Abstract
The aim of this review is to give a summary of natural lignocellulose-degrading systems focusing mainly on animal digestive tracts of wood-feeding insects and ruminants in order to find effective strategies that can be applied to improve anaerobic digestion processes in engineered systems. Wood-feeding animals co-evolved with symbiotic microorganisms to digest lignocellulose-rich biomass in a very successful way. Considering the similarities between these animal gut systems and the lignocellulose-based biotechnological processes, the gut with its microbial consortium can be a perfect model for an advanced lignocellulose-degrading biorefinery. The physicochemical properties and structure of the gut may provide a scheme for the process design, and the microbial consortium may be applied as genetic resource for the up-scaled bioreactor communities. Manipulation of the gut microbiota is also discussed in relation to the management of the reactor communities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In order to mitigate global climate issues caused by increasing anthropogenic greenhouse gas (GHG) emissions, fossil energy carrier consumptions must be drastically reduced and organic municipal, agricultural, and agro-industrial wastes should be properly treated (Aguirre-Villegas and Larson 2017; Aneja et al. 2009; Bogner et al. 2008). Utilization of plant biomass for both material and energetic use can be considered an important strategy to contribute to these aims, but further improvements of the current state of technology are necessary (Sawatdeenarunat et al. 2015). The major structural component of plant cells is lignocellulose, a complex matrix composed of cellulose (D-glucose homo-polysaccharide), hemicellulose (hetero-polysaccharide containing both C6 and C5 sugars, such as arabinose, galactose, glucose, mannose, and xylose, as well as their uronic acids), lignin (hetero-polymer of mainly aromatic compounds, such as p-coumaryl alcohol, coniferyl alcohol, and sinapyl alcohol), pectin (hetero-polysaccharide rich in galacturonic acid) and various minor components (proteins, terpenic oils, fatty acids, fatty acid esters, and inorganic components). For more details about the composition of lignocellulose, see the following references (Rubin 2008; Scharf and Tartar 2008). Annually, approximately 200 billion tons of lignocellulosic biomass is produced (da Silva et al. 2012) and a considerable part of it is treated as waste, such as agricultural wastes or green cuts from parks and gardens. Despite the energy potential conserved in the polysaccharide structure, the cost-effective utilization of lignocellulose is hampered by its recalcitrant nature. The plant cell-wall material has evolved complex structural and chemical mechanisms to withstand external digestion by microbes and animals, which makes the hydrolysis of these main biopolymers the rate-limiting step of biochemical conversion processes. Nature offers already successful and established systems that decompose dead plant material relatively fast, as a result of complex interactions of numerous bacteria, fungi, protists, and a variety of wood-feeding animals (Cragg et al. 2015). Compared to conventional anaerobic digesters, lignocellulosic biomass breakdown in rumen of cow is estimated to be three times more effective (Bayane and Guiot 2011). In the digestion system of wood-feeding termites, the rate of biomass biodegradation is even higher (Okwakol 1980). Therefore, more detailed knowledge on the digestion mechanisms of specific herbivores and the contribution of co-evolved microorganisms in their gut might be applied in the design of novel biorefinery concepts. Biomimicry or biomimetics is an emerging strategy studying the principles in biological systems that have been evolved over geological times to apply this knowledge to create novel technologies for the purpose of solving complex challenges (Dicks 2017; Vincent et al. 2006). The aim of this review is to give an overview of effective natural lignocellulose-degrading systems with high biotechnological application potentials in biorefineries focusing mainly on animal digestive tracts of wood-feeding insects and ruminants with a major focus on their symbiotic microbial partners. The potential utilization of such microbiota in engineered systems for biomass conversion will also be discussed. Besides biological treatment and enhancement options, the structure and physiology of the digestive tract of effective herbivorous animals may provide a better design for future lignocellulose-utilizing biorefinery systems. Due to the important role of the microbiota in nutrition and health of the host, potential manipulation of the microbial community has attracted lots of research in case of livestock animals. Management of bioreactor microbiota is similar in many aspects; therefore, the review will give also an overview on these topics.
Structural and functional differences between animal gut systems and biogas reactors
During evolution, animals developed highly sophisticated digestive systems to get maximum benefit from various food sources to gain energy, to sustain their lives, and to be successful in reproduction. These “ecosystem engineers” have completed the optimization period of the digestion process for all kinds of feedstock (Godon et al. 2013). Compared to the digestion systems in animals, human-engineered anaerobic digestion (AD) is a relatively new technology, which still deals with challenges especially regarding limited hydrolysis rates of lignocellulosic biomass.
In nature, there are many invertebrate (e.g., termites or some beetle larvae) and vertebrate animals (e.g., ruminants) that evolved digestive tracts having the ability to effectively degrade lignocellulosic substrates thanks to the specific structure and the symbiotic interactions with a wide variety of microorganisms colonizing the digestion tract (Auer et al. 2017; Bayane and Guiot 2011; Ozbayram et al. 2018d). The gut systems of these animals are considered as efficient bioreactors (Brune 2007). Furthermore, it is assumed that the smaller body size of the animals enables higher concentrations of cellulolytic enzymes contributing to faster degradation of these substrates.
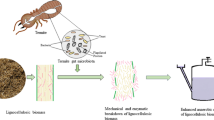
Biomass digestion in insects highly depends on the symbiotic relationships with microorganisms, whereas the microorganisms provide various compounds, such as digestive enzymes and nutrients (amino acids, vitamins, etc.) (Andert et al. 2010; Berasategui et al. 2016). Members of the family Scarabaeidae (order Coleoptera) are abundant in grassland environments, and larvae of this family are mostly herbivorous and digest plant material very effectively (Huang et al. 2010). They have unique digestive tracts with three compartments: the first part is a foregut, the second part is a long midgut with alkaline conditions, and the third part is an expanded hindgut, also called paunch or fermentation chamber (Fig. 1a) (Engel and Moran 2013; Huang et al. 2010).
Simplified schemes of (a) the gut of the sun beetle larva, (b) the gut of a ruminant represented by a cow, (c) a current biogas reactor digesting plant biomass, and (d) a future biorefinery based on the gut system of herbivorous animals. CHP, combined heat and power system; SCFAs, short-chain fatty acids; MCFAs, medium-chain fatty acids
A considerable amount of energy is spent on chewing and grinding the food particles to smaller pieces to improve their digestibility. As part of the foregut, strong mandibles as well as a proventriculus region (also called gizzard) with teeth-like cuticle structures and a strongly developed muscle layer around it have been evolved for the more effective mechanical treatment/grinding of the plant biomass. The so-called crop is a flexible part of the foregut between the esophagus and the proventriculus, with the function of temporary storage. Another typical characteristic of the scarab larvae is a long midgut with high pH conditions and an expanded hindgut with lower pH environment. Such pH gradients play a role in the digestive process for effective decomposition and absorption of nutrients. The high pH in the midgut is a key driver for enhancing the solubility of various polymers, such as hemicellulose and lignin (Huang et al. 2010). Alkaline conditions of the midgut enable dissolution of lignin and de-esterification of intermolecular ester bonds, increase the surface area and porosity, and decrease crystallinity of the biomass (Kim et al. 2016). Rows of caeca circle the midgut tract with hypothesized functions related to digestion, nutrient and fluid reabsorption, and ion homeostasis. The entry of Malpighian tubes, which are involved in excretion and osmoregulation, marks the transition from midgut to hindgut. A highly muscular pyloric sphincter separates these two sections and enables the food transfer from midgut to hindgut (Huang et al. 2010). The dilated hindgut has lobe-like structures and is considered the main region for digestion of (hemi)celluloses. The pH of the lumen content is closer to neutral, but the redox potential is more negative compared to the midgut (Cazemier et al. 2003; Lemke et al. 2003) providing appropriate conditions even for methanogenesis. However, only the central part of the paunch is completely anoxic due to the oxygen diffusion through the epithelial tissue.
