Abstract
Salmonella Typhimurium, a common Gram-negative foodborne pathogen, threatens public health and hinders the development of the food industry. In this study, we evaluated the antibiofilm activity of coenzyme Q0 (CoQ0) against S. Typhimurium. Besides, the inhibition of the S. Typhimurium’s adhesion to and invasion of Caco-2 cells and its survival and replication in RAW 264.7 cells by CoQ0 were also explored. The minimum inhibitory concentrations and minimal bactericidal concentrations of CoQ0 against Salmonella were both 100–400 μg/mL. Salmonella Typhimurium biofilm formation was effectively inhibited by subinhibitory concentrations (SICs) of CoQ0. The CoQ0-affected biofilm morphology was observed with light microscopy and field-emission scanning electron microscopy. CoQ0 at SICs reduced the swimming motility and quorum sensing of S. Typhimurium and repressed the transcription of critical virulence-related genes. CoQ0 at SICs also clearly reduced the adhesion of S. Typhimurium to and its invasion of Caco-2 cells and reduced its survival and replication within RAW 264.7 macrophage cells. These findings suggest that CoQ0 has strong antibiofilm activity and can be used as an anti-infectious agent against Salmonella.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salmonella is a ubiquitous, Gram-negative, flagellated bacillus belonging to the family Enterobacteriaceae (Mathur et al. 2012). Among its more than 2600 serovars, defined according to its surface antigens, Salmonella enterica serovar Typhimurium is one of the commonest causes of human infection (Fàbrega and Vila 2013). The ingestion of food products contaminated with S. Typhimurium, such as poultry, beef, pork, eggs, milk, seafood, and fresh produce, can cause many diseases, including gastroenteritis, with symptoms such as diarrhea, vomiting, cramps, fever, and headache (Gal-Mor 2019). It is estimated that food contaminated with Salmonella species is responsible for 94 million cases of gastroenteritis annually, which result in 155,000 deaths globally every year (Ros-Chumillas et al. 2017). In total, 94,530 cases of Salmonella infection were confirmed by the European Union in 2016, with an incidence of 20.4 cases per 100,000 (European Food Safety Authority (EFSA) 2017).
Biofilm formation always renders foodborne pathogens more persistent and resistant to antimicrobial stress, limited nutrient supply, and inappropriate pH or temperature, increasing their threat to public health and posing a huge challenge for food industries worldwide (Fàbrega and Vila 2013). Salmonella can form biofilms on various surfaces, including poultry skin, stainless steel, concrete, tile, glass, granite, rubber, and polyvinyl surfaces (Cappitelli et al. 2014; Dhowlaghar et al. 2018). Stepanović et al. (2004) isolated 122 Salmonella spp. from humans, animals, and food and confirmed that all of the strains had the capacity to form a biofilm. The formation of biofilms by pathogens is involved in over 80% of microbial infections (Bjarnsholt et al. 2018). The ability to form a biofilm is also thought to be a major virulence factor in Salmonella and is closely related to quorum sensing (QS) and motility (Chakroun et al. 2018). The chemical signaling molecules of the QS system, called “autoinducers” (AIs), are reported to activate the signal transduction cascade that regulates biofilm formation (Ng and Bassler 2009). The initial attachment of pathogens to the surfaces of biotic or abiotic objects depends on their motility, and these adherent cells constitute the basis of the biofilm (Srey et al. 2013).
Pathogenic Salmonella strains are quite different from nonpathogenic Salmonella strains because they express virulence genes, which are usually organized into regions known as “Salmonella pathogenicity islands” (SPIs) (Kaur and Jain 2012). SPI-1 encodes a needle-like structure, termed “type III secretion system 1” (T3SS-1). For S. Typhimurium, T3SS-1 is directly related to the biofilm formation, the bacterium’s ability to invade the host intestinal epithelial cells, and the initiation of inflammation (Roche et al. 2018). “Type III secretion system 2” (T3SS-2), encoded by SPI-2, contributes to the survival of Salmonella in macrophages, the spread of the bacterium, and systemic infections (Zhao et al. 2016).
Fluoroquinolones, trimethoprim-sulfamethoxazole, ampicillin, and expanded-spectrum cephalosporins are used to efficiently treat Salmonella (Fàbrega and Vila 2013). However, antibiotic resistance has disseminated rapidly for many reasons, including the overuse or misuse of antibiotics and some complex ancient antibiotic resistance mechanisms bacteria (Bao et al. 2018). Consequently, it is essential to find a substitute for antibiotics to control bacterial contamination. Natural plant products have the potential to inhibit bacterial virulence rather than the viability of the pathogen, resulting in less-severe infections and more repaid immune responses in their hosts (Silva et al. 2016).
Coenzyme Q is a member of the ubiquinone compound family, consists of a benzoquinone ring conjugated to an isoprenoid chain. The number of isoprenoid units determines the name of the coenzyme (Q0-Q10) (Fan et al. 2018). Coenzyme Q0 (CoQ0, 2,3-dimethoxy-5-methyl-1,4-benzoquinone) is a natural compound containing no isoprenoid unit extracted from the filtrates of submerged cultures of Antrodia cinnamomea (Chung et al. 2014). Zhao et al. (2014) and Fan et al. (2018) demonstrated that CoQ0 has antimicrobial activity against Staphylococcus aureus and Listeria monocytogenes, respectively. Yang et al. (2016) proved that CoQ0 inhibited LPS-induced inflammation and the redox imbalance in RAW 264.7 cells and mice. Chung et al. (2014) reported that CoQ0 induced reactive oxygen species generation and apoptosis in A549 human lung cancer cells. Moreover, CoQ0 has been demonstrated to have anti-metastatic (Yang et al. 2019) and anti-angiogenic (Yang et al. 2015) activities.
Although the antimicrobial characteristics of CoQ0 have been extensively investigated, little research has been directed towards its antibiofilm and anti-infection activities against Salmonella. In this study, the effects of CoQ0 on the biofilm formation, QS, and motility of S. Typhimurium were evaluated. The effects of CoQ0 treatment on the capacities of S. Typhimurium to adhere to and invade Caco-2 cells and to survive and replicate in RAW 264.7 cells were also investigated. Finally, the mechanism by which CoQ0 regulates the transcription levels of critical virulence-related genes of S. Typhimurium was examined.
Materials and methods
Reagents
Coenzyme Q0 (HPLC > 99%, CAS 605-94-7) was purchased from J&K Scientific Co., Ltd (Beijing, China). Before each assay, CoQ0 was dissolved in dimethyl sulfoxide (DMSO) and vortexed for 30 s at room temperature. The final concentration of DMSO in all of the sample solutions (treatment and control samples) was 0.1% (v/v), which has no apparent effect on the growth of S. Typhimurium.
