Abstract
Introduction
For regional quantification of nuclear brain imaging data, defining volumes of interest (VOIs) by hand is still the gold standard. As this procedure is time-consuming and operator-dependent, a variety of software tools for automated identification of neuroanatomical structures were developed. As the quality and performance of those tools are poorly investigated so far in analyzing amyloid PET data, we compared in this project four algorithms for automated VOI definition (HERMES Brass, two PMOD approaches, and FreeSurfer) against the conventional method. We systematically analyzed florbetaben brain PET and MRI data of ten patients with probable Alzheimer’s dementia (AD) and ten age-matched healthy controls (HCs) collected in a previous clinical study.
Methods
VOIs were manually defined on the data as well as through the four automated workflows. Standardized uptake value ratios (SUVRs) with the cerebellar cortex as a reference region were obtained for each VOI. SUVR comparisons between ADs and HCs were carried out using Mann-Whitney-U tests, and effect sizes (Cohen’s d) were calculated. SUVRs of automatically generated VOIs were correlated with SUVRs of conventionally derived VOIs (Pearson’s tests).
Results
The composite neocortex SUVRs obtained by manually defined VOIs were significantly higher for ADs vs. HCs (p=0.010, d=1.53). This was also the case for the four tested automated approaches which achieved effect sizes of d=1.38 to d=1.62. SUVRs of automatically generated VOIs correlated significantly with those of the hand-drawn VOIs in a number of brain regions, with regional differences in the degree of these correlations. Best overall correlation was observed in the lateral temporal VOI for all tested software tools (r=0.82 to r=0.95, p<0.001).
Conclusion
Automated VOI definition by the software tools tested has a great potential to substitute for the current standard procedure to manually define VOIs in β-amyloid PET data analysis.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
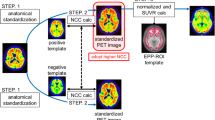
For regional quantification of brain PET data, it is still the gold standard to define the volumes of interest (VOIs) manually on individual MRIs. This conventional process has to deal, however, with two problems: It is time-consuming and operator-dependent [1, 2]. These limitations have stimulated the search for alternative solutions, which allow identifying brain VOIs automatically [3–5]. Respective software solutions, which vary widely in underlying principles and workflows, are now at a “ready to use” stage. Underlying essential principals of these tools are intensity-based segmentation [6] and spatial normalization to a template space [7, 8]. Some software tools combine both approaches generatively with additional statistical classifiers leading to maximum-a-posteriori solutions for each VOI label. [9, 10].
One desired application of such automated approaches to delineate anatomical VOIs is in β-amyloid PET imaging. Recently, three 18F-labeled β-amyloid plaque-targeting PET tracers ([18F]florbetapir, [18F]florbetaben, [18F]flutemetamol) were approved for clinical routine use [11]. In β-amyloid PET imaging, especially for supporting standard visual assessment in borderline cases and for follow-up imaging, quantification of the PET signal by anatomical VOI analysis is employed. So far, however, there is a lack of systematic comparative investigations of the different software tools available to perform these quantifications for amyloid PET data.
This situation inspired the present research which dealt with the question of whether the automated neuroanatomical VOI definition tools tested are capable of substituting the current gold standard approach of manual VOI definition to analyze [18F]florbetaben β-amyloid PET images.
Materials and methods
Study population
The chosen software tools for automated VOI definition were tested on the data of the European [18F]florbetaben phase 0 proof of mechanism trial [12]. The dataset included ten patients with mild to moderate probable Alzheimer’s dementia (AD; 69±7 yrs; two females; mini–mental state examination (MMSE) score: 19±7; clinical dementia rating (CDR) score: 1.5 ± 0.5) and ten sex- and age-matched healthy controls (HCs; 67±8 years; two females; MMSE score: 29±1; CDR score 0±0).
Image data acquisition
After the i.v. administration of 300±60 MBq of [18F]florbetaben, PET images were acquired in 3-D mode using an ECAT EXACT HR+ scanner (Siemens, Erlangen, Germany). Brain MRI data were obtained on a 1.5 T Siemens Magnetom Symphony scanner. For that project, standardized T1-weighted volumetric magnetization prepared rapid acquisition with gradient echo (MPRAGE) sequences were used.
