Abstract
Background
Circular RNA migration and invasion inhibitory protein (circMIIP) is reported to be upregulated in non-small cell lung cancer (NSCLC) tissues compared with normal tissues. However, the role and working mechanism of circMIIP in NSCLC progression remain largely unclear.
Methods
Cell proliferation ability was analyzed by colony formation assay, cell counting kit-8 (CCK-8) assay, and 5-ethynyl-2′-deoxyuridine assay. Cell apoptosis was assessed by flow cytometry. Transwell assays were performed to analyze the migration and invasion abilities of NSCLC cells. The interaction between microRNA-766-5p (miR-766-5p) and circMIIP or family with sequence similarity 83A (FAM83A) was validated by dual-luciferase reporter assay and RNA immunoprecipitation assay. Xenograft tumor model was established to analyze the role of circMIIP on tumor growth in vivo.
Results
CircMIIP was highly expressed in NSCLC tissues and cell lines. CircMIIP knockdown restrained the proliferation, migration and invasion and induced the apoptosis of NSCLC cells. CircMIIP acted as a molecular sponge for miR-766-5p, and circMIIP silencing-mediated anti-tumor effects were largely overturned by the knockdown of miR-766-5p in NSCLC cells. miR-766-5p interacted with the 3’ untranslated region (3′UTR) of FAM83A, and FAM83A overexpression largely reversed miR-766-5p accumulation-induced anti-tumor effects in NSCLC cells. CircMIIP competitively bound to miR-766-5p to elevate the expression of FAM83A in NSCLC cells. CircMIIP knockdown significantly restrained xenograft tumor growth in vivo.
Conclusion
CircMIIP promoted cell proliferation, migration and invasion and suppressed cell apoptosis in NSCLC cells through mediating miR-766-5p/FAM83A axis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-small cell lung cancer (NSCLC) accounts for 85% of all lung cancer cases and is one of the leading causes of cancer-associated deaths [1, 2]. Although significant advances have been made in the treatment of NSCLC, including surgery, radiotherapy and chemotherapy, the five-year survival rate of NSCLC patients remains only 16% [3]. Therefore, an in-depth understanding of the molecular mechanisms of NSCLC progression is helpful for NSCLC therapy.
Circular RNAs (circRNAs) are a class of endogenous non-coding RNAs with covalently closed loop structure [4]. Accumulating evidence have demonstrated that circRNAs play important roles in multiple malignancies, including colorectal cancer [5], ovarian cancer [6], hepatocellular carcinoma [7], and NSCLC [8]. CircRNA migration and invasion inhibitory protein (circMIIP, ID: hsa_circ_0009932) is derived from exon 4–10 of the MIIP gene, whose mature spliced sequence length is 946 nt. Circular RNA microarray analysis by a previous study showed that circMIIP expression is notably upregulated in NSCLC tissues compared with normal tissues [9]. However, the role and mechanism of circMIIP in NSCLC progression remain largely undefined.
In this study, we aimed to investigate the expression and mechanism of circMIIP in NSCLC progression, with a view to finding new therapeutic targets for NSCLC.
Materials and Methods
Clinical Tissue Samples
Sixty-five pairs of NSCLC tumor tissues and adjacent normal tissues (at least 2 cm away from tumor border) were collected from NSCLC patients who underwent surgical resection at Affiliated Hospital of Guizhou Medical University, and all tissues were examined by two pathologists. None of the patients had received radiotherapy or chemotherapy before the surgery. All tissue samples were immediately frozen in liquid nitrogen and then stored in a − 80 °C refrigerator. This research was approved by the Ethics Committee of Affiliated hospital of Guizhou medical university. All patients had signed the written informed consent before the surgery.
Cell Culture and Transfection
Human NSCLC cell lines (NCI-H23 and A549) and normal human lung epithelial cell line BEAS-2B were purchased from Chinese Academy of Sciences (Shanghai, China) and cultured with Dulbecco’s Modified Eagle Medium (DMEM; Invitrogen, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS; Solarbio, Shanghai, China) at 37 °C with 5% CO2.
CircMIIP overexpression plasmid in pCD5-ciR vector (circMIIP), pCD5-ciR empty vector (vector), small interfering RNA (siRNA) targeting circMIIP (si-circMIIP#1, si-circMIIP#2, or si-circMIIP#3), and si-NC were purchased from Invitrogen. microRNA-766-5p (miR-766-5p) mimics or inhibitor (miR-766-5p or anti-miR-766-5p) and the controls (miR-NC or anti-NC) were obtained from RiboBio (Guangzhou, China). Family with sequence similarity 83A (FAM83A) overexpression plasmid (FAM83A) was constructed by cloning its complementary DNA (cDNA) into pcDNA3.1 vector (Invitrogen). Cell transfection was conducted using lipofectamine 2000 reagent (Invitrogen).
Reverse Transcription-Quantitative Polymerase Chain Reaction (RT-qPCR)
RNA samples were extracted from tissues and cells with Trizol reagent (Invitrogen), and complementary DNA (cDNA) was synthesized using the PrimeScript RT polymerase Kit (Qiagen, Frankfurt, Germany) or miScript II RT Kit (Qiagen). The Plus SYBR real-time PCR mixture (LMAI Bio, Shanghai, China) was utilized to perform qPCR with cDNA as the template. The expression levels were normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH; for circMIIP and FAM83A) or U6 small nuclear RNA (for miR-766-5p) and calculated by the 2−ΔΔCt method. The primers are shown in Sup. Table 1.
Colony Formation Assay
Transfected NCI-H23 and A549 cells were seeded onto the 12-well plates at 150 cells/well. Cells were incubated for 14 d to form visible colonies, and the culture medium was replenished every 4 d. After immobilizing with 4% paraformaldehyde (Sigma, St. Louis, MO, USA) and staining with 0.1% crystal violet solution (Solarbio), the colonies were washed, air-fried, imaged and then manually counted.
