Abstract
As a primary cause of food contamination and human diseases, Salmonella Typhimurium can easily form a biofilm that is difficult to remove from food surfaces, and often causes significant invisible threats to food safety. Although berberine has been widely used as an anti-infective drug in traditional medicine, some basic principles underlying its mechanism, especially the interaction between berberine and type I fimbriae genes, has not been verified yet. In this study, two strains of major fimbrial gene mutants (ΔfimA and ΔfimH) were constructed to demonstrate the possible action of berberine on type I fimbriae genes. The broth microdilution method was used to determine the antibacterial activity of berberine against selected strains (WT, ΔfimA, and ΔfimH). Cell agglutination experiments revealed that the number of S. Typhimurium type I fimbriae reduced after berberine treatment, which was consistent with transmission electron microscopy results. Quantitative real-time PCR experiments also confirmed that berberine reduced fimA gene expression, indicating a certain interaction between berberine and fimA gene. Furthermore, confocal laser scanning microscopy imaging of biofilm clearly revealed that berberine prevents biofilm formation by reducing the number of type I fimbriae. Overall, it is well speculated for us that berberine could be an excellent combating-biofilm drug in clinical microbiology and food preservation.
Key points
• Reduce the number of fimbriae.
• Berberine targeting fimA.
• Effective biofilm inhibitor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With food safety becoming an issue of increasing importance, people have begun to focus on food safety and health (Andino and Hanning 2015). Food spoilage caused by foodborne bacterial contamination has been identified as a key problem in food safety (Eran et al. 2020). Salmonella Typhimurium belongs to the genus Salmonella; it is a Gram-negative, Enterobacteriaceae bacterium, determined to be responsible for majority of human food poisoning cases worldwide. It often causes serious illnesses, including gastroenteritis, septicemia, and some suppurative infections, thus posing serious challenges to food safety and public health (Kuehn 2019). According to reports, approximately 40,000 cases of salmonellosis are reported every year in the USA. As no milder cases have been reported, the actual number may be 30 times or more than what has been reported (Fàbrega and Vila 2013). S. Typhimurium is considered to be highly infectious due to its complex virulence factors. Adhesion and invasion on the host are the prerequisite for its pathogenicity, which are also a prerequisite for bacterial infection. The initial adhesion and invasion are often accomplished by the movement mediated by flagella and fimbria (Ledeboer et al. 2006). Among these structures, the bacterium depends on its body surface fimbria and its adhesion to complete the initial colonization and adhesion (Zeng et al. 2017). Therefore, bacterial fimbriae have been verified to play a vital role in infection and pathogenesis, among which type I fimbriae, the major fimbriae of S. Typhimurium, are the most widely examined. The majority of these fimbriae are composed of structural protein subunits that are short rod-like structures existing on the surface of bacteria and are manipulated and encoded in fim gene clusters (Korhonen et al. 1980). It has been reported to be expressed in all Salmonella serotypes detected under static conditions (Wang et al. 2012). fimA is the major protein subunit of type I fimbria, which forms its main component and exerts an effect on host orientation (Meng et al. 2019). fimH is located at the top of the fimbria and determines the specific adhesion of different serotypes I fimbria to host cells (Uchiya et al. 2019). Moreover, type I fimbriae may be directly related to the formation of biofilm.
S. Typhimurium often colonizes the host’s intestinal tract through adhesion and invasion, forming a dense biofilm to enhance its pathogenicity (Zeng et al. 2017). The biofilm is a special structure formed by bacteria colonizing on biological or nonbiological surfaces to resist the external environment (Flemming et al. 2016). The biofilm formation prolongs the duration of contaminating the food and, at the same time, increases the hidden threat to food safety. Currently, traditional sterilization methods have several issues in combating bacteria and their biofilms (Wolfmeier et al. 2018). In other words, antibiotics have emerged as the most important approach to fight bacteria. Nevertheless, in the past few years, antibiotic abuse has led to the emergence of several resistant super bacteria (Kuehn 2019; Stapels et al. 2018). These resistant bacteria can withstand the effects of traditional antibiotics such as penicillin, azithromycin, tetracycline, chlorine, and rifamycin (Michael and Schwarz 2016). Only a few antibiotics such as vancomycin have a killing effect on them. Therefore, exploring for new and novel effective antibacterial agents has become an urgent task for us. In recent years, natural active products have attracted much attention due to their strong antimicrobial effects and fewer side effects. In this context, a recent research has demonstrated berberine to possess more medicinal value as a traditional Chinese medicine ingredient (Habtemariam 2020). It has been identified as a benzylisoquinoline alkaloid that exists as an active ingredient in several medicinal plants and possesses a variety of pharmacological properties. Earlier research has reported that berberine has excellent effects such as anti-Plasmodium, antidiarrheal, and anti-trachoma activities (Kumar et al. 2015). Furthermore, some clinical and preclinical studies have demonstrated that berberine can improve several disorders, including metabolic, neurological, and cardiac diseases (Imenshahidi and Hosseinzadeh 2016; Liu et al. 2020). At the same time, it also possesses good antibacterial activity. Therefore, berberine can be considered as a dual-use biologically active substance with an irreplaceable role in the prevention of food poisoning (Aswathanarayan and Vittal 2018; Chu et al. 2014; Liu et al. 2015). Considering food safety becoming an issue of increasing importance, berberine use may become more widespread in the future.
