Abstract
Objectives
Texture analysis performed on MRI images can provide additional quantitative information that is invisible to human assessment. This study aimed to evaluate the feasibility of texture analysis on preoperative conventional MRI images in predicting early malignant transformation from low- to high-grade glioma and compare its utility to histogram analysis alone.
Methods
A total of 68 patients with low-grade glioma (LGG) were included in this study, 15 of which showed malignant transformation. Patients were randomly divided into training (60%) and testing (40%) sets. Texture analyses were performed to obtain the most discriminant factor (MDF) values for both training and testing data. Receiver operating characteristic (ROC) curve analyses were performed on MDF values and 9 histogram parameters in the training data to obtain cutoff values for determining the correct rates of discrimination between two groups in the testing data.
Results
The ROC analyses on MDF values resulted in an area under the curve (AUC) of 0.90 (sensitivity 85%, specificity 84%) for T2w FLAIR, 0.92 (86%, 94%) for ADC, 0.96 (97%, 84%) for T1w, and 0.82 (78%, 75%) for T1w + Gd and correctly discriminated between the two groups in 93%, 100%, 93%, and 92% of cases in testing data, respectively. In the astrocytoma subgroup, AUCs were 0.92 (88%, 83%) for T2w FLAIR and 0.90 (92%, 74%) for T1w + Gd and correctly discriminated two groups in 100% and 92% of cases. The MDF outperformed all 9 of the histogram parameters.
Conclusion
Texture analysis on conventional preoperative MRI images can accurately predict early malignant transformation of LGGs, which may guide therapeutic planning.
Key Points
• Texture analysis performed on MRI images can provide additional quantitative information that is invisible to human assessment.
• Texture analysis based on conventional preoperative MR images can accurately predict early malignant transformation from low- to high-grade glioma.
• Texture analysis is a clinically feasible technique that may provide an alternative and effective way of determining the likelihood of early malignant transformation and help guide therapeutic decisions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Glioma can be classified as low (LGG, WHO grades I–II) or high (HGG, WHO grade III/IV) grade [1], with low-grade gliomas typically having a longer survival time. The median survival of patients with low-grade gliomas (WHO grade II) can range between 2 and 20 years, depending on the presence or absence of a 1p19q co-deletion and isocitrate dehydrogenase (IDH) mutational status [2,3,4,5]. However, even within 1p19q and IDH mutational subgroups, the prognosis is highly variable, with some growing slowly for years, while others progress rapidly to high-grade (grade III/IV) gliomas. The time of malignant transformation differs among patients, with a 5-year malignancy-free survival rate of 30 to 70% [6, 7]. Advances in MRI and PET techniques have improved grading of preoperative gliomas [8,9,10], which is important in guiding treatment and predicting prognosis, but predicting early malignant transformation of LGG still remains a challenge and is important for clinical management.
Standard-of-care neuroradiological monitoring of LGG relies on visual inspection on MRI images, which can vary based on radiologists’ training, experience, and expertise. Employing a machine-learning operator-independent algorithm, such as the basic texture analysis, may identify unique MRI features that reflect the underlying pathophysiology of LGGs that eventually undergo malignant transformation. In recent years, texture analysis has gained increasing interest in clinical studies, as it enables quantification of spatial variations in the gray-level patterns, pixel interrelationships, and spectral properties of an image [11]. Although some characteristics can be observed qualitatively, many features remain imperceptible to the human eye. Texture features also account for a large and important part of radiomics analysis, which allows for more power in facilitating better clinical decision-making in the care of patients with cancer [12]. Texture analysis has been performed on MRI images to grade brain glioma [13,14,15,16], differentiate among brain metastases [17, 18], and predict glioma phenotype and overall survival [19], with promising results. Textural parameters consist of various statistical classes, including histogram analysis, gray-level co-occurrence matrices (GLCM), and run-length matrices (RLM) [20]. Histogram analysis alone can represent a gray-level distribution without spatial information and is increasingly used in quantitative analysis of gliomas in clinical studies [21, 22]. It is considered preferable to operator-dependent ROI analysis in describing tumor heterogeneity and may guide tumor classification and assessment of progression. We hypothesize that textural parameters of various statistical classes, some of which are invisible to the human eye, can predict malignant transformation of LGG based on conventional MR images.
