Abstract
Objectives
To evaluate radiomics studies according to radiomics quality score (RQS) and Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis (TRIPOD) to provide objective measurement of radiomics research.
Materials and methods
PubMed and Embase were searched for studies published in high clinical imaging journals until December 2018 using the terms “radiomics” and “radiogenomics.” Studies were scored against the items in the RQS and TRIPOD guidelines. Subgroup analyses were performed for journal type (clinical vs. imaging), intended use (diagnostic vs. prognostic), and imaging modality (CT vs. MRI), and articles were compared using Fisher’s exact test and Mann-Whitney analysis.
Results
Seventy-seven articles were included. The mean RQS score was 26.1% of the maximum (9.4 out of 36). The RQS was low in demonstration of clinical utility (19.5%), test-retest analysis (6.5%), prospective study (3.9%), and open science (3.9%). None of the studies conducted a phantom or cost-effectiveness analysis. The adherence rate for TRIPOD was 57.8% (mean) and was particularly low in reporting title (2.6%), stating study objective in abstract and introduction (7.8% and 16.9%), blind assessment of outcome (14.3%), sample size (6.5%), and missing data (11.7%) categories. Studies in clinical journals scored higher and more frequently adopted external validation than imaging journals.
Conclusions
The overall scientific quality and reporting of radiomics studies is insufficient. Scientific improvements need to be made to feature reproducibility, analysis of clinical utility, and open science categories. Reporting of study objectives, blind assessment, sample size, and missing data is deemed to be necessary.
Key Points
• The overall scientific quality and reporting of radiomics studies is insufficient.
• The RQS was low in demonstration of clinical utility, test-retest analysis, prospective study, and open science.
• Room for improvement was shown in TRIPOD in stating study objective in abstract and introduction, blind assessment of outcome, sample size, and missing data categories.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Radiomics research has been rapidly expanding ever since Gilles et al declared “images are data” [1]. Sophisticated bioinformatics tools are applied to reduce data dimensionality and select features from high-dimensional data, and models with potential diagnostic or prognostic utility are typically developed [1,2,3]. Although radiomics research shows great potential, its current use is confined to the academic literature, without real-world clinical applications. High quality in science and reporting may present strategies for radiomics to become an effective imaging biomarker able to cross the “translational gap” [4, 5] for use in guiding clinical decisions.
The quality of scientific research articles consists of two elements: the quality of the science and the quality of the report [6], and deficiencies in either may hamper translation of biomarkers to patient care [7]. With regard to the quality of the science, a system of metrics in the form of the radiomics quality score (RQS) was developed by the expert opinions of Lambin et al [2], to determine the validity and completeness of radiomics studies. The RQS consists of 16 components that consider radiomics-specific high-dimensional data and modeling and accounts for image protocol and feature reproducibility, biologic/clinical validation and utility, performance index, high level of evidence, and open science. With regard to the quality of reporting, radiomics research is a model-based approach and reporting according to the Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis (TRIPOD) initiative [8] is desirable.
To our knowledge, the quality of the science and reporting in radiomics research studies is largely unknown. A RQS study from the score developer [3] reported an average score of less than 50% over 41 radiomics studies, but the RQS score is underutilized because many investigators and peer reviewers are unfamiliar with it. Prediction model studies in the clinical domain showed suboptimal quality of reporting according to TRIPOD [9], but whether the radiomics domain is good or bad at reporting has not been studied. In this study, we evaluated radiomics studies using RQS and TRIPOD items to evaluate their scientific quality and assessed whether the score and degree of adherence depend on the study design or journal type. The purpose of the study was therefore to evaluate the quality of the science and reporting of radiomics studies according to RQS and TRIPOD.
Materials and methods
Article search strategy and study selection
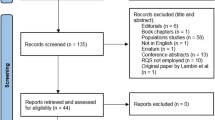
A search was conducted for all potentially relevant original research papers using radiomics analysis published up until December 3, 2018. The search terms used to find radiomics studies were “radiomic” OR “radiogenomic” in the MEDLINE (National Center for Biotechnology Information, NCBI) and EMBASE databases. The eligible articles were high-impact factor medical journals ranked higher than 7.0 according to the 2018 edition of the Journal Citation reports, as well as those published in radiology journal of Radiology and European Radiology. The impact factor of 7.0 was chosen as it was considered that articles published in journals above 7.0 would be representative of the reporting of high-quality clinical studies on radiomics analysis. Imaging journals were chosen because they are the highest-ranked US and non-US general radiology, given the impact and status of the two journals. The inclusion process is shown in Fig. 1. Study selection and data extraction are shown in Supplementary Materials 1.