In recent years, with the continuous deepening of research, an increasing number of antibacterial mechanisms of berberine have been elucidated (Shi et al. 2018). Its multitarget mechanism against organisms includes inhibiting the activity of tonic isomerase, thereby affecting the synthesis of DNA and the function of the FtsZ cell division protein (Boberek et al. 2010). A previous study reported that berberine has an inhibitory effect on Escherichia coli pili in urinary tract infections (Sun et al. 1988). Chaperone–usher pili are the most common type of fimbriae produced by Gram-negative bacteria. They included type I and P pili, which exhibit several similarities in their biogenesis and structure (Proft and Baker 2008). Therefore, based on the above-described information, we speculated that berberine can also affect type I fimbriae. To verify this speculation, we conduct this work to explore the interaction of berberine as a biologically active substance with S. Typhimurium type I fimbriae, elucidate the antibacterial mechanism of berberine on biofilm, and provide a novel perspective for food preservation and infection prevention.
Materials and methods
Reagents
Berberine (HPLC ≥ 98%, CAS 633–65-8) was purchased from Sangon Biotech Co., Ltd. (Shanghai, China). It was dissolved in dimethyl sulfoxide (DMSO) to prepare a 2 mg/mL mother liquor. The final concentration of DMSO in the sample and that in its control treatment solution used in the experiment was 10% (v/v) each. It has no effect on S. Typhimurium.
Bacterial strains and growth conditions
S. Typhimurium wild-type (WT) strain CMCC50115 is identified as a S. Typhimurium LT2 derivative. All strains and plasmids used in this research are described in Table 1. The strains used in the experiment were stored at −80 °C. Bacteria were cultured in Luria–Bertani (LB) agar or LB broth at 37 °C. The biofilm was then cultured in 1% tryptone media (10 g/L NaCl, 15 g/L agar, 5 g/L yeast extract, and 10 g/L tryptone) at 28 °C.
Construction of mutants
Bacterial mutants were constructed through allelic replacement and homologous recombination (Datsenko and Wanner 2000). A chloramphenicol fragment with two FRT sites was amplified by polymerase chain reaction (PCR) using pKD3 as a vector. After digestion, it was electroporated into bacteria carrying pKD46. The first recombinant strain was obtained by screening positive clones on a chloramphenicol agar plate. Next, the plasmid pCP20 was electroporated into the first recombinant strain, and the second recombinant strain was obtained by screening positive clones on ampicillin agar plates (42 °C, 24 h) and confirmed by sequencing analysis.
Determination of minimum inhibitory concentration
The minimum inhibitory concentrations (MICs) of berberine against Salmonella strains were determined via the broth dilution method. After adding 100 μL of the bacterial suspension at a concentration of approximately 106 CFU/mL to each well of a 96-well plate, 100 μL of berberine (at concentrations of 2000, 1000, 900, 500, 300, or 0 μg/mL) was added to each well. As a negative control, 10% DMSO LB broth was used. The 96-well plate was then incubated at 37 °C for 24 h, after which the effect of berberine on different test bacteria was examined.
Killing curve of berberine
The bacterial suspension was diluted to 106 CFU/mL using LB broth. Equal volumes of LB broth containing different concentrations of berberine were added to individual wells in order to achieve final berberine concentrations of 0.9, 0.45, 0.225, 0.1125, or 0.05625 mg/mL. LB broth containing 10% DMSO was then used as a negative control. The samples were further cultured at 37 °C for 24 h, and the OD600 nm was monitored every 1 h.
Guinea pig erythrocyte hemagglutination test
Salmonella and its mutants were cultivated overnight in LB broth. These overnight cultures were diluted at 1:100 in 5 mL of LB. To express more fimbriae, the strains were cultivated thrice, each for 48 h. Then, the bacteria were diluted in LB medium to OD600 = 0.8. Next, 1 mL of the bacterial suspension was taken, washed three times, and resuspended in PBS. A 3% red blood cell solution (v/v) was prepared in PBS. The hemagglutination assay was then performed.
