Abstract
Key message
Six foxtail millet ASR genes were regulated by various stress-related signals. Overexpression of ASR1 increased drought and oxidative tolerance by controlling ROS homeostasis and regulating oxidation-related genes in tobacco plants.
Abstract
Abscisic acid stress ripening (ASR) proteins with ABA/WDS domains constituted a class of plant-specific transcription factors, playing important roles in plant development, growth and abiotic stress responses. However, only a few ASRs genes have been characterized in crop plants and none was reported so far in foxtail millet (Setaria italic), an important drought-tolerant crop and model bioenergy grain crop. In the present study, we identified six foxtail millet ASR genes. Gene structure, protein alignments and phylogenetic relationships were analyzed. Transcript expression patterns of ASR genes revealed that ASRs might play important roles in stress-related signaling and abiotic stress responses in diverse tissues in foxtail millet. Subcellular localization assays showed that SiASR1 localized in the nucleus. Overexpression of SiASR1 in tobacco remarkably increased tolerance to drought and oxidative stresses, as determined through developmental and physiological analyses of germination rate, root growth, survival rate, relative water content, ion leakage, chlorophyll content and antioxidant enzyme activities. Furthermore, expression of SiASR1 modulated the transcript levels of oxidation-related genes, including NtSOD, NtAPX, NtCAT, NtRbohA and NtRbohB, under drought and oxidative stress conditions. These results provide a foundation for evolutionary and functional characterization of the ASR gene family in foxtail millet.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Adaptation to drought stress in plants is coupled with multifaceted strategies, including morphological, physiological and biochemical changes (Shinozaki and Yamaguchi-Shinozaki 2007). It was proposed that drought stress could provoke complex and tailored signaling transduction to a regulatory network that perceived and responded to adverse conditions by modulating the expressions of downstream effectors. It was predicted that modulation of signaling regulators would be a promising method for improving stress tolerance of plants. Transcription factors were implicated to be essential and critical points in regulating the expression of downstream genes (Golldack et al. 2011).
Abscisic acid stress ripening (ASR) proteins with abscisic acid (ABA)/water deficit stress (WDS) domains constitute a class of plant-specific transcription factors, that play important roles in biotic and abiotic stress (González and Iusem 2014). They occur in species ranging from gymnosperms to angiosperms and from monocots to dicots, but no homologous gene were found in the Arabidopsis genome (Carrari et al. 2004). It seemed that ASR genes were also missing from the Brassicaceae family (Shkolnik and Bar-Zvi 2008). ASR proteins are encoded by a small gene family represented by five genes in tomato, four genes in loblolly pine and banana, six genes in rice (González and Iusem 2014) and nine genes in maize (Virlouvet et al. 2011).
The ASR gene family functionally participates in various abiotic stress signaling pathways (Iusem et al. 1993; Konrad and Bar-Zvi 2008; Hsu et al. 2011). Overexpression of tomato SlASR1 decreased rates of water loss during drought stress in tobacco (Ricardi et al. 2014). Overexpressing lily LLA23 or banana MpASR conferred drought tolerance in Arabidopsis (Yang et al. 2008; Liu et al. 2010). Maize ZmASR1 increased crop productivity under field drought tests based on the expression of primary and/or cellular metabolic genes in the vegetative stages (Virlouvet et al. 2011). The extreme halophyte Suaeda liaotungensis K. gene SbASR1 improved tolerance to salt, drought and freezing stresses in transgenic Arabidopsis (Hu et al. 2014). Overexpression of rice OsASR1 increased cold tolerance through reactive photosynthetic rate in rice (Kim et al. 2009). Recently, it was reported that rice OsASR5 is involved in response to aluminum (Al) ion stress (Arenhart et al. 2014). Soybean ASR protein exhibited antioxidant activity in vitro, but understanding the mechanistic understanding is still in its infancy (Li et al. 2013). Nevertheless, a collective understanding of the ASR family in foxtail millet (Setaria italica), an elite drought-tolerant crop (Lata et al. 2013), has not been established. This necessitated further work to explore their functions in this species.
The release of foxtail millet genome information shed light on the acquisition of ASR family members on a genome-wide scale (Zhang et al. 2012). In this study, six putative ASR genes were identified in the foxtail millet genome. Their sequence phylogeny, genomic structure, chromosomal location and promoter elements were analyzed by bioinformatics methods. Temporal and spatial expression profiles of ASRs in different tissues and in response to different hormone and stress treatments were investigated through quantitative real-time PCR (qRT-PCR). We found that SiASR1 could positively regulate drought and oxidative stress in transgenic tobacco plants.
Materials and methods
Identification of foxtail millet ASR genes
To obtain all the ASRs in foxtail millet, we used rice, maize, Brachypodium and sorghum ASRs as queries to identify homologous peptides from foxtail millet by BLASTP searches (for homologous peptides listed in Supplemental Table S1). The protein sequences were downloaded from the PHYTOZOME v10.1 database (http://www.phytozome.org/). The keywords ‘‘ASR’’, ‘‘Abscisic acid stress ripening protein’’ and the HMM profiles of the ASR domain PF02496 were also utilized for identification of foxtail millet ASR and ASR-like sequences. Redundant sequences were removed via the decrease redundancy tool (http://web.expasy.org/decrease_redundancy/). Structural analysis of conserved regions of each non-redundant sequence was executed by SMART (http://smart.embl-heidelberg.de/) (Letunic et al. 2012), Pfam (http://pfam.sanger.ac.uk/) and conserved domain database (CDD) (Marchler-Bauer et al. 2005) searches.