Bacterial strains and culture conditions
Salmonella Typhimurium SL1344 (DSM 24522) was obtained from the German Collection of Microorganisms and Cell Cultures (DSMZ, Braunschweig, Germany), S. Typhimurium ATCC 14028, and Chromobacterium violaceum ATCC 12472 were obtained from the American Type Culture Collection (ATCC; Manassas, VA, USA). Salmonella Typhimurium CMCC 50115 was obtained from the National Center for Medical Culture Collections (CMCC, Beijing, China). Nine other Salmonella strains were from our laboratory strain collection and were originally isolated from various raw chicken samples in China. All of the Salmonella isolates were used in minimum inhibitory concentration (MIC) and minimal bactericidal concentration (MBC) assays. Salmonella Typhimurium SL1344 was used for further experiments because it is commonly used in Salmonella virulence studies and its phenotypic and genotypic characteristics are well-documented (Li et al. 2014). Because S. Typhimurium ATCC 14028 displayed stronger biofilm production, it was used to perform the assays related to biofilm. Chromobacterium violaceum ATCC 12472 was used as the QS indicator strain to evaluate the effect of CoQ0 on the QS system (Ma et al. 2018).
Before each assay, Luria–Bertani (LB) agar was streak-inoculated with the Salmonella strains, which were cultured at 37 °C for 12 h to activate the bacteria. To obtain fresh overnight cultures, 30 mL of LB broth was inoculated with a single colony and incubated at 37 °C for 12 h with shaking at 130 rpm. The culture was then diluted in LB broth to an optical density at a wavelength of 600 nm (OD600 nm) of 0.5 (approximately 109 CFU/mL).
Determination of MICs and MBCs
The MICs of CoQ0 against the test Salmonella strains were determined with the broth dilution method based on the Clinical and Laboratory Standards Institute guidelines (CLSI 2009). Each well of a 96-well plate was inoculated with 200 μL of diluted bacterial strain culture, with a final bacterial concentration of approximately 5 × 105 CFU/mL. CoQ0 solution was added to each well to final concentrations of 800, 400, 200, 100, 50, 25, or 0 μg/mL. LB broth containing 0.1% DMSO was used as the negative control. The samples were then incubated at 37 °C for 24 h. The MIC of CoQ0 was the lowest concentration of CoQ0 that resulted in no visible Salmonella growth. To determine the MBCs, 100 μL of bacterial solution from each well was plated on an LB agar plate and cultured for 24 h. The MBC of CoQ0 was defined as the minimum concentration of CoQ0 that killed 99.9% of the test bacteria.
Determination of the subinhibitory concentrations (SICs)
The SICs of CoQ0 against S. Typhimurium SL1344 and S. Typhimurium ATCC 14028 were determined with the Bioscreen C Automated Microbiology Growth Curve Analysis System (Labsystems, Helsinki, Finland). The bacterial suspension was diluted to 5 × 105 CFU/mL with LB broth and 125 μL of the diluted culture was added to the individual cells of 100-well plates. Equal volumes of LB broth containing different concentrations of CoQ0 were added to the individual wells to achieve final CoQ0 concentrations of 400, 200, 100, 50, 25, 12.5, 6.25, 3.125, 1.5625, 0.78125, or 0 μg/mL. LB broth containing 0.1% DMSO was used as the negative control. The samples were further cultured at 37 °C for 24 h, and the OD600 nm was monitored automatically every 1 h. Concentrations of CoQ0 that showed no inhibitory effect on bacterial growth were deemed to be SICs.
Motility assay
A motility assay was performed on medium containing 20 mL of LB broth and 0.3% (v/v) agar, as described by Li et al. (2014) with some modifications. CoQ0 was added to the medium at a final concentration of 25, 12.5. 6.25, 3.125, or 0 μg/mL, and the plates were left to dry for 1 h. The center of this semisolid medium surface was inoculated with 5 μL of overnight culture (OD600 nm = 0.5) and the samples were incubated upright at 37 °C for 7 h. Images of the motile bacteria were collected with the Gel Imaging System (Bio-Rad, USA), and the ImageJ software was used to measure the diameters of the areas of motility.
Specific biofilm formation (SBF) inhibition assay
To evaluate the effect of CoQ0 on the SBF of S. Typhimurium SL1344 and ATCC 14028, the crystal violet staining method was used, with some modifications (Shi et al. 2017). Tryptic soy broth (TSB, 30 mL) was inoculated with a single colony of S. Typhimurium and incubated at 37 °C overnight. CoQ0 was added to the overnight bacterial suspension (OD600 nm = 1) to a concentration of 25, 12.5, 6.25, or 0 μg/mL, and then 200 μL of the incubated mixture was added to the wells of a 96-well plate. TSB containing 0.1% DMSO was used as the negative control. The samples were incubated at 37 °C for 24, 48, or 72 h. After incubation, the absorbance of each culture was measured at 630 nm and the wells were washed with distilled water. The plates were air-dried for 30 min, and 200 μL of 0.1% (w/v) crystal violet was added to each well for 20 min at room temperature to stain the biofilm. The uncombined dye was then removed by washing the wells three times with 300 μL of distilled water. After the wells were dried, the bound dye was solubilized in 200 μL of 33% (v/v) glacial acetic acid and the plates were shaken for 20 min at 25 °C. The ODs were read at 570 nm with a microtiter spectrophotometer. SBF was determined by calibrating the OD570 nm with the OD630 nm.
Light microscope analysis
To visualize the effect of CoQ0 on biofilm formation by S. Typhimurium ATCC 14028, the method of Bai et al. (2019) was used. Briefly, glass slides of the same size were placed in each well of a 24-well plate, and were covered with S. Typhimurium ATCC 14028 bacterial suspension (OD600 nm = 1) treated with 25, 12.5, 6.25, or 0 μg/mL CoQ0. After incubation at 37 °C for 72 h, the fluid in each well was removed and the plate was washed twice with phosphate-buffered saline (PBS). The glass slides were removed and stained with 0.4% crystal violet solution for 20 min. They were then washed three times with 300 μL of distilled water to remove any excess stain and air-dried. A light microscope (BX53, Olympus, Tokyo, Japan) was used to observe the stained biofilms.
Field-emission scanning electron microscopy (FESEM)
Biofilms were formed on glass slides as described in "Light microscope analysis" section. After the removal of planktonic cells, PBS containing 2.5% glutaraldehyde was added to fix the biofilm for 12 h at 4 °C. The glass slides were then washed twice with PBS and dehydrated with a series of ethanol (50%, 70%, 80%, 90%, and 100%) to fix the cells. The glass slides were dried at room temperature and coated with gold, and the samples were examined under a field-emission scanning electron microscope (S-4800, Hitachi, Tokyo, Japan).