Image data processing
PET data frames in the earliest part of the plateau phase of tracer accumulation (70-90 min p.i.) were chosen for further analysis [13]. The data were corrected as described elsewhere [12] and iteratively reconstructed (ten iterations, 17 subsets, Gaussian filter with a full-width at half-maximum cut-off of 7.1 mm). Corresponding PET images were coregistered to the individual MRIs employing the normalized mutual information criterion implemented in the PMOD software [14] (PMOD version 3.3, PMOD Technologies Ltd., Zurich, Switzerland) using the default settings.
Conventional VOI definition
The conventional VOI dataset (“Leipzig region map”) consisted of 25 VOIs (frontal cortex (right/left), lateral temporal cortex (r/l), mesial temporal cortex (r/l), parietal cortex (r/l), occipital cortex (r/l), anterior cingulate cortex (r/l), posterior cingulate cortex and precuneus (r/l), head of caudate nucleus (r/l), putamen (r/l), thalamus (r/l), white matter (r/l), cerebellar cortex (r/l), and pons/midbrain) (Fig. 1). For each subject, the individual 3-D T1 MPRAGE MRI scan was reoriented perpendicular to the anterior-posterior commissure (AC-PC) line, and the above VOIs were manually defined by an experienced neurobiologist in three adjacent transversal slices with a thickness of 2.5 mm per slice using the VOI tool implemented in the PMOD software.
Representative individual 3-D T1-weighted MRI data set with display of the conventional (manually defined) volume of interest (VOI) set. Five paradigmatic transversal slices reoriented perpendicular to the anterior commissure-posterior commissure (AC-PC) line. VOIs were defined for a cerebellar cortex (red) and pons/midbrain (yellow), b mesial (red) and lateral temporal cortex (blue), c occipital cortex (blue), d frontal cortex (blue), caudate head (yellow), putamen (green) and thalamus (turquoise) and e parietal cortex (blue), anterior (purple) and posterior cingulate cortex/precuneus (yellow) and white matter (pink)
Automated VOI definition
The automated neuroanatomical VOI definition tools tested in this project included one PET-based and one MRI-based normalization as well as one volumetric and one spherical hybrid algorithm. The specifications of the four software tools tested are displayed in Table 1.
One automated VOI set was created in the HERMES Brass software (version 2.5, Hermes Medical Solutions, Stockholm, Sweden) [15–18]. A [18F]florbetaben HERMES BRASS normal database was created in-house utilizing the [18F]florbetaben PET images of 93 β-amyloid PET-negative healthy subjects. The corresponding anatomical VOI atlas was created by manual delineation of 25 VOIs on the ICBM152 standard template MRI defining the same brain regions as for the conventional VOI set.
Furthermore, we tested the PMOD Normalization algorithm, which implements the SPM 5 normalization algorithm [7] and an adjusted version of Tzourio-Mazoyer’s AAL atlas [4] edited by PMOD Technologies Ltd. Within the PFUS tool of PMOD (version 3.308), the subject’s T1 MPRAGE MRI was spatially normalized to PMOD’s brain template ‘MR T1 HFS’ using the default settings. The same transformation was applied to the coregistered PET image resulting in the spatially normalized PET image of the subject.
Furthermore, VOI datasets were created in PMOD (version 3.403 beta) using the ‘Maximum Probability’ workflow integrated in PNEURO. This workflow is based on the SPM5 segmentation and normalization algorithm [9] and resulted in VOIs of the Hammers N30R83 Maximum Probability atlas [19]. Parameter settings were: ‘MNI T1’ – template, no cropping, mask threshold of 0.3 (default) for intersection of the atlas regions with the gray matter probability map, masking of basal nuclei, ‘Individual – MR’ as result space.
Another automated anatomical VOI definition was achieved by using the open source software FreeSurfer (version v5.1.0, Athinoula A. Martinos Center for Biomedical Imaging, Charlestown, Massachusetts, USA) [20–22]. After import of every subject’s MRI into the the FreeSurfer environment, the fully automated ‘autorecon’ – workflow (command: ‘recon-all -all’) was applied. The resulting output aparc.annot files which refer to parcellations of the Desikan-Killiany [23] atlas were used for further analysis.