Termites and their evolutionary sister group, the cockroach family Cryptocercidae, have similar digestive systems and can digest wood effectively in their hindgut by cooperating with microbial symbionts (Bauer et al. 2015; Watanabe and Tokuda 2010). Generally, the digestive systems are composed of the mouth, esophagus, salivary glands, foregut, midgut, and hindgut. The mandibles are used to grind solid particles, and, according to the size of the animal, the mandible’s size gets smaller from cockroaches to termites. Whereas lower termites and cockroaches have a large anterior part in the form of a crop in their foreguts, higher termites have a reduced crop. The volume of the midgut in termites is smaller compared to that of cockroaches. Moreover, cockroaches have bigger gastric ceca in the anterior midgut than lower termites. In cockroaches, the hindgut is divided into two compartments, namely ileum and rectum. However, termites have more specialized hindguts composed of ileum, enteric valve, paunch, colon, and rectum (Watanabe and Tokuda 2010). Termites are very efficient in cellulolytic biomass degradation with estimated degradation rates of 74–99% for cellulose and 65–87% for hemicellulose carried out mainly by microbial symbionts (Watanabe and Tokuda 2010).
The ruminants are considered the most important foregut fermenters having a special four-chamber stomach (reticulum, rumen, omasum, and abomasum) (Fig. 1b). The feed fermentation and the absorption of the resulting volatile fatty acids (VFAs) take place in the first three chambers, including rumen, reticulum, and omasum, collectively named forestomach. The fourth part is called a true stomach (abomasum) regarding its acidic conditions. The rumen microbiota is responsible to convert ingested biomass to VFAs, providing 70–85% of the nutrients absorbed by ruminants (Noel et al. 2017). The produced VFAs are continuously absorbed by the rumen epithelial cells, thus maintaining stable conditions for microbial activities (Bayane and Guiot 2011). The most distinct feature of the ruminants is a mechanism called rumination, which means that the animals regurgitate the large particles of the semi-digested substrate and chew again for long periods to enhance the surface of the feed for enzymatic attacks (Welch 1982).
AD is a good example of a biotechnological application in the field of renewable energies as well as in waste management (De Vrieze and Verstraete 2016). The conversion of biomass into methane-rich biogas is a complex biochemical process occurring in four steps, namely hydrolysis, acidogenesis, acetogenesis, and methanogenesis. It is completely carried out by synergistic interactions among the members of a diverse microbial community (Wei 2016). The process occurs in a controlled engineered system, in which operational parameters, such as temperature, mixing, hydraulic retention time, and solid retention time, are maintained in the reactor (Fig. 1c).
There are two major types of ideal continuous reactors, the plug flow reactor (PFR) and the continuous stirred tank reactor (CSTR). The PFR has usually a cylindrical geometry where the reactor content progresses from the inlet in the axial direction to the outlet as a series of infinitely thin coherent “plugs.” A key assumption is that the reactor content is perfectly mixed in the radial but not in the axial direction and each plug is considered a separate small batch reactor. CSTRs consist of a well-stirred tank, which is fed (semi)continuously and at the same time the digestate stream is removed. If all four AD steps take place in a single reactor, the system is called as one-stage digestion. In contrast, two-stage digestion systems consist of two sequential reactors to separate hydrolysis and acidogenesis in the first stage from acetogenesis and methanogenesis in the second stage (Nkemka et al. 2015). The main functional difference between the animal gut and the current AD systems is that the process in the gut is optimized toward production of VFAs that are utilized by the animal, while methane is just a side product of hydrogenotrophic or methylotrophic methanogenesis (Breznak 1982; Brune and Dietrich 2015; Mason and Stuckey 2016). In current engineered AD systems, a high degree of biomass degradation is only achieved because methanogenesis is a major sink of VFAs and acetoclastic methanogenesis is involved in addition to hydrogenotrophic methanogenesis. Besides hydrogenotrophic methanogenesis, homoacetogenesis, in which hydrogen is used to reduce carbon dioxide to acetate, is another hydrogen sink in the gut of wood-feeding termites (Tholen and Brune 1999; Tholen and Brune 2000) and ruminants (Gagen et al. 2015; Henderson et al. 2010). Although homoacetogenic bacteria are also found in biogas reactors treating biomass (Demirel and Scherer 2008), due to thermodynamic reasons, they do not play a major role in reductive acetogenesis but rather in the reverse process of syntrophic acetate oxidation, which is coupled to hydrogenotrophic methanogenesis (Schnürer et al. 1999; Westerholm et al. 2019). Nevertheless, Siriwongrungson and co-workers found that under altered conditions these bacteria can indeed perform homoacetogenesis (Siriwongrungson et al. 2007). In biomethanation systems utilizing extra hydrogen generated from excess electricity of other renewables, this process is supposed to play an important role (Omar et al. 2018; Wahid et al. 2019; Zabranska and Pokorna 2018).
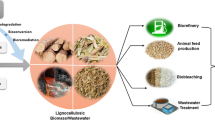
The high efficiency of lignocellulose degradation by specialized animals is also due to their unique characteristics of the digestive systems. Different mechanisms in the digestive systems, such as enzymatic attacks (e.g., by cellulases, xylanases, esterases, ligninases), mechanical grinding, and chemical conditions (e.g., alkaline or acidic conditions), contribute to the successful solubilization of lignocellulosic compounds. The fermentation products are removed continuously by different mechanisms to promote degradation, for instance VFAs by absorption on epithelial surface and hydrogen by methanogenesis or homoacetogenesis. The compartmentalization in biomass-feeding animals, such as ruminants, enables both homogenization and stratification of digestate during fermentation. Rumination is the main characteristic of these vertebrate animals, a process creating new surface areas for microorganisms to degrade polymers. Moreover, radial and/or axial oxygen gradients along the digestive system may contribute to delignification of these compounds. Some microorganisms are located on the epithelium or trapped in the mucus retaining them in the digestive tract, and adhesion on the lignocellulosic particles facilitates the contact with the feed (Bayane and Guiot 2011; Mason and Stuckey 2016). Since hydrolysis rate in these animals is faster than in a typical anaerobic digester, biomimicry is an important approach for successful innovations (Mason and Stuckey 2016). Thus, in order to enhance the digestion performance, some additional approaches, such as pre-treatment (chemical, physical, enzymatic), bioaugmentation, and co-digestion strategies, should be integrated in future sophisticated AD systems. Moreover, the biorefinery concept can be incorporated in such AD systems by including compartmentalization and utilization of carboxylic acids as additional products besides biogas (Fig. 1d).
Major differences in microbial communities from gut and current engineered AD systems
Recent trends in high-throughput amplicon sequencing and metagenome analysis of microbial communities and decreasing sequencing costs have led to proliferation of studies investigating lignocellulose-degrading communities in various natural and engineered environments.
The gut microorganisms with direct or indirect roles in lignocellulose degradation belong to Bacteria, Archaea, and Eukarya, such as protists and fungi. Figure 2 shows key families of the three domains of life found as abundant members of the microbiota of termite gut, beetle larvae gut, rumen, and AD systems. There is a strong link between the phylogenetic classification of the host and the microbial community of arthropods (Ley et al. 2008). The gut of scarab beetle larvae, Pachnoda spp. (Coleoptera: Scarabaeidae), harbors diverse bacterial communities involved in the degradation of plant materials. Previous studies pointed out that Bacteroidetes, Proteobacteria, and Firmicutes are the predominant phyla in the hindgut compartment, while Actinobacteria members are most abundant in the midgut (Andert et al. 2010; Egert et al. 2003). Due to the alkaline conditions in the midgut of beetle larvae, the bacterial richness is much lower than that of the hindgut, where diverse microbial processes and high concentrations of fermentation products occur (Andert et al. 2010). A new species, Promicromonospora pachnodae, excreting xylanases and endoglucanases, was also isolated from the hindgut of Pachnoda marginata larvae (Cazemier et al. 2003).
