Abstract
Milbemycins are used commercially as insect repellents and acaricides; however, their high cost remains a significant challenge to commercial production. Hence, improving the titer of milbemycins for commercial application is an urgent priority. The present study aimed to effectively increase the titer of milbemycins using a combination of genome re-sequencing and metabolic engineering. First, 133 mutation sites were identified by genome re-sequencing in the mutagenized high-yielding strain BC04. Among them, three modifiable candidate genes (sbi_04868 encoding citrate synthase, sbi_06921 and sbi_06922 encoding alpha and beta subunits of acetyl-CoA carboxylase, and sbi_04683 encoding carbon uptake system gluconate transporter) related to primary metabolism were screened and identified. Next, the DNase-deactivated Cpf1–based integrative CRISPRi system was used in S. bingchenggensis to downregulate the transcription level of gene sbi_04868. Then, overexpression of the potential targets sbi_06921-06922 and sbi_04683 further facilitated milbemycin biosynthesis. Finally, those candidate genes were engineered to produce strains with combinatorial downregulation and overexpression, which resulted in the titer of milbemycin A3/A4 increased by 27.6% to 3164.5 mg/L. Our research not only identified three genes in S. bingchenggensis that are closely related to the production of milbemycins, but also offered an efficient engineering strategy to improve the titer of milbemycins using genome re-sequencing.
Key points
• We compared the genomes of two strains with different titers of milbemycins.
• We found three genes belonging to primary metabolism influence milbemycin production.
• We improved titer of milbemycins by a combinatorial engineering of three targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The main secondary metabolites produced by Streptomyces are economically valuable in medicine, veterinary, and agriculture (Bérdy 2004). Since secondary metabolites are not essential for cell growth, during evolution, the yields of secondary metabolites produced by cells become low and are only required to satisfy their special physiological functions (Pickens et al. 2011). However, this low output does not reach the requirements for industrialization.
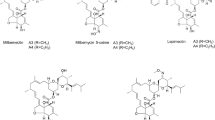
Milbemycins, produced by Streptomyces bingchenggensis, are used commercially as insect repellents and acaricides in agriculture and animal husbandry (Hayes et al. 2015; Monod et al. 2019; Wang et al. 2014). Milbemycins are similar to avermectins, doramectins, and moxidectins in structure, sharing a similar lactone ring (Takiguchi et al. 1983). Notably, milbemycin A3/A4 and their derivatives have advantages of higher acaricidal activity and lower animal toxicity compared with those of avermectins, which rank first in terms of sales. Nevertheless, the low extraction yield and high production price restrict the wide application of milbemycins (Merola and Eubig 2018).
Initially, the biosynthetic pathway of milbemycin was proposed based on physicochemical characterization, bioconversion data, and some studies on the biosynthesis of avermectin and meilingmycin (He et al. 2010; Nonaka et al. 1999a, b). Derived from acetate or propionate, 7 malonyl-CoA units and 5 methylmalonyl-CoA units are condensed to form the starting unit, catalyzed by the polyketide synthases (PKSs) to form the polyketide backbone, in a stepwise process (Fig. 1). During the past 10 years, many efforts have been made to increase the titer of milbemycin A3/A4, such as random mutations (Wang et al. 2014, 2009), removing analogous by-products (Wang et al. 2020a; Zhang et al. 2013), transcription regulator engineering (He et al. 2018; Zhang et al. 2016), as well as transporter engineering (Jin et al. 2020). Moreover, using an avermectin high-producing strain as a heterologous host could also enhance production of milbemycin A3/A4 (Kim et al. 2017). Nevertheless, the current yield of milbemycins in S. bingchenggensis is much lower than that of avermectin B1a (Wang et al. 2020b). In recent years, rapid progress has been made in sequencing technology, which has enabled genomic and post-genomic technologies to further promote strain improvement using genetic engineering methods (Palazzotto et al. 2019; Peano et al. 2014; Dai and Nielsen 2015). The combination of omics data and known information can help to further recognize and identify gene targets for priority modification at the transcriptional and metabolic levels (Doroghazi and Metcalf 2013; Hu et al. 2011; Illeghems et al. 2013; Wang et al. 2018). Meanwhile, because of its low price, genomic re-sequencing has also been widely applied in research to complete the analysis of different species together with reference genomic data. Increasing numbers of potentially valuable mutated genes have been identified using this method. Together with metabolic engineering, this technique can be used to modify mutations or transfer them to other host strains to obtain desired strains (Duan et al. 2017; Zhang et al. 2019). Hence, re-sequencing the high-yielding mutated strain and fully mining the mutated genes might be a promising strategy for rational metabolic engineering of S. bingchenggensis for titer, and therefore, yield increase of milbemycins.
In the present work, we obtained previously unidentified targets based on genome re-sequencing and implemented and engineered these targets to improve the titer of milbemycin A3/A4 (Fig. 2). First, we re-sequenced the genome of the high-yielding mutant strain BC04 and obtained 133 mutation sites. Canonical pathways and enrichment analysis by Kyoto Encyclopedia of Genes and Genomes (KEGG) showed that the proteins encoded by the mutated genes of BC04 were distributed in different metabolic pathways, among which primary metabolic pathways accounted for 58.14%. Then, we focused on the mutant genes enriched in the primary metabolic pathways that provide milbemycin biosynthesis precursors. Finally, three targets (citrate synthase (GltA), acetyl-CoA carboxylase (ACC), and carbon uptake system gluconate transporter (GntP)) related to primary metabolic pathways were downregulated using the CRISPRi system or overexpressed to increase the titer of milbemycin A3/A4. This work provided new targets to redirect metabolic fluxes from primary metabolism to the milbemycin biosynthetic pathway for yield improvement. This genome re-sequencing–based metabolic engineering strategy showed great potential to obtain high-yielding strains of milbemycins in S. bingchenggensis.
Materials and methods
Bacterial strains and growth conditions
All bacterial strains and plasmids used in this study are listed in Table S1. S. bingchenggensis BC-101-4, the producer of milbemycin A3/A4, has been deposited at the China General Microbiology Culture Collection Center (accession no. CGMCC1734). S. bingchenggensis BC04 was derived from S. bingchenggensis BC-101-4 by random mutagenesis using ultraviolet mutagenesis and N-methyl-N’-nitroso-N-nitrosoguanidine (NTG) mutagenesis (Zhang et al. 2016). For sporulation and conjugation, S. bingchenggensis strains were grown on SKYM (0.4% sucrose, 0.1% skimmed milk powder, 0.2% yeast extract, and 0.5% malt extract pH 7.2) agar plate and mannitol soya flour (MS) agar medium at 28 °C (Wang et al. 2014; Kieser et al. 2000). The media and methods used for milbemycin production were the same as those reported previously (Zhang et al. 2016; He et al. 2018). All Escherichia coli strains were grown at 37 °C on Luria-Bertani (LB) agar (Russell and Sambrook 2001) or in LB liquid.