VOI data post-processing
For all the conventional and automated anatomical VOI definition approaches, regional tracer uptake values were obtained. VOIs which were not fully covered by the PET camera field of view were not considered. Three of the tested automated atlases contained more VOIs than the conventional VOI dataset (PMOD Normalization, PMOD Maximum Probability, FreeSurfer). To correct for that, several VOIs were pooled by the experienced neurobiologist calculating the volume-weighted tracer uptake averages. All regional uptake values were divided by the cerebellar cortex uptake values, which resulted in regional standardized uptake value ratios (SUVRs). Furthermore, composite SUVRs were calculated for every subject in slide modification from the method published by Rowe et al. [24] by calculating the volume-weighted mean value of the SUVRs from the frontal, parietal, lateral temporal, anterior and posterior cingulate, and occipital cortices.
Statistics
Statistical analyses were performed using the IBM SPSS Statistics software (version 20.0, IBM Corp., Armonk, NY, USA) and SigmaPlot software (version 9.01, Systat Software GmbH, Erkrath, Germany). Hemispherical SUVR differences within identical VOIs were evaluated with the Wilcoxon test and Bonferroni correction for multiple comparisons. Group differences in SUVR data were tested for significance using the Mann-Whitney U test. Effect sizes of the SUVR differences between groups were expressed as Cohen’s d. Correlations between manual and automated VOI datasets were calculated with linear regression analysis and Pearson’s test. Inter-rater reliability was expressed as Cohen’s kappa. Unless stated, data are mean value ±1 standard deviation. Due to the explorative nature of this study, no further corrections for multiple comparisons were made. A p value of <0.05 was considered significant.
Results
Paired comparisons of SUVRs obtained for the left and the right hemisphere revealed significant interhemispherical differences in five regions for the PMOD Normalization method (parietal, occipital, anterior cingulate, posterior cingulate cortex, and caudate nucleus), in two regions for the HERMES Brass method (occipital cortex, thalamus) and for one pair using the PMOD Maximum Probability tool (anterior cingulate cortex). No significant interhemispherical differences in tracer uptake were obtained by the FreeSurfer algorithm and by the conventional VOI definition method.
For the conventional as well as for all tested automated anatomical VOI definition methods, SUVRs were significantly higher in the AD patients as compared to the HCs in a number of brain regions (Table 2).
The regional distribution of the SUVR effect sizes for the group discrimination between the AD patients and HCs for all anatomical VOI definition methods is illustrated in Fig. 2. It becomes evident that, in a number of neocortical VOIs, group discrimination was better by different automated methods as compared to the conventional method (Fig. 2). On a regional level, best group discrimination was achieved by the PMOD Maximum Probability method for the left lateral temporal cortex SUVRs (Cohen’s d=1.68). Also of interest, for three brain VOIs in which the conventional method did not reveal significant SUVR differences between the AD patients and HCs, different automated anatomical VOI definition methods resulted in significant group differences: left occipital cortex (HERMES Brass, PMOD Maximum Probability), left mesial temporal cortex (PMOD Normalization, FreeSurfer), and left thalamus (PMOD Normalization) (Table 2). In keeping with that, the effect sizes of the composite SUVRs were higher for the PMOD Maximum Probability than for the conventional method (Fig. 3).
Effect sizes of [18F]florbetaben PET standardized uptake value ratio differences between patients with Alzheimer’s disease and healthy controls displayed as color-coded Cohen’s d for every analyzed brain region (representative axial slices on hippocampus level (top), level of the basal ganglia (middle) and level of the corpus callosum (bottom)) for the five tested neuroanatomical volume of interest definition approaches. d: Cohen’s d, CC: Cerebellar cortex, LTC: Lateral temporal cortex, MTC: Mesial temporal cortex, OC: Occipital cortex, FC: Frontal cortex, PC: Parietal cortex, BG: Basal ganglia, GCA: Anterior cingulate cortex, GCP: Posterior cingulate cortex
Composite [18F]florbetaben PET standardized uptake value ratios (SUVRs) of patients with Alzheimer’s disease (AD) and healthy controls (HC) obtained by the conventional manual and four tested automated procedures. Box plots (median, 25 % and 75 % quartile) with whiskers at highest/lowest value within the1.5 * inter-quartile range of the closest quartile and points for identified outliers. p: p value in Mann-Whitney U test, d: Cohen’s d
To investigate the potential of individual composite SUVRs as obtained by the different VOI definition methods to discriminate between the AD patients and the HCs, respective cut-off values were defined by receiver operating characteristic (ROC) analyses. The resulting discrimination parameters are provided in Table 3: For all automated VOI definition methods, sensitivities (80 % for all analyses), specificities (range: 80 % - 100 %), and the area under the ROC curve (AUC) (range: 0.79 – 0.83) were similar to those of the conventional method.