Potential key families found as abundant members of the microbiota of termite gut, beetle larvae gut, rumen, and AD systems. The data for the tree was derived from the Taxonomy database in NCBI using common tree option (https://www.ncbi.nlm.nih.gov/Taxonomy/CommonTree/wwwcmt.cgi). All the families mentioned in the review were added manually and saved as phylip tree format. Then, the tree was drawn using iTOL v4 software (Letunic and Bork 2019). The colors represent the three domains as follows: purple for Bacteria, pink for Archaea, and green for Eukarya
The digestive tract of beetle larvae, termites, and ruminants harbors also archaeal communities dominated by methanogens (Brune 2014; Cunha et al. 2011; Hook et al. 2010; Paul et al. 2017; Shi et al. 2015). There are two main pathways to produce methane. During acetoclastic methanogenesis, methane is produced by conversion of acetic acid to methane by Methanosarcinales comprising the families Methansarcinaceae and Methanotrichaceae (formerly Methanosaetaceae). The members of the genus Methanothrix are strictly acetoclastic methanogens and use only acetate as a substrate for methanogenesis (Oren 2014). The members of Methansarcinaceae are usually not predominant but frequently found in the gut of insects and ruminants, and the presence of the strict acetotroph Methanothrix has not yet been reported (Janssen and Kirs 2008).
In contrast, hydrogenotrophic methanogens, such as Methanococcales, Methanobacteriales, Methanomicrobiales, and Methanocellales, produce methane by reduction of carbon dioxide with hydrogen (Bayane and Guiot 2011; Buan 2018; Christy et al. 2014). Methane formation can also be carried out through the methylotrophic pathway, in which methylated compounds, such as methanol, methylamines or methylated thiols, are converted into methane by Methanomassiliicoccales, Methanobacteriales, or Methanosarcinales (Enzmann et al. 2018). Methylotrophic methanogens are classified into two groups based on the way the reducing equivalents are provided. In case of Methanosarcina, one methyl-CoM is oxidized to CO2 via the reverse hydrogenotrophic pathway to generate the reducing equivalents for three methyl-CoM to methane (Enzmann et al. 2018). Hydrogen-dependent methylotrophs, such as members of the Methanomassiliicoccales, cannot oxidize the methyl groups to CO2; therefore, they require hydrogen as electron donor for methanogenesis (Lang et al. 2015). In the beetle larvae, Crenarchaeota is the most abundant archaeal phylum in the midgut, while species belonging to the Euryarchaeota (mostly Methanobacteriaceae) are dominant in the hindgut (Egert et al. 2003).
Termites (order Blattodea) comprise diverse species and are divided into lower and higher termites (Inward et al. 2007). Lower termites contain symbiotic protists in their hindguts, such as Trichomonadida, Hypermastigida (class Parabasalea), and Oxymonadida (class Oxymonadea), which excrete cellulases for plant biomass degradation (Ohkuma 2003). These symbionts can be horizontally transferred between the individuals in a colony (Kitade 2004; Ohkuma 2003). Moreover, each colony of termites may have different microbial communities due to the diet and/or living environment (Minkley et al. 2006). Whereas the microbiota of lower termites comprise flagellated protist symbionts, the hindgut in higher termites (family Termitidae), which constitute the major part of all termite species, has different physicochemical conditions not suitable for these protozoan symbionts (He et al. 2013; Warnecke et al. 2007). Generally, the hindgut microbial community is dominated by bacteria. Besides, in some higher termites, amoebae were detected in the gut system playing a role in cellulose digestion (Brune and Ohkuma 2010). The bacterial hindgut community of higher termites is considered a great source for motile bacteria. Species belonging to the Spirochaetes and Fibrobacteres are dominant in this environment. Treponema has been described to be the most abundant genus in the hindgut of higher termites (Warnecke et al. 2007). In the lower termites, the hindgut community is dominated by species belonging to the Spirochaetes, Bacteroidetes, and Proteobacteria. It was found that termites contribute substantially to global methane emissions, which is estimated around 3–15 Tg CH4 yr−1 (Saunois et al. 2016). The methane is mostly produced by the hydrogenotrophic pathway, whereas acetoclastic methanogenesis was not yet detected in termites. The methane production rate in lower termites is dependent on the activity of hydrogen-producing gut flagellates, while hydrogen in higher termites is mostly produced by fermenting bacteria. Methanobrevibacter is the most abundant archaeal genus in the hindgut of lower termites. In higher termites, members of the Methanosarcinales (genus Methanimicrococcus) and the Methanomicrobiales in addition to Methanobacteriales (genus Methanobrevibacter) were also detected (Hongoh et al. 2003; Ohkuma 2003). Furthermore, Thermoplasmatales and Crenoarchaeota were also found in some termites (Ohkuma 2003). More information about the gut microbiota of the termites can be found in detailed reviews by Brune (2014) and Brune and Dietrich (2015).
As in the beetle larvae, termites, and wood roaches, a diverse microbial community in ruminants carries out the plant material degradation. In the rumen, protozoa comprise almost 50% of the total rumen microbial biomass and produce similar fermentation products as bacteria (Choudhury et al. 2015). A total of 30–40% of fiber degradation in the rumen is accomplished by ciliates (Bayane and Guiot 2011). The bacteria associated with protozoa can accomplish various functions, such as serving electron sinks through nitrogen fixation, acetogenesis, or methanogenesis, and can provide nutrients for the protozoan host organisms. This relationship can be ecto- or endosymbiotic having benefits both to the protozoan host and symbiotic prokaryotes (Levy and Jami 2018). Furthermore, nearly 20% of the rumen microbial biomass is composed of anaerobic fungi, mostly of the phylum Neocallimastigomycota, that play an active role in the degradation of lignified plant biomass (Choudhury et al. 2015). There are numerous studies that investigated the microbial community structure, dynamics, and functions of rumen microbiota (Ozbayram et al. 2018b; Pitta et al. 2014; Söllinger et al. 2018; Zened et al. 2013). The bacterial community of the rumen fluid is dominated by the phyla Bacteroidetes, Firmicutes, and Proteobacteria in different proportions depending on the animal (Ozbayram et al. 2018b; Pitta et al. 2014; Söllinger et al. 2018; Zened et al. 2013). Additionally, the phylum Fibrobacteres includes several important rumen bacteria, such as Fibrobacter succinogenes and Fibrobacter intestinalis, which contribute to the degradation of plant material in the rumen environment (Ozbayram et al. 2018b; Ransom-Jones et al. 2012). Moreover, some members of this phylum were recently found in termites (Rahman et al. 2016). As a significant feature, Prevotella (order Bacteroidales, phylum Bacteroidetes) is the most abundant genus in the rumen, playing a key role in the breakdown of proteins and carbohydrates and excreting cellulolytic enzymes like carboxymethylcellulase and xylanase (Nyonyo et al. 2014). Henderson et al. (2015) described Butyrivibrio and Ruminococcus as well as unclassified Lachnospiraceae, Ruminococcaceae, Bacteroidales, and Clostridiales belonging to the core rumen bacterial community. Methane production in the rumen is a secondary microbial activity while the major function of rumen fermentation is to produce VFAs (Bayane and Guiot 2011). However, ruminants are considered one of the major contributors to methane emissions. The emissions vary according to the ruminant species. Whereas 26–497 g methane per day is emitted from a dairy cattle, the daily values in beef cattle and Suffolk sheep were found as 161–396 and 22–25 g, respectively (Broucek 2014). The genera Methanobrevibacter, Methanobacterium, and Methanomicrobium are dominant methanogens in the rumen environment and thus have been defined as characteristic hydrogenotrophic rumen methanogens (Bayane and Guiot 2011). In a recent study, the order Methanomassiliicoccales, which comprises hydrogen-dependent methylotrophic methanogens, was found to be abundant in the rumen fluid (Jin et al. 2017; Ozbayram et al. 2018b; Söllinger et al. 2018).
