Abstract
To identify plant growth promotion ability of phosphorus-solubilizing native bacteria, we have examined a collection of isolates representing the diversity of culturable phosphate-solubilizing bacteria from acid soils of the northeast of Argentina. Assays in growth medium supplemented with tricalcium phosphate revealed different phosphorus solubilization activity and temporal patterns of solubilization. Acidification of the broth medium coincided with phosphorus solubilization. The isolates were grouped according to their Rep fingerprinting profiles and phylogenetically classified by 16S rDNA and biochemical analyses. These isolates were assigned to the genera Enterobacter, Pantoea, Pseudomonas, Acinetobacter, Burkholderia, and Exiguobacterium. Four isolates showing high phosphorus solubilizing activity in in vitro assays were inoculated on common beans (Phaseolus vulgaris); some of them promoted plant growth and increased photosynthesis and the P and N content of leaves. The results indicated that the ability to in vitro solubilize P is not necessarily associated to the promotion of plant growth.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Phosphorus (P) is an essential mineral nutrient required for plant growth because it is a major component of important molecules (Bielski 1973). Although a substantial reserve of P is present in soil, a large proportion is unavailable to plants and a considerable part of the fertilizer phosphates applied to soil is immobilized after application (Rodríguez and Fraga 1999). In acid soils such as Ultisols, soluble P forms are mainly fixed by aluminum and iron-free oxides and hydroxides (Fearnside 1998), and this limits P availability and crops in tropical soils (Tiessen et al. 1996). A crop usually grown in acid soils is the common bean, the most important food legume especially in Latin America and Africa and a P-demanding plant. Many efforts have been made in order to obtain plants able to grow in P-deficient soils (Broughton et al. 2003).
Soil and rhizosphere bacteria, known as phosphate-solubilizing bacteria (PSB), solubilize insoluble inorganic P compounds mainly by producing low-molecular-weight organic acids such as gluconic and keto-gluconic acids (Rodríguez et al. 2006). The association between PSB and plant roots plays a key role in P nutrition in many agro-ecosystems, particularly in P-deficient soils (Goldstein 2007; Jorquera et al. 2008). Although several PSB occur in soil and in plant rhizospheres, the amount of P released by these bacteria is generally not sufficient to fulfill the requirements of growing plants (Rodríguez et al. 2006). The inoculation of plants with selected PSB to increase native population can mobilize P from poorly available sources and therefore improve plant nutrition (Richardson 2001; Guiñazu et al. 2010). The widespread use of PSB inoculants remains limited because of the inconsistent results in different environments, but the potentialities of these resources have not been completely explored in terms of bacterial species efficient in mobilizing P with crops of economic and social significance. Particularly, the P-solubilizing bacteria activity, determined by in vitro assays, should be correlated with the ability to promote plant growth under P-limited conditions, and the inoculated bacteria should be able to colonize roots.
The aim of this research was to establish a collection of PSB adapted to P-deficient acid soils and to compare their in vitro P solubilization activity with the promotion of plant growth. For this purpose, we characterized efficient in vitro P-solubilizing bacteria isolated from an acidic lateritic soil associated with yerba mate (Ilex paraguariensis St. Hil.) and evaluated their effects on the growth of Phaseolus vulgaris (common beans), a species highly dependent on P availability and sensitive to P deficiency (Hernández et al. 2007). We found that only some isolates promoted the growth of P. vulgaris, although all of them solubilized P in in vitro conditions. Our data revealed that the in vitro solubilization activity of PSB not necessarily correlates with the ability to promote plant growth.
Materials and methods
Site description and soil sampling
The study site was a yerba mate plot located in the northeast of Argentina (28°02′ S, 56°00′ W). The orchard was established in 2002 with a plant density of 4,000 plants/ha. The region has a mean annual rainfall of approximately 1,800 mm, distributed mainly during spring and autumn. The mean annual temperature is 22°C with scarce frosts. The soil is an Ultisol (Kandihumultes family) with high clay (kaolinite) in the B horizon and acid pH concomitantly with high aluminum content and low available P. The soil contains 65.4% clay, 14.3% silts, and 20.3% sands and 3.55% organic content, 1.8 mg g−1 N, 3.9 mg kg−1 P, 5.1 mg kg−1 Ca, 0.66 mg kg−1 Mg, 0,65 mg kg−1 K, and 1.08 meq/100 g Al, pH 4.1.
Samples associated with yerba mate used for bacteria isolation were root, rhizosphere, and root-free soil. Triplicate samples were collected from the rhizosphere and roots of randomly selected plants. Soil samples were also collected from an area free of root near the orchard. All samples were taken from a depth of 2–20 cm, kept in plastic bags in an icebox, and carried to the laboratory at the Universidad Nacional del Nordeste. The samples were stored at 4°C for 12 h until processing.
Isolation and screening of PSB
The first step for the bacterial isolation was specific for each type of sample. Rhizosphere bacteria were obtained from the roots previously washed under running tap water to remove the soil. The soil samples from the root-free zone were thoroughly mixed and sieved with a 100-mesh net. For the endophytic bacteria isolation, roots were surfaced-sterilized with ethanol 95% for 1 min, followed by 1% sodium hypochlorite for 20 min, washed extensively with sterile water, and crushed with a sterile mortar and pestle. After the different treatments, 5 g of roots, soil, and crushed roots was suspended in 40 mL of phosphate-buffered saline solution (10 mM K2PO4–KH2PO4, 0.14 M NaCl, pH 7.2) plus 20 g of glass beads and shaken for 30 min. After 4 h, debris and soil particles in solution were removed by centrifugation (2 min at 500×g) and the cells collected from the supernatant.