The inter-rater results between the different automated and the conventional VOI definition method are also provided in Table 3. Here, very high inter-rater reliability (Cohen’s kappa ≥ 0.8) was observed for all methods.
The regional SUVRs of most VOIs as obtained by the different automated anatomical VOI definition methods were strongly correlated with those of the conventional VOI definition method (Electronic Supplementary Material 1). The SUVRs as obtained by PMOD Maximum Probability correlated significantly with those obtained by the conventional method in 95 % of the VOIs analyzed. For FreeSurfer, HERMES Brass and PMOD Normalization these portions were 92 %, 88 %, and 76 % (Electronic Supplementary Material 1). Closest SUVR correlations with the values of the conventional method were observed in the frontal cortex (Pearson’s r=0.96; p<0.0001 (l) / r=0.93; p<0.0001 (r); PMOD Maximum Probability), posterior cingulate cortex (r=0.95; p<0.0001 (l) / r=0.94; p<0.0001 (r); FreeSurfer) and lateral temporal cortex (r=0.95; p<0.0001 (r) / r=0.92; p<0.0001 (l); PMOD Maximum Probability). The SUVRs in these VOIs together with those SUVRs of the anterior cingulate cortex (r/l), parietal cortex (r/l), occipital cortex (r/l), and putamen (r/l) consistently showed significant correlations with those obtained by the conventional method in all automated analysis methods. This was also the case for the composite SUVRs (Fig. 4).
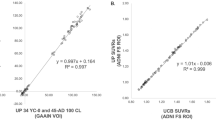
Correlations of the composite [18F]florbetaben PET standardized uptake value ratios (SUVRs) between different automated neuroanatomical definition methods and the conventional standard method for Alzheimer’s disease (AD) patients and healthy controls (HC). Grey lines represent regression lines with 95 % confidence interval. Correlation coefficients are expressed as Pearson’s r with its respective p values
Associating the regional SUVRs as obtained by the different automatic VOI definition methods with those of the conventional method separately for the AD patients and the HCs revealed an additional feature: When comparing the Pearson’s r values between the two groups on a VOI leveI, the correlation with SUVRs as obtained with the conventional method was better in AD patients than in HCs in the majority of regions (Table 4): For the PMOD Maximum Probability tool this was true for 82 % of the tested VOIs. The respective portions for the HERMES Brass, FreeSurfer, and PMOD Normalization algorithms were 75 %, 75 %, and 67 %, respectively. The absolute amount of VOIs in which the automatically obtained SUVRs were significantly correlated to the conventional SUVRs was higher in AD patients compared to HCs for the three algorithms HERMES Brass, PMOD Maximum Probability and FreeSurfer, the opposite was the case for the PMOD Normalization algorithm.
Discussion
This work aimed at evaluating different software tools for automated neuroanatomical VOI definition of [18F]florbetaben amyloid PET data. Discriminative power of the resulting SUVRs to differentiate AD patients from age-matched healthy controls was compared with that of the current gold standard manual VOI definition approach.
Automated neuroanatomical VOI definition approaches have been broadly applied in prior β-amyloid PET research projects [25–29]. However, there is a lack of studies systematically evaluating a set of fundamentally different software algorithms against the standard method of conventional VOI definition on the same data. To our knowledge, comparative studies in the field of β-amyloid PET imaging included only a maximum of two basically different automated analysis methods so far [25, 29, 30]. A subset of recently published studies evaluated automated analysis methods against other automated algorithms [28, 31] and against visual read [32], but not against the current gold-standard approach of manual VOI definition.
