li and Liu 2016). While a fivefold increase of proline was observed in the D1, it was 50-fold after D2 in pea plants (Lahuta et al. 2022). Expression of the proline biosynthesis gene P5CS was higher in subsequent droughts after acclimation in potato (Chen et al. 2020). Switchgrass exposed to multiple droughts upregulated the trehalose and proline biosynthesis genes. Trehalose plays a vital role in the osmotic adjustment of cells during secondary stress (Zhang et al. 2018a, b). Similarly, the levels of another osmolyte myoinositol increased with drought exposure but decreased after re-watering (Galiano et al. 2017). Plants produce glycine-rich hydrophilic LEA proteins that accumulate in the intracellular spaces to tolerate drought stress by stabilizing the cell membrane and protein structure (Magwanga et al. 2018), a common mechanism to handle dehydration in all life forms (Hundertmark and Hincha 2008). LEA genes were induced to higher levels under D2 in potato, indicating their roles in drought acclimation (Chen et al. 2020).
Role of phytohormones in drought memory
ABA levels increase under drought, playing critical roles in tolerance response. These include regulation of stomatal closure to prevent water loss and triggering signal cascades that lead to the transcriptional activation of downstream genes via the ABA-responsive cis-element (ABRE) in their promoters (Osakabe et al. 2014). The genes downstream to ABA encode proteins that lead to membrane protection and detoxification of ROS and osmolytes, which synergistically protect the plant cell from damage (Todaka et al. 2019). Many ABA-responsive downstream genes, e.g., RD29B, encoding LEA-like proteins acting as warning signals (Msanne et al. 2011) and RAB18, encoding a cell-protective dehydrin (Hallouin et al. 2002) exhibited [+/+] memory that persisted 5–7 days in Arabidopsis and maize (Ding et al. 2013, 2014). Several ABA-responsive LEA-like genes that code for hydrophilic proteins, protecting plants from desiccation tolerance (Hand et al. 2011), had increasing levels of transcripts and proteins in subsequent droughts and recovery intervals in the desiccation-tolerant plant Craterostigma plantagineum (Liu et al. 2019). Interestingly, in an invasive CAM xerophyte, Aptenia cordifolia, and the C4 switchgrass, even higher ABA levels were produced under D2 than in D1, suggesting significant roles of ABA in drought memory (Fleta-Soriano et al. 2015; Zhang et al. 2018a, b). Following ABA levels, the ABA biosynthesis NCED gene family transcripts in switchgrass displayed superinduced [+/+] behavior. In contrast, the ABA catabolism gene CYP703A3 was downregulated in D1 and D2 (Zhang et al. 2018a, b). ABA is primarily synthesized in the roots, which sense drought in the soil and is transported through the xylem to the leaves to induce stomatal closure (Zhu 2002). This is why rootstock genotypes strongly influence re-iterated drought acclimation in grafted citrus plants due to the difference in their ABA synthesis capacities (Neves et al. 2017).
ABA regulates drought memory of stomatal guard cells
The guard cells surrounding the stomatal aperture play critical roles in stomatal closure and display different transcriptional memory than the surrounding mesophyll cells (Virlouvet and Fromm 2015). Stomata close on every drought exposure to minimize water loss, but interestingly, stomatal reopening is partial in R1, showing evidence of drought memory (Fig. 2). Under prolonged water availability, however, this memory is lost. Guard cell memory is dependent on ABA levels as the transpiration rates from stomata are the same in D1 and D2 in the ABA-deficient aba2 mutant (Nambara et al. 1998). A higher expression of NCED3 and AAO3 and resultant higher ABA synthesis in guard cells leads to higher levels of ABA during R1 than the surrounding mesophyll cells. Even though mesophyll cell transcript levels of these genes return to W values during R1, the guard cell-specific transcript levels remain high, explaining partial stomatal closure in R1. On the other hand, higher transcript levels of ABA-dependent TFs regulating stomatal closure (Li et al. 2008) and lower transcript levels of TFs regulating stomatal opening (Oh et al. 2011; Rusconi et al. 2013) in guard cells during R1 bring about the partial stomatal closure. Finally, the downstream ion transporters for stomatal opening have lower transcript levels in D1 in guard cells, decreasing stomatal opening in D1. During R1, their transcript levels return to that in W. On the other hand, a subset of ion transporter genes for stomatal closure has higher R1-expression in guard cells, facilitating partial stomatal closure. Moreover, different ABA-regulated drought-tolerance genes have higher transcript levels during D2 in guard cells. (Virlouvet and Fromm 2015).
Role of abscisic acid and jasmonate in transcriptional drought memory. The early steps of abscisic acid (ABA) and jasmonate (JA) synthesis occur in the chloroplast from the mevalonate and α-linolenate pathways. The first drought exposure (D1) leads to induction in ABA biosynthesis genes NSY and NCED with respect to the watered control condition (W). After a water recovery interval (R1), a second drought (D2) leads to even higher transcript levels in these genes. This is indicative of a superinduced [+/+] memory behavior. Similar memory behavior is observed in the downstream genes of ABA signaling, viz. PP2C and SnRK2. In the absence of ABA, PP2C negatively regulates the signaling Kinase SnRK2. ABA binds to its receptor PYL and the co-receptor PP2C and this removes the negative regulation of PP2C on SnRK2 kinases. Once SnRK2 is activated by phosphorylation, it can activate downstream target proteins. The dynamics of SnRK2-activated ion transporters in the guard cells lead to stomatal closure in D1 and D2, but only partial closure in R1 (see main text). The transcript levels of the transcription factors (TFs) regulated by ABA via SnRK2 are reduced in D2 than in D1 [+/−]. These TFs binds to the ABA-responsive element (ABRE) in the promoter of +/+ downstream drought-response genes, e.g., RAB18, RD29B, LEA, etc. whose gene products lead to protection of the cell under drought. In some plant species, ABA levels are reduced in D2, concomitant with the [+/−] memory behavior of NCED and SnRK2 (shown in blue). In addition, drought-induced changes in the chromatin remodellers ATX1, SNF/BRM and SWI3B lead to their activation or abolition of inhibitory activity, causing activation of the transcription of NCED, ABI5 SnRK2 genes, respectively. The upstream JA biosynthesis genes follow [+/−] behavior, but the downstream genes, whose products are localized in the peroxisome, observe [+/+] patterns. The net effect is the increased levels of JA in D1 but the reduction in D2. The TF MYC2 is positioned at the crossroads of both ABA and JA-mediated signaling. MYC2 follows a [+/−] memory pattern or progressive downregulation in both D1 and D2 [−/−] (shown in purple) and is regulated by JA through the JAR, COI-1 and JAZ-mediated pathway. ABA also regulates MYC2 via PYL6 and by another unknown transcription factor which is supressed in D1 but induced in D2 [−/+] (shown in green). ABA triggers JA biosynthesis via MYC2, regulating the upstream biosynthesis genes of JA, AOC and AOS. The expression of JA biosynthesis genes of the α-linolenate pathway in the chloroplast are induced in D1 but supressed in D2 [+/−], or remain at constant levels in D2 after their initial induction in D1 [+/=], while downstream JA biosynthesis genes in the peroxisome follow [+/+] patterns, with a net effect of increased JA levels in D1 followed by a decrease in D2. Again, JA is also found to trigger ABA biosynthesis. JA also causes recruitment of the transcriptional machinery to the promoters of ABA-responsive genes via MYC2 and mediator protein complexes. But the RNA polymerase II is stalled in R1, marked by Serine 5 phosphorylation of the C-terminal tail of Pol II (Ser5P), and an unknown additional factor produced during D1 but activated only in D2 through an ABA-independent pathway triggers activation of the Pol II in forwarding motion, marked by serine 2 phosphorylation of Pol II C-terminal tail (Ser2P). Direct regulations are shown by bold lines and indirect regulations by dashed lines
ABA regulates signaling and transcription for drought memory
ABA exerts its role in the memory patterns of drought-responsive genes, e.g., RD29B, through the ABRE (Uno et al. 2000). Deleting the ABREs from the promoter led to the abolition of transcriptional memory of RD29B. However, ABRE-flanking sequences have some role in the variation in memory behavior of two genes, both containing ABRE in their promoters (Virlouvet et al. 2014). Analysis of memory patterns in Arabidopsis ABA-signaling mutants suggests that ABA controls drought memory through the SnRK2/ABF/ABRE signaling pathway (Fujii et al. 2009; Ding et al. 2013; Fujita et al. 2013; Karle et al. 2020). Corroborating these results, several SNRK2 orthologs displayed memory behavior in rice and switchgrass, increasing transcripts in D2 and decreasing in R1 (Zhang et al. 2018a, b; Li et al. 2019). On the other hand, the ABA-independent SnRK2s exert opposite repressive roles to transcriptional memory (Fujii et al. 2011; Virlouvet et al. 2014). Protein levels of TFs, ABF3, and ABF4 slightly increased in D2, and after posttranslational modifications, they regulate memory of RD29B. In switchgrass, ABF genes behaved as non-memory gene, i.e., maintained at constant levels after their induction during D1 [+/=] (Zhang et al. 2018a, b). ABA, once bound to its co-receptors PYL and PP2C, prevents the inhibitory action of PP2Cs, leading to the activation of SnRKs (Umezawa et al. 2009; Karle et al. 2020). Three PP2C orthologs behaved as a superinduced memory gene, i.e., with progressively higher expression in D1 and D2 [+/+] in maize, switchgrass, and soybean, while some other orthologs showed induction in D1, but suppression in D2 [+/−] or transcript levels maintained equally in D2 after their induction in D1 [+/=] (Ding et al. 2014; Zhang et al. 2018a, b; Kim et al. 2020). LRR-like kinases of ABA signaling showed [+/−] memory in coffee (Guedes et al. 2018). LRR-proteins interact with heat shock proteins (Lopes-Caitar et al. 2016), which although suppressed in D1, was induced during D2 [−/+] in coffee, indicating their function in the proper folding of proteins during drought acclimation (Guedes et al. 2018). Different ABA-responsive MYB TFs displayed memory under re-iterated drought. In Arabidopsis, MYBs showed [+/+] memory (Ding et al. 2013). In coffee, drought and ABA-suppressed transcriptional activator MYBs displayed [−/+] behavior, while repressor MYBs, showed a [+/−] pattern, being suppressed in D2 (Guedes et al. 2018).