Genome re-sequencing
The genomic DNA of S. bingchenggensis BC04 was extracted, purified, and randomly fragmented. DNA fragments of approximately (~500 bp) were collected, and a single A base was added to the 3′ end of these fragments and then used to ligate the fragments into the library plasmid via an overhanging T base. The 500-bp genomic library of S. bingchenggensis BC04 was thus prepared. Then, the library was sequenced by paired-end 125 (PE125) sequencing using a HiSeq 2000 instrument (Illumina, San Diego, CA, USA).
Genome annotation and bioinformatic analysis
The alignment of reads is the basis of re-sequencing analysis. The information obtained from the alignment between the reads from the sample and the specified reference sequence was used to analyze the differences between the sample and the reference sequence. Reads were compared to the reference sequence using the BWA software. The re-sequencing quality of S. bingchenggensis BC04 is shown in Table S2. Compared with the reference strain BC-101-4 (GenBank: CP002047), the coverage of the reference genome BC-101-4 and BC04 was compared by BWA and SAMTOOLS to determine their relationship (Li 2014; Li et al. 2009). The statistics are shown in Table S3. The sequence similarity between the BC04 sequencing data and the reference sequence was 99.7%. Meanwhile, the 20× coverage of the BC04 genome was 91.6% compared with that of the BC-101-4 genome (Fig. S1). To validate that the mutations truly exist in the BC04 genome, mutation sites were randomly selected and conducted by PCR amplification and Sanger sequencing, which produced consistent results with those of the whole genome re-sequencing (data not shown). For functional annotation, the assembled unigenes that could possibly encode proteins were used as search queries against the nr (http://www.ncbi.nlm.nih.gov/), SWISS-PROT (http://www.expasy.ch/sprot/), KEGG (http://www.genome.jp/kegg/), and COG (http://www.ncbi.nlm.nih.gov/cog/) databases using the BLASTX algorithm.
Construction of recombinant strains
Gene disruption experiments in S. bingchenggensis were performed as described previously (Zhang et al. 2016). To construct the gltA null mutant, the 2177-bp upstream and 2021-bp downstream arms flanking gltA (sbi_04868) were amplified from the genome of S. bingchenggensis BC-101-4 and BC04 using KOD plus polymerase and the primers listed in Table S4. The kanamycin-resistant gene (neo) was obtained using the plasmid pUC119::neo by PCR amplification. The E. coli-Streptomyces shuttle vector pKC1139 was used to construct recombinant plasmids for gene disruption, which contains a temperature-sensitive origin of replication from pSG5 (Bierman et al. 1992). The plasmid pKCgltAneo was generated by three-piece Gibson assembly with the pKC1139 backbone (digested with HindIII and XbaI) and homologous arms using ClonExpress MultiS (Vazyme Biotech Co, Ltd, Nanjing, China) according to the manufacturer’s instructions. Then, pKCgltAneo was introduced into the S. bingchenggensis strains BC-101-4 and BC04 using conjugal transfer.
pSET152 was used to create recombinant plasmids, which can integrate into the Streptomyces chromosome by site-specific recombination at the phage ФC31 (Bierman et al. 1992). Based on pSETddCpf1 constructed in a previous study (Li et al. 2018), we constructed CRISPRi ddCpf1 plasmids by introducing a 23-nt gene-specific spacer sequence into the crRNA scaffold of pSETddCpf1. In brief, using pSETddCpf1 as the template, the crRNA expression cassettes were amplified using the ddCpf1-F primer 5′-CACTAGTN23ATCTACAACAGTAGAAATTTGG-3′ (N23 represents the 23-nt gene-specific spacer sequence) and the ddCpf1-R primer (crRNA-rev). The PCR products of sgRNAgltA1 and sgRNAgltA2 were digested with NdeI and SpeI and ligated to NdeI/SpeI–digested pSETddCpf1. The primers used are listed in Table S4 in the supplementary material. This yielded pSETddCpf1::gltAsgRNA1 and pSETddCpf1::gltAsgRNA2, which were introduced into the S. bingchenggensis strains BC-101-4 and BC04 using conjugal transfer.
The 500-bp promoter region upstream of gene sbi_06921-06922 amplified from the genome of BC-101-4 and BC04 of S. bingchenggensis by primer pair Pacc-F/R and Pmacc-F/R were cloned to generate Pacc and Pmacc. The plasmid with the reporter green fluorescent (sfgfp) was cloned from plasmid pIJ-Potr using primers PIJ-F/R (Wang et al. 2016). The PCR product was ligated with Pacc and Pmacc by Gibson assembly to obtain Pacc-GFP and Pmacc-GFP. Strains BpaccG and BpmaccG were obtained by integrated Pacc-GFP and Pmacc-GFP into the genome of BC-101-4, respectively. The fragment containing the coding region of alpha and beta subunits of acc (sbi_06921 and sbi_06922) and the upstream region 500-bp (Pmacc) was amplified from BC04 genomic DNA using primers ACCs-F/R and then cloned into pSET152 using Gibson assembly, generating pSET152::ACC. The integrative shuttle vector pSET152:: ACC was introduced into appropriate S. bingchenggensis BC-101-4 and BC04 strains to obtain BCACC and B4ACC, respectively. Similarly, the fragment containing the coding region of gntP (sbi_04683) was amplified from the BC-101-4 and BC04 genomes using primers gntPkasO*-F/R. The fragments containing the KasO* promoter was amplified using the primers listed in Table S4, and then the plasmid pSET152::PkasO*gntP was generated by three-piece Gibson assembly with the pSET152 backbone digested with EcoRI and XbaI and homologous arms using ClonExpress MultiS according to the manufacturer’s instructions. The plasmid pSET152::PkasO*gntP was introduced into appropriate S. bingchenggensis BC-101-4 and BC04 strains by intergeneric conjugation. The coding sequences (CDS) region of sbi_04683, the fragment containing PkasO* and the fragment containing sbi_06921-06922 under the control of Pmacc promoter, was connected with integrative shuttle vector pIJ10500::PkasO*gntP::ACC by Gibson assembly. Then, the plasmid was introduced using intergeneric conjugation into strain B4gltAsgRNA1, which comprised pSETddCpf1::gltAsgRNA1 in strain S. bingchenggensis BC04.
RNA isolation and quantitative real-time PCR assay
As previously described, RNAs were isolated from S. bingchenggensis strains grown at 28 °C at 2, 3, and 6 days for experiments (Wang et al. 2013). The quality and concentration of RNA were determined by agarose gel electrophoresis and UV spectroscopy. As previously mentioned, quantitative real-time PCR (qRT-PCR) analysis was performed (Zhang et al. 2016). The primers used are listed in Table S4.
HPLC analysis of milbemycin A3/A4 production and residual sugar measurement
Milbemycins were analyzed using high-performance liquid chromatography (HPLC) as described previously (Wang et al. 2014). HPLC was performed using an Agilent 1260 Infinity II LC system (Agilent Technologies Inc., Beijing, China) using a Zorbax SB-C18 column (4.6 mm × 250 mm, 5 μm, Agilent) at a flow rate of 1.0 mL/min with a linear gradient from 0 to 100 % of solvent B in 15 min (solvent A: acetonitrile-H2O-methanol (350:50:100, v/v/v); solvent B: methanol), and milbemycins were detected at 242 nm.
