Abstract
Main conclusion
OsbZIP42 is a positive regulator of ABA signaling and drought stress tolerance. The activation of OsbZIP42 depends on stress-/ABA-activated protein kinase 4 (SAPK4) and an additional ABA-dependent modification of OsbZIP42.
Basic leucine zipper transcription factors (bZIP TFs) play important roles in the ABA signaling pathway in plants. Rice OsbZIP42 is a member of the group E bZIP, which is an ortholog of Arabidopsis group A bZIP. This latter group includes abscisic acid-responsive element (ABRE)-binding factors (ABFs) involved in abiotic stress tolerance. The expression of OsbZIP42 was induced by ABA treatment, although it was not induced by drought and salt stresses. Unlike other bZIP TFs, OsbZIP42 contained two transcriptional activation domains. Although the full-length OsbZIP42 protein did not, the N-terminus of the protein interacted with SAPK4. Our results suggest that the activation of OsbZIP42 by SAPK4 requires another ABA-dependent modification of OsbZIP42. Transgenic rice overexpressing OsbZIP42 (OsbZIP42-OX) exhibited a rapidly elevated expression of the ABA-responsive LEA3 and Rab16 genes and was hypersensitive to ABA. Analyses of the OsbZIP42-OX plants revealed enhanced tolerance to drought stress. These results suggest that OsbZIP42 is a positive regulator of ABA signaling and drought stress tolerance depending on its activation, which is followed by an additional ABA-dependent modification. We propose that OsbZIP42 is an important player in rice for conferring ABA-dependent drought tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Abiotic and biotic stresses seriously affect crop yield. Abiotic stresses particularly cause an average yield loss of more than 50% in most major crop plants (Fujita et al. 2006). In particular, rice (Oryza sativa L.) production has been significantly affected by drought stress (Tuong and Bouman 2003). Rice is a notoriously drought-susceptible crop due in part to its small root system (Hirasawa 1999). Thus, understanding drought stress responses in rice is important for improving yield and quality. The ABA signal transduction pathway plays an important role in abiotic stress responses that modulate stress-responsive gene expression (Shinozaki and Yamaguchi-Shinozaki 1997; Verslues and Zhu 2005). In Arabidopsis, the ABA signaling model has been well established. The ABA and receptor (PYR/PYL/RCAR) complex inhibits type 2C protein phosphatases (PP2C), which dephosphorylate/inactivate SNF1-related protein kinase 2 (SnRK2) (Umezawa et al. 2009; Vlad et al. 2009; Bhaskara et al. 2012). This deactivation leads to the activation of SnRK2, which phosphorylates/activates bZIP transcription factors (TFs; Schütze et al. 2008; Cutler et al. 2010). This type of posttranslational modification is one of the mechanisms that regulate the activities of bZIP TFs. bZIP TFs are defined by a conserved bZIP domain (Landschulz et al. 1988) and are involved in numerous response pathways in plants, such as light signaling, abiotic stress responses, biotic stress defense, seed maturation and flower development (Jakoby et al. 2002; Smeekens et al. 2010; Zg et al. 2014). bZIP TFs are present in all eukaryotes and are expressed by multiple genes. Arabidopsis encodes approximately 75 bZIP TFs (Jakoby et al. 2002), and rice encodes approximately 89 bZIP TFs (Nijhawan et al. 2008). The bZIP family is subdivided into ten groups (A–I plus S in Arabidopsis and A–J in rice) based on sequence similarity and conserved motifs.
bZIP TFs are involved in responses to ABA signaling and play important roles in ABA-dependent stress responses (Umezawa et al. 2010; Llorca et al. 2014). The Arabidopsis group A bZIP TFs include ABF1, ABF2/AREB1, ABF3, ABF4/AREB2, ABI5, GBF4, DPBF2, and DPBF4 (Jakoby et al. 2002). ABF1–ABF4 and ABI5 are expressed in vegetative tissues, and their expression is induced by various abiotic stresses (Choi et al. 2000; Finkelstein and Lynch 2000; Uno et al. 2000; Kang et al. 2002; Fujita et al. 2005). They play important roles in cooperatively regulating ABRE-dependent ABA signaling involved in abiotic stress tolerance (Uno et al. 2000; Kang et al. 2002; Yoshida et al. 2010). Furthermore, ABF2/AREB1, ABF4/AREB2, and ABF3 are the key TFs for ABA-dependent stress responses (Yoshida et al. 2010). These TFs are highly homologous to the members of rice group E bZIP TFs (Xiang et al. 2008; Lu et al. 2009). Members of rice group E, such as OsbZIP10/OsABI5 (Zou et al. 2008), OsbZIP12 (Amir Hossain et al. 2010; Joo et al. 2014), OsbZIP23 (Xiang et al. 2008), OsbZIP46 (Tang et al. 2012), OsbZIP66/TRAB1 (Hobo et al. 1999), and OsbZIP72 (Lu et al. 2009), are also involved in ABA signal transduction and abiotic stress responses. In addition, the members of rice group E bZIP TFs contain motifs 10–14, which contain potential phosphorylation sites (Nijhawan et al. 2008). These motifs have been identified in members of group A bZIP TFs in Arabidopsis. The activation of Arabidopsis group A bZIP TFs, which are key transcriptional regulators of ABA responses, is known to be regulated by ABA- and abiotic stress-induced protein kinases (Schütze et al. 2008). Rice group E bZIP TRAB1 is also phosphorylated in response to ABA stress. SnRK2 kinase directly phosphorylates TRAB1 in response to ABA (Kobayashi et al. 2005). The phosphorylation of bZIP proteins can modify homo-/heterodimerization and DNA-binding properties. Phosphorylation of bZIP proteins can also modify dimerization specificity (Lee et al. 2010), DNA-binding properties (Deppmann et al. 2003), and subcellular localization (Djamei et al. 2007; Ishida et al. 2008).
In this study, we characterized OsbZIP42, which belongs to subgroup II of group E. We determined the functional analysis of OsbZIP42 by investigating the ABA and sugar sensitivity and the stress tolerance of transgenic plants overexpressing OsbZIP42. An analysis of the cis-acting elements in the promoter region and an expression profiling analysis were also performed. The results indicate that OsbZIP42 is a positive regulator of ABA signaling and confers ABA-dependent drought stress tolerance to rice.
