Abstract
Salmonella spp. can cause animal and human salmonellosis. In this study, we established a simple method to detect all Salmonella species by amplifying a specific region within the flgE gene encoding the flagellar hook protein. Our preliminary sequence analysis among flagella-associated genes of Salmonella revealed that although Salmonella Gallinarum and Salmonella Pullorum are lacking flagella, they did have flagella-associated genes, including flgE. To investigate in detail, a comparative flgE sequence analysis was conducted using different bacterial strains including flagellated and non-flagellated Salmonella as well as non-Salmonella strains. Two unique regions (481–529 bp and 721–775 bp of the reference sequence) within the flgE open reading frame were found to be highly conserved and specific to all Salmonella species. Next, we designed a pair of PCR primers (flgE-UP and flgE-LO) targeting the above two regions, and performed a flgE-tailored PCR using as template DNA prepared from a total of 76 bacterial strains (31 flagellated Salmonella strains, 26 non-flagellated Salmonella strains, and 19 other non-Salmonella bacteria strains). Results showed that specific positive bands with expected size were obtained from all Salmonella (including flagellated and non-flagellated Salmonella) strains, while no specific product was generated from non-Salmonella bacterial strains. PCR products from the positive bands were confirmed by DNA sequencing. The minimum detection amount for genomic DNA and bacteria cells reached 18.3 pg/μL and 100 colony-forming unit (CFU) per PCR reaction, respectively. Using the flgE-PCR method to detect Salmonella in artificially contaminated milk samples, as low as 1 CFU/mL Salmonella was detectable after an 8-h pre-culture. Meanwhile, the flgE-tailored PCR method was applied to evaluate 247 clinical samples infected with Salmonella from different chicken breeding farms. The detection results indicated that flgE-PCR could be used to specifically detect Salmonella in concordance with the traditional bacterial culture-based detection method. It is worthwhile noticed that identification results using flgE-tailored PCR should be completed within less than 1 day, expanding the result of much faster than the standard method, which took more than 5 days. Overall, the flgE-tailored PCR method can specifically detect flagellated and non-flagellated Salmonella and can serve as a powerful tool for rapid, simple, and sensitive detection of Salmonella species.
Key points
• Targeting flgE gene for all Salmonella spp. found.
• The established PCR assay is used to specifically detect all Salmonella spp.
• The PCR method is applied to detect clinical Salmonella spp. samples within less than 1 day.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a member of Enterobacteriaceae, Salmonella is a group of Gram-negative bacterial pathogens and more than 2 600 different serovars have been reported until now (Issenhuth-Jeanjean et al. 2014). Some Salmonella species can cause gastroenteritis or systemic typhoid fever in several animal hosts including human. These pathogenic Salmonella are potential risk factors for public health safety (Foley and Lynne 2008). Accurate identification of this pathogen is one of the key processes to control salmonellosis. Therefore, it is imperative to develop a rapid, accurate, and convenient method for Salmonella detection in order to improve the security of human and animal health.
There have been numerous reports on Salmonella detection methods. Conventional methods based on bacterial culture, isolation, biochemical tests, and serotyping have been regarded as a “gold standard” (Cho and Ku 2017). However, the traditional Salmonella detection process is time-consuming and labor-intensive. According to White-Kauffmann-Le Minor (WKLM) scheme (Issenhuth-Jeanjean et al. 2014) and based on determination somatic (O) and flagellar (H) antigens, Salmonella can be serotyped by slide agglutination test (SAT). Although the SAT is simple, rapid, and the most important diagnostic tool under on-the-spot inspection conditions, it has the drawbacks of low detection sensitivity, easy to produce false positive results, and not economical because of the expensive commercial specific antisera. These shortcomings limit its usefulness as an ideal detection method.
Currently, the polymerase chain reaction (PCR) has been proved as a sensitive, rapid, and specific method, which is widely used for the detection of Salmonella (Cho and Ku 2017). Many PCR detection methods employed various genes as a target for the detection of Salmonella. Some of them use housekeeping genes such as 16S rDNA and gyrB as detection targets (Lin et al. 2004; Ye et al. 2011), while other methods are using virulence-associated genes as a target, such as invA (Rahn et al. 1992), hilA (Pathmanathan et al. 2003), stn (Makino et al. 1999), and ompC (Kwang et al. 1996).
In addition, some genes coding for bacterial surface antigens, such as flagellum and fimbriae, have been introduced as genetic targets for the identification of Salmonella (Doran et al. 1993; Hirose et al. 2002; Munir et al. 2015; Zhang et al. 2014). Among them, flagellum-associated genes have been proved to be potential targets suitable for Salmonella identification, for example, flagellin protein coding genes (fliC) have been reported as a target for the detection of Salmonella (Chiu et al. 2005; Khan et al. 2012). These methods can identify one or several particular serotype serovars of Salmonella. For instance, Chiu et al. (2005) developed a PCR method based on the fliC gene for detection of Salmonella enterica serovar Choleraesuis; Khan et al. (2012) reported a nested PCR targeting the flagellin gene for the detection of Salmonella Typhi. However, to our knowledge, there are no reports about using only single pair of primers targeting the flagellar gene, to identify all Salmonella species, including flagellated and non-flagellated Salmonella strains. We believe that this is due to the non-flagellated phenotype of Salmonella Pullorum and Salmonella Gallinarum, which limits the idea of using flagellum-associated genes to identify non-flagellum bacteria. Since the infections of Salmonella Pullorum and Salmonella Gallinarum, both of which are non-flagellated, remain an important problem for the poultry industry in many countries and regions (Barrow and Freitas Neto 2011), it is necessary to conduct improvement for the existing detection methods.