According to the hologenome concept, multicellular organisms should be considered holobionts (host plus symbionts) with their hologenome (host genome plus metagenome of the symbionts) rather than individuals as a level of selection in evolution (Rosenberg and Zilber-Rosenberg 2011, 2016; Zilber-Rosenberg and Rosenberg 2008). The gut microbiota as part of the holobiont co-evolved with their hosts over millions of years, while engineered AD systems are typically ad hoc inoculated without much consideration of selecting the most effective microbiota (Godon et al. 2013). The co-evolution of gut symbionts with their host requires specific transfer mechanisms from parents to offspring, which will be discussed later in this review. The bacterial communities of biogas systems are mostly dominated by Firmicutes and Bacteroidetes members (Kröber et al. 2009; Lucas et al. 2015; Lv et al. 2019), and a diverse microbial community takes a part in each step of biogas production. Hydrolytic and fermenting bacteria, namely Clostridia, Micrococcus, Bacteroides, Butyrivibrio, Fusobacterium, Selenomonas, and Streptococcus excrete various enzymes, such as cellulases, cellobiases, xylanases, amylases, proteases, and lipases, for hydrolyzing the insoluble biomass polymers into smaller units. Moreover, anaerobic microorganisms can excrete extracellular enzyme complexes, so-called cellulosomes, to degrade plant cell walls and form various fermentation products, such as ethanol and organic acids (Doi and Kosugi 2004). Right after hydrolysis, sugars, long-chain fatty acids, and amino acids are converted into VFAs, such as acetic, propionic, butyric, and other short-chain carboxylic acids, alcohols, H2, and CO2 by fermentative bacteria, such as Streptococcus, Lactobacillus, and Bacillus, during acidogenesis, which is considered the fastest step during AD (Christy et al. 2014). VFAs longer than two carbon atoms and alcohols longer than one carbon atom are further converted to acetate, hydrogen, and carbon dioxide in the acetogenesis step (Schink 1997) involving, e.g., the genera Smithella, Pelotomaculum, Syntrophobacter, and Syntrophomonas (Mathai et al. 2015). This syntrophic oxidation process should be distinguished from homoacetogenesis, which is acetate formation by using hydrogen to reduce carbon dioxide to acetate. Acetate is a central metabolite of the AD process that is either converted directly to CH4 and CO2 by acetoclastic methanogens or is oxidized to H2 and CO2 or formate by syntrophic acetate-oxidizing bacteria (SAOB). However, the latter process is only possible if the hydrogen partial pressure is kept low by hydrogenotrophic methanogens (Westerholm et al. 2016).
The predominant methanogenic pathway for biogas production in anaerobic digesters is dependent on the feedstock and operating conditions (Karakashev et al. 2005; Nettmann et al. 2010). The concentrations of ammonia and VFAs have an effect on the composition of the methanogenic community (Karakashev et al. 2005). Ziganshin et al. (2016) highlighted that the genus Methanoculleus was positively correlated with the NH3 concentration, whereas the prevalence of Methanocorpusculum, Methanobacterium, and Methanosaeta was negatively correlated with the NH3 level in biogas reactors. In another study, the same group showed that the organic loading rate shapes the methanogenic community in anaerobic digesters treating distillers’ grains (Ziganshin et al. 2011). Methanosarcina species were abundant in the reactors operated at a high organic loading rate and supplemented with iron hydroxide. However, acetoclastic methanogens of the genus Methanothrix (formerly Methanosaeta) dominated the well-performing reactors operated at lower organic loading rates. In large-scale agricultural biogas reactors, Methanobacterium, Methanosaeta and Methanoculleus were found as the most abundant genera (Lucas et al. 2015). The methanogenic communities in agricultural biogas plants were dominated by the orders Methanomicrobiales, Methanosarcinales, and Methanobacteriales (Nettmann et al. 2008; Rastogi et al. 2008). Fungi (such as Ascomycetes and Aspergillus) and protozoa (such as Amoeboflagellates, Cyclidium, Naeglaria, Rhynchomonas, Vorticella, Trichomonas) were detected in anaerobic digesters in lower abundances compared to rumen and termites. However, their role in the substrate degradation remains unknown (Bayane and Guiot 2011; Güllert et al. 2016).
Transfer of gut microbiota between generations
The microorganisms in the gut of herbivorous animals either belong to the core microbiota as obligate endosymbionts or just transient members of the community as non-core, facultative endosymbionts acquired from the environment. According to the hologenome concept, the obligate endosymbionts co-evolved with their hosts (Rosenberg and Zilber-Rosenberg 2011, 2016, 2018). The host is mainly defined at the species level; however, intra-species variations might exist as discussed earlier. Therefore, effective transfer mechanisms of beneficial microbes from the parent to the offspring are important. The way the gut endosymbionts are transmitted from one generation to the next is often related to the importance of the service(s) that they provide for the host (Shapira 2016). It is especially challenging in case of insects with different lifestyles between the larva and the imago stage. In many cases, the effective transfer of these microorganisms requires the bacteria to persist in the environment for a longer period, which might allow their potential transfer and utilization also in engineered systems.
The vertical transfer of beneficial microorganisms is prevailing in case of social insects as was shown in a study assessing the gut microbiota of two social termites (Mastotermes darwiniensis, Heterotermes aureus), a social wood roach (Cryptocercus punctulatus) and a non-social cockroach (Periplaneta americana) (Sabree et al. 2012). The gut of social termites harbors a more conserved microbial community similar to that of other termites, whereas cockroaches harbor more variable gut communities that differ among individual hosts and contain more typically environmental microbes. Wood roaches are the sister group to termites and display an early stage of sociality, and their gut community structures are similar among individuals (Sabree et al. 2012). Proctodeal trophallaxis is considered the main transfer mechanism of the microbiota in case of termites (Hongoh et al. 2005; Kohler et al. 2012) and wood-feeding cockroaches of the genus Cryptocercus (Nalepa 2015). A recent study showed that rather a “mixed mode” of transmission combining vertical (colony-to-offspring) and horizontal (colony-to-colony) transfer has been the major driving force shaping the gut microbiota of termites (Bourguignon et al. 2018). Another example for strict vertical transmission of symbionts is found in many stinkbug species where the mother lays eggs covered by feces-derived capsules (egg-smearing) containing the endosymbionts. The juveniles ingest the capsules after hatching and inoculate themselves with the symbionts (Fukatsu and Hosokawa 2002; Hosokawa et al. 2006). Reed beetles also use egg-smearing for the vertical transfer of symbionts (Kolsch and Pedersen 2010). An alternative way to acquire gut endosymbionts is via horizontal transmission, in which symbionts are derived from the environment as found in crickets and solitary cockroaches (Engel and Moran 2013). Environmentally acquired symbionts represent higher genetic variation, which could provide more opportunities for adaptation, but also allows the settlement of non-beneficial “cheater” microorganisms (Shapira 2016).
In conclusion, the major mechanisms of endosymbiont transfer of insects include direct strict vertical transmission through producing specific symbiont-containing capsules for the eggs via defecation (egg-smearing), forms of coprophagy or proctodeal trophallaxis, defecating and feeding in the same habitat, and acquiring microorganisms from the environment, or a kind of mixed mode of transmission.
In case of mammals, the major inoculation happens during the transition through the birth canal (vaginal and anal microbiota) and during breast feeding (colostrum, milk, and skin microorganisms) (Fernandez et al. 2013; Rodriguez et al. 2015). The colonization of the rumen of various livestock species (cattle, sheep, and goats) was investigated by many studies because of the prospect of effective manipulation of microbiota to improve the capacity to harvest energy from the feed (better utilization of forage with less methane production) (Hook et al. 2010; Yanez-Ruiz et al. 2015). The whole gastrointestinal tract, including the rumen, was assumed to be sterile at birth but is rapidly colonized by microorganisms within the first day of life (Guzman et al. 2015; Ziolecki and Briggs 1961). All major types of rumen microorganisms are already present during the pre-ruminant period in milk-feeding calves (Li et al. 2012). Jami et al. also detected some rumen bacteria essential for mature rumen function as early as one day after birth (Jami et al. 2013). Rey and co-workers showed that the microbial community establishment is rapid after birth and sequential (Rey et al. 2014). Proteobacteria are dominant in the early phase, but, then, they are gradually replaced by Bacteroidetes as the main phylum. Later, between days 3 and 12, the bacterial community includes most of the bacteria present in the developed rumen; however, in different relative abundances. Ciliates are detected in the rumen of young ruminants within two weeks after birth and supposed to be transferred by the saliva of the mother (Eadie 1962). This was proved by isolation of infants from their mother, which led to the lack of protozoa establishment in the rumen (Bryant and Small 1960; Eadie 1962). Anaerobic fungi (e.g., Neocallimastix frontalis) are also able to develop in the rumen before feeding on solid substrates as shown previously (Fonty et al. 1987). Early establishment (days 2 to 4) of methanogenic archaea long before the first solid feeding was shown in lamb rumen (Fonty et al. 1987). By the application of modern molecular techniques, the detection of methanogens at an even earlier stage (first day or neonatal stage) of dairy calves was possible (Guzman et al. 2015). Transition from liquid (milk) to solid feed has a major impact on the community structure as shown by many studies (Jami et al. 2013; Rey et al. 2014). It can be concluded that the ruminal microbial community establishment happens before intake of solid food, but, then, the arrival of solid substrate shapes the community structure (Rey et al. 2014) and the anatomic development occurring at last (Jiao et al. 2015). Factors influencing early life colonization (presence of mother vs. artificial feeding etc.) of the rumen have been reviewed in detail by Yanez-Ruiz et al. (2015).