Sequence alignment, phylogenetic, promoter and gene structure analyses
Multiple alignments of ASR protein sequences were carried out with ClustalX2.0. Phylogenetic trees were constructed by the neighbor-joining (NJ) method using MEGA5.10 software (Tamura et al. 2011) and the bootstrap tests were performed with 1000 replications. Information on the number and composition of stress-mediated regulatory elements in SiASR promoters were obtained from PLACE (http://www.dna.affrc.go.jp/PLACE/). Structures of SiASRs were analyzed by the GSDS tool (gsds.cbi.pku.edu.cn/).
Homology modeling of ASR protein
To identify the best template having a similar sequence and known three-dimensional structure, BLASTP searches were made against the Protein Data Bank (PDB) (http://www.rcsb.org/pdb/). Structures of ASR proteins were modeled by homology under the “intensive” mode at 90 % confidence, using the Phyre2 Server (http://www.sbg.bio.ic.ac.uk/phyre2). Subcellular localizations of ASRs were predicted by YLoc (Briesemeister et al. 2010).
Plant material, stress treatment and qRT-PCR analysis
Seeds of foxtail millet cultivar Yugu 1 were imbibed for 2 days and planted in peat/vermiculite mixture (1:1 v/v) (28 °C day/20 °C night, 16 h photoperiod, 65 % relative humidity). Three-week-old seedlings were immersed in 10 % PEG6000, 150 mM NaCl, 200 mM mannitol, 100 µM ABA, 10 mM H2O2 and 10 % sugar solutions for 3 h. Non-stressed plants were maintained as controls. Samples from roots, stems and leaves were frozen in liquid nitrogen and stored at −80 °C until RNA extraction. For inhibitor treatment, the H2O2 scavenger dimethyl thiourea (DMTU) was dissolved in distilled water to make a 10-M stock solution. Foxtail millet seedlings were pretreated with DMTU for 6 h, followed by exposure to PEG and mannitol treatments for 3 or 6 h. Actin-RNA was used as the internal reference. The 2 −ΔΔC t method was used to evaluate the relative amounts of transcript accumulated for the SiASR genes. Each experiment was repeated three times.
De novo transcriptome assembly
For drought treatment, 3-week-old foxtail millet seedlings were not watered for 1 week in soil (28 °C day/20 °C night, 16 h photoperiod, 65 % relative humidity) and the well-watered seedlings were used as the control. Construction of subtracted cDNA libraries, sequencing, data analysis of ESTs, differential screening of ESTs by microarray analysis and statistical analysis were performed as the procedures described by Puranik for exploring differential gene expression in response salinity stress (Puranik et al. 2011). The normalized data were subjected to fold difference calculation. ESTs that showed fold difference ≥2 were considered significant.
Subcellular localization
Subcellular localizations of green fluorescent protein (GFP) tags were used for protein localization analysis (Xu et al. 2007). The SiASR coding sequence was amplified and fused to the N-terminal end of GFP under control of the CaMV 35S promoter (for primers listed in Supplemental Table S2). The GFP fusion vector was transformed into onion epidermal cells and GFP fluorescence was observed by fluorescence microscopy (Liu et al. 2013).
Tobacco transformation
For tobacco transformation, the ORF of SiASR was cloned into the pBI121 vector to create a pBI121-SiNF-Y construct under the control of the CaMV 35S promoter (for primers listed in Supplemental Table S2). Transformation of tobacco (W38 genetic background) was performed using the Agrobacterium-mediated transformation method (Xu et al. 2009). Seeds from transformed tobacco plants were plated in 50 mg/L kanamycin selection medium in a growth chamber (16 h light/8 h darkness, 70 % relative humidity, 20 °C).
Abiotic stress tolerance assays
For the seed germination assay, 50 sterile seeds were transferred to and cultured on MS agar plates containing 2 % PEG, 100 mM mannitol or 0.5 µM methyl viologen (MV). Germination rates were scored at radicle emergence. To examine root morphologies, 5-day-old tobacco seedlings were cultured vertically on MS agar plates containing 4 % PEG, 150 mM mannitol or 2 µM MV for 5 days under the above conditions. Primary root lengths were measured. For drought treatment in soil, 2-week-old seedlings were not watered for 10 days and then re-watered for 20 days under the above conditions. Accumulation of H2O2 and O2− was detected using 3, 3′-diaminobenzidine (DAB) and nitroblue tetrazolium (NBT) as previously described (Yan et al. 2014). For oxidative treatment in soil, 2-week-old seedlings were watered with 10 µM MV for 10 days and then re-watered normally for 20 days. Survival rates were analyzed. For expression analysis of stress-responsive genes, leaves were sampled for qRT-PCR assays (for primers listed in Supplemental Table S2). All experiments were triplicated.
Physiological and biochemical detection
Before re-watering, leaves were harvested for measurement of relative water content (RWC), ion leakage, chlorophyll content, superoxide dismutase (SOD), catalase (CAT), peroxidase (POD) and malonaldehyde (MDA) activities (Wu et al. 2008; Hu et al. 2013; Zhang et al. 2014a). All experiments were triplicated.
Results
Identification of ASR family genes in foxtail millet
Six ASR genes were identified and isolated from foxtail millet (Table 1; Dataset 1). The ASR proteins consisted of 100–200 amino acids (Garay-Arroyo et al. 2000), with predicted molecular masses of 11–25 kDa and isoelectronic points (pI) 6–10 (Table 1). The genes were mapped to three chromosomes, with chromosome 8 harboring one locus, chromosome 7 harboring two loci and chromosome 5 harboring three loci. On chromosome 7, SiASR1 and SiASR6 were separated by 13,503 bp. On chromosome 5, SiASR3 and SiASR4 were separated by 652 bp, whereas SiASR3 and SiASR5 were separated by 2740 bp.