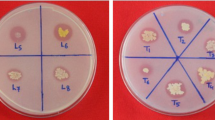
For bacterial isolation, each sample was serially diluted, plated on trypticase soy agar medium supplemented with 50 μg mL−1 cycloheximide, and incubated at 30°C for 2–10 days. Colonies of the predominant morphology were selected at random from each plate, and 900 isolates from each type were purified and screened for their phosphate-solubilizing ability. A total of 50 colonies per plate were incubated for 2 weeks at 30°C on National Botanical Research Institute’s phosphate growth medium (NBRIP) with 5 g L−1 of tricalcium phosphate (TCP) as the exclusive P source (Nautiyal 1999). Colonies were selected on the basis of the appearance of a clear halo. After this initial screening, four selected isolates per plate were inoculated in the same medium and incubated at 30°C for 2 weeks. The sizes of the halo and colony diameter were measured every 2 days. The solubilization index (SI) was calculated by subtracting the colony diameter from the total diameter of the halo. Strains displaying an SI equal to or higher than 1.4 in three replicates were selected for further analysis.
The efficient PSB strain Pseudomonas RHP3 was used as reference in this study (Mehta and Nautiyal 2001).
Rep-PCR analysis
The isolated PSB community was characterized by repetitive extragenic palindromic sequence PCR (Rep-PCR) using ERIC1R-ERIC2 primers (Versalovic et al. 1994). DNA was prepared from bacterial cells using a Chelex 100 resin (Bio-Rad) method as described by Alippi and Aguilar (1998). PCR amplifications were performed according to Versalovic et al. (1994). For the analysis of amplification products, 10 μL of each PCR reaction was run on a 2% (w/v) agarose gel in Tris-borate-EDTA electrophoresis buffer. A digital image of each gel was analyzed using the Gelcompare software version 4.0 (Applied Maths BVBA, Belgium). A similarity of 70% between isolates was defined to determine groups. The Shannon–Weaver index was determined for the estimation of diversity (H′) in populations from soil, root, and rhizosphere samples based on the number of isolates belonging to each Rep-PCR profile.
Physiological characterization of the PSB
One isolate of each different profile generated by Rep-PCR, as described in the preceding section, was selected for the quantitative P solubilization assay. Briefly, 150-mL flasks containing 25 mL NBRIP medium with 5 g L−1 TCP were inoculated with an aliquot of pure bacterial culture to yield an initial optical density at 600 nm of 0.1. Flasks were shaken at 28°C for 10 days, and bacterial growth (by optical density at 600 nm) and pH were periodically determined. Quantitative analysis of P solubilization was estimated in the culture supernatant using the molybdenum blue method (Murphy and Riley 1962). Non-inoculated NBRIP medium was included in the experiments as control.
Strains showing efficient TCP solubilization were selected for further physiological characterization. The ability to grow under stressed conditions such as acidity (pH 4.5, 5.0, and 5.5) and high temperature (37°C, 39°C, and 42°C) was analyzed in cells growing in NBRIP plates supplemented with KH2PO4 (10 g L−1) and CaCl2⋅2H2O (11 mg L−1). To evaluate the effect of high temperature in the solubilizing activity, strains were grown at 42°C in NBRIP plates with 5 g L−1 TCP. Solubilizing activity of other forms of insoluble P was determined in NBRIP broth medium using FePO4 (1 g L−1) or AlPO4 (2 g L−1) instead of Ca3(PO4)2. To evaluate the solubilizing activity in the presence of soluble P, bacterial cells were cultured in NBRIP plates supplemented with Ca3(PO4)2 (5 g l−1) and KH2PO4 (2.7 g l−1).
Identification of phosphate-solubilizing strains
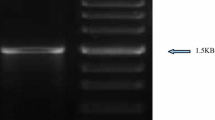
Phylogenetic assignment of isolates was carried out by sequence analysis of the 16S rRNA gene and by analysis of phenotypic traits using the API system (BioMerieux). For the analysis of the 16S rRNA gene, a fragment of about 1.5 Kb—which spans almost the full length of the gene—was PCR-amplified using the universal primers rD1 and fD1 (Weisburg et al. 1991). The reactions were performed using the following cycling conditions: one step of 95°C for 10 min, then 30 cycles of 95°C for 30 s, 58°C for 30 s, and 72°C for 1 min, followed by one final step of 72°C for 10 min. PCR products were cloned using pGEM-T easy (Promega) and sequenced by Macrogen Inc., Seoul, Korea. Nucleotide sequences were compared with the database, and pairwise sequence similarities were determined with the EzTaxon server (Chun et al. 2007).
Gram-negative isolates were characterized using the biochemical test API 20E for Enterobacteriaceae, whereas the 20 NE was used for non-Enterobacteriaceae according to the manufacturer’s instructions (BioMérieux). The APIWEB software was used for identification, and assignment was considered acceptable when it scored at least 80% similarity (BioMérieux).
Assignment at the level of genus or species was carried out according to the degree of similarity in 16S rDNA sequences. Identification was considered at the genus level when the sequence similarity to a reference sequence was between 95% and 99%. The isolate was classified at the species level when similarity was equal to or higher than 99%. However, if the second most similar species showed <0.5% additional sequence divergence, the isolate was categorized as a species with low delimitation regarding the closest species.
According to Bosshard et al. (2006), if the identification by 16S rDNA and API analyses diverged from each other, the sequence data were examined in more detail. When the identity between the sequence of the isolate and that of the species assigned by the API system resulted lower than 97%, it was concluded that the phenotypic identification was incorrect and the isolate was identified according to 16S rDNA analysis, whereas when the sequence identity was equal to or higher than 97%, the isolate was categorized as unresolved.