Management of the microbiota of various gut and AD systems
Due to the important role of the microbiota in nutrition of various ruminant livestock species, the possibility to influence the microbial community in order to improve the performance of the animals (better utilization of the fodder) has attracted lots of research in the last decades (Chalupa 1977; Hart et al. 2008; Yanez-Ruiz et al. 2015). Some part of knowledge obtained from engineering attempts of animal gut microbiota might be transferred to AD systems.
Weimer et al. (2010) investigated the stability and host specificity of the ruminal bacterial communities of a cow following a massive exchange of the rumen content including active microbiota derived from another cow. The microbial community structure was altered but returned to a state resembling more to the original structure than the one from the donor. Mitigation of methane emission is an important issue in case of ruminants, not just to reduce the GHG emission from the animal husbandry sector but also to better utilize the carbon in the fodder converting it into meat rather than into methane (Hook et al. 2010). In AD reactors, the enhancement of methane production is usually aimed; however, such reactors can also be utilized to produce carboxylates as value-added products (Agler et al. 2011; Janke et al. 2016; Urban et al. 2017). In such cases, the inhibition of methanogenesis is desired. A review on the potential utilization of plant extracts, such as essential oils, saponins, and organosulfurous compounds, to manipulate rumen fermentation was published previously (Hart et al. 2008). Bromochloromethane (BCM), as a well-known inhibitor of methanogenesis, was applied to young goats to alter their methanogenic community structures (Abecia et al. 2013, 2014). The treatment resulted in a reduction of methane emission but could not completely eliminate it, and 4 months after the treatment the community structure was similar to the ones in non-treated specimens (Abecia et al. 2014). Fonty et al. (2007) inoculated gnotobiotic lambs with a functional rumen microbiota lacking methanogens. Reductive acetogenesis was an effective H2 sink and sustained a functional rumen; however, H2 utilization was much lower than in lambs with ruminal methanogens. Additional examples and more details about the potential re-programming of the rumen microbiota can be found in the detailed review by Yanez-Ruiz et al. (2015). A not yet fully confirmed hypothesis of that review is that once the rumen community is more or less constant, there is little room for alteration. Similar difficulties might arise during attempts of microbial community modification of stable (steady state) anaerobic digesters. Application of living microorganisms by addition to the fodder (direct-fed microbes) is applied for ruminants to improve feed utilization (Klieve et al. 2003; Martin and Nisbet 1992) or to mitigate methane emission (Jeyanathan et al. 2014). This approach is similar to the application of probiotics in human medicine. Probiotics are defined as live microorganisms that, when administered in adequate amounts, confer a health benefit on the host (Hill et al. 2014). The application of probiotics in animal production was previously reviewed (Kmet et al. 1993; Musa et al. 2009).
Bioaugmentation is a well-established biotechnological method to introduce microorganisms to a bioprocess. It is very similar to the application of probiotics used for health improvement via influencing the gut microbiota. There are many requirements for an effective probiotic treatment, and, in an analogous way, similar prerequisites can be listed for strains used for bioaugmentation (Table 1)
There are many studies at laboratory-scale demonstrating positive effects of bioaugmentation (Table 2), but only few strains or complex cultures fulfill the criteria described in Table 1.
Successful recovery and increase of methane production compared to the control was achieved by adding a propionate-degrading enrichment culture to an overloaded reactor (Tale et al. 2011) involved in degradation of propionate or long-chain fatty acids. Remediation of ammonia toxicity of biogas reactors was also possible by adding either a pure culture of the ammonia-tolerant methanogen Methanoculleus bourgensis MS2 or a mixed culture containing M. bourgensis as a predominant methanogen (Fotidis et al. 2017; Fotidis et al. 2014).
Improving the AD of lignocellulose with microorganisms having excellent degradation properties has been proposed and investigated by many studies (Cater et al. 2015). Adding living microorganisms instead of free enzymes is, in theory, more efficient, because microorganisms can regenerate and produce various useful enzymes at the same time. Bagi and co-workers tested the enhancement potential of two H2-producing strains, Enterobacter cloacae and Caldicellulosiruptor saccharolyticus, using a complex substrate in batch experiment. Biogas production increased by 160–170% compared to the control, which was partially due to the cellulolytic activity of the second strain (Bagi et al. 2007). The enhancement potential of these two strains was later also tested in a series of continuous feeding experiments in CSTRs applying various substrates and reactor conditions (Ács et al. 2015; Herbel et al. 2010; Kovacs et al. 2013) (Table 2). Another strain of the genus Caldicellulosiruptor, namely Caldicellulosiruptor bescii, was successfully used to enhance methane production from steam-explosion treated birch in a batch experiment (Mulat et al. 2018). Another target genus of bioaugmentation is Clostridium. The mesophilic Clostridium cellulolyticum (Desvaux 2005) and the thermophilic Clostridium thermocellum (Akinosho et al. 2014), species well adapted to a cellulolytic lifestyle, were successfully applied to enhance AD of various lignocellulosic biomass (Lü et al. 2013; Öner et al. 2018; Peng et al. 2014; Tsapekos et al. 2017) (Table 2). These strains produce cellulosomes and due to their major fermentation products consisting mainly of formic, acetic, lactic, and succinic acids, and ethanol besides carbon dioxide and hydrogen, they are also perfect candidates for consolidated bioprocessing (CBP). CBP applications combine the enzyme production, hydrolysis, and fermentation stages into a single step, which, in theory, improve the process efficiency by eliminating the need for addition of exogenous hydrolytic enzymes. The enhancement potential of anaerobic fungi was also demonstrated in batch experiments, but less pronounced effects were observed in continuous experiments (Nkemka et al. 2015; Prochazka et al. 2012) (Table 2).
Comparison of such enhancement experiments is not easy, because the control inoculum plays also an important role in the extent of enhancement. The microbiota in an inoculum from a system not treating lignocellulosic substrates is less accommodated to lignocellulose degradation, and, therefore, a more pronounced enhancement is expected via bioaugmentation. A well-operating biogas reactor treating partially or solely lignocellulosic biomass for a long period can be less effectively enhanced further. Although many successful bioaugmentation efforts were demonstrated in small batch scale, the applications of the same strains in continuous experiments were less efficient and the observed methane yield enhancement was lower and often transient. Similar to the human probiotic applications, this is probably due to the fact that the introduced strains do not became stable members of the AD community. Martin-Ryals and co-workers demonstrated that frequent and repeated bioaugmentation can be rather effective (Martin-Ryals et al. 2015). However, such continuous addition of cultivated microorganisms would be extremely expensive at large-scale AD plants. Another approach is to use mixtures of strains or even more complex microbial communities. Ozbayram and co-workers demonstrated the applicability of enrichment cultures derived from the rumen microbial communities of sheep (Ozbayram et al. 2017), goat, and cow (Ozbayram et al. 2018c). An important finding was that a considerable amount of bioaugmentation culture compared to the indigenous microbiota is necessary for an effective enhancement, and only a minor fraction of the bioaugmentation culture could establish in the final community at the end of the experiments. De Vrieze and Verstraete suggested in a recent review paper (De Vrieze and Verstraete 2016) that an effective strategy could be the introduction of so-called keystone species, which are not involved in the process itself but are required to be present for structural organization or enhanced proliferation and activity of the community members with the desired (hemi)cellulolytic activity. As a recent example, Ács and co-workers demonstrated that addition of a non-cellulolytic Enterobacter cloacae strain successfully enhanced the methane production from maize silage, which was partially achieved by the increased abundance of the polymer-degrading Clostridiales derived from the standard inoculum (Ács et al. 2015). Addition of manure with its complex microbiota, including effective fiber-degrading strains or ensiled plant biomass containing lactobacilli, has many beneficial effects observed by biogas plant operators. However, the positive effects of the introduced complex microbiota cannot be clearly separated from the ones originating from the substrate (trace elements, lactate, pre-digested polymers). According to the strict definition of probiotics, foods containing potentially beneficial live and active cultures should not be called probiotics (Hill et al. 2014).