Materials and methods
Plant materials and growth conditions
Both the transgenic and wild-type (WT) rice plants used in this study were of the Oryza sativa subsp. japonica cv. Nakdong background (seeds obtained from Professor Ju-kon Kim, Graduate School of International Agricultural Technology and Crop Biotechnology Institute/GreenBio Science and Technology, Seoul National University, Pyeongchang, Korea). Husked seeds were germinated in a half-strength Murashige–Skoog (MS) solid medium in a growth chamber, transplanted into soil in pots and grown in a greenhouse until further use (Joo et al. 2013b). The samples from each rice organ were prepared as described below. Seeds from the transgenic and WT rice were germinated in half-strength MS solid medium in a growth chamber at 28 °C for 3 days in the dark, followed by 3 days in the light. After germination, the seedlings were grown in a greenhouse for 3 weeks to obtain leaves, internodes, and roots for analysis. Panicles were obtained from field-grown rice plants during the pre- and postheading stages. The young panicles were harvested from the sheath, measured and categorized into three groups (P3 3–5 cm, P4 10–15 cm and P5 15–20 cm) based on the length of the panicle. The rice seeds were tagged on the day of pollination (0 DAP) and collected on each DAP from 0 to 29 DAP (S1 0–2 DAP, S2 3–4 DAP, S3 5–10 DAP, S4 11–20 DAP and S5 25–29 DAP).
Quantitative real-time PCR analysis
Total RNA was isolated from the rice tissue samples using TRI Reagent® (Molecular Research Center) according to the manufacturer’s instructions. For quantitative real-time PCR (qRT-PCR), first-strand cDNA synthesis was performed using oligo(dT)18 primers and 5 μg of total RNA as a template according to the manufacturer’s instructions (RevertAid™ First-Strand cDNA Synthesis Kit, Fermentas). A one-third dilution of the cDNA synthesis reaction mixture was used, and 1 μL of the diluted cDNA mixture was added as a template for the subsequent real-time PCR analyses using 2 × Real-time PCR Pre-Mix with Evagreen (SolGent). Thermocycling and fluorescence detection were performed using an Mx3000p real-time PCR machine (Stratagene). qRT-PCR was performed in triplicate, and each experiment was repeated three times. OsUbi1 was used as a control to normalize the expression data. The primers for the qRT- PCR analyses are listed in Supplementary Table S1.
Promoter analysis
For the promoter analyses, 2-kb segments from the 5′ regulatory regions of the OsbZIP42 gene were obtained from Oryzabase and scanned for the presence of putative cis-acting elements identical or highly similar to the motifs registered in PLACE (http://www.dna.affrc.go.jp/PLACE/). The accession number of OsbZIP42 in NCBI is Os05g0489700.
Transactivation and two-hybrid assays in yeast
A transactivation assay was performed using the vector pDEST32 (Invitrogen) and the yeast strain Mav203 (Invitrogen). The full-length OsbZIP42 or fragments of OsbZIP42 were fused in frame with the yeast GAL4 DNA-binding domain in the pDEST32 vector using Gateway LR ClonaseTM II enzyme mix (Invitrogen). A colony-lift filter assay (X-gal assay) was performed according to the manufacturer’s manual (Invitrogen). A yeast two-hybrid assay was performed using a ProQuest two-hybrid system (Invitrogen). The coding region of OsbZIP42 was cloned into the pDEST22 vector, and SAPKs were cloned into the pDEST32 vector using Gateway technology (Invitrogen) to generate the bait and prey vectors. The two constructed vectors were cotransformed into the Mav203 yeast strain. The transformed yeast strains were plated on minimal medium–Leu/–Trp and incubated for 4 days, and a colony-lift filter assay (X-gal assay) was performed as described by the manufacturer (Invitrogen). The primers for the plasmid construction are listed in Supplementary Table S1.
Plasmid construction and transformation of rice
The overexpression plasmids contained the bar gene under the control of the cauliflower mosaic virus 35S promoter for herbicide-based selection. The rice cytochrome c promoter was used to drive constitutive expression. The coding region of the OsbZIP42 gene used in this study was PCR-amplified using rice total RNA and a pair of primers containing the attB sequence for the Gateway® recombination site. The primers for construction are listed in Supplementary Table S1. The attB-PCR products were inserted into pMJ101 through BP and LR recombination reactions according to the manufacturer’s instructions (Invitrogen). The plasmid for OsbZIP42-OX was confirmed by a sequencing analysis. The plasmids were introduced into Agrobacterium tumefaciens LBA4404 through triparental mating, and embryogenic calli from mature rice seeds were transformed. Callus induction, cocultivation with A. tumefaciens and the selection of transformed calli were performed (Joo et al. 2013b).
Abiotic stress treatments and chlorophyll fluorescence measurement
For the abiotic stress treatments, 3-week-old WT plants (grown in a greenhouse) were washed to remove the soil from their roots and transferred to a growth chamber (16-h light/8-h dark cycle at 28 °C) for water adaptation for 3 days. After adaptation, the plants were treated with a solution containing 100 µM ABA, 250 mM NaCl, 100 μM gibberellin (GA), 100 mM glucose (Glc) or 100 mM sucrose (Suc) or were air-dried (Dry). The samples were obtained at the indicated times. To test the ABA sensitivity of the transgenic plants at the germination stage, the transgenic lines and the WT seeds were germinated in half-strength MS medium containing ABA (0, 2, and 6 μM), and the germination rate of the treated seeds was calculated after 5 days. To evaluate the responses of the OsbZIP42-OX plants to abiotic stress during germination, the seeds of WT and transgenic plants were germinated in water containing 400 mM mannitol or 250 mM NaCl, and then the germination rates were measured. To carry out the drought-resistance assays, the germinated transgenic OsbZIP42-OX and WT seedlings that were grown in half-strength MS solid medium in a growth chamber were transplanted into soil. Fifteen to twenty seedlings from each transgenic and WT plants were grown in pots (3 × 3 × 5 cm; one plant per pot) for 4 weeks before carrying out the drought stress experiments. To test for resistance to drought stress, 4-week-old WT and transgenic plants (grown in pots) were subjected to 2 days of drought followed by 10 days of watering in a greenhouse. Chlorophyll fluorescence was measured in 4-week-old WT and transgenic plants using a pulse modulation fluorometer (mini-PAM, Walz). For the leaf disc tests, the green portions of approximately ten seedlings were cut using scissors prior to in vitro stress treatments. Under continuous light at 150 µmol m−2 s−1, the leaf discs were air dried for 2 h (for drought stress). For cold stress, the leaf discs were incubated in a 4 °C water bath for 5 h under the same light conditions as described above. After the stress treatments, the leaf discs were dark-adapted for 10 min, and the minimal fluorescence level (Fo) was measured. After the application of a saturating light pulse, the maximal fluorescence level (Fm) was measured, and the ratio of Fv to Fm (Fv/Fm = Fm − F0/Fm), which represents the activity of photosystem II, was used to assess functional damage to the plants (Joo et al. 2013a).