Quantification of indole acetic acid
Chromatographic determination of indole acetic acid (IAA) produced by bacterial cultures was performed according to Perrig et al. (2007). Bacterial cultures (20 mL) in exponential growth phase were centrifuged at 8,000×g for 10 min at 4°C and acidified at pH 2.5 with 1% acetic acid. Subsequently, 100 ng of 2H5-IAA was added to the supernatants, and 10 μL of the mixtures was analyzed by an Alliance 2695 chromatograph (Water Inc., USA). Each aliquot was analyzed by an MS-MS Quatro Ultima™ (Micromass, UK) coupled to the HPLC. The amount of free IAA was calculated by comparing the molecular mass 174/179 and the transition mass 130/135 for endogenous/deuterons, respectively. Free IAA was quantified by integrating the area under the peak with the authentic indole compound (Sigma Chem. Co., USA) as standard.
Inoculation of common beans with phosphate-solubilizing strains
The inoculation effects of PSB on plant growth and P uptake was assayed on common beans (P. vulgaris) cv. Nag12. Seeds were surface-sterilized with ethanol 95% for 1 min, followed by 1% sodium hypochlorite for 30 min, washed extensively with sterile water, and germinated in darkness on water agar plates at 28°C. Seedlings were grown in 500-mL plastic pots with each pot filled with sterile perlite irrigated with Hoagland solution (Arnon and Hoagland 1940). The experiment was performed under controlled environmental conditions (27 ± 1/22 ± 2°C day/night temperature, 14-h photoperiod at 400 μmol m−2 s−1 photosynthetic photon flux density (PPFD) from fluorescent lamps, and 55–60% relative humidity).
Seven-day-old plantlets were inoculated with bacterial cells resuspended in 10 mM MgSO4 at a density of about 108 colony forming units (cfu) per milliliter. To assess P mobilization in plant, soluble (KH2PO4, 1 mM) and insoluble (TCP 160 mg L−1 corresponding to PO 3−4 1 mM) P forms were supplied as a micronutrient solution (Hoagland’s) every 3 days. Treatments were designed in a randomized complete block as follows: (1) non-inoculated without P fertilization; (2) non-inoculated with TCP; (3) non-inoculated with KH2PO4; (4) inoculated and supplemented with TCP; and (5) inoculated and supplemented with KH2PO4. In treatments with insoluble P, the irrigation solution was modified by reducing the 1 mM KH2PO4 concentration to 5 μM. In order to maintain an equivalent potassium concentration, 2 mM of KCl was added.
At the beginning of the flowering phase (35-day-old plants), the plants were harvested and the root colonization by the inoculated strains was assayed. The roots were removed aseptically and fresh weights determined. Root segments were crushed with a sterile mortar and pestle and resuspended in phosphate buffer saline solution. Serial dilutions of these suspensions were plated on NBRIP medium. The colonies were analyzed by Rep-PCR and the fingerprint pattern was compared with those of the pure culture inoculated strains.
Photosynthesis and nutrient contents
Photosynthetic rates and stomatal conductance were measured on intact mature leaves (no. 3 from the apex) at the beginning of the flowering phase using an open-flow gas exchange system Li-Cor LI-6400 (Li-Cor Inc., Nebraska). Measurements were taken at CO2 concentrations ranging from 50 to 800 μmol mol−1 and 1,500 μmol m−2 s−1 PPFD. Leaf temperature was kept at 30°C. Subsequently, the plants were harvested and shoot-and-root dry weights recorded for each individual plant. Leaf area was determined using an area meter analyzer Li-Cor Li-3000. The Kjeldahl (Bremmer and Mulvaney 1982) and vanadomolybdate yellow colorimetric (Hesse 1971) methods were respectively applied to determine N and P contents in leaves.
Nucleotide sequence accession numbers
The sequences obtained in this study were deposited in the GenBank nucleotide sequence database under accession numbers GQ478255 to GQ478276.
Results
Isolation of PSB from acid soils
A core collection of 518 PSB was established after screening 2,700 colonies for in vitro P solubilization activity. The collection comprised isolates from soil (15%), rhizosphere (46%), and roots of yerba mate plants (38%). The culturable PSB community showed diverse P solubilization activity; therefore, isolates were grouped according to their temporal pattern of activity into two groups. Isolates that showed activity early in culture conformed a group, whereas the other group of isolates showed significant activity at the late stages of growth. Each group represented half of the isolates of the collection, which comprised SI levels that ranged from 1 (inconspicuous halo) to higher than that produced by the efficient strain RHP3 used for comparison.
Rep-PCR fingerprinting assays of the isolates with efficient solubilization activity revealed a high diversity (H′ = 3.7) with 58 distinct Rep-PCR profiles (haplotypes) among the 252 isolates examined (Fig. 1). The rhizosphere, root, and soil samples did not share Rep-PCR profiles, except for profiles H and S (R4M-H and B4M-S in Table 1), which were observed in isolates from rhizosphere and soil samples. The highest diversity was found in the rhizosphere samples (H′ = 3.1) with 32 haplotypes out of 118 isolates examined, whereas soil (H′ = 2.05) and endophytic (H′ = 2.4) populations showed a lower diversity, with 11 and 17 haplotypes out of 52 and 82 isolates analyzed, respectively.