Cell Counting Kit-8 (CCK-8) Assay
NCI-H23 and A549 cells were seeded onto 96-well plates to settle down. NSCLC cells were incubated with 10 µL CCK-8 reagent (Beyotime, Shanghai, China) after transfection for 24 h, 48 h, or 72 h. The absorbance at 450 nm was examined using a microplate reader (Olympus, Tokyo, Japan).
5-Ethynyl-2’-Deoxyuridine (EdU) Assay
After transfection for 48 h, NCI-H23 and A549 cells were incubated with 50 μM EdU (RiboBio) for 2 h. After immobilizing with 4% paraformaldehyde and 2 mg/mL glycine, NSCLC cells were incubated with 1 × Apollo staining solution for 30 min. Cell nuclei was stained with DAPI, and the fluorescence images were captured under a fluorescence microscope (Leica, Wetzlar, Germany). The fluorescence images were merged using Adobe Photoshop software. ImageJ software (Bio-Rad, Hercules, CA, USA) was used to quantify the numbers of DAPI-positive cells and EdU-positive cells, and we calculated the percentage of EdU-positive cells as the number of EdU-positive cells/the number of DAPI-positive cells.
Flow Cytometry
Annexin V-fluorescein isothiocyanate (FITC)/propidium iodide (PI) apoptosis detection kit (Invitrogen) was utilized in this assay. After transfection for 48 h, 1 × 106 NSCLC cells were collected and suspended in 100 μL binding buffer. Then, 10 μL Annexin V-FITC and 10 μL PI were added to the binding buffer to incubate with NSCLC cells at 37 °C for 15 min in the dark. The apoptosis rate (the percentage of NSCLC cells with FITC+ and PI±) was analyzed by BD FACS Calibur flow cytometer (BD Biosciences, San Diego, CA, USA).
Transwell Assays
Transwell chambers (Costar, Corning, Switzerland) coated with or without Matrigel (BD Biosciences) were used for cell invasion or migration assay, respectively. A total of 100 μL cell suspension (serum-free medium) was added to the upper chamber, while the bottom chambers were filled with 500 μL complete culture medium. After incubation for 24 h, the migrated and invaded cells were fixed with 4% paraformaldehyde (Sigma) and stained with hematoxylin (Solarbio). Finally, the numbers of migrated and invaded cells in five random fields were counted under a light microscope (Olympus).
Western Blot Assay
NSCLC cells were disrupted with radio-immunoprecipitation assay (RIPA) buffer (Solarbio), and the concentrations of protein samples were determined by the bicinchoninic acid (BCA) method. Equal amounts of protein samples (30 μg) were separated by 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a polyvinylidene fluoride (PVDF) membrane (Millipore, Billerica, MA, USA). The membrane was blocked with 5% skimmed milk for 1 h at 37 °C. Subsequently, the membrane was incubated with primary antibodies overnight at 4 °C, including anti-ki67 (1:3000, ab16667, Abcam, Cambridge, MA, USA), anti-cleaved-caspase 3 (1:8000, ab2302, Abcam), anti-MMP9 (1:500, ab137867, abcam), anti-FAM83A (1:3000, PA5-46441, Thermo Fisher Scientific, Rochester, NY, USA) and GAPDH (1:20,000, ab9485, Abcam). The next day, the membrane was incubated with the secondary antibody (1:3000; Abcam) for 2 h at room temperature, the immune-reactive protein bands were visualized using the enhanced chemiluminescence system kit (Beyotime). ImageJ software (Bio-Rad) was used to analyze the intensities of protein bands.
Dual-Luciferase Reporter Assay
The partial sequence of circMIIP or FAM83A 3′ untranslated region (3′UTR), including the wild-type or mutant type binding sites with miR-766-5p, was inserted into pmirGLO vector (Promega, Madison, WI, USA). The luciferase reporter was co-transfected into NCI-H23 and A549 cells with miR-766-5p mimics or miR-NC. After transfection for 48 h, the luciferase activity was examined by Dual-Luciferase reporter Assay Kit (Solarbio).
RNA Immunoprecipitation (RIP) Assay
A Magna RNA immunoprecipitation kit (Millipore) was used to perform RIP assay. NSCLC cells were disrupted with ice-cold RIP lysis buffer. Then, cell lysates were incubated with magnetic beads labeled with the antibody of Ago2 (ab186733, Abcam) or IgG (ab19047, Abcam). After that, beads were washed with phosphate buffer saline (PBS) twice, and RT-qPCR was conducted to determine RNA enrichment in precipitated complex.
Xenograft Tumor Model
A549 cell line stably expressing sh-NC or sh-circMIIP was established. BALB/c nude mice (5-week-old) were randomly divided into two groups (n = 6/group, Charles River Labs, Beijing, China). A549 cells (1 × 106 cells) stably expressing sh-NC or sh-circMIIP were subcutaneously injected into the right flank of mice. Tumor volume was measured every 3 d as (length × width2)/2. After inoculation for 21 d, mice were killed, and the xenograft tumors were excised and weighed. The animal experiment was authorized by the Animal Welfare Committee of Affiliated hospital of Guizhou medical university.
Statistical Analysis
Data were expressed as mean ± standard deviation. Student’s t test and one-way analysis of variance followed by Tukey’s test were used to compare the difference in two groups or multiple groups using GraphPad Prism 7 software (GraphPad, La Jolla, CA, USA). The linear correlation was assessed by Pearson’s correlation coefficient. P < 0.05 indicated the statistically significant differences.