Despite promising laboratory-scale demonstration of bioaugmentation, successful large-scale applications are still missing. This is probably due to the fact that alteration of a stable microbial community by introducing allochthonous microbiota is difficult, similarly as it applies to established gut microbiota. In case of process failure or underperformance, the addition of still a large number of cells would be needed for effective establishment of the bioaugmented microorganisms.
Genome mining for lignocellulose-modifying enzymes appropriate for industrial applications
Although gut symbionts play crucial roles in lignocellulose degradation, many animals can also produce endogenous enzymes that contribute in a complementary way to the overall biomass utilization. The host input was mainly investigated in case of lower termites (Cairo et al. 2011). In several studies, the whole digestome, defined as the pool of host and symbiont genes that collaborate for high efficiency lignocellulose digestion, was investigated by metagenomics or metatranscriptomics (Tartar et al. 2009). While the symbionts play major roles in the hindgut, the host enzymes are produced mainly in the salivary glands and act in the foregut and middle gut. Relatively high levels of oxygen can be detected in these compartments compared to the hindgut, which promotes lignin modification (Ke and Chen 2013; Ke et al. 2010). Tartar and co-workers identified several genes with potential roles in either lignin degradation or protection from toxic metabolites (e.g., reactive oxygen species) generated during lignin degradation in the lower termite Reticulitermes flavipes via metatranscriptomic approach (Tartar et al. 2009). Laccases, catalases, esterases, cytochrome P450, superoxide dismutases, epoxide hydrolases, and glutathione peroxidases are produced by the host, and they are probably involved in the degradation of lignin and its metabolites. In addition, genes encoding endogenous carbohydrate active enzymes associated with cellulose degradation and related to various glycosyl hydrolase families (GHF) were also detected. These enzymes are potential targets of bioprospecting. Especially, addition of lignin-modifying enzymes could be part of pretreatment strategies in gut-inspired biorefinery systems. There are many enzyme preparations commercially available for biorefinery applications, mainly for bioethanol production, but all are similar in composition and have been mostly optimized for acid-pretreated corn stover (Banerjee et al. 2010). However, mimicking gut systems would require other pretreatment chemistry and developed enzyme cocktails should be compatible with mimicked gut conditions (e.g., alkaline conditions in case of insect larvae systems). Besides endogenous enzymes, microbial symbionts are also potential sources of novel enzymes that can be derived by cultivating them. However, obtaining pure cultures is quite complicated in case of many anaerobic microorganisms. Molecular techniques provide an alternative approach, i.e., genes can be obtained by metagenomics and metatranscriptomics and then expressed in heterologous systems. Subsequently, their enzymatic activity can be screened and potential well-performing lignin- or carbohydrate-active candidates can be used for mass production. This approach was demonstrated by Hess and co-workers by sequencing and analyzing 268 Gb of metagenomic DNA from microbes adherent to plant fibers incubated in cow rumen (Hess et al. 2011). From this dataset, 15 metagenome-assembled microbial genomes were reconstructed together with 27,755 genes encoding putative carbohydrate-active enzymes. A selection of 90 candidate genes were expressed, of which the majority produced proteins that were enzymatically active against cellulosic substrates, some of them with low sequence similarity to known enzymes. The general approach presented in this study is applicable to other gut systems and various environmental and engineered systems. A similar metagenomics approach was used for the investigation of camel rumen (Gharechahi and Salekdeh 2018), moose rumen (Svartström et al. 2017), Vietnamese native goat rumen (Do et al. 2018), Indian buffalo rumen (Singh et al. 2014), goat rumen (Lim et al. 2013), and cow rumen (Dai et al. 2015; Stewart et al. 2018) . However, recent studies showed that ruminant feces are probably a poor proxy for the lignocellulolytic potential of the host (Al-Masaudi et al. 2017); therefore, its utilization as inoculum or as target of genome mining is limited. Heterologous expression of the novel genes and testing of enzyme activity were only performed in few cases, including heterologous expression of lignocellulolytic proteins cow rumen (Del Pozo et al. 2012) and buffalo rumen (Shah et al. 2017).
Besides the bottlenecks of finding novel enzymes with good biotechnological potential (Ferrer et al. 2016), the production costs should be drastically reduced for economic application in AD systems. An alternative approach is the production of lignocellulolytic enzymes directly within the plant biomass via the so called in-planta expression approach (Abdeev et al. 2003; Borkhardt et al. 2010; Harrison et al. 2011, 2014; Jiang et al. 2011). Enzyme expression can be regulated in a way that they are expressed at a particular stage of development or specifically induced. The potential of in-planta lignocellulolytic enzyme production was reviewed elsewhere (Willis et al. 2016).
Potential biomimicry of the gut in advanced reactor engineering
We should be careful when copying the concept of the digestive tract into an engineered system because various gut systems have no uniform features, such as shape, flux, and mixing. The idea to simulate animal digestive tracts in engineered reactors is not new. Many attempts were made to enhance the methane yield in biogas reactors inspired by ruminants. The most well-known systems are the rumen simulation technique (RUSITEC) (Czerkawski and Breckenridge 1977) and the rumen-derived anaerobic digestion system (RUDAD) (Gijzen et al. 1988). Briefly, an anaerobic digester is seeded by rumen-based microbial inoculum in the RUSITEC system, whereas RUDAD is a two-stage system composed of an acidogenic rumen reactor and a high rate methanogenic reactor to sustain optimum conditions for both hydrolysis and methanogenesis. Two-stage anaerobic digester systems are quite similar to RUDAD and show better performance compared to single-stage digesters (Akobi et al. 2016; Lindner et al. 2016). It was reported that a few RUDAD systems were constructed and operated in industrial scale to treat municipal solid wastes (Deublein and Steinhauser 2008).
In a recent study carried out by Bize and co-workers, a biomimetic approach was integrated in anaerobic digesters, which the authors designated as bovid-like engineered digestive system (Bize et al. 2015). They achieved better performance in COD removal in the systems inoculated with cow rumen inoculum. Due to their powerful skills in lignocellulose degradation, a considerable literature has grown up around the theme of termite digestive tract for almost a century (Brune and Dietrich 2015). However, most of the work carried out on industrial application failed to scale-up this cellulolytic system. While termites have mastication, we have pre-treatment steps for size reduction in the engineered systems, which play a vital role for effective degradation rates in the further steps. It is an energy-consuming step contributing the greatest share to the operational costs (Watanabe and Tokuda 2010).
Lignin has a complex structure resisting biochemical degradation and limits lignocellulose degradation. Oxygen is necessary for lignin degradation/modification, which acts as co-substrate during the oxidative enzymatic breakdown (Breznak and Brune 1994; Scharf and Tartar 2008). Ke and co-workers showed that termites can also modify and/or break down lignin compounds through their gut system. Degradation starts in the foregut and then continues in the midgut of the termites, which is not completely anaerobic (Ke et al. 2011). Oxic treatment steps can be integrated in the biomass utilization systems as a first step, or a biomimicking reactor system might include a pre-digestion reactor that is not strictly anoxic.
Godon and his colleagues suggested some points to be taken into consideration while biomimicking animal digestive tracts (Godon et al. 2016). In summary, adjusting mesophilic temperature, extreme pH hydrolysis (acidic for vertebrates, alkaline for insects), using grinding approach rather than cutting, specific enzyme addition, and including oxidative enzymes could be incorporated into gut-inspired future AD systems. Alkaline pre-treatment was integrated in AD processes, and results revealed significant enhancement on methane yield (Janke et al. 2017; Sambusiti et al. 2013; Sträuber et al. 2015; Zheng et al. 2009). As another approach, size reduction has positive effects on biogas yield in digesters treating lignocellulosic feedstock as it promotes the hydrolysis rate by increasing the surface area (Leite et al. 2015; Silva et al. 2012). Enzyme addition revealed 10–34% enhancement of the methane yield (Bruni et al. 2010; Vervaeren et al. 2010). However, in another study, the biogas yield and methane yield were not affected by enzyme addition (Romano et al. 2009).