ABA alone is not sufficient for drought memory
Surprisingly, the aba2 mutant retains drought memory, although it fails to produce ABA (Virlouvet et al. 2014). Also, D1 plants cannot trigger memory if treated only with ABA but not D2. These observations, coupled with no observed specific cis-element for transcriptional memory, suggest that although necessary, ABA is insufficient for triggering memory by itself. In Arabidopsis, rice, maize, and soybean, ABA levels increased during D1, D2, D3, etc. but returned to W levels in the water recovery intervals. In these cases, ABA levels in D2 and D3 were not higher than D1 (Ding et al. 2014; Virlouvet et al. 2018; Li et al. 2019; Thuy Quynh Nguyen et al. 2020). This is due to a [+/+] behavior of the ABA catabolism genes CYP707A3 and CYP76C2 in these species (Ding et al. 2014). There was a sharp spike in ABA levels in D1, but ABA levels reduced in the subsequent droughts in rice. Similarly, all ABA-signaling genes, including NCED3 and the rice orthologs of SNRK2s, SAPKs, followed [+/−] behavior, indicating a species-specific alteration in drought memory behavior (Li et al. 2019).
The memory of superinduced genes was lost if the recovery intervals lasted more than seven days, even though ABA levels were retained (Ding et al. 2012). ABA levels were slightly higher than W, but insufficient to maintain the memory in R1, R2, and R3. In this case, neither ABA nor the ABA-dependent TFs (Yoshida et al. 2010) can explain the memory behavior, although essential for the superinduction of memory genes. It is speculated that ABA generated from D1 triggers an unknown factor activated by ABA-independent signaling, which leads to the superinduction of memory genes in D2 (Virlouvet et al. 2014). Even though transcription of the memory genes goes down in R1, R2, and R3 back to W levels, certain epigenetic marks are still maintained, and the RNA polymerase is stalled, only to be re-activated into transcription by some unknown ‘pause-breaking’ factor in D2 (Fig. 2) (Ding et al. 2012).
Jasmonic acid and other phytohormones regulate drought memory
Jasmonic acid (JA) is another phytohormone rapidly produced during drought protecting plants from oxidative damage (Sasaki-Sekimoto et al. 2005; Ding et al. 2013; Wang et al. 2021). JA signals the TF MYC2 through the COI-1-JAR1-JAZ pathway to activate the [+/−] JA-related drought memory genes (Fig. 2) (Ding et al. 2014; Liu et al. 2016). JA accumulates during D1 in most plants but is formed in low levels in R1 and D2, equivalent to W. JA is produced from α-linolenate in chloroplasts, catalyzed by enzymes allene oxide cyclase (AOC) and allene oxide synthase (AOS), which followed [+/−] memory patterns in switchgrass (Zhang et al. 2018a, b), and [+/=] non-memory behavior in maize (Ding et al. 2014). But the peroxisome-localized downstream enzymes of JA biosynthesis were encoded by [+/+] superinduced genes. Although there is a discrepancy between the transcript levels of the downstream biosynthesis genes and the JA levels in D2, unidirectional [+/+], [−/−], and [+/−] memory response of many JA-responsive genes indicates that JA has a vital role in drought memory (Ding et al. 2013). On the other hand, the chloroplast-localized upstream JA biosynthesis genes, AOC and AOS, are regulated by the bHLH TF MYC2 in S1, but not in S2, because MYC2 orthologs follow [+/−] and [−/−] memory patterns. This can partially explain why JA levels reduce in D2 despite the [+/+] behavior of the downstream genes of JA biosynthesis in the peroxisome. Most JA signaling genes, including an AP2/ERF family TF, showed conserved [+/−] memory behavior in most plants (Ding et al. 2014). It is speculated that an unknown ABA-responsive TF activated by drought, possibly having an opposite [−/+] memory, induces MYC2 in D1 but suppresses it in D2, leading to [+/−] memory behavior of most of the JA-related genes (Liu et al. 2016; Avramova 2018). The cross-talk of ABA and JA in drought memory is explained graphically in Fig. 2.
MYC2 is a focal point of the cross-talk between various phytohormone signaling, ABA, JA, salicylate (SA), gibberellic acid (GA), and auxin (Kazan and Manners 2013). ABA triggers the JA biosynthesis genes through MYC2, which physically binds the ABA receptor PYL6 (Adie et al. 2007; Aleman et al. 2016; Avramova 2018). Among other phytohormones, the biosynthesis genes of GA, auxin, ethylene, and zeatin (cytokinin) of potato, switchgrass, and the resurrection plant Boea hygrometrica display superinduced memory behavior (Zhang et al. 2018a, b; Chen et al. 2020; Sun et al. 2021). This coupled with the observed increase of auxin levels (Neves et al. 2017) and reduction in salicylic acid in roots under re-iterated drought (Miura et al. 2013), indicates the roles of a myriad of phytohormones in drought memory. In B. hygrometrica, along with CTK genes encoding cytokinin biosynthesis enzymes, high-affinity nitrate transporters were superinduced in D2 (Sun et al. 2021). Nitrate signals upregulate cytokinin biosynthesis affecting plant development under stress (Sakakibara et al. 2006; Wang et al. 2016). In A. cordifolia, GA levels reduced progressively in D1 and D2, which synergistically with the steady increase in ABA levels in D1 and D2, increase production of ROS for stress signaling without causing photo-oxidative damage (Fleta-Soriano et al. 2015).
Epigenetic regulation of drought memory
Epigenetic modifications include heritable changes in gene expression via chromatin modifications through DNA methylation and histone modifications, e.g., methylation and phosphorylation, and post-transcriptional regulation of RNAs via RNA interference (Hirayama and Shinozaki 2010; Santos et al. 2011). DNA methylation, chromatin modifications, and small interfering RNAs (siRNAs) play vital roles in the epigenetic regulation of genes involved in stress memory. Memory can be inherited to the offspring through mitotic cell divisions, as plants develop germ cells late in their life cycle (Kinoshita and Seki 2014). So, any epigenetic modifications occurring in the juvenile phase can be transferred to the next generations. But maladaptive epigenetic marks at one part of the plant are prevented from getting inherited by future generations if the plant produces seeds many times in its life cycle. Also, seeds arising from different meristems add to the variability of the epialleles (Galviz et al. 2020).