Results
Transcriptional activity of OsbZIP42
We studied the molecular function of OsbZIP42 in subgroup II of the group E bZIP TFs. The amino acid sequence of OsbZIP42 was analyzed using an online motif scan tool (http://hits.isb-sib.ch/cgi-bin/PFSCAN), which revealed that it contained the bZIP domain, a nuclear localization signal (NLS), and five conserved phosphorylation sites (Fig. 1a). First, we checked the transactivation activity of OsbZIP42 in yeast. The full-length OsbZIP42 (42-FL) fused to the GAL4 DNA-binding domain had no transactivation activity in yeast (Fig. 1b). To map the transcriptional activation domains within OsbZIP42, we fused the GAL4 DNA-binding domain to the N-terminus or C-terminus with or without domain D. Domain D is a conserved 23-amino acid stretch that is only in the members of group E bZIP TFs (domain 13 in Nijhawan et al. 2008; Tang et al. 2012). A deletion assay of OsbZIP46 indicated that transactivation activity was detected only when domain D was absent. It was proposed that domain D is a negative regulatory domain and has a pivotal role in the regulation of OsbZIP46 activity (Tang et al. 2012). The N-terminus (42-N1, residues 1–128 without domain D) activated the expression of reporter genes, but the 42-N2 (residues 1–164 with domain D) did not. Interestingly, the C-terminus of OsbZIP42 (42-C1, residues 132–335 with domain D) activated the expression of reporter genes, but the 42-C2 (residues 159–335 without domain D) did not. These results show that OsbZIP42 contains two potential transcriptional activation domains.
Gene structures and transactivation activity assay of OsbZIP42. a The predicted domains and motifs of OsbZIP42 are shown. bZIP indicates the basic leucine zipper domain. NLS indicates a nuclear localization signal. The five arrowheads represent the conserved phosphorylation sites. The numbers indicate the amino acid position. b Fusion proteins of the GAL4 DNA-binding domain and five portions of OsbZIP42 were checked for their transactivation activity in yeast strain Mav203. The left panel displays schematic diagrams of the various constructs used for the transactivation activity assays. The results of a colony-lift filter assay are shown on the right. 42-FL shows the construct inserted with the full-length CDS of OsbZIP42. 42-N1 (1–128), 42-N2 (1–164), 42-C1 (132–335), and 42-C2 (159–335). 3-AT 3-amino-1,2,4-triazole, D domain D (domain 13)
Analysis of the cis-acting elements in the 5′ regulatory regions of OsbZIP42 and the expression profiles of OsbZIP42
The characterization of the cis-acting region bound by TFs that control gene expression often provides essential information regarding gene function. Thus, we checked the promoter sequence of the OsbZIP42 gene (1500 bp upstream of the putative transcription start site) using the PLACE database (http://www.dna.affrc.go.jp/PLACE/) (Table 1). The expression of the OsbZIP42 gene was analyzed following drought, high salinity and ABA treatments using qRT-PCR (Fig. 2). The OsbZIP42 gene showed the highest expression in spikelets (Fig. 2a), and its promoter region contained two cis-acting elements associated with seed expression, the prolamin box and the RY repeat (Table 1). We performed qRT-PCR analysis at various stages of panicle and seed development (Fig. 2b). The transcript levels of OsbZIP42 showed the highest levels during the early stage and decreased gradually during seed development.
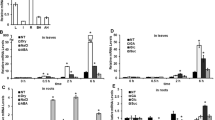
Expression analyses of the OsbZIP42 gene. a The transcript levels of OsbZIP42 in different organs under normal conditions were analyzed using qRT-PCR. The relative transcript abundances of the tested genes were quantified relative to those in the leaves. The indicated organs, young leaves (L), internodes (I), young roots (R), prepollinated spikelets (BH), and postpollinated spikelets (AH), are shown. b The transcript levels of OsbZIP42 at different reproductive stages were determined using qRT-PCR. The relative transcript abundances of the OsbZIP42 gene were quantified relative to the S4 stage. The young panicles and developmental seeds were divided into three (P3 3–5 cm, P4 10–15 cm, P5 15–20 cm of panicle length) and five groups (S1 0–2 DAP, S2 3–4 DAP, S3 5–10 DAP, S4 11–20 DAP, S5 5–29 DAP), respectively. The transcript levels of OsbZIP42 under stress conditions were analyzed in the leaves (c) and roots (d) using qRT-PCR. The relative transcript abundances of the tested genes were quantified relative to those of the NT (nontreated control) at 0 h. Three-week-old plants grown in soil were hydroponically adapted in water for 3 days and then transferred to fresh water containing either 250 mM NaCl (NaCl) or 100 µM ABA (ABA) in a greenhouse, or air-dried (Dry). NT indicates nontreated control plants. Total RNA was prepared during each condition at the indicated time points. The OsUbi1 gene was used as a reference gene. The data are presented as the mean ± SE values (n = 3) from three independent experiments. Asterisks indicate statistically significant differences compared to the NT using Student’s t test (*P < 0.05)
Several functionally significant cis-acting elements associated with ABA and stress responses, such as the abscisic acid-responsive element (ABRE), the drought-responsive element (DRE), the GCC box, the MYB recognition site (MYBRS), the MYC recognition site (MYCRS), and the WRKY recognition site (WBOX), were identified in the OsbZIP42 gene promoter. Under ABA treatment, the transcript level of OsbZIP42 was increased in the leaves and roots (Fig. 2c, d). Unlike other members of rice subgroup II bZIP TFs (Lu et al. 2009), the expression of OsbZIP42 was only slightly repressed by drought and high-salinity stress. The cis-acting elements associated with sugar responses, such as the sugar-repressive element (SRE), the sucrose-responsive element (SURE), and the TATCCAY motif, were identified in the OsbZIP42 gene promoter. To verify the regulation of OsbZIP42 gene expression by GA and sugar, we performed qRT-PCR under exogenous 100 μM GA, 100 mM glucose or 100 mM sucrose treatments (Fig. 3). The expression of OsbZIP42 was slightly affected by the exogenous GA and sugar treatments.