The purpose of this study was to evaluate the feasibility of a clinically implementable texture analysis package, based on conventional preoperative MRI images (T2w FLAIR, ADC, T1w, and post-Gadolinium T1w (T1w + Gd)), in predicting early malignant transformation from low- to high-grade glioma, and compare its utility to histogram analysis alone.
Materials and methods
Patient selection
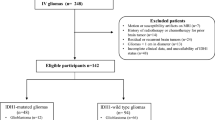
This retrospective study was approved by our local institutional review board, which waived written informed consent. Ninety-five consecutive patients who underwent MR imaging in our institution for low-grade glioma between June 2000 and December 2017 were identified. Of these patients, 68 met the following inclusion criteria: (1) pathologically proven (after resection or biopsy) grade II tumor according to the WHO Classification of Tumors of the Central Nervous System [1, 23]; (2) follow-up until pathologically proven malignant transformation to high-grade glioma or at least 2 years of clinical/radiological follow-up demonstrating tumor stability; (3) a preoperative MRI scan that included T2w FLAIR, ADC, T1w, or T1w + Gd sequences.
MRI protocol
All brain MRIs were performed on our clinical scanners (GE Signa HDxt 1.5 T and 3.0 T, GE SIGNA EXCITE 1.5 T). Parameters for T2w FLAIR are as follows: field of view = 24 cm, TR = 8802–10,000 ms, TE = 113–321 ms, TI = 1660–2300 ms, flip angle = 90–111°, slice thickness = 3–5 mm, matrix = (256–352) × (192–160), pixel bandwidth = 75–893. ADC maps were reconstructed from the DWI sequence, whose parameters are as follows: field of view = 24 cm, TR = 4500–10,500 ms, TE = 62–121 ms, flip angle = 90–180°, slice thickness = 3–5 mm, matrix = 128 × 128, pixel bandwidth = 484–1421, b-values = 0, 1000 s/mm2. Image parameters for T1w and T1w + Gd are as follows: field of view = 24 cm, TR = 367–450 ms, TE = 7–18 ms, flip angle = 90–130°, slice thickness = 3–5 mm, matrix = (256–320) × (192–224), pixel bandwidth = 61.
Texture analysis
Texture analysis was performed using open-source MaZda software (version 4.6.0, Institute of Electronics, Technical University of Lodz) [24, 25]. Training (using 60% of the data) and testing sets (40%) were randomly generated, both including astrocytomas and oligodendrogliomas that did or did not undergo early malignant transformation. MRI images were loaded into the MaZda software; then, the tumors were manually segmented on each image slice of the entire abnormal area on T2w FLAIR images by a neuroradiologist (6 years of experience) and reassessed by another senior neuroradiologist (20 years of experience). The segmented tumor was then overlaid onto other co-registered images (ADC, T1w, and T1w + Gd), which were further edited using the add/erase tool to better match the lesion geometry when necessary. Any cystic components of the tumor with similar signal intensity to CSF on T2w and T2w FLAIR images were excluded. No evidence of necrosis or hemorrhage was found in our patient cohort, as expected in grade II tumors. All the image analyses were performed slice per slice on each individual preoperative MRI image, and the outcome of each image slice was classified as the same as the patient’s outcome. The number of image slices within the tumor ranged from 2 to 14 (mean 7.7). In order to evaluate reproducibility, we randomly selected 10 cases (57 image slices) for repeat segmentation, more than 3 months after the initial analysis. The tumoral area calculated from the repeat image analysis was compared to the initial area using linear regression and Bland-Altman analysis.
Since there were a limited number of oligodendroglioma and astrocytoma studies that did not include ADC and T1w images (Table 1), subgroup texture analysis was performed only on astrocytomas with T2w FLAIR and T1w + Gd images.