In general, relevant differences in tracer uptake between the two groups (in favor of the AD patients) and high agreement between manually and automatically derived SUVRs were reported. With regard to the different software tools currently available for neuroanatomical VOI definition, in prior partly non-amyloid-related nuclear brain imaging research, good to excellent accordance between automatically and manually generated VOIs was reported for SPM-based procedures [26, 27, 33], FreeSurfer applications [23, 34, 35] and HERMES Brass [18, 36, 37]. In accordance with that, we observed reliable group discrimination between AD patients and HCs and significant correlation between the automatically and conventionally derived SUVRs for all software tools tested for the composite and for most regional SUVRs. The composite SUVR effect sizes were in three of the four tested automated approaches only slightly lower than that of manual VOI definition approach, while they were even higher for the PMOD Maximum Probability tool. As one promising result, we showed one automated algorithm (PMOD Maximum Probability) to reach 100 % classificational accord and equal diagnostic accuracy compared to manual VOI definition of the tested dataset.
Also, in concordance with the findings in the literature, our testing revealed high or very high degrees of correlation between the SUVRs obtained by manual and automatic procedures in the majority of brain regions. Most of the cited studies, however, compared either with a partly automatically generated conventional VOI set [34] or worked with a large volume or limited VOI set [33, 35]. Lastly, some of the reported studies involved an excessive parameter setting refinement [18, 36]. As we aimed to compare automatically generated VOIs against conventional native-space VOIs in a clinical routine default parameter environment this could explain eventual lower degrees of correlation in our testing.
Two general tendencies related to the quality of automated neuroanatomical VOI identification were observed in our project: (1) Overall correlation to the SUVRs obtained by manual VOI definition was better in AD patients than in HC in all tested tools. (2) In deep brain structures the two algorithms with higher computational effort (PMOD Maximum Probability and FreeSurfer), which include segmentation techniques, showed higher correlation to the results of manual VOI definition than the algorithms which rely only on normalization to spatial space. Statistically significant correlation to conventionally derived SUVRs (p<0.01) was found in caudate nucleus, thalamus and putamen of both hemispheres for the PMOD Maximum Probability and FreeSurfer algorithms. In contrast, no statistically significant correlation with SUVRs as obtained with the conventional method could be demonstrated for caudate nucleus (r/l, PMOD Normalization, HERMES Brass) and thalamus (r, PMOD Normalization) using the normalization-based algorithms (Electronic Supplementary Material 1). Of interest, in cortical VOIs which are commonly used in β-amyloid PET data quantification, the correlation to conventionally derived SUVRs was less affected by the underlying algorithm than in deep brain structures.
One aspect which could explain the better correlation with manually derived SUVRs in AD patients might be a wider spreading of uptake values compared to the HC subjects. As the finding of better correlation in AD patients than in HCs was most prominent for the PMOD Maximum Probability tool, this algorithm might be the method of choice in case of deviations from standard neuroanatomy on the regional level. However, we identified all four automated procedures to work reliably even under the pathological anatomical conditions encountered in AD, at least for neocortical brain structures.
Several constraints of the automated procedures were observed during the conduct of this study, which require consideration for future application: (1) We observed unexpected statistically significant differences in tracer uptake between the left and right hemisphere in three of four software tools, which were not observed in manual SUVR analysis. This finding was more prominent in the two tools based on spatial normalization (PMOD Normalization and HERMES Brass) compared to the more computational effort requiring tools (FreeSurfer and PMOD Maximum Probability). However, no hemispherical predominance was observed rendering this finding coincidental. Furthermore, (2) brain regions with highest and lowest SUVR correlation with the results of the manual analysis were not totally consistent across and within the software tools. Although multiple-region-conjunct analysis offered stable correlation to manual uptake values, our work showed single-region comparisons to be still more prone to software-inherent deviations. From this, it is evident that future implementation of automated procedures as routine procedure will need continued careful evaluation against gold standard methods.