The expression patterns of the OsbZIP42 gene under gibberellin, glucose, and sucrose treatments were analyzed using qRT-PCR. The relative transcript abundances of the tested genes in the leaves (a) and roots (b) were quantified relative to those of the NT at 0 h. Three-week-old plants grown in soil were hydroponically adapted in water for 3 days and then transferred to fresh water containing either 100 μM gibberellin (GA), 100 mM glucose (Glc) or 100 mM sucrose (Suc) in a greenhouse. NT indicates nontreated control plants. Total RNA was prepared during each condition at the indicated time points. The OsUbi1 gene was used as a reference gene. The data are presented as the mean ± SE values (n = 3) from three independent experiments. Asterisks indicate statistically significant differences compared to the NT using Student’s t test (*P < 0.05)
Yeast two-hybrid assays of OsbZIP42 and SAPK members
The stress-/ABA-activated protein kinases (SAPKs) or SnRK2 has been suggested to phosphorylate ABFs/AREBs in Arabidopsis (Furihata et al. 2006; Fujii and Zhu 2009) and OsbZIP46 in rice (Tang et al. 2012). We investigated the interaction between OsbZIP42 and ten putative rice SAPK family members (Kobayashi et al. 2004) using yeast two-hybrid assays. OsbZIP42 did not interact with any SAPKs (Supplementary Fig. S1). Unlike other bZIP TFs, the OsbZIP42 protein contained two transcriptional activation domains (Fig. 1b); therefore, we investigated the interaction activity of 42-N2 and 42-C2 separately (Fig. 4). Only 42-N2 could interact with SAPK4. This result suggests that OsbZIP42 may require another modification and/or conformational change mechanism to interact with SAPK4. To explain this result, the three-dimensional protein structure of OsbZIP42 was predicted using the I-TASSER online server (http://zhanglab.ccmb.med.umich.edu/I-TASSER/) (Supplementary Fig. S2). The predicted models of OsbZIP12, 42, 46, and 62 TFs exhibited very similar protein structures. In contrast, OsbZIP42 showed a different structure from those other models (Supplementary Fig. S2). OsbZIP12, 46, and 62 contained several well-developed long alpha helical structures, but OsbZIP42 contained very short alpha helical structures. This contrast in structure could lead to different interaction activities with SAPKs.
Interaction between OsbZIP42 and rice SAPKs. Yeast two-hybrid assays of OsbZIP42-N2/OsbZIP42-C2 and SAPK members were performed. The results of a colony-lift filter assay are shown (X-gal assay). Control P and control N indicate the positive and negative controls, respectively. 3-AT 3-amino-1,2,4-triazole, SD-LTH minimal medium–Leu–Trp–His
Overexpression of OsbZIP42 leads to seedlings that are hypersensitive to ABA
To examine the roles of OsbZIP42 in rice plants, we constructed rice transformation plasmids (Fig. 5a), and transgenic rice plants overexpressing OsbZIP42 (OsbZIP42-OX) were obtained via the Agrobacterium-mediated transformation method. T1–T4 generation seeds were collected from individual transgenic plants, and two independent homozygous T4 generation lines were subjected to further analysis. The ectopic expression of the transgenes in the OsbZIP42-OX plants was confirmed using qRT-PCR (Fig. 5b). The transcript levels of OsbZIP42 were enhanced in the individual transgenic lines relative to the WT control.
Increased ABA sensitivity of OsbZIP42 overexpression plants during germination and seedling stages. a The overexpression plasmid consisting of the constitutively expressed OsCc1 promoter linked to the OsbZIP42 coding region, which is transcribed with the 3′ UTR region of the potato proteinase inhibitor II gene (3′ pinII), also includes a Basta resistance (Bar) expression cassette containing the 35S promoter, the bar-coding region and the 3′ UTR region of the nopaline synthase gene (3′ nos). b The transcript levels of the OsbZIP42 gene in the leaves of T4 transgenic (OsbZIP42-OX) and control rice plants (WT) were analyzed using qRT-PCR. Those in the transgenic lines were quantified relative to those in the WT plants. The OsUbi1 gene was used as a reference gene. The data are presented as the mean ± SE values (n = 3) from two independent experiments. c Germination assay of OsbZIP42-OX transgenic and WT seeds in half-strength MS medium containing 0, 2, and 6 μM ABA. The germination rate of the treated seeds was calculated after 5 days. d The OsbZIP42-OX transgenic and WT control plants were grown in MS0 and MS0 containing 5 µM ABA. Shoot (e) and root length (f) measurements of the rice plants are shown in panel c. The data are presented as the mean ± SE values (n = 15). Asterisks indicate statistically significant differences compared to the WT using Student’s t test (*P < 0.05)
The bZIP TFs are involved in various developmental and physiological processes in response to ABA signaling and play important roles in ABA-dependent stress responses (Llorca et al. 2014). To test whether ABA has an effect on the germination of OsbZIP42-OX plants, WT and transgenic seeds were germinated in half-strength MS medium with ABA (0, 2, and 6 μM). The germination rate of the OsbZIP42-OX transgenic lines was similar to that of WT control at 0 μM ABA (Fig. 5c). However, the germination rate of the OsbZIP42-OX transgenic lines was significantly decreased at 2 and 6 μM ABA. These results suggested that the sensitivity of seed germination of OsbZIP42-OX transgenic rice plants to ABA was increased. Next, we tested the effect of ABA on seedling development in OsbZIP42-OX plants. WT and transgenic seedlings were grown for 2 weeks in half-strength MS solid media with or without ABA (Fig. 5d), and the lengths of the shoots and roots were measured (Fig. 5e, f). There were no significant differences in shoot length between the WT and transgenic rice plants grown in the medium without ABA. The shoot and root lengths of the WT plants grown in the medium with 5 µM ABA were reduced by approximately 22% and 25% of those of the control plants grown in the medium without ABA, respectively. However, the shoot and root lengths of the OsbZIP42-OX transgenic lines were significantly reduced by 6–12% and 8–16%, respectively, compared to those of the control. These data indicate that the overexpression of OsbZIP42 makes transgenic seedlings hypersensitive to ABA in comparison to WT plants, and thus OsbZIP42 is involved in ABA signaling in rice.