Analysis of method quality based on RQS
The RQS score with 16 components is defined in Supplementary Table 1 [2]. The reviewers extracted the data using a predetermined RQS evaluation according to six domains. Domain 1 covers the protocol quality and reproducibility in image and segmentation: well-documented image protocols (1 point) and/or usage of public image protocols (1 point), multiple segmentations (1 point), phantom study (1 point), and test-retest analysis with imaging at multiple time points (1 point). Domain 2 covers the reporting of feature reduction and validation: feature reduction or adjustment for multiple testing (3 or − 3 points) and validation (− 5 to 5 points). Domain 3 covers the reporting of the performance index: reporting of discrimination statistics (1 point) with resampling (1 point), calibration statistics (1 point) with resampling (1 point), and application of cut-off analyses (1 point). Domain 4 covers the reporting of biological/clinical validation and utility: multivariate analysis with non-radiomics features (1 point), biological correlates (1 point), comparison with the gold standard (2 points), and potential clinical utility (2 points). Domain 5 covers the demonstration of a higher level of evidence: by conducting a prospective study (7 points) or cost-effectiveness analysis (2 points). The final domain (domain 6) covers open science, with open availability of source code and data (4 points).
The six domains and topics which were subject to further discussions until a consensus was reached were in Supplementary Materials 1.
Analysis of reporting completeness based on TRIPOD statement
The TRIPOD checklist was applied to each article to determine the completeness of reporting. The details of the checklist are described elsewhere [8], but it consists of 22 main criteria with 37 items. First, the type of prediction model was decided, whether the radiomics model was development only (type 1a), development and validation using resampling (type 1b), random split-sample validation (type 2a), nonrandom split-sample validation (type 2b), validation using separate data (type 3), or validation only (type 4). The details for TRIPOD checklist and data extraction are shown in Supplementary Materials 1.
Analysis of the role of radiologists
To demonstrate the role of radiologists in the radiomics studies, the analysis was undertaken to calculate the position and number of radiologist among the author lists. The radiologists include general radiologists and nuclear medicine radiologists. First, the main authors, either first or corresponding author, were checked. When the radiologists are not main authors, the position and number of radiologists among the author’s lists were checked. The position is checked for the first appearance (i.e., 3rd and 5th author are radiologists among 8 authors, the position was checked as 3/8, 0.37).
Statistical analysis
For the six domains in the RQS (protocol quality and segmentation, feature selection and validation, biologic/clinical validation and utility, model performance index, high level of evidence, and open science and data), basic adherence was assigned when a score of at least 1 point was obtained without minus points. The basic adherence to RQS (for 0–16 criteria) and each item scored in TRIPOD were counted (range, 0–35 items) and calculated in a descriptive manner using proportions (%). The TRIPOD item 5c (“if done” item) and the validation items 10c, 10e, 12, 13c, 17, and 19a were excluded from both the numerator and denominator when the overall adherence rate was calculated. For all included articles, the total RQS score was calculated (score range, − 8 to 36) and expressed as mean ± standard deviation. A graphical display for the proportion of studies was adopted from the suggested graphical display for Quality Assessment of Diagnostic Accuracy Studies-2 results [10].
Subgroup analyses were performed to determine whether the reporting quality differed according to intended use (diagnostic or prognostic), journal type (clinical or imaging journal), and imaging modality (CT or MRI). Additionally, we compared RQS between radiogenomics studies and non-radiogenomics studies. Before subgroup analysis, the RQS was plotted for each journal to observe whether there was a systematic difference between journals (Supplementary Figure 2). As no systematic difference was observed between journals, the journal was not adjusted for in the analysis. The nonparametric Mann-Whitney U test was used to compare the RQS score in each group. Fisher’s exact test was used to compare proportions in RQS and TRIPOD for small sample sizes in each group. All statistical analyses were performed using SPSS (SPSS version 22; SPSS) and R (R version 3.3.3; R Foundation for Statistical Computing), and a p value < .05 was considered statistically significant.
Results
Characteristics of the included studies
Seventy-seven articles [11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87] were finally analyzed. The journal, impact factor, study topic, intended use, imaging modality, number of patients, and model type are summarized in Supplementary Table 2. The number and characteristics of the included radiomics studies are provided in Table 1 and Fig. 2. The mean patient number was 232 (standard deviation, 248.7; range, 38–2029). The studies were published in 2014 (1 article), 2016 (8 articles), 2017 (16 articles), and 2018 (52 articles). There were 25 articles published in high IF clinical journals and 52 articles in imaging journals (14 in Radiology and 38 in European Radiology). Most articles were oncologic studies (90.9%). Radiomics analysis was most frequently studied as a diagnostic biomarker (80.5%), then as a prognostic (19.5%) biomarker. MRI was the most studied modality (66.0%), followed by CT (26.0%), and PET or US (each 4.0%). Analysis of the validation methods revealed that external validation was missing in 63 out of 77 studies (81.8%). In the oncologic studies, the study purposes most frequently included histopathologic grade and differential diagnosis (51.9%), followed by molecular or genomic classification (21.4%), survival prediction (12.8%), and assessment of treatment response (11.4%).