Quantitative QS inhibition assay
The QS indicator strain C. violaceum ATCC 12472 was used to determine the effect of CoQ0 on the QS-inhibitory activity of CoQ0 by quantifying its violacein production (Ma et al. 2018). First, the effects of SICs of CoQ0 on the growth of C. violaceum ATCC 12472 were determined using the method described in "Determination of the subinhibitory concentrations" Section. The three highest concentrations of CoQ0 that had no effect on the growth of C. violaceum ATCC 12472 were selected as the SICs for subsequent experiments.
An overnight culture of C. violaceum ATCC 12472, diluted to OD600 nm = 0.2, was added to LB broth containing CoQ0 at a concentration of 12.5, 6.25, 3.125, or 0 μg/mL. The samples were cultured at 30 °C for 24 h with shaking at 130 rpm. Violacein was extracted and quantified according to the method described by Choo et al. (2006) with some modifications. Briefly, 3 mL of culture was centrifuged (5000×g, 5 min, 4 °C) to precipitate the insoluble violacein. The liquid supernatant was discarded and 1 mL of DMSO was added to the centrifuge tube. After shaking, the mixture was centrifuged (10,000×g, 10 min, 4 °C) to remove the cells. The violacein solution (200 μL) was added to 96-well microtiter plates and the OD585 nm was read.
Adhesion and invasion of Caco-2 cells
In this study, the human colon carcinoma cell line, Caco-2, was obtained from the American Type Culture Collection and cultured according to the method described by Shi et al. (2017). The effects of CoQ0 on bacterial adhesion and invasion were determined according to the method described by Fan et al. (2018) with some modifications. Briefly, Caco-2 cells were seeded in a 24-well plate, grown in DMEM with 10% FBS (105 cells per well) at 37 °C and 5% CO2 for 18 h, and then rinsed twice with PBS. S. Typhimurium SL1344 bacterial strain culture was added to LB broth containing different concentrations (25, 12.5, 6.25, or 0 μg/mL) of CoQ0 and cultured at 37 °C for 12 h with shaking at 130 rpm before Caco-2 cells adhesion and invasion assays. The bacterial suspensions were centrifuged and resuspended in PBS to remove any residual CoQ0. After then, the samples were diluted 1000 times with DMEM and added to each well at a multiplicity of infection (MOI) of 10. The samples were incubated at 37 °C in a humidified 5% CO2 incubator for 1 h.
For the adhesion assay, the infected monolayers of cells were washed twice with PBS and lysed with 1 mL of 0.1% Triton X-100. The number of S. Typhimurium SL1344 cells was counted after they were serially diluted, plated onto LB agar, and incubated for 24 h at 37 °C. For the invasion assay, the Caco-2 monolayers were washed once with PBS and incubated for 1 h with 1 mL of DMEM containing gentamicin (100 μg/mL) to kill the extracellular bacteria. The monolayers were then washed three times and lysed with 0.1% Triton X-100 at 4 °C. The number of bacteria was counted by colony plating, as described for the adhesion assay. The adhesion and invasion rates of S. Typhimurium in the control groups were taken to be 100%, and those for the treatment groups were calculated as a percentage of the control value.
Survival and intracellular replication in macrophages
In this study, the murine macrophage cell line, RAW 264.7, was obtained from the American Type Culture Collection. The intracellular survival and replication of S. Typhimurium SL1344 in RAW 264.7 were examined according to the method described by Ryan et al. (2018) with minor modifications. Briefly, RAW 264.7 cells were seeded in a 24-well plate and cultured in DMEM with 10% FBS (105 cells per well) at 37 °C and 5% CO2 for 12 h. S. Typhimurium SL1344 bacterial suspension was pre-treated with or without CoQ0 (12.5, 6.25, 3.125, or 0 μg/mL) before adding into each well containing RAW 264.7 cells. After incubation overnight, the bacterial suspension was washed twice with PBS to remove any extra CoQ0, and diluted to a concentration of OD600 nm = 0.5. After that, the samples were diluted 1000 times with DMEM, yielding a final bacterial concentration of approximately 106 CFU/mL. The bacterial suspension (1 mL, MOI = 10) was added to each well containing RAW 264.7 cells. After subsequent incubation for 45 min, the cells were washed once with PBS and incubated for 30 min with 1 mL of DMEM containing gentamicin (100 μg/mL) to kill any extracellular bacteria.
For the survival assay, the macrophages were lysed by the addition of 0.1% Triton X-100 for 20 min at 4 °C and then serially diluted and plated on LB agar. To estimate intracellular replication, 1 mL of DMEM containing 10 μg/mL gentamicin was added to each well. After incubation for 24 or 72 h, the cells were lysed and the appropriate dilution was plated on LB agar plates, as described in the survival assay. The number of bacteria was counted after incubation for 24 h.
RNA extraction and reverse transcription–quantitative PCR (RT–qPCR)
To determine the effect of CoQ0 on the transcription of motility-, biofilm formation–, invasion-, and virulence-related genes (Table 1), S. Typhimurium SL1344 was grown in LB broth containing different concentrations (25, 12.5, or 0 μg/mL) of CoQ0 at 37 °C for 8 h. Total RNA was extracted with the Tiangen RNAprep Pure Cell/Bacteria Kit (Tiangen, Beijing, China), according to the manufacturer’s instructions. The RNA quality and concentration were determined with a nucleic acid and protein spectrophotometer (Nano-200, Aosheng Instrument Co., Ltd, Hangzhou, China). The Takara PrimeScript™ RT Reagent Kit (Takara, Kyoto, Japan) was used to reverse transcribe the RNA into cDNA, according to the manufacturer’s instructions. The cDNA samples were stored at − 20 °C until analysis. The primer sequences used for RT–qPCR are listed in Table 1.
The qPCR reactions (25 μL) with SYBR® Premix Ex Taq™ II (Takara) were performed using the IQ™5 system (Bio-Rad) with the following cycling conditions: initial denaturation at 95 °C for 30 s, followed by 40 cycles of denaturation at 95 °C for 5 s, annealing at 60 °C for 30 s, and a dissociation step at 95 °C for 15 s and 60 °C for 30 s. The relative gene transcription in the samples was analyzed using the 2−ΔΔCt method, relative to the reference gene 16s rRNA.