Transmission electron microscopy analysis
The fimbrial morphology was visualized by transmission electron microscopy (TEM) with negative staining. A single colony was picked and inoculated into fresh LB medium, and the OD600 value of the bacteria was adjusted to approximately 0.3. The cells were washed with deionized water and resuspended in sterile water. Next, 5 μL of the bacterial solution was added dropwise to a copper mesh with a carbon film and dried at room temperature. Using 1% phosphotungstic acid (pH 7.4) for negative staining for 30 s, the excess solution was blotted with filter paper and observed by electron microscopy (TecnaiG2 F20, FEI, America).
Total RNA isolation and quantitative real-time PCR
The determination method is as described previously (Yang et al. 2019). To determine the effect of berberine on the transcription of fimbriae-related genes, S. Typhimurium CMCC50115 was cultured in LB broth containing different concentrations of berberine at 37 °C for 24 h. Total RNA was extracted using the Tiangen RNAprep Pure Cell/Bacteria Kit (Tiangen, Beijing, China), according to the manufacturer’s instructions. The Takara PrimeScript™ RT Reagent Kit (Takara, Kyoto, Japan) was used to reverse-transcribe the RNA into cDNA, according to the manufacturer’s instructions. The obtained cDNA samples were stored at −20 °C until further use. The primer sequences used for qPCR are listed in Table 2. The qPCR assays (10 μL) with SYBR® Premix Ex Taq™II (Takara) were performed using the IQ™5 system (Bio-Rad).
Biofilm formation and quantification assay
Biofilm formation was observed as described previously with some modifications (Baugh et al. 2012). Bacteria were cultured overnight in LB medium without NaCl and diluted to 106 CFU/mL. Then, 200 μL of the bacterial suspension was added to each well of a 96-well plate and cultured at 28 °C for 24, 48, and 72 h, respectively. The total amount of biofilm biomass was quantified by crystal violet. The medium was removed in each well and was later washed with deionized water three times to avoid the interference of suspended bacteria on the biofilm. The biofilm was then fixed with formaldehyde for 10 min, and 0.5% crystal violet was added to stain the biofilm for 15 min. The excess crystal violet was blotted and washed three times, the remaining crystal violet was dissolved in 200 μL of 33% glacial acetic acid per well, and then the OD595 value was measured using an ELISA reader (Thermo Scientific Labsystems 354). All experiments were repeated three times at different time points.
Confocal laser scanning microscopy imaging of biofilm
The test bacteria were cultured overnight at 28 °C on a shaker and then suspended in a 50 mL NaCl-free LB Erlenmeyer flask to maintain the final concentration of 107 CFU/mL. A sterile glass slide was then placed in the Erlenmeyer flask, followed by incubation for 24, 48, and 72 h at 28 °C. After completing the culture, the glass slide was placed in a clean Petri dish and rinsed with deionized water three times to remove excess medium. Then, the slide was stained with 100 μg/mL, green fluorescence, labeled bacterial cells, and Alexa Fluor 647 (50 μg/mL, red fluorescence, labeled EPS) solution in the dark at room temperature for 30 min. Confocal microscopic images were obtained using a Leica DMi8 microscope with a × 40 objective lens. The confocal microscope was operated at excitation/emission wavelengths of Alexa Fluor 647 and SYTO 9 of 650/668 and 480/500 nm, respectively. Stacked images were obtained by scanning the biofilm along the Z-axis at intervals of 0.5 μm.
Results
MICs
As has been depicted in Fig. 1, the MICs of berberine against WT and mutant strains (ΔfimA and ΔfimH) were evaluated in the concentration range of 0.3–2 mg/mL. The MIC of berberine on WT and ΔfimH was found to be 0.9 mg/mL, whereas the MIC for ΔfimA was 1 mg/mL. We selected 0.9 mg/mL concentration of berberine as the MIC for subsequent experiments.
Killing curve of berberine
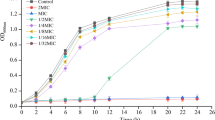
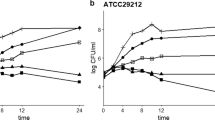
The 24 h growth curves of WT, fimbrial mutant ΔfimA, and ΔfimH were separately determined. As shown in Fig. 2a, the growth curves of the mutant and WT strains were basically similar. The WT and mutant strains were treated with different concentrations (1/32, 1/16, 1/8, 1/4, and 1/2, and 1 MIC) of berberine, and the untreated group was used as the control. As shown in Fig. 2b, c, and d, 1/32 and 1/16 MIC of berberine showed no significant difference in the growth of bacterial population. Therefore, 1/16 MIC of berberine was selected as the subinhibitory concentration to test the effect of berberine on the fimbrial genes.
a Twenty-four-hour growth curve of WT and mutant (ΔfimA, ΔfimH). b Effects of different sub-inhibitory concentrations of berberine on WT. c Effects of different sub-inhibitory concentrations of berberine on mutants (ΔfimA). d Effects of different sub-inhibitory concentrations of berberine on mutants (ΔfimH). Error bars represent the standard deviations of three replicate experiments
TEM analysis
TEM was used to further verify the microscopic morphological changes of WT and its mutants. As presented in Fig. 3a–f, a large number of fimbriae have been determined to exist on the surface of WT. However, neither ΔfimA nor ΔfimH observed the production of type I fimbriae. It was confirmed that the deletion of fimA or fimH gene would obstruct the ability to produce type I fimbriae. Compared with untreated WT, the strains treated with 1/16 MIC of berberine showed a reduced number of type I fimbriae on their surface (Fig. 3g, h).
