Abstract
GBM is a markedly heterogeneous brain tumor consisting of three main volumetric phenotypes identifiable on magnetic resonance imaging: necrosis (vN), active tumor (vAT), and edema/invasion (vE). The goal of this study is to identify the three glioblastoma multiforme (GBM) phenotypes using a texture-based gray-level co-occurrence matrix (GLCM) approach and determine whether the texture features of phenotypes are related to patient survival. MR imaging data in 40 GBM patients were analyzed. Phenotypes vN, vAT, and vE were segmented in a preprocessing step using 3D Slicer for rigid registration by T1-weighted imaging and corresponding fluid attenuation inversion recovery images. The GBM phenotypes were segmented using 3D Slicer tools. Texture features were extracted from GLCM of GBM phenotypes. Thereafter, Kruskal–Wallis test was employed to select the significant features. Robust predictive GBM features were identified and underwent numerous classifier analyses to distinguish phenotypes. Kaplan–Meier analysis was also performed to determine the relationship, if any, between phenotype texture features and survival rate. The simulation results showed that the 22 texture features were significant with p value <0.05. GBM phenotype discrimination based on texture features showed the best accuracy, sensitivity, and specificity of 79.31, 91.67, and 98.75 %, respectively. Three texture features derived from active tumor parts: difference entropy, information measure of correlation, and inverse difference were statistically significant in the prediction of survival, with log-rank p values of 0.001, 0.001, and 0.008, respectively. Among 22 features examined, three texture features have the ability to predict overall survival for GBM patients demonstrating the utility of GLCM analyses in both the diagnosis and prognosis of this patient population.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Glioblastoma multiforme (GBM) is the most common and most aggressive primary brain malignancy in adults [39]. The inability to perform complete surgical tumor resection and poor drug delivery to the brain contributes notably to the lack of effective treatment and poor prognosis. Generally, tumors such as GBM are heterogeneous with intratumoral spatial variation in both cellularity and areas of necrosis [4]. Tumors with high intratumoral heterogeneity have been shown to have a poorer prognosis likely secondary to their intrinsically aggressive biology [38]. Tumors are classified into four grades of glioma, namely grade 1 (juvenile pilocytic astrocytoma; best prognosis), grade 2 (low-grade glioma), grade 3 (anaplastic astrocytoma), and grade 4 (glioblastoma: the most aggressive type). It is mainly located in the cerebral hemispheres, and the average patient survival with GBM is around 14.6 months [30, 39].
Given the poor patient survival in GBM, computer-aided detection techniques may play a role in early, accurate diagnosis. To this end, brain tumor diagnosis using the image processing and analysis has been an area of increased research interest in recent years allowing for partial automation of GBM identification [45]. This identification is based on the tumor heterogeneity which is a pattern feature of malignancy that represents areas of high cell density or GBM phenotypes such as: active tumor and edema parts, with low cell density of necrotic parts [6, 40, 43]. These phenotypes can be segmented manually or semiautomatically by image processing tools applied to MR images. Moreover, they can specifically be identified as active tumor (vAT) and necrosis (vN) using T1-weighted imaging (T1–WI) images and edema (vE) by fluid attenuation inversion recovery (FLAIR) sequences [12]. However, not all the parts of phenotypes can be clearly identified even though this identification is clinically and prognostically useful. Therefore, it is important to assess the GBM heterogeneity by quantifying its phenotypes and this is represented by the employment of many feature types including Gaussian mixture model (GMM) features [8], histogram-based features [17], wavelet-based features [28], and texture-based features [3, 5, 7, 28].
GMM-based features have been successfully employed to classify the normal from abnormal brain heterogeneity [13]. Similarly, histogram-based features have shown that GBM has two Gaussian distributions in FLAIR sequence which are represented by vAT and vE phenotypes. Among nine statistical features for classifying vAT and vE, kurtosis and skewness have shown a highest range of 58.33−75.00 % accuracy classifier [9]. Wavelet-based features have been used to extract space–frequency textures in order to predict MGMT gene methylation status in GBM [28]. Texture features based on gray-level co-occurrence matrix (GLCM) have been employed to differentiate between pathological and healthy tissue in different organs. Thus, texture features based on the GLCM have been extracted and can evaluate the relationships of gray-level intensity in the image by second-order statistics. In this context, GLCM-based textures can compute the gray-level intensity within an image and provide additional descriptors of tumor heterogeneity [7]. These feature types can enhance standard reporting techniques and help to more accurately characterize tumor heterogeneity [15].