Statistical analysis
All statistical analyses were performed with SPSS 23.0 (IBM, New York, NY, USA). The data are presented as means ± standard deviations (SD) and the differences between the means were tested with the independent-sample Student’s t test. P < 0.05 and P < 0.01 were considered statistically significant and extremely significant, respectively. All experiments were measured independently three times.
Results
MICs and MBCs
The MICs and MBCs of CoQ0 against Salmonella spp. are listed in Table 2. The MICs of CoQ0 against Salmonella ranged from 100 to 400 μg/mL. The MBC values were equal to the MIC values.
SICs
As can be seen in Fig. 1a and b, the growth conditions of S. Typhimurium SL1344 were similar to those for S. Typhimurium ATCC 14028. When the concentrations of CoQ0 ranged from 50 to 100 μg/mL, the lag phase of S. Typhimurium was lengthened. When the concentration of CoQ0 was reduced to 25 μg/mL or below, there was no apparent effect on the growth of S. Typhimurium. Therefore, concentrations of 25, 12.5, 6.25, and 3.125 μg/mL CoQ0 were selected as SICs to study the effects of CoQ0 on the virulence of S. Typhimurium.
Motility
The motility of S. Typhimurium SL1344 was reduced by CoQ0 (Fig. 2a–f). The area of motility of untreated S. Typhimurium SL1344 was 16.75 ± 0.06 cm2, whereas those of the strains treated with 25, 12.5, 6.25, or 3.125 μg/mL, CoQ0 were 7.27 ± 0.04 cm2, 9.57 ± 0.08 cm2, 11.93 ± 0.05 cm2, or 12.02 ± 0.06 cm2, respectively (about 43.37%, 57.14%, 71.24%, or 71.75% of the control motility area, respectively).
Swimming motility of S. Typhimurium on soft agar plates containing different concentrations of CoQ0. a Untreated cells; b, c, d, e cells treated with 3.125, 6.25, 12.5, or 25 mg/mL CoQ0, respectively. f Quantification of S. Typhimurium SL1344 swimming motility in the presence of CoQ0. Relative swimming motility area of the strain was measured after treatment with CoQ0. Values are normalized to the 100% motility area measured in the absence of CoQ0. Bars indicate means ± the standard deviation. *P < 0.05, **P < 0.01
Inhibition of biofilm formation by CoQ0
As shown in Fig. 3a and b, the formation of biofilm by S. Typhimurium SL1344 and ATCC 14028 grown at 37 °C for 24, 48, or 72 h was effectively and concentration-dependently inhibited by CoQ0 at SICs. For S. Typhimurium ATCC 14028, the biofilm biomass treated with 6.25, 12.5, or 25 μg/mL CoQ0 was reduced by 21.93%, 39.53%, or 48.70% of the control level, respectively, after 24 h. At 25 μg/mL, CoQ0 inhibited the biofilm formation of S. Typhimurium SL1344 after 24, 48, or 72 h by 55.39–60.77% compared with that of the control.
Light microscopic and FESEM observations
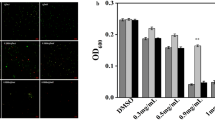
The S. Typhimurium ATCC 14028 biofilm was observed at a × 400 magnification by light microscopy (Fig. 4a–d). The biofilms of CoQ0-treated S. Typhimurium were markedly reduced on the glass slides. The biofilm biomass was also dose-dependently reduced by CoQ0 treatment.
Light microscopic images of S. Typhimurium ATCC 14028 biofilm in the presence of CoQ0 at concentrations of 0 (a), 3.125 (b), 6.25 (c), or 12.5 mg/mL (d). Scanning electron microscopic images at a × 1500 magnification of S. Typhimurium ATCC 14028 biofilm after treatment with CoQ0 at 0 (e), 3.125 (f), 6.25 (g), or 12.5 mg/mL (h). Scanning electron microscopic images at × 4000 magnification of S. Typhimurium ATCC 14028 biofilm after treatment with CoQ0 at 0 (g), 3.125 (h), 6.25 (i), or 12.5 mg/mL (l)
As shown in Fig. 4e–l, FESEM observations at × 1500 and × 4000 magnification demonstrated that the formation of S. Typhimurium ATCC 14028 biofilm on the surfaces of glass slides was inhibited by CoQ0 at SICs, consistent with the crystal violet staining results. After incubation for 72 h, untreated S. Typhimurium showed a dense biofilm layer, and most cells had gathered into large clusters and multilayer structures (Fig. 4e). In contrast to the control, as the concentration of CoQ0 increased, Salmonella displayed severe damage to the typical biofilm structure and lower biofilm cell densities (Fig. 4b–d).
Anti-QS activity of CoQ0
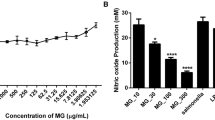
Figure 5a shows that the concentrations of CoQ0 used in this study (25, 12.5, and 6.25 μg/mL) had no apparent inhibitory effect on the growth of C. violaceum. As seen in Fig. 5b, the production of violacein decreased to about 89.4%, 87.0%, or 68.8% of the control after treatment with 3.125, 6.25, or 12.5 μg/mL CoQ0, respectively.
a Growth of C. violaceum ATCC 12472 in LB with various concentrations of CoQ0. Error bars represent the standard deviations of three replicate experiments. b Inhibition of violacein production by C. violaceum ATCC 12472 at different concentrations of CoQ0. Bars represent the means ± standard deviations (n = 3). *P < 0.05, **P < 0.01
Adhesion to and invasion of Caco-2 cells
The effects of CoQ0 on the adhesion of S. Typhimurium to Caco-2 cells are shown in Fig. 6a. Compared with the control, 6.25, 12.5, and 25 μg/mL CoQ0 inhibited the adhesion of S. Typhimurium to 59.1%, 50.9%, and 52.3% of the control value, respectively. The invasion of Caco-2 cells by S. Typhimurium was also inhibited by CoQ0, decreasing to 84.3%, 69.8%, and 40.6% of the control value, respectively, at the SICs (Fig. 6b).
Survival and replication within macrophage cells
As shown in Fig. 7, only 25 μg/mL CoQ0 significantly inhibited the intracellular survival of S. Typhimurium in RAW264.7 cells (P < 0.01), and the quantity of intracellular bacteria decreased by 42.9% of the control level. However, the intracellular replication of S. Typhimurium in RAW264.7 cells was significantly reduced by all of the SICs of CoQ0 in a dose-dependent manner relative to the control after 72 h (P < 0.01). Salmonella Typhimurium replication in RAW264.7 cells decreased by 80.7–87.1% in the presence of 6.25, 12.5, or 25 μg/mL CoQ0.