Guinea pig erythrocyte hemagglutination test for type 1 fimbriae
Type I fimbria present on the surface of S. Typhimurium has been determined to contain lectin that is sensitive to mannose and can agglutinate the red blood cells of guinea pigs. Results showed that WT has an agglutination effect on the red blood cells of guinea pigs, whereas the fimbrial mutants ΔfimA and ΔfimH could not agglutinate. Agglutination experiment was then performed simultaneously after treating the WT strains with 1/16 MIC of berberine, which showed that the agglutination effect and the number of type I fimbriae in the experimental group treated with berberine were reduced compared with the WT, as shown in Table 3.
Quantitative real-time PCR
Total RNA was prepared, and quantitative real-time PCR was performed in order to determine the expression of the fimbrial subunit gene fimA and the fimbrial adhesin gene fimH. The expression of Grby gene was used as control. First, the expression profile of a single gene was standardized according to the Grby gene expression and was then later compared with the expression levels of fimA and fimH in WT treated with different subinhibitory concentrations of berberine. As depicted in Fig. 4a, the fimA gene is not expressed in the ΔfimA strain, and the expression level of fimH is determined to be 0.81. In the ΔfimH strain, the expression level of fimH was 0, and the expression level of the fimA gene was only 0.51. As the concentration of berberine increases, the expression of fimA gene decreases in a concentration-dependent manner, and the expression of fimH gene increases in a concentration-dependent manner (Fig. 4b, c).
Biofilm formation and quantification assay
The biofilm formation processes of WT, ΔfimA, and ΔfimH were observed by crystal violet staining after culturing at 28 °C for 24, 48, and 72 h, and their absorbance were measured at OD595 nm, respectively. As depicted in Fig. 5, the number of biofilm has positively correlated with time. The biofilm formation rates of ΔfimA and ΔfimH were judged based on that of WT at different time periods. The biofilm formation rates of ΔfimA and ΔfimH at 24 and 48 h were 81.12%, 77.7%, and 54.22% and 56.58%, 99.59%, and 99.74%, respectively.
a ΔfimA, ΔfimH, and WT were grown for 24 h, 48 h, and 72 h in the borosilicate tubules. b Quantify the biofilm of WT, fimA, and fimH at 24 h, 48 h, and 72 h. Data are averages of three replicates (n = 3). Error bars represent the standard deviation. The data were analyzed using Student’s two-tailed t test. *P < 0.05
The biofilms of WT, ΔfimA, and ΔfimH were further observed after culturing at 28 °C for 48 h (Fig. 6a), with the value of WT being considered as the standard. The biofilm formation rates of ΔfimA and ΔfimH were 54.22% and 56.58%, respectively. The formed biofilms of WT, ΔfimA, and ΔfimH strains after treatment with 1/16 MIC of berberine were inhibited to varying degrees of 66.29%, 7.58%, and 26.59%, respectively, compared with the control group.
a Forty-eight-hour crystal violet stained S. Typhimurium biofilm was formed on the wells of a 96-well polystyrene plate. The first well has medium alone (Media), followed by WT, 1/16MIC berberine-treated WT, ΔfimA, 1/16MIC berberine-treated ΔfimA, ΔfimH, and 1/16MIC berberine-treated ΔfimH. b WT, 1/16MIC berberine-treated WT, ΔfimA, 1/16MIC berberine-treated ΔfimA, ΔfimH, and 1/16MIC berberine-treated ΔfimH were grown for 48 h on the glass, stained with SYTO 9 and Con-Alexa Fluor, and 3D images were acquired by CLSM, scale bar: 10 μm. c The bacteria of biofilm thickness in strains WT, 1/16MIC berberine-treated WT, ΔfimA, 1/16MIC berberine-treated ΔfimA, ΔfimH, and 1/16MIC berberine-treated ΔfimH are respectively indicated. Data are averages of three replicates (n = 3). Error bars represent the standard deviation. The data were analyzed using Student’s two-tailed t test. *P < 0. 05, **P < 0.005
CLSM imaging of biofilm
Confocal laser scanning microscopy (CLSM) was used to further characterize the biofilm formation ability of the different bacterial strains. As shown in Fig. 6b, the biofilm thickness reflects the ability of biofilm formation. In six sets of groups (WT, ΔfimA, ΔfimH, 1/16 MIC WT, 1/16 MIC ΔfimA, and 1/16 MIC ΔfimH), the thicknesses of the biofilm were 21.52, 11.32, 11.09, 6.07, 10.47, and 7.81 μm, respectively. This result was consistent with that obtained by crystal violet staining. The WT strain treated with 1/16 MIC of berberine showed inhibition of biofilm formation, but it had no significant effect on the biofilm formation ability of ΔfimA treated with 1/16 MIC of berberine.