In addition to their diagnostic utility, imaging features have also been associated with survival in GBM. For example, contrast-enhanced MR image features provided novel prognostic information and were accurate in predicting survival times in patients with advanced gliomas [34]. Additionally, two newer studies have been considered a standardized lexicon of imaging features derived from MRI [known Visually Accessible Rembrandt Images (VASARI)] which have been demonstrated to be feasible predictors of survival [21, 29]. New texture analysis of GBM phenotypes with texture-based survival prediction could be useful in clinical practice. Recently, it showed that the texture feature ratios from contrast-enhancing, non-enhancing lesions and kinetic texture analysis obtained from perfusion parametric maps provide useful information for predicting survival in patients with GBM [27].
This paper focuses on GBM phenotype analysis using texture feature extraction based on GLCM from MR images. This approach further show the effect of radiomics analysis for GBM tumors using descriptors (features) that may be subsequently employed in automated glioma diagnosis with phenotype characterization, and demonstrate the feasibility of texture features to be associated with survival in GBM. They may also provide a more accurate assessment of the patient prognosis and underlying genomic composition.
2 Methods
2.1 Patients population and data acquisition
In this study, data were collected from The Cancer Imaging Archive (TCIA) (http://cancerimagingarchive.net/), and 40 GBM patients were used to validate the proposed method. The GBM data were acquired prior to any treatment from patients with brain tumors that were subsequently diagnosed as GBM. The GBM diagnosis was based on histological examination. These patients were visually assessed as having sufficient quality and as containing the phenotypes [necrosis parts (vN), contrast enhancement/active tumor (vAT), and edema/invasion (vE)]. For visual assessment, 3D Slicer software was used for illustrating the GBM phenotypes and testing that the patient images can be correctly registered using T1–WI and FLAIR sequence. Table 1 shows the characteristics of GBM patients by representing the average, median, minimum, and maximum of the phenotypes (vN, vAT, and vE), age, and overall survival.
The imaging protocol used whole-brain T1–WI and FLAIR scanning using a 3T MRI scanner (GE Healthcare). T1–WI scans were acquired based on the following parameters: slice thickness (ST) = 5 mm, spatial resolution (SR) = 1.04 mm, pixel spacing (PS) = 0.78 mm, repetition time (TR) = 650 ms, echo time (TE) = 9 ms, and flip angle (FA) = 90°. And, FLAIR scans were acquired using the following parameters: ST = 5 mm, SR = 1.24 mm, PS = 0.78 mm, TR = 10,002 ms, TE = 147 ms, FA = 90°, and acquisition time 10:24 min.
2.2 Data preprocessing
The entire sequences available for the patient MRI set were obtained, yet only post-contrast T1–WI and FLAIR were used for texture analyses. Moreover, from the available database 40 patients’ data were randomly chosen to obtain full GBM tumor imaging MRI sets. All of the images had 512 × 512 pixel acquisition matrices and were converted into grayscale before further processing. Note that the standard imaging parameters were used for each of the sequences as noted in the TCIA database. Standardization was employed by the linear normalization of each image. MRI raw data were filtered by an average filter (spatial filter) with size window of 3 × 3 before further processing to minimize the effects of noise in images and other external factors. Thereafter, registration, segmentation, and texture feature extraction were employed on GBM phenotypes for data collection.
2.3 Registration and segmentation based on 3D Slicer
Registration obtains more information from different scan angles by using different slices and voxel thickness. It registers two corresponding scans or images and rigidly aligns them to each other to make an accurate registration. This operation affects the computation time which is a necessary obligation as we finish the registration step. The computation time changes depending on the image size and angle rotations. It can be more sophisticated with the three-dimensional data (3D), and the percentage accuracy may be decreased [18, 42]. Eventually, transformation of locations in one image to new locations in another image requires regulator parameters. The step of determining the correct transformation parameters is required during the image registration process. Since the FLAIR scans and T1–WI post-contrast scans obtained with different slice parameters, angles, and voxel thickness, we rigidly aligned and registered the scans to each other. Moreover, most of the voxel size of the FLAIR and T1–WI images was similar and was simply registered. However, in case that the voxel size was dissimilar, we resampled the FLAIR volume to the matrix of T1–WI voxel size. The patient’s images which have complex rotation modifications and registrations were not considered in order to achieve an error less than 2 mm. The average of computational time necessary to complete each volume registration is 40–50 s.