For residual sugar measurement, the fermentation broth of S. bingchenggensis and its derivatives were extracted using two volumes of methyl alcohol. Residual sugars were quantified using HPLC. HPLC was performed using a Shimadzu LC-20AT (Zorbax, Carbohydrate column, 4.6 mm × 250 mm, 5 μm) at a flow rate of 1.0 mL/min 80% acetonitrile and detected using a refractive index detector.
Extraction and tandem mass spectroscopy (LC-MS/MS) analysis of intracellular acyl-CoA esters
LC-MS/MS was used to determine the concentrations of intracellular acyl-CoA esters in cells as described previously (Lu et al. 2016). The mycelia were washed rapidly with ice-cold 30% (v/v) methanol, then washed twice with ice-cold water, and then centrifuged at 12,000 rpm at 4 °C for 3 min. The washed mycelium was ground in liquid nitrogen and extracted with acetonitrile/methanol/0.1% glacial acetate (45:45:10, v/v/v) to a final volume of 1 mL at 20 °C by intermittent vortexing for 15 min. After centrifugation at 12,000 rpm, 4 °C, a Waters Acquity UPLC BEH C18 column (1.7 μm, 2.1 ×100 mm) was used to inject the supernatant for UPLC-MS/MS analysis (Waters XEVOTM TQ-S; Waters, Milford, MA, USA). At a flow rate of 300 μL/min, the analytes were eluted using a gradient from 2 to 35 % B (mobile phase A: 25 mM ammonium acetate in 0.5% glacial acetic acid, mobile phase B: 25 mM ammonium acetate in methanol) over 3 min. As previously described, acyl-CoAs were detected in the multiple reaction monitoring (MRM) mode (Lu et al. 2016).
Determination of cell dry weight
Two milliliters of cell culture were collected using vacuum filtration and dried at 55 °C to a constant weight to determine the dry cell weight.
Statistical analysis
All experiments were executed independently at least three times, and result was shown as mean value ± standard deviation (SD). Significance was analyzed by Student’s t-test. p < 0.05 is considered as a standard criterion of statistical significance.
Results
Phenotypic comparison of BC-101-4 and BC04 of S. bingchenggensis
BC04, a high-yielding milbemycin-producing mutant, was obtained after multiple rounds of random mutagenesis of S. bingchenggensis BC-101-4 using ultraviolet mutagenesis and N-methyl-N’-nitroso-N-nitrosoguanidine (NTG) mutagenesis (Zhang et al. 2016). As shown in Fig. 3a, in comparison with the parental strain S. bingchenggensis BC-101-4, on SKYM solid medium, BC04 cannot produce the yellow-green soluble pigment, but the inky hyphae could be observed. Meanwhile, the cell growth (Fig. S2), the milbemycin titer, and yield of two strains were quantitatively compared in fermentation medium. We observed that milbemycin B2 was significantly reduced compared to BC-101-4 (Fig. 3b), and the titer and yield of milbemycin A3/A4 were enhanced significantly (Fig. 3c); the titer increased from 1460.7 mg/L in BC-101-4 to 2480.0 mg/L in BC04. Furthermore, the ratio of milbemycin A3 and A4 was markedly different (Fig. S3). It is well known that acetyl-CoA and propionyl-CoA are used as the starting units of milbemycin A3 and A4, respectively (Fig. 1). Hence, we analyzed the concentrations of the four precursors (i.e., acetyl-CoA, malonyl-CoA, propionyl-CoA, and melthylmalonyl-CoA). The results showed that all of the precursors in strain BC04 were higher than those in BC-101-4 (Fig. 3d). In summary, the increase of precursors had important effect on the production and ratio of milbemycin A3/A4 in the high-yielding mutant BC04. Since primary metabolism is the main source of precursors for secondary metabolite biosynthesis, thus, identifying the mutant genes related to primary metabolism for further pathway engineering might be a promising way to achieve high-yielding milbemycin-producing strains.
Phenotype comparison between mutant strain BC04 and wild-type BC-101-4. a The phenotypes of BC-101-4 and BC04 on SKYM plates at 28 °C for 7days. b HPLC analysis of milbemycins. c Time course of milbemycin A3/A4 titer and yield of BC-101-4 and BC04. d Concentrations of acyl-CoAs in BC-101-4 and BC04 in different fermentation stages. Error bars depict the standard deviation of three independent experiments. Differences were analyzed by Student’s t-test. ***p < 0.001, **p < 0.01, *p < 0.05; “ns” means not significant.
Genome re-sequencing analysis of S. bingchenggensis BC04
Next, we re-sequenced the genome sequence of the mutant strain BC04 (Fig. 4a). As shown in Table 1, 80 single nucleotide polymorphisms (SNPs) and 53 insertions and deletions (InDels) were identified in the BC04 genome. Among them, 77 mutations were identified in open reading frame (ORF) regions (Table S5), including 50 nonsynonymous mutations causing the change of amino acids, which might affect the function of corresponding proteins. We also noted that 56 mutations of BC04 were located in intergenic regions and 31 of them were located in possible promoters (within 0.5 kb upstream of a start codon) and terminators (within 0.2 kb downstream of a stop codon) (Table S6).
Genome re-sequencing of BC04 and pathway specificity analysis of mutated genes. a Genomic distribution of SNPs and InDels of BC04. b KEGG pathways that were significantly enriched among the CDS mutant genes in BC04. c COG functional classification of BC04 CDS mutant genes. d Schematic diagram of some essential metabolic pathways in S. bingchenggensis. Three key nodes that were selected as the targets are shown in yellow. Cit, citric acid; Isocit, isocitrate; 2OG, 2-oxoglutarate; Suc CoA, succinyl-CoA; Suc, succinate; Fum, fumarate; Mal, malate; OAA, oxaloacetate
Then, the ORFs with mutations were classified according to the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis and the Clusters of Orthologous Groups (COG) protein database (Fig. 4b and c). The results showed that the mutated genes of BC04 were distributed in different metabolic pathways, among which primary metabolic pathways accounted for 58.14%. Meanwhile, we noted a SNP mutation (178 A>G) in the CDS region of C5-O-methyltransferase (encoded by sbi_00790, milD) catalyzing α-class (A3/A4) into C5-O-methylmilbemycins B2/B3, resulting the change from Lys47 to Glu. Notably, disruption of milD could achieve milbemycin A3/A4 titer improvement and the abolition of milbemycin B2/B3 and β1/β2 production (Zhang et al. 2013). Thus, in BC04, the loss of milbemycin B2 production was caused by mutation of milD.