A flowchart of the patient groups and texture analysis procedure is shown in Fig. S1. The average time for lesion segmentation is approximately 10 min for each case, and the MaZda software took less than 5 min to perform the texture analysis.
Training data set
All segmented tumors for each image slice on T2w FLAIR, ADC, T1w, and T1w + Gd images were loaded into the MaZda package to perform texture analysis; as many as 279 features were generated within each ROI. These texture features were derived from 6 different statistical image descriptors: histogram features, gradient features, run-length matrix (RLM), co-occurrence matrix (COM), autoregressive model (AR), and wavelet transform. A detailed description of these textural features can be found in previous literature [25]. Before texture analysis, image intensities were normalized between μ ± 3σ (μ indicates the mean value of the gray levels within the ROI; σ the standard deviation); the range obtained was quantized to 6 bits/pixel. This procedure, used by previous studies [26,27,28], reduces brightness and contrast variations and minimizes the influence of interscanner as well as field strength differences, in order to generate ideal classifications. Since analyses on all 279 texture features are clinically impractical, the MaZda software provides three feature reduction algorithms: mutual information (MI), Fisher coefficient (F), and classification error probability and average correlation coefficients (POE + ACC, PA). Each algorithm determines the 10 best distinguishable texture features resulting in a combined total of up to 30 top-ranked features for further analysis. These 30 features were then loaded into the statistical B11 texture analysis package; a linear discriminant analysis (LDA) model with the lowest misclassification rate was selected to obtain the most discriminant factor (MDF) values [29], which served as a comprehensive variable for discrimination. MDF defines which feature contributes the most significant amount of prediction to help separating the groups. The values of the 9 histogram parameters (mean, variance, skewness, kurtosis, percent 1%, percent 10%, percent 50%, percent 90%, and percent 99%) were separately saved from the previously described histogram features (one of 6 different statistical image descriptors used for texture analysis), in order to compare with texture analysis.
Receiver operating characteristic (ROC) curves were performed on the generated MDF values and 9 histogram parameters for each image slice using SPSS for Windows (version 19.0). The area under the curve (AUC) and the optimal cutoff values from the maximum Youden index, as well as the corresponding sensitivity and specificity for discriminating between LGGs that had early malignant transformation and those that did not, were obtained from ROC curve analysis. The MDF values of the two groups were compared using independent sample t test, where p < 0.05 indicates statistical significance.
Testing data set
ROIs for T2w FLAIR, ADC, T1w, and T1w + Gd images in the testing data were loaded into the MaZda software and processed in the same way as the training data in order to generate the 279 features. None of the training data were used in the analysis of the testing data. The same 30 features as in the training data were selected and loaded into the B11 analysis package; the same LDA model was used to generate the MDF values for each image slice. The MDF values and 9 histogram parameters were used to classify the testing samples into either an early malignant transformation group or without malignant transformation group based on the optimal cutoff value predefined in the training data. A correct rate of discriminating the two groups was consequently determined. The equation below, as did in a previous study [29], was used to calculate the weighted MDF values and histogram parameters on multiple image slices for each tumor. They were also tested on the predefined cutoff value to define a percent accuracy of discriminating between two groups.
where \( \overline{x} \) is the overall weighted mean value for each parameter, w1 is the area of the first ROI, x1 is the mean value of the first ROI, w2 is the area of the second ROI, x2 is the mean value of the second ROI, and so forth.
Results
Patient characteristics
Out of a total of 95 subjects, 68 LGG patients (31 males, 37 females) that met the inclusion criteria were included in this study: 15 (22%) showed malignant transformation to higher grade (8 transformed to grade III and 7 transformed to grade IV) after 1 to 13 years of follow-up (mean 4.18 years, median 3.29 years) (Fig. 1). Fifty-three of them (78%) did not show malignant transformation after 2–12 years of follow-up (mean 4.82 years, median 4.32 years). Twenty-seven patients were excluded either because of short follow-up time (less than 2 years, n = 10) or lack of available MRI images (n = 17). Patient details are listed in Table 1.