As a limitation of this study, the sample size analyzed was limited. We decided to not extend the sample size in this project, as in the subsequent florbetaben trials following the phase 0 study of which the data were used in this investigation, since the PET acquisition time-point was modified (from 70-90 min p.i. to 90-110 min p.i.). As known from investigating tracer dynamics [13], we cannot exclude an influence on the study results by this modification. However, more work with PET and MRI data of other clinical trials and/or as obtained in clinical routine, potentially also including other amyloid tracers and other-quality MRI data, is required to fully uncover the potential of the available automated neuroanatomical VOI definition approaches in this context.
Regarding the clinical usability, the tool with highest impact on time saving, economic effectiveness, and analyzer convenience might be the modality of choice. Keeping this in mind, automated analysis of β-amyloid PET images without the need of acquiring additional MR images would be able to provide faster and cheaper diagnosis than double-modality workflows given that diagnostic accuracy would not be severely affected. Prior research has addressed this topic by comparing the performance of MRI templates and PET templates in automated image analysis. Even though MRI-based approaches were reported to be advantageous in terms of quality in small region comparisons [30], a subset of studies indicated reliable performance of tracer-specific PET templates [28, 31, 33, 38–40]. These findings are in good agreement with the results of this work: Composite SUVR analysis with the PET-based tool HERMES Brass had only minimally lower diagnostic accuracy than both conventional SUVR analysis and the best performing automated program. The correlations between the data obtained by the HERMES Brass software with those of the manual VOI definition approach were slightly less strong than the correlations for the other algorithms. However, this MR-less normalization algorithm tool showed accurate AD vs. HC group discrimination, rendering it an interesting solution, at least in cases in which individual MRI data are not available.
Conclusion
In this [18F]florbetaben PET analysis of AD patient and age-matched HC data, all tested software tools for automated neuroanatomical VOI definition revealed results very similar to those of the current gold standard, the manual approach. While the diagnostic potential of the composite SUVRs was to main parts not dependent on the particular software approach employed, slight differences in the AD vs HC discrimination by the obtained regional SUVRs as well as in the degree of correlation between the SUVRs of the automatic and manual approaches were observed. Taken together, regardless of whether individual MRI data are available, there is a great potential for automated neuroanatomical VOI definition tools to simplify and objectivize regional and global β-amyloid PET quantification.
References
White DR, Houston AS, Sampson WF, Wilkins GP. Intra- and interoperator variations in region-of-interest drawing and their effect on the measurement of glomerular filtration rates [eng]. Clin Nucl Med. 1999;24(3):177–81.
Destrieux C, Fischl B, Dale A, Halgren E. Automatic parcellation of human cortical gyri and sulci using standard anatomical nomenclature. NeuroImage. 2010;53(1):1–15.
Evans AC, Marrett S, Neelin P, Collins L, Worsley K, Dai W, et al. Anatomical mapping of functional activation in stereotactic coordinate space. NeuroImage. 1992;1(1):43–53.
Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N, et al. Automated Anatomical Labeling of Activations in SPM Using a Macroscopic Anatomical Parcellation of the MNI MRI Single-Subject Brain. NeuroImage. 2002;15(1):273–89.
Fischl B, Salat DH, Busa E, Albert M, Dieterich M, Haselgrove C, et al. Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain [eng]. Neuron. 2002;33(3):341–55.
Balafar MA, Ramli AR, Saripan MI, Mashohor S. Review of brain MRI image segmentation methods. Artif Intell Rev. 2010;33(3):261–74.
Ashburner J, Friston KJ. Nonlinear spatial normalization using basis functions [eng]. Hum Brain Mapp. 1999;7(4):254–66.
Collins DL, Neelin P, Peters TM, Evans AC. Automatic 3D intersubject registration of MR volumetric data in standardized Talairach space [eng]. J Comput Assist Tomogr. 1994;18(2):192–205.
Ashburner J, Friston KJ. Unified segmentation [eng]. Neuroimage. 2005;26(3):839–51.
Fischl B. Automatically parcellating the human cerebral cortex. Cereb Cortex. 2004;14(1):11–22.
Barthel H, Seibyl J, Sabri O. The role of positron emission tomography imaging in understanding Alzheimer’s disease [eng]. Expert Rev Neurother. 2015;15(4):395–406.