As mentioned previously, BC04 and BC-101-4 showed differences in concentrations of cellular precursors for milbemycin biosynthesis (Fig. 3d). We thus focused on the mutant genes enriched in the primary metabolic pathways that provide these precursors. As shown in Table 2, four mutated genes were located on the key nodes in the carbon flux (Fig. 4d), such as sbi_04868 (encoding citrate synthase GltA), sbi_06507 (encoding NAD-glutamate dehydrogenase GdhB), sbi_04616 (encoding flavoprotein disulfide reductase FDR), and sbi_03212 (encoding arginine deiminase ADI). Sbi_04868 contained a SNP mutation, in which Thr126 was changed to Ile owing to the mutation of a C to a T. In fact, citrate synthase catalyzes the formation of citric acid from acetyl-CoA and oxaloacetic acid, which is the first step of the tricarboxylic acid (TCA) cycle involved in regulating the consumption of acetyl-CoA (Tian et al. 2020). It is known that GdhB links amino acid metabolism to the TCA cycle through the conversion of L-glutamate to 2-oxoglutarate via oxidative deamination (Sharkey et al. 2013). Sbi_06507 encodes 1663 amino acids, and it was affected by the InDel (2536_2537insC) mutation. The insertion of this site causes the protein to terminate expression after Ala590, which may cause the inactivation of GdhB. Thus, we speculated that mutations in gdhB might reduce the entry of acetyl-CoA into the TCA cycle, thereby increasing the titer of milbemycins from an elevated precursor pool. In addition to the two above genes, the other two mutated genes related to precursor pathways are sbi_04616 and sbi_03212. The pyridine nucleotide–dependent reduction of a variety of substrates can be catalyzed by FDR (encoded by sbi_04616) (Argyrou and Blanchard 2004), which is also involved in leucine, isoleucine, or valine degradation metabolism that is an important source of propionyl-CoA (Liu and Reynolds 2001). And ADI (encoded by sbi_03212) can convert arginine to ornithine and produce the NH3, CO2, and ATP (Abdelal 1979; Beck et al. 2019). However, their direct effect on antibiotic production has not been reported, and this may be the interesting targets in future work.
We also performed pathway enrichment analysis on genes affected by mutations in putative promoters and terminators in the BC04 intergenic regions. KEGG analysis showed that the proteins encoded by these mutated genes are also distributed in pathways associated with primary metabolism (Fig. S4), such as the acetyl-CoA carboxylase alpha and beta subunits coding genes sbi_06921 and sbi_06922, affected by an insertion of 16 bases (CTGCCTGTGGTTCTGG) in the promoter region, and the insertion position was at the 31st base (within upstream of the start codon) (Fig. S5a). It is known that ACC can catalyze the conversion from acetyl-CoA to malonyl-CoA. qRT-PCR results showed that sbi_06921-06922 transcription levels were upregulated in BC04, indicating that the yield of milbemycins may be affected by the mutation of this promoter region (Fig. S5b). As the research background of citrate synthase and acetyl-CoA carboxylase are relatively clear (Tao et al. 2017; Zabala et al. 2013), GltA and ACC could be precedently selected as potential targets to divert related precursor pools toward milbemycin biosynthesis.
Identification of mutated genes in the biosynthesis of acyl-CoA metabolic pathways of S. bingchenggensis
Based on previous analyses, citrate synthase gltA was the first to be recognized as candidate gene in the precursor pathway for milbemycin overproduction. However, the encoded protein of sbi_04868, GltA, is located at the key node of carbon metabolism; therefore, knockout of sbi_04868 decreased the growth of the strain (Fig. S6a and b). Hence, CRISPRi with sgRNA1 and sgRNA2 was employed to downregulate the expressions of gltA (Fig. 5a) (Tian et al. 2020). The highest milbemycin accumulation by sgRNA1 with its gltA gene repressed via CRISPRi was increased by approximately 26.9% and 6.2% compared with that by BC-101-4 and BC04 containing the CRISPRi vector without target, respectively (Fig. 5b). And the milbemycin yield of engineered strains were also increased compared with parental strains (Fig. S6c). To further characterize the effect of CRISPRi repression, we analyzed the transcription levels of gltA in strains using different sgRNAs. The transcript levels of gltA in BC-101-4 and BC04 with CRISPRi system using sgRNA1 were downregulated to 14.7–25.5% and 18.1–24.9%, respectively, and using sgRNA2, they were downregulated by 27.2–42.3% and 26.4–48.4%, respectively, compared with that of the control strain throughout the tested time course of 2, 3, and 6 days (Fig. S7). These results indicated that, compared with knockout strains, titer of milbemycins could be optimized by downregulating the transcription level of gltA to an appropriate level.
Identify modifiable metabolic pathways to enhance beneficial mutations for milbemycin titer improvement. a Design of the ddCpf1-mediated genome editing plasmid for gltA. Two strong promoters, PermE* and PkasO*, were used to drive the expression of ddCpf1 and crRNA, respectively. Schematic representation of the sgRNA target sites for the targeted genomic loci with yellow and blue lines for gltA. Sequence shows the sgRNA1 and sgRNA2 with their matching regions in the gene. b Influence of disruption and repression gltA on the titer of milbemycin A3/A4 in BC-101-4 and BC04. The titer of milbemycins was obtained at 9 days. BCDgltA and B4DgltA are strains in which gltA was disrupted by homologous recombination. BCddCpf1 and B4ddCpf1 have integrated the plasmid without the sgRNA of gltA. c Milbemycin A3/A4 titer in BC-101-4, BC04, and the alpha and beta subunits of acc overexpressing strains, BCACC and B4ACC. d Comparison of the concentrations of intracellular CoAs (6 days) between BC-101-4, BCgltAsgRNA1, BCACC, BC04, B4gltAsgRNA1, and B4ACC. Error bars depict the standard deviation of three independent experiments. Differences were analyzed by Student’s t-test. ***p< 0.001, **p < 0.01, *p < 0.05; “ns” means not significant.
Next targets were sbi_06921 and sbi_06922 (encoding the acetyl/propionyl–CoA carboxylase (ACC) alpha and beta subunits), in which mutation occurred in the promoter region. To further evaluate mutated promoter profile, we determined the time course transcriptional level of the reporter green fluorescent (sfgfp) using qRT-PCR. The mutated promoter showed better effect on transcriptional level of sfgfp than native promoter (Fig. S8a). Therefore, to increase the cellular concentration of malonyl-CoA, sbi_06921-06922 was overexpressed both in BC-101-4 and BC04 under the control of mutated promoter Pmacc yielding BCACC and B4ACC. The results showed that the titer of milbemycin A3/A4 was increased by 23.3% and 11.7% in BCACC and B4ACC, respectively (Fig. 5c), and the yield of milbemycin A3/A4 was increased by 27.6% and 11.5% (Fig. S8b). Based on the results, we further tested the effect of downregulating gltA and overexpressing acc (the alpha and beta subunits) on the concentrations of the intracellular acyl-CoAs in BC-101-4 and BC04 during the period of culture when milbemycins are largely produced (6 days). As shown in Fig. 5d, the overexpression of acc (the alpha and beta subunits) turned more acetyl-CoA into malonyl-CoA; however, the total of four intracellular acyl-CoAs showed no significant difference. Meanwhile, we observed that the concentrations of precursors were increased by varying degrees in gltA-downregulated strains. Among them, the increase of the concentrations of acetyl-CoA in BC-101-4 with gltA downregulated was greater than that in BC04 with gltA downregulated. This indicated that gltA was the beneficial mutation in BC04 and its mutation might weaken the TCA cycle and increase the metabolic flux of milbemycins. In all, GltA and ACC were adjustable targets in BC04 to increase malonyl-CoA levels to improve the titer of milbemycins.