Histone modifications
Histones are essential components of the chromatin structure. Histone variations and posttranslational modifications play crucial roles in chromatin structure and accessibility of the RNA polymerase, regulating the transcriptional status of genes. The linker histone H1 variations play a significant role in chromatin remodeling. The variant H1.3 increases under stress, essential for normal stomatal function and adaptive developmental responses to light and water stress. ABA-mediated drought signaling causes H1.3 to be expressed in stomatal guard cells (Rutowicz et al. 2015).
Plants remember drought by modifying histones associated with numerous drought-induced genes. Enhanced H3K4 methylation marks the recently activated drought and ABA-response genes for enhanced reactivation in response to recurrent droughts (Kim et al. 2008; Zhang et al. 2009; Van et al. 2010; Levine 2011). Under drought stress, H3K4me3 modification redundantly by the methylase ATX1 triggers NCED3 expression for ABA production (Ding et al. 2011). As these epigenetic marks are enriched during D1 and sustained throughout R1, they play a role in transcriptional memory (He and Li 2018).
Interestingly out of the [+/+] Arabidopsis superinduced genes, transcript levels of a subset remained high at R1. But, transcripts of the other subgroup decreased in R1. This is due to subset-specific histone modification by CLF, a subunit of polycomb-group proteins. Although the histone modifications H3K27me3 and H3K4me3 antagonize each other in developmental genes, these modifications were not antagonistic in the drought memory genes. H3K27me3 could not prevent transcription, possibly due to the effects of other activating TFs produced under drought, but limit the full expression potential of memory genes. This suggests that histone modifications have completely different regulatory dynamics between developmental versus memory genes (Liu et al. 2014).
In addition, a linker histone variant H1-S was upregulated by drought through ABA signaling in tomato (Scippa et al. 2004; Chinnusamy and Zhu 2009). The SWI/SNF nucleosome remodeling protein alters gene expression of ABA-signaling genes. In the absence of stress, SWI3B interacts with HAB1, a PP2C inhibiting ABA signaling. The SNF/Brahma chromatin remodeling complexes negatively regulate drought tolerance by repression of ABI5 (Han et al. 2012). Drought stress inhibits SWI/SNF complexes leading to the activation of ABA-regulated stress response genes. Thus, ABA is intricately linked with histone modifications for drought memory (Mehrotra et al. 2014). This relationship of ABA with histone modifications is graphically explained in Fig. 2.
DNA methylation changes for short-term drought memory
DNA methylation at the cytosine residues is an epigenetic mark that regulates gene expression and is one of the means for drought memory to be transmitted to the next (intergenerational memory) or more subsequent generations (transgenerational memory). DNA methylation in plants occurs by various methyltransferases, at CG sequences (gene body methylation), at CHG (H = A, C or T), and methylation de novo at asymmetrical CHH sequences triggered by the RNA-directed DNA methylation (RdDM) pathway. Methylation at the regulatory regions of a gene shuts off gene expression, while demethylation by demethylases turns it on (Zhang et al. 2018a). Drought induces a change in methylation patterns of the genomic DNA in various plant species (Wojtyla et al. 2020). Consistent with DNA hypermethylation under drought (Labra et al. 2002), in drought-tolerant clones of coffee, a methyltransferase showed [−/+] transcriptional memory (Guedes et al. 2018).
By contrast, the promoters of proline biosynthesis genes are demethylated, activating transcription, leading to higher proline contents for enhanced protection under D2 (Zhang et al. 2013). Drought-tolerant rice genotypes were associated with cytosine demethylation, whereas the sensitive genotypes showed hypermethylation (Gayacharan 2013; Kou et al. 2021). DNA methylation in Arabidopsis and rice was found to have a wide variation with the genotype, and the sequence variations within specific methylases and argonautes, regulating this trait, were identified by a genome-wide association study (GWAS) (Kawakatsu et al. 2016; Rajkumar et al. 2020; Zhang et al. 2020). The sensitive genotypes of rice had more differentially methylated regions (DMRs), indicating the role of methylation in the short-term drought memory of rice. Interestingly most of these DMRs were within the CHH sequences and in transposons, showing the most significant role of the RdDM pathway, which affects CHH methylation. However, repeated drought stress led to the stable maintenance of methylation patterns sculpted by the first drought exposure. Besides transposons, memory DMRs were also associated with osmolyte biosynthesis and JA signaling (Li et al. 2019). Cytosine methylation in B. hygrometrica is more robustly maintained than Arabidopsis and other crops. DNA demethylation was found in drought-acclimated plants at all symmetrical sites, viz. CG and CHG, but the asymmetrical CHH sequences remained hypermethylated. DNA demethylation was observed in the promoters of memory genes, guiding alternative splicing under stress, producing raffinose, and maintaining homeostasis of amino acids between the vacuole and cytoplasm, leading to the upregulation of these genes in D2 (Fig. 3) (Sun et al. 2021).
Maintenance of DNA methylation for drought memory. Drought (D1) causes demethylation in most plants' CG and CHH sequence contexts, but in the CG and CHG sequences in Boea hygrometrica. The change in methylation status in the promoter of protein-coding genes leads to their activation causing drought tolerance. A second drought (D2) does not change the methylation status further, which is maintained. The methylation patterns of stressed plants are inherited to the next generation (S1) through the maternal parent. No further change in methylation is observed, and this status is inherited by the second generation of stressed (S2) or non-stressed plants (S1C1). On the other hand, the hypermethylated status of control non-stressed plants of the starting generation is transmitted to subsequent generations C1 and C2. Methylation levels of S1, S2 and S1C1 plants are lower than the control C1 and C2 plants. The role of siRNA-mediated methylation in the CHH context through the RNA-directed DNA methylation pathway is important for the transgenerational drought memory
Apart from the influence of the genotype, drought alone can shape epigenomic diversity. In genetically identical poplar clones grown in varied geographic conditions, variation in DNA methylation due to prior drought exposures was observed. This variation guided differential transcriptomic changes determining the drought adaptability of the poplar clones (Raj et al. 2011). Environmentally-associated differences in methylation (epialleles) have also been identified in Arabidopsis (He et al. 2018).
Role of DNA methylation in transgenerational drought memory
Ganguly et al. (2017) found transgenerational changes in DNA methylation are not correlated with drought memory in Arabidopsis. Sometimes, memory development requires drought exposure for a minimum of two generations (Kron et al. 2008; Pecinka et al. 2012). But, in Polygonum persicaria, the inherited methylation patterns in the progeny of drought-exposed parents determined transgenerational drought memory. In this plant, the progenies of the drought-exposed parents were better equipped to counter drought, possessing longer roots than the progeny of the non-stressed parents, a phenotype abolished by methylation inhibitors. Different genotypes resulted in methylation and memory variations in the progeny, indicating genetic control of the process (Herman and Sultan 2016).
In Arabidopsis, homologous recombination frequency (HRF) and DNA methylation decreased by drought in the progeny of stressed plants (S1) relative to the progeny of the unstressed plants (C1). These changes in HRF were preserved in the next generation of stressed S1 plants (S2) but not in the second-generation progeny of the unstressed S1 plants (S1C1). However, the changes in DNA methylation due to drought stress exposure in the starting generation were maintained equally in all subsequent generations, S1, S2, and S1C1 regulated by RdDM-triggered CHH methylation (explained graphically in Fig. 3) (Boyko et al. 2010; Walter et al. 2011; Luna et al. 2012). Transgenerational memory is passed on to the offspring only from the maternal parent because a DNA glycosylase produced from the pollen vegetative cell and a central cell in the female gametophyte inhibits paternal inheritance (Choi et al. 2002; Park et al. 2020) through demethylation of transposons leading to the production of siRNAs which starts deactivating neighboring paternal genes through RNA-induced transcriptional silencing (Calarco et al. 2012; Zhang et al. 2018a, b; Park et al. 2020).
Non-coding RNAs regulate drought memory
Small non-coding RNAs are also the vehicles that transmit the drought memory to the next generation (Brosnan and Voinnet 2011). Systemic movement of drought-triggered small RNAs via the symplast and vascular tissues to the meristem results in DNA methylation by the RdDM pathway (Melnyk et al. 2011).