Results
CircMIIP Knockdown Suppresses the Proliferation, Migration, and Invasion While Promotes the Apoptosis of NSCLC Cells
CircMIIP (hsa_circ_0009932) is derived from exon 4–10 of its host gene MIIP, with a length of 946 nt (Sup. Fig. 1A). The results of RT-qPCR assay showed that circMIIP expression was remarkably increased in NSCLC tissues and cell lines (NCI-H23 and A549) compared with adjacent healthy tissues and normal human lung epithelial cell line (BEAS-2B) (Sup. Fig. 1B, C).
To investigate the biological role of circMIIP in NSCLC progression, loss-of-function experiments were performed. The results of RT-qPCR revealed that si-circMIIP#3 had the highest knockdown efficiency among three circMIIP-specific siRNAs (Fig. 1A). Then, si-circMIIP#3 was selected for further experiments. Colony formation assay, CCK-8 assay, and EdU assay together suggested that circMIIP knockdown suppressed the proliferation of NSCLC cells. (Sup. Fig. 2A, B; Fig. 1B; Sup. Fig. 3A). Flow cytometry showed that circMIIP absence induced the apoptosis of NSCLC cells (Fig. 1C; Sup. Fig. 3B). Transwell chambers coated with or without Matrigel were used to analyze cell invasion or migration ability, respectively. Transwell assays showed that circMIIP knockdown inhibited the migration and invasion abilities of NSCLC cells (Fig. 1D, E; Sup. Fig. 3C, D). Previous studies reported that ki67 expression can be detected in the nuclei of cells at G1, S, G2 phase and mitosis, but not in the nuclei of quiescent cells at G0 phase [10, 11]. Hence, ki67 level suggests the status of cell proliferation. Interestingly, ki67 is reported to be highly expressed in cancer cells and has been identified as a prognostic indicator of multiple cancers [12, 13]. MMP9 is one of the most widely investigated matrix metalloproteinases (MMPs), and it plays a vital role in ECM remodeling and membrane protein cleavage [14]. Increasing articles have demonstrated the MMP9 is closely associated with the pathology of multiple malignancies, including but not limited to metastasis and angiogenesis [15,16,17]. Currently, MMP9 has been widely used as a cell motility-related indicator in multiple cancers [18,19,20]. We measured the protein levels of proliferation-associated indicator ki67, apoptosis-associated indicator cleaved-caspase3, and motility-associated indicator MMP9 in transfected NSCLC cells. The protein levels of ki67 and MMP9 were decreased, while cleaved-caspase-3 level was increased in circMIIP-silenced NSCLC cells (Fig. 1F), further demonstrating that circMIIP silencing suppressed the proliferation and motility and induced the apoptosis of NSCLC cells. Taken together, circMIIP knockdown suppressed NSCLC progression in vitro.
CircMIIP knockdown suppresses NSCLC cell growth, migration, and invasion in vitro. A CircMIIP knockdown efficiency was confirmed in NCI-H23 and A549 cells introduced with si-NC, si-circMIIP#1, si-circMIIP#2, or si-circMIIP#3. B–F NCI-H23 and A549 cells were introduced with si-NC or si-circMIIP#3. B Cell proliferation ability was analyzed by EdU assay. C Flow cytometry was conducted to analyze cell apoptosis rate. D and E Transwell assays were conducted to analyze the migration and invasion abilities of NSCLC cells. F Western blot assay was performed to analyze the protein levels of ki67, clesved-caspase3, and MMP9 in transfected NSCLC cells. *P < 0.05
CircMIIP Acts as a Molecular Sponge for miR-766-5p
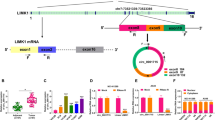
Bioinformatics databases starbase and circbank were used to predict the potential miRNA targets of circMIIP, and there were 4 miRNAs (miR-138-5p, miR-6783-3p, miR-766-5p, and miR-1343-3p) that were predicted to be targets of circMIIP by both databases (Fig. 2A). Only the expression of miR-766-5p was reduced by the overexpression of circMIIP in A549 cells (Fig. 2A). Thus, miR-766-5p was chosen for further experiments. The complementary sites between circMIIP and miR-766-5p are shown in Fig. 2B. miR-766-5p expression was notably reduced in NSCLC tissues compared with adjacent normal tissues (Fig. 2C), and the expression of miR-766-5p in NSCLC tissues was negatively correlated with circMIIP expression (Fig. 2D). Also, circMIIP expression was decreased in NSCLC cell lines relative to BEAS-2B cell line (Fig. 2E). RT-qPCR confirmed that the transfection efficiencies of miR-766-5p and anti-miR-766-5p were significant in NSCLC cells (Fig. 2F). Dual-luciferase reporter assay showed that miR-766-5p overexpression significantly reduced the luciferase activity of wild-type reporter (WT-circMIIP), but not that of mutant reporter (MUT-circMIIP) (Fig. 2G), suggesting that miR-766-5p was a direct target of circMIIP in NSCLC cells. RIP assay suggested that circMIIP and miR-766-5p were simultaneously enriched in Ago2 antibody group compared with IgG antibody group (Fig. 2H), further demonstrating the target relation between circMIIP and miR-766-5p. Taken together, circMIIP directly targeted miR-766-5p and negatively regulated miR-766-5p expression in NSCLC cells.