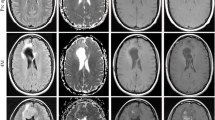
Registration for each patient’s data was done by using T1–WI and its corresponding FLAIR sequence using 3D Slicer (Fig. 1) [1]. Phenotypes were segmented based on T1–WI, FLAIR, and its corresponding registration image by using the 3D Slicer (example in Fig. 1d). Histogram analysis clearly shows the difference in shape of phenotype data fitting. For example, distribution shape in the pixel intensity axis for each phenotype was not similar (Fig. 1). Moreover, phenotype was segmented manually slice-by-slice and organized in order to extract the texture features. Then, the texture features of necrosis (vN) and active tumor parts (vAT) were computed from T1–WI, while texture feature of edema/invasion (vE) was extracted from FLAIR sequence. Texture features extracted for each phenotype were employed on each axial slice using the GLCM technique.
Preprocessing of texturized MRI image from GBM patients. a T1–WI axial image, b corresponding axial FLAIR image, c corresponding axial registration, and d 2D segmentation (label map) of GBM phenotypes. Phenotypes segmentation: illustration of 3D segmentation of phenotype vN, vAT, and vE. And, two-dimensional histogram distribution of normalized phenotypes: associated pixel histogram distribution of vN, vAT, and vE
2.4 Gray-level co-occurrence matrix computation
Texture feature extraction based on GLCM is a second-order texture that is based on the joint co-occurrence of gray values for pairs of pixels at a given distance d and direction θ [10, 23]. Traditionally, the co-occurrence matrix P d,θ (i, j) of a given two-dimensional (2D) image I of size N × N can be defined as
where dx and dy specify the distance between the pixel of interest and its neighbor, along the x and y axis of an image, respectively. Note that the GLCM is a square matrix of size N g × N g, where N g represents the number of gray levels of the image.
For 2D images, typical values used for “d” are {1, 2, 3, 4} and those for “θ” equal {0o, 45°, 90°, 135°}. The GLCMs corresponding to the additional directions {180°, 225°, 270°, 315°} added to the specification of the texture associated with combinations of the aforementioned four offsets and four directions. These additional directions were considered to cover the whole spatial relationship of offset d. In fact, 4 offsets with 4 symmetric directions may provide 16 GLCMs.
For each phenotype slice, 16 GLCMs are quantified by 22 descriptors with each slice being represented by a feature vector of 22 features. Each of these feature values is the average of 16 descriptor values.
For whole GBM tumor, a feature vector of 22 features represents the average of each descriptor in whole slice in each GBM patient. Note that these descriptors (features) have been proposed by Haralick [23] and an additional feature type based on GLCM originally described for sea ice texture analysis [37]. Table 2 shows the texture feature names and explains the mathematical formula representing the texture descriptors. All the 22 texture functions are analyzed and discussed in the result section.
2.5 Processing and analysis of features
The Kruskal–Wallis test was applied to assess the statistical significance between texture features and GBM phenotypes [31]. For this statistical test, a p value < 0.05 was considered to be significant. Thus, z score normalization was employed on each of the feature vectors which converted to zero mean and unit variance. Note that the mean and standard deviation (σ) of the feature vector were calculated as follows:
where r is the original value, r n is the new value, and the mean and σ are the average and standard deviation of the original data, respectively.
2.6 Classifier setting
We implemented four classifier techniques, namely discriminant analysis (DA) [20], naïve Bayes (NB) [2], decision trees (DT) [35], and support vector machine (SVM) [24] using the Statistics and Machine Learning Toolbox in MATLAB software. The implementation of DA was performed using pseudo-inverse which is equivalent to approximating the solution using a least-squares solution method. The implementation of NB considered a kernel estimation method which approximated the complex distributions of data.