Multiple alignments and phylogenetic tree analysis of the ASR family
Multiple alignments showed that SiASR proteins contained a highly conserved ABA/WDS domain (PF02496) at the C-terminus (Supplemental Figs. S1A, S2). Two pairs of closely related proteins were identified in the phylogenetic tree analysis: (1) SiASR1 and SiASR2, and (2) SiASR3 and SiASR4 (Supplemental Fig. S1B). Multiple alignments showed that SiASR nucleotide sequence identities ranged from 58.1 to 86.4 % (Supplemental Fig. S3).
To evaluate the similarity/diversity of ASRs, the motif distribution among SiASR proteins was compared with other plant ASR proteins. Multiple alignments and MEME analysis indicated that ASR proteins were present and well conserved in both dicotyledonous and monocotyledonous (Supplemental Figs. S4, S5, S6). All ASR proteins contained conserved ABA/WDS domains (motifs 1 and 3) and bipartite nuclear localization signals (motif 2) (Supplemental Fig. S5). Motifs 1 and 3 contained amino acid groups determined to be Zn2+-dependent DNA-binding activity domains and a sequence possibly hindering DNA binding of ASR proteins. In addition, some ASRs contained a small N-terminal consensus containing a sequence of six His residues (motif 5).
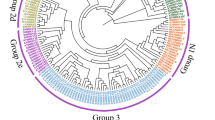
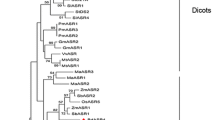
The phylogenetic tree revealed that ASRs consisted of three major groups, namely coniferous, dicotyledonous and monocotyledonous groups (Fig. 1). Dicotyledonous ASRs formed group A. Monocotyledonous ASRs formed groups B and C. SiASR1 was very close to ZmASR1 and TaASR1 in monocot group B. SiASR2 and SiASR5 were highly homologous to OsASR5 and ZmASR4, respectively, in monocot group B. SiASR3, SiASR4 and SiASR6 were highly homologous to OsASR2, OsASR1 and ZmASR5, respectively, in monocot group C.
Gene and promoter region structures of the ASR family
SiASR1 possessed two introns (Supplemental Fig. S1C); SiASR3 and SiASR4 had single exons; and SiASR2, SiASR5 and SiASR6 had one intron (Supplemental Fig. S1C).
The promoter sequences in the 2.0-kb region upstream of the SiASR genes were also analyzed. DRE (response to drought and ABA) boxes, CURECORECR (response to oxygen) boxes, MYB boxes (response to drought), MYC (response to drought) boxes and GT1GMSCAM4 (response to salt) cassettes were overrepresented in upstream regions of all ASR genes (Supplemental Table S3). In these five types of elements, the average numbers for each gene were DRE (2.8), CURECORECR (18.6), MYB (9.0), MYC (21.0) and GT1GMSCAM4 (5.0). This indicated that the ASR gene family in foxtail millet might be actively involved in drought and oxidative signal transduction.
Tissue expression profiles of the ASR family under abiotic stress
Analysis of tissue expression profiles of 6 ASRs by qRT-PCR for different tissues showed that SiASRs were expressed in all tissues tested, including roots, stems and leaves. SiASR1, SiASR2, SiASR3- and SiASR6 were preferentially expressed in the roots, whereas SiASR4 and SiASR5 were predominantly expressed in leaves (Fig. 2).
The expression patterns of SiASRs during PEG, NaCl and mannitol abiotic stresses in the three tissues were analyzed by qRT-PCR. Transcripts of ASRs were strongly up-regulated under PEG, NaCl and mannitol stresses in roots and leaves, but down-regulated in stems. As shown in Fig. 3, SiASR1 and SiASR5 were up-regulated in leaves, and SiASR3, SiASR4 and SiASR6 were expressed at high levels in roots. The highest increase in leaves under PEG and mannitol stresses was observed for SiASR1; the largest increase in roots under NaCl stress was observed for SiASR4.
Expression profiles of six SiASR genes under different stress treatments. Stresses included PEG, NaCl or mannitol treatment. Tissues included roots, stems and leaves. The gene expression level in stems under control condition for each ASR gene is seen as 1. Values are mean ± SE of three independent experiments
Drought, salt and osmotic responses interacted with various signal molecules, so we also examined the effect of ABA, H2O2 and sucrose on SiASRs transcription. After ABA, H2O2 and sucrose treatments, transcripts of SiASRs were induced to high levels in roots and leaves (Fig. 4). The highest increase was observed for SiASR1 with all three signaling molecules (Fig. 4).
Expression profiles of six SiASR genes under stress-related signal treatments. Stresses included ABA, H2O2 or sucrose treatment. Tissues included roots, stems and leaves. The gene expression level in stems under control condition for each ASR gene is seen as 1. Values are mean ± SE of three independent experiments
Each ASR gene member showed a differential expression pattern. For example, SiASR1 was mainly up-regulated in leaves under PEG, mannitol and H2O2 treatments. SiASR3 was mainly up-regulated under PEG and ABA treatments. SiASR4 was mainly up-regulated in roots under PEG and NaCl treatments. SiASR5 showed notable changes in leaves under all stress treatments. SiASR6 was mainly induced in roots under PEG, mannitol and ABA treatments.