Discussion
Berberine has been widely used as an anti-infective drug in traditional medicine. Its antibacterial activity has been confirmed against a variety of bacteria, including Streptococcus and Actinomyces (Chen et al. 2016). Although several studies have been conducted to explore the possible antibacterial action mechanism of berberine, there is no research describing its inhibitory effect on S. Typhimurium type I fimbriae. The structure of type I fimbriae of S. Typhimurium is as conservative as in E. coli (Hospenthal et al. 2017). The aggregation of S. Typhimurium type I fimbriae is dependent on the transcription and assembly of fim gene clusters (Proft and Baker 2008). The primary body of S. Typhimurium type I fimbriae has been determined to be composed of fimA and fmH. fimA constitutes the major component of the fimbrial structure, and fimH is the adhesin located at the top of the fimbria. Therefore, using the two primary structures of fimbriae, we used homologous recombination in this study to construct the fimbrial gene mutants ΔfimA and ΔfimH in order to examine their function. Next, we investigated the MIC of berberine on the bacteria. Results revealed an MIC of 1 mg/mL on ΔfimA, which was greater than the MIC of 0.9 mg/mL on WT and ΔfimH strains (Fig. 1). Hence, we speculated that the fimA gene could be one of the action sites of berberine against the bacteria.
Berberine has been confirmed to be a natural active substance with multiple targets (Karaosmanoglu et al. 2013). We constructed the time-killing curve of berberine concentration gradient to explore its killing effect on the bacterial strains (WT, ΔfimA, and ΔfimH) (Fig. 2b–d), and the results demonstrated that 1/16 and 1/32 MIC of berberine had no quantitative effect on the bacteria, which confirmed that the bacteria can still grow normally at this concentration. Therefore, 1/16 MIC of berberine was selected as the subinhibitory concentration to treat the bacteria. Previous research has demonstrated that both fimA and fimH are required for the aggregation of type I fimbriae of S. Typhimurium and that the complete fimbrial structure cannot be successfully assembled in the absence of fimA or fimH (Zeiner et al. 2012). We then confirmed that finding by TEM, which demonstrated a large number of actinomorphous, short rod-like fimbriae on the surface of WT strain that contrasted clearly with the flagella on the body. However, the absence of the genes of fimA and fimH resulted in a nonfimbriate phenotype with no fimbriae detected by direct observation by electron microscopy (Fig. 3). S. Typhimurium type I fimbriae are sensitive to mannose due to its adhesin fimH, which can mediate the presence, aggregation, and adhesion of bacteria on the intestinal tract (Uchiya et al. 2019).
A previous study also demonstrated that type I fimbriae can adhere to Saccharomyces cerevisiae and guinea pig red blood cells with mannose residues (Wang et al. 2012). The agglutination test using yeast or guinea pig red blood cells can be used to characterize the number of type I fimbriae on the surface of S. Typhimurium (Korhonen 1979; Kuan and Yeh 2019). Our results showed that the mutants ΔfimA and ΔfimH appeared to have lost their ability to agglutinate guinea pig red blood cells (Table 3). Because of the absence of fimbriae on their body surface, guinea pig red blood cells cannot bind to bacteria. The agglutination ability of WT strain treated with 1/16 MIC of berberine was only half of that of the untreated WT strain, which again confirmed that berberine can inhibit fimbrial aggregation and reduce their number.
To further examine the relationship between berberine and fimA gene at the genetic level, we evaluated the relative expression levels of fimA and fimH genes of WT strain after treating with different concentrations of berberine (Fig. 4). In ΔfimA, the gene expression of fimH was 0.811, which was inhibited to a certain extent. In ΔfimH, the expression of fimA gene was only 0.512 (Fig. 4a). This is consistent with a previously reported result (Zeiner et al. 2012). The expression of fimA gene in WT strain decreased in a concentration-dependent manner with increasing concentrations of berberine. The expression of fimA gene was suppressed with berberine treatment (Fig. 4b). Simultaneously, the same experiment was performed on fimH, and it was observed that berberine has exerted a promoting effect on the expression of fimH gene. With the increase in berberine concentration, the expression of fimH gene was also increased (Fig. 4c). The biogenesis of type I fimbriae in S. Typhimurium is companion-guided and generally starts from the adhesin site at the top of the fimbriae. Therefore, we speculated that berberine affects the expression of fimA gene. The decrease in fimA gene expression stimulates the overexpression of fimH gene at the top of the fimbria. This is consistent with the experimental results. Through fluorescence quantitative PCR, we again confirmed the targeted inhibition of berberine on fimA gene.