For training based on DT classifier, two conceptual phases is considered: a “growing” phase where training examples are recursively split based on their attributes and a “pruning” phase which simplifies the tree by removing low discriminating branches of the tree. One of the most important components of decision trees is the split criterion, which selects for each node of the tree an attribute to separate examples along the branches of this node. The choice for best attribute splitting can be based on several techniques. For this work, we considered the Gini index (I G(t)) for splitting data and identify the feature subset leading to the highest accuracy. I G(t) is an impurity-based criterion that measures the divergence between the probability distributions of the attribute’s values
where p i is the relative frequency of class i at node t and node t represents any node at which a given split is performed. p i is determined by dividing the total number of observations in the class by the total number of observations.
SVM function (c) uses a model to identify support vectors (s i ), weighted (α i ), and bias (b) that are used to classify vectors (x) according to the following equation
where K is a kernel function (radial basis function).
Note that the SVM is a binary classifier which can be extended by fusing several of its kind into a multiclass classifier [16]. For 3 classes as we have the phenotypes (vN, vAT, and vE), 3 classifiers are necessary: one SVM classifies vN from vAT and vE, a second SVM classifies vAT from vN and vE, and a third SVM classifies vE from vN and vAT. The decision then is a combination of targets of all the separate SVMs. For example, vectors from classes vN, vAT, and vE have codes (1, −1, −1), (−1, 1, −1), and (−1, −1, 1), respectively. If each of the separate SVMs classifies a phenotype correctly, it means that no error for this phenotype classifier. However, if at least one of the SVMs misclassifies the phenotype, the class selected for this phenotype is the one its target code closest in the Hamming distance sense to the actual output code and this may be an erroneous decision. The reason to use these specific classifier methods is to achieve the trade-off performance metrics.
The low number of patient samples was a limitation to prove the technical method and the medical analysis. In this case, leave-one-out cross-validation can be a solution to evaluate the performance metric by the swapping test and training sample data [36]. In this way, all of the GBM phenotypes were used for both training and testing. Thus, we computed the classifier accuracy, sensitivity, and specificity to evaluate the classifier feature. In addition, receiver operating characteristic (ROC) curve was employed which provides the true positive versus (vs) false positive rates. ROC curves are widely employed to evaluate the performance of a medical test or model and are associated with area under the curve (AUC). An AUC value close to 1 shows better classification performance [22, 33].
Moreover, survival curves were plotted by Kaplan–Meier method and compared by the log-rank test [25]. We considered the median of each feature to grouping the patients in two groups based on cutoff with the threshold being the median feature value. For survival analysis, we considered p value <0.01 to be statistically significant.
3 Results
Proposed approaches were implemented in 3D Slicer [1] and MATLAB 2013b (version 8.2 [44]) and performed on GBM brain tumor MR images. The GBM images were registered, and phenotypes were segmented and identified by our considered techniques.
3.1 GLCM-based texture features
The 22 texture features exhibited p values <0.05 which formed the basis for their use in further analysis (Table 2). Each of the 22 texture functions provided a unique value for each GBM phenotype. Texture functions f 17 , f 18 , f 19, and f 20 show high values among the 22 features, and across the three GBM phenotypes, with maximum values of texture function seen for f 19. Note that the values for each feature take into consideration the average of the values derived from different phases and offsets for that particular feature and also the average derived from multiple samples. This correlation of texture feature values with the three GBM heterogeneity phenotypes allows these phenotypes to be distinguished using their distinct texture features.
3.2 Classification
The Kruskal–Wallis test showed that whole 22 texture feature extracted from GLCMs were significant with p value <0.05 (Table 2). Performance metrics of the texture feature classifier showed a maximum accuracy, sensitivity, and specificity of 79.31, 91.67, and 98.75 %, respectively, using DA technique, while the classifier accuracy value was decreased with SVM (77.59 %), NB (75.00 %), and DT (70.69 %), respectively (Table 3).
Moreover, from 36 vN, 40 vAT, and 40 vE phenotypes, confusion matrix showed that 33 vN and 32 vE were correctly classified using DA, while the highest vAT samples (29 vAT) were correctly classified using SVM classifier (Table 4).