In order to determine which members of the ASR family were up-regulated by drought stress, we compared de novo transcriptome assembly data of foxtail millet with and without drought treatment and found that all six ASRs had significantly different expression levels (Supplemental Table S4). Among six ASRs, SiASR1 showed the most significant up-regulation after drought treatment (~4.6 fold). Based on the de novo transcriptome assembly data and expression profiles (Figs. 3, 4), SiASR1 was selected for further investigation.
H2O2 signaling is involved in the induction of SiASR1 by PEG and mannitol treatments
Under PEG treatment, SiASR1 expression was significantly up-regulated (11.3 fold) after 3 h, followed by a decrease (Fig. 5a). To explore whether H2O2 was involved in up-regulation of SiASR1 under PEG treatment, foxtail millet seedlings were pretreated with the H2O2 inhibitor DMTU, prior to PEG treatment. Induction of SiASR1 expression was suppressed by pretreatment with DMTU in PEG-treated foxtail millet seedlings (Fig. 5b).
Effects of inhibitor of H2O2 on the SiASR1 transcripts under PEG and mannitol treatments. a SiASR1 transcript under PEG treatment, b Effect of inhibitor of H2O2 on the SiASR1 transcript under PEG treatment, c SiASR1 transcript under mannitol treatment, d Effect of inhibitor of H2O2 on the SiASR1 transcript under mannitol treatment. Values are mean ± SE of three independent experiments
Under mannitol treatment, SiASR1 was induced 3.4-fold at 1 h and reached the highest level at 3 h (5.9-fold) followed by a decrease (Fig. 5c). To explore whether H2O2 was involved in up-regulation of SiASR1 under mannitol treatment, foxtail millet seedlings were pretreated with the H2O2 inhibitor DMTU, prior to mannitol treatment. In a similar way, induction of SiASR1 expression was suppressed by pretreatment with DMTU in mannitol-treated foxtail millet seedlings (Fig. 5d). Moreover, cis-regulatory elements (CURECORECR, MYB and MYC boxes) detected upstream of the translation initiation site of SiASR1 (Supplemental Table S3) further implied involvement in drought, osmotic and oxidative responses. The results suggested that H2O2 was involved in up-regulation of SiASR1 under PEG and mannitol treatments.
Sequence and subcellular localization analysis of SiASR1
The SiASR1 cDNA consisted of 945 bp, with a 603-bp open reading frame (ORF) and a 342-bp 3′ untranslated region. SiASR1 encoded 200 amino acids with a calculated molecular mass of 22.5 kDa and a predicted pI of 6.3 (Table 1; Supplemental Dataset S1). Homology modeling showed that the secondary structure of SiASR1 protein was predominantly comprised of α-helices and coils (Supplemental Fig. S7). Cellular localization showed that SiASR1-GFP fusion protein was confined to the nucleus, whereas GFP was uniformly distributed throughout the cell in the control (Fig. 6).
Overexpression of SiASR1 enhanced drought and osmotic tolerances
Transgenic tobacco plants overexpressing SiASR1 were generated to investigate the biological function of SiASR1. To examine whether SiASR1 mediated drought stress response, we compared WT and SiASR1 overexpressing plants in a drought tolerance assay. When sown on MS media containing PEG or mannitol, higher germination rates were observed in SiASR1 transgenic seeds than in WT (Fig. 7). After 5 days of PEG stress, 80.02–82.28 % of SiASR1 transgenic seeds had germinated compared to 36.50 % for WT seeds germinated (Fig. 7). Under mannitol stress, 59.30–61.36 % of SiASR1 transgenic seeds germinated, compared to 24.61 % for WT seeds (Fig. 7). Seedling root growth of SiASR1 transgenic tobacco was also investigated on MS medium treated with PEG and mannitol. As shown in Fig. 8, when the seedlings were placed on MS media containing PEG and mannitol for 5 days, growth of primary roots of WT was significantly inhibited, whereas the roots of transgenic seedlings exhibited stronger and longer growth than WT (Fig. 8). Under PEG stress treatment, the primary roots were reduced to 50–53 % in transgenic seedlings, whereas primary roots in the WT seedlings was 37 %, compared with that in the untreated seedlings (Fig. 8). Under mannitol stress treatment, the primary roots were reduced to 80–83 % in transgenic seedlings, whereas primary roots in the WT seedlings was reduced to 42 %, compared with that in the untreated seedlings (Fig. 8).
Analysis of root elongation in WT and SiASR1 transgenic seedlings exposed to drought and oxidative stresses. a Root elongation of tobacco plants after treatment with PEG, mannitol and MV. b Statistical analysis of the root length. Scale bars 1 cm. Values are mean ± SE of three independent experiments
Overexpression of ASR1 enhanced oxidative tolerance
Expression pattern analysis in foxtail millet revealed that SiASR1 was involved in response to oxidative stress. To examine the role of SiASR1 in oxidative response, we compared WT and SiASR1 transgenic plants in an oxidative-tolerance assay (Fig. 7). MV is a herbicide that causes chlorophyll degradation and cell membrane leakage through ROS production (Yan et al. 2014). In the MS medium supplemented with MV, germination of WT seeds was inhibited, whereas transgenic seeds continued to germinate (Fig. 7). The germination rate of WT was 40.20 % compared to 80.50–86.52 % for transgenic seeds (Fig. 7). Similarly, in the presence of MV, the decrease in the primary length of the transgenic seedlings was more remarkable than that in the WT seedlings (Fig. 8). Moreover, for plant growth, the growth on WT seedlings was retarded and the WT seedlings exhibited a greater reduction in the whole seedling fresh weight and dry weight compared with the transgenic seedlings (Fig. 8).