Barthel H, Luthardt J, Becker G, Patt M, Hammerstein E, Hartwig K, et al. Individualized quantification of brain β-amyloid burden: results of a proof of mechanism phase 0 florbetaben PET trial in patients with Alzheimer’s disease and healthy controls. Eur J Nucl Med Mol Imaging. 2011;38(9):1702–14.
Becker GA, Ichise M, Barthel H, Luthardt J, Patt M, Seese A, et al. PET Quantification of 18F-Florbetaben Binding to -Amyloid Deposits in Human Brains. J Nucl Med. 2013;54(5):723–31.
D’Agostino E, Maes F, Vandermeulen D, Suetens P. A viscous fluid model for multimodal non-rigid image registration using mutual information [eng]. Med Image Anal. 2003;7(4):565–75.
Slomka PJ, Hurwitz GA, Stephenson J, Cradduck T. Automated alignment and sizing of myocardial stress and rest scans to three-dimensional normal templates using an image registration algorithm [eng]. J Nucl Med. 1995;36(6):1115–22.
Slomka PJ, Radau P, Hurwitz GA, Dey D. Automated three-dimensional quantification of myocardial perfusion and brain SPECT [eng]. Comput Med Imaging Graph. 2001;25(2):153–64.
Radau PE, Linke R, Slomka PJ, Tatsch K. Optimization of automated quantification of 123I-IBZM uptake in the striatum applied to parkinsonism [eng]. J Nucl Med. 2000;41(2):220–7.
Radau PE, Slomka PJ, Julin P, Svensson L, Wahlund LO. Evaluation of linear registration algorithms for brain SPECT and the errors due to hypoperfusion lesions [eng]. Med Phys. 2001;28(8):1660–8.
Hammers A, Allom R, Koepp MJ, Free SL, Myers R, Lemieux L, et al. Three-dimensional maximum probability atlas of the human brain, with particular reference to the temporal lobe. Hum Brain Mapp. 2003;19(4):224–47.
Fischl B. FreeSurfer. NeuroImage. 2012;62(2):774–81.
Fischl B, Sereno MI, Dale AM. Cortical surface-based analysis. II: inflation, flattening, and a surface-based coordinate system [eng]. NeuroImage. 1999;9(2):195–207.
Dale AM, Fischl B, Sereno MI. Cortical surface-based analysis. I. Segmentation and surface reconstruction [eng]. NeuroImage. 1999;9(2):179–94.
Desikan RS, Ségonne F, Fischl B, Quinn BT, Dickerson BC, Blacker D, et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. NeuroImage. 2006;31(3):968–80.
Rowe CC, Ackerman U, Browne W, Mulligan R, Pike KL, O’Keefe G, et al. Imaging of amyloid β in Alzheimer’s disease with 18F-BAY94-9172, a novel PET tracer: proof of mechanism. Lancet Neurol. 2008;7(2):129–35.
Schain M, Varnäs K, Cselényi Z, Halldin C, Farde L, Varrone A. Evaluation of two automated methods for PET region of interest analysis [eng]. Neuroinformatics. 2014;12(4):551–62.
Aalto S, Scheinin NM, Kemppainen NM, Någren K, Kailajärvi M, Leinonen M, et al. Reproducibility of automated simplified voxel-based analysis of PET amyloid ligand [11C]PIB uptake using 30-min scanning data. Eur J Nucl Med Mol Imaging. 2009;36(10):1651–60.
Rosario BL, Weissfeld LA, Laymon CM, Mathis CA, Klunk WE, Berginc MD, et al. Inter-rater reliability of manual and automated region-of-interest delineation for PiB PET. NeuroImage. 2011;55(3):933–41.
Hutton C, Declerck J, Mintun MA, Pontecorvo MJ, Devous MD, Joshi AD. Quantification of (18)F-florbetapir PET: comparison of two analysis methods [ENG]. Eur J Nucl Med Mol Imaging. 2015;42(5):725–32.
Landau SM, Thomas BA, Thurfjell L, Schmidt M, Margolin R, Mintun M, et al. Amyloid PET imaging in Alzheimer’s disease: a comparison of three radiotracers [eng]. Eur J Nucl Med Mol Imaging. 2014;41(7):1398–407.