Representative low-grade glioma which underwent malignant transformation. On the preoperative images, the tumor is in the right frontal lobe and genu of the corpus callosum. The pathological diagnosis was oligodendroglioma (WHO grade II). The red color indicates the ROIs segmented on each individual image. Four months after resection, no obvious residual/recurrent tumor was found. Four years later, recurrent tumor was confirmed on biopsy to be an anaplastic oligodendroglioma (WHO grade III)
Reproducibility analysis
Tumor segmentation was repeated in 10 randomly selected cases (57 image slices); there was a strong correlation between two times area measurements (R2 = 0.982). Bland-Altman analysis revealed a small bias of 12.5 mm2 and 95% limits of agreement of [− 25.7, 50.6] mm2 (Fig. S2).
Training data set
Detailed group information for T2w FLAIR, ADC, T1w, and T1w + Gd in the training data is shown in the flowchart in Fig. S1. The MDF values generated from the LDA model when performing B11 analysis were significantly different between the two groups (all p < 0.001). ROC analyses on these MDF values resulted in an AUC of 0.90 (95% CI, 0.86–0.95) for T2w FLAIR, 0.92 (95% CI, 0.87–0.97) for ADC, 0.96 (95% CI, 0.93–0.99) for T1w, and 0.82 (95% CI, 0.77–0.87) for T1w + Gd (Figs. 2 and 3). Cutoff values of 0.01 (T2w FLAIR), − 0.015 (ADC), 0.0025 (T1w), and 0.02 (T1w + Gd) with corresponding high sensitivities and specificities (T2w FLAIR 77%, 93%; ADC 86%, 94%; T1w 97%, 84%; T1w + Gd 75%, 77%) were obtained. The MDF from texture analysis outperformed all the other histogram parameters, which had lower AUC (T2w FLAIR range 0.51–0.72, ADC 0.55–0.71, T1w 0.51–0.68, T1w + Gd 0.51–0.63), sensitivities, and specificities (Tables S1–S4).
Texture analysis results on individual T2w FLAIR and ADC image slices and tested on the testing data set. a, d Output of B11 analysis using the linear discriminant analysis (LDA) model performed on all image slices from training data. The misclassification rates of T2w FLAIR and ADC were 17% and 18%, respectively. The most discriminant factor (MDF) is shown for two groups, where the red 1 label represents early malignant transformation and the green 2 label indicates without early malignant transformation. b, e Receiver operating characteristic (ROC) curve performed on MDF and 9 histogram parameters to discriminate between the two groups, where the red solid line indicates MDF obtained from texture analysis and dotted lines represent the 9 histogram parameters. The area under the curve (AUC) of the texture-based MDF value outperformed the 9 other parameters. c, f The MDF cutoff value obtained from ROC analysis on the training data was used to test the weighed value within each tumor in the testing data set. For T2w FLAIR, 3 in 27 cases (shown in red) were misclassified when differentiating two groups using the same training set cutoff value of 0.01 (c), and for ADC, there were no misclassifications when using the cutoff value of − 0.015 (f)
Texture analysis results on individual T1w and T1w + Gd image slices and tested on the testing data set. a, d Output of B11 analysis using the linear discriminant analysis (LDA) model performed on all image slices from training data. The misclassification rates of T1w and T1w + Gd were 12% and 31%, respectively. The most discriminant factor (MDF) is shown for two groups, where the red 1 label represents early malignant transformation and the green 2 label indicates without early malignant transformation. b, e Receiver operating characteristic (ROC) curve performed on MDF and 9 histogram parameters to discriminate between two groups, where the red solid line indicates MDF obtained from texture analysis and dotted lines represent the 9 histogram parameters. The area under the curve (AUC) of texture-based MDF value outperformed the 9 other parameters. c, f The results of using MDF cutoff value obtained from ROC analysis on training data to test the weighed value within each tumor in testing data set. For T1w, 1 in 14 cases (shown in red) was misclassified using the cutoff value of 0.0025 for differentiating two groups (c), and for ADC, 2 in 26 cases were misclassified using the same cutoff value of 0.02 (f)
For the astrocytoma subgroup analysis using T2w FLAIR and T1w + Gd images, the MDF values were significantly different between the two groups (all p < 0.001). ROC analyses on these MDF values resulted in an AUC of 0.92 (95% CI, 0.88–0.97) for T2w FLAIR and 0.89 (95% CI, 0.84–0.95) for T1w + Gd (Fig. S3). Cutoff values of 0.001 (T2w FLAIR) and − 0.008 (T1w + Gd), with corresponding high sensitivities and specificities (T2w FLAIR 90%, 83%; T1w + Gd 78%, 90%), were obtained. The MDF from texture analysis outperformed all the other histogram parameters, which had lower AUC (T2w FLAIR range 0.53–0.68, T1w + Gd 0.51–0.54), sensitivities, and specificities (Tables S5 and S6).