Overexpression of SiASR1 decreased H2O2 accumulation and improved antioxidant enzyme activities under drought and oxidative stresses
To examine the role of SiASR1 in drought response in soil, WT and SiASR1 transgenic seedlings were withheld from water. After drought treatment, the leaves of the WT wilted severely and approached death (Fig. 9a). After re-watering, some WT plants became dark and died. By comparison, SiASR1 transgenic plants displayed better growth and more green leaves than WT plants. In addition, drought-treated transgenic plants showed lower accumulations of H2O2 and O2− than WT in the leaves as evidenced by brown (DAB staining) and blue (NBT staining) pigments (Fig. 9b). Survival rates were 33.33 % for WT plants compared to 94.40–97.13 % of SiASR1 transgenic lines (Fig. 9c). Physiological traits related to drought stress tolerance were also analyzed. Under normal conditions, there was no difference in RWC and ion leakage between WT and transgenic seedlings, but under drought conditions, transgenic plants showed higher RWC and lower ion leakage compared to WT seedlings (Fig. 9c). Drought stress markedly increased SOD, CAT and POD activities in transgenic plants, compared to WT plants (Fig. 9d).
Overexpression of SiASR1 decreased ROS production and oxidative damage under drought stress conditions. a Sensitivity of WT and transgenic plants to drought stress. b In situ detection of H2O2 and O2− by DAB and NBT staining of WT and transgenic seedlings under drought stress. c Survival rate, ion leakage and RWC contents. d Activities of SOD, POD and CAT. Values are mean ± SE of three independent experiments
WT plants grown in soil irrigated with MV also showed weaker development and more severe chlorosis than transgenic plants (Fig. 10a). Detached leaf pieces from the transgenic lines appeared healthier and showed less chlorophyll degradation than those from WT plants (Fig. 10b). Physiological differences were also examined under oxidative stress and normal conditions. Without oxidative stress, no differences were detected between transgenic lines and WT with respect to ion leakage, chlorophyll content, MDA content and SOD, POD and CAT activities (Fig. 10c, d). However, under oxidative stress, chlorophyll content in transgenic seedlings was 1.8–1.9 mg g−1 higher than that in WT (0.7 mg g−1) (Fig. 10c). MDA content was increased by 5.3 µM g−1 in WT plants, compared to 3.5–4.0 µM g−1 in transgenic plants (Fig. 10c). Transgenic plants had markedly higher SOD, CAT and POD activities than WT plants (Fig. 10d).
Oxidative stress tolerance of SiASR1 transgenic plants. a Representative images showing leaf senescence during oxidative treatment. b Leaf discs obtained from WT and transgenic seedlings are incubated in MV (50 μM) for 72 h and then they are photographed. c RWC, chlorphyll and MDA contents. d Activities of SOD, POD and CAT. Values are mean ± SE of three independent experiments
Overexpression of SiASR1 altered expression levels of oxidation-related genes under stress conditions
To determine the possible mechanism of SiASR1 in ROS scavenging, the expression levels of genes that encode ROS-scavenging enzymes [ascorbate peroxidase (APX), CAT and SOD] and ROS producers, i.e. the respiratory burst oxidase homologs (RbohA and RbohB) were examined. No obvious differences in expression were detected between transgenic and WT plants under normal conditions (Fig. 11), but following drought and oxidative stresses, the expressions of APX, CAT and SOD were clearly increased in transgenic plants; transcript accumulations of RbohA and RbohB were considerably lower in transgenic compared to WT plants (Fig. 11a, b).
Gene expression in leaves of SiASR1 transgenic plants. Expression levels of ROS-scavenging and producing genes in leaves of WT and transgenic plants during drought (a) and oxidative (b) stress responses as determined by qRT-PCR. The β-tubulin gene was used as an internal reference. V. Asterisk significant at P = 0.05
Discussion
Foxtail millet possesses several salient attributes, such as a small genome, short life-cycle and drought tolerance, that stand to become more important in a potentially dryer future environment (Li and Brutnell 2011; Muthamilarasan and Prasad 2015). Plant cells respond to dehydration through triggering several signal transduction pathways that result in accumulation of certain proteins, sugar molecules and lipophilic antioxidants (Huang et al. 2012). Among the proteins, the ancestral ASR gene family served as an example of positive selection during colonization of arid regions by adaptive evolution following gene duplication (Zhang et al. 2015).
Varied expression of ASRs in response to stress-related signals and abiotic stresses
Abscisic acid stress ripening transcripts accumulated under different abiotic stresses and ASR proteins acted as downstream components of a transduction pathway (Cakir et al. 2003; Ricardi et al. 2014; Zhang et al. 2015). Systematic analyses of expression patterns showed that all six ASR genes in foxtail millet were induced by various abiotic stresses and stress-related signals (Figs. 3, 4). The widespread induction of ASR genes in response to diverse stress stimuli suggested that ASRs play important crosstalk roles in diverse stress tolerances (Cakir et al. 2003; Cortés et al. 2012; Kim et al. 2012).
Many recent reports describe crosstalk between drought and salt stresses and signal molecules in the model plant of Arabidopsis. However, studies in crop species were still limited (Alam et al. 2015; Chen et al. 2015; Yan et al. 2014; Zhang et al. 2014b). Drought-imposed osmotic stress elicits ROS production (Krasensky and Jonak 2012). ROS act as important signal transduction molecules mediating various stress responses (Golldack et al. 2014). As for TaASR1 (Hu et al. 2013), the H2O2 inhibitor assay confirmed that H2O2 was an important signal involved in the induction of SiASR1 transcript to drought and osmotic stresses. These results implied that stress-related signals contribute to an interactive network that coordinates the responses of foxtail millet ASR genes to different stresses.