Salmonella Pullorum and Salmonella Gallinarum serovars are known as non-flagellated bacteria (Barrow and Freitas Neto 2011). However, when we performed bioinformatic and subsequent experimental sequencing analysis, we found that there are flagella coding genes in the genome of the two serotypes of Salmonella, including flgE gene, which encodes the flagellate hook protein hook. Besides, we found that flgE gene sequences from Salmonella Pullorum and Salmonella Gallinarum showed a very high sequence identity (100% sequence identity) when comparing with the flgE gene sequence from flagellated Salmonella Enteritidis. These unexpected discoveries were encouraging to investigate the possibility of using the flagellum-associated gene flgE as target to detect flagellated and non-flagellated Salmonella species.
In this work, based on the bioinformatics and sequencing analysis, we established a rapid PCR method to identify both flagellated and non-flagellated Salmonella simultaneously. The upstream and downstream primers were designed to specifically target unique sequences regions of Salmonella flgE gene. The specificity and sensitivity of the PCR method were determined in this study. Result showed that the flgE-PCR method was specific and sensitive enough to detect Salmonella spp. In addition, by detecting 247 samples of chicken embryos from breeding farms, the flgE-PCR method was proved to be capable for Salmonella detection in clinical samples. In summary, the PCR method was suitable for an accurate, rapid, and convenient detection of all Salmonella spp.
Materials and methods
Bacterial strains
A total of 76 bacteria strains including 31 flagellated Salmonella strains, 26 non-flagellated Salmonella strains, and 19 other non-Salmonella bacteria strains (Table 1) were used to characterize the specificity of flgE-PCR method. The Salmonella Enteritidis strain CMCC (B) 50336 (abbreviated as SE50336 in subsequent text) was kindly offered by Professor Xinan Jiao in Yangzhou University. Salmonella Enteritidis strains T48 and T64; Salmonella Typhimurium strains W32, T14, U27, W2, A12; Salmonella Typhi W33; Salmonella Choleraesuis U80, U81, U82; and Salmonella Gallinarum U20 were kindly offered by Professor Shulin Liu, Harbin Medical University. Salmonella Pullorum strains CVCC523, CVCC526, CVCC535, and CVCC540 were from China Veterinary Culture Collection Center (CVCC). Escherichia coli strains CE7 was provided by Professor Chengping Lu in Nanjing Agricultural University. The Staphylococcus strain was a gift from the Institute of Animal Husbandry and Veterinary Medicine, Beijing Academy of Agriculture and Forestry Sciences. The other strains were preserved in our laboratory. The SE50336 strain was used as a reference strain to perform the optimization of PCR conditions and sensitivity of the PCR assay. All the strains were grown in Luria-Bertani (LB) broth (NaCl [Sinopharm Chemical Reagent Co., Ltd, Beijing, China] 10 g/L, Tryptone [Oxoid, Hampshire, UK] 10.0 g/L, Yeast extract [Oxoid, Hampshire, UK] 5.0 g/L) or on LB agar plates at 37 °C.
Genomic DNA extraction
Genomic DNA was extracted from cultured bacterial cells by boiling method as previously described (Zhang et al. 2014). Briefly, 1.5 mL of overnight bacteria culture was centrifuged at 10000 rpm for 5 min, and the pellet was washed with sterilized double-distilled water (DDW) twice. Then, 200 μL of DDW was added and the mixture was suspended by pipetting then boiled at 100 °C for 10 min. After centrifuging at 10000 rpm for 5 min, the supernatants were transferred into a clean Eppendorf tube. The concentration of genomic DNA was measured using NanoDrop 2000 (Thermo Scientific, Wilmington, DE, USA). These DNA were stored at − 20 °C and then served as templates for PCR amplification.
Bioinformatics analysis
Multiple sequence alignments were performed using flgE gene sequences from different flagellated and non-flagellated Salmonella strains and non-Salmonella strains. The flgE sequences were downloaded from the GenBank database. The flgE nucleotide sequence from Salmonella Enteritidis strain P125109 (AM933172.1), a serovar harbor flagella-associated genes and has the nearest genetic relationships with Salmonella Pullorum and Salmonella Gallinarum, was selected as a reference sequence. The presence of the flgE gene in non-flagellated Salmonella strains, including Salmonella Gallinarum strains (T63, T88, T89, T90, U20, SG9R, and SG01) and Salmonella Pullorum strains (CVCC523, CVCC526, CVCC535, CVCC540, S07, S08, S10, S11, S78, S12-1, S12-2, 45SP13, SP68, SP73, SP79, SP80, SP82, SP90, and SP95) (Table 1), was confirmed by PCR amplification. Primers for PCR-amplifying the complete sequence of flgE were described in Table 2. Then, the PCR products were purified and cloned into pMD19-T vector (Takara Biotechnology Co., Dalian, China) and sequenced by GENEWIZ (Suzhou, China). The sequenced data were analyzed and aligned by ClustalW method using the MEGA 7 software (Version 7.0.26) and Jalview (Version 2.11.0) (Waterhouse et al. 2009).
PCR procedure
The PCR assay was performed in a 25 μL reaction volume containing: 2.5 μL of 10 × PCR reaction buffer (Mg2+ free) (Takara Biotechnology Co., Dalian, China), 2 μL of dNTPs (2.5 mM each), 1 μL of upstream and downstream primer (10 μM), 0 mM to 2.0 mM of final concentrations of MgCl2, 1.5 U of rTaq DNA polymerase (Takara Biotechnology Co., Dalian, China), and 1 μL(183 ng/μL) of extracted genomic DNA template. Positive and negative controls were the DNA of reference strain SE50336 and DDW, respectively. Amplifications were carried out using a DNA thermocycler (Applied Biosystems, Foster City, CA, USA) and the following amplification program: initial denaturation at 94 °C for 4 min, followed by 30 cycles of denaturation at 94 °C for 1 min, annealing at gradient temperature (from 50 °C to 60 °C) for 45 s, and extension at 72 °C for 45 s, with a final extension at 72 °C for 10 min.
Amplification products were analyzed by electrophoreses on 2% agarose gel with a DL2000 ladder (Takara Biotechnology Co., Dalian, China) as molecular weight marker. The gels were imaged by a gel image system (BIO-RAD, Hercules, CA, USA).