Physiological characterization of the PSB
A subsample of isolates representing each Rep-PCR fingerprint pattern was quantitatively assessed for P solubilization activity. Twenty-one strains showed P solubilization activity above 300 μg P mL−1, similar to that of the reference strain RHP3 (Table 1). Twenty-six strains were less active, with maximal activity between 100 and 300 μg P mL−1. Even lower activity was displayed by five of the strains (<100 μg P mL−), which were considered inefficient solubilizing bacteria (e.g., strain R4M16 in Table 1).
All phosphate-solubilizing bacteria assayed decreased the pH of the medium, and the relative time course coincided with the increase in the P solubilization activity. The lowest pH values were scored during the growth phase in which maximal solubilization activity was detected (Table 1). Then, alkalinization of the medium occurred together with the decrease of soluble P in the supernatant.
According to the P solubilization activity during the bacterial growth, PSB can be classified as: (1) early solubilizers displaying significant solubilizing activity during the first hours of culture and with maximal activity within the first 72 h of growth. In this case, the culture reached the maximal cell density (about 109 cfu mL−1) 24 h after culture initiation (Fig. 2a). (2) Late solubilizers showing low activity during the first 72 h and reaching maximal activity between the fourth and fifth days (Fig. 2b). These strains showed slow growth in NBRIP medium, reaching a maximum cell density after 72 h of culture. It was noteworthy that the population of early solubilizers fell mainly in the group of isolates from the rhizosphere.
Phosphate solubilization activity of early (a) and late (b) solubilizing bacterial isolates growing in NBRIP broth at 30°C after 24, 72, 120, and 168 h of culture. Strains R4M16 and RHP3 (dot lines) were included as control of inefficient and efficient solubilizers, respectively. Each value is a mean of three independent replicates
Phosphate-solubilizing activity can be repressed by the presence of soluble P in the medium (Gyaneshwar et al. 1999; Mikanová and Nováková 2002). To assess the effect of readily available P on the solubilization activity by the PSB isolated, the strains were individually grown in NBRIP medium supplemented with KH2PO4 (20 mM). We found that the solubilizing activity of most strains (13 out of 21) decreased in the presence of KH2PO4, with some strains being highly affected (SI = 1; e.g., E4M-B and B4M-N strains in Table 1). Other strains were intermediately affected and presented a solubilizing activity significantly lower (SI > 1) than that detected in the NBRIP control (R4M-D strain in Table 1).
The strains were tested for their ability to solubilize other salts such as AlPO4 and FePO4 in NBRIP broth medium. All the strains assayed decreased the medium pH but displayed a low capacity to solubilize these phosphate salts, with maximal activity of 20 μg P mL−1 (E4M-B strain) and 12 μg P mL−1 (B4M-T strain) for AlPO4 and FePO4, respectively.
Efficient TCP-solubilizing bacteria were further screened for their ability to grow under stressed conditions such as low pH and high temperature. All strains could grow on acid conditions (pH 4.5), and more than a half (13 strains) were able to grow at 42°C (Table 1). However, the SI values were low, showing that high temperatures negatively affect TCP solubilization activity.
Identification of phosphate-solubilizing strains
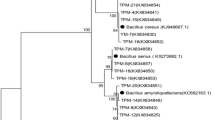
Twenty-two selected isolates were further characterized by applying molecular and physiological analysis. All but three of the isolates were identified by both API and 16S rDNA analyses, and the results were in accordance with each other at the level of genus. As shown in Table 2, 63% of the isolates were assigned to the γ-proteobacteria subdivision, the majority assigned to Enterobacteriales (Enterobacter and Pantoea) and the rest to Pseudomonadales (Pseudomonas and Acinetobacter) and Xanthomonadales (Stenotrophomonas). The β-proteobacteria, which belonged to the order Burkholderiales (Burkholderia and Variovorax genus), was the second most abundant group (33%). Only one of the isolates was Gram-positive, namely, strain R4M-H, which was similar to the genus Exiguobacterium (order Baciliales).
It is noteworthy that isolates from all the orders described above were detected in soil and rhizosphere, except for the Enterobacteriales which were detected only in root samples.
The 16S rDNA sequences (1.5 Kb) of 12 isolates showed similarities higher than 99% to a reference sequence. However, the phylogenetic assignment remained ambiguous in the case of the R4M-V, R4M-W, and R4M-P strains because difference between species was low (i.e., comparison of sequence revealed <0.5% additional difference with the second most similar species). Thus, nine isolates were unambiguously assigned to a single species by molecular analysis of 16S rDNA, and four isolates showed identical results by applying API analysis (Table 2). In the case of the isolates identified by API analysis as Burkholderia cepacia, results diverged between API and 16S rDNA analyses. The sequences of the R4M-I, R4M-O, and R4M-G isolates were very similar to each other (99–99.9%) as well as to that of Burkholderia caledonica (99.2–100%). Since the 16S rDNA sequence of members of this group and that of the B. cepacia complex showed <96% similarity, we concluded that our isolates should be assigned to B. caledonica rather than to B. cepacia. In contrast, R4M-F isolate, which is closely related to Burkholderia ambifaria according to the 16S rDNA sequence (99.2% similarity), could not be assigned to the species level because of its high similarity (98.2%) with B. cepacia.
Most of the enterobacterial isolates were identified at the genus level since sequence similarity to the reference species was <99%. R4M-C, R4M-Q, B4M-N, B4M-S, E4M-U, and E4M-P isolates showed the highest sequence similarity among them (99.7–99.8%) and with the type strains that corresponded to Enterobacter hormaechei, Enterobacter asburiae, Enterobacter cancerogenus, and Enterobacter ludwigii (98.3–99.36%) species, which are included in the Enterobacter cloacae complex.