SiASR1 enhanced drought and oxidative tolerance through antioxidant enzymes
Abscisic acid stress ripening proteins were shown to possess zinc-dependent DNA-binding activity and to modulate the expression of other genes. For example, grape VvMSA directly targeted the promoter of a sugar transporter gene (Cakir et al. 2003). Several putative targets of rice OsASR5 in roots related to Al tolerance, such as an ABC transporter, were found through chromatin immunoprecipitation (ChIP) assays (Arenhart et al. 2014). Tomato SlASR1 bound to a large number of water transport and cell wall remodeling genes (Ricardi et al. 2014). Wheat TaASR1 modulated the expressions of antioxidant defense-associated genes in tobacco (Hu et al. 2013).
Abscisic acid stress ripening proteins also play a critical role as toxic ROS scavengers and chaperone-like proteins by preventing protein denaturation in response to exogenous stimuli until other protective proteins or mechanisms were activated (Kim et al. 2012). These protective proteins included metabolic, antioxidant enzyme and chaperone proteins which affected the redox state and proteostasis in plant cell. For example, rice OsASR1 protein was shown to act more highly synergistically with the osmolyte glycine-betaine in regulating chaperone-mediated protein disaggregation than proline and threhalose under stress conditions and to regulate branched-chain amino acid biosynthesis (Konrad and Bar-Zvi 2008).
In the present study, SiASR1 functioned in drought stress by regulating cellular levels of ROS and being involved in ROS signaling. DAB and NBT histochemical staining assay and detection of antioxidant enzymes activities indicated SiASR1 transgenic plants decreased ROS accumulation under drought stress (Fig. 9). The enhanced activities of antioxidant enzymes protected plants against ROS damage due to drought stress. The lower oxidative injury of SiASR1-overexpressing plants under MV stress further demonstrated the involvement of SiASR1 in ROS signaling which was an integral part of acclimation of plants to drought (Suzuki et al. 2011). The expression levels of antioxidant genes (SOD, CAT and APX) significantly increased in SiASR1 transgenic plants, while the expression levels of ROS-producing genes (RbohA and RbohB) significantly decreased in SiASR1 transgenic plants under drought and oxidative stresses (Fig. 11). Although the hypothesis that SiASR1 regulates these oxidation-related genes directly needs to be confirmed in detail, our results indicate that SiASR1 protein acts as ROS regulators by increasing the transcription of SiASR1-regulated genes or chaperon activity during abiotic stress.
Author contribution statement
Z.S.X. coordinated the project, conceived and designed the experiments and edited the manuscript. Z.J.F. performed experiments, analyzed data, and wrote the first draft. J.T.S. conducted the bioinformatic work and analyzed the data. M.C. and L.C.L. managed reagents and provided analytical tools. Y.Z.M., G.Y.H. and G.X.Y. contributed with valuable discussions. All authors have read and approved the final manuscript.
Abbreviations
- ASR:
-
Abscisic acid stress ripening
- CAT:
-
Catalase
- DAB:
-
3,3′-Diaminobenzidine
- GFP:
-
Green fluorescent protein
- MDA:
-
Malonaldehyde
- MV:
-
Methyl viologen
- NBT:
-
Nitroblue tetrazolium
- POD:
-
Peroxidase
- qRT-PCR:
-
Quantitative real-time PCR
- RWC:
-
Relative water content
- SOD:
-
Superoxide dismutase
References
Alam MM, Tanaka T, Nakamura H, Ichikawa H, Kobayashi K, Yaeno T, Yamaoka N, Shimomoto K, Takayama K, Nishina H, Nishiguchi M (2015) Overexpression of a rice heme activator protein gene (OsHAP2E) confers resistance to pathogens, salinity and drought, and increases photosynthesis and tiller number. Plant Biotechnol J 13:85–96
Arenhart RA, Bai Y, Valter de Oliveira LF, Bucker Neto L, Schunemann M, Maraschin Fdos S, Mariath J, Silverio A, Sachetto-Martins G, Margis R, Wang ZY, Margis-Pinheiro M (2014) New insights into aluminum tolerance in rice: the ASR5 protein binds the STAR1 promoter and other aluminum-responsive genes. Mol Plant 7:709–721
Briesemeister S, Rahnenfuhrer J, Kohlbacher O (2010) YLoc-an interpretable web server for predicting subcellular localization. Nucl Acids Res 38:W497–W502
Cakir B, Agasse A, Gaillard C, Saumonneau A, Delrot S (2003) A grape ASR protein involved in sugar and abscisic acid signaling. Plant Cell 15:2165–2180
Carrari F, Fernie AR, Iusem ND (2004) Heard it through the grapevine ABA and sugar cross-talk: the ASR story. Trends Plant Sci 9:57–59
Chen M, Zhao Y, Zhuo C, Lu S, Guo Z (2015) Overexpression of a NF-YC transcription factor from bermudagrass confers tolerance to drought and salinity in transgenic rice. Plant Biotechnol J 13:482–491
Cortés AJ, Chavarro MC, Madriñán S, This D, Blair MW (2012) Molecular ecology and selection in the drought-related Asr gene polymorphisms in wild and cultivated common bean (Phaseolus vulgaris L.). BMC Genet 13:58–71