The texture feature classifier of phenotypes was then evaluated based on ROC which showed that the AUC value based on DT classifier is better than the DA, SVM, and NB classifier (Fig. 2). AUC of (vN vs. vE) is greater than (vN vs. vAT) and (vAT vs. vE) of 99.23, 97.63, and 94.12 %, respectively (Fig. 2b). We choose the AUC of (vN vs. vE), (vN vs. vAT), and (vAT vs. vE) because the ROC must be operated between two classes.
3.3 Survival analysis
Kaplan–Meier curves were significantly different for three texture features: difference entropy f 18 (p value = 0.001), information measure of correlation f 20 (p value = 0.001), and inverse difference f 21 (p value = 0.008) (Fig. 3; Table 5). Kaplan–Meier analysis confirmed that three features based on active tumor phenotype were predictors of overall survival. For edema and necrosis parts, the texture features were not statistically significant for predicting overall survival with considered the p value <0.01. We observed that the most texture feature statistically significant to predict the survival time of GBM patient was in the active tumor parts (contrast enhancement, vAT), and this is confirmed if we considered the p value = 0.01. In these terms, we can also consider the features (f 1, f 2, f 3, f 4, f 17, f 19, and f 22) to predict the survival time of GBM patient (Table 5).
Considered p value <0.01, median survival time based on texture feature is 273.5 days (f 18 ≤ 0.36; f 20 ≤ 0.37; f 21 ≤ 0.35), 443 days (f 18 > 0.36; f 20 > 0.37), and 325 days (f 21 > 0.35). These texture features were statistically significant based on active tumor phenotype, while the features of edema and necrosis parts were not significant using Kaplan–Meier and log-rank test (Fig. 3; Table 5).
4 Discussion
The goal of this study was to provide high discrimination accuracy of GBM phenotypes using texture feature extraction from GLCM computation and find the texture features that predict the overall survival of GBM patients. The results of our study provide further evidence that texture image features that describe tumor spatial variations are useful in describing the GBM phenotypes and in predicting survival.
Data fitting showed that the histogram shape was distinct in data distribution of GBM phenotypes (Fig. 1). Moreover, the average, median, minimum, and maximum function of GBM phenotypes showed maximum, medium, and minimum values in vE, vAT, and vN, respectively (Table 1). Similarly, texture features relating to GBM phenotypes showed maximum, medium, and minimum values in vE, vAT, and vN, respectively (Table 2). This suggests that each GBM phenotype has higher values of texture vE than vAT and vN. Test of significance using texture feature to discriminate between GBM phenotypes was obtained using Kruskal–Wallis test. Furthermore, we observed that the whole GLCM-based feature set showed high significance (p value <0.05) which is represented by 22 texture features (Table 2). This suggests that spatial textures based on the GLCM provide significant features to distinguish between phenotypes.
For classification of the GBM phenotypes, we observed that the trade-off between performance and accuracy was achieved using DA classifier. The misclassification rate is resulted from some texture features that have common characteristics between GBM phenotypes which is represented by an AUC value between vAT versus vE in four classifier techniques (Fig. 2). This is further demonstrated by higher coefficient of correlation values between phenotype features. The heat map of the phenotype feature correlation showed that edema parts have some correlation with necrosis and active tumor features (Fig. 4). Specifically, middle correlation coefficient range of 0.4–0.6 presented certain texture features between vAT and vE as illustrated in Fig. 4. The middle range of correlation coefficient values demonstrated a middle similarity between vAT and vE texture features.
Heat map of correlation coefficients between texture features of GBM phenotypes. vN, vAT, and vE are the necrosis, active tumor, and edema parts, respectively. Black rectangle is the part of correlation coefficient between (vE) and (vAT and vN). Correlation coefficient value >0.6, <0.2–0.6>, and <0.2 represents high, middle, and low correlation, respectively
In reality, overlap between GBM phenotypes can be represented by the misclassification (≈20 %), and thus, it is not surprising that a certain texture feature shares a similar value. Note that the link between the texture feature and raw data is not directly correlated. For instance, a raw image with regular texture generates an irregular GLCM matrix which provides low value of the entropy (f 9). Looking at the entropy in GBM phenotypes, vE has a maximum value followed by vAT and vN. It can be conditional that the edema phenotype is more irregular texture than vAT and vN, respectively (Table 2). This technique was successfully employed in abnormal colon cell discrimination [11], and the prediction of the clinical and pathological response to neoadjuvant chemotherapy (NAC) in patients with locally advanced breast cancer before NAC is started [41].