CircMIIP was a sponge of miR-766-5p in NSCLC cells. A Schematic illustration exhibiting overlapping of the target miRNAs of circMIIP predicted by starbase and circbank, and luciferase activity was detected in A549 cells co-transfected with wild-type circMIIP reporter and mimics of miR-766-5p miR-138-5p, miR-6783-5p or miR-1343-3p using the dual-luciferase reporter assay. B The predicted binding sites of circMIIP and miR-766-5p were exhibited. C CircMIIP expression was measured in NSCLC tissues and normal tissues using RT-qPCR. D Correlation between circMIIP and miR-766-5p in NSCLC samples. E CircMIIP expression was measured in NSCLC cell lines (NCI-H23 and A549) and human lung epithelial cell line (BEAS-2B) using RT-qPCR. F Transfection efficiencies detection of miR-766-5p mimics, inhibitor or their controls in NCI-H23 and A549 cells using RT-qPCR. G Luciferase activity was detected in NCI-H23 and A549 cells co-transfected with wild-type or mutated circMIIP reporter and miR-766-5p mimics or mimics control using the dual-luciferase reporter assay. H RT-qPCR analysis of circMIIP and miR-766-5p levels in NCI-H23 and A549 cells pulled down by anti-Ago antibody or IgG control antibody. *P < 0.05
CircMIIP Knockdown Suppresses NSCLC Progression by Sponging miR-766-5p
To explore whether circMIIP functioned by targeting miR-766-5p in NSCLC cells, rescue experiments were conducted by transfecting NSCLC cells with si-circMIIP#3 alone or together with miR-766-5p inhibitor. RT-qPCR analysis suggested that the introduction of miR-766-5p inhibitor attenuated circMIIP knockdown-induced upregulation of miR-766-5p in NSCLC cells (Fig. 3A). Rescue experiments showed that miR-766-5p inhibitor abolished circMIIP knockdown-mediated suppressive effects on the proliferation (Fig. 3B; Sup. Fig. 4A, B), migration (Fig. 3D) and invasion (Fig. 3E) and promoting effect on the apoptosis (Fig. 3C) of NSCLC cells. Furthermore, miR-766-5p knockdown overturned the effects of circMIIP knockdown on the protein levels of ki67, MMP9 and cleaved-caspase-3 in NSCLC cells (Fig. 3F). Altogether, circMIIP absence suppressed the malignant behaviors of NSCLC cells largely by upregulating miR-766-5p.
CircMIIP knockdown suppresses NSCLC cell growth, migration, and invasion via miR-766-5p. A–F NCI-H23 and A549 cells were co-transfected with si-NC, si-circMIIP#3, si-circMIIP#3 + anti-NC or si-circMIIP#3 + anti-miR-766-5p. A RT-qPCR assay was conducted to analyze miR-766-5p expression in transfected NSCLC cells. B Cell proliferation capacity was analyzed by EdU assay. C Flow cytometry was carried out to assess the apoptosis of transfected NSCLC cells. D and E Transwell assays were conducted to analyze the migration and invasion abilities of NSCLC cells. F Western blot assay was implemented to measure the protein levels of ki67, clesved-caspase-3 and MMP9 in NSCLC cells. *P < 0.05
CircMIIP Positively Regulates FAM83A Expression by Sponging miR-766-5p in NSCLC Cells
Four bioinformatics databases (miRDB, starbase, targetscan and GEPIA) were used to predict the potential messenger RNA (mRNA) targets of miR-766-5p, and only FAM83A was predicted to be the target of miR-766-5p by all four databases (Fig. 4A). The complementary sites between the 3’UTR of FAM83A and miR-766-5p were shown in Fig. 4B. The results of dual-luciferase reporter assay revealed that miR-766-5p overexpression markedly decreased the luciferase activity of wild-type reporter (WT-FAM83A 3’UTR) rather than mutant reporter (MUT-FAM83A 3’UTR), confirming the binding relationship between FAM83A and miR-766-5p via the predicted sites (Fig. 4C). FAM83A mRNA and protein expression was conspicuously increased in NSCLC tissues relative to adjacent normal tissues, and the mRNA expression of FAM83A in NSCLC tissues was negatively correlated with miR-766-5p expression (Fig. 4D, F). Besides, immunohistochemistry (IHC) assay showed that the staining intensity of FAM83A was significantly elevated in NSCLC tissues compared with adjacent normal tissues (Fig. 4G), further demonstrating the upregulation of FAM83A in NSCLC tissues. FAM83A protein expression was also upregulated in NSCLC cell lines relative to BEAS-2B cell line (Fig. 4H). Moreover, miR-766-5p overexpression decreased FAM83A expression in mRNA and protein levels, while miR-766-5p knockdown increased FAM83A mRNA and protein expression in NCI-H23 and A549 cells (Fig. 4I). CircMIIP overexpression elevated the protein expression of FAM83A in NSCLC cells, and this effect was reversed by the addition of miR-766-5p mimics (Fig. 4J). Overall, these results demonstrated that circMIIP positively regulated FAM83A expression by sponging miR-766-5p in NSCLC cells.
FAM83A is target of miR-766-5p, and circMIIP positively regulates FAM83A via miR-766-5p. A Schematic illustration exhibiting overlapping of the target genes of miR-766-5p predicted by miRDB, starbase, targetscan and GEPIA. B The predicted binding sites of FAM83A and miR-766-5p were exhibited. C Luciferase activity was detected in NCI-H23 and A549 cells co-transfected with wild-type or mutated FAM83A reporter and miR-766-5p mimics or mimics control using the dual-luciferase reporter assay. D FAM83A mRNA expression was measured in NSCLC tissues and normal tissues using RT-qPCR. E Correlation between FAM83A and miR-766-5p in NSCLC samples. F FAM83A protein level was measured in NSCLC tissues and normal tissues using western blot. G IHC staining of FAM83A level in NSCLC tissues and normal tissues. H Western blot analysis of FAM83A levels in NSCLC cell lines (NCI-H23 and A549) and BEAS-2B cells. I Western blot analysis of FAM83A levels in NCI-H23 and A549 cells transfected with miR-766-5p mimics, inhibitor or their controls. J Western blot analysis of FAM83A levels in NCI-H23 and A549 cells transfected with vector, circMIIP, circMIIP + miR-NC or circMIIP + miR-766-5p. *P < 0.05
miR-766-5p Overexpression Suppresses the Malignant Behaviors of NSCLC Cells by Targeting FAM83A
To investigate whether miR-766-5p regulated the biological behaviors of NSCLC cells by targeting FAM83A, compensation experiments were conducted. Western blot assay confirmed that the overexpression efficiency of FAM83A plasmid was significant (Fig. 5A). miR-766-5p overexpression reduced the protein level of FAM83A in NSCLC cells, which was largely rescued by the addition of FAM83A plasmid (Fig. 5B). Functionally, we found that miR-766-5p overexpression suppressed cell proliferation (Fig. 5C; Sup. Fig. 5A, B), induced cell apoptosis (Fig. 5D), as well as inhibited cell migration and invasion (Fig. 5E, F) in NSCLC cells, whereas these effects were largely counteracted by FAM83A overexpression. In addition, FAM83A overexpression largely overturned the effects of miR-766-5p accumulation on the protein levels of ki67, MMP9 and cleaved-caspase-3 in NSCLC cells (Fig. 5G, H). Overall, these results demonstrated that miR-766-5p overexpression restrained the malignant behaviors of NSCLC cells largely by downregulating FAM83A.