RT–qPCR analysis of virulence-related genes
CoQ0 at SICs downregulated the transcription of 13 genes in S. Typhimurium SL1344 that are associated with its virulence (Table 1). CoQ0 downregulated the transcription of genes fljB and flhD (critical for flagellar regulation); fimD (motility); arcZ, sroC, csrB, csgD, and adrA (biofilm formation); hilA, invA, and invH (adherence and invasion); pipB and orf245 (T3SS); sdiA and srgE (QS); and sodC (survival in macrophages) to various degrees.
Discussion
Motility allows Salmonella to form biofilm, to approach the host cell, and to initiate adhesion and invasion processes, so that it can successfully infect mammalian cells (Fàbrega and Vila 2013). In this study, the swimming motility of Salmonella was significantly inhibited by CoQ0 at SICs (P < 0.05) (Fig. 2). Similarly, a previous study reported that the swimming motility of Escherichia coli O157:H7 was dose-dependently blocked by grape seed extract (Zhu et al. 2015). Li et al. (2014) reported that the areas of motility were reduced to 23.17% and 39.01% of the control area by 15.625 and 31.25 μg/mL punicalagin, respectively, while the flagellum-associated genes were downregulated. The flagellum is a component of Salmonella that is essential for its motility, and also contributes to its chemotaxis, adherence to and invasion of host cells, colonization, and even subsequent innate immune signaling (Horstmann et al. 2017). The fijB gene encodes the flagellin protein, which forms the filament structure of the flagellar apparatus and is considered a primary proinflammatory determinant of Salmonella (Zeng et al. 2003). The flhD gene encodes the flagellar switch protein FlhD4C2, which is the flagellar operon master regulator and the transcriptional activator of all of the flagellar genes. The fimD gene partly encodes the type 1 fimbriae, which reportedly play roles in adherence, invasion, biofilm formation, and the immune response (Li et al. 2017). Our results show that all three genes, fijB, flhD, and fimD, were repressed at the transcriptional level by CoQ0 at different SICs. Therefore, it can be inferred that CoQ0 restricts the motility of Salmonella by mediating the function of the flagellum, which may influence other features of infection, such as adherence, invasion, and biofilm formation.
Salmonella can form biofilms on various surfaces, including stainless steel, aluminum, plastic, and glass (Merino et al. 2019). The great damage caused by Salmonella to the food processing industry is mainly attributable to the persistence and resistance of mature biofilms (Shi and Zhu 2009). Our results show that CoQ0 at SICs exerts a significant inhibitory effect on the biofilm formation of S. Typhimurium on polystyrene and glass surfaces (Figs. 3 and 4). These results are similar to those of a study of S. aureus, in which its biofilm formation was effectively inhibited by 0.3125 mg/mL shikimic acid, as determined by confocal laser scanning microscopy, light microscopy, and scanning electron microscopy, which were used to analyze the biofilm biomass, the viability of the biofilm cells, and the biofilm architecture in different strains (Bai et al. 2019). Similarly, Shi et al. (2017) reported that thymoquinone prevented biofilm formation by Cronobacter sakazakii by inhibiting the production of cellulose and the flagellum. Cellulose is a crucial component of the Salmonella biofilm (Steenackers et al. 2012) and is encoded by csgD and adrA. Some Salmonella small RNAs (sRNAs) have been shown to play a role in biofilm formation (Ryan et al. 2017). In the present study, CoQ0 at SICs downregulated the transcription of various genes, including csgD and adrA, and some sRNAs, including arcZ, sroC, and csrB, which trigger biofilm formation by regulating biofilm-associated genes, such as csgD and the flagellar genes (Fuentes et al. 2015). Therefore, CoQ0 may control the biofilm formation of S. Typhimurium by mediating cellulose and the function of related sRNAs.
QS is a chemical cell-to-cell communication system relying on the small signaling molecules called AIs. By the production, release, and detection of AIs, Salmonella’s biofilm formation, motility, and production of virulence factors could be regulated (Defoirdt et al. 2013). N-Acyl homoserine lactone (AHL), a type of membrane-permeable small chemical compound, is classified as one of these signaling molecules and combines with cytoplasmic receptors to play a regulatory role (Thoendel et al. 2011). In this study, by measuring the concentration of violacein pigment in C. violaceum, we showed that CoQ0 markedly interferes with the production of the QS signaling molecule AHL (Fig. 5). Similarly, Borges et al. (2014) showed that six phenolic products extracted from natural sources reduced AHL-regulated violacein pigment to disturb the QS system. Vinothkannan et al. (2018) reported that fructose furoic acid ester curbed the QS system of E. coli by downregulating sdiA expression and that it is a potential QS inhibitor. The gene sidA encodes an AHL receptor, SdiA (a LuxR homolog), that activates srgE expression and helps S. Typhimurium sense and respond to the AHLs produced by other bacteria (Dyszel et al. 2010). Our RT–qPCR assay demonstrated that CoQ0 significantly downregulated the transcription of sdiA and srgE in Salmonella. We assume that CoQ0 reduced the Salmonella QS system through inhibiting the synthesis of AHL receptor protein. The intracellular physiological state affected by CoQ0 will be investigated in further studies. The pathway or receptor protein by which external CoQ0 enter into the cell will be also studied.
The adhesion to and invasion of host epithelial cells by Salmonella depends on functions encoded by SPI-1. Salmonella Typhimurium SPI-1 T3SS shifts effector proteins into host cells across the plasma membrane so that the bacterium can invade the cells and induce an intestinal inflammatory response (LaRock et al. 2015). Baxter and Jones (2015) reported that hilA, the master transcriptional activator of SPI-1, regulates at least 10 activators and eight repressors, and that cell invasion requires the upregulation of hilA transcription. The invA and invH genes encoded by S. Typhimurium SPI-1 are responsible for stimulating the inflammatory responses in cultured epithelial cells and causing inflammation in the gut (Pati et al. 2013). In the present study, the abilities of Salmonella to adhere to and invade Caco-2 cells were reduced by CoQ0 at SICs, and CoQ0 significantly downregulated hilA, invA, and invH transcription (P < 0.05) (Fig. 6). In a previous study, Wu et al. (2018) reported that 20 μg/mL baicalin inhibited the invasion of Caco-2 cells by S. Typhimurium and downregulated its transcription of the sopB, sopE, and sopE2 genes, which are associated with SPI-1. Likewise, 30 μg/mL methyl gallate inhibited 43.75% of S. Typhimurium adhesion to and invasion of Caco-2 cells and downregulated the transcription of the cheY, ompD, sipB, lexA, and ompF genes, which are essential for its invasion and adhesion. Our results demonstrate that CoQ0 has potential utility as an antibiotic substitute to reduce Salmonella adhesion to and invasion of enterocytes and to ultimately alleviate inflammation in the gut.