Specificity analysis of the flgE-PCR
The specificity of the flgE-PCR was assessed using genomic DNA from 57 Salmonella strains (including 31 flagellated Salmonella strains and 26 non-flagellated Salmonella strains) and 19 non-Salmonella bacterial strains (Table 1).
Sensitivity of the flgE-PCR
To determine the minimum amount of DNA that can be detected by flgE-PCR method, genomic DNA extracted from Salmonella Enteritidis strain SE50336 was 10-fold serially diluted with DDW from 183 ng/μL to 183 fg/μL and then served as template for the PCR reaction. To determine the minimum amount of bacteria cells that can be detected by the flgE-PCR method, 30 μL of overnight culture for Salmonella Enteritidis strain SE50336 was subcultured into 3 mL of LB at 37 °C with shaking at 200 revolutions per minute (RPM) for about 1.5 h. The culture was first adjusted to an optical density of 1.0 at 600 nm (OD600nm = 1.0), and then prepared for serial 10-fold dilutions in PBS. An aliquot (1 μL) of each dilution was directly used as templates to perform the PCR test. In parallel, an aliquot (100 μL) of each dilution was plated onto a LB agar plate, then incubated overnight at 37 °C to determine the colony-forming unit (CFU) of SE50336.
Milk sample was obtained from the local market and confirmed negative for Salmonella. Then, 0.5 ml of milk was mixed with 4.5 ml of selenite cystine (SC) broth (Qingdao Hope Bio-Technology Co., Ltd, Qingdao, China). To these mixtures, 0 to 1 × 103 CFU/mL of Salmonella SE50336 cells was inoculated, and the mixtures were incubated at 37 °C for 6–10 h. DNA was then extracted from 1 mL of each culture and used as template for PCR assays following DNA amplification conditions described above.
Salmonella detection of the samples from the chicken embryos in the breeding farms
A total of 247 samples were collected from healthy and dead chicken embryos in breeding farms located in Guangxi province during 2019. Samples were pre-enriched in Buffered Peptone Water (BPW) for 18 hours at 37 °C, then selectively enriched in SC broth for 18–24 h at 37 °C. After pre-enrichment, the presence of Salmonella in samples was determined using both the described flgE-PCR method and a modified method from standard microbiological analysis procedure of China (GB 4789. 4-2010). Positive samples were confirmed on Xylose Lysine Deoxycholate (XLD) agar and MacConkey (MAC) agar (Qingdao Hope Bio-Technology Co., Ltd, Qingdao, China), and serotyped by SAT using a commercial kit of Salmonella-specific O and H antisera (Tianjin biochip corporation, Tianjin, China) based on WKLM scheme.
Results
Bioinformatics analysis of the flgE gene of the flagellated Salmonella
We began our study by nucleotide sequences alignment of flgE gene from bacterial strains belonging to Salmonella (24 serovars of flagellated Salmonella) and non-Salmonella (13 strains) genera (Supplementary Fig. S1). Results showed that the flgE gene is distributed in all flagellated Salmonella strains (100 %) used in this study. In order to find a unique region for identification of Salmonella, the difference of the flgE gene between Salmonella and non-Salmonella was analyzed. The result showed that, two regions of the flgE gene, corresponding to nucleotides 481–529 and 721–775 of the flgE open reading frame, were found to have the least identity with non-Salmonella sequences (Supplementary Fig. S1). Since these two regions of the flgE gene were both specific and unique for Salmonella strains, they were considered suitable regions to design a specific pair of primer for the identification of Salmonella strains.
Bioinformatics analysis of the flgE gene of non-flagellated Salmonella (i.e., Salmonella Gallinarum/Pullorum)
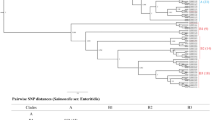
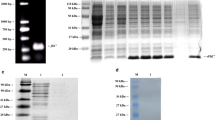
Although Salmonella Gallinarum and Salmonella Pullorum did not contain flagella, complete genome sequence analysis of these two serovars showed that the genomes of both Salmonella species harbor flagella-associated genes, including the flgE gene (Table 3). We downloaded the flgE gene sequences from the complete genome data (listed in Table 3) and compared them with the reference sequences from a confirmed flagellated Salmonella Enteritidis strain P125109 (AM933172.1) (Supplementary Fig. S2). Furthermore, in order to evaluate whether the flgE gene was present in all the non-flagellated Salmonella, PCR assays were carried out using the genome of Salmonella Gallinarum and Pullorum strains as template which were collected strains in our laboratory (Table 1), and flgE-N and flgE-C primer pairs were used (Table 2). Then, the sequences of flgE gene from the above strains were sequenced and compared with the reference sequence (AM933172.1). Results suggested that the flgE gene is present in all the tested 7 strains of Salmonella Gallinarum and 19 strains of Salmonella Pullorum (Fig. 1). In addition, the flgE sequence from Salmonella Pullorum/Gallinarum showed a very high sequence identity (100%) with the flgE sequence from reference strain P125109 (Supplementary Fig. S2). These results suggested that the flagellated Salmonella-specific regions in the flgE gene also exist in non-flagellated strains and were conserved among all the analyzed Salmonella strains.
The presence and identification of the flagella hook gene flgE in non-flagellated Salmonella (Salmonella Pullorum and Salmonella Gallinarum) strains. Lane M, 2 K plus DNA ladder; lanes 1–26, genomic DNA samples from different Salmonella Pullorum and Salmonella Gallinarum strains which were listed in Table 1. Lane 27, positive control (PC) use genomic DNA from a known positive for flgE gene strain (Salmonella Enteritidis str. SE50336) as template, Lane 28, negative control (NC), the genomic DNA was replaced with equal-volume of DDW. The flgE gene presents in all strains of non-flagellated Salmonella tested
Primer designs for detecting Salmonella spp.