Additionally, survival analysis of GBM patients based on texture feature derived from GLCM demonstrated that the three features (difference entropy, information measure of correlation, and inverse difference) of vAT phenotype were statistically significant with median overall survival range of 273.5–443 days (Fig. 3). We observed that the texture in vAT phenotype may predict overall survival of GBM patients. Recently, a new study based on spatial habitat features has shown 14 spatial features associated with molecular subtype and 12-month survival GBM [26].
Our approach can be a potentially valuable tool in estimating characteristics invisible to the radiologist on inspection. To our knowledge, this is the first study looking into the discrimination of GBM phenotypes using texture feature extraction based on GLCM and to show the ability of the technique to predict overall survival of GBM patients based on their phenotype features. Many studies, however, have been carried out to discriminate between tumor types. For example, texture analysis proved to differentiate benign from malignant based on T1–WI and similarly to discriminate pleomorphic adenomas and warthin tumors [19]. Also, texture analysis has been utilized effectively in the characterization of posterior fossa tumors of children [32]. Moreover, a recent study using MRI showed that texture features derived from GLCM were significant features in differentiating true progression from pseudo-progression in GBM [14].
The influence of increased diagnostic accuracy is still limited to establish clinical use. However, one potential advantage of the texture analysis is that the methods applied and their results are not limited to simple GBM phenotype discrimination. Our results, in addition to considering the optimal method of GBM phenotype discrimination, are also promising in their ability to characterize GBM heterogeneity, while the selection of texture features helps to accurately predict overall survival of GBM patients.
5 Conclusions
In this study, a novel approach based on texture feature extraction has been presented for GBM phenotype discrimination and predicting of overall patient survival. We have demonstrated the potential of texture features extracted from GLCM to characterize GBM phenotypes. The most important finding of this work is that the whole 22 texture feature set has been found to significantly classify GBM phenotypes, and three features were statistically significant for predicting overall survival. This study provides preliminary information of GBM phenotypes characterization and survival analysis of GBM patients. We note that a larger prospective trial is needed to fully evaluate the performance metrics of the proposed approach. Considering all the texture types extracted from GBM phenotypes that are under investigation, we posit that we are on the verge of a watershed moment in the identification and prediction of each phenotype by their texture features.
References
“3D Slicer.” http://www.slicer.org/. Accessed 20 Oct 2014
Aggarwal CC (2014) Data classification: algorithms and applications. CRC Press, Boca Raton
Al-Kadi OS, Watson D (2008) Texture analysis of aggressive and nonaggressive lung tumor CE CT images. IEEE Trans Biomed Eng 55(7):1822–1830
Bonavia R, Inda M-M, Cavenee WK, Furnari FB (2011) Heterogeneity maintenance in glioblastoma: a social network. Cancer Res 71(12):4055–4060
Brown RA, Frayne R (2008) A comparison of texture quantification techniques based on the Fourier and S transforms. Med Phys 35(11):4998–5008
Cancer Genome Atlas Research Network (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455(7216):1061–1068
Castellano G, Bonilha L, Li LM, Cendes F (2004) Texture analysis of medical images. Clin Radiol 59(12):1061–1069
Chaddad A (2015) Automated feature extraction in brain tumor by magnetic resonance imaging using gaussian mixture models. Int J Biomed Imaging 2015:e868031
Chaddad A, Tanougast C (2015) High-throughput quantification of phenotype heterogeneity using statistical features. Adv Bioinform 2015:e728164