Fimbriae are essential for host identification, colonization, and biofilm formation during bacterial infection (Steenackers et al. 2012). Compared with planktonic bacteria, bacterial cells in biofilms are considered to be more resistant to most antibacterial agents and host defenses (Koo et al. 2017). Therefore, efficiently removing the biofilm is becoming more difficult. However, the development of biofilms is a dynamic process, which is primarily divided into three stages as follows: attachment, maturation, and dispersion (Roy et al. 2018; Wolfmeier et al. 2018). Regarding the biofilm formation mechanism of bacteria, fimbria-mediated adhesion is generally considered to be the first step to achieve initial contact, formation of microcolonies, invasion, and chronic infection (Flemming et al. 2016). Through the crystal violet staining, the ability to form biofilm of WT, ΔfimA, and ΔfimH at different times (24 h, 48 h, 72 h) were explored. The results showed that at 24 h, the biofilm formation rates of ΔfimA and ΔfimH that were compared to WT were 81.12% and 77.7%. At 48 h, the biofilm formation rates of ΔfimA and ΔfimH were only 54.22% and 56.58% relative to WT (Fig. 5). Deletion of fimA and fimH genes has reduced the biofilm formation ability of the strain, reaching the lowest rate at 48 h. Then, after maturation and accumulation of biofilm, the amount of biofilm of these bacteria did not exhibit an obvious difference. It is speculated that the fimbria-mediated adhesion played a vital role in the early and middle stages of the formation of the biofilm. Upon further maturation, the bacterial stack becomes dense, during which the fimbriae may exhibit a weaker effect on contributing to the development of the biofilm.
Type I fimbriae are required for establishing the initial attachment to cells (Lukaszczyk et al. 2019). Therefore, S. Typhimurium type I fimbriae are a good target against its biofilm formation. Based on previous experimental results, we speculated that type I fimbriae could be inhibited by berberine (Aswathanarayan and Vittal 2018). To further confirm the antibacterial ability of berberine on biofilm, we explored the relationship between biofilms in different treatment groups at 48 h (Fig. 6a). This was again characterized by CLSM (Fig. 6b, c). After treating the WT strain with 1/16 MIC of berberine, the thickness of the biofilm was only 6.07 μm, indicating that a low berberine concentration can prevent the biofilm formation. Comparing ΔfimA (without berberine treatment) with ΔfimA treated by berberine, we observed that the lack of fimA gene made berberine to lose its antibacterial ability on the biofilm. However, for ΔfimH, its biofilm can still be inhibited by berberine. It can be inferred that berberine affects S. Typhimurium type I fimA gene, thereby inhibiting the production of biofilms. This is consistent with our previous conclusions.
In summary, berberine affects the expression of fimA gene of type I fimbriae, reduces the number of type I fimbriae, and thence decreases the activity and adhesion of bacteria. This property renders type I fimbriae to lose their major function and further prevent bacterial aggregation to decrease biofilm formation. Therefore, berberine can be considered as an effective natural drug against S. Typhimurium and its biofilm.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Andino A, Hanning I (2015) Salmonella enterica: survival, colonization, and virulence differences among serovars. Sci World J 2015:520179–520179. https://doi.org/10.1155/2015/520179
Aswathanarayan JB, Vittal RR (2018) Inhibition of biofilm formation and quorum sensing mediated phenotypes by berberine in Pseudomonas aeruginosa and Salmonella Typhimurium. RSC Adv 8(63):36133–36141. https://doi.org/10.1039/C8RA06413J
Baugh S, Ekanayaka AS, Piddock LJV, Webber MA (2012) Loss of or inhibition of all multidrug resistance efflux pumps of Salmonella enterica serovar Typhimurium results in impaired ability to form a biofilm. J Antimicrob Chemoth 67(10):2409–2417. https://doi.org/10.1093/jac/dks228
Boberek JM, Stach J, Good L (2010) Genetic evidence for inhibition of bacterial division protein FtsZ by berberine. PLoS One 5(10):e13745–e13745. https://doi.org/10.1371/journal.pone.0013745