Moreover, there are successful applications of co-inoculation of anaerobic digesters with ruminal microbiota. The level on the enhancement of methane yield differed according to the study (Deng et al. 2017; Ozbayram et al. 2018a; Quintero et al. 2012; Wall et al. 2015). Furthermore, our recent studies showed that bioaugmentation with rumen-derived microbial communities also enhanced the methane production in biogas reactors operated in batch mode (Ozbayram et al. 2017, 2018c). However, another attempt to use rumen fluid as co-inoculum was not significantly effective on cellulose degradation compared to sludge from a municipal solid waste digester alone (Chapleur et al. 2014), which means that the enhancement effect is system-dependent.
Perspectives and future challenges
Lignocellulosic biomass has still a great and largely untapped potential for the biorefinery approach producing high-value chemicals and energy carriers. Current engineered systems are underperforming, especially in comparison to the gut systems of few herbivorous animals specialized for lignocellulosic biomass consumption. The new generation of meta-omics technologies enable us to better understand the background of this superior performance and hopefully provide us useful information for the significant improvement of our current technological systems.
However, different criteria apply for evolution driven by natural selection and economic considerations for industrial applications. Biomimicry of the gut systems might bring us closer to faster and significantly better lignocellulose to biofuel conversion; however, the invested cost will not necessarily be covered by the higher profit as a result of increased biogas or carboxylate production. Nevertheless, it would be still beneficial to establish and explore such improved engineered systems based on gut biomimicry. Later on, some costly treatment elements of the gut-based engineered system could be omitted and the decrease of the conversion rate could be investigated for further economic optimization. We can expect biotechnological developments in other fields that may contribute to a more economic animal gut mimicking system development. For example, a full implementation of the animal gut strategies might require addition of external enzymes, but current relatively high prizes of (hemi)cellulolytic enzymes prevent such strategies. Alternative solutions could be the implementation of in-planta expression of particular enzymes in energy plants as a strategy for accelerated lignocellulose digestion (Mir et al. 2017; Willis et al. 2016). Despite the challenges, the gut structure and inhabiting microbiota, the interaction of the community members, the enzymes and enzyme complexes evolved over millions of years will provide inspirations and valuable resources for process engineers to improve lignocellulose biorefineries.
References
Abdeev RM, Goldenkova IV, Musiychuk KA, Piruzian ES (2003) Expression of a thermostable bacterial cellulase in transgenic tobacco plants. Russ J Genet 39(3):300–305
Abecia L, Martin-Garcia AI, Martinez G, Newbold CJ, Yanez-Ruiz DR (2013) Nutritional intervention in early life to manipulate rumen microbial colonization and methane output by kid goats postweaning. J Anim Sci 91(10):4832–4840. https://doi.org/10.2527/jas.2012-6142
Abecia L, Waddams KE, Martinez-Fernandez G, Martin-Garcia AI, Ramos-Morales E, Newbold CJ, Yanez-Ruiz DR (2014) An antimethanogenic nutritional intervention in early life of ruminants modifies ruminal colonization by Archaea. Archaea:Artn 841463. https://doi.org/10.1155/2014/841463
Ács N, Bagi Z, Rakhely G, Minarovics J, Nagy K, Kovacs KL (2015) Bioaugmentation of biogas production by a hydrogen-producing bacterium. Bioresour Technol 186:286–293
Agler MT, Wrenn BA, Zinder SH, Angenent LT (2011) Waste to bioproduct conversion with undefined mixed cultures: the carboxylate platform. Trends Biotechnol 29(2):70–78. https://doi.org/10.1016/j.tibtech.2010.11.006
Aguirre-Villegas HA, Larson RA (2017) Evaluating greenhouse gas emissions from dairy manure management practices using survey data and lifecycle tools. J Clean Prod 143:169–179. https://doi.org/10.1016/j.jclepro.2016.12.133
Akinosho H, Yee K, Close D, Ragauskas A (2014) The emergence of Clostridium thermocellum as a high utility candidate for consolidated bioprocessing applications. Front Chem 2:Article 66. https://doi.org/10.3389/fchem.2014.00066
Akobi C, Yeo H, Hafez H, Nakhla G (2016) Single-stage and two-stage anaerobic digestion of extruded lignocellulosic biomass. Appl Energy 184:548–559
Al-Masaudi S, El Kaoutari A, Drula E, Al-Mehdar H, Redwan EM, Lombard V, Henrissat B (2017) A metagenomics investigation of carbohydrate-active enzymes along the gastrointestinal tract of Saudi sheep. Front Microbiol 8
Andert J, Marten A, Brandl R, Brune A (2010) Inter- and intraspecific comparison of the bacterial assemblages in the hindgut of humivorous scarab beetle larvae (Pachnoda spp.). FEMS Microbiol Ecol 74(2):439–449. https://doi.org/10.1111/j.1574-6941.2010.00950.x
Aneja VP, Schlesinger WH, Erisman JW (2009) Effects of agriculture upon the air quality and climate: research, policy, and regulations. Environ Sci Technol 43(12):4234–4240. https://doi.org/10.1021/es8024403
Auer L, Lazuka A, Sillam-Dusses D, Miambi E, O'Donohue M, Hernandez-Raquet G (2017) Uncovering the potential of termite gut microbiome for lignocellulose bioconversion in anaerobic batch bioreactors. Front Microbiol 8:ARTN 2623. https://doi.org/10.3389/fmicb.2017.02623
Bagi Z, Acs N, Balint B, Horvath L, Dobo K, Perei KR, Rakhely G, Kovacs KL (2007) Biotechnological intensification of biogas production. Appl Microbiol Biotechnol 76(2):473–482
Banerjee G, Scott-Craig JS, Walton JD (2010) Improving enzymes for biomass conversion: A basic research perspective. Bioenerg Res 3(1):82–92. https://doi.org/10.1007/s12155-009-9067-5
Bauer E, Lampert N, Mikaelyan A, Kohler T, Maekawa K, Brune A (2015) Physicochemical conditions, metabolites and community structure of the bacterial microbiota in the gut of wood-feeding cockroaches (Blaberidae: Panesthiinae). FEMS Microbiol Ecol 91(2). https://doi.org/10.1093/femsec/fiu028
Bayane A, Guiot SR (2011) Animal digestive strategies versus anaerobic digestion bioprocesses for biogas production from lignocellulosic biomass. Rev Environ Sci Biotechnol 10(1):43–62. https://doi.org/10.1007/s11157-010-9209-4
Berasategui A, Shukla S, Salem H, Kaltenpoth M (2016) Potential applications of insect symbionts in biotechnology. Appl Microbiol Biotechnol 100(4):1567–1577. https://doi.org/10.1007/s00253-015-7186-9
Bize A, Cardona L, Desmond-Le Quemener E, Battimelli A, Badalato N, Bureau C, Madigou C, Chevret D, Guillot A, Monnet V, Godon JJ, Bouchez T (2015) Shotgun metaproteomic profiling of biomimetic anaerobic digestion processes treating sewage sludge. Proteomics 15(20):3532–3543
Bogner J, Pipatti R, Hashimoto S, Diaz C, Mareckova K, Diaz L, Kjeldsen P, Monni S, Faaij A, Gao QX, Zhang T, Ahmed MA, Sutamihardja RTM, Gregory R (2008) Mitigation of global greenhouse gas emissions from waste: conclusions and strategies from the Intergovernmental Panel on Climate Change (IPCC) Fourth Assessment Report. Working Group III (Mitigation). Waste Manag Res 26(1):11–32. https://doi.org/10.1177/0734242x07088433