Identification of mutated genes in the carbon source uptake system of S. bingchenggensis
Effective utilization of carbon sources and the redirection of metabolic flux toward secondary metabolism is a prerequisite for titer improvement. We thus focused on the mutated genes that might be responsible for carbohydrate transport. Six mutated genes were identified, and two of them, a tartrate transporter encoded by sbi_00454 and a putative gluconate transporter encoded by sbi_04683 (Fig. 6a), belonged to the predicted as carbon source uptake proteins in S. bingchenggensis (Jin et al. 2020). S. bingchenggensis uses sucrose as the main carbon source for milbemycin production in liquid medium, which does not contain tartaric acid. Thus, tartrate transporter does not qualify as a candidate gene for engineering. It is known that gluconate is produced from glucose through a simple dehydrogenation reaction catalyzed by a glucose oxidase; then, it is imported into the cell by the gluconate transporter (GntP) and phosphorylated to gluconate-6-phosphate by gluconokinase. Finally, it enters the pentose phosphate pathway (PP pathway) (Romero-Rodriguez et al. 2016; Avignone Rossa et al. 2002; Letek et al. 2006). Meanwhile, many transporters exhibit substrate promiscuity; gntP in E. coli was identified encoding a fructuronate transporter (Bates Utz et al. 2004). Hence, we overexpressed gntP (sbi_04683) with the strong promoter kasOp*. The results showed that the overexpression of gntP yielding BCgntP and B4gntP in BC-101-4 and BC04 improved the titer of milbemycin A3/A4, by 33.3% and 18.6%, respectively (Fig. 6b). And the yield of milbemycin A3/A4 was also increased by 37.3% and 19.4% in BCgntP and B4gntP (Fig. S9). Consequently, we tested the consumption rate of glucose and fructose; the consumption rates were increased by 33.2% and 23.7%, respectively, in B4gntP compared with those in BC04 (Fig. 6c and d). This showed overexpression of gntP ultimately enhanced the overall consumption of glucose and fructose. These results demonstrated that it was advantageous to optimize carbon flux toward the desired secondary metabolites in S. bingchenggensis.
Evaluation of the carbon source uptake system of the GntP transporter. a The number of identified carbon source uptake systems (orange) and other functions (gray) in the carbohydrate transport and metabolism pathway. b Increased milbemycin production in BCgntP and B4gntP compared with that in BC-101-4 and BC04 by gntP overexpression under the control of the kasO* promoter. c Comparison of the glucose consumption rate (144–216 h) between BC04 and B4gntP. d Comparison of the fructose consumption rate (144–216 h) between BC04 and B4gntP. For (b, c, and d), the data were obtained from three independent experiments. Error bars depict the standard deviation of three independent experiments. Differences were analyzed by Student’s t-test. ***p < 0.001, **p < 0.01, *p < 0.05; “ns” means not significant.
Reconstruction of a milbemycin overproducer
As described above, downregulation of the key node gltA of the TCA cycle pathway and overexpression of gntP and alpha and beta subunits of acc achieved enhanced milbemycin titers, respectively. To further engineer a high-yielding strain, we attempted to overexpress gntP and alpha and beta subunits of acc in strain B4gltAsgRNA1 (repressed for gltA expression) (Fig. 7a). As expected, we observed that the titer and yield of milbemycin A3/A4 were increased in engineered strain B4HS (Fig. 7b and S10a), which produced the highest milbemycin A3/A4 titer of 3164.5 mg/L. Meanwhile, the concentrations of precursors were measured at 6 days of fermentation, which showed that concentrations of intracellular acyl-CoAs were improved (Fig. S10b), which explains the increased titer of milbemycins. At the same time, this implied that the identification and genetic manipulation of key nodes regulating carbon flux through the metabolic network of central carbon metabolism can lead to an increase in the availability of precursors. The dramatically improved milbemycin production in these strains demonstrated that downregulation or overexpression of the targets identified through combination of genome re-sequencing and metabolic engineering strategies can efficiently and synergistically increase the production of target product (Fig. 7c). Finally, we increased milbemycin A3/A4 titer by 27.6% from 2480.0 mg/L in BC04 to 3164.5 mg/L in B4HS.
Milbemycin engineering in strain BC04. a Schematic representation of the overexpression acc (the alpha and beta subunits) under the control of Pmacc promoter and gntP under the control of the PkasO* promoter in high-yielding strain B4gltAsgRNA1. b Production of total milbemycins in B4HS and B4gltAsgRNA1 compared with those in BC04. c Schematic representation of the metabolic engineering rationale to enhance carbon flux into the milbemycin biosynthetic pathway. Candidate genes in the precursor and carbon transporter parts of the milbemycin biosynthetic pathway are visualized in red. The CRISPRi system repressive or inhibitory steps are indicated by green lines ending with a bar. Error bars depict the standard deviation of three independent experiments. Differences were analyzed by Student’s t-test. ***p < 0.001, **p < 0.01, *p< 0.05; “ns” means not significant.
Discussions
The main secondary metabolites produced by Streptomyces have important economic value. However, the low output does not reach the requirements of industrialization. The traditional random mutagenesis technique is still an important means of industrial breeding. For the overproduction of secondary metabolites, comparative genomic analysis of excellent producer strains with their progenitors allows the identification of beneficial mutations that could be combined with metabolic engineering to generate a producer strain with a high yielding using targeted modification (Conrad et al. 2009; Lee and Palsson 2010). Hence, re-sequencing the high-yielding mutated strain and fully mining the mutated genes was a promising strategy for rational metabolic engineering of S. bingchenggensis for titer and yield increase of milbemycins. In the present study, we applied genome re-sequencing and identified three targets (citrate synthase (GltA), acetyl-CoA carboxylase (ACC), and carbon uptake system gluconate transporter (GntP)), which related to primary metabolic pathways were downregulated using the CRISPRi system or overexpressed to increase the titer of milbemycins. Meanwhile, this is the first report to apply the CRISPERi technology in S. bingchenggensis. The results of the present study not only offered a comprehensive strategy to obtain high-yielding strains for milbemycin production, but also the mutations introduced in this study will provide a reference for subsequent research.
It should also be emphasized that precursor competition exists between the primary metabolic pathway (particularly central carbon metabolism) and the biosynthesis of natural products. Insufficient carbon flux and precursor supply may hinder titer improvement of desired products. Hence, we identified and engineered the targets in S. bingchenggensis to promote precursor supply as well as the titer and yield of milbemycins on the basis of randomly mutagenized high-yielding strains. There have been many studies using gene knockout or overexpression to modify the corresponding primary metabolic pathways, in order to increase the intracellular pool of precursors and redirect the flux toward the biosynthesis of a desired metabolite (Borodina et al. 2008; Butler et al. 2002; Jung et al. 2011; Olano et al. 2008; Reeves et al. 2007; Ryu et al. 2006; Wattanachaisaereekul et al. 2008). However, the key nodes of the TCA cycle, which are essential for cell growth, cannot be deleted from the genome (Tao et al. 2017; Viollier et al. 2001). In recent years, effective genome editing has become increasingly easy and convenient, paving the way for inhibition of the expression of key node genes (Tao et al. 2017; Tian et al. 2020; Lian et al. 2017). Our work achieved the appropriate expression of gltA (encoded citrate synthase) in S. bingchenggensis to improve the titer of milbemycins by CRISPRi system. In agreement with our observation, repression of gltA channeled more acetyl-CoA from the TCA cycle to poly (3-hydroxybutyrate) (PHB) synthesis in Halomonas sp. TD01 (Tao et al. 2017). This also proved that genes involved in the TCA cycle are essential and the identification and genetic manipulation of key nodes regulating carbon flux through the metabolic network of central carbon metabolism can lead to an increase in the availability of precursors. It implied that the continuous development and application of the CRISPRi system can be used to regulate essential gene expression in S. bingchenggensis for achieving multiple metabolic engineering goals.