Chen L, Bu Q, Xu H, Liu Y, She P, Tan R, Wu Y (2016) The effect of berberine hydrochloride on Enterococcus faecalis biofilm formation and dispersion in vitro. Microbiol Res 186-187:44–51. https://doi.org/10.1016/j.micres.2016.03.003
Chu M, Ding R, Z-y C, M-b Z, X-y L, Xie S-h, Y-j Z, Wang Y-d (2014) Role of berberine in anti-bacterial as a high-affinity LPS antagonist binding to TLR4/MD-2 receptor. BMC Complem Altern M 14:89–89. https://doi.org/10.1186/1472-6882-14-89
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97(12):6640–6645. https://doi.org/10.1073/pnas.120163297
Eran Z, Akçelik M, Yazıcı BC, Özcengiz G, Akçelik N (2020) Regulation of biofilm formation by marT in Salmonella Typhimurium. Mol Biol Rep 47(7):5041–5050. https://doi.org/10.1007/s11033-020-05573-6
Fàbrega A, Vila J (2013) Salmonella enterica serovar Typhimurium skills to succeed in the host: virulence and regulation. Clin Microbiol Rev 26(2):308–341. https://doi.org/10.1128/CMR.00066-12
Flemming H-C, Wingender J, Szewzyk U, Steinberg P, Rice SA, Kjelleberg S (2016) Biofilms: an emergent form of bacterial life. Nat Rev Microbiol 14(9):563–575. https://doi.org/10.1038/nrmicro.2016.94
Habtemariam S (2020) Berberine pharmacology and the gut microbiota: a hidden therapeutic link. Pharmacol Res 155:104722. https://doi.org/10.1016/j.phrs.2020.104722
Hospenthal MK, Costa TRD, Waksman G (2017) A comprehensive guide to pilus biogenesis in gram-negative bacteria. Nat Rev Microbiol 15(6):365–379. https://doi.org/10.1038/nrmicro.2017.40
Imenshahidi M, Hosseinzadeh H (2016) Berberis vulgaris and berberine: an update review. Phytother Res 30(11):1745–1764. https://doi.org/10.1002/ptr.5693
Karaosmanoglu K, Sayar NA, Kurnaz IA, Akbulut BS (2013) Assessment of berberine as a multi-target antimicrobial: a multi-omics study for drug discovery and repositioning. OMICS: OMICS 18(1):42–53. https://doi.org/10.1089/omi.2013.0100
Koo H, Allan RN, Howlin RP, Stoodley P, Hall-Stoodley L (2017) Targeting microbial biofilms: current and prospective therapeutic strategies. Nat Rev Microbiol 15(12):740–755. https://doi.org/10.1038/nrmicro.2017.99
Korhonen TK (1979) Yeast cell agglutination by purified enterobacterial pili. FEMS Microbiol Lett 6(6):421–425. https://doi.org/10.1111/j.1574-6968.1979.tb03756.x
Korhonen TK, Lounatmaa K, Ranta H, Kuusi N (1980) Characterization of type 1 pili of Salmonella Typhimurium LT2. J Bacteriol 144(2):800–805. https://doi.org/10.1128/JB.144.2.800-805.1980
Kuan N-L, Yeh K-S (2019) Arginine within a specific motif near the N-terminal of FimY is critical for the maximal production of type 1 fimbriae in Salmonella enterica serovar Typhimurium. MICROBIOLOGYOPEN 8(9):e00846–e00846. https://doi.org/10.1002/mbo3.846
Kuehn B (2019) Multidrug-resistant Salmonella. JAMA 322(14):1344–1344. https://doi.org/10.1001/jama.2019.15309
Kumar A, Ekavali CK, Mukherjee M, Pottabathini R, Dhull DK (2015) Current knowledge and pharmacological profile of berberine: an update. Eur J Pharmacol 761:288–297. https://doi.org/10.1016/j.ejphar.2015.05.068
Ledeboer NA, Frye JG, McClelland M, Jones BD (2006) Salmonella enterica serovar Typhimurium requires the Lpf, Pef, and Tafi fimbriae for biofilm formation on HEp-2 tissue culture cells and chicken intestinal epithelium. Infect Immun 74(6):3156–3169. https://doi.org/10.1128/IAI.01428-05
Liu Q, Niu H, Zhang W, Mu H, Sun C, Duan J (2015) Synergy among thymol, eugenol, berberine, cinnamaldehyde and streptomycin against planktonic and biofilm-associated food-borne pathogens. Lett Appl Microbiol 60(5):421–430. https://doi.org/10.1111/lam.12401
Liu Y, Liu X, Zhang N, Yin M, Dong J, Zeng Q, Mao G, Song D, Liu L, Deng H (2020) Berberine diminishes cancer cell PD-L1 expression and facilitates antitumor immunity via inhibiting the deubiquitination activity of CSN5. Acta Pharm Sin B 10:2299–2312. https://doi.org/10.1016/j.apsb.2020.06.014
Lukaszczyk M, Pradhan B, Remaut H (2019) The biosynthesis and structures of bacterial pili. In: Kuhn A (ed) Bacterial cell walls and membranes. Springer International Publishing, Cham, pp 369–413
Meng X, Meng X, Wang J, Wang H, Zhu C, Ni J, Zhu G (2019) Small non-coding RNA STnc640 regulates expression of fimA fimbrial gene and virulence of Salmonella enterica serovar Enteritidis. BMC Vet Res 15(1):319–319. https://doi.org/10.1186/s12917-019-2066-7