During its infection of the gastrointestinal tract, S. Typhimurium targets both epithelial cells and macrophages (Hautefort et al. 2008). After entering the small intestine, Salmonella invades and penetrates the intestinal epithelium, which results in the systemic spread of the bacteria, which are taken up by macrophages (Jiang et al. 2017). Birhanu et al. (2018) have shown that the intracellular survival of S. Typhimurium in RAW 264.7 cells was effectively reduced by 30 μg/mL methyl gallate. In the present study, CoQ0 reduced the ability of S. Typhimurium to survive and reproduce intracellularly (Fig. 7). Our results indicate that CoQ0 reduces the ability of S. Typhimurium to utilize macrophages to evade the immune response and invade bodily organs. Cu, Zn superoxide dismutase has been shown to be expressed in several bacteria, including Salmonella, and confers protection against extracellular reactive oxygen species (Pacello et al. 2008). A deficiency in Cu, Zn superoxide dismutase, which is encoded by the sodC gene, reduces Salmonella survival in macrophages and attenuates its virulence in mice (DeGroote et al. 1997). In the present study, CoQ0 significantly downregulated the transcription of sodC (Table 1). Therefore, we hypothesize that CoQ0 inhibits the survival and replication of S. Typhimurium within macrophages by inhibiting Cu, Zn superoxide dismutase expression, ultimately attenuating Salmonella infection.
In conclusion, SICs of CoQ0 effectively prevent the formation of Salmonella biofilm by reducing the biofilm biomass, collapsing the biofilm architecture, and downregulating the expression of biofilm-associated genes and sRNAs. SICs of CoQ0 also inhibit the swimming motility and QS of S. Typhimurium, its adhesion to and invasion of Caco-2 cells, and its intracellular survival in RAW 264.7 macrophage cells, and repress its transcription of critical virulence-associated genes. Our results demonstrate that CoQ0 has potential utility as an antibiofilm and anti-infectious agent against Salmonella. However, further studies are required to confirm the anti-infection effects of CoQ0 in vivo before its practical application.
References
Bai JR, Zhong K, Wu YP, Elena G, Gao H (2019) Antibiofilm activity of shikimic acid against Staphylococcus aureus. Food Control 95:327–333. https://doi.org/10.1016/j.foodcont.2018.08.020
Bao KF, Yuan WY, Ma CB, Yu X, Wang L, Hong M, Xi XP, Zhou M, Chen TB (2018) Modification targeting the “Rana Box” motif of a novel nigrocin peptide from Hylarana latouchii enhances and broadens its potency against multiple bacteria. Front Microbiol 9:2846. https://doi.org/10.3389/fmicb.2018.02846
Baxter MA, Jones BD (2015) Two-component regulators control hilA expression by controlling fimZ and hilE expression within Salmonella enterica serovar Typhimurium. Infect Immun 83(3):978–985. https://doi.org/10.1128/IAI.02506-14
Birhanu BT, Park NH, Lee SJ, Hossain MA, Park SC (2018) Inhibition of Salmonella Typhimurium adhesion, invasion, and intracellular survival via treatment with methyl gallate alone and in combination with marbofloxacin. Vet Res 49(1):101. https://doi.org/10.1186/s13567-018-0597-8
Bjarnsholt T, Buhlin K, Dufrene YF, Gomelsky M, Moroni A, Ramstedt M, Rumbaugh KP, Schulte T, Sun L, Akerlund B, Romling U (2018) Biofilm formation-what we can learn from recent developments. J Intern Med 284(4):332–345. https://doi.org/10.1111/joim.12782
Borges A, Serra S, Abreu AC, Saavedra MJ, Salgado A, Simoes M (2014) Evaluation of the effects of selected phytochemicals on quorum sensing inhibition and in vitro cytotoxicity. Biofouling 30(2):183–195. https://doi.org/10.1080/08927014.2013.852542
Cappitelli F, Polo A, Villa F (2014) Biofilm formation in food processing environments is still poorly understood and controlled. Food Eng Rev 6(1-2):29–42. https://doi.org/10.1007/s12393-014-9077-8
Chakroun I, Mahdhi A, Morcillo P, Cordero H, Cuesta A, Bakhrouf A, Mahdouani K, Esteban MÁ (2018) Motility, biofilm formation, apoptotic effect and virulence gene expression of atypical Salmonella Typhimurium outside and inside Caco-2 cells. Microb Pathog 114:153–162. https://doi.org/10.1016/j.micpath.2017.11.010
Choo JH, Rukayadi Y, Hwang JK (2006) Inhibition of bacterial quorum sensing by vanilla extract. Lett Appl Microbiol 42(6):637–641. https://doi.org/10.1111/j.1472-765X.2006.01928.x
Chung CH, Yeh SC, Chen CJ, Lee KT (2014) Coenzyme Q0 from Antrodia cinnamomea in submerged cultures induces reactive oxygen species-mediated apoptosis in A549 human lung cancer cells. Evid Based Complement Alternat Med 2014(4):246748–246710. https://doi.org/10.1155/2014/246748
Clinical and Laboratory Standards Institute (2009) Clinical and Laboratory Standards Institute CLSI; CLSI Publishes 2009 Antimicrobial susceptibility testing standards. Atlanta, p 12
Defoirdt T, Brackman G, Coenye T (2013) Quorum sensing inhibitors: how strong is the evidence? Trends Microbiol 21(12):619–624. https://doi.org/10.1016/j.tim.2013.09.006
DeGroote MA, Ochsner UA, Shiloh MU, Nathan C, McCord JM, Dinauer MC, Libby SJ, VazquezTorres A, Xu YS, Fang FC (1997) Periplasmic superoxide dismutase protects Salmonella from products of phagocyte NADPH-oxidase and nitric oxide synthase. Proc Natl Acad Sci U S A 94(25):13997–14001. https://doi.org/10.1073/pnas.94.25.13997
Dhowlaghar N, Abeysundara PDA, Nannapaneni R, Schilling MW, Chang S, Cheng WH, Sharma CS (2018) Biofilm formation by Salmonella spp. in catfish mucus extract under industrial conditions. Food Microbiol:70. https://doi.org/10.1016/j.fm.2017.09.016