miR-766-5p suppresses NSCLC cell growth, migration, and invasion via FAM83A. A Western blot analysis of FAM83A level in NCI-H23 and A549 cells transfected with pcNDA3.1 or FAM83A. B–H NCI-H23 and A549 cells were transfected with miR-NC, miR-766-5p, miR-766-5p + pcNDA3.1, or miR-766-5p + FAM83A. B Western blot analysis of FAM83A levels in cells. C Cell proliferation was evaluated by EdU assay. D Flow cytometry was utilized to analyze cell apoptosis rate. E and F Transwell assays were performed to analyze cell motility in transfected NSCLC cells. G and H The protein levels of ki67, clesved-caspase-3 and MMP9 were detected by western blot assay. *P < 0.05
CircMIIP Knockdown Suppresses Xenograft Tumor Growth In Vivo
Considering the oncogenic role of circMIIP in NSCLC cells in vitro, we further explored whether circMIIP played a similar role on tumor growth in vivo. As shown in Fig. 6A, B, circMIIP silencing inhibited the growth of xenograft tumors. RT-qPCR and western blot analysis revealed that the levels of circMIIP and FAM83A mRNA and protein were decreased while miR-766-5p expression was elevated in circMIIP-silenced tumor tissues (Fig. 6C, D). IHC analysis revealed that the staining intensities of ki67 and MMP9 were both reduced in circMIIP-silenced tumor tissues compared with sh-NC group (Fig. 6E). These results demonstrated that circMIIP silencing inhibited xenograft tumor growth at least partly by targeting miR-766-5p/FAM83A axis in vivo.
CircMIIP knockdown suppresses NSCLC tumor growth and migration in vivo. A Tumor volume was measured every 3 days after injection. B The xenograft tumors were excised and weighed after mice were sacrificed. C, D The expression of circMIIP, miR-766-5p and FAM83A was examined in xenograft tissues by RT-qPCR and western blot, respectively. E IHC staining of ki67 and MMP9 in xenograft tissues. *P < 0.05
We also analyzed the roles of circMIIP and miR-766-5p on normal human lung epithelial cell line BEAS-2B. We found that circMIIP absence or miR-766-5p overexpression had almost no effect on the proliferation of BEAS-2B cells (Sup. Figs. 6A, B and 7A, B). Dual-luciferase reporter assay and RIP assay were performed to validate the interaction between miR-766-5p and circMIIP or FAM83A in BEAS-2B cells. The results revealed that circMIIP/miR-766-5p/FAM83A axis can also be established in BEAS-2B cells (Sup. Fig. 6C, F).
Discussion
CircRNAs are characterized by closed circular structure, which endows them high stability [21, 22]. Accumulating evidence demonstrated that circRNAs are widely dysregulated in multiple malignancies, and they play important roles in the patho-mechanism of cancers [23, 24]. In the present study, we found that circMIIP was strikingly upregulated in NSCLC tissues and cell lines compared with adjacent normal tissues and BEAS-2B cell line, implying that circMIIP expression might be associated with NSCLC pathology. The abnormal high expression pattern of circMIIP in HCC was consistent with the previous study [9]. Loss-of-function experiments suggested that circMIIP knockdown restrained the proliferation, migration, and invasion and induced the apoptosis of NSCLC cells in vitro. Moreover, xenograft tumor model showed that circMIIP knockdown hampered the growth of xenograft tumors in vivo. Taken together, these results suggested that circMIIP played an oncogenic role in NSCLC.
It is well established that circRNAs can act as molecular sponges for miRNAs to regulate target gene expression, thereby influencing tumor initiation and progression [25, 26]. Increasing articles have reported that miRNAs play vital regulatory roles in NSCLC carcinogenesis [27, 28]. We found that miR-766-5p was a direct target of circMIIP in NSCLC cells. Previous studies reported that miR-766-5p is a functional miRNA and is implicated in the pathology of various cancers. Furthermore, the function of miR-766-5p may be completely opposite in different cancers, which may due to different tumor microenvironments. For example, miR-766-5p is identified as an onco-miR in colorectal cancer [29], cervical cancer [30], and glioma [31], and it is reported to be a tumor suppressor in lung cancer [32, 33]. We found that miR-766-5p was downregulated in NSCLC tissues and cell lines. Consistent with former articles [32, 33], miR-766-5p overexpression restrained the malignant phenotypes of NSCLC cells. Moreover, we observed that miR-766-5p knockdown largely reversed the anti-cancer effects induced by circMIIP knockdown in NSCLC cells, suggesting that circMIIP knockdown restrained NSCLC progression largely by upregulating miR-766-5p in vitro.