Plant assays
Effect of inoculation on plant growth and nutrient contents
The effect of inoculation on plant growth was assessed on P. vulgaris under environmentally controlled conditions. The following PSB, showing a good performance in TCP solubilization, were selected for the plant assay: Enterobacter aerogenes R4M-A and Burkholderia spp. R4M-F, both isolated from the rhizosphere; Acinetobacter baumannii B4M-L, isolated from the soil; and E. aerogenes E4M-B isolated from root. Thirty days after inoculation, R4M-A and R4M-F strains effectively colonized the root system of P. vulgaris with 2.7 × 105 and 1.3 × 105 cfu g−1 roots, respectively; both B4M-L and E4M-B strains showed lower root colonization (1.2 × 102 and 1.7 × 102 cfu g−1 roots, respectively). Results from plant inoculation experiments with E4M-B, R4M-L, and R4MF strains were similar to each other; therefore, only data on the colonizing strain R4M-F are compared with strain R4M-A.
Non-inoculated plants grown under insoluble P conditions or without P fertilization showed lower shoot dry weight and leaf area and higher root/shoot ratio than plants grown with soluble P (Fig. 3a–d). Inoculation with E. aerogenes R4M-A strain increased plant dry biomass and leaf area regardless of the P form supplied in the medium. The root/shoot ratio of R4M-A-inoculated plants grown with TCP and that of plants grown with soluble P did not significantly differ from each other, suggesting that plants showed no P deficiency symptoms (Hernández et al. 2007). Under insoluble P conditions, inoculation with E. aerogenes R4M-A strain significantly increased plant leaf area (P < 0.001), shoot dry mass (P < 0.001), and root dry mass (P < 0.005), whereas no effect was detected in plants inoculated with Burkholderia spp. R4M-F strain (Fig. 3a–c). In addition, under this condition, shoot dry mass and leaf area of R4M-A-inoculated plants were similar (P > 0.05) to those of non-inoculated plants grown with soluble P (Fig. 3a, b). On the other hand, the root dry mass was significantly higher than that of non-inoculated or R4M-F-inoculated plants (1.7 and 2.2 times, respectively) grown in soluble P (Fig. 3c).
Growth of P. vulgaris after 6 weeks of incubation under the environmental conditions described in “Materials and methods.” Dry shoot mass (a), foliar area (b), dry root mass (c), and root/shoot ratio for inoculated (with E. aerogenes R4M-A or Burkholderia spp. R4M-F strains) and non-inoculated plants (NI) grown either with soluble P (gray bars) or insoluble P (white bars) sources. Bars indicate standard error (n = 7). Histograms with different letters are significantly different (P < 0.05) according to Tukey’s multiple comparison test
The P content in fully expanded leaves was affected by the bacterial inoculation as well as by the chemical form of P supplied to the plants (Fig. 4). Non-inoculated plants grown with insoluble P and without P fertilization had P levels similar to each other and significantly lower than those grown with soluble P (1.25 ± 0.13, 1.16 ± 0.09, 6.23 ± 0.66, respectively). Inoculation with E. aerogenes R4M-A strain significantly increased the P content under insoluble P conditions. The P level of leaves was about two times higher than those of the other treatments (Fig. 4). In contrast, under high P availability, neither inoculation with R4M-A nor that with R4M-F affected the P content (Fig. 4). This indicates that R4M-A inoculation is associated with higher levels of P content in common bean leaves under insoluble P conditions, which could be related to the bacterial P solubilization activity.
Effect of inoculation on P and N content in leaves. Inoculated plants (with E. aerogenes R4M-A or Burkholderia spp. R4M-F strains) and non-inoculated plants (NI) were grown in either soluble (gray bars) or insoluble (white bars) P sources. Bars indicate standard error (n = 3). Histograms with different letters are significantly different (P < 0.05) according to Tukey’s multiple comparison test
The N levels in the trifoliate leaves of common beans were higher in the plants inoculated with E. aerogenes R4M-A strain than in those inoculated with Burkholderia spp. R4M-F and non-inoculated plants (Fig. 4) regardless of the form of P supplied in the medium (P < 0.01 and P < 0.05 for insoluble and soluble phosphates, respectively). The leaf N/P ratio of plants grown under P-limited conditions was markedly lower in plants inoculated with R4M-A (11 ± 1.5) than in those under the other treatments (17.6 ± 1.9 and 17.8 ± 1.3 for non-inoculated and inoculated with R4M-F, respectively).
Effect of inoculation on photosynthesis
The E. aerogenes R4M-A strain significantly affected leaf photosynthetic capacity (A max) of plants grown under insoluble P (P < 0.0001; Fig. 5) by increasing A max from 0.9 ± 0.07 to 24.7 ± 1.5 μmol CO2 m−2 s−1 when the CO2 supply to the leaf sample varied from 50 to 800 μmol mol−1 (Fig. 5a). Similar results were obtained with plants grown under soluble P. The A max of R4M-A-inoculated plants at 400 μmol CO2 mol−1 was 1.3–1.6-fold higher than that of non-inoculated plants subjected to insoluble P. When A max was related to leaf P concentration, we found that photosynthetic rates increased 1.5 times for leaf P concentration increasing from 1.4 to 2.8 mg g−1 leaf dry weight (Fig. 5b), showing a linear positive relationship (Pearson r = 0.59, P = 0.0097). At higher leaf P content, photosynthesis remained unchanged. On the other hand, the stomatal conductance was negatively affected by the leaf P concentration independently of the inoculation treatment. Although the reduction of stomatal conductance in plants subjected to TCP did not differ significantly with that of plants subjected to soluble P, the reduction was lower in R4M-A-inoculated plants (10%) than in R4M-F-inoculated (20.2%) and non-inoculated ones (37.8%).