RQS according to the six key domains
Table 2 summarizes the results. The averaged RQS of the 77 studies expressed as a percentage of the ideal score according to the six key domains is shown in Fig. 3. The mean RQS score of the 36 studies was 9.40 (standard deviation, 5.60), which was 26.1% of the ideal score of 36. The lowest score was − 5, and the highest score was 21 (58.3% of the ideal quality score).
In domain 1, all studies except one reported well-documented image acquisition protocols or the use of publicly available image databases. Multiple segmentations by two readers were performed in 35 of the 77 studies (45.4%), including six studies with automatic segmentation. Notably, only five studies [11, 12, 20, 26, 47] conducted imaging at multiple time points and tested feature robustness. No articles conducted a phantom study.
In domain 2, most studies adopted appropriate feature reduction or adjustment for multiple testing (74/77, 96.1%). The studies used either false discovery rate with univariate logistic regression or two-sample t tests (for binary outcomes), and a variety of statistical and machine learning methods such as lasso, elastic net, random forest, recursive feature elimination, and support vector machine. Validation was performed without retraining from the same or a different institute in 70.1% of studies (54 out of 77).
In domain 3, all studies used discriminative statistics, but one study [23] provided hazard ratios and p values from a log-rank test for survival analysis instead of the C-index.
In domain 4, half of the studies evaluated relationships between the radiomics features and non-radiomics features (50.6%), but only 28.6% of studies found biological correlates of radiomics to provide a more holistic model and imply biological relevance. Less than half of the studies (46.7%) compared results with an existing gold standard. By contrast, in terms of clinical utility, only 15 studies (19.5%) analyzed a net improvement in health outcomes using decision curve analysis or other statistical tools.
Surprisingly, studies were deficient in demonstrating a high level of evidence such as a prospective design or cost-effectiveness analysis. Only three studies [21, 48, 55] (3.9%) included prospective validation, and no studies conducted cost-effective analysis. For domain 6, only three studies [33, 34, 46] (3.9%) made their code and/or data publicly available.
Both feature reduction and validation were missing from the study [25] with the lowest score. Meanwhile, seven studies with the highest scores [12, 14, 21, 26, 48, 54, 55] (three articles with a RQS score of 16, 1 article with 18, 1 article with 19, and 1 article with 21) earned additional points by using publicly available images [12], multiple segmentation [12, 14, 26], test-retest analysis [12, 26], and validation using three or more datasets [12, 21, 26, 55], demonstrating potential clinical utility using decision curve analysis [14, 54] and conducting prospective validation [21, 48, 55], with all studies fulfilling requirements for image protocol quality, feature reduction, and use of a discrimination index.
Completeness in reporting a radiomics-based multivariable prediction model using TRIPOD
The mean number of TRIPOD items reported was 18.51 ± 3.96 (standard deviation; range, 11–26) when all 35 items were considered. The adherence rate for TRIPOD was 57.8% ± 10.9% (standard deviation; range, 33–78%) when “if relevant” and “if done” items were excluded from both the numerator and denominator. The completeness of reporting individual TRIPOD items is shown in Table 3. The detailed results are shown in Supplementary Materials 2.
Subgroup analysis
The results of the subgroup analysis are shown in Table 4. Prognostic studies showed a trend for a higher RQS score than diagnostic studies (11.83 ± 5.03 vs. 8.93 ± 5.52), but this was not statistically significant. Prognostic studies received a higher score than diagnostic studies in comparison with a “gold standard” (p < .001) and using cut-off analysis (p < .001). This was reflected in the TRIPOD items, with the prognostic studies showing higher adherence rates in “describing risk group” (p = .007) and “report unadjusted association between predictors and outcome” (if done, p = .017).
Studies in clinical journals also showed significantly higher RQS scores than those in imaging journals (12.2 ± 5.23 vs. 8.03 ± 5.17, p = .001). They achieved a higher score in protocol quality (p = .018), test-retest analysis (p < .001), validation (p = .012), multivariable analysis with non-radiomics features (p = .036), finding biologic correlates (p = .009), and conducted prospective study (p = .011). In the reporting quality, studies in clinical journals well reported the study design or source of data (p = .047) and reported unadjusted association. Meanwhile, studies in imaging journal more frequently reported blind assessment of predictors (p = .038), the flow of participants (p = .012), and number of predictors and outcomes (p = .001).
Studies utilizing CT tended to have higher RQS than those using MRI (11.8 ± 3.71 vs. 9 ± 5.2), but this trend was not statistically significant. Studies using CT received a higher score in test-retest analysis (p = .028), discrimination statistics with resampling or cross-validation (p = .011), and cut-off analysis (p = .039) than those using MRI. In the TRIPOD items, studies using CT clearly stated study objective and setting in the abstract (p = .006) and described both discrimination and calibration (p = .017) and more provided the full prediction model (p = .014) than those using MRI.
There were 15 articles studied radiogenomics (19.6% among total, 21.4% among oncologic studies). There was no significant difference in radiomics quality score (Mann-Whitney U test, p = .862) between that of radiomics studies (median 10.5, interquartile range 5.0–13.0) and radiogenomics studies (median 10.0, interquartile range 4.25–14.7) and according to each domain.