Dyszel JL, Smith JN, Lucas DE, Soares JA, Swearingen MC, Vross MA, Young GM, Ahmer BMM (2010) Salmonella enterica serovar Typhimurium can detect acyl homoserine lactone production by Yersinia enterocolitica in mice. J Bacteriol 192(1):29–37. https://doi.org/10.1128/JB.01139-09
European Food Safety Authority (2017) The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2016. https://doi.org/10.2903/j.efsa.2017.5077
Fàbrega A, Vila J (2013) Salmonella enterica serovar Typhimurium skills to succeed in the host: virulence and regulation. Clin Microbiol Rev 26(2):308–341. https://doi.org/10.1128/CMR.00066-12
Fan Q, Zhang Y, Yang H, Wu Q, Shi C, Zhang C, Xia X, Wang X (2018) Effect of coenzyme Q0 on biofilm formation and attachment-invasion efficiency of Listeria monocytogenes. Food Control 90:274–281. https://doi.org/10.1016/j.foodcont.2018.02.047
Fuentes DN, Calderon PF, Acuna LG, Rodas PI, Paredes-Sabja D, Fuentes JA, Gil F, Calderon IL (2015) Motility modulation by the small non-coding RNA SroC in Salmonella Typhimurium. FEMS Microbiol Lett 362(17):fnv135. https://doi.org/10.1093/femsle/fnv135
Gal-Mor O (2019) Persistent infection and long-term carriage of typhoidal and nontyphoidal Salmonellae. Clin Microbiol Rev 32:e00088–e00018. https://doi.org/10.1128/CMR.00088-18
Hautefort I, Thompson A, Eriksson-Ygberg S, Parker ML, Lucchini S, Danino V, Bongaerts RJM, Ahmad N, Rhen M, Hinton JCD (2008) During infection of epithelial cells Salmonella enterica serovar Typhimurium undergoes a time-dependent transcriptional adaptation that results in simultaneous expression of three type 3 secretion systems. Cell Microbiol 10(4):958–984. https://doi.org/10.1111/j.1462-5822.2007.01099.x
Horstmann JA, Zschieschang E, Truschel T, de Diego J, Lunelli M, Rohde M, May T, Strowig T, Stradal T, Kolbe M, Erhardt M (2017) Flagellin phase-dependent swimming on epithelial cell surfaces contributes to productive Salmonella gut colonisation. Cell Microbiol 19:e12739. https://doi.org/10.1111/cmi.12739
Jiang L, Feng L, Yang B, Zhang W, Wang P, Jiang X, Wang L (2017) Signal transduction pathway mediated by the novel regulator LoiA for low oxygen tension induced Salmonella Typhimurium invasion. PLoS Pathog 13(6):e1006429–e1006429. https://doi.org/10.1371/journal.ppat.1006429
Kaur J, Jain SK (2012) Role of antigens and virulence factors of Salmonella enterica serovar Typhi in its pathogenesis. Microbiol Res 167(4):199–210. https://doi.org/10.1016/j.micres.2011.08.001
Lamas A, Regal P, Vazquez B, Miranda JM, Cepeda A, Franco CM (2018) Influence of milk, chicken residues and oxygen levels on biofilm formation on stainless steel, gene expression and small RNAs in Salmonella enterica. Food Control 90:1–9. https://doi.org/10.1016/j.foodcont.2018.02.023
LaRock DL, Chaudhary A, Miller SI (2015) Salmonellae interactions with host processes. Nat Rev Microbiol 13(4):191–205. https://doi.org/10.1038/nrmicro3420
Li GH, Yan CH, Xu YF, Feng YQ, Wu Q, Lv XY, Yang BW, Wang X, Xia XD (2014) Punicalagin inhibits Salmonella virulence factors and has anti-quorum-sensing potential. Appl Environ Microbiol 80(19):6204–6211. https://doi.org/10.1128/AEM.01458-14
Li BQ, Yue YY, Yuan ZL, Zhang FY, Li P, Song NN, Lin W, Liu Y, Yang YL, Li ZH, Gu LC (2017) Salmonella STM1697 coordinates flagella biogenesis and virulence by restricting flagellar master protein FlhD(4)C(2) from recruiting RNA polymerase. Nucleic Acids Res 45(17):9976–9989. https://doi.org/10.1093/nar/gkx656
Ma ZP, Song Y, Cai ZH, Lin ZJ, Lin GH, Wang Y, Zhou J (2018) Anti-quorum sensing activities of selected coral symbiotic bacterial extracts from the South China Sea. Front Cell Infect Microbiol 8:144. https://doi.org/10.3389/fcimb.2018.00144
Mathur R, Oh H, Zhang DK, Park SG, Seo J, Koblansky A, Hayden MS, Ghosh S (2012) A mouse model of Salmonella Typhi infection. Cell 151(3):590–602. https://doi.org/10.1016/j.cell.2012.08.042
Merino L, Procura F, Trejo FM, Bueno DJ, Golowczyc MA (2019) Biofilm formation by Salmonella sp. in the poultry industry: detection, control and eradication strategies. Food Res Int 119:530–540. https://doi.org/10.1016/j.foodres.2017.11.024
Ng WL, Bassler BL (2009) Bacterial quorum-sensing network architectures. Annu Rev Genet 43:197–222. https://doi.org/10.1146/annurev-genet-102108-134304
Pacello F, Ceci P, Ammendola S, Pasquali P, Chiancone E, Battistoni A (2008) Periplasmic Cu, Zn superoxide dismutase and cytoplasmic Dps concur in protecting Salmonella enterica serovar Typhimurium from extracellular reactive oxygen species. Biochim Biophys Acta 1780(2):226–232. https://doi.org/10.1016/j.bbagen.2007.12.001
Pati NB, Vishwakarma V, Jaiswal S, Periaswamy B, Hardt WD, Suar M (2013) Deletion of invH gene in Salmonella enterica serovar Typhimurium limits the secretion of Sip effector proteins. Microbes Infect 15(1):66–73. https://doi.org/10.1016/j.micinf.2012.10.014
Roche SM, Holbert S, Trotereau J, Schaeffer S, Georgeault S, Virlogeux-Payant I, Velge P (2018) Salmonella Typhimurium invalidated for the three currently known invasion factors keeps its ability to invade several cell models. Front Cell Infect Microbiol 8:273. https://doi.org/10.3389/fcimb.2018.00273
Ros-Chumillas M, Garre A, Mate J, Palop A, Periago PM (2017) Nanoemulsified D-limonene reduces the heat resistance of Salmonella Senftenberg over 50 times. Nanomaterials (Basel) 7(3). https://doi.org/10.3390/nano7030065
Ryan D, Mukherjee M, Suar M (2017) The expanding targetome of small RNAs in Salmonella Typhimurium. Biochimie 137:69–77. https://doi.org/10.1016/j.biochi.2017.03.005