Photosynthetic rates determined in mature leaves of common beans. Plant inoculation and determination of photosynthesis rate were performed as described in “Materials and methods.” a Photosynthesis CO2 response curves. Photosynthesis measurements were made on the third fully expanded leaf at an irradiance of 1,500 μmol photon m−2 s−1 (λ, 400–700 nm). b Photosynthetic rate as related to leaf P concentration. The measurements were made at 350 μmol mol−1 CO2 in the sample leaf and at the same light conditions as described in a. Filled and open squares Non-inoculated; Filled and open circles E. aerogenes R4M-A; Filled and open diamonds, Burkholderia spp. R4M-F. Black and white symbols indicate soluble and insoluble phosphate forms, respectively. In all cases, the bars indicate error standard (n = 3, P < 0.05, Tukey’s multiple comparisons test). In b, bars are standard errors for leaf P concentration (x-axis) and rate of photosynthesis (y-axis)
Discussion
Distinct and diverse populations of phosphate-solubilizing bacteria in soil and plant rhizosphere have been isolated and characterized (Rodríguez et al. 2006). However, the information available on the occurrence of PSB in acid soils is limited (Pal 1998; Pérez et al. 2007). In addition, most reports related to PSB diversity have focused on particular groups or taxa instead of considering the whole community (Rivas et al. 2006; Gulati et al. 2008; Naik et al. 2008). In the present study, we have assessed the genetic diversity of the entire culturable PSB community isolated from roots, rhizosphere, and non-rhizosphere soil of yerba mate plants growing in an acid ultisol soil. The high number of genotypes identified with efficient P solubilization showed that the community composition of PSB is highly diverse, probably because the bacteria were isolated from low P soil. This diversity of species sharing the ability to mobilize P is an essential bacterial property to inhabit microenvironments with P-limited soil conditions (Goldstein et al. 1999).
Analysis of the PSB allowed taxonomic assignment to previously reported mineral P solubilizers (Kampfer 2007). The majority of the isolates belong to the Enterobacteriaceae family and showed high P solubilization activity which was repressed by soluble P. Our results showed Enterobacteriaceae as the predominant group displaying high P solubilization activity in acid soils with P deficiency (Chung et al. 2005; Son et al. 2006; Pérez et al. 2007).
Structure of the bacterial community was diverse as found in similar studies, but without isolates identified in a tropical soil of Taiwan (Chen et al. 2006) and with a few genotypes isolated from acid soils of Venezuela (Pérez et al. 2007). Environmental factors such as soil texture, moisture, management of soil, and type of crop can affect the composition of the PSB community. High variability was observed in the P solubilization ability among the isolates. Although the activity of most strains decreased in the presence of soluble P, the inhibition level was variable ranging from total to low inhibition. In Erwinia herbicola and B. cepacia, the P solubilization activity was repressed by elevated levels of exogenous P (Goldstein and Liu 1987; Rodríguez and Fraga 1999), whereas that of Rhizobium leguminosarum was not affected (Halder et al. 1990). Mikanová and Nováková (2002) reported a number of PSB with different sensitivities to soluble P.
An important issue for the potential use of PSB is the growth stimulation when plants are inoculated. In addition, in the case of the beneficial effect, it is important to find out which specific trait of the plant is affected by the inoculation. Four efficient P-solubilizing isolates out of 518 P-solubilizing bacteria isolated from an acid soil were tested for their effects on growth of bean plants. One of these isolates, R4M-A, assigned to species E. aerogenes, stimulated the plant growth and increased leaf P and N concentration under P-insoluble conditions. In contrast, inoculation with R4M-F strain had no positive effects despite the in vitro assay showing that both bacterial strains solubilized P. In addition, since the inoculation with R4M-A strain also increased the root dry mass of plants growing either on soluble or insoluble P, it is possible that factors other than P solubilization activity might be involved. Therefore, we tested E. aerogenes R4M-A and Burkholderia spp R4M-F strains for IAA production in culture medium and found that both produced and released free IAA. IAA production by R4M-A strain was significantly higher (7.3 ± 0.4 μg mL−1) than that by R4M-F (2.3 ± 0.5 μg mL−1) and comparable to that produced by Azospirillum brasilense Cd (Li et al. 2000; Mirza et al. 2001; Perrig et al. 2007). IAA produced by several bacteria can stimulate the development and proliferation of roots, with increases in uptake of water and nutrients (Bashan and de-Bashan 2005). Therefore, the positive effect on growth with non-soluble P may result from the synergic combination of both bacterial capacities for IAA production and P mobilization. The stimulation of the root system, the ability to solubilize inorganic P, and the increase in the leaf N content could be an important trait for R4M-A strain to improve plant growth under limited P availability in soil.
Nitrogen and P deficiency limit plant growth by restricting leaf area development and consequently by reducing the photosynthetic rate (Reich et al. 2009). Phosphorus-deficient conditions can affect several plant processes (Rao and Terry 1995; Plaxton 1998; Campbell and Sage 2006), including N partitioning (Warren et al. 2005), and thus, there is an interactive effect between N and P availability (Whitehead et al. 2005). Recently, Reich et al. (2009) concluded that in a P-limited ecosystem, as in the subtropical soils, the A max/N ratio may be constrained by low P. Our results demonstrated the beneficial effect of bacterial inoculation on the photosynthetic rate of plants grown in the presence of insoluble P forms, which could be partly explained by a better leaf N/P status and an improvement in stomatal conductance.