Carbon sources are used by microorganisms as important nutrient molecules. Considering the significant contribution of sugar uptake transporters to the flux toward products, it is important to investigate sugar uptake transporters. Many transporters exhibit substrate promiscuity, for example, gntP encodes the gluconate transporter in Streptomyces coelicolor (Tsypik et al. 2017), but encodes a fructuronate transporter in E. coli (Bates Utz et al. 2004). In the present study, we demonstrated that gluconate transporter overexpression could promote the uptake of glucose and fructose as well as increase milbemycin production. Recently, sugar uptake systems from S. bingchenggensis genome have been shown to be effective in improving the titer of milbemycin A3/A4, avermectin B1a, and nemadectin (Jin et al. 2020). The wide application foreground and potential of the sugar uptake systems indicates that gntP may have an effect on the biosynthesis of other secondary metabolites. In brief, we achieved the goal of increasing the yield of the target product by enhancing the uptake of another carbon source, gluconate.
In summary, we provided a simple workflow, which applied genome re-sequencing to obtain interesting mutation-related targets. Moreover, CRISPRi mediated repression of genes encoding the key nodes of primary metabolism, and transporter overexpression redirected the metabolic flux in a manner that supported high secondary metabolite productivity. The present study not only identified suitable modification pathways but also provided a more meaningful reference for subsequent research.
Data availability
The re-sequence data related to this article were deposited in NCBI BioProject database with an accession number PRJNA664656.
References
Abdelal AT (1979) Arginine catabolism by microorganisms. Annu Rev Microbiol 33:139–168. https://doi.org/10.1146/annurev.mi.33.100179.001035
Argyrou A, Blanchard JS (2004) Flavoprotein disulfide reductases: advances in chemistry and function. Prog Nucleic Acid Res Mol Biol 78:89–142. https://doi.org/10.1016/S0079-6603(04)78003-4
Avignone Rossa C, White J, Kuiper A, Postma PW, Bibb M, Teixeira de Mattos MJ (2002) Carbon flux distribution in antibiotic-producing chemostat cultures of Streptomyces lividans. Metab Eng 4(2):138–150. https://doi.org/10.1006/mben.2001.0217
Bates Utz C, Nguyen AB, Smalley DJ, Anderson AB, Conway T (2004) GntP is the Escherichia coli fructuronic acid transporter and belongs to the UxuR regulon. J Bacteriol 186(22):7690–7696. https://doi.org/10.1128/JB.186.22.7690-7696.2004
Beck MH, Flaiz M, Bengelsdorf FR, Durre P (2019) Induced heterologous expression of the arginine deiminase pathway promotes growth advantages in the strict anaerobe Acetobacterium woodii. Appl Microbiol Biotechnol 104:687–699. https://doi.org/10.1007/s00253-019-10248-9
Bérdy J (2004) Bioactive microbial metabolites. J Antibiot (Tokyo) 58(1):1–26. https://doi.org/10.1038/ja.2005.1
Bierman M, Logan R, O'Brien K, Seno ET, Rao RN, Schoner RN (1992) Plasmid cloning vectors for the conjugal transfer of DNA from Escherichia coli to Streptomyces spp. Gene 116:43–49. https://doi.org/10.1016/0378-1119(92)90627-2
Borodina I, Siebring J, Zhang J, Smith CP, van Keulen G, Dijkhuizen L, Nielsen J (2008) Antibiotic overproduction in Streptomyces coelicolor A3(2) mediated by phosphofructokinase deletion. J Biol Chem 283(37):25186–25199. https://doi.org/10.1074/jbc.M803105200
Butler MJ, Bruheim P, Jovetic S, Marinelli F, Postma PW, Bibb MJ (2002) Engineering of primary carbon metabolism for improved antibiotic production in Streptomyces lividans. Appl Environ Microbiol 68(10):4731–4739. https://doi.org/10.1128/AEM.68.10.4731-4739.2002
Conrad TM, Joyce AR, Applebee MK, Barrett CL, Palsson BØJGB (2009) Whole-genome resequencing of Escherichia coli K-12 MG1655 undergoing short-term laboratory evolution in lactate minimal media reveals flexible selection of adaptive mutations. Genome Biol 10(10):R118. https://doi.org/10.1186/gb-2009-10-10-r118
Dai Z, Nielsen J (2015) Advancing metabolic engineering through systems biology of industrial microorganisms. Curr Opin Biotechnol 36:8–15. https://doi.org/10.1016/j.copbio.2015.08.006
Doroghazi JR, Metcalf WW (2013) Comparative genomics of actinomycetes with a focus on natural product biosynthetic genes. BMC Genomics 14:611. https://doi.org/10.1186/1471-2164-14-611
Duan N, Bai Y, Sun H, Wang N, Ma Y, Li M, Wang X, Jiao C, Legall N, Mao L, Wan S, Wang K, He T, Feng S, Zhang Z, Mao Z, Shen X, Chen X, Jiang Y, Wu S, Yin C, Ge S, Yang L, Jiang S, Xu H, Liu J, Wang D, Qu C, Wang Y, Zuo W, Xiang L, Liu C, Zhang D, Gao Y, Xu Y, Xu K, Chao T, Fazio G, Shu H, Zhong GY, Cheng L, Fei Z, Chen X (2017) Genome re-sequencing reveals the history of apple and supports a two-stage model for fruit enlargement. Nat Commun 8(1):249. https://doi.org/10.1038/s41467-017-00336-7
Hayes B, Schnitzler B, Wiseman S, Snyder DE (2015) Field evaluation of the efficacy and safety of a combination of spinosad and milbemycin oxime in the treatment and prevention of naturally acquired flea infestations and treatment of intestinal nematode infections in dogs in Europe. Vet Parasitol 207(1):99–106. https://doi.org/10.1016/j.vetpar.2014.11.011
He Y, Sun Y, Liu T, Zhou X, Bai L, Deng Z (2010) Cloning of separate meilingmycin biosynthesis gene clusters by use of acyltransferase-ketoreductase didomain PCR amplification. Appl Environ Microbiol 76(10):3283–3292. https://doi.org/10.1128/AEM.02262-09
He H, Ye L, Li C, Wang H, Guo X, Wang X, Zhang Y, Xiang W (2018) SbbR/SbbA, an important ArpA/AfsA-like system, regulates milbemycin production in Streptomyces bingchenggensis. Front Microbiol 9:1064. https://doi.org/10.3389/fmicb.2018.01064
Hu S, Zheng H, Gu Y, Zhao J, Zhang W, Yang Y, Wang S, Zhao G, Yang S, Jiang W (2011) Comparative genomic and transcriptomic analysis revealed genetic characteristics related to solvent formation and xylose utilization in Clostridium acetobutylicum EA 2018. BMC Genomics 12:93. https://doi.org/10.1186/1471-2164-12-93