Testing data set
The same cutoff MDF value generated in the training data set was used to define the early transformation status in the testing data. Percent accuracies of 80% (157/196) for T2w FLAIR, 85% (103/121) for ADC, 89% (89/100) for T1w, and 93% (172/185) for T1w + Gd were obtained when evaluated on each image slice. The weighted MDF value on all image slices for each tumor resulted in percent accuracies of 89% (24/27) for T2w FLAIR, 100% (15/15) for ADC, 93% (13/14) for T1w, and 93% (24/26) for T1w + Gd. The MDF values from texture analysis had a higher percent accuracy over histogram parameters when evaluated on individual image slices (T2w FLAIR range 29–68%, 57–134/196; ADC range 41–82%, 49–99/121; T1w range 49–74%, 49–74/100; T1w + Gd range 28–57%, 52–106/185) or on weighted values for each tumor (T2w FLAIR range 22–78%, 6–21/27; ADC range 33–80%, 5–12/15; T1w range 43–79%, 6–11/14; T1w + Gd range 23–69%, 6–18/26) (Tables S1–S4, Figs. 2 and 3).
For the astrocytoma subgroup analysis using T2w FLAIR and T1w + Gd images, percent accuracies of 90% (69/77) for T2w FLAIR and 96% (70/73) for T1w + Gd were obtained when evaluated on each image slice. The weighted MDF value on all image slices for each tumor resulted in percent accuracies of 100% (12/12) for T2w FLAIR and 100% (12/12) for T1w + Gd. The MDF values from texture analysis had a higher percent accuracy over histogram parameters when evaluated on individual image slices (T2w FLAIR range 44–61%, 34–47/77; T1w + Gd range 47–71%, 34–52/73) or on weighted values for each tumor (T2w FLAIR range 33–75%, 4–9/12; T1w + Gd range 50–67%, 6–8/12) (Tables S5 and S6, Fig. S3).
Discussion
Our study shows that texture analysis on conventional preoperative MRI images has high sensitivity, specificity, and accuracy in discriminating between low-grade gliomas that subsequently underwent early malignant transformation and those that did not. Texture analysis outperformed all histogram parameters and may provide additional information that could guide clinical management decisions.
Texture analysis on MRI images can provide quantitative information to help characterize tumor heterogeneity and serve as a potential prognosticator for tumor malignancy in a wide range of tumor types [18, 30, 31]. In this study, we used the MaZda open-source software [29, 32] to obtain high sensitivity, specificity, and accuracy in predicting LGGs that would undergo early malignant transformation in each conventional MRI sequence. By analyzing the top 30 selected features, the texture analysis package generated an overall discriminator MDF, which can perform better than individual features for identifying early malignant transformation in LGGs.
T2w FLAIR is the most commonly used sequence to evaluate brain pathology [33] and shows the extent of infiltrative tumor. ADC maps, obtained from DWI, demonstrate Brownian motion of water molecules and are an efficient tool in discriminating between low- and high-grade gliomas [9, 34]. T1w and T1w + Gd can provide anatomic information, with tissue enhancement reflecting an increase in blood-brain barrier permeability. But, the primary purpose of these anatomical sequences is to identify a lesion as well as its location and potential for surgical intervention. Serial imaging can identify new areas of contrast enhancement or a significant change in tumor size, which may signify transformation to a higher grade. However, this approach might cause a delay for necessary treatment changes as multiple scans must be done.