Role of radiologists in radiomics studies
The results are shown in the Supplementary Table 3. There were 18 articles (23.4%) that radiologists were not the main authors. Three articles (3.9%) did not have radiologists in the author list. When radiologists were not the main authors, the relative position of radiologist in the author list was 0.5, which indicates middle position in the entire author lists.
Discussion
In this study, radiomics studies were evaluated in respect to the quality of both the science and the reporting, using RQS and TRIPOD guidelines. Radiomics studies were insufficient in regard to both the quality of the science and the reporting, with an average score of 26.1% of the ideal RQS and 57.8% of the maximum adherence rate to the TRIPOD reporting guidelines. No study conducted a phantom study or cost-effective analysis and a high level of evidence for radiomics studies, with further limitations being demonstrated in the openness to data and code. Half of the items that the TRIPOD statement deems necessary to report in multivariable prediction model publications were not completely recorded in the radiomics studies. Our results imply that radiomics studies require significant improvement in both scientific and reporting quality.
The six key RQS domains used in this study were designed to support the integration of the RQS in radiomics approaches. Adopted from the consensus statement of the FDA-NIH Biomarker Working Group [4], the three aspects of technical validation, biological/clinical validation, and assessment of cost-effectiveness for imaging biomarker standardization were included in domains 1 (image protocol and feature reproducibility), 4 (biologic/clinical validation), and 5 (high level of evidence), respectively. With regard to technical validation, radiomics approaches are yet to become a reliable measure for the testing of hypotheses in clinical cancer research, with insufficient data supporting their precision or technical bias. Precision analysis using repeatability and reproducibility testing was conducted in one study [47], but reproducibility needs to be tested using different geographical sites and different equipment. Furthermore, none of the evaluated studies reported analysis of technical bias using a phantom study, which describes the systemic difference between the measurements of a parameter and its real values [88]. For clinical validation, prospective testing of an imaging biomarker in clinical populations is required [89], and only three reports covered a prospective study in the field of neuro-oncology. After biological/clinical validation, the cost-effectiveness of radiomics needs to be studied to ensure that it provides good value for money compared with the other currently available biomarkers. From the current standpoint, the time when radiomics will achieve this end seems far away, and technical and clinical validation is still required.
Validation in TRIPOD pursues external validation, which was performed in only 18.2% of the reports covered in the present study, while independent validation including internal validation is acceptable in RQS, which accounts for 70.1% of the studies. In the reporting of radiomics studies, the highly problematic TRIPOD items existed. In the title, only two radiomics studies explicitly wrote the term “development” or “validation” with the target population and outcome. There are several elements that should be present in the abstract, and one of these was missing in 92.2% of studies, as they did not explicitly describe “development” or “validation” in the study objective, or did not describe whether the study design was a randomized controlled trial, a cohort study, or a case-control design. Furthermore, reporting of the sample size calculation and the handling of missing data were often poorly conducted. These results are similar to the findings of a previous systematic review of TRIPOD adherence that examined publications using clinical multivariable prediction models [9]. Differing from the findings on the clinical multivariable prediction models, the radiomics studies were excellent (100%) in the criterion involving the definition of all predictors and how quantitative features are handled. However, the reporting of blindness to the outcome was insufficient (32.1%). The results for blindness were similar to the adherence to the Standards for Reporting of Diagnostic Accuracy Studies (STARD) [90, 91], which stated that blinding of both predictors and outcome is still insufficient in both clinical and imaging studies.
Subgroup analysis may provide more specific guidance in radiomics research. Studies in clinical journals showed significantly higher RQS scores, especially in test-retest analysis, multivariable analysis with non-radiomics features, finding biologic correlates, and pursuit of prospective study design. In the TRIPOD items, they more clearly defined the data source and study design, i.e., a consecutive retrospective design or case-control design, and the study setting, i.e., tertiary hospital or general population. In terms of validation, all three prospective studies were in the clinical journal, and both external and independent validation was performed more frequently in the clinical journal group. These findings imply that high-impact clinical journals pursue precision in research, clarification of epidemiological background, and independent validation, which demonstrates the room for improvement in radiomics studies.
Of note, radiologists played the main role as either first or corresponding authors in the radiomics studies. There were 23.4% articles that radiologists were not the main authors, but most studies work collaboratively among radiologists, other clinicians, and physicists.
This study has some potential limitations. The first is the relatively small sample size, especially with an impact factor below 7. This was placed to permit in-depth analysis of the radiomics applications. Second, radiomics is still a developing imaging biomarker, and the suggested RQS may be too “ideal.” The criteria of phantom study and multiple imaging acquisitions may be unrealistic for clinical situations. Third, the adoption of TRIPOD items to radiomics studies can be rather strict. For example, most studies are case-control and retrospective study designs, and a clear description of “case-control” is not commonly given in imaging journals. Nonetheless, clear stating of the participants and study setting is important for study transportability, and studies in imaging journals need to pursue this. Fourth, we considered internal validation with a random sample or split sample as independent validation, while the TRIPOD statement only considers external validation for validation of a pre-existing model. When the rates of open science and open data increase in the field of radiomics, the true validation of a model should become easier to perform.