Chaddad A, Tanougast C, Dandache A, Bouridane A (2011) Extracted haralick’s texture features and morphological parameters from segmented multispectrale texture bio-images for classification of colon cancer cells. WSEAS Trans Biol Biomed 8(2):39–50
Chaddad A, Tanougast C, Dandache A et al (2011) Improving of colon cancer cells detection based on Haralick’s features on segmented histopathological images. In: 2011 IEEE international conference on computer applications and industrial electronics (ICCAIE), pp 87–90
Chaddad A, Zinn PO, Colen RR (2014) Quantitative texture analysis for Glioblastoma phenotypes discrimination. In: 2014 International conference on control, decision and information technologies (CoDIT), pp 605–608
Chaddad A, Zinn PO, Colen RR (2014) Brain tumor identification using Gaussian mixture model features and decision trees classifier. In: 2014 48th annual conference on information sciences and systems (CISS), pp 1–4
Chen X, Wei X, Zhang Z, Yang R, Zhu Y, Jiang X (2015) Differentiation of true-progression from pseudoprogression in glioblastoma treated with radiation therapy and concomitant temozolomide by GLCM texture analysis of conventional MRI. Clin Imaging 39(5):775–780
Davnall F, Yip CSP, Ljungqvist G, Selmi M, Ng F, Sanghera B, Ganeshan B, Miles KA, Cook GJ, Goh V (2012) Assessment of tumor heterogeneity: an emerging imaging tool for clinical practice? Insights Imaging 3(6):573–589
Dietterich TG, Bakiri G (1995) Solving multiclass learning problems via error-correcting output codes. J Artif Int Res 2(1):263–286
Downey K, Riches SF, Morgan VA, Giles SL, Attygalle AD, Ind TE, Barton DPJ, Shepherd JH, deSouza NM (2013) Relationship between imaging biomarkers of stage I cervical cancer and poor-prognosis histologic features: quantitative histogram analysis of diffusion-weighted MR images. AJR Am J Roentgenol 200(2):314–320
Fright WR, Linney AD (1993) Registration of 3-D head surfaces using multiple landmarks. IEEE Trans Med Imaging 12(3):515–520
Fruehwald-Pallamar J, Czerny C, Holzer-Fruehwald L, Nemec SF, Mueller-Mang C, Weber M, Mayerhoefer ME (2013) Texture-based and diffusion-weighted discrimination of parotid gland lesions on MR images at 3.0 Tesla. NMR Biomed 26(11):1372–1379
Guo Y, Hastie T, Tibshirani R (2007) Regularized linear discriminant analysis and its application in microarrays. Biostatistics 8(1):86–100
Gutman DA, Cooper LAD, Hwang SN, Holder CA, Gao J, Aurora TD, Dunn WD, Scarpace L, Mikkelsen T, Jain R, Wintermark M, Jilwan M, Raghavan P, Huang E, Clifford RJ, Mongkolwat P, Kleper V, Freymann J, Kirby J, Zinn PO, Moreno CS, Jaffe C, Colen R, Rubin DL, Saltz J, Flanders A, Brat DJ (2013) MR imaging predictors of molecular profile and survival: multi-institutional study of the TCGA glioblastoma data set. Radiology 267(2):560–569
Hand DJ, Till RJ (2001) A simple generalisation of the area under the ROC curve for multiple class classification problems. Mach Learn 45(2):171–186
Haralick RM, Shanmugam K, Dinstein I (1973) Textural features for image classification. IEEE Trans Syst Man Cybern SMC-3(6):610–621
Hearst MA, Dumais ST, Osman E, Platt J, Scholkopf B (1998) Support vector machines. IEEE Intel Syst Their Appl 13(4):18–28
Kleinbaum DG, Klein M (2012) Kaplan–Meier survival curves and the log-rank test. In: Survival analysis, 3rd edn. Springer, New York, pp 55–96
Lee J, Narang S, Martinez J, Rao G, Rao A (2015) Spatial habitat features derived from multiparametric magnetic resonance imaging data are associated with molecular subtype and 12-month survival status in glioblastoma multiforme. PLoS ONE 10(9):1–13
Lee J, Jain R, Khalil K, Griffith B, Bosca R, Rao G, Rao A (2016) Texture feature ratios from relative CBV maps of perfusion MRI are associated with patient survival in glioblastoma. Am J Neuroradiol 37(1):37–43
Levner I, Drabycz S, Roldan G, De Robles P, Cairncross JG, Mitchell R (2009) Predicting MGMT methylation status of glioblastomas from MRI texture. Medical image computing and computer-assisted intervention on MICCAI international conference of medical image computing and computer-assisted intervention, vol 12 (Pt 2), pp 522–530