Long non-coding RNA
Long non-coding RNAs (lncRNAs) operate as precursors of microRNAs (miRNAs) and other siRNAs or as miRNA target mimics, regulating many biological processes (Kim and Sung 2012). Also, they influence chromatin dynamics and alternative splicing of pre-mRNA (Liu et al. 2015). Antisense lncRNAs were enriched in regions differentially methylated recurrent osmotic stress, suggesting the combined role of DNA methylation and lncRNA-mediated gene regulation for memory (Mozgova et al. 2019). In rice, 6% of lncRNAs show transcriptional drought memory. Among them, TCONS_00028567 and its product miRNA showed strong induction at D2 but reduced levels after R3 and following droughts including ABA and ethylene signaling via SAPK10 (Li et al. 2019). In switchgrass, levels of ABA and trehalose and that of the lncRNAs targeting their biosynthesis increased progressively in D1 and D2. However, ethylene signaling and its regulator lncRNAs were suppressed in D2, promoting plant development and preventing leaf senescence (Zhang et al. 2018a, b). Differentially expressed lncRNAs have also been identified under drought in other species (Urquiaga et al. 2021), wherein they post-transcriptionally regulate several drought-responsive TFs (Suksamran et al. 2020) or behave as target mimics of miRNAs (Shuai et al. 2014). In particular, the identified roles of lncRNAs in the regulation of DNA methylation via the RdDM pathway (Heo and Sung 2011; Yong-Villalobos et al. 2015; Ariel et al. 2020) place them as potent regulators of drought memory (Urquiaga et al. 2021).
Micro RNA
MiRNAs regulate cellular processes of drought adaptation like development, phytohormone signaling, antioxidant defense, osmoprotection, photosynthesis, etc., through the upregulation and downregulation of positive and negative regulatory molecules. MiRNAs participate in stress memory through directed DNA methylation and various histone modifications (Sunkar and Zhu 2004; Sun et al. 2021). MiRNAs associate with Argonaute proteins to target complementary mRNA cleavage or translational inhibition (Gelaw and Sanan-Mishra 2021). MiRNAs amplify the silencing signal by generating siRNAs via RNA-dependent RNA polymerases (Creasey et al. 2014). Several miRNAs are upregulated under drought, targeting drought-responsive TFs (Liu et al. 2008; Jacques et al. 2021). In a unique example, grafting rapeseed scions into a more -tolerant turnip improved its drought memory, possibly due to the movement of miRNAs between the rootstock and scion (Pagliarani et al. 2017; Luo et al. 2020). Several miRNAs revealed distinct patterns of transcriptional memory behavior under repeated drought cycles in coffee and exerted complex regulation with their targets, the memory MYBs (Guedes et al. 2018).
Resetting drought memory
Short-term drought memory is usually reset after a prolonged watered recovery. Like heat stress, autophagy genes may have critical roles in resetting short-term drought memory, possibly by clearing protective gene products like heat shock proteins (Sedaghatmehr et al. 2019). Moreover, since each generation of plants faces stress conditions different from the previous generation, the inherited memory is erased in the new generation in the early stages of development to ensure normal growth of the plant. Although the mechanism of erasing trans-generational drought memory is unknown, a screening of Arabidopsis mutants impaired in resetting stress memory indicated the involvement of chromatin accessibility factors and siRNA-mediated silencing (Yokthongwattana et al. 2010; Zhou et al. 2010; Iwasaki and Paszkowski 2014) in preventing transgenerational memory. However, how these proteins erase the epigenetic marks are not well understood. Histone demethylases may also play some role in resetting drought memory, as an ortholog of this enzyme in barley was induced under drought (Papaefthimiou and Tsaftaris 2012). In the sperm cells, histone methylases are silenced, demethylases are induced, and methylation-resistant histone variants are recruited to the chromatin to re-configure the paternal epigenome (Borg et al. 2020).
Ecosystem-level drought memory
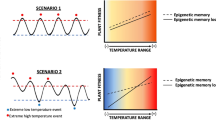
An intriguing question is whether plants remember a drought as an individual or a community. Recent studies hint towards the existence of ecological memory for drought. Entire juniper forests growing for centuries at drier sites of the Tibetan plateau are more susceptible to drought-induced damages than forests growing in wetter regions without prior history of droughts (Fang et al. 2021). This may be because detrimental effects of ecological drought memory last several decades after exposure, as observed in Mediterranean holm oaks (Lopez et al. 2009). Again, positive adaptations of an individual to drought can quickly be passed on to the community through seed dispersal leading to community adaptation to recurrent droughts. In the ecosystem, some plants growing at the edge of the forest or crop field serve as ‘sentinels,’ sending warning signals to their neighbors about the upcoming water deficit scenario (Ribeiro and Torres 2018). The detailed mechanisms of this type of community signaling remain largely unknown. Canarini et al. (2021) found recurrent drought-induced and ecosystem-level memory altering the soil microbiome. While drought can change plant exudates, affecting microbial stress response and survival, on the other hand, rainfall-regulated CO2 emissions from the soil microbes can affect the photosynthesis efficiency of the plant community. Moreover, the epigenetic changes due to drought could be spread in whole plant communities in the ecosystem through seed dispersal from the maternal parent (Walter et al. 2013). Further research is needed in natural ecosystems to understand community drought memory in plants.
The gap in our knowledge and where we are headed
We are just beginning the scientific understanding and appreciation of memory in plants, despite the perplexing absence of a nervous system in plants. The memory of drought and other stresses is of utmost importance for plant survival in the face of erratic precipitation caused due to global climate change. Several mechanisms have been identified that partly explain both short- and long-term drought memory of plants. Some are at the crossroads of other stresses, heat, pathogen attack, and herbivory (Liu et al. 2016).
GWAS is a robust forward genetic tool to identify novel genomic loci regulating complex stress phenotypes, including gene expression-level variation in plants (Kobayashi et al. 2016; Sadhukhan et al. 2017, 2021; Zhu et al. 2022). GWAS on the methylome variation in many Arabidopsis natural accessions under control conditions revealed a strong association of the nucleotide sequence variation within several epigenetic modifiers (Kawakatsu et al. 2016). GWAS on the short-term and transgenerational drought phenotypes, including memory gene expression-level polymorphisms, will be powerful ways to identify new loci regulating drought stress memory.
Plants are more prone to inheriting environment-triggered epimutations due to the late development of germ cells by dedifferentiation of somatic cells under stress (He et al. 2018; Zhang et al. 2020). Moreover, the epigenetic states are not entirely erased from the germline (Minow and Colasanti 2020). But some questions remain. How do epimutations evolve under drought? Are inherited epigenetic marks and DNA polymorphisms associated, and do they co-segregate? Epigenome-wide association studies (EWAS) have recently been reported in plants to identify the epigenetic basis of complex phenotypes (Denkena et al. 2021). EWAS under drought conditions will throw more light on the epigenome evolution for a clear understanding of drought stress memory. On the other hand, the specific mechanisms of epigenetic drift and resetting drought memory in the next generation are unknown, needing future research.
We need to understand plant epigenome evolution influenced by drought to manipulate it to develop drought-resilient crops to meet future demand. Recent breeding programs have partly succeeded in selecting epigenetic states (Raju et al. 2018). Advancement of genome editing, including fusion of an epigenetic modifier with modified CRISPR-associated nuclease (dCas9), can guide favorable chromatin modifications in a gene-specific manner. Recent application of this technology has led to the development of drought-resistant plants without any changes in the nucleotide sequence (Paixão et al. 2019). Thus, epigenome engineering holds immense promise to develop plants with more robust memory responses. Research on plants naturally adapted to extreme environments such as deserts will further unravel novel molecular mechanisms regulating drought memory.
Author contribution statement
AS and SSP wrote the article with inputs from the rest of the authors. AS prepared the figures. All authors read and approved the manuscript.