The bioinformatic analysis showed that the flagellar hook gene flgE was distributed in all Salmonella species including flagellated and non-flagellated Salmonella strains. In addition, results suggested that the above two regions of the flgE gene (481–529 bp and 721–775 bp of the reference sequence) were conserved, and present in all Salmonella strains. These evidences supported the potential idea of using these two unique regions of the flgE gene to design primers for Salmonella spp. detection. By taking advantage of this feature, we designed upstream primer (flgE-UP) within the region one and downstream primer (flgE-LO) within the region two (Fig. 2, Table 2). This pair of primer was then utilized to exploit a PCR method to rapidly and efficiently detect flagellated and non-flagellated Salmonella.
PCR procedure
The PCR amplification system was developed and the reaction conditions were optimized, including annealing temperature and Mg2+ concentration. The results showed that under the conditions of annealing temperature 50–61 °C (Fig. 3a), and concentration of 0.4–2.0 mM Mg2+ (Fig. 3b), target fragments can be amplified very well, which means that the established reaction system allows for PCR at a wide range of annealing temperature and Mg2+ concentrations. The primers yield to the expected 262-bp amplicon (Fig. 3). Based on the conditions described in the “Materials and methods” section, the optimal conditions for the two other parameters were determined as follows: an annealing temperature of 56 °C and a final concentration of 2.0 mM MgCl2. The determined parameters were used in subsequent experiments.
The effects of the annealing temperature and MgCl2 concentration on flgE-PCR reaction. Lane M, DL2000 DNA ladder. a The effects of the annealing temperature on flgE-PCR reaction. Lane 1–12, the PCR reaction at the different annealing temperature (50 to 61 °C). b The effects of the MgCl2 concentrations on flgE-PCR reaction. Lanes 1–11, PCR amplification system with different final concentrations of MgCl2 (0, 0.2, 0.4, 0.6, 0.8, 1.0, 1.2, 1.4, 1.6, 1.8, 2.0 mM)
The flgE-PCR method proved to be both unique and specific to all the Salmonella spp.
The specificity of the flgE-PCR system was determined by testing 31 flagellated Salmonella strains, 26 non-flagellated Salmonella strains, and 19 non-Salmonella bacterial strains (Table 1). The results revealed that all the flagellated Salmonella strains generated the 262-bp of specific target band. In addition, all the non-flagellated Salmonella (Salmonella Pullorum/Gallinarum) strains also generated the specific target band (Fig. 4, Table 1). In contrast, no amplification products were observed in 19 strains of different non-Salmonella pathogens (Fig. 4, Table 1). This result shows that the flgE-PCR method has the potential to specifically detect flagellated Salmonella (lanes 1–31) and the non-flagellated Salmonella (lanes 32–57), without any detectable cross-reactivity with non-Salmonella bacteria strains.
Specificity of the flgE-PCR for detecting all Salmonella spp. The PCR assays were conducted using genomic DNA as template from different flagellated, non-flagellated Salmonella strains, and non-Salmonella bacterial strains. Lane M, DL2000 DNA ladder; lane 1–31, PCR products amplified from flagellated Salmonella strains. Lane 32–57, PCR products amplified from non-flagellated Salmonella strains. Lane 58–76, PCR products amplified from non-Salmonella strains. Positive control (PC), using genomic DNA from a known positive strain (SE50336) as template. Detailed strain information were listed in Table 1
The sensitivity of the flgE gene-PCR method for detecting all Salmonella spp.
To evaluate the sensitivity of the PCR method, genomic DNA of Salmonella Enteritidis strain SE50336 was consecutively diluted and used as templates for PCR reaction. The target DNA fragment was amplified using template DNA concentrations ranging from of 183 ng/μL to 183 fg/μL. The results showed that the minimum amount of DNA that could be detected by this method was 18.3 pg/μL of genomic DNA (Fig. 5a). Furthermore, the minimum of bacterial cells of Salmonella Enteritidis that could be detected using this flgE-PCR method was determined using different dilutions of SE50336 cells. After analyzed by the flgE-PCR method, the least bacterial cells that could be detected were validated as 100 CFU per reaction (Fig. 5b). Milk samples artificially contaminated with different cell numbers of Salmonella SE50336 were analyzed through flgE-PCR after 6, 8, and 10-h incubation. Our data showed that it was possible to detect an inoculation concentration of 1 CFU/mL of Salmonella after 8 h of enrichment with the SC mixture (Fig. 6).
Sensitivity of the flgE-PCR assay for detection of bacterial genomic DNA and bacterial cells from Salmonella Enteritidis (str. SE50336). The PCR amplified a product of 262 bp. Lane M, DL2000 DNA ladder. a The PCR detection for the bacterial genomic DNA, lanes 1–7, 183 ng/μL, 18.3 ng/μL, 1.83 ng/μL, 183 pg/μL, 18.3 pg/μL, 1.83 pg/μL, 183 fg/μL of bacterial genomic DNA from SE50336 were used as template. b The PCR detection for the Salmonella Enteritidis cells, lanes 1–6, the number of bacteria cells per PCR reaction was 106 CFU, 105 CFU, 104 CFU, 103 CFU, 102 CFU, and 10 CFU
Sensitivity of the flgE-PCR for detection of Salmonella Enteritidis CMCC (B) 50336 in milk samples with pre-enrichment for 6 h (A), 8 h (B), or 10 h (C). Lane M, DL2000 DNA ladder, lane 1–4, milk samples spiked with bacterial culture serially diluted 10-fold from 103 (lane 1) to 100 (lane 4) CFU/mL, lane 5, unspiked (0 CFU/mL) milk sample as PCR template, lane 6, positive control (PC) use genomic DNA from CMCC (B) 50336 as template, lane 7, negative control (NC), equal-volume of DDW as PCR template
The flgE-PCR method is adequate for the Salmonella detection in clinical samples
A total number of 247 chicken embryos samples were tested for the presence of Salmonella using both the flgE-PCR method and the traditional standard method. The results were listed in Table 4. After pre-enrichment and selective enrichment (less than 1 day), 46 of the 247 samples were tested positive for the presence of Salmonella using flgE-PCR method (less than 3 h). An identical number of Salmonella positive samples (46 out of 247) was obtained when performing the traditional microbiological identification process (about 5 days, Fig. 7). Taken together our designed PCR method, which is highly sensitive and in complete agreement with the traditional identification method, could have potential application as molecular microbiology tool for rapid Salmonella detection in aviculture.