Mazurowski MA, Desjardins A, Malof JM (2013) Imaging descriptors improve the predictive power of survival models for glioblastoma patients. Neuro Oncol 15(10):1389–1394
McCarthy BJ, Kruchko C, Dolecek TA (2013) The impact of the Benign Brain Tumor Cancer Registries Amendment Act (Public Law 107-260) on non-malignant brain and central nervous system tumor incidence trends. J Regist Manag 40(1):32–35
McKight PE, Najab J (2010) Kruskal–Wallis Test. In: The Corsini encyclopedia of psychology. Wiley, New York
Orphanidou-Vlachou E, Vlachos N, Davies NP, Arvanitis TN, Grundy RG, Peet AC (2014) Texture analysis of T1- and T2-weighted MR images and use of probabilistic neural network to discriminate posterior fossa tumours in children. NMR Biomed 27(6):632–639
Park SH, Goo JM, Jo C-H (2004) Receiver operating characteristic (ROC) Curve: practical review for radiologists. Korean J Radiol 5(1):11–18
Pope WB, Sayre J, Perlina A, Villablanca JP, Mischel PS, Cloughesy TF (2005) MR imaging correlates of survival in patients with high-grade gliomas. AJNR Am J Neuroradiol 26(10):2466–2474
Rokach L (2007) Data mining with decision trees: theory and applications. World Scientific, Singapore
Shao J (1993) Linear model selection by Cross-validation. J Am Stat Assoc 88(422):486–494
Soh L-K, Tsatsoulis C (1999) Texture analysis of SAR sea ice imagery using gray level co-occurrence matrices. IEEE Trans Geosci Remote Sens 37(2):780–795
Sottoriva A, Spiteri I, Piccirillo SGM, Touloumis A, Collins VP, Marioni JC, Curtis C, Watts C, Tavaré S (2013) Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc Natl Acad Sci USA 110(10):4009–4014
Stupp R, Hegi ME, van den Bent MJ, Mason WP, Weller M, Mirimanoff RO, Cairncross JG, European Organisation for Research and Treatment of Cancer Brain Tumor and Radiotherapy Groups, National Cancer Institute of Canada Clinical Trials Group (2006) Changing paradigms–an update on the multidisciplinary management of malignant glioma. Oncologist 11(2):165–180
Stupp R, Hegi ME, van den Bent MJ, Mason WP, Weller M, Mirimanoff RO, Cairncross JG (2006) Changing paradigms—an update on the multidisciplinary management of malignant glioma. Oncologist 11(2):165–180
Teruel JR, Heldahl MG, Goa PE, Pickles M, Lundgren S, Bathen TF, Gibbs P (2014) Dynamic contrast-enhanced MRI texture analysis for pretreatment prediction of clinical and pathological response to neoadjuvant chemotherapy in patients with locally advanced breast cancer. NMR Biomed 27(8):887–896
Turkington TG, Hoffman JM, Jaszczak RJ, MacFall JR, Harris CC, Kilts CD, Pelizzari CA, Coleman RE (1995) Accuracy of surface fit registration for PET and MR brain images using full and incomplete brain surfaces. J Comput Assist Tomogr 19(1):117–124
Verhaak RGW, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O’Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17(1):98–110
Wallisch P, Lusignan ME, Benayoun MD, Baker TI, Dickey AS, Hatsopoulos NG (2014) MATLAB for neuroscientists: an introduction to scientific computing in MATLAB. Academic Press, Cambridge
Zacharaki EI, Wang S, Chawla S, Soo Yoo D, Wolf R, Melhem ER, Davatzikos C (2009) Classification of brain tumor type and grade using MRI texture and shape in a machine learning scheme. Magn Reson Med 62(6):1609–1618
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this article.
Ethical statement
The materials are in compliance with all applicable laws, regulations, and policies for the protection of medical data, and any necessary approvals, authorizations, and informed consent documents were obtained.
Rights and permissions
About this article
Cite this article
Chaddad, A., Tanougast, C. Extracted magnetic resonance texture features discriminate between phenotypes and are associated with overall survival in glioblastoma multiforme patients. Med Biol Eng Comput 54, 1707–1718 (2016). https://doi.org/10.1007/s11517-016-1461-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-016-1461-5