References
Adie BAT, Pérez-Pérez J, Pérez-Pérez MM, Godoy M, Sánchez-Serrano J-J, Schmelz EA, Solano R (2007) ABA is an essential signal for plant resistance to pathogens affecting JA biosynthesis and the activation of defenses in Arabidopsis. Plant Cell 19:1665–1681. https://doi.org/10.1105/tpc.106.048041
Aleman F, Yazaki J, Lee M et al (2016) An ABA-increased interaction of the PYL6 ABA receptor with MYC2 transcription factor: a putative link of ABA and JA signaling. Sci Rep 6:28941. https://doi.org/10.1038/srep28941
Alves RDFB, Menezes-Silva PE, Sousa LF et al (2020) (2020) Evidence of drought memory in Dipteryx alata indicates differential acclimation of plants to savanna conditions. Sci Rep 101(10):1–16. https://doi.org/10.1038/s41598-020-73423-3
Ariel F, Lucero L, Christ A et al (2020) R-loop mediated trans action of the APOLO long non-coding RNA. Mol Cell 77:1055–1065. https://doi.org/10.1016/j.molcel.2019.12.015
Auler PA, Souza GM, da Silva Engela MRG et al (2021) Stress memory of physiological, biochemical and metabolomic responses in two different rice genotypes under drought stress: the scale matters. Plant Sci 311:110994. https://doi.org/10.1016/J.PLANTSCI.2021.110994
Avramova Z (2018) Defence-related priming and responses to recurring drought: two manifestations of plant transcriptional memory mediated by the ABA and JA signalling pathways. Plant Cell Environ 42:983–997. https://doi.org/10.1111/pce.13458
Borg M, Jacob Y, Susaki D et al (2020) Targeted reprogramming of H3K27me3 resets epigenetic memory in plant paternal chromatin. Nat Cell Biol 22:621–629. https://doi.org/10.1038/s41556-020-0515-y
Boyko A, Blevins T, Yao Y et al (2010) Transgenerational adaptation of Arabidopsis to stress requires DNA methylation and the function of dicer-like proteins. PLoS ONE 5(3):e9514. https://doi.org/10.1371/journal.pone.0009514
Brosnan CA, Voinnet O (2011) Cell-to-cell and long-distance siRNA movement in plants: mechanisms and biological implications. Curr Opin Plant Biol 14(5):580–587. https://doi.org/10.1016/j.pbi.2011.07.011
Buzas DM, Tamada Y, Kurata T (2012) FLC: a hidden Polycomb response element shows up in silence. Plant Cell Physiol 53:785–793. https://doi.org/10.1093/pcp/pcr163
Calarco JP, Borges F, Donoghue MT et al (2012) Reprogramming of DNA methylation in pollen guides epigenetic inheritance via small RNA. Cell 151(1):194–205. https://doi.org/10.1016/j.cell.2012.09.001
Canarini A, Schmidt H, Fuchslueger L et al (2021) Ecological memory of recurrent drought modifies soil processes via changes in soil microbial community. Nat Commun 12(1):1–14. https://doi.org/10.1038/s41467-021-25675-4
Chen Y, Li C, Yi J et al (2020) Transcriptome response to drought, rehydration and re-dehydration in potato. Int J Mol Sci 21:159. https://doi.org/10.3390/ijms21010159
Chinnusamy V, Zhu J-K (2009) Epigenetic regulation of stress responses in plants. Curr Opin Plant Biol 12:133–139. https://doi.org/10.1016/j.pbi.2008.12.006
Choi Y, Gehring M, Johnson L et al (2002) DEMETER, a DNA glycosylase domain protein, is required for endosperm gene imprinting and seed viability in Arabidopsis. Cell 110(1):33–42. https://doi.org/10.1016/s0092-8674(02)00807-3
Creasey KM, Zhai J, Borges F, Van Ex F, Regulski M, Meyers BC, Martienssen RA (2014) miRNAs trigger widespread epigenetically activated siRNAs from transposons in Arabidopsis. Nature 508:411–415. https://doi.org/10.1038/nature13069
de Guedes FAF, Nobres P, Rodrigues Ferreira DC et al (2018) Transcriptional memory contributes to drought tolerance in coffee (Coffea canephora) plants. Environ Exp Bot 147:220–233. https://doi.org/10.1016/j.envexpbot.2017.12.004
de Urquiaga MCO, Thiebaut F, Hemerly AS, Ferreira PCG (2021) From trash to luxury: the potential role of plant lncRNA in DNA methylation during abiotic stress. Front Plant Sci 11:2130. https://doi.org/10.3389/fpls.2020.603246
Denkena J, Johannes F, Colomé-Tatché M (2021) Region-level epimutation rates in Arabidopsis thaliana. Heredity (edinb) 127:190–202. https://doi.org/10.1038/s41437-021-00441-w
Ding Y, Avramova Z, Fromm M (2011) The Arabidopsis trithorax-like factor ATX1 functions in dehydration stress responses via ABA-dependent and ABA-independent pathways. Plant J 66:735–744. https://doi.org/10.1111/j.1365-313X.2011.04534.x
Ding Y, Fromm M, Avramova Z (2012) Multiple exposures to drought “train” transcriptional responses in Arabidopsis. Nat Comm 3:740. https://doi.org/10.1038/ncomms1732
Ding Y, Liu N, Virlouvet L et al (2013) Four distinct types of dehydration stress memory genes in Arabidopsis thaliana. BMC Plant Biol 13:229. https://doi.org/10.1186/1471-2229-13-229
Ding Y, Virlouvet L, Liu N, Riethoven JJ, Fromm M, Avramova Z (2014) Dehydration stress memory genes of Zea mays; comparison with Arabidopsis thaliana. BMC Plant Biol 14:141. https://doi.org/10.1186/1471-2229-14-141
Fang O, Zhang QB, Vitasse Y, Zweifel R, Cherubini P (2021) The frequency and severity of past droughts shape the drought sensitivity of juniper trees on the Tibetan plateau. For Ecol Manag 486:118968. https://doi.org/10.1016/j.foreco.2021.118968
Fleta-Soriano E, Munné-Bosch S (2016) Stress memory and the inevitable effects of drought: a physiological perspective. Front Plant Sci 7:143. https://doi.org/10.3389/FPLS.2016.00143/BIBTEX
Fleta-Soriano E, Pinto-Marijuan M, Munne-Bosch S (2015) Evidence of drought stress memory in the facultative CAM, Aptenia cordifolia: possible role of phytohormones. PLoS ONE 10:e0135391. https://doi.org/10.1371/journal.pone.0135391
Fujii H, Zhu JK (2009) Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc Nat Acad Sci USA 106(20):8380–8385. https://doi.org/10.1073/pnas.0903144106
Fujii H, Verslues PE, Zhu JK (2011) Arabidopsis decuple mutant reveals the importance of SnRK2 kinases in osmotic stress responses in vivo. Proc Nat Acad Sci USA 108(4):1717–1722. https://doi.org/10.1073/pnas.1018367108
Fujita Y, Yoshida T, Yamaguchi-Shinozaki K (2013) Pivotal role of the AREB/ABF-SnRK2 pathway in ABRE-mediated transcription in response to osmotic stress in plants. Physiol Plant 147(1):15–27. https://doi.org/10.1111/j.1399-3054.2012.01635.x
Galiano L, Timofeeva G, Saurer M et al (2017) The fate of recently fixed carbon after drought release: towards unravelling C storage regulation in Tilia platyphyllos and Pinus sylvestris. Plant Cell Environ. https://doi.org/10.1111/pce.12972
Galviz YCF, Ribeiro RV, Souza GM (2020) Yes, plants do have memory. Theor Exp Plant Physiol 32:195–202. https://doi.org/10.1007/s40626-020-00181-y
Ganguly D, Crisp PA, Eichten SR, Pogson BJ (2017) The Arabidopsis DNA methylome is stable under transgenerational drought stress. Plant Physiol 175(4):1893–1912. https://doi.org/10.1104/pp.17.00744
Gayacharan JAJ (2013) Epigenetic responses to drought stress in rice (Oryza sativa L.). Physiol Mol Biol Plants 19(3):379–387. https://doi.org/10.1007/s12298-013-0176-4
Gelaw TA, Sanan-Mishra N (2021) Non-coding RNAs in response to drought stress. Int J Mol Sci. https://doi.org/10.3390/ijms222212519
Godwin J, Farrona S (2020) Plant epigenetic stress memory induced by drought: a physiological and molecular perspective. Methods in molecular biology. Humana, New York, pp 243–259
Hallouin M, Ghelis T, Brault M et al (2002) Plasmalemma abscisic acid perception leads to RAB18 expression via phospholipase D activation in Arabidopsis suspension cells. Plant Physiol 130:265–272. https://doi.org/10.1104/pp.004168