Edison P, Carter S, Rinne J, Gelosa G, Herholz K, Nordberg A, et al. Comparison of MRI based and PET template based approaches in the quantitative analysis of amyloid imaging with PIB-PET. NeuroImage. 2013;70:423–33.
Saint-Aubert L, Nemmi F, Péran P, Barbeau EJ, Payoux P, Chollet F, et al. Comparison between PET template-based method and MRI-based method for cortical quantification of florbetapir (AV-45) uptake in vivo [eng]. Eur J Nucl Med Mol Imaging. 2014;41(5):836–43.
Thurfjell L, Lilja J, Lundqvist R, Buckley C, Smith A, Vandenberghe R, et al. Automated quantification of 18F-flutemetamol PET activity for categorizing scans as negative or positive for brain amyloid: concordance with visual image reads [eng]. J Nucl Med : Off Publ Soc Nucl Med. 2014;55(10):1623–8.
Lundqvist R, Lilja J, Thomas BA, Lotjonen J, Villemagne VL, Rowe CC, et al. Implementation and validation of an adaptive template registration method for 18F-flutemetamol imaging data. J Nucl Med. 2013;54(8):1472–8.
Su Y, D’Angelo GM, Vlassenko AG, Zhou G, Snyder AZ, Marcus DS, et al. Quantitative Analysis of PiB-PET with FreeSurfer ROIs. PLoS ONE. 2013;8(11):e73377.
Barnes J, Foster J, Boyes R, Pepple T, Moore E, Schott J, et al. A comparison of methods for the automated calculation of volumes and atrophy rates in the hippocampus. NeuroImage. 2008;40(4):1655–71.
Pöpperl G, Radau P, Linke R, Hahn K, Tatsch K. Diagnostic performance of a 3-D automated quantification method of dopamine D2 receptor SPECT studies in the differential diagnosis of parkinsonism [eng]. Nucl Med Commun. 2005;26(1):39–43.
Koch W, Radau PE, Hamann C, Tatsch K. Clinical testing of an optimized software solution for an automated, observer-independent evaluation of dopamine transporter SPECT studies [eng]. J Nucl Med. 2005;46(7):1109–18.
Fripp J, Bourgeat P, Raniga P, Acosta O, Villemagne V, Jones G, et al. MR-less high dimensional spatial normalization of 11C PiB PET images on a population of elderly, mild cognitive impaired and Alzheimer disease patients [eng]. Med Image Comput Comput Assist Interv. 2008;11(Pt 1):442–9.
Kuhn FP, Warnock GI, Burger C, Ledermann K, Martin-Soelch C, Buck A. Comparison of PET template-based and MRI-based image processing in the quantitative analysis of C11-raclopride PET. EJNMMI Res. 2014;4(1):7.
Meyer JH, Gunn RN, Myers R, Grasby PM. Assessment of spatial normalization of PET ligand images using ligand-specific templates [eng]. NeuroImage. 1999;9(5):545–53.
Acknowledgments
We would like to thank the statistical counselling service of IMISE, University of Leipzig for their support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This project received research support by Piramal Imaging, HERMES Medical solutions, and PMOD Technologies Ltd.
Conflict of interest
OS and HB received speaker and consulting honoraria from Piramal Imaging and Siemens Healthcare. MR received travel expenses from Piramal Imaging. SH received travel grants and honoraria from General Electric (GE) Healthcare and Bayer Schering Pharma.
Ethical approval
All procedures performed involving human participants were in accordance with the ethical standards of the 1964 Helsinki declaration and its later amendments and with the standards of the institutional and national research committee. This article does not contain any studies with animals performed by any of the authors.
Informed consent
Written informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 290 kb)
Rights and permissions
About this article
Cite this article
Tuszynski, T., Rullmann, M., Luthardt, J. et al. Evaluation of software tools for automated identification of neuroanatomical structures in quantitative β-amyloid PET imaging to diagnose Alzheimer’s disease. Eur J Nucl Med Mol Imaging 43, 1077–1087 (2016). https://doi.org/10.1007/s00259-015-3300-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-015-3300-6