Ryan D, Mukherjee M, Nayak R, Dutta R, Suar M (2018) Biological and regulatory roles of acid-induced small RNA RyeC in Salmonella Typhimurium. Biochimie 150:48–56. https://doi.org/10.1016/j.biochi.2018.05.001
Salaheen S, Jaiswal E, Joo J, Peng M, Ho R, Oconnor D, Adlerz K, Aranda-Espinoza JH, Biswas D (2016) Bioactive extracts from berry byproducts on the pathogenicity of Salmonella Typhimurium. Int J Food Microbiol 237:128–135. https://doi.org/10.1016/j.ijfoodmicro.2016.08.027
Shi X, Zhu X (2009) Biofilm formation and food safety in food industries. Trends Food Sci Technol 20(9):407–413. https://doi.org/10.1016/j.tifs.2009.01.054
Shi C, Yan CH, Sui Y, Sun Y, Guo D, Chen YF, Jin T, Peng XL, Ma LL, Xia XD (2017) Thymoquinone inhibits virulence related traits of Cronobacter sakazakii ATCC 29544 and has anti-biofilm formation potential. Front Microbiol 8:2220. https://doi.org/10.3389/fmicb.2017.02220
Silva LN, Zimmer KR, Macedo AJ, Trentin DS (2016) Plant natural products targeting bacterial virulence factors. Chem Rev 116(16):9162–9236. https://doi.org/10.1021/acs.chemrev.6b00184
Srey S, Jahid IK, Ha SD (2013) Biofilm formation in food industries: a food safety concern. Food Control 31(2):572–585. https://doi.org/10.1016/j.foodcont.2012.12.001
Steenackers H, Hermans K, Vanderleyden J, De Keersmaecker SCJ (2012) Salmonella biofilms: an overview on occurrence, structure, regulation and eradication. Food Res Int 45(2):502–531. https://doi.org/10.1016/j.foodres.2011.01.038
Stepanović S, Cirković I, Ranin L, Svabić-Vlahović M (2004) Biofilm formation by Salmonella spp. and Listeria monocytogenes on plastic surface. Lett Appl Microbiol 38(5):428–432. https://doi.org/10.1111/j.1472-765X.2004.01513.x
Thoendel M, Kavanaugh JS, Flack CE, Horswill AR (2011) Peptide signaling in the Staphylococci. Chem Rev 111(1):117–151. https://doi.org/10.1021/cr100370n
Vinothkannan R, Tamizh MM, Raj CD, Princy SA (2018) Fructose furoic acid ester: an effective quorum sensing inhibitor against uropathogenic Escherichia coli. Bioorg Chem 79:310–318. https://doi.org/10.1016/j.bioorg.2018.05.009
Wu SC, Fu BD, Chu XL, Su JQ, Fu YX, Cui ZQ, Xu DX, Wu ZM (2016) Subinhibitory concentrations of phloretin repress the virulence of Salmonella Typhimurium and protect against Salmonella Typhimurium infection. Antonie Van Leeuwenhoek 109(11):1503–1512. https://doi.org/10.1007/s10482-016-0752-z
Wu SC, Chu XL, Su JQ, Cui ZQ, Zhang LY, Yu ZJ, Wu ZM, Cai ML, Li HX, Zhang ZJ (2018) Baicalin protects mice against Salmonella Typhimurium infection via the modulation of both bacterial virulence and host response. Phytomedicine 48:21–31. https://doi.org/10.1016/j.phymed.2018.04.063
Yang HL, Korivi M, Lin MW, Chen SC, Chou CW, Hseu YC (2015) Anti-angiogenic properties of coenzyme Q0 through downregulation of MMP-9/NF-κB and upregulation of HO-1 signaling in TNF-α-activated human endothelial cells. Biochem Pharmacol 98(1):144–156. https://doi.org/10.1016/j.bcp.2015.09.003
Yang HL, Lin MW, Korivi M, Wu JJ, Liao CH, Chang CT, Liao JW, Hseu YC (2016) Coenzyme Q0 regulates NFκB/AP-1 activation and enhances Nrf2 stabilization in attenuation of LPS-induced inflammation and redox imbalance: evidence from in vitro and in vivo studies. Biochim Biophys Acta 1859(2):246–261. https://doi.org/10.1016/j.bbagrm.2015.11.001
Yang HL, Thiyagarajan V, Shen PC, Mathew DC, Lin KY, Liao JW, Hseu YC (2019) Anti-EMT properties of CoQ0 attributed to PI3K/AKT/NFKB/MMP-9 signaling pathway through ROS-mediated apoptosis. J Exp Clin Cancer Res 38(1):186. https://doi.org/10.1186/s13046-019-1196-x
Zeng H, Carlson AQ, Guo YW, Yu YM, Collier-Hyams LS, Madara JL, Gewirtz AT, Neish AS (2003) Flagellin is the major proinflammatory determinant of enteropathogenic Salmonella. J Immunol 171(7):3668–3674. https://doi.org/10.4049/jimmunol.171.7.3668
Zhao XC, Liu ZH, Li WL, Li X, Shi C, Meng RZ, Cheng W, Jin KQ, Yang ZQ, Shi XC, Guo N, Yu L (2014) In Vitro synergy of nisin and coenzyme Q0 against Staphylococcus aureus. Food Control 46:368–373. https://doi.org/10.1016/j.foodcont.2014.05.051
Zhao YY, Gorvel JP, Meresse S (2016) Effector proteins support the asymmetric apportioning of Salmonella during cytokinesis. Virulence 7(6):669–678. https://doi.org/10.1080/21505594.2016.1173298
Zhu MJ, Olsen SA, Sheng L, Xue Y, Yue W (2015) Antimicrobial efficacy of grape seed extract against Escherichia coli O157:H7 growth, motility and Shiga toxin production. Food Control 51:177–182. https://doi.org/10.1016/j.foodcont.2014.11.024
Acknowledgments
We thank all the partners and laboratory members for their kind help.
Funding
This work was financially supported by the Natural Science Foundation of China (31801659), the Fundamental Research Funds for the Central Universities (2452017228), and General Financial Grant from the China Postdoctoral Science Foundation (No. 2017M623256).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethical statements
This paper is our original work. It has not been submitted elsewhere, and it is not under consideration in any other Journal. This article does not contain any studies with human participants or animals performed by any of the authors. All the authors have seen the manuscript and approved its submission to Applied Microbiology and Biotechnology.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yang, Y., Li, J., Yin, Y. et al. Antibiofilm activity of coenzyme Q0 against Salmonella Typhimurium and its effect on adhesion–invasion and survival–replication. Appl Microbiol Biotechnol 103, 8545–8557 (2019). https://doi.org/10.1007/s00253-019-10095-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-10095-8