Han SK, Sang Y, Rodrigues A et al (2012) The SWI2/SNF2 chromatin remodeling ATPase BRAHMA represses abscisic acid responses in the absence of the stress stimulus in Arabidopsis. Plant Cell 24(12):4892–4906. https://doi.org/10.1105/tpc.112.105114
Hand SC, Menze MA, Toner M, Boswell L, Moore D (2011) LEA proteins during water stress: not just for plants anymore. Ann Rev Physiol 73:115–134. https://doi.org/10.1146/annurev-physiol-012110-142203
He Y, Li Z (2018) Epigenetic environmental memories in plants: establishment, maintenance, and reprogramming. Trends Genet 34:856–866. https://doi.org/10.1016/j.tig.2018.07.006
He L, Wu W, Zinta G et al (2018) A naturally occurring epiallele associates with leaf senescence and local climate adaptation in Arabidopsis accessions. Nature Commun 9(1):1–11. https://doi.org/10.1038/s41467-018-02839-3
Heo JB, Sung S (2011) Vernalization-mediated epigenetic silencing by a long intronic non-coding RNA. Science 331:76–79. https://doi.org/10.1126/science.1197349
Herman JJ (1838) Sultan SE (2016) DNA methylation mediates genetic variation for adaptive transgenerational plasticity. Proc R Soc b Biol Sci 283:20160988. https://doi.org/10.1098/rspb.2016.0988
Hirayama T, Shinozaki K (2010) Research on plant abiotic stress responses in the post-genome era: past, present and future. Plant J 61:1041–1052. https://doi.org/10.1111/j.1365-313X.2010.04124.x
Huettel B, Kanno T, Daxinger L et al (2007) RNA-directed DNA methylation mediated by DRD1 and Pol IVb: a versatile pathway for transcriptional gene silencing in plants. Biochim Biophys Acta 1769:358–374. https://doi.org/10.1016/j.bbaexp.2007.03.001
Hundertmark M, Hincha DK (2008) LEA (Late Embryogenesis Abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genom 9:1–22. https://doi.org/10.1186/1471-2164-9-118/FIGURES/5
Iwasaki M, Paszkowski J (2014) Identification of genes preventing transgenerational transmission of stress-induced epigenetic states. Proc Nat Acad Sci USA 111(23):8547–8552. https://doi.org/10.1073/pnas.1402275111
Jacques C, Salon C, Barnard RL, Vernoud V, Prudent M (2021) Drought stress memory at the plant cycle level: a review. Plants 10(9):1873. https://doi.org/10.3390/plants10091873
Karle SB, Nair B, Kumar K (2020) SnRK2s: part and parcel of ABA signaling in plants. Protein kinases and stress signaling in plants: functional genomic perspective, pp. 133–46 https://doi.org/10.1002/9781119541578.ch6
Kawakatsu T, Huang SSC, Jupe F et al (2016) Epigenomic diversity in a global collection of Arabidopsis thaliana accessions. Cell 166(2):492–505. https://doi.org/10.1016/j.cell.2016.06.044
Kazan K, Manners JM (2013) MYC2: the master in action. Mol Plant 6:686–703. https://doi.org/10.1093/mp/sss128
Kim ED, Sung S (2012) Long non-coding RNA: unveiling hidden layer of gene regulatory networks. Trends Plant Sci 17(1):16–21. https://doi.org/10.1016/j.tplants.2011.10.008
Kim J-M, To TK, Ishida J et al (2008) Alterations of lysine modifications on the histone H3 N-tail under drought stress conditions in Arabidopsis thaliana. Plant Cell Physiol 49:1580–1588. https://doi.org/10.1093/pcp/pcn133
Kim YK, Chae S, Oh NI, Nguyen NH, Cheong JJ (2020) Recurrent drought conditions enhance the induction of drought stress memory genes in Glycine max L. Front Genet. https://doi.org/10.3389/fgene.2020.576086
Kinoshita T, Seki M (2014) Epigenetic memory for stress response and adaptation in plants. Plant Cell Physiol 55:1859–1863. https://doi.org/10.1093/pcp/pcu125
Kobayashi Y, Sadhukhan A, Tazib T et al (2016) Joint genetic and network analyses identify loci associated with root growth under NaCl stress in Arabidopsis thaliana. Plant Cell Environ 39:918–934. https://doi.org/10.1111/pce.12691
Kou S, Gu Q, Duan L et al (2021) Genome-wide bisulphite sequencing uncovered the contribution of DNA methylation to rice short-term drought memory formation. J Plant Growth Regul. https://doi.org/10.1007/s00344-021-10483-3
Kron AP, Souza GM, Ribeiro RV (2008) Water deficiency at different developmental stages of Glycine max can improve drought tolerance. Bragantia 67:43–49. https://doi.org/10.1590/S0006-87052008000100005
Labra M, Ghiani A, Citterio S et al (2002) Analysis of cytosine methylation pattern in response to water deficit in pea root tips. Plant Biol 4(6):694–699. https://doi.org/10.1055/s-2002-37398
Lahuta LB, Szablińska-Piernik J, Horbowicz M (2022) Changes in metabolic profiles of pea (Pisum sativum L.) as a result of repeated short-term soil drought and subsequent re-watering. Int J Mol Sci 23:1704. https://doi.org/10.3390/IJMS23031704
Leng G, Hall J (2019) Crop yield sensitivity of global major agricultural countries to droughts and the projected changes in the future. Sci Total Environ 654:811–821. https://doi.org/10.1016/j.scitotenv.2018.10.434
Leufen G, Noga G, Hunsche M (2016) Drought stress memory in sugar beet: mismatch between biochemical and physiological parameters. J Plant Growth Regul 353(35):680–689. https://doi.org/10.1007/S00344-016-9571-8
Levine M (2011) Paused RNA polymerase II as a developmental checkpoint. Cell 145:502–511. https://doi.org/10.1016/j.cell.2011.04.021
Li X, Liu F (2016) Drought stress memory and drought stress tolerance in plants: biochemical and molecular basis. In: Drought stress tolerance in plants, vol 1. Springer, Cham, pp 17–44. https://doi.org/10.1007/978-3-319-28899-4_2
Li WX, Oono Y, Zhu J et al (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 20(8):2238–2251. https://doi.org/10.1105/tpc.108.059444
Li P, Yang H, Wang L et al (2019) Physiological and transcriptome analyses reveal short-term responses and formation of memory under drought stress in rice. Front Genet 10:55. https://doi.org/10.3389/fgene.2019.00055
Liu HH, Tian X, Li YJ, Wu CA, Zheng CC (2008) Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 14(5):836–843. https://doi.org/10.1261/rna.895308
Liu N, Fromm M, Avramova Z (2014) H3K27me3 and H3K4me3 chromatin environment at super-induced dehydration stress memory genes of Arabidopsis thaliana. Mol Plant 7:502–513. https://doi.org/10.1093/mp/ssu001
Liu X, Hao L, Li D, Zhu L, Hu S (2015) Long non-coding RNAs and their biological roles in plants. In: Genomics, proteomics and bioinformatics, vol 13, issue 3. Elsevier, Oxford, pp 137–147. https://doi.org/10.1016/j.gpb.2015.02.003
Liu N, Staswick PE, Avramova Z (2016) Memory responses of jasmonic acid-associated Arabidopsis genes to a repeated dehydration stress. Plant Cell Environ 39:2515–2529. https://doi.org/10.1111/pce.12806
Liu X, Challabathula D, Quan W et al (2019) Transcriptional and metabolic changes in the desiccation tolerant plant Craterostigma plantagineum during recurrent exposures to dehydration. Planta 249:1017–1035. https://doi.org/10.1007/s00425-018-3058-8
Lopes-Caitar VS, Silva SMH, Marcelino-Guimaraes FC (2016) Plant small heat shock proteins and its interactions with biotic stress. In: Asea A, Kaur P, Calderwood S (eds) Heat shock proteins and plants. Heat shock proteins, vol 10. Springer, Cham. https://doi.org/10.1007/978-3-319-46340-7_2
Lopez BC, Gracia CA, Sabate S, Keenan T (2009) Assessing the resilience of Mediterranean holm oaks to disturbances using selective thinning. Acta Oecol (montrouge) 35:849–854. https://doi.org/10.1016/j.actao.2009.09.001
Luna E, Bruce TJA, Roberts MR, Flors V, Ton J (2012) Next-generation systemic acquired resistance. Plant Physiol 158:844–853. https://doi.org/10.1104/pp.111.187468