Expression analysis of ABA signal pathway marker genes in the transgenic plants
The ABA-dependent phosphorylation of bZIP TFs by SnRK2 or SAPK family protein kinases and dephosphorylation by PP2C-type protein phosphatases (PP2C) regulate their activation in plants. The expression of late-embryogenesis-abundant (LEA) genes, which participate in the acquisition of desiccation tolerance, could be induced by the activated ABF/ABRE system (Umezawa et al. 2010). To specify whether OsbZIP42 affects ABA signaling, three genes (PP2C, LEA3 and Rab16) involved in ABA signal transduction were selected for expression level comparison between the WT and transgenic plants under ABA treatment; these included a PP2C family gene (ABI2, Os01g0583100) and two downstream genes (LEA3, Os05g0542500; Rab16, Os11g0454300). Under normal conditions, the three genes showed no differences in transcript levels between the WT and OsbZIP42-OX lines. Under 100 µM ABA, the transcript levels of ABI2 (PP2C family gene) were decreased in the OsbZIP42-OX transgenic lines compared to those in the WT control (Fig. 6a). This result suggests that OsbZIP42 negatively regulates PP2C in the ABA signaling pathway. In the OsbZIP42-OX lines, the transcript levels of LEA3 were 9.5- and 6.5-fold increased within 2 h compared to those of the WT controls. In contrast, these levels were 0.41- and 0.54-fold decreased at 6 h compared to those of the WT controls (Fig. 6b). The transcript levels of Rab16 were 10.9- and 14.5-fold increased within 2 h compared to those of the WT controls, and they showed no significant difference at 6 h (Fig. 6c). These results support the involvement of OsbZIP42 in the ABA signal transduction pathway in rice. Furthermore, OsbZIP42 is involved in early ABA-responsive gene induction.
Expression analysis of the ABA signaling pathway marker genes in OsbZIP42-OX transgenic plants. The relative transcript abundances of the ABI2 (a), LEA3 (b) and Rab16 (c) genes in the leaves of the OsbZIP42-OX transgenic and WT plants under 100 µM ABA treatment were quantified relative to those in the WT plants at 0 h. Total RNA templates were prepared at the indicated time points. The OsUbi1 gene was used as a reference gene. The data are presented as the mean ± SE values (n = 3) from three independent experiments. Asterisks indicate statistically significant differences compared to the WT using Student’s t test (*P < 0.05)
Abiotic stress tolerance of transgenic plants
To examine the responses of the OsbZIP42-OX plants to abiotic stress during the germination stage, the seeds of WT and transgenic plants were germinated in water containing 400 mM mannitol or 250 mM NaCl, and then the germination rates were measured (Fig. 7). There were no differences in the germination rates for WT and OsbZIP42-OX seeds under normal and 250 mM NaCl conditions. Under 400 mM mannitol, the OsbZIP42-OX seeds exhibit increased germination rates compared to the WT.
To evaluate the responses of the OsbZIP42-OX plants to water deficit, 4-week-old WT and OsbZIP42-OX seedlings were subjected to drought stress for 2 days, followed by rewatering. The WT and transgenic plants both exhibited leaf rolling and other stress-induced symptoms by 2 days of drought stress. During rewatering, the transgenic lines showed a faster recovery from drought stress and more stimulated growth than did the severely injured WT plants (Fig. 8a, R3 and R10). The survival rate of the OsbZIP42-OX transgenic lines was 83–94% (Fig. 8b). In contrast, the survival rate of the WT plants was only 40%. To further verify the stress tolerance of the OsbZIP42-OX plants, we measured the variations in the chlorophyll fluorescence ratio (Fv/Fm) after drought and cold stress treatments. Healthy plants typically achieve a maximum Fv/Fm value of approximately 0.8, and lower values are observed in plants that are exposed to abiotic stress factors. The values of Fv/Fm were 1.39- to 1.62-fold higher (P < 0.001) in the OsbZIP42-OX plants than in the WT plants under drought stress conditions (Fig. 8c). Under cold stress conditions, the OsbZIP42-OX plants showed 1.33- to 1.7-fold higher values for Fv/Fm than the WT plants (Fig. 8d). The results of the phenotypic and chlorophyll fluorescence analyses of the OsbZIP42-OX plants suggest that OsbZIP42 confers drought tolerance to rice.
Overexpression of OsbZIP42 enhanced abiotic stress tolerance in rice. a The OsbZIP42-OX transgenic plants had an increased tolerance to abiotic stresses during the vegetative stage. The rice plants were subjected to drought stress by draining the water out of the pots (D0), and water was withheld for 2 days (D2) in a greenhouse. After 2 days of drought stress, the rice plants were rehydrated by resupplying water (R0). R3 and R10 indicate water supply for 3 and 10 days, respectively. Photographs were taken on the indicated days. b The survival rates of the OsbZIP42-OX transgenic and WT plants after drought stress. The data are presented as the mean ± SE values (n = 20) from two independent experiments. Changes in the chlorophyll fluorescence (Fv/Fm) of the leaf discs under drought (c) or cold stress (d). The data are presented as the mean ± SE values (n = 9) from two independent experiments. Asterisks indicate statistically significant differences compared to the WT using Student’s t test (*P < 0.05, ***P < 0.001)
Discussion
Plant bZIP TFs have been reported to be involved in various biological processes, including environmental stress responses. The Arabidopsis bZIP TFs, ABFs/AREBs, which play important roles in cooperatively regulating ABRE-dependent ABA signaling involved in abiotic stress tolerance, are classified as group A (Jakoby et al. 2002; Kang et al. 2002; Yoshida et al. 2010). The rice group E bZIP TFs are highly homologous to the Arabidopsis group A bZIP family members. Rice group E includes 11 members and is divided into three subgroups (Nijhawan et al. 2008; Lu et al. 2009). The members of subgroup I, OsABF1 (OsbZIP12), and subgroup III, OsABI5 (OsbZIP10), OsbZIP23, ABL1 (OsbZIP46), TRAB1 (OsbZIP66), and OsbZIP72, have been previously studied. Arabidopsis DPBF4 (AtbZIP12) and AREB3 (AtbZIP66) are classified as subgroup II members (Lu et al. 2009). However, the functions of subgroup II have not been well studied. In this study, we report on the functional characterization of OsbZIP42, which belongs to subgroup II. Our data suggest that OsbZIP42 is involved in ABA signaling and drought stress responses and is regulated through transcriptional control and posttranslational modification in rice.
Our results show that OsZIP42 contains two potential activation domains that are located in the N-terminus (residues 1–128) and C-terminus (residues 132–335). It has been reported that human LZIP or Luman, which is a basic leucine zipper protein that interacts with host cell factor-1, contains two transcriptional activation domains located in the N and C-termini (Luciano and Wilson 2000). However, the function of the C-terminal transcriptional activation domain has not been studied. In yeast assays, the full-length OsbZIP42 and N-terminal fragment 42-N1, which contains domain D, showed no transactivation activity in yeast (Fig. 1). Domain D is proposed as a negative regulatory domain in OsbZIP46, as the deletion of domain D from OsbZIP46 results in constitutive transactivation activity in yeast. It has been suggested that deletion of domain D may cause a conformational change that mimics the changes resulting from posttranslational modification (Tang et al. 2012). In contrast, the C-terminal fragment 42-C1, which contains domain D and the bZIP domain, exhibited transactivation activity in yeast. These results suggest that domain D may act only on the N-terminal transcriptional activation domain, which is commonly found in bZIP TFs, and/or has another function in addition to being a negative regulator.