Illeghems K, De Vuyst L, Weckx S (2013) Complete genome sequence and comparative analysis of Acetobacter pasteurianus 386B, a strain well-adapted to the cocoa bean fermentation ecosystem. BMC Genomics 14:526. https://doi.org/10.1186/1471-2164-14-526
Jin P, Li S, Zhang Y, Chu L, He H, Dong Z, Xiang W (2020) Mining and fine-tuning sugar uptake system for titer improvement of milbemycins in Streptomyces bingchenggensis. Synth Syst Biotechnol 5(3):214–221. https://doi.org/10.1016/j.synbio.2020.07.001
Jung WS, Yoo YJ, Park JW, Park SR, Han AR, Ban YH, Kim EJ, Kim E, Yoon YJ (2011) A combined approach of classical mutagenesis and rational metabolic engineering improves rapamycin biosynthesis and provides insights into methylmalonyl-CoA precursor supply pathway in Streptomyces hygroscopicus ATCC 29253. Appl Microbiol Biotechnol 91(5):1389–1397. https://doi.org/10.1007/s00253-011-3348-6
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical streptomyces genetics, vol 412. John Innes Foundation, Norwich
Kim MS, Cho WJ, Song MC, Park SW, Kim K, Kim E, Lee N, Nam SJ, Oh KH, Yoon Y (2017) Engineered biosynthesis of milbemycins in the avermectin high-producing strain Streptomyces avermitilis. Microb Cell Factories 16(1):9. https://doi.org/10.1186/s12934-017-0626-8
Lee DH, Palsson BØ (2010) Adaptive evolution of Escherichia coli K-12 MG1655 during growth on a nonnative carbon source, L-1,2-Propanediol. Appl Environ Microbiol 76(13):4158–4168. https://doi.org/10.1128/AEM.00373-10
Letek M, Valbuena N, Ramos A, Ordóñez E, Gil JA, Mateos LM (2006) Characterization and use of catabolite-repressed promoters from gluconate genes in Corynebacterium glutamicum. J Bacteriol 188(2):409–423. https://doi.org/10.1128/JB.188.2.409-423.2006
Li H (2014) Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 30(20):2843–2851. https://doi.org/10.1093/bioinformatics/btu356
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, Genome Project Data Processing S (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li L, Wei K, Zheng G, Liu X, Chen S, Jiang W, Lu Y (2018) CRISPR-Cpf1-Assisted multiplex genome editing and transcriptional repression in Streptomyces. Appl Environ Microbiol 84(18):00827–00818. https://doi.org/10.1128/AEM.00827-18
Lian J, HamediRad M, Hu S, Zhao H (2017) Combinatorial metabolic engineering using an orthogonal tri-functional CRISPR system. Nat Commun 8(1):1688. https://doi.org/10.1038/s41467-017-01695-x
Liu H, Reynolds KA (2001) Precursor supply for polyketide biosynthesis: the role of crotonyl-CoA reductase. Metab Eng 3(1):40–48. https://doi.org/10.1006/mben.2000.0169
Lu C, Zhang X, Jiang M, Bai L (2016) Enhanced salinomycin production by adjusting the supply of polyketide extender units in Streptomyces albus. Metab Eng 35:129–137. https://doi.org/10.1016/j.ymben.2016.02.012
Merola VM, Eubig PA (2018) Toxicology of avermectins and milbemycins (macrocyclic lactones) and the role of P-Glycoprotein in dogs and cats. Vet Clin North Am Small Anim Pract 48(6):991–1012. https://doi.org/10.1016/j.cvsm.2018.07.002
Monod M, Feuermann M, Salamin K, Fratti M, Makino M, Alshahni MM, Makimura K, Yamada T (2019) Trichophyton rubrum azole resistance mediated by a new ABC transporter, TruMDR3. Antimicrob Agents Chemother 63(11):e00863–e00819. https://doi.org/10.1128/aac.00863-19
Nonaka K, Kumasaka C, Okamoto Y, Maruyama F, Yoshikawa H (1999a) Bioconversion of milbemycin-related compounds: biosynthetic pathway of milbemycins. J Antibiot (Tokyo) 57(2):109–116. https://doi.org/10.7164/antibiotics.52.109
Nonaka K, Tsukiyama T, Sato K, Kumasaka C, Maruyama F, Yoshikawa H (1999b) Bioconversion of milbemycin-related compounds: isolation and utilization of non-producer, strain RNBC-5-51. J Antibiot (Tokyo) 52(7):620–627. https://doi.org/10.7164/antibiotics.52.620
Olano C, Lombo F, Mendez C, Salas JA (2008) Improving production of bioactive secondary metabolites in actinomycetes by metabolic engineering. Metab Eng 10(5):281–292. https://doi.org/10.1016/j.ymben.2008.07.001
Palazzotto E, Tong Y, Lee SY, Weber T (2019) Synthetic biology and metabolic engineering of actinomycetes for natural product discovery. Biotechnol Adv 37(6):107366. https://doi.org/10.1016/j.biotechadv.2019.03.005
Peano C, Damiano F, Forcato M, Pietrelli A, Palumbo C, Corti G, Siculella L, Fuligni F, Tagliazucchi GM, De Benedetto GE, Bicciato S, De Bellis G, Alifano P (2014) Comparative genomics revealed key molecular targets to rapidly convert a reference rifamycin-producing bacterial strain into an overproducer by genetic engineering. Metab Eng 26:1–16. https://doi.org/10.1016/j.ymben.2014.08.001
Pickens LB, Tang Y, Chooi YH (2011) Metabolic engineering for the production of natural products. Annu Rev Chem Biomol Eng 2:211–236. https://doi.org/10.1146/annurev-chembioeng-061010-114209
Reeves AR, Brikun IA, Cernota WH, Leach BI, Gonzalez MC, Weber JM (2007) Engineering of the methylmalonyl-CoA metabolite node of Saccharopolyspora erythraea for increased erythromycin production. Metab Eng 9(3):293–303. https://doi.org/10.1016/j.ymben.2007.02.001
Romero-Rodriguez A, Rocha D, Ruiz-Villafan B, Tierrafria V, Rodriguez-Sanoja R, Segura-Gonzalez D, Sanchez S (2016) Transcriptomic analysis of a classical model of carbon catabolite regulation in Streptomyces coelicolor. BMC Microbiol 16:77. https://doi.org/10.1186/s12866-016-0690-y
Russell DW, Sambrook J (2001) Molecular cloning: a laboratory manual. Cold Spring Harbour Laboratory Press, New York
Ryu Y-G, Butler MJ, Chater KF, Lee KJ (2006) Engineering of primary carbohydrate metabolism for increased production of actinorhodin in Streptomyces coelicolor. Appl Environ Microbiol 72(11):7132–7139. https://doi.org/10.1128/AEM.01308-06
Sharkey MA, Oliveira TF, Engel PC, Khan AR (2013) Structure of NADP(+)-dependent glutamate dehydrogenase from Escherichia coli--reflections on the basis of coenzyme specificity in the family of glutamate dehydrogenases. FEBS J 280(18):4681–4692. https://doi.org/10.1111/febs.12439