Texture features can provide quantitative information in addition to the qualitative information identified through neuroradiological assessment from conventional MR images. Of note, texture analysis based on T1w images demonstrated a relative higher AUC than the other three sequences. One explanation for this might be that T1 relaxation times have the largest range in different tissue types when compared with T2 relaxation times. Since T1w images are the best for obtaining contrast between tissue types, it was anticipated to have more informative data for extraction using texture analysis. Surprisingly, we also found that T1w + Gd, which is useful for visual evaluation of tumor, had the lowest AUC of all the MRI sequences. Overall, texture analysis provided us with useful and valuable information for predicting which LGGs would have early malignant transformation and should be closely followed and aggressively treated.
Furthermore, MaZda software can be easily implemented in the clinic without additional professional technical input, allowing it to serve as an alternative method in routine clinical surveillance of LGGs. Our texture analysis results also form the basis for further radiomics analyses, which extract innumerable quantitative features from various kinds of digital images, and are a rapidly expanding research area [35, 36]. Radiogenomics, which correlates image features with underlying genetic data such as 1p19q co-deletion and IDH mutation, has proved to be useful in providing surrogate biomarkers for predicting tumor biology and therapy response, making it an important tool in advancing personalized treatment of glioma [37, 38]. Future studies using texture analysis to differentiate astrocytoma from oligodendroglioma and correlating results with genotypic information may have clinical implications in treatment.
Histogram parameters have been investigated to evaluate tumor heterogeneity, glioma grading, and prediction of malignant transformation [21, 39]. However, in this texture analysis, we combined three levels of statistical methods; the texture analysis–based MDF outperformed each individual histogram parameter in discriminating LGGs that underwent early malignant transformation from those that did not. This suggests that texture analysis can provide a more comprehensive evaluation of clinical MR images and is better than histogram analysis alone.
There are several limitations in our study: First, the sample size of the early malignant transformation group was small due to the low incidence of LGG malignant transformation in this cohort. In addition, subgroup analysis of oligodendrogliomas and astrocytomas with available ADC and T1w images could not be performed as a result of having limited numbers of cases in the malignant transformation group. We believe that having an increased number of malignant transformation cases would enhance the accuracy of texture analysis in future studies. Second, we performed texture analysis on individual image slices of each tumor instead of a volumetric analysis, again due to small sample size. We believe that further studies based on a volumetric analysis will make texture analysis more reliable. Third, given the fact that some inherent process of texture analysis remains unknown, there may exist some unexpected outlier correlations driving the good outcome, and normalization of image intensities in order to minimize interscanner differences might be seen as a potential source of bias, so our results should be further validated using other independent data. Fourth, the diffusion model in our study is based on only two b-values, but having multiple b-values [40] for ADC maps, might increase accuracy of texture analysis. Another concern is the treatment heterogeneity: LGG patients received radiotherapy and/or chemotherapy or no additional therapy after initial resection; these treatments may influence malignant transformation. Finally, the status of 1p19q deletions and IDH mutations was not available for all tumors in our study, which is a part of the 2016 WHO tumor classification [1]. So for these, we had to use the old WHO classification [23].
In conclusion, texture analysis based on conventional preoperative MR images can accurately predict early malignant transformation of LGGs. Texture analysis is a clinically feasible technique that may provide an alternative and effective way of determining the likelihood of early malignant transformation and help guide therapeutic decisions.
Abbreviations
- AUC:
-
Area under the curve
- LDA:
-
Linear discriminant analysis
- LGG:
-
Low-grade glioma
- MDF:
-
Most discriminant factor
- ROC:
-
Receiver operating characteristic
References
Louis DN, Perry A, Reifenberger G et al (2016) The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 131:803–820
Claus EB, Walsh KM, Wiencke JK et al (2015) Survival and low-grade glioma: the emergence of genetic information. Neurosurg Focus 38:E6
Okamoto Y, Di Patre PL, Burkhard C et al (2004) Population-based study on incidence, survival rates, and genetic alterations of low-grade diffuse astrocytomas and oligodendrogliomas. Acta Neuropathol 108:49–56
Pignatti F, van den Bent M, Curran D et al (2002) Prognostic factors for survival in adult patients with cerebral low-grade glioma. J Clin Oncol 20:2076–2084
Olar A, Wani KM, Alfaro-Munoz KD et al (2015) IDH mutation status and role of WHO grade and mitotic index in overall survival in grade II-III diffuse gliomas. Acta Neuropathol 129:585–596.