Full-length OsbZIP42 showed no transactivation activity in the yeast assays (Fig. 1b). The OsbZIP42 gene was induced only by ABA treatment in the leaves and roots (Fig. 2c, d). Furthermore, the target genes for OsbZIP42, LEA3 and Rab16, were significantly more induced in the OsbZIP42-OX plants compared to in the WT plants under ABA treatment but showed no difference under normal conditions (Fig. 6). These results suggest that the transactivation activities of OsbZIP42 rely primarily on ABA-dependent modifications. Several Arabidopsis group A and rice group E bZIP TFs were reported to be activated by ABA-dependent phosphorylation (Uno et al. 2000; Kagaya et al. 2002; Schütze et al. 2008; Tang et al. 2012). These reports suggested that OsbZIP42 may also require ABA-dependent phosphorylation for transactivation activity. In the ABA signaling pathway, the phosphorylation of ABF/ABEB by SnRK2 in Arabidopsis and of TRAB1 by SAPK in rice have been well characterized (Kobayashi et al. 2005; Furihata et al. 2006; Fujii and Zhu 2009). TRAB1 can be activated via ABA-dependent phosphorylation. Its Ser-102 residue is critical for this function, and this residue is phosphorylated in response to ABA or hyperosmotic stress (Kagaya et al. 2002; Kobayashi et al. 2005). OsbZIP42 contains phosphorylation residues, and these conserved phosphorylation sites have been shown to be related to the ABA response and stress signaling.
Our results show that full-length OsbZIP42 did not interact with any SAPKs in yeast, but the N-terminal fragment 42-N2 (without domain D) interacted with SAPK4 (Supplementary Fig. S1, Fig. 4). This result suggests that OsbZIP42 likely requires additional modifications and/or conformational change mechanisms to interact with SAPK4. To investigate the effect of the structure of OsbZIP42 on its interactions with SAPKs, the predicted protein structure of OsbZIP42 was compared to the structures of other OsbZIP TFs using the I-TASSER online server (http://zhanglab.ccmb.med.umich.edu/I-TASSER/) (Supplementary Fig. S2). OsbZIP12, 46, and 62 contained several well-developed long alpha helical structures, but OsbZIP42 contained very short alpha helical structures. The results suggest that OsbZIP42 may have a different structure from those of other OsbZIP TFs. In addition, this difference in structure could require an additional modification and/or conformational change mechanism for OsbZIP42 to interact with SAPK4.
Several members of the rice group E bZIP TFs have been reported to be involved in the ABA signaling pathway. OsbZIP23, 46, and 72 are positive regulators of ABA signaling (Xiang et al. 2008; Lu et al. 2009; Tang et al. 2012). The overexpression of OsbZIP42 induced the transgenic rice plants to become hypersensitive to ABA, indicating that OsbZIP42 plays a role in the ABA signaling pathway. Under ABA conditions, the LEA3 and Rab16 genes, which are known to be downstream of ABFs in the ABA signaling pathway, were rapidly induced by ABA in the OsbZIP42-OX transgenic plants compared to in the WT plants (Fig. 6). These genes showed no significant difference or decrease at 6 h. These results suggest that OsbZIP42 may be involved early in the ABA signaling pathway. Our results show that the overexpression of OsbZIP42 gene in transgenic rice plants confers increased tolerance to drought stress during germination and in seedlings (Figs. 7, 8). Differences in the chlorophyll fluorescence analysis of the transgenic plants further support the increased drought tolerance phenotypes. The SnRK2 family is known to comprise the principal positive regulators of ABA and osmotic stress that act via the phosphorylation of AREB/ABF TFs (Fujita et al. 2013). The SnRK2 family consists of ten genes classified into three subclasses in Arabidopsis and rice (Kobayashi et al. 2004). SnRK2 subclass I member SAPK4, which interacts with OsbZIP42 (Fig. 4), has been reported to regulate stress-responsive gene expression in rice (Kobayashi et al. 2004; Diédhiou et al. 2008). In rice, only SnRK2 subclass III members are activated by ABA, and subclass I and II members are activated only by osmotic and salt stress (Kobayashi et al. 2004). Thus, OsbZIP42 could be phosphorylated in response to ABA by some other kinases. Calcium-dependent protein kinases (CPKs) have been found to phosphorylate ABF/AREBs (Choi et al. 2005; Zhu et al. 2007). CPKs are central components of ABA signal transduction (Choi et al. 2005). A SnRK3‐ type protein kinase CIPK11 is also found to phosphorylate ABI5 to regulate ABA response (Zhou et al. 2015). SAPK4 has been reported to interact with OsABF1/OsABI5/OREB1, which is highly phosphorylated in response to ABA and abiotic stress signals, in yeast and plants (Ding et al. 2009). The overexpression of SAPK4 resulted in improved germination, growth and development under salt stress in both seedlings and mature plants. SAPK4-regulated genes are involved in ion homeostasis and oxidative stress response, vacuolar H+-ATPase, the Na+/H+ antiporter NHX1, the Cl− channel OsCLC1 and a catalase (Diédhiou et al. 2008). The germination assay showed that overexpression of OsbZIP42 resulted in improved germination under mannitol treatment but not under NaCl treatment (Fig. 8). These results suggest that OsbZIP42 is activated via ABA-dependent phosphorylation by SAPK4 and plays a role in ABA-dependent drought stress responses.
It has been determined that bZIP TFs bind to DNA predominantly as homo- or heterodimers (Landschulz et al. 1988; Nijhawan et al. 2008) and that homo- and heterodimers of bZIPs control gene expression during a variety of stress responses in plants (Schütze et al. 2008; Dietrich et al. 2011). Intact OsbZIP42 did not form homodimers (Supplementary Fig. S3). However, it may form heterodimers, and modified OsbZIP42 could form homodimers. This result could explain the enhancement in drought tolerance due to OsbZIP42 overexpression, even though the OsbZIP42 gene is not induced by drought and high-salt stresses. Further studies are required to elucidate the heterodimerization and modification mechanisms of OsbZIP42 that are responsible for drought signal transduction.
Author contribution statement
YHL and SIS conceived and designed the experiments; JJ and YHL performed the experiments; JJ and SIS wrote the manuscript.