Garay-Arroyo A, Colmenero-Flores JM, Garciarrubio A, Covarrubias AA (2000) Highly hydrophilic proteins in prokaryotes and eukaryotes are common during conditions of water deficit. J Biol Chem 275:5668–5674
Golldack D, Lüking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30:1383–1391
Golldack D, Li C, Mohan H, Probst N (2014) Tolerance to drought and salt stress in plants: unraveling the signaling networks. Front Plant Sci 5:151
González RM, Iusem ND (2014) Twenty years of research on Asr (ABA-stress-ripening) genes and proteins. Planta 239:941–949
Hsu YF, Yu SC, Yang CY, Wang CS (2011) Lily ASR protein-conferred cold and freezing resistance in Arabidopsis. Plant Physiol Biochem 49:937–945
Hu W, Huang C, Deng X, Zhou S, Chen L, Li Y, Wang C, Ma Z, Yuan Q, Wang Y, Cai R, Liang X, Yang G, He G (2013) TaASR1, a transcription factor gene in wheat, confers drought stress tolerance in transgenic tobacco. Plant Cell Environ 36:1449–1464
Hu YX, Yang X, Li XL, Yu XD, Li QL (2014) The SlASR gene cloned from the extreme halophyte Suaeda liaotungensis K. enhances abiotic stress tolerance in transgenic Arabidopsis thaliana. Gene 549:243–251
Huang GT, Ma SL, Bai LP, Zhang L, Ma H, Jia P, Liu J, Zhong M, Guo ZF (2012) Signal transduction during cold, salt, and drought stresses in plants. Mol Biol Rep 39:969–987
Iusem ND, Bartholomew DM, Hitz WD, Scolnik PA (1993) Tomato (Lycopersicon esculentum) transcript induced by water deficit and ripening. Plant Physiol 102:353–1354
Kim SJ, Lee SC, Hong SK, An K, An G, Kim SR (2009) Ectopic expression of a cold-responsive OsAsr1 cDNA gives enhanced cold tolerance in transgenic rice plants. Mol Cells 27:449–458
Kim IS, Kim YS, Yoon HS (2012) Rice ASR1 protein with reactive oxygen species scavenging and chaperone-like activities. Mol Cells 33:285–293
Konrad Z, Bar-Zvi D (2008) Synergism between the chaperone-like activity of the stress regulated ASR1 protein and the osmolyte glycine-betaine. Planta 227:1213–1219
Krasensky J, Jonak C (2012) Drought, salt and temperature stress-induced metabolic rearrangements and regulatory networks. J Exp Bot 63:1593–1608
Lata C, Gupta S, Prasad M (2013) Foxtail millet: a model crop for genetic, genomic studies in bioenergy grasses. Crit Rev Biotechnol 33:328–343
Letunic I, Doerks T, Bork P (2012) SMART 7: recent updates to the protein domain annotation resource. Nucl Acids Res 40:D302–D305
Li P, Brutnell TP (2011) Setaria viridis and Setaria italica, model genetic systems for the Panicoid grasses. J Exp Bot 62:3031–3037
Li RH, Liu GB, Wang H, Zheng YZ (2013) Effects of Fe3+ and Zn2+ on the structural and thermodynamic properties of a soybean ASR protein. Biosci Biotechnol Biochem 77:475–481
Liu HY, Dai JR, Feng DR, Liu B, Wang HB, Wang JF (2010) Characterization of a novel plantain Asr gene, MpAsr, that is regulated in response to infection of Fusarium oxysporum f. sp. cubense and abiotic stresses. J Integr Plant Biol 52:315–323
Liu P, Xu ZS, Lu PP, Hu D, Chen M, Li LC, Ma YZ (2013) A wheat PI4K gene whose product possesses threonine autophophorylation activity confers tolerance to drought and salt in Arabidopsis. J Exp Bot 64:2915–2927
Marchler-Bauer A, Anderson JB, Cherukuri PF, DeWweese-Scott C, Geer LY, Gwadz M, He S, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Liebert CA, Liu C, Lu F, Marchler GH, Mullokandov M, Shoemaker BA, Simonyan V, Song JS, Thiessen PA, Yamashita RA, Yin JJ, Zhang D, Bryant SH (2005) CDD: a conserved domain database for protein classification. Nucl Acids Res 33:D192–D196
Muthamilarasan M, Prasad M (2015) Advances in Setaria genomics for genetic improvement of cereals and bioenergy grasses. Theor Appl Genet 128:1–14
Puranik S, Jha S, Srivastava PS, Sreenivasulu N, Prasad M (2011) Comparative transcriptome analysis of contrasting foxtail millet cultivars in response to short-term salinity stress. J Plant Physiol 168:280–287
Ricardi MM, González RM, Zhong S, Domínguez PG, Duffy T, Turjanski PG, Salgado Salter JD, Alleva K, Carrari F, Giovannoni JJ, Estévez JM, Iusem ND (2014) Genome-wide data (ChIP-seq) enabled identification of cell wall-related and aquaporin genes as targets of tomato ASR1, a drought stress-responsive transcription factor. BMC Plant Biol 14:1–29
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227
Shkolnik D, Bar-Zvi D (2008) Tomato ASR1 abrogates the response to abscisic acid and glucose in Arabidopsis by competing with ABI4 for DNA binding. Plant Biotechnol J 6:368–378
Suzuki N, Miller G, Morales J, Shulaev V, Torres MA, Mittler R (2011) Respiratory burst oxidases: the engines of ROS signaling. Curr Opin Plant Biol 14:691–699
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Virlouvet L, Jacquemot MP, Gerentes D, Corti H, Bouton S, Gilard F, Valot B, Trouverie J, Tcherkez G, Falque M, Damerval C, Rogowsky P, Perez P, Noctor G, Zivy M, Coursol S (2011) The ZmASR1 protein influences branched-chain amino acid biosynthesis and maintains kernel yield in maize under water-limited conditions. Plant Physiol 157:917–936