Discussion
Salmonella induces infections in humans and a broad range of animals, which remains a serious health problem in human and veterinary medicine (Foley and Lynne 2008; Kirk et al. 2015; Rukambile et al. 2019; Threlfall 2002). Traditional culture-based detection procedure is still the most commonly used routine method for the identification of Salmonella. This diagnosis method for Salmonella detection is based on the use of selective culture media such as SC, XLD, MAC, Tetrathionate broth (TTB), Bismuth Sulfite (BS), Deoxycholate Hydrogen Sulfide Lactose (DHL), and serological tests according to the White-Kauffmann-Le Minor scheme and biochemical features analysis. The culture-based detection is time-consuming and laborious because it always takes up to about 5–7 days to complete the detection process (Cho and Ku 2017). Nucleic acid-based analytical methods have undergone considerable improvements in sensitivity, specificity, and speed during the past decades. Among these methods, PCR serves as a powerful tool to detect Salmonella in a simple, rapid, and convenient manner (Ricke et al. 2018).
Bacterial flagellum is a motile organelle, which provide bacteria swimming or swarming motilities and has multiple biological functions on bacterial pathogenicity, such as cell adhesion, biofilm formation, and host cell invasion (Chaban et al. 2015; Duan et al. 2013; Zhou et al. 2015). A typical bacterial flagellum consists of three parts: a basal body, a filament, and a hook (Chaban et al. 2015). The basal body is located in the cytoplasmic membrane and function as a motor. The flagellar filament is a long tubular structure and functions as a helical propeller that enables bacterial motility. Flagellin, which is encoded by the fliC gene, is a subunit protein of flagellar filament. The bacterial flagellin is a well-studied and typical pathogen-associated molecular pattern (PAMP), which can be recognized by pattern recognition receptors (PRRs), such as Toll-like receptor (TLR) and NOD-like receptors (NLRs), and subsequently trigger the innate immunity (Cui et al. 2018; Hayashi et al. 2001; Hajam et al. 2017). In addition, as a strong immunogen, flagellin can also activate the adaptive immune response (Honko and Mizel 2005). Therefore, it has been widely recognized that flagellin can serve as an immune adjuvant to enhance antigenic immunogenicity of the vaccines (Gupta et al. 2014; Kim et al. 2018).
The flagellar hook is a short, highly curved tubular structure and serves as a universal joint that connects the basal body to the flagellar filament. The gene flgE codes a 403 amino acid (AA) hook protein FlgE, which forms the hook of the Salmonella flagellum. Previous studies are mainly focused on the structural feature of hook subunit protein FlgE and hook assembling mechanisms (Matsunami et al. 2016; Moriya et al. 2011; Saijo-Hamano et al. 2019; Samatey et al. 2004). However, recent research has revealed that the flagellar hook and hook monomer protein FlgE of Pseudomonas aeruginosa have the potency of pro-inflammation or immune-stimulation (Shen et al. 2017; Li et al. 2019). This means that the flagellar hook and its monomer protein FlgE are novel key factors, which contribute to host-bacterial interactions. Thus, just like flagellin, the flagellar hook and protein FlgE have the potential to serve as a vaccine adjuvant (Shen et al. 2017; Li et al. 2019).
In this study, we developed a PCR method for detecting all Salmonella spp. based on targeting the flagellar hook protein coding gene flgE. The primers were designed within two unique regions (481–529 bp and 721–775 bp of the reference sequence) in flgE gene, which were conserved and unique to Salmonella species. Interestingly, although the two non-flagellated Salmonella serovars (Salmonella Gallinarum and Salmonella Pullorum) did not contain flagella, we noticed that their genome harbors flagella-associated genes, including flgE gene. Subsequently, we confirmed that flgE gene was present in all the Salmonella Gallinarum/Pullorum strains tested in this study. In addition, the flgE sequences from Salmonella Pullorum/Gallinarum are 100% identical to the flgE sequence from the reference strain. This means that the new clue described in this study is competent for the detection of non-flagellated Salmonella. Subsequently, to validate the proof of concept of our designed method, we tested various Salmonella Gallinarum and Salmonella Pullorum strains available in our laboratory using this flgE-PCR method. In addition, the flgE-PCR method was also carried out in clinical chicken embryo sample detection. The result of PCR has a high accordance (100%) with traditional identification methods. Besides, the flgE-PCR method showed low time consumption compared with the standard method.
Several molecular targets have been applied for Salmonella detection (Table 5). Salmonella have been detected using conventional PCR, quantitative real-time PCR (qPCR), multiplex real-time PCR (Multiplex qPCR), etc. (Malorny et al. 2009). The sensitivity of the different methods is varied. Generally, the qPCR assay was more sensitive than the conventional PCR assay. However, the higher cost of the equipment and reagent of the qPCR method somewhat limited its wide application, despite high sensitivity of identification. Conventional PCR shows advantages in simple operation and cost-effectiveness; therefore, it is more suitable for low cost routine analysis. Our flgE-PCR method was capable to detect as few as 100 CFU/PCR of the pure culture of Salmonella SE50336 directly. This result showed that our method is significantly more sensitive than the previous PCR assays established by Kwang et al. (1996) and Pathmanathan et al. (2003), in which a minimum of 400 CFU and 120 CFU was respectively required for detecting Salmonella ompC and hilA genes. However, using our flgE-PCR method, the minimum amount for bacterial cell required for Salmonella detection was 10- and 100-fold behind the methods reported by Lin et al. (2004) and Ye et al. (2011), respectively. In their reports, 1–9 CFU (by targeting 16S rDNA gene) and 3.2 CFU (by targeting gyrB gene) of bacteria cells could be detected. A pre-culture step of a few hours is always necessary before performing PCR to fulfil the detection requirements of national legislations (GB 4789. 4-2010). The minimum detection amount of bacterial cells in our flgE-PCR method was improved to 1 CFU/mL in contaminated milk after 8 h of pre-culture. The results are consistent with the early research that the PCR demonstrated increased sensitivity when pre-enrichment was performed prior to the PCR detection (Nam et al. 2005). In short, the flgE-PCR method can be used for routine detection of Salmonella with a reasonable sensitivity.