Rotariu D, Gaivas S, Faiyad Z, Haba D, Iliescu B, Poeata I (2010) Malignant transformation of low grade gliomas into glioblastoma a series of 10 cases and review of the literature. Rom Neurosurg 4:403–412
Bogdańska MU, Bodnar M, Piotrowska MJ et al (2017) A mathematical model describes the malignant transformation of low grade gliomas: prognostic implications. PLoS One 12:e0179999
Jiang R, Jiang J, Zhao L et al (2015) Diffusion kurtosis imaging can efficiently assess the glioma grade and cellular proliferation. Oncotarget 6:42380–42393
Arevalo-Perez J, Peck KK, Young RJ, Holodny AI, Karimi S, Lyo JK (2015) Dynamic contrast-enhanced perfusion MRI and diffusion-weighted imaging in grading of gliomas. J Neuroimaging 25:792–798
Bulik M, Jancalek R, Vanicek J, Skoch A, Mechl M (2013) Potential of MR spectroscopy for assessment of glioma grading. Clin Neurol Neurosurg 115:146–153
Kassner A, Thornhill RE (2010) Texture analysis: a review of neurologic MR imaging applications. AJNR Am J Neuroradiol 31:809–816
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577
Xie T, Chen X, Fang J et al (2018) Textural features of dynamic contrast-enhanced MRI derived model-free and model-based parameter maps in glioma grading. J Magn Reson Imaging 47:1099–1111
Skogen K, Schulz A, Dormagen JB, Ganeshan B, Helseth E, Server A (2016) Diagnostic performance of texture analysis on MRI in grading cerebral gliomas. Eur J Radiol 85:824–829
Reza SM, Mays R, Iftekharuddin KM (2015) Multi-fractal Detrended Texture Feature for Brain Tumor Classification. Proc SPIE Int Soc Opt Eng 9414:941410.
Ryu YJ, Choi SH, Park SJ, Yun TJ, Kim JH, Sohn CH (2014) Glioma: application of whole-tumor texture analysis of diffusion-weighted imaging for the evaluation of tumor heterogeneity. PLoS One 9:e108335
Béresová M, Larroza A, Arana E, Varga J, Balkay L, Moratal D (2018) 2D and 3D texture analysis to differentiate brain metastases on MR images: proceed with caution. MAGMA 31:285–294
Li Z, Mao Y, Li H, Yu G, Wan H, Li B (2016) Differentiating brain metastases from different pathological types of lung cancers using texture analysis of T1 postcontrast MR. Magn Reson Med 76:1410–1419
Chaddad A, Tanougast C (2016) Extracted magnetic resonance texture features discriminate between phenotypes and are associated with overall survival in glioblastoma multiforme patients. Med Biol Eng Comput 54:1707–1718
Castellano G, Bonilha L, Li LM, Cendes F (2004) Texture analysis of medical images. Clin Radiol 59:1061–1069
Just N (2014) Improving tumour heterogeneity MRI assessment with histograms. Br J Cancer 111:2205–2213
Aggarwal N, Agrawal R (2012) First and second order statistics features for classification of magnetic resonance brain images. J Signal Inf Process 3:146
Louis DN, Ohgaki H, Wiestler OD et al (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114:97–109
Strzelecki M, Szczypinski P, Materka A, Klepaczko A (2013) A software tool for automatic classification and segmentation of 2D/3D medical images. Nucl Inst Methods Phys Res A 702:137–140
Szczypiński PM, Strzelecki M, Materka A, Klepaczko A (2009) MaZda--a software package for image texture analysis. Comput Methods Prog Biomed 94:66–76
Yan PF, Yan L, Hu TT et al (2017) The potential value of preoperative MRI texture and shape analysis in grading meningiomas: a preliminary investigation. Transl Oncol 10:570–577
Collewet G, Strzelecki M, Mariette F (2004) Influence of MRI acquisition protocols and image intensity normalization methods on texture classification. Magn Reson Imaging 22:81–91