Abbreviations
- SAPK:
-
Stress-/ABA-activated protein kinase
- SnRK2:
-
SNF1-related type 2 protein kinase
- TF:
-
Transcription factor
References
Amir Hossain M, Lee Y, Cho JI, Ahn CH, Lee SK, Jeon JS, Kang H, Lee CH, An G, Park PB (2010) The bZIP transcription factor OsABF1 is an ABA responsive element binding factor that enhances abiotic stress signaling in rice. Plant Mol Biol 72(4–5):557–566. https://doi.org/10.1007/s11103-009-9592-9
Bhaskara GB, Nguyen TT, Verslues PE (2012) Unique drought resistance functions of the highly ABA-induced clade A protein phosphatase 2Cs. Plant Physiol 160(1):379–395. https://doi.org/10.1104/pp.112.202408
Choi H, Hong J, Ha J, Kang J, Kim SY (2000) ABFs, a family of ABA-responsive element binding factors. J Biol Chem 275(3):1723–1730
Choi HI, Park HJ, Park JH, Kim S, Im MY, Seo HH, Kim YW, Hwang I, Kim SY (2005) Arabidopsis calcium-dependent protein kinase AtCPK32 interacts with ABF4, a transcriptional regulator of abscisic acid-responsive gene expression, and modulates its activity. Plant Physiol 139(4):1750–1761
Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679. https://doi.org/10.1146/annurev-arplant-042809-112122
Deppmann CD, Thornton TM, Utama FE, Taparowsky EJ (2003) Phosphorylation of BATF regulates DNA binding: a novel mechanism for AP-1 (activator protein-1) regulation. Biochem J 374(Pt 2):423–431. https://doi.org/10.1042/BJ20030455
Diédhiou CJ, Popova OV, Dietz KJ, Golldack D (2008) The SNF1-type serine-threonine protein kinase SAPK4 regulates stress-responsive gene expression in rice. BMC Plant Biol 8:49. https://doi.org/10.1186/1471-2229-8-49
Dietrich K, Weltmeier F, Ehlert A, Weiste C, Stahl M, Harter K, Dröge-Laser W (2011) Heterodimers of the Arabidopsis transcription factors bZIP1 and bZIP53 reprogram amino acid metabolism during low energy stress. Plant Cell 23(1):381–395. https://doi.org/10.1105/tpc.110.075390
Ding X, Richter T, Chen M et al (2009) A rice kinase–protein interaction map. Plant Physiol 149(3):1478–1492
Djamei A, Pitzschke A, Nakagami H, Rajh I, Hirt H (2007) Trojan horse strategy in Agrobacterium transformation: abusing MAPK defense signaling. Science 318(5849):453–456. https://doi.org/10.1126/science.1148110
Finkelstein RR, Lynch TJ (2000) The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 12(4):599–609
Fujii H, Zhu JK (2009) Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc Natl Acad Sci USA 106(20):8380–8385. https://doi.org/10.1073/pnas.0903144106
Fujita Y, Fujita M, Satoh R, Maruyama K, Parvez MM, Seki M, Hiratsu K, Ohme-Takagi M, Shinozaki K, Yamaguchi-Shinozaki K (2005) AREB1 is a transcription activator of novel ABRE-dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 17(12):3470–3488. https://doi.org/10.1105/tpc.105.035659
Fujita M, Fujita Y, Noutoshi Y, Takahashi F, Narusaka Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Crosstalk between abiotic and biotic stress responses: a current view from the points of convergence in the stress signaling networks. Curr Opin Plant Biol 9(4):436–442. https://doi.org/10.1016/j.pbi.2006.05.014
Fujita Y, Yoshida T, Yamaguchi-Shinozaki K (2013) Pivotal role of the AREB/ABF-SnRK2 pathway in ABRE-mediated transcription in response to osmotic stress in plants. Physiol Plant 147(1):15–27. https://doi.org/10.1111/j.1399-3054.2012.01635.x
Furihata T, Maruyama K, Fujita Y, Umezawa T, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2006) Abscisic acid-dependent multisite phosphorylation regulates the activity of a transcription activator AREB1. Proc Natl Acad Sci USA 103(6):1988–1993. https://doi.org/10.1073/pnas.0505667103
Hirasawa T (1999) Physiological characterization of rice plant for tolerance of water deficit. In: Ito O, O’Toole JC, Hardy B (eds) Genetic improvement of rice for water-limited environments. International Rice Research Institute, Los Banos, pp 89–98
Hobo T, Kowyama Y, Hattori T (1999) A bZIP factor, TRAB1, interacts with VP1 and mediates abscisic acid-induced transcription. Proc Natl Acad Sci USA 96(26):15348–15353
Ishida S, Yuasa T, Nakata M, Takahashi Y (2008) A tobacco calcium-dependent protein kinase, CDPK1, regulates the transcription factor REPRESSION OF SHOOT GROWTH in response to gibberellins. Plant Cell 20(12):3273–3288. https://doi.org/10.1105/tpc.107.057489
Jakoby M, Weisshaar B, Dröge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7(3):106–111
Joo J, Choi HJ, Lee YH, Kim YK, Song SI (2013a) A transcriptional repressor of the ERF family confers drought tolerance to rice and regulates genes preferentially located on chromosome 11. Planta 238(1):155–170. https://doi.org/10.1007/s00425-013-1880-6
Joo J, Lee YH, Kim YK, Nahm BH, Song SI (2013b) Abiotic stress responsive rice ASR1 and ASR3 exhibit different tissue-dependent sugar and hormone-sensitivities. Mol Cells 35(5):421–435. https://doi.org/10.1007/s10059-013-0036-7
Joo J, Lee YH, Song SI (2014) Overexpression of the rice basic leucine zipper transcription factor OsbZIP12 confers drought tolerance to rice and makes seedlings hypersensitive to ABA. Plant Biotechnol Rep 8(6):431–441
Kagaya Y, Hobo T, Murata M, Ban A, Hattori T (2002) Abscisic acid-induced transcription is mediated by phosphorylation of an abscisic acid response element binding factor, TRAB1. Plant Cell 14(12):3177–3189
Kang JY, Choi HI, Im MY, Kim SY (2002) Arabidopsis basic leucine zipper proteins that mediate stress-responsive abscisic acid signaling. Plant Cell 14(2):343–357
Kobayashi Y, Yamamoto S, Minami H, Kagaya Y, Hattori T (2004) Differential activation of the rice sucrose nonfermenting1–related protein kinase2 family by hyperosmotic stress and abscisic acid. Plant Cell 16(5):1163–1177. https://doi.org/10.1105/tpc.019943