Overall, through bioinformatics analysis of the flgE gene and subsequent experimental verification, a practical PCR method for identification of Salmonella was provided in this study. Furthermore, the detailed sequence analysis of the flgE gene in this work will provide the basis and the necessary information for further study of the function of the Salmonella FlgE protein. Future research can focus on, but not limited to, the following aspects: designing experiments to identify the major domains of Salmonella FlgE protein that are responsible for stimulating the inflammatory response of host cells, and formulating strategies to evaluate the potential of the FlgE protein to serve as an immune adjuvant, and so on.
In conclusion, we found two unique regions within the flagellar hook protein encoding gene flgE, which are both conserved and unique to the Salmonella genus. By taking advantage of this characteristic, one pair of PCR primers was designed within these two regions. Based on this pair of primers, a direct PCR assay specific for all Salmonella spp. was developed. The specificity and sensitivity of the flgE-PCR method were proved by detecting different Salmonella and non-Salmonella strains preserved in our laboratory. Furthermore, flgE-PCR method was also proved to be competent for the Salmonella detection in clinical samples collecting from chicken farms. Compared with the traditional methods, this flgE-PCR method was rapid, accurate, and easy to operate since the entire Salmonella detection process can be rapidly completed without intensive labor force. Due to the potential of our flgE-PCR method, it is worth further investigations to optimize and apply this PCR assay in the clinical detection and surveillance of Salmonella in the near future.
References
Afroj S, Aldahami K, Reddy G, Guard J, Adesiyun A, Samuel T, Abdela W (2017) Simultaneous detection of multiple Salmonella serovars from milk and chicken meat by real-time PCR using unique genomic target regions. J Food Prot:1944–1957. https://doi.org/10.4315/0362-028X.JFP-17-133
Barrow PA, Freitas Neto OC (2011) Pullorum disease and fowl typhoid--new thoughts on old diseases: a review. Avian Pathol 40(1):1–13. https://doi.org/10.1080/03079457.2010.542575
Calvo L, Martinez-Planells A, Pardos-Bosch J, Garcia-Gil LJ (2008) A new real-time PCR assay for the specific detection of Salmonella spp. targeting the bipA gene. Food Anal Methods 1(4):236–242. https://doi.org/10.1007/s12161-007-9008-x
Chaban B, Hughes HV, Beeby M (2015) The flagellum in bacterial pathogens: for motility and a whole lot more. Semin Cell Dev Biol 46:91–103. https://doi.org/10.1016/j.semcdb.2015.10.032
Chen S, Yee A, Griffiths M, Larkin C, Yamashiro CT, Behari R, Paszko-Kolva C, Rahn K, De Grandis SA (1997) The evaluation of a fluorogenic polymerase chain reaction assay for the detection of Salmonella species in food commodities. Int J Food Microbiol 35(3):239–250. https://doi.org/10.1016/s0168-1605(97)01241-5
Chen W, Martinez G, Mulchandani A (2000) Molecular beacons: a real-time polymerase chain reaction assay for detecting Salmonella. Anal Biochem 280(1):166–172. https://doi.org/10.1006/abio.2000.4518
Chen J, Zhang L, Paoli GC, Shi C, Tu SI, Shi X (2010) A real-time PCR method for the detection of Salmonella enterica from food using a target sequence identified by comparative genomic analysis. Int J Food Microbiol 137(2-3):168–174. https://doi.org/10.1016/j.ijfoodmicro.2009.12.004
Chiu TH, Pang JC, Hwang WZ, Tsen HY (2005) Development of PCR primers for the detection of Salmonella enterica serovar Choleraesuis based on the fliC gene. J Food Prot 68(8):1575–1580. https://doi.org/10.4315/0362-028x-68.8.1575
Cho IH, Ku S (2017) Current technical approaches for the early detection of foodborne pathogens: challenges and opportunities. Int J Mol Sci 18(10). https://doi.org/10.3390/ijms18102078
Cui BF, Liu XS, Fang YZ, Zhou P, Zhang YG, Wang YL (2018) Flagellin as a vaccine adjuvant. Expert Rev Vaccines 17(4):335–349. https://doi.org/10.1080/14760584.2018.1457443
de Almeida MV, Silva A Jr, Nero LA (2014) Evaluation of target sequences for the polymerase chain reaction-based detection of Salmonella in artificially contaminated beef. Foodborne Pathog Dis 11(2):111–118. https://doi.org/10.1089/fpd.2013.1623
Ding T, Suo Y, Zhang Z, Liu D, Ye X, Chen S, Zhao Y (2017) A multiplex RT-PCR assay for S. aureus, L. monocytogenes, and Salmonella spp. detection in raw milk with pre-enrichment. Front Microbiol 8:989. https://doi.org/10.3389/fmicb.2017.00989
Doran JL, Collinson SK, Burian J, Sarlos G, Todd EC, Munro CK, Kay CM, Banser PA, Peterkin PI, Kay WW (1993) DNA-based diagnostic tests for Salmonella species targeting agfA, the structural gene for thin, aggregative fimbriae. J Clin Microbiol 31(9):2263–2273
Duan Q, Zhou M, Zhu L, Zhu G (2013) Flagella and bacterial pathogenicity. J Basic Microbiol 53(1):1–8. https://doi.org/10.1002/jobm.201100335