In conclusion, the overall scientific quality and reporting of radiomics studies is insufficient, with the scientific quality showing the greatest deficiencies. Scientific improvements need to be made to feature reproducibility, analysis of clinical utility, and open science. Reporting of study objectives, blind assessment, sample size, and missing data is deemed to be necessary. Our intention is to promote the quality of radiomics research studies as diagnostic and prognostic prediction models, and the above criteria and items need to be pursued for radiomics to become a viable tool for medical decision-making.
Abbreviations
- RQS:
-
Radiomics quality score,
- TRIPOD:
-
Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis
References
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577
Lambin P, Leijenaar RTH, Deist TM et al (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14:749–762
Sanduleanu S, Woodruff HC, de Jong EEC et al (2018) Tracking tumor biology with radiomics: a systematic review utilizing a radiomics quality score. Radiother Oncol 127:349–360
O'Connor JP, Aboagye EO, Adams JE et al (2017) Imaging biomarker roadmap for cancer studies. Nat Rev Clin Oncol 14:169–186
Sung NS, Crowley WF Jr, Genel M et al (2003) Central challenges facing the national clinical research enterprise. JAMA 289:1278–1287
Choi YJ, Chung MS, Koo HJ, Park JE, Yoon HM, Park SH (2016) Does the reporting quality of diagnostic test accuracy studies, as defined by STARD 2015, affect citation? Korean J Radiol 17:706–714
Waterton JC, Pylkkanen L (2012) Qualification of imaging biomarkers for oncology drug development. Eur J Cancer 48:409–415
Moons KG, Altman DG, Reitsma JB et al (2015) Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): explanation and elaboration. Ann Intern Med 162:W1–W73
Heus P, Damen JAAG, Pajouheshnia R et al (2018) Poor reporting of multivariable prediction model studies: towards a targeted implementation strategy of the TRIPOD statement. BMC Med 16:120
Whiting PF, Rutjes AW, Westwood ME et al (2011) QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med 155:529–536
Aerts HJ, Velazquez ER, Leijenaar RT et al (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun 5:4006
Hawkins S, Wang H, Liu Y et al (2016) Predicting malignant nodules from screening CT scans. J Thorac Oncol 11:2120–2128
Huang Y, Liu Z, He L et al (2016) Radiomics signature: a potential biomarker for the prediction of disease-free survival in early-stage (I or II) non-small cell lung cancer. Radiology 281:947–957
Huang YQ, Liang CH, He L et al (2016) Development and validation of a radiomics nomogram for preoperative prediction of lymph node metastasis in colorectal cancer. J Clin Oncol 34:2157–2164
Kickingereder P, Bonekamp D, Nowosielski M et al (2016) Radiogenomics of glioblastoma: machine learning-based classification of molecular characteristics by using multiparametric and multiregional MR imaging features. Radiology 281:907–918
Kickingereder P, Burth S, Wick A et al (2016) Radiomic profiling of glioblastoma: identifying an imaging predictor of patient survival with improved performance over established clinical and radiologic risk models. Radiology 280:880–889
Kickingereder P, Gotz M, Muschelli J et al (2016) Large-scale Radiomic profiling of recurrent glioblastoma identifies an imaging predictor for stratifying anti-angiogenic treatment response. Clin Cancer Res 22:5765–5771
Li H, Zhu Y, Burnside ES et al (2016) MR imaging radiomics signatures for predicting the risk of breast cancer recurrence as given by research versions of MammaPrint, oncotype DX, and PAM50 gene assays. Radiology 281:382–391
Nie K, Shi L, Chen Q et al (2016) Rectal cancer: assessment of neoadjuvant chemoradiation outcome based on radiomics of multiparametric MRI. Clin Cancer Res 22:5256–5264
Coroller TP, Agrawal V, Huynh E et al (2017) Radiomic-based pathological response prediction from primary tumors and lymph nodes in NSCLC. J Thorac Oncol 12:467–476
Grossmann P, Narayan V, Chang K et al (2017) Quantitative imaging biomarkers for risk stratification of patients with recurrent glioblastoma treated with bevacizumab. Neuro Oncol 19:1688–1697
Hu LS, Ning S, Eschbacher JM et al (2017) Radiogenomics to characterize regional genetic heterogeneity in glioblastoma. Neuro Oncol 19:128–137
Liu TT, Achrol AS, Mitchell LA et al (2017) Magnetic resonance perfusion image features uncover an angiogenic subgroup of glioblastoma patients with poor survival and better response to antiangiogenic treatment. Neuro Oncol 19:997–1007