Wu L, Zhang Z, Zhang H, Wang XC, Huang R (2008) Transcriptional modulation of ethylene response factor protein JERF3 in the oxidative stress response enhances tolerance of tobacco seedlings to salt, drought, and freezing. Plant Physiol 148:1953–1963
Xu ZS, Xia LQ, Chen M, Cheng XG, Zhang RY, Li LC, Zhao YX, Lu Y, Ni ZY, Liu L, Qiu ZG, Ma YZ (2007) Isolation and molecular characterization of the Triticum aestivum ethylene-responsive factor 1 (TaERF1) that increases multiple stress tolerance. Plant Mol Biol 65:719–732
Xu ZS, Xiong TF, Ni ZY, Chen XP, Chen M, Li LC, Ma YZ (2009) Isolation and identification of two genes encoding leucine-rich repeat (LRR) proteins differentially responsive to pathogen attack and salt stress in tobacco. Plant Sci 176:38–45
Yan HR, Jia HH, Chen XB, Hao LL, An HL, Guo X (2014) The cotton WRKY transcription factor GhWRKY17 functions in drought and salt stress in transgenic Nicotiana benthamiana through ABA signaling and the modulation of reactive oxygen species production. Plant Cell Physiol 55:2060–2076
Yang CY, Wu CH, Jauh GY, Huang JC, Lin CC, Wang CS (2008) The LLA23 protein translocates into nuclei shortly before desiccation in developing pollen grains and regulates gene expression in Arabidopsis. Protoplasma 223:241–254
Zhang G, Liu X, Quan Z, Cheng S, Xu X, Pan S, Xie M, Zeng P, Yue Z, Wang W, Tao Y, Bian C, Han C, Xia Q, Peng X, Cao R, Yang X, Zhan D, Hu J, Zhang Y, Li H, Li H, Li N, Wang J, Wang C, Wang R, Guo T, Cai Y, Liu C, Xiang H, Shi Q, Huang P, Chen Q, Li Y, Wang J, Zhao Z, Wang J (2012) Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nat Biotechnol 30:549–554
Zhang D, Jiang S, Pan J, Kong X, Zhou Y, Liu Y, Li D (2014a) The overexpression of a maize mitogen-activated protein kinase gene (ZmMPK5) confers salt stress tolerance and induces defence responses in tobacco. Plant Biol (Stuttg) 16:558–570
Zhang XY, Wang LM, Xu XY, Cai CP, Guo WZ (2014b) Genome-wide identification of mitogen-activated protein kinase gene family in Gossypium raimondii and the function of their corresponding orthologs in tetraploid cultivated cotton. BMC Plant Biol 14:345–361
Zhang L, Hu W, Wang Y, Feng R, Zhang Y, Liu J, Jia C, Miao H, Zhang J, Xu B, Jin Z (2015) The MaASR gene as a crucial component in multiple drought stress response pathways in Arabidopsis. Funct Integr Genom 15:247–260
Acknowledgments
This research was financially supported by the National Transgenic Key Project of Ministry of Agriculture (2014ZX08009-016B and 2014ZX08002-003B). We are grateful to Dr. Xianmin Diao, Institute of Crop Science, Chinese Academy of Agricultural Sciences, for kindly providing foxtail millet seeds.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by M. Menossi.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1
Supplemental Fig. S1 (A) Conserved domains of SiASR proteins. (B) Phylogenetic relationship of SiASRs. (C) Structure of SiASR genes. (TIFF 414 kb)
Supplementary material 2
Supplemental Fig. S2 Multiple sequence alignments of SiASR full-length CDs. (TIFF 318 kb)
Supplementary material 3
Supplemental Fig. S3 Multiple sequence alignments of SiASR full-length proteins. (TIFF 1261 kb)
Supplementary material 4
Supplemental Fig. S4 Multiple alignments of SiASR proteins with other ASR proteins. (TIFF 2426 kb)
Supplementary material 5
Supplemental Fig. S5 Variation in motif clades for ASR proteins from foxtail millet with other ASR proteins. The MEME motifs are shown as differently colored boxes. (TIFF 766 kb)
Supplementary material 6
Supplemental Fig. S6 Conserved motifs identified for ASR proteins by MEME software. (TIFF 823 kb)
Supplementary material 7
Supplemental Fig. S7 Predicted secondary structure of SiASR1 protein by Phyre2. (TIFF 69 kb)
Supplementary material 8
Supplemental Table S1. Nomenclature of ASRs in foxtail millet and other species. (XLS 36 kb)
Supplementary material 9
Supplemental Table S2. Primers used in this study. (XLS 24 kb)
Supplementary material 10
Supplemental Table S3. Prediction of cis-elements in SiASR from database analysis. The data were obtained from PLACE (http://www.dna.affrc.go.jp/PLACE/). (XLS 12 kb)
Supplementary material 11
Supplemental Table S4. Drought-induced foxtail millet ASR genes from the de novo transcriptome assembly sequencing data. The whole foxtail millet seedlings were used for the experiment. FDR: false discovery rate. (XLS 27 kb)
Rights and permissions
About this article
Cite this article
Feng, ZJ., Xu, ZS., Sun, J. et al. Investigation of the ASR family in foxtail millet and the role of ASR1 in drought/oxidative stress tolerance. Plant Cell Rep 35, 115–128 (2016). https://doi.org/10.1007/s00299-015-1873-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-015-1873-y