FAM83A, also known as tumor-specific gene BJ-TSA-9, is a member of the eight-member FAM83 family [34]. Previous studies reported that FAM83A is highly expressed in bladder cancer, breast cancer, and pancreatic cancer [35,36,37]. FAM83A expression is also reported to be upregulated in lung cancer, and high expression of FAM83A is associated with poor outcome of lung cancer patients [38, 39]. FAM83A is reported to facilitate the proliferation and invasion of lung cancer cells by activating Wnt signaling and blocking Hippo signaling [40]. Hu et al. showed that FAM83A contributes to NSCLC progression by promoting cell proliferation and motility [41]. In this study, FAM83A was identified as a target of miR-766-5p, and we found that circMIIP served as a molecular sponge for miR-766-5p to elevate FAM83A expression in NSCLC cells. FAM83A overexpression largely abolished the anti-cancer effects induced by miR-766-5p overexpression in NSCLC cells, indicating that miR-766-5p overexpression restrained NSCLC progression largely by downregulating FAM83A in vitro.
In conclusion, circMIIP promoted cell proliferation, migration, and invasion and inhibited cell apoptosis in NSCLC cells through mediating miR-766-5p/FAM83A axis (Sup. Fig. 8), which provided novel potential therapeutic targets for NSCLC.
Data Availability
The analyzed data sets generated during the present study are available from the corresponding author on reasonable request.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(6):394–424. https://doi.org/10.3322/caac.21492
Relli V, Trerotola M, Guerra E, Alberti S (2019) Abandoning the notion of non-small cell lung cancer. Trends Mol Med 25(7):585–594
Wang J, Zhou T, Wang T, Wang B (2018) Suppression of lncRNA-ATB prevents amyloid-beta-induced neurotoxicity in PC12 cells via regulating miR-200/ZNF217 axis. Biomed Pharmacother 108:707–715. https://doi.org/10.1016/j.biopha.2018.08.155
Rong G, Wang H, Bowlus CL, Wang C, Lu Y, Zeng Z, Qu J, Lou M, Chen Y, An L, Yang Y, Gershwin ME (2015) Incidence and risk factors for hepatocellular carcinoma in primary biliary cirrhosis. Clin Rev Allergy Immunol 48(2–3):132–141. https://doi.org/10.1007/s12016-015-8483-x
Hsiao KY, Lin YC, Gupta SK, Chang N, Yen L, Sun HS, Tsai SJ (2017) Noncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res 77(9):2339–2350. https://doi.org/10.1158/0008-5472.can-16-1883
Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T, Mesteri I, Grunt TW, Zeillinger R, Pils D (2015) Correlation of circular RNA abundance with proliferation–exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep 5:8057. https://doi.org/10.1038/srep08057
Huang XY, Huang ZL, Xu YH, Zheng Q, Chen Z, Song W, Zhou J, Tang ZY, Huang XY (2017) Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-100338/miR-141-3p pathway in hepatitis B-related hepatocellular carcinoma. Sci Rep 7(1):5428. https://doi.org/10.1038/s41598-017-05432-8
Wang T, Wang X, Du Q, Wu N, Liu X, Chen Y, Wang X (2019) The circRNA circP4HB promotes NSCLC aggressiveness and metastasis by sponging miR-133a-5p. Biochem Biophys Res Commun 513(4):904–911. https://doi.org/10.1016/j.bbrc.2019.04.108
Jiang MM, Mai ZT, Wan SZ, Chi YM, Zhang X, Sun BH, Di QG (2018) Microarray profiles reveal that circular RNA hsa_circ_0007385 functions as an oncogene in non-small cell lung cancer tumorigenesis. J Cancer Res Clin Oncol 144(4):667–674. https://doi.org/10.1007/s00432-017-2576-2
du Manoir S, Guillaud P, Camus E, Seigneurin D, Brugal G (1991) Ki-67 labeling in postmitotic cells defines different Ki-67 pathways within the 2c compartment. Cytometry 12(5):455–463. https://doi.org/10.1002/cyto.990120511
Gerdes J, Lemke H, Baisch H, Wacker HH, Schwab U, Stein H (1984) Cell cycle analysis of a cell proliferation-associated human nuclear antigen defined by the monoclonal antibody Ki-67. J Immunol 133(4):1710–1715
Rioux-Leclercq N, Turlin B, Bansard J, Patard J, Manunta A, Moulinoux JP, Guillé F, Ramée MP, Lobel B (2000) Value of immunohistochemical Ki-67 and p53 determinations as predictive factors of outcome in renal cell carcinoma. Urology 55(4):501–505. https://doi.org/10.1016/s0090-4295(99)00550-6
Visapää H, Bui M, Huang Y, Seligson D, Tsai H, Pantuck A, Figlin R, Rao JY, Belldegrun A, Horvath S, Palotie A (2003) Correlation of Ki-67 and gelsolin expression to clinical outcome in renal clear cell carcinoma. Urology 61(4):845–850. https://doi.org/10.1016/s0090-4295(02)02404-4
Huang H (2018) Matrix metalloproteinase-9 (MMP-9) as a cancer biomarker and MMP-9 biosensors: recent advances. Sensors 18:3249. https://doi.org/10.3390/s18103249
Gialeli C, Theocharis AD, Karamanos NK (2011) Roles of matrix metalloproteinases in cancer progression and their pharmacological targeting. FEBS J 278(1):16–27. https://doi.org/10.1111/j.1742-4658.2010.07919.x
Farina AR, Mackay AR (2014) Gelatinase B/MMP-9 in tumour pathogenesis and progression. Cancers 6(1):240–296. https://doi.org/10.3390/cancers6010240
Xu D, McKee CM, Cao Y, Ding Y, Kessler BM, Muschel RJ (2010) Matrix metalloproteinase-9 regulates tumor cell invasion through cleavage of protease nexin-1. Can Res 70(17):6988–6998. https://doi.org/10.1158/0008-5472.can-10-0242