In summary, novelties of our work are the demonstration that the abilities to solubilize P in vitro and promote plant growth are not necessarily associated with each other and that plant photosynthesis affected by P deficiency can be relieved by inoculating PSB. We believe that our results provide the basis to consider some PSB representatives of a great potential for agro-biotechnological applications, in particular for the use in acid soils. Further studies will be needed to validate promoting effects of inoculation under field conditions.
References
Alippi AM, Aguilar OM (1998) Characterization of isolates of Paenibacillus larvae subsp. larvae from diverse geographical origin by the polymerase chain reaction and BOX primers. J Invertebr Pathol 72:21–27
Arnon DJ, Hoagland DR (1940) Crop production in artificial culture solution with special reference to factors influencing yields absorption of inorganic nutrients. Soil Sci 60:463–485
Bashan Y, de-Bashan LE (2005) Bacteria/plant growth-promotion. In: Hillel D (ed) Encyclopedia of soils in the environment. Elsevier, Oxford, UK, pp 103–115
Bielski RL (1973) Phosphate pools, phosphate transport, and phosphate availability. Annu Rev Plant Physiol 24:225–252
Bosshard PP, Zbinden R, Abels S, Böddinghaus B, Altwegg M, Böttger EC (2006) 16S rRNA gene sequencing versus the API 20 NE System and the VITEK 2 ID-GNB Card for identification of non-fermenting gram-negative bacteria in the clinical laboratory. J Clin Microbiol 44:1359–1366
Bremmer JM, Mulvaney CS (1982) Total nitrogen. In: Page AL, Miller RH, Keeney DR (eds) Methods of soil analysis, 2nd edn. Agronomy, ASA, Madison, pp 595–623
Broughton WJ, Hernandez G, Blair M, Beebe S, Gepts P, Vanderleyden J (2003) Beans (Phaseolus spp.): model food legumes. Plant Soil 249:291–363
Campbell CD, Sage RF (2006) Interactions between the effect of atmospheric CO2 content and P nutrition on photosynthesis in white lupin (Lupinus albus L.). Plant Cell Environ 29:844–853
Chen YP, Rekha PD, Arunshen AB, Lai WA, Young CC (2006) Phosphate solubilizing bacteria from subtropical soil and their tricalcium phosphate solubilizing abilities. Appl Soil Ecol 34:33–41
Chun J, Lee JH, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Chung H, Park M, Madhaiyan M, Seshadri S, Song J, Cho H, Sa T (2005) Isolation and characterization of phosphate solubilizing bacteria from the rhizosphere of crop plants of Korea. Soil Biol Biochem 37:1970–1974
Fearnside PM (1998) Phosphorous and human carrying capacity in Brazilian Amazonia. In: Lynch JP, Deikman J (eds) Phosphorous in plant biology: regulatory roles in molecular, cellular, organismic, and ecosystem processes. American Society Plant Physiology, Rockville, pp 94–108
Goldstein AH (2007) Future trends in research on microbial phosphate solubilization: one hundred years of insolubility. In: Velazquez E, Rodriguez-Barrueco C (eds) First International Meeting on Microbial phosphate solubilization. Springer, Dordrecht, pp 91–96
Goldstein AH, Liu ST (1987) Molecular cloning and regulation of a mineral phosphate solubilizing gene from Erwinia herbicola. Bio/Technology 5:72–74
Goldstein AH, Braverman K, Osorio N (1999) Evidence for mutualism between a plant growing in a phosphate-limited desert environment and a mineral phosphate solubilizing (MPS) rhizobacterium. FEMS Microbiol Ecol 30:295–300
Guiñazu LB, Andres JA, Del Papa MF, Pistorio M, Rosas SB (2010) Response of alfalfa (Medicago sativa L.) to single and mixed inoculation with phosphate-solubilising bacteria and Sinorhizobium meliloti. Biol Fertil Soils 46:185–190
Gulati A, Rahi P, Vyas P (2008) Characterization of phosphate-solubilizing fluorescent Pseudomonads from the rhizosphere of Seabuckthorn growing in the cold deserts of Himalayas. Curr Microbiol 56:73–79
Gyaneshwar P, Parekh LJ, Archana G, Poole PS, Collins MD, Hutson RA, Naresh Kumar G (1999) Involvement of a phosphate starvation inducible glucose dehydrogenase in soil phosphate solubilisation by Enterobacter asburiae. FEMS Microbiol Lett 171:223–229
Halder AK, Mishra AK, Bhattacharyya P, Chakrabartty PK (1990) Solubilization of rock phosphate by Rhizobium and Bradyrhizobium. J Gen Appl Microbiol 36:81–92
Hernández G, Ramírez M, Valdés-López O, Tesfaye M, Graham M, Czechowski T, Schlereth A, Wandrey M, Erban A, Cheung F, Wu HC, Lara M, Town CD, Kopka J, Udvardi MK, Vance CP (2007) Phosphorus stress in common bean: root transcript and metabolic responses. Plant Physiol 144:752–767
Hesse PR (1971) Soil phosphorus: its measurements and its uptake by plants. Aust J Soil Res 35:227–239
Jorquera MA, Hernandez MT, Rengel Z, Marschner P, Mora M (2008) Isolation of culturable phosphobacteria with both phytate-mineralization and phosphate-solubilization activity from the rhizosphere of plants grown in a volcanic soil. Biol Fertil Soils 44:1025–1034
Kampfer P (2007) Taxonomy of phosphate solubilizing bacteria. In: Velazquez E, Rodriguez-Barrueco C (eds) First International Meeting on Microbial Phosphate Solubilization. Springer, Dordrecht, pp 101–106