Kobayashi Y, Murata M, Minami H, Yamamoto S, Kagaya Y, Hobo T, Yamamoto A, Hattori T (2005) Abscisic acid-activated SNRK2 protein kinases function in the gene-regulation pathway of ABA signal transduction by phosphorylating ABA response element-binding factors. Plant J 44(6):939–949. https://doi.org/10.1111/j.1365-313X.2005.02583.x
Landschulz WH, Johnson PF, McKnight SL (1988) The leucine zipper: a hypothetical structure common to a new class of DNA binding proteins. Science 240(4860):1759–1764
Lee S, Shuman JD, Guszczynski T, Sakchaisri K, Sebastian T, Copeland TD, Miller M, Cohen MS, Taunton J, Smart RC, Xiao Z, Yu LR, Veenstra TD, Johnson PF (2010) RSK-mediated phosphorylation in the C/EBPβ leucine zipper regulates DNA binding, dimerization, and growth arrest activity. Mol Cell Biol 30(11):2621–2635. https://doi.org/10.1128/MCB.00782-09
Llorca CM, Potschin M, Zentgraf U (2014) bZIPs and WRKYs: two large transcription factor families executing two different functional strategies. Front Plant Sci 5:169. https://doi.org/10.3389/fpls.2014.00169
Lu G, Gao C, Zheng X, Han B (2009) Identification of OsbZIP72 as a positive regulator of ABA response and drought tolerance in rice. Planta 229(3):605–615. https://doi.org/10.1007/s00425-008-0857-3
Luciano RL, Wilson AC (2000) N-terminal transcriptional activation domain of LZIP comprises two LxxLL motifs and the host cell factor-1 binding motif. Proc Natl Acad Sci USA 97(20):10757–10762. https://doi.org/10.1073/pnas.190062797
Nijhawan A, Jain M, Tyagi AK, Khurana JP (2008) Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol 146(2):333–350. https://doi.org/10.1104/pp.107.112821
Schütze K, Harter K, Chaban C (2008) Post-translational regulation of plant bZIP factors. Trends Plant Sci 13(5):247–255. https://doi.org/10.1016/j.tplants.2008.03.002
Shinozaki K, Yamaguchi-Shinozaki K (1997) Gene expression and signal transduction in water-stress response. Plant Physiol 115(2):327–334
Smeekens S, Ma J, Hanson J, Rolland F (2010) Sugar signals and molecular networks controlling plant growth. Curr Opin Plant Biol 13(3):274–279. https://doi.org/10.1016/j.pbi.2009.12.002
Tang N, Zhang H, Li X, Xiao J, Xiong L (2012) Constitutive activation of transcription factor OsbZIP46 improves drought tolerance in rice. Plant Physiol 158(4):1755–1768. https://doi.org/10.1104/pp.111.190389
Tuong TP, Bouman BAM (2003) Rice production in water scarce environments. In: Kijne JW, Barker R, Molden D (eds) Water productivity in agriculture: limits and opportunities for improvement. CAB International, Wallingford, pp 53–67
Umezawa T, Sugiyama N, Mizoguchi M, Hayashi S, Myouga F, Yamaguchi-Shinozaki K, Ishihama Y, Hirayama T, Shinozaki K (2009) Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. Proc Natl Acad Sci USA 106(41):17588–17593. https://doi.org/10.1073/pnas.0907095106
Umezawa T, Nakashima K, Miyakawa T, Kuromori T, Tanokura M, Shinozaki K, Yamaguchi-Shinozaki K (2010) Molecular basis of the core regulatory network in ABA responses: sensing, signaling and transport. Plant Cell Physiol 51(11):1821–1839. https://doi.org/10.1093/pcp/pcq156
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97(21):11632–11637. https://doi.org/10.1073/pnas.190309197
Verslues PE, Zhu JK (2005) Before and beyond ABA: upstream sensing and internal signals that determine ABA accumulation and response under abiotic stress. Biochem Soc Trans 33(Pt 2):375–379. https://doi.org/10.1042/BST0330375
Vlad F, Rubio S, Rodrigues A, Sirichandra C, Belin C, Robert N, Leung J, Rodriguez PL, Laurière C, Merlot S (2009) Protein phosphatases 2C regulate the activation of the Snf1-related kinase OST1 by abscisic acid in Arabidopsis. Plant Cell 21(10):3170–3184. https://doi.org/10.1105/tpc.109.069179
Xiang Y, Tang N, Du H, Ye H, Xiong L (2008) Characterization of OsbZIP23 as a key player of the basic leucine zipper transcription factor family for conferring abscisic acid sensitivity and salinity and drought tolerance in rice. Plant Physiol 148(4):1938–1952. https://doi.org/10.1104/pp.108.128199
Yoshida T, Fujita Y, Sayama H, Kidokoro S, Maruyama K, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2010) AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J 61(4):672–685. https://doi.org/10.1111/j.1365-313X.2009.04092.x
Zg E, Zhang YP, Zhou JH, Wang L (2014) Mini review roles of the bZIP gene family in rice. Genet Mol Res 13(2):3025–3036. https://doi.org/10.4238/2014.April.16.11
Zhou X, Hao H, Zhang Y, Bai Y, Zhu W, Qin Y, Yuan F, Zhao F, Wang M, Hu J, Xu H, Guo A, Zhao H, Zhao Y, Cao C, Yang Y, Schumaker KS, Guo Y, Xie CG (2015) SOS2-LIKE PROTEIN KINASE5, an SNF1-RELATED PROTEIN KINASE3-type protein kinase, is important for abscisic acid responses in Arabidopsis through phosphorylation of ABSCISIC ACID-INSENSITIVE5. Plant Physiol 168(2):659–676
Zhu SY, Yu XC, Wang XJ, Zhao R, Li Y, Fan RC, Shang Y, Du SY, Wang XF, Wu FQ, Xu YH, Zhang XY, Zhang DP (2007) Two calcium-dependent protein kinases, CPK4 and CPK11, regulate abscisic acid signal transduction in Arabidopsis. Plant Cell 19(10):3019–3036
Zou M, Guan Y, Ren H, Zhang F, Chen F (2008) A bZIP transcription factor, OsABI5, is involved in rice fertility and stress tolerance. Plant Mol Biol 66(6):675–683. https://doi.org/10.1007/s11103-008-9298-4
Acknowledgements
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2017R1D1A1B03030725).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Joo, J., Lee, Y.H. & Song, S.I. OsbZIP42 is a positive regulator of ABA signaling and confers drought tolerance to rice. Planta 249, 1521–1533 (2019). https://doi.org/10.1007/s00425-019-03104-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-019-03104-7