Takiguchi Y, Ono M, Muramatsu S, Ide J, Mishima H, Terao M (1983) Milbemycins, a new family of macrolide antibiotics. Fermentation, isolation and physico-chemical properties of milbemycins D, E, F, G, and H. J Antibiot (Tokyo) 36(5):502. https://doi.org/10.7164/antibiotics.36.502
Tao W, Lv L, Chen GQ (2017) Engineering Halomonas species TD01 for enhanced polyhydroxyalkanoates synthesis via CRISPRi. Microb Cell Factories 16(1):48. https://doi.org/10.1186/s12934-017-0655-3
Tian J, Yang G, Gu Y, Sun X, Lu Y, Jiang W (2020) Developing an endogenous quorum-sensing based CRISPRi circuit for autonomous and tunable dynamic regulation of multiple targets in Streptomyces. Nucleic Acids Res 48(14):8188–8202. https://doi.org/10.1093/nar/gkaa602
Tsypik O, Makitrynskyy R, Bera A, Song L, Wohlleben W, Fedorenko V, Ostash B (2017) Role of gntR family regulatory gene SCO1678 in gluconate metabolism in Streptomyces coelicolor M145. Biomed Res Int 2017:9529501–9529509. https://doi.org/10.1155/2017/9529501
Viollier PH, Minas W, Dale GE, Folcher M, Thompson CJ (2001) Role of acid metabolism in Streptomyces coelicolor morphological differentiation and antibiotic biosynthesis. J Bacteriol 183(10):3184–3192. https://doi.org/10.1128/JB.183.10.3184-3192.2001
Wang X, Wang X, Xiang W (2009) Improvement of milbemycin-producing Streptomyces bingchenggensis by rational screening of ultraviolet- and chemically induced mutants. World J Microbiol Biotechnol 25(6):1051–1056. https://doi.org/10.1007/s11274-009-9986-5
Wang X, Zhang B, Yan Y, An J, Zhang J, Liu C, Xiang W (2013) Characterization and analysis of an industrial strain of Streptomyces bingchenggensis by genome sequencing and gene microarray. Genome 56(11):677–689. https://doi.org/10.1139/gen-2013-0098
Wang H, Zhang J, Zhang Y, Zhang B, Liu CX, He H, Wang X, Xiang W (2014) Combined application of plasma mutagenesis and gene engineering leads to 5-oxomilbemycins A3/A4 as main components from Streptomyces bingchenggensis. Appl Microbiol Biotechnol 98(23):9703–9712. https://doi.org/10.1007/s00253-014-5970-6
Wang W, Yang T, Li Y, Li S, Yin S, Styles K, Corre C, Yang K (2016) Development of a synthetic oxytetracycline-inducible expression system for streptomycetes using de novo characterized genetic parts. ACS Synth Biol 5:765–773. https://doi.org/10.1021/acssynbio.6b00087
Wang G, Shi T, Chen T, Wang X, Wang Y, Liu D, Guo J, Fu J, Feng L, Wang Z, Zhao X (2018) Integrated whole-genome and transcriptome sequence analysis reveals the genetic characteristics of a riboflavin-overproducing Bacillus subtilis. Metab Eng 48:138–149. https://doi.org/10.1016/j.ymben.2018.05.022
Wang H, Cheng X, Liu Y, Li S, Zhang Y, Wang X, Xiang W (2020a) Improved milbemycin production by engineering two cytochromes P450 in Streptomyces bingchenggensis. Appl Microbiol Biotechnol 104(7):2935–2946. https://doi.org/10.1007/s00253-020-10410-8
Wang W, Li S, Li Z, Zhang J, Fan K, Tan G, Ai G, Lam S, Shui G, Yang Z, Lu H, Jin P, Li X, Xia X, Liu X, Dannelly K, Yang C, Yang Y, Zhang S, Alterovitz G, Xiang W, Zhang L (2020b) Harnessing the intracellular triacylglycerols for titer improvement of polyketides in Streptomyces. Nat Biotechnol 38(1):76–83. https://doi.org/10.1038/s41587-019-0389-3
Wattanachaisaereekul S, Lantz AE, Nielsen ML, Nielsen J (2008) Production of the polyketide 6-MSA in yeast engineered for increased malonyl-CoA supply. Metab Eng 10(5):246–254. https://doi.org/10.1016/j.ymben.2008.04.005
Zabala D, Brana AF, Florez AB, Salas JA, Mendez C (2013) Engineering precursor metabolite pools for increasing production of antitumor mithramycins in Streptomyces argillaceus. Metab Eng 20:187–197. https://doi.org/10.1016/j.ymben.2013.10.002
Zhang J, An J, Wang J, Yan Y, He H, Wang X, Xiang W (2013) Genetic engineering of Streptomyces bingchenggensis to produce milbemycins A3/A4 as main components and eliminate the biosynthesis of nanchangmycin. Appl Microbiol Biotechnol 97(23):10091–10101. https://doi.org/10.1007/s00253-013-5255-5
Zhang Y, He H, Liu H, Wang H, Wang X, Xiang W (2016) Characterization of a pathway-specific activator of milbemycin biosynthesis and improved milbemycin production by its overexpression in Streptomyces bingchenggensis. Microb Cell Factories 15(1):152. https://doi.org/10.1186/s12934-016-0552-1
Zhang X, Liu B, Zou F, Shen D, Yin Z, Wang R, He F, Wang Y, Tyler BM, Fan W, Qian W, Dou D (2019) Whole genome re-sequencing reveals natural variation and adaptive evolution of Phytophthora sojae. Front Microbiol 10:2792. https://doi.org/10.3389/fmicb.2019.02792
Acknowledgements
We would like to thank Professor Weihong Jiang (Institute of Plant Physiology and Ecology, Chinese Academy of Sciences, Shanghai, China) and Professor Yinhua Lu (Shanghai Normal University, Shanghai, China) for providing pSET-ddCpf1, Professor Mark Buttner (John Innes Centre, Norwich, UK) for providing plasmid pIJ10500, and Professor Weishan Wang (Chinese Academy of Sciences, Beijing, China) for providing plasmid PIJ-Potr and promoter kasOp*.
Funding
This work received financial support from the National Natural Science Foundation of China (Grant Nos: 31972291, 31872936 and 31572070).
Author information
Authors and Affiliations
Contributions
WX, XW, and YL designed the research. Material preparation, data collection, and analysis were performed by YL and HW. XC assisted with experiments. The first draft of the manuscript was written by YL, WX, XW, HW, and SL, and YZ modified the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 743 kb).
Rights and permissions
About this article
Cite this article
Liu, Y., Wang, H., Li, S. et al. Engineering of primary metabolic pathways for titer improvement of milbemycins in Streptomyces bingchenggensis. Appl Microbiol Biotechnol 105, 1875–1887 (2021). https://doi.org/10.1007/s00253-021-11164-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11164-7