Li J, Ovakim DH, Charles TC, Glick BR (2000) An Acc deaminase minus mutant of Enterobacter cloacae UW4 no longer promotes root elongation. Curr Microbiol 41:101–105
Mehta S, Nautiyal CS (2001) An efficient method for qualitative screening of phosphate-solubilizing bacteria. Curr Microbiol 43:51–56
Mikanová O, Nováková J (2002) Evaluation of the P-solubilizing activity of soil microorganisms and its sensitivity to soluble phosphate. Rostl Výroba 48:97–400
Mirza MS, Ahmad W, Latif F, Haurat J, Bally R, Normand P, Malik KA (2001) Isolation, partial characterization, and the effect of plant growth-promoting bacteria (PGPB) on micro-propagated sugarcane in vitro. Plant Soil 237:47–54
Murphy J, Riley JP (1962) A modified single solution method for the determination of phosphate in natural waters. Anal Chim Acta 27:31–36
Naik PR, Raman G, Narayanan KB, Sakthivel N (2008) Assessment of genetic and functional diversity of phosphate solubilizing fluorescent pseudomonads isolated from rhizospheric soil. BMC Microbiol 8:230. doi:10.1186/1471-2180-8-230
Nautiyal C (1999) An efficient microbiological growth medium for screening phosphate solubilizing microorganisms. FEMS Microbiol Lett 170:265–270
Pal S (1998) Interactions of an acid tolerant strain of phosphate solubilizing bacteria with a few acid tolerant crops. Plant Soil 198:169–177
Pérez E, Sulbarán M, Ball M, Yarzábal L (2007) Isolation and characterization of mineral-phosphate-solubilizing bacteria naturally colonizing a limonitic crust in the south-eastern Venezuelan region. Soil Biol Biochem 39:2905–2914
Perrig D, Boiero L, Masciarelli O, Penna C, Cassán F, Luna V (2007) Plant growth promoting compounds produced by two agronomically important strains of Azospirillum brasilense, and their implications for inoculant formulation. Appl Microbiol Biotechnol 75:1143–1150
Plaxton WC (1998) Metabolic aspects of phosphate starvation in plants. In: Lynch JP, Deikman J (eds) Phosphorous in plant biology: regulatory roles in molecular, cellular, organismic, and ecosystem processes. American Society Plant Physiology, Rockville, pp 229–241
Rao IM, Terry N (1995) Leaf phosphate status, photosynthesis, and carbon partitioning in sugar beet. IV. Changes with time following increased supply of phosphate to low-phosphate plants. Plant Physiol 107:1313–1321
Reich P, Oleksyn J, Wright I (2009) Leaf phosphorus influences the photosynthesis–nitrogen relation: a cross-biome analysis of 314 species. Oecologia 160:207–212. doi:10.1007/s00442-009-1291-3
Richardson A (2001) Prospect for using soil microorganisms to improve the acquisition of phosphorous by plants. Aust J Plant Physiol 28:897–906
Rivas R, Reix A, Mateos PF, Trujillo ME, Martínez-Molina E, Velázquez E (2006) Biodiversity of populations of phosphate solubilizing rhizobia that nodulates chickpea in different Spanish soils. Plant Soil 287:23–33
Rodríguez H, Fraga R (1999) Phosphate solubilizing bacteria and their role in plant growth promotion. Biotechnol Adv 17:319–339
Rodríguez H, Fraga R, Bashan Y (2006) Genetics of phosphate solubilization and potential applications for improving plant growth-promoting bacteria. Plant Soil 287:15–21
Son HJ, Park GT, Cha MS, Heo MS (2006) Solubilization of insoluble inorganic phosphates by a novel salt- and pH-tolerant Pantoea agglomerans R-42 isolated from soybean rhizosphere. Bioresour Technol 97:204–210
Tiessen H, Lo Monaco S, Ramirez A, Santos MDC, Shang C (1996) Phosphate minerals in a lateritic crust from Venezuela. Biogeochemistry 34:8–17
Versalovic J, Schneider M, de Bruijn FJ, Lupski JR (1994) Genomic fingerprinting of bacteria using repetitive sequence-based polymerase chain reaction. Meth Mol Cell Biol 5:25–40
Warren CR, McGrath JF, Adams MA (2005) Differential effects of N, P and K on photosynthesis and partitioning of N in Pinus pinaster needles. Ann For Sci 62:1–8
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Whitehead D, Boelman NT, Turnbull MH, Griffin KL, Tissue DT, Barbour MM, Hunt JE, Richardson SJ, Peltzer DA (2005) Photosynthesis and reflectance indices for rainforest species in ecosystems undergoing progression and retrogression along a soil fertility chronosequence in New Zealand. Oecologia 144:233–244
Acknowledgments
The authors thank Dr Chandra Nautiyal, NBRI, Lucknow, India, for kindly gifting us Pseudomonas RHP3 strain. Thanks also to Belén Burdyn for the technical support. This work was supported by a grant from Agencia de Promoción Científica y Tecnológica de Argentina. M.M.C., P.A.S., L.A.M., and O.M.A. were supported by The National Scientific and Technical Research Council-CONICET, Argentina.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Collavino, M.M., Sansberro, P.A., Mroginski, L.A. et al. Comparison of in vitro solubilization activity of diverse phosphate-solubilizing bacteria native to acid soil and their ability to promote Phaseolus vulgaris growth. Biol Fertil Soils 46, 727–738 (2010). https://doi.org/10.1007/s00374-010-0480-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-010-0480-x