Foley SL, Lynne AM (2008) Food animal-associated Salmonella challenges: pathogenicity and antimicrobial resistance. J Anim Sci 86(14 Suppl):E173–E187. https://doi.org/10.2527/jas.2007-0447
Gupta SK, Bajwa P, Deb R, Chellappa MM, Dey S (2014) Flagellin a Toll-like receptor 5 agonist as an adjuvant in chicken vaccines. Clin Vaccine Immunol 21(3):261–270. https://doi.org/10.1128/Cvi.00669-13
Hadjinicolaou AV, Demetriou VL, Emmanuel MA, Kakoyiannis CK, Kostrikis LG (2009) Molecular beacon-based real-time PCR detection of primary isolates of Salmonella Typhimurium and Salmonella Enteritidis in environmental and clinical samples. BMC Microbiol 9:97. https://doi.org/10.1186/1471-2180-9-97
Hajam IA, Dar PA, Shahnawaz I, Jaume JC, Lee JH (2017) Bacterial flagellin-a potent immunomodulatory agent. Exp Mol Med 49(9):e373. https://doi.org/10.1038/emm.2017.172
Hayashi F, Smith KD, Ozinsky A, Hawn TR, Yi EC, Goodlett DR, Eng JK, Akira S, Underhill DM, Aderem A (2001) The innate immune response to bacterial flagellin is mediated by Toll-like receptor 5. Nature 410(6832):1099–1103. https://doi.org/10.1038/35074106
Hirose K, Itoh K, Nakajima H, Kurazono T, Yamaguchi M, Moriya K, Ezaki T, Kawamura Y, Tamura K, Watanabe H (2002) Selective amplification of tyv (rfbE), prt (rfbS), viaB, and fliC genes by multiplex PCR for identification of Salmonella enterica serovars Typhi and Paratyphi A. J Clin Microbiol 40(2):633–636. https://doi.org/10.1128/jcm.40.02.633-636.2002
Honko AN, Mizel SB (2005) Effects of flagellin on innate and adaptive immunity. Immunol Res 33(1):83–101. https://doi.org/10.1385/IR:33:1:083
Issenhuth-Jeanjean S, Roggentin P, Mikoleit M, Guibourdenche M, de Pinna E, Nair S, Fields PI, Weill FX (2014) Supplement 2008-2010 (no. 48) to the White-Kauffmann-Le Minor scheme. Res Microbiol 165(7):526–530. https://doi.org/10.1016/j.resmic.2014.07.004
Khan S, Harish BN, Menezes GA, Acharya NS, Parija SC (2012) Early diagnosis of typhoid fever by nested PCR for flagellin gene of Salmonella enterica serotype Typhi. Indian J Med Res 136(5):850–854
Kim MI, Lee C, Park J, Jeon BY, Hong M (2018) Structure-guided fusion-protein designs using Bacillus flagellin as a vaccine adjuvant. Acta Crystallogr A 74:A134–A134. https://doi.org/10.1107/S0108767318098653
Kirk MD, Pires SM, Black RE, Caipo M, Crump JA, Devleesschauwer B, Dopfer D, Fazil A, Fischer-Walker CL, Hald T, Hall AJ, Keddy KH, Lake RJ, Lanata CF, Torgerson PR, Havelaar AH, Angulo FJ (2015) World health organization estimates of the global and regional disease burden of 22 foodborne bacterial, protozoal, and viral diseases, 2010: a data synthesis. PLoS Med 12(12):e1001921. https://doi.org/10.1371/journal.pmed.1001921
Kwang J, Littledike ET, Keen JE (1996) Use of the polymerase chain reaction for Salmonella detection. Lett Appl Microbiol 22(1):46–51. https://doi.org/10.1111/j.1472-765x.1996.tb01106.x
Li Y, Shen Y, Lin D, Zhang H, Wang T, Liu H, Wang Y (2019) Neutrophils and IL17A mediate flagellar hook protein FlgE-induced mouse acute lung inflammation. Cell Microbiol 21(3):e12975. https://doi.org/10.1111/cmi.12975
Lin CK, Hung CL, Hsu SC, Tsai CC, Tsen HY (2004) An improved PCR primer pair based on 16S rDNA for the specific detection of Salmonella serovars in food samples. J Food Prot 67(7):1335–1343. https://doi.org/10.4315/0362-028x-67.7.1335
Makino S, Kurazono H, Chongsanguam M, Hayashi H, Cheun H, Suzuki S, Shirahata T (1999) Establishment of the PCR system specific to Salmonella spp. and its application for the inspection of food and fecal samples. J Vet Med Sci 61(11):1245–1247. https://doi.org/10.1292/jvms.61.1245
Malorny B, Paccassoni E, Fach P, Bunge C, Martin A, Helmuth R (2004) Diagnostic real-time PCR for detection of Salmonella in food. Appl Environ Microbiol 70(12):7046–7052. https://doi.org/10.1128/AEM.70.12.7046-7052.2004
Malorny B, Huehn S, Dieckmann R, Krmer N, Helmuth R (2009) Polymerase chain reaction for the rapid detection and serovar identification of Salmonella in food and feeding stuff. Food Anal Methods 2(2):81–95. https://doi.org/10.1007/s12161-008-9057-9
Matsunami H, Barker CS, Yoon YH, Wolf M, Samatey FA (2016) Complete structure of the bacterial flagellar hook reveals extensive set of stabilizing interactions. Nat Commun 7:13425. https://doi.org/10.1038/ncomms13425
Moriya N, Minamino T, Imada K, Namba K (2011) Genetic analysis of the bacterial hook-capping protein FlgD responsible for hook assembly. Microbiol-SGM 157:1354–1362. https://doi.org/10.1099/mic.0.047100-0
Munir T, Lodhi M, Ali S, Hussain Zaidi SB, Razak S (2015) Early diagnosis of typhoid by PCR for fliC-d gene of Salmonella Typhi in patients taking antibiotics. J Coll Physicians Surg Pak 25(9):662–666
Nam HM, Srinivasan V, Gillespie BE, Murinda SE, Oliver SP (2005) Application of SYBR green real-time PCR assay for specific detection of Salmonella spp. in dairy farm environmental samples. Int J Food Microbiol 102(2):161–171. https://doi.org/10.1016/j.ijfoodmicro.2004.12.020