Liu Z, Zhang XY, Shi YJ et al (2017) Radiomics analysis for evaluation of pathological complete response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Clin Cancer Res 23:7253–7262
Lohmann P, Stoffels G, Ceccon G et al (2017) Radiation injury vs. recurrent brain metastasis: combining textural feature radiomics analysis and standard parameters may increase (18)F-FET PET accuracy without dynamic scans. Eur Radiol 27:2916–2927
Rios Velazquez E, Parmar C, Liu Y et al (2017) Somatic mutations drive distinct imaging phenotypes in lung cancer. Cancer Res 77:3922–3930
Song SH, Park H, Lee G et al (2017) Imaging phenotyping using Radiomics to predict micropapillary pattern within lung adenocarcinoma. J Thorac Oncol 12:624–632
Wang J, Wu CJ, Bao ML, Zhang J, Wang XN, Zhang YD (2017) Machine learning-based analysis of MR radiomics can help to improve the diagnostic performance of PI-RADS v2 in clinically relevant prostate cancer. Eur Radiol 27:4082–4090
Wu S, Zheng J, Li Y et al (2017) A radiomics nomogram for the preoperative prediction of lymph node metastasis in bladder cancer. Clin Cancer Res 23:6904–6911
Yu J, Shi Z, Lian Y et al (2017) Noninvasive IDH1 mutation estimation based on a quantitative radiomics approach for grade II glioma. Eur Radiol 27:3509–3522
Yuan M, Zhang YD, Pu XH et al (2017) Comparison of a radiomic biomarker with volumetric analysis for decoding tumour phenotypes of lung adenocarcinoma with different disease-specific survival. Eur Radiol 27:4857–4865
Zhang B, Tian J, Dong D et al (2017) Radiomics features of multiparametric MRI as novel prognostic factors in advanced nasopharyngeal carcinoma. Clin Cancer Res 23:4259–4269
Zhou H, Vallieres M, Bai HX et al (2017) MRI features predict survival and molecular markers in diffuse lower-grade gliomas. Neuro Oncol 19:862–870
Akbari H, Bakas S, Pisapia JM et al (2018) In vivo evaluation of EGFRvIII mutation in primary glioblastoma patients via complex multiparametric MRI signature. Neuro Oncol 20:1068–1079
Bae S, Choi YS, Ahn SS et al (2018) Radiomic MRI phenotyping of glioblastoma: improving survival prediction. Radiology 289:797–806
Beukinga RJ, Hulshoff JB, Mul VEM et al (2018) Prediction of response to neoadjuvant chemotherapy and radiation therapy with baseline and restaging (18)F-FDG PET imaging biomarkers in patients with esophageal cancer. Radiology 287:983–992
Bickelhaupt S, Jaeger PF, Laun FB et al (2018) Radiomics based on adapted diffusion kurtosis imaging helps to clarify most mammographic findings suspicious for cancer. Radiology 287:761–770
Chen T, Ning Z, Xu L et al (2018) Radiomics nomogram for predicting the malignant potential of gastrointestinal stromal tumours preoperatively. Eur Radiol 29:1074–1082
Chen Y, Chen TW, Wu CQ et al (2018) Radiomics model of contrast-enhanced computed tomography for predicting the recurrence of acute pancreatitis. Eur Radiol 29:4408–4417
Cui Y, Yang X, Shi Z et al (2018) Radiomics analysis of multiparametric MRI for prediction of pathological complete response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Eur Radiol 29:1211–1220
Dong F, Li Q, Xu D et al (2018) Differentiation between pilocytic astrocytoma and glioblastoma: a decision tree model using contrast-enhanced magnetic resonance imaging-derived quantitative radiomic features. Eur Radiol 29:3968–3975
Dong Y, Feng Q, Yang W et al (2018) Preoperative prediction of sentinel lymph node metastasis in breast cancer based on radiomics of T2-weighted fat-suppression and diffusion-weighted MRI. Eur Radiol 28:582–591
Guo J, Liu Z, Shen C et al (2018) MR-based radiomics signature in differentiating ocular adnexal lymphoma from idiopathic orbital inflammation. Eur Radiol 28:3872–3881
Horvat N, Veeraraghavan H, Khan M et al (2018) MR imaging of rectal cancer: radiomics analysis to assess treatment response after neoadjuvant therapy. Radiology 287:833–843
Hu HT, Wang Z, Huang XW et al (2018) Ultrasound-based radiomics score: a potential biomarker for the prediction of microvascular invasion in hepatocellular carcinoma. Eur Radiol 29:2890–2901
Kang D, Park JE, Kim YH et al (2018) Diffusion radiomics as a diagnostic model for atypical manifestation of primary central nervous system lymphoma: development and multicenter external validation. Neuro Oncol 20:1251–1261
Kickingereder P, Neuberger U, Bonekamp D et al (2018) Radiomic subtyping improves disease stratification beyond key molecular, clinical, and standard imaging characteristics in patients with glioblastoma. Neuro Oncol 20:848–857
Kim JY, Park JE, Jo Y et al (2018) Incorporating diffusion- and perfusion-weighted MRI into a radiomics model improves diagnostic performance for pseudoprogression in glioblastoma patients. Neuro Oncol 21:404–414