Luo L, Zheng Y, Gao Z et al (2020) Grafting improves drought stress memory by increasing the P5CS1 gene expression in Brassica rapa. Plant Soil 452:61–72. https://doi.org/10.1007/s11104-020-04547-8
Magwanga RO, Lu P, Kirungu JN et al (2018) Characterization of the late embryogenesis abundant (LEA) proteins family and their role in drought stress tolerance in upland cotton. BMC Genet. https://doi.org/10.1186/S12863-017-0596-1
Mantyla E, Lang V, Palva ET (1995) Role of abscisic acid in drought-induced freezing tolerance, cold acclimation, and accumulation of LT178 and RAB18 proteins in Arabidopsis thaliana. Plant Physiol 107(1):141–148. https://doi.org/10.1104/pp.107.1.141
Martorell S, Medrano H, Tomàs M et al (2015) Plasticity of vulnerability to leaf hydraulic dysfunction during acclimation to drought in grapevines: a+n osmotic-mediated process. Physiol Plant 153:381–391. https://doi.org/10.1111/PPL.12253
Meena M, Divyanshu K, Kumar S et al (2019) Regulation of L-proline biosynthesis, signal transduction, transport, accumulation and its vital role in plants during variable environmental conditions. Heliyon 5:e02952. https://doi.org/10.1016/J.HELIYON.2019.E02952
Mehrotra R, Bhalothia P, Bansal P, Basantani MK, Bharti V, Mehrotra S (2014) Abscisic acid and abiotic stress tolerance–different tiers of regulation. J Plant Physiol 171(7):486–496. https://doi.org/10.1016/j.jplph.2013.12.007
Melnyk CW, Molnar A, Baulcombe DC (2011) Intercellular and systemic movement of RNA silencing signals. EMBO J 30(17):3553–3563. https://doi.org/10.1038/emboj.2011.274
Minow MA, Colasanti J (2020) Does variable epigenetic inheritance fuel plant evolution? Genome 63(5):253–262. https://doi.org/10.1139/gen-2019-0190
Miura K, Okamoto H, Okuma E et al (2013) SIZ1 deficiency causes reduced stomatal aperture and enhanced drought tolerance via controlling salicylic acid-induced accumulation of reactive oxygen species in Arabidopsis. Plant J 73:91–104. https://doi.org/10.1111/tpj.12014
Mozgova I, Mikulski P, Pecinka A, Farrona S (2019) Epigenetic mechanisms of abiotic stress response and memory in plants. Epigenetics in plants of agronomic importance: fundamentals and applications: transcriptional regulation and chromatin remodelling in plants, 2nd edn, pp 1–64. https://doi.org/10.1007/978-3-030-14760-0_1
Msanne J, Lin J, Stone JM, Awada T (2011) Characterization of abiotic stress-responsive Arabidopsis thaliana RD29A and RD29B genes and evaluation of transgenes. Planta 234(1):97–107. https://doi.org/10.1007/s00425-011-1387-y
Neves DM, Almeida LAdH, Santana-Vieira DDS et al (2017) Recurrent water deficit causes epigenetic and hormonal changes in citrus plants. Sci Rep 7:13684. https://doi.org/10.1038/s41598-017-14161-x
Oh JE, Kwon Y, Kim JH, Noh H, Hong SW, Lee H (2011) A dual role for MYB60 in stomatal regulation and root growth of Arabidopsis thaliana under drought stress. Plant Mol Biol 77(1):91–103. https://doi.org/10.1007/s11103-011-9796-7
Osakabe Y, Osakabe K, Shinozaki K, Tran LSP (2014) Response of plants to water stress. Front Plant Sci 5:86. https://doi.org/10.3389/fpls.2014.00086
Pagliarani C, Vitali M, Ferrero M, Vitulo N, Incarbone M, Lovisolo C, Valle G, Schubert A (2017) The accumulation of miRNAs differentially modulated by drought stress is affected by grafting in grapevine. Plant Physiol 173(4):2180–2195. https://doi.org/10.1104/pp.16.01119
Paixão JFR, Gillet FX, Ribeiro TP et al (2019) Improved drought stress tolerance in Arabidopsis by CRISPR/dCas9 fusion with a Histone Acetyl Transferase. Sci Rep 9(1):1–9. https://doi.org/10.1038/s41598-019-44571-y
Papaefthimiou D, Tsaftaris AS (2012) Significant induction by drought of HvPKDM7–1, a gene encoding a jumonji-like histone demethylase homologue in barley (H. vulgare). Acta Physiol Plant 34(3):1187–1198. https://doi.org/10.1007/s11738-011-0915-5
Park K, Lee S, Yoo H et al (2020) DEMETER-mediated DNA demethylation in gamete companion cells and the endosperm, and its possible role in embryo development in arabidopsis. J Plant Biol 63:321–329. https://doi.org/10.1007/s12374-020-09258-2
Pecinka A, Mittelsten Scheid O (2012) Stress-induced chromatin changes: a critical view on their heritability. Plant Cell Physiol 53(5):801–808. https://doi.org/10.1093/pcp/pcs044
Pikaard CS (2006) Cell biology of the Arabidopsis nuclear siRNA pathway for RNA-directed chromatin modification. Cold Spring Harb Symp Quant Biol 71:473–480. https://doi.org/10.1101/sqb.2006.71.046
Raj S, Bräutigam K, Hamanishi ET et al (2011) Clone history shapes Populus drought responses. Proc Nat Acad Sci USA 108(30):12521–12526. https://doi.org/10.1073/pnas.1103341108
Rajkumar MS, Shankar R, Garg R, Jain M (2020) Bisulphite sequencing reveals dynamic DNA methylation under desiccation and salinity stresses in rice cultivars. Genomics 112(5):3537–3548. https://doi.org/10.1016/j.ygeno.2020.04.005
Raju SKK, Shao MR, Sanchez R, Xu YZ, Sandhu A, Graef G, Mackenzie S (2018) An epigenetic breeding system in soybean for increased yield and stability. Plant Biotechnol J 16(11):1836–1847. https://doi.org/10.1111/pbi.12919
Ribeiro RV, Torres RDS (2018) Sentinel plants as programmable processing units: insights from a multidisciplinary perspective about stress memory and plant signaling and their relevance at community level. Plant Signal Behav 13(10):e1526001. https://doi.org/10.1080/15592324.2018.1526001
Rusconi F, Simeoni F, Francia P et al (2013) The Arabidopsis thaliana MYB60 promoter provides a tool for the spatio-temporal control of gene expression in stomatal guard cells. J Exp Bot 64(11):3361–3371. https://doi.org/10.1093/jxb/ert180
Rutowicz K, Puzio M, Halibart-Puzio J et al (2015) A specialized histone H1 variant is required for adaptive responses to complex abiotic stress and related DNA methylation in Arabidopsis. Plant Physiol 169:2080–2101. https://doi.org/10.1104/pp.15.00493
Sadhukhan A, Kobayashi Y, Nakano Y, Iuchi S, Kobayashi M, Sahoo L, Koyama H (2017) Genome-wide association study reveals that the aquaporin NIP1;1 contributes to variation in hydrogen peroxide sensitivity in Arabidopsis thaliana. Mol Plant 10:1082–1094
Sadhukhan A, Agrahari RK, Wu L, Watanabe T, Nakano Y, Panda SK, Koyama H, Kobayashi Y (2021) Expression genome-wide association study identifies that phosphatidylinositol-derived signaling regulates ALUMINIUM SENSITIVE3 expression under aluminium stress in the shoots of Arabidopsis thaliana. Plant Sci 302:110711
Sakakibara H, Takei K, Hirose N (2006) Interactions between nitrogen and cytokinin in the regulation of metabolism and development. Trends Plant Sci 11(9):440–448. https://doi.org/10.1016/j.tplants.2006.07.004
Sala A, Woodruff DR, Meinzer FC (2012) Carbon dynamics in trees: feast or famine? Tree Physiol 32:764–775. https://doi.org/10.1093/TREEPHYS/TPR143
Salinger MJ (2005) Climate variability and change: past, present and future—an overview. Increasing climate variability and change. Clim Change 70:9–29. https://doi.org/10.1007/s10584-005-5936-x
Santos AP, Serra T, Figueiredo DD, Barros P, Lourenço T, Chander S, Oliveira MM, Saibo NJM (2011) Transcription regulation of abiotic stress responses in rice: a combined action of transcription Factors and Epigenetic Mechanisms. OMICS J Integr Biol 15:839–857. https://doi.org/10.1089/omi.2011.0095
Sasaki-Sekimoto Y, Taki N, Obayashi T et al (2005) Coordinated activation of metabolic pathways for antioxidants and defence compounds by jasmonates and their roles in stress tolerance in Arabidopsis. Plant J 44:653–668. https://doi.org/10.1111/j.1365-313X.2005.02560.x