Pathmanathan SG, Cardona-Castro N, Sanchez-Jimenez MM, Correa-Ochoa MM, Puthucheary SD, Thong KL (2003) Simple and rapid detection of Salmonella strains by direct PCR amplification of the hilA gene. J Med Microbiol 52(Pt 9):773–776. https://doi.org/10.1099/jmm.0.05188-0
Rahn K, De Grandis SA, Clarke RC, McEwen SA, Galan JE, Ginocchio C, Curtiss R 3rd, Gyles CL (1992) Amplification of an invA gene sequence of Salmonella Typhimurium by polymerase chain reaction as a specific method of detection of Salmonella. Mol Cell Probes 6(4):271–279. https://doi.org/10.1016/0890-8508(92)90002-f
Ricke SC, Kim SA, Shi Z, Park SH (2018) Molecular-based identification and detection of Salmonella in food production systems: current perspectives. J Appl Microbiol 125(2):313–327. https://doi.org/10.1111/jam.13888
Rukambile E, Sintchenko V, Muscatello G, Kock R, Alders R (2019) Infection, colonization and shedding of Campylobacter and Salmonella in animals and their contribution to human disease: a review. Zoonoses Public Health 66(6):562–578. https://doi.org/10.1111/zph.12611
Saijo-Hamano Y, Matsunami H, Namba K, Imada K (2019) Architecture of the bacterial flagellar distal rod and hook of Salmonella. Biomole 9(7). https://doi.org/10.3390/biom9070260
Samatey FA, Matsunami H, Imada K, Nagashima S, Shaikh TR, Thomas DR, Chen JZ, Derosier DJ, Kitao A, Namba K (2004) Structure of the bacterial flagellar hook and implication for the molecular universal joint mechanism. Nature 431(7012):1062–1068. https://doi.org/10.1038/nature02997
Shen Y, Chen L, Wang M, Lin D, Liang Z, Song P, Yuan Q, Tang H, Li W, Duan K, Liu B, Zhao G, Wang Y (2017) Flagellar hooks and hook protein FlgE participate in host microbe interactions at immunological level. Sci Rep 7(1):1433. https://doi.org/10.1038/s41598-017-01619-1
Threlfall EJ (2002) Antimicrobial drug resistance in Salmonella: problems and perspectives in food- and water-borne infections. FEMS Microbiol Rev 26(2):141–148. https://doi.org/10.1111/j.1574-6976.2002.tb00606.x
Waterhouse AM, Procter JB, Martin DM, Clamp M, Barton GJ (2009) Jalview Version 2--a multiple sequence alignment editor and analysis workbench. Bioinformatics 25(9):1189–1191. https://doi.org/10.1093/bioinformatics/btp033
Ye XH, Wang YM, Lin XG (2011) A gyrB-targeted PCR for rapid identification of Salmonella. Curr Microbiol 63(5):477–483. https://doi.org/10.1007/s00284-011-0007-1
Zhang JY, Dong LW, Ren Q, Wang XZ, Yang Y, Zhou W, Zhu CH, Meng X, Zhu GQ (2014) Simple and rapid detection of Salmonella by direct PCR amplification of gene fimW. Curr Microbiol 69(4):429–435. https://doi.org/10.1007/s00284-014-0602-z
Zhou MX, Yang Y, Chen PL, Hu HJ, Hardwidge PR, Zhu GQ (2015) More than a locomotive organelle: flagella in Escherichia coli. Appl Microbiol Biotechnol 99(21):8883–8890. https://doi.org/10.1007/s00253-015-6946-x
Zhou B, Liang T, Zhan Z, Liu R, Li F, Xu H (2017) Rapid and simultaneous quantification of viable Escherichia coli O157:H7 and Salmonella spp. in milk through multiplex real-time PCR. J Dairy Sci 100(11):8804–8813. https://doi.org/10.3168/jds.2017-13362
Acknowledgments
The authors gratefully acknowledge Professor Xinan Jiao in Yangzhou University, Professor Chengping Lu in Nanjing Agricultural University, Professor Shulin Liu from Harbin Medical University and the Institute of Animal Husbandry and Veterinary Medicine, Beijing Academy of Agriculture and Forestry Sciences for providing the bacteria strains for this study.
Funding
This work was supported by the grants from the no. 2016YFD0500905 and 2017YFD0500105 from the National Key Research and Development Program of China, and a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
YY and PW performed the experiments, analyzed the data, and wrote the manuscript. PX, BY, PD, TH, and JL participated in the data analysis and wrote the paper. QS contributed to the experiments designing, manuscript writing, and language polishing. GZ and XM conceived and designed the study, participated in experimental work, and wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Ethics approval and consent to participate
This article does not contain any studies with human participants performed by any of the authors. The current study was approved by the by the Institutional Animal Care and Use Committee of the Yangzhou University College of Veterinary Medicine of China.
Consent for publication
All authors listed on this manuscript have read and agreed to the publication of this research.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 3078 kb)
Rights and permissions
About this article
Cite this article
Yang, Y., Wang, P., Xia, P. et al. Rapid detection of flagellated and non-flagellated Salmonella by targeting the common flagellar hook gene flgE. Appl Microbiol Biotechnol 104, 9719–9732 (2020). https://doi.org/10.1007/s00253-020-10925-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10925-0