Kniep HC, Madesta F, Schneider T et al (2018) Radiomics of brain MRI: utility in prediction of metastatic tumor type. Radiology 180946
Multiparametric ultrasomics of significant liver fibrosis: a machine learning-based analysis. Eur Radiol 29:1496–1506
Li Y, Liu X, Qian Z et al (2018) Genotype prediction of ATRX mutation in lower-grade gliomas using an MRI radiomics signature. Eur Radiol 28:2960–2968
Li Y, Liu X, Xu K et al (2018) MRI features can predict EGFR expression in lower grade gliomas: a voxel-based radiomic analysis. Eur Radiol 28:356–362
Li ZC, Bai H, Sun Q et al (2018) Multiregional radiomics features from multiparametric MRI for prediction of MGMT methylation status in glioblastoma multiforme: a multicentre study. Eur Radiol 28:3640–3650
Liang W, Yang P, Huang R et al (2018) A combined nomogram model to preoperatively predict histologic grade in pancreatic neuroendocrine tumors. Clin Cancer Res 25:584–594
Liu H, Zhang C, Wang L et al (2018) MRI radiomics analysis for predicting preoperative synchronous distant metastasis in patients with rectal cancer. Eur Radiol 29:4418–4426
Lu CF, Hsu FT, Hsieh KL et al (2018) Machine learning-based radiomics for molecular subtyping of gliomas. Clin Cancer Res 24:4429–4436
Lv W, Yuan Q, Wang Q et al (2018) Robustness versus disease differentiation when varying parameter settings in radiomics features: application to nasopharyngeal PET/CT. Eur Radiol 28:3245–3254
Meng X, Xia W, Xie P et al (2018) Preoperative radiomic signature based on multiparametric magnetic resonance imaging for noninvasive evaluation of biological characteristics in rectal cancer. Eur Radiol 29:3200–3209
Naganawa S, Enooku K, Tateishi R et al (2018) Imaging prediction of nonalcoholic steatohepatitis using computed tomography texture analysis. Eur Radiol 28:3050–3058
Niu J, Zhang S, Ma S et al (2018) Preoperative prediction of cavernous sinus invasion by pituitary adenomas using a radiomics method based on magnetic resonance images. Eur Radiol 29:1625–1634
Ortiz-Ramón R, Larroza A, Ruiz-España S, Arana E, Moratal D (2018) Classifying brain metastases by their primary site of origin using a radiomics approach based on texture analysis: a feasibility study. Eur Radiol 28:4514–4523
Park YW, Oh J, You SC et al (2018) Radiomics and machine learning may accurately predict the grade and histological subtype in meningiomas using conventional and diffusion tensor imaging. Eur Radiol 29:4068–4076
She Y, Zhang L, Zhu H et al (2018) The predictive value of CT-based radiomics in differentiating indolent from invasive lung adenocarcinoma in patients with pulmonary nodules. Eur Radiol 28:5121–5128
Shi Z, Zhu C, Degnan AJ et al (2018) Identification of high-risk plaque features in intracranial atherosclerosis: initial experience using a radiomic approach. Eur Radiol 28:3912–3921
Su C, Jiang J, Zhang S et al (2018) Radiomics based on multicontrast MRI can precisely differentiate among glioma subtypes and predict tumour-proliferative behaviour. Eur Radiol 29:1986–1996
Suh HB, Choi YS, Bae S et al (2018) Primary central nervous system lymphoma and atypical glioblastoma: differentiation using radiomics approach. Eur Radiol 28:3832–3839
Sun H, Chen Y, Huang Q et al (2018) Psychoradiologic utility of MR imaging for diagnosis of attention deficit hyperactivity disorder: a radiomics analysis. Radiology 287:620–630
Truhn D, Schrading S, Haarburger C, Schneider H, Merhof D, Kuhl C (2018) Radiomic versus convolutional neural networks analysis for classification of contrast-enhancing lesions at multiparametric breast MRI. Radiology 181352
Wu M, Tan H, Gao F et al (2018) Predicting the grade of hepatocellular carcinoma based on non-contrastenhanced MRI radiomics signature. Eur Radiol 29:2802–2811
Yang L, Dong D, Fang M et al (2018) Can CT-based radiomics signature predict KRAS/NRAS/BRAF mutations in colorectal cancer? Eur Radiol 28:2058–2067
Yin P, Mao N, Zhao C et al (2018) Comparison of radiomics machine-learning classifiers and feature selection for differentiation of sacral chordoma and sacral giant cell tumour based on 3D computed tomography features. Eur Radiol 29:1841–1847
Zhang S, Song G, Zang Y et al (2018) Non-invasive radiomics approach potentially predicts non-functioning pituitary adenomas subtypes before surgery. Eur Radiol 28:3692–3701
Zhang Y, Zhang B, Liang F et al (2018) Radiomics features on noncontrast-enhanced CT scan can precisely classify AVM-related hematomas from other spontaneous intraparenchymal hematoma types. Eur Radiol 29:2157–2165