Dong DD, Zhou H, Li G (2015) ADAM15 targets MMP9 activity to promote lung cancer cell invasion. Oncol Rep 34(5):2451–2460. https://doi.org/10.3892/or.2015.4203
Chen Y, Jiang T, Mao A, Xu J (2014) Esophageal cancer stem cells express PLGF to increase cancer invasion through MMP9 activation. Tumour Biol 35(12):12749–12755. https://doi.org/10.1007/s13277-014-2601-x
Jian H, Zhao Y, Liu B, Lu S (2014) SEMA4b inhibits MMP9 to prevent metastasis of non-small cell lung cancer. Tumour Biol 35(11):11051–11056. https://doi.org/10.1007/s13277-014-2409-8
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO (2012) Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7(2):e30733. https://doi.org/10.1371/journal.pone.0030733
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S (2014) circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56(1):55–66. https://doi.org/10.1016/j.molcel.2014.08.019
Chen J, Li Y, Zheng Q, Bao C, He J, Chen B, Lyu D, Zheng B, Xu Y, Long Z, Zhou Y, Zhu H, Wang Y, He X, Shi Y, Huang S (2017) Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. Cancer Lett 388(2):08–219. https://doi.org/10.1016/j.canlet.2016.12.006
Feng Y, Wang Q, Shi C, Liu C, Zhang Z (2019) Does circular RNA exert significant effects in ovarian cancer? Crit Rev Eukaryot Gene Expr 29(2):161–170. https://doi.org/10.1615/CritRevEukaryotGeneExpr.2019025941
Chen X, Yu J, Tian H, Shan Z, Liu W, Pan Z, Ren J (2019) Circle RNA hsa_circRNA_100290 serves as a ceRNA for miR-378a to regulate oral squamous cell carcinoma cells growth via glucose transporter-1 (GLUT1) and glycolysis. J Cell Physiol 234(11):19130–19140. https://doi.org/10.1002/jcp.28692
Weng W, Wei Q, Toden S, Yoshida K, Nagasaka T, Fujiwara T, Cai S, Qin H, Ma Y, Goel A (2017) Circular RNA ciRS-7-A promising prognostic biomarker and a potential therapeutic target in colorectal cancer. Clin Cancer Res 23(14):3918–3928. https://doi.org/10.1158/1078-0432.ccr-16-2541
Bica-Pop C, Cojocneanu-Petric R, Magdo L, Raduly L, Gulei D, Berindan-Neagoe I (2018) Overview upon miR-21 in lung cancer: focus on NSCLC. Cell Mol Life Sci 75(19):3539–3551. https://doi.org/10.1007/s00018-018-2877-x
Chen Y, Min L, Ren C, Xu X, Yang J, Sun X, Wang T, Wang F, Sun C, Zhang X (2017) miRNA-148a serves as a prognostic factor and suppresses migration and invasion through Wnt1 in non-small cell lung cancer. PLoS ONE 12(2):e0171751. https://doi.org/10.1371/journal.pone.0171751
Jia B, Xia L, Cao F (2018) The role of miR-766-5p in cell migration and invasion in colorectal cancer. Exp Ther Med 15(3):2569–2574. https://doi.org/10.3892/etm.2018.5716
Cai Y, Zhang K, Cao L, Sun H, Wang H (2020) Inhibition of microrna-766-5p attenuates the development of cervical cancer through regulating SCAI. Technol Cancer Res Treat 19:1533033820980081. https://doi.org/10.1177/1533033820980081
Mao Y, Shen G, Su Z, Du J, Xu F, Yu Y (2020) RAD21 inhibited transcription of tumor suppressor MIR4697HG and led to glioma tumorigenesis. Biomed Pharmacother 123:109. https://doi.org/10.1016/j.biopha.2019.109759
Wang M, Liao Q, Zou P (2020) PRKCZ-AS1 promotes the tumorigenesis of lung adenocarcinoma via sponging miR-766-5p to modulate MAPK1. Cancer Biol Ther 21(4):364–371. https://doi.org/10.1080/15384047.2019.1702402
Bai Y, Zhang G, Cheng R, Yang R, Chu H (2019) CASC15 contributes to proliferation and invasion through regulating miR-766-5p/ KLK12 axis in lung cancer. Cell Cycle 18(18):2323–2331. https://doi.org/10.1080/15384101.2019.1646562
Li Y, Dong X, Yin Y, Su Y, Xu Q, Zhang Y, Pang X, Zhang Y, Chen W (2005) BJ-TSA-9, a novel human tumor-specific gene, has potential as a biomarker of lung cancer. Neoplasia 7(12):1073–1080. https://doi.org/10.1593/neo.05406
Cipriano R, Miskimen KL, Bryson BL, Foy CR, Bartel CA, Jackson MW (2014) Conserved oncogenic behavior of the FAM83 family regulates MAPK signaling in human cancer. Mol Cancer Res 12(8):1156–1165. https://doi.org/10.1158/1541-7786.mcr-13-0289
Bartel CA, Jackson MW (2017) HER2-positive breast cancer cells expressing elevated FAM83A are sensitive to FAM83A loss. PLoS ONE 12(5):e0176778. https://doi.org/10.1371/journal.pone.0176778
Chen S, Huang J, Liu Z, Liang Q, Zhang N, Jin Y (2017) FAM83A is amplified and promotes cancer stem cell-like traits and chemoresistance in pancreatic cancer. Oncogenesis 6(3):e300. https://doi.org/10.1038/oncsis.2017.3
Richtmann S, Wilkens D, Warth A, Lasitschka F, Winter H, Christopoulos P, Herth FJF, Muley T, Meister M, Schneider MA (2019) FAM83A and FAM83B as prognostic biomarkers and potential new therapeutic targets in NSCLC. Cancers. https://doi.org/10.3390/cancers11050652
Zhang J, Sun G, Mei X (2019) Elevated FAM83A expression predicts poorer clincal outcome in lung adenocarcinoma. Cancer Biomark 26(3):367–373. https://doi.org/10.3233/cbm-190520
Zheng YW, Li ZH, Lei L, Liu CC, Wang Z, Fei LR, Yang MQ, Huang WJ, Xu HT (2020) FAM83A promotes lung cancer progression by regulating the Wnt and hippo signaling pathways and indicates poor prognosis. Front Oncol 10:180. https://doi.org/10.3389/fonc.2020.00180
Hu H, Wang F, Wang M, Liu Y, Wu H, Chen X, Lin Q (2020) FAM83A is amplified and promotes tumorigenicity in non-small cell lung cancer via ERK and PI3K/Akt/mTOR pathways. Int J Med Sci 17(6):807–814. https://doi.org/10.7150/ijms.33992
Acknowledgements
Not applicable.