Vallières M, Freeman CR, Skamene SR, El Naqa I (2015) A radiomics model from joint FDG-PET and MRI texture features for the prediction of lung metastases in soft-tissue sarcomas of the extremities. Phys Med Biol 60:5471–5496
Brown AM, Nagala S, McLean MA et al (2016) Multi-institutional validation of a novel textural analysis tool for preoperative stratification of suspected thyroid tumors on diffusion-weighted MRI. Magn Reson Med 75:1708–1716
Lisson CS, Lisson CG, Flosdorf K et al (2018) Diagnostic value of MRI-based 3D texture analysis for tissue characterisation and discrimination of low-grade chondrosarcoma from enchondroma: a pilot study. Eur Radiol 28:468–477
Fan M, Cheng H, Zhang P et al (2018) DCE-MRI texture analysis with tumor subregion partitioning for predicting Ki-67 status of estrogen receptor-positive breast cancers. J Magn Reson Imaging 48:237–247
Meyer HJ, Schob S, Hohn AK, Surov A (2017) MRI texture analysis reflects histopathology parameters in thyroid cancer - a first preliminary study. Transl Oncol 10:911–916
Ly KI, Gerstner ER (2018) The role of advanced brain tumor imaging in the care of patients with central nervous system malignancies. Curr Treat Options Oncol 19:40
Wang S, Meng M, Zhang X et al (2018) Texture analysis of diffusion weighted imaging for the evaluation of glioma heterogeneity based on different regions of interest. Oncol Lett 15:7297–7304
Hassan I, Kotrotsou A, Bakhtiari AS et al (2016) Radiomic texture analysis mapping predicts areas of true functional MRI activity. Sci Rep 6:25295
Aerts HJ (2016) The potential of radiomic-based phenotyping in precision medicine: a review. JAMA Oncol 2:1636–1642
Gevaert O, Mitchell LA, Achrol AS et al (2014) Glioblastoma multiforme: exploratory radiogenomic analysis by using quantitative image features. Radiology 273:168–174
Jamshidi N, Diehn M, Bredel M, Kuo MD (2014) Illuminating radiogenomic characteristics of glioblastoma multiforme through integration of MR imaging, messenger RNA expression, and DNA copy number variation. Radiology 270:1–2
Tofts PS, Benton CE, Weil RS et al (2007) Quantitative analysis of whole-tumor Gd enhancement histograms predicts malignant transformation in low-grade gliomas. J Magn Reson Imaging 25:208–214
McKinnon ET, Jensen JH, Glenn GR, Helpern JA (2017) Dependence on b-value of the direction-averaged diffusion-weighted imaging signal in brain. Magn Reson Imaging 36:121–127
Acknowledgements
The authors thank Kelly McCabe Gillen, PhD, for her assistance in manuscript editing.
Funding
This study has received funding by grants from the National Institutes of Health of United States (R01 NS095562, R01 NS090464) and National Natural Science Foundation of China (No. 81730049, 81801666).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Guarantor
The scientific guarantor of this publication is Shun Zhang.
Conflict of interest
The authors of this manuscript declare no relationships with any companies whose products or services may be related to the subject matter of the article.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• retrospective
• observational
• performed at one institution
Electronic supplementary material
ESM 1
(DOCX 1194 kb)
Rights and permissions
About this article
Cite this article
Zhang, S., Chiang, G.CY., Magge, R.S. et al. Texture analysis on conventional MRI images accurately predicts early malignant transformation of low-grade gliomas. Eur Radiol 29, 2751–2759 (2019). https://doi.org/10.1007/s00330-018-5921-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-018-5921-1