Zhang Z, Yang J, Ho A et al (2018) A predictive model for distinguishing radiation necrosis from tumour progression after gamma knife radiosurgery based on radiomic features from MR images. Eur Radiol 28:2255–2263
Zhu X, Dong D, Chen Z et al (2018) Radiomic signature as a diagnostic factor for histologic subtype classification of non-small cell lung cancer. Eur Radiol 28:2772–2778
Zinn PO, Singh SK, Kotrotsou A et al (2018) A coclinical radiogenomic validation study: conserved magnetic resonance radiomic appearance of periostin-expressing glioblastoma in patients and xenograft models. Clin Cancer Res 24:6288–6299
Choe J, Lee SM, Do KH et al (2019) Prognostic value of radiomic analysis of iodine overlay maps from dual-energy computed tomography in patients with resectable lung cancer. Eur Radiol 29:915–923
Hu T, Wang S, Huang L et al (2019) A clinical-radiomics nomogram for the preoperative prediction of lung metastasis in colorectal cancer patients with indeterminate pulmonary nodules. Eur Radiol 29:439–449
Ji GW, Zhang YD, Zhang H et al (2019) Biliary tract cancer at CT: a radiomics-based model to predict lymph node metastasis and survival outcomes. Radiology 290:90–98
Kontos D, Winham SJ, Oustimov A et al (2019) Radiomic phenotypes of mammographic parenchymal complexity: toward augmenting breast density in breast cancer risk assessment. Radiology 290:41–49
Qu J, Shen C, Qin J et al (2019) The MR radiomic signature can predict preoperative lymph node metastasis in patients with esophageal cancer. Eur Radiol 29:906–914
Tan X, Ma Z, Yan L, Ye W, Liu Z, Liang C (2019) Radiomics nomogram outperforms size criteria in discriminating lymph node metastasis in resectable esophageal squamous cell carcinoma. Eur Radiol 29:392–400
Wei J, Yang G, Hao X et al (2019) A multi-sequence and habitat-based MRI radiomics signature for preoperative prediction of MGMT promoter methylation in astrocytomas with prognostic implication. Eur Radiol 29:877–888
Bonekamp D, Kohl S, Wiesenfarth M et al (2018) Radiomic machine learning for characterization of prostate lesions with MRI: comparison to ADC values. Radiology 289:128–137
Park H, Lim Y, Ko ES et al (2018) Radiomics signature on magnetic resonance imaging: association with disease-free survival in patients with invasive breast cancer. Clin Cancer Res 24:4705–4714
Sun R, Limkin EJ, Vakalopoulou M et al (2018) A radiomics approach to assess tumour-infiltrating CD8 cells and response to anti-PD-1 or anti-PD-L1 immunotherapy: an imaging biomarker, retrospective multicohort study. Lancet Oncol 19:1180–1191
Wang K, Lu X, Zhou H et al (2018) Deep learning radiomics of shear wave elastography significantly improved diagnostic performance for assessing liver fibrosis in chronic hepatitis B: a prospective multicentre study. Gut 68:729–741
Kessler LG, Barnhart HX, Buckler AJ et al (2015) The emerging science of quantitative imaging biomarkers terminology and definitions for scientific studies and regulatory submissions. Stat Methods Med Res 24:9–26
McShane LM, Altman DG, Sauerbrei W et al (2005) Reporting recommendations for tumor marker prognostic studies (REMARK). J Natl Cancer Inst 97:1180–1184
Korevaar DA, van Enst WA, Spijker R, Bossuyt PM, Hooft L (2014) Reporting quality of diagnostic accuracy studies: a systematic review and meta-analysis of investigations on adherence to STARD. Evid Based Med 19:47–54
Korevaar DA, Wang J, van Enst WA et al (2015) Reporting diagnostic accuracy studies: some improvements after 10 years of STARD. Radiology 274:781–789
Funding
This research was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIP) (grant number: NRF-2017R1A2A2A05001217 and grant number: NRF-2017R1C1B2007258).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Jeong Hoon Kim.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
One of the authors has significant statistical expertise (Seo Young Park, 8 years of experience).
Informed consent
Written informed consent was not required because of the nature of our study, which was a study based on research articles.
Ethical approval
Institutional Review Board approval was not required because of the nature of our study, which was a study based on research articles.
Methodology
• retrospective
• cross-sectional study
• performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 260 kb)
Rights and permissions
About this article
Cite this article
Park, J.E., Kim, D., Kim, H.S. et al. Quality of science and reporting of radiomics in oncologic studies: room for improvement according to radiomics quality score and TRIPOD statement. Eur Radiol 30, 523–536 (2020). https://doi.org/10.1007/s00330-019-06360-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-019-06360-z