Funding
No funding was received.
Author information
Authors and Affiliations
Contributions
All authors made substantial contribution to conception and design, acquisition of the data, or analysis and interpretation of the data; take part in drafting the article or revising it critically for important intellectual content; gave final approval of the revision to be published; and agree to be accountable for all aspect of the work. Conceptualization, Methodology, Formal analysis and Data curation: XZ and KW; Validation and Investigation: TW and XZ; Writing—original draft preparation and Writing—review and editing: TW, XZ and KW; Approval of final manuscript: all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
The present study was approved by the ethical review committee of Affiliated Hospital of Guizhou Medical University. Written informed consent was obtained from all enrolled patients.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
408_2021_500_MOESM2_ESM.tif
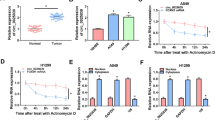
Supplementary file2 Sup. Fig 1. CircMIIP is highly expressed in NSCLC tissues and cells. (A) Schematic illustration showing the genomic location of circMIIP generated from its host gene MIIP. (B, C) CircMIIP expression was measured in NSCLC tissues and normal tissues, as well as in NSCLC cell lines (NCI-H23 and A549) and human lung epithelial cell line (BEAS-2B) using RT-qPCR. *P < 0.05 (TIF 3420 kb)
408_2021_500_MOESM3_ESM.tif
Supplementary file3 Sup. Fig 2. CircMIIP silencing restrains the proliferation of NSCLC cells. (A and B) NSCLC cells were transfected with si-NC or si-circMIIP#3. Cell proliferation ability was analyzed by colony formation assay and CCK-8 assay. *P < 0.05 (TIF 3017 kb)
408_2021_500_MOESM4_ESM.tif
Supplementary file4 Sup. Fig 3. The representative images of EdU assay, flow cytometry, and transwell assays in Figure 1. (A) The representative images of EdU assay in Figure 1B. (B) The representative images of flow cytometry in Figure 1C. (C and D) The representative images of transwell assays in Figure 1D and 1E (TIF 22548 kb)
408_2021_500_MOESM5_ESM.tif
Supplementary file5 Sup. Fig 4. CircMIIP absence restrains the proliferation of NSCLC cells largely by up-regulating miR-766-5p. (A and B) NSCLC cells were transfected with si-circMIIP#3 alone or together with anti-miR-766-5p. Colony formation assay and CCK-8 assay were conducted to analyze the proliferation ability of NSCLC cells. *P < 0.05 (TIF 761 kb)
408_2021_500_MOESM6_ESM.tif
Supplementary file6 Sup. Fig 5. miR-766-5p overexpression suppresses the proliferation of NSCLC cells largely by down-regulating FAM83A. (A and B) NSCLC cells were transfected with miR-766-5p alone or together with FAM83A. Cell proliferation ability was analyzed by colony formation assay and CCK-8 assay. *P < 0.05 (TIF 753 kb)
408_2021_500_MOESM7_ESM.tif
Supplementary file7 Sup. Fig 6. CircMIIP/miR-766-5p/FAM83A axis in established in BEAS-2B cells. (A) The knockdown efficiencies of three circMIIP-targeted siRNAs were analyzed in BEAS-2B cells by RT-qPCR. (B) BEAS-2B cells were transfected with si-NC or si-circMIIP#3, and CCK-8 assay was conducted to analyze the proliferation ability of transfected BEAS-2B cells. (C and D) Dual-luciferase reporter assay and RIP assay were conducted to verify the target relationship between circMIIP and miR-766-5p in BEAS-2B cells. (E and F) The interaction between miR-766-5p and FAM83A in BEAS-2B cells was confirmed by dual-luciferase reporter assay and RIP assay. *P < 0.05 (TIF 1432 kb)
408_2021_500_MOESM8_ESM.tif
Supplementary file8 Sup. Fig 7. miR-766-5p overexpression has almost no effect on the proliferation of BEAS-2B cells. (A and B) BEAS-2B cells were transfected with miR-NC or miR-766-5p. (A) RT-qPCR was conducted to analyze the overexpression efficiency of miR-766-5p mimics in BEAS-2B cells. (B) CCK-8 assay was performed to assess the proliferation of BEAS-2B cells. *P < 0.05 (TIF 460 kb)
408_2021_500_MOESM9_ESM.tif
Supplementary file9 Sup. Fig 8. A schematic diagram shows the role of circMIIP/miR-766-5p/FAM83A axis on the malignant behaviors of NSCLC cells (TIF 3065 kb)
Rights and permissions
About this article
Cite this article
Wang, T., Zhu, X. & Wang, K. CircMIIP Contributes to Non-Small Cell Lung Cancer Progression by Binding miR-766-5p to Upregulate FAM83A Expression. Lung 200, 107–117 (2022). https://doi.org/10.1007/s00408-021-00500-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00408-021-00500-3