Scippa GS, Di Michele M, Onelli E et al (2004) The histone-like protein H1-S and the response of tomato leaves to water deficit. J Exp Bot 55:99–109. https://doi.org/10.1093/jxb/erh022
Sedaghatmehr M, Thirumalaikumar VP, Kamranfar I, Marmagne A, Masclaux-Daubresse C, Balazadeh S (2019) A regulatory role of autophagy for resetting the memory of heat stress in plants. Plant Cell Environ 42:1054–1064. https://doi.org/10.1111/pce.13426
Shuai P, Liang D, Tang S, Zhang Z, Ye CY, Su Y et al (2014) Genome-wide identification and functional prediction of novel and drought-responsive lincRNAs in Populus trichocarpa. J Exp Bot 65:4975–4983. https://doi.org/10.1093/jxb/eru256
Suksamran R, Saithong T, Thammarongtham C, Kalapanulak S (2020) Genomic and transcriptomic analysis identified novel putative cassava lncRNAs involved in cold and drought stress. Genes (basel) 11:366. https://doi.org/10.3390/genes11040366
Sun C, Ali K, Yan K et al (2021) Exploration of epigenetics for improvement of drought and other stress resistance in crops: a review. Plants 10:1226. https://doi.org/10.3390/plants10061226
Sunkar R, Zhu JK (2004) Novel and stress-regulated MicroRNAs and other small RNAs from Arabidopsis. Plant Cell 16(8):2001–2019. https://doi.org/10.1105/tpc.104.022830
Szabados L, Savouré A (2010) Proline: a multifunctional amino acid. Trends Plant Sci 15:89–97. https://doi.org/10.1016/J.TPLANTS.2009.11.009
Thuy Quynh Nguyen T, Thanh Huyen Trinh L, Bao Vy Pham H et al (2020) Evaluation of proline, soluble sugar and ABA content in soybean Glycine max (L.) under drought stress memory. AIMS Bioeng 7:114–123. https://doi.org/10.3934/bioeng.2020011
Todaka D, Takahashi F, Yamaguchi-Shinozaki K, Shinozaki K (2019) ABA-responsive gene expression in response to drought stress: cellular regulation and long-distance signaling. In Advances in botanical research. Elsevier, pp. 83–113. https://doi.org/10.1016/bs.abr.2019.05.001
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97:11632–11637. https://doi.org/10.1073/pnas.190309197
Van DK, Ding Y, Malkaram S et al (2010) Dynamic changes in genome-wide histone H3 lysine 4 methylation patterns in response to dehydration stress in Arabidopsis thaliana. BMC Plant Biol 10:238. https://doi.org/10.1186/1471-2229-10-238
Virlouvet L, Avenson TJ, Du Q, Zhang C, Liu N, Fromm M, Avramova Z, Russo SE (2018) Dehydration stress memory: Gene networks linked to physiological responses during repeated stresses of zea mays. Front Plant Sci 9:1058. https://doi.org/10.3389/fpls.2018.01058
Virlouvet L, Fromm M (2015) Physiological and transcriptional memory in guard cells during repetitive dehydration stress. New Phytol 205(2):596–607. https://doi.org/10.1111/nph.13080
Virlouvet L, Ding Y, Fujii H, Avramova Z, Fromm M (2014) ABA signaling is necessary but not sufficient for RD29B transcriptional memory during successive dehydration stresses in Arabidopsis thaliana. Plant J 79:150–161. https://doi.org/10.1111/tpj.12548
Walter J, Nagy L, Hein R et al (2011) Do plants remember drought? Hints towards a drought-memory in grasses. Environ Exp Bot 71:34–40. https://doi.org/10.1016/j.envexpbot.2010.10.020
Walter J, Jentsch A, Beierkuhnlein C, Kreyling J (2013) Ecological stress memory and cross stress tolerance in plants in the face of climate extremes. Environ Exp Bot 94:3–8. https://doi.org/10.1016/j.envexpbot.2012.02.009
Wang X-L, Wang J-J, Sun R-H et al (2016) Correlation of the corn compensatory growth mechanism after post-drought rewatering with cytokinin induced by root nitrate absorption. Agric Water Manag 166:77–85. https://doi.org/10.1016/j.agwat.2015.12.007
Wang X, Li Q, Xie J et al (2021) Abscisic acid and jasmonic acid are involved in drought priming-induced tolerance to drought in wheat. Crop J 9:120–132. https://doi.org/10.1016/j.cj.2020.06.002
Ward D (2016) The biology of deserts. Oxford Univ Press. https://doi.org/10.1093/acprof:oso/9780199211470.001.0001
Wojtyla Ł, Paluch-Lubawa E, Sobieszczuk-Nowicka E, Garnczarska M (2020) Drought stress memory and subsequent drought stress tolerance in plants. Prim Mediat Stress Cross-Stress Tolerance Crop Plants. https://doi.org/10.1016/B978-0-12-817892-8.00007-6
Woodruff DR, Meinzer FC (2011) Water stress, shoot growth and storage of non-structural carbohydrates along a tree height gradient in a tall conifer. Plant Cell Environ 34:1920–1930. https://doi.org/10.1111/J.1365-3040.2011.02388.X
Yancey PH (2005) Organic osmolytes as compatible, metabolic and counteracting cytoprotectants in high osmolarity and other stresses. J Exp Biol 208:2819–2830. https://doi.org/10.1242/JEB.01730
Yokthongwattana C, Bucher E, Caikovski M et al (2010) MOM1 and Pol-IV/V interactions regulate the intensity and specificity of transcriptional gene silencing. EMBO J 29(2):340–351. https://doi.org/10.1038/emboj.2009.328
Yong-Villalobos L, González-Morales SI, Wrobel K et al (2015) Methylome analysis reveals an important role for epigenetic changes in the regulation of the Arabidopsis response to phosphate starvation. Proc Natl Acad Sci USA 112:E7293–E7302. https://doi.org/10.1073/pnas.1522301112
Yoshida T, Fujita Y, Sayama H et al (2010) AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J 61:672–685. https://doi.org/10.1111/j.1365-313X.2009.04092.x
Zhang X, Bernatavichute YV, Cokus S, Pellegrini M, Jacobsen SE (2009) Genome-wide analysis of mono-, di- and trimethylation of histone H3 lysine 4 in Arabidopsis thaliana. Genome Biol 10:R62. https://doi.org/10.1186/gb-2009-10-6-r62
Zhang C, Wang N, Zhang Y, Feng Q, Yang C, Liu B (2013) DNA methylation involved in proline accumulation in response to osmotic stress in rice (Oryza sativa). Genet Mol Res 12(2):1269–1277. https://doi.org/10.4238/2013.April.17.5
Zhang C, Tang G, Peng X, Sun F, Liu S, Xi Y (2018a) Long non-coding RNAs of switchgrass (Panicum virgatum L.) in multiple dehydration stresses. BMC Plant Biol 18(1):1–15. https://doi.org/10.1186/s12870-018-1288-3
Zhang H, Lang Z, Zhu JK (2018b) Dynamics and function of DNA methylation in plants. Nat Rev Mol Cell Biol 19(8):489–506. https://doi.org/10.1038/s41580-018-0016-z
Zhang Y, Wendte JM, Ji L, Schmitz RJ (2020) Natural variation in DNA methylation homeostasis and the emergence of epialleles. Proc Nat Acad Sci 117(9):4874–4884. https://doi.org/10.1073/pnas.1918172117
Zhou Y, Zhang J, Lin H, Guo G, Guo Y (2010) MORPHEUS’ MOLECULE1 is required to prevent aberrant RNA transcriptional read-through in Arabidopsis. Plant Physiol 154(3):1272–1280. https://doi.org/10.1104/pp.110.162131
Zhu J-K (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Physiol Plant Mol Biol 53:247–273. https://doi.org/10.1146/annurev.arplant.53.091401.143329
Zhu F, Ahchige MW, Brotman Y, Alseekh S, Zsögön A, Fernie AR (2022) Bringing more players into play: Leveraging stress in genome wide association studies. J Plant Physiol 271:153657. https://doi.org/10.1016/j.jplph.2022.153657
Acknowledgements
The authors are grateful to their respective institutions for providing infrastructure support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Gerhard Leubner.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sadhukhan, A., Prasad, S.S., Mitra, J. et al. How do plants remember drought?. Planta 256, 7 (2022). https://doi.org/10.1007/s00425-022-03924-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-022-03924-0