Michael GB, Schwarz S (2016) Antimicrobial resistance in zoonotic nontyphoidal Salmonella: an alarming trend? Clin Microbiol Infec 22(12):968–974. https://doi.org/10.1016/j.cmi.2016.07.033
Proft T, Baker EN (2008) Pili in gram-negative and gram-positive bacteria — structure, assembly and their role in disease. Cell Mol Life Sci 66(4):613–635. https://doi.org/10.1007/s00018-008-8477-4
Roy R, Tiwari M, Donelli G, Tiwari V (2018) Strategies for combating bacterial biofilms: a focus on anti-biofilm agents and their mechanisms of action. VIRULENCE 9(1):522–554. https://doi.org/10.1080/21505594.2017.1313372
Shi C, Li M, Muhammad I, Ma X, Chang Y, Li R, Li C, He J, Liu F (2018) Combination of berberine and ciprofloxacin reduces multi-resistant Salmonella strain biofilm formation by depressing mRNA expressions of luxS, rpoE, and ompR. J Vet Sci 19(6):808–816. https://doi.org/10.4142/jvs.2018.19.6.808
Stapels DAC, Hill PWS, Westermann AJ, Fisher RA, Thurston TL, Saliba A-E, Blommestein I, Vogel J, Helaine S (2018) Salmonella persisters undermine host immune defenses during antibiotic treatment. Science 362(6419):1156–1160. https://doi.org/10.1126/science.aat7148
Steenackers H, Hermans K, Vanderleyden J, De Keersmaecker SCJ (2012) Salmonella biofilms: an overview on occurrence, structure, regulation and eradication. Food Res Int 45(2):502–531. https://doi.org/10.1016/j.foodres.2011.01.038
Sun D, Abraham SN, Beachey EH (1988) Influence of berberine sulfate on synthesis and expression of pap fimbrial adhesin in uropathogenic Escherichia coli. Antimicrob Agents CH 32(8):1274–1277. https://doi.org/10.1128/aac.32.8.1274
Uchiya K-I, Kamimura Y, Jusakon A, Nikai T (2019) Salmonella fimbrial protein FimH is involved in expression of proinflammatory cytokines in a toll-like receptor 4-dependent manner. Infect Immun 87(3):e00881–e00818. https://doi.org/10.1128/IAI.00881-18
Wang K-C, Hsu Y-H, Huang Y-N, Yeh K-S (2012) A previously uncharacterized gene stm0551 plays a repressive role in the regulation of type 1 fimbriae in Salmonella enterica serotype Typhimurium. BMC Microbiol 12:111–111. https://doi.org/10.1186/1471-2180-12-111
Wolfmeier H, Pletzer D, Mansour SC, Hancock REW (2018) New perspectives in biofilm eradication. ACS Infect DIS 4(2):93–106. https://doi.org/10.1021/acsinfecdis.7b00170
Yang Y, Li J, Yin Y, Guo D, Jin T, Guan N, Shi Y, Xu Y, Liang S, Xia X, Shi C (2019) Antibiofilm activity of coenzyme Q0 against Salmonella Typhimurium and its effect on adhesion–invasion and survival–replication. Appl Microbiol Biot 103(20):8545–8557. https://doi.org/10.1007/s00253-019-10095-8
Zeiner SA, Dwyer BE, Clegg S (2012) FimA, FimF, and FimH are necessary for assembly of type 1 fimbriae on Salmonella enterica serovar Typhimurium. Infect Immun 80(9):3289–3296. https://doi.org/10.1128/IAI.00331-12
Zeng L, Zhang L, Wang P, Meng G (2017) Structural basis of host recognition and biofilm formation by Salmonella Saf pili. Elife 6:e28619. https://doi.org/10.7554/eLife.28619
Funding
This work was supported by the National Natural Science Foundation of China (31770109) and (21775036).
Author information
Authors and Affiliations
Contributions
CX designed, supervised the experiments, analyzed the results, revised the first draft, and prepared the last draft of the manuscript. LD performed part of the experiments, contributed significantly to analysis and manuscript preparation, and revised the last draft of the manuscript. FH and FW performed part of the experiments, analyzed the data, and participated in the first draft of the manuscript. DH analyzed the data and helped perform the analysis with constructive discussions. MY collaborated in the design of the experiments and revised different versions of the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no conflict of interest.
Additional information
This paper is our original work. It has not been submitted elsewhere, and it is not under consideration in any other Journal.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xu, C., Wang, F., Huang, F. et al. Targeting effect of berberine on type I fimbriae of Salmonella Typhimurium and its effective inhibition of biofilm. Appl Microbiol Biotechnol 105, 1563–1573 (2021). https://doi.org/10.1007/s00253-021-11116-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11116-1