Borkhardt B, Harholt J, Ulvskov P, Ahring BK, Jorgensen B, Brinch-Pedersen H (2010) Autohydrolysis of plant xylans by apoplastic expression of thermophilic bacterial endo-xylanases. Plant Biotechnol J 8(3):363–374
Bourguignon T, Lo N, Dietrich C, Sobotnik J, Sidek S, Roisin Y, Brune A, Evans TA (2018) Rampant host switching shaped the termite gut microbiome. Curr Biol 28(4):649–654. https://doi.org/10.1016/j.cub.2018.01.035
Breznak JA (1982) Intestinal microbiota of termites and other xylophagous insects. Annu Rev Microbiol 36:323–343. https://doi.org/10.1146/annurev.mi.36.100182.001543
Breznak JA, Brune A (1994) Role of microorganisms in the digestion of lignocellulose by termites. Annu Rev Entomol 39:453–487
Broucek J (2014) Production of methane emissions from ruminant husbandry: a review. J Environ Prot 5:1482–1493. https://doi.org/10.4236/jep.2014.515141
Brune A (2007) Microbiology—Woodworker's digest. Nature 450(7169):487–488. https://doi.org/10.1038/450487a
Brune A (2014) Symbiotic digestion of lignocellulose in termite guts. Nat Rev Microbiol 12(3):168–180. https://doi.org/10.1038/nrmicro3182
Brune A, Dietrich C (2015) The gut microbiota of termites: digesting the diversity in the light of ecology and evolution. Annu Rev Microbiol 69:145–166. https://doi.org/10.1146/annurev-micro-092412-155715
Brune A, Ohkuma M (2010) Role of the termite gut microbiota in symbiotic digestion. In: Bignell DE, Roisin Y, Lo N (eds) Biology of termites: a modern synthesis. Springer, Dordrecht, pp 439–475
Bruni E, Jensen AP, Angelidaki I (2010) Comparative study of mechanical, hydrothermal, chemical and enzymatic treatments of digested biofibers to improve biogas production. Bioresour Technol 101(22):8713–8717
Bryant MP, Small N (1960) Observations on the ruminal microorganisms of isolated and inoculated calves. J Dairy Sci 43(5):654–667. https://doi.org/10.3168/jds.S0022-0302(60)90216-2
Buan NR (2018) Methanogens: pushing the boundaries of biology. Emerg Top Life Sci 2(4):629–646. https://doi.org/10.1042/etls20180031
Cairo JPLF, Leonardo FC, Alvarez TM, Ribeiro DA, Buchli F, Costa-Leonardo AM, Carazzolle MF, Costa FF, Leme AFP, Pereira GAG, Squina FM (2011) Functional characterization and target discovery of glycoside hydrolases from the digestome of the lower termite Coptotermes gestroi. Biotechnol Biofuels 4
Cater M, Fanedl L, Malovrh S, Logar RM (2015) Biogas production from brewery spent grain enhanced by bioaugmentation with hydrolytic anaerobic bacteria. Bioresour Technol 186:261–269
Cazemier AE, Verdoes JC, Reubsaet FA, Hackstein JH, van der Drift C, Op den Camp HJ (2003) Promicromonospora pachnodae sp. nov., a member of the (hemi)cellulolytic hindgut flora of larvae of the scarab beetle Pachnoda marginata. Antonie Van Leeuwenhoek 83(2):135–148
Chalupa W (1977) Manipulating Rumen Fermentation. J Anim Sci 45(3):585–599
Chapleur O, Bize A, Serain T, Mazeas L, Bouchez T (2014) Co-inoculating ruminal content neither provides active hydrolytic microbes nor improves methanization of 13 C-cellulose in batch digesters. FEMS Microbiol Ecol 87(3):616–629
Choudhury PK, Salem AZM, Jena R, Kumar S, Singh R, Puniya AK (2015) Rumen microbiology: an overview. In: Puniya AK, Singh R, Kamra DN (eds) Rumen microbiology: from evolution to revolution. Springer, New Delhi, pp 3–16
Christy PM, Gopinath LR, Divya D (2014) A review on anaerobic decomposition and enhancement of biogas production through enzymes and microorganisms. Renew Sust Energ Rev 34:167–173
Cragg SM, Beckham GT, Bruce NC, Bugg TDH, Distel DL, Dupree P, Etxabe AG, Goodell BS, Jellison J, McGeehan JE, McQueen-Mason SJ, Schnorr K, Walton PH, Watts JEM, Zimmer M (2015) Lignocellulose degradation mechanisms across the tree of life. Curr Opin Chem Biol 29:108–119. https://doi.org/10.1016/j.cbpa.2015.10.018
Cunha IS, Barreto CC, Costa OYA, Bomfim MA, Castro AP, Kruger RH, Quirino BF (2011) Bacteria and Archaea community structure in the rumen microbiome of goats (Capra hircus) from the semiarid region of Brazil. Anaerobe 17(3):118–124. https://doi.org/10.1016/j.anaerobe.2011.04.018
Czerkawski JW, Breckenridge G (1977) Design and development of a long-term rumen simulation technique (RUSITEC). Brit J Nutr 38(3):371–384
da Silva SS, Chandel AK, Wickramasinghe SR, Dominguez JMG (2012) Fermentative production of value-added products from lignocellulosic biomass. J Biomed Biotechnol: Artn 826162. https://doi.org/10.1155/2012/826162
Dai X, Tian Y, Li JT, Su XY, Wang XW, Zhao SG, Liu L, Luo YF, Liu D, Zheng HJ, Wang JQ, Dong ZY, Hu SN, Huang L (2015) Metatranscriptomic analyses of plant cell wall polysaccharide degradation by microorganisms in the cow rumen. Appl Environ Microbiol 81(4):1375–1386
De Vrieze J, Verstraete W (2016) Perspectives for microbial community composition in anaerobic digestion: from abundance and activity to connectivity. Environ Microbiol 18(9):2797–2809. https://doi.org/10.1111/1462-2920.13437
Del Pozo MV, Fernandez-Arrojo L, Gil-Martinez J, Montesinos A, Chernikova TN, Nechitaylo TY, Waliszek A, Tortajada M, Rojas A, Huws SA, Golyshina OV, Newbold CJ, Polaina J, Ferrer M, Golyshin PN (2012) Microbial beta-glucosidases from cow rumen metagenome enhance the saccharification of lignocellulose in combination with commercial cellulase cocktail. Biotechnol Biofuels 5
Demirel B, Scherer P (2008) The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Rev Environ Sci Biotechnol 7(2):173–190. https://doi.org/10.1007/s11157-008-9131-1
Deng YY, Huang ZX, Ruan WQ, Zhao MX, Miao HF, Ren HY (2017) Co-inoculation of cellulolytic rumen bacteria with methanogenic sludge to enhance methanogenesis of rice straw. Int Biodeterior Biodegradation 117:224–235
Desvaux M (2005) Clostridium cellulolyticum: model organism of mesophilic cellulolytic clostridia. FEMS Microbiol Rev 29(4):741–764
Deublein D, Steinhauser A (2008) Biogas from waste and renewable resources: an introduction. Wiley, Hoboken
Dicks H (2017) A new way of valuing nature: Articulating biomimicry and ecosystem services. Environ Ethics 39(3):281–299
Do TH, Dao TK, Nguyen KHV, Le NG, Nguyen TMP, Le TL, Phung TN, van Straalen NM, Roelofs D, Truong NH (2018) Metagenomic analysis of bacterial community structure and diversity of lignocellulolytic bacteria in Vietnamese native goat rumen. Asian Austral J Anim 31(5):738–747
Doi RH, Kosugi A (2004) Cellulosomes: plant-cell-wall-degrading enzyme complexes. Nat Rev Microbiol 2(7):541–551
Eadie JM (1962) Development of rumen microbial populations in lambs and calves under various conditions of management. J Gen Microbiol 29(4):563. https://doi.org/10.1099/00221287-29-4-563
Egert M, Wagner B, Lemke T, Brune A, Friedrich MW (2003) Microbial community structure in midgut and hindgut of the humus-feeding larva of Pachnoda ephippiata (Coleoptera: Scarabaeidae). Appl Environ Microbiol 69(11):6659–6668. https://doi.org/10.1128/Aem.69.11.6657-6668.2003
Engel P, Moran NA (2013) The gut microbiota of insects—diversity in structure and function. FEMS Microbiol Rev 37(5):699–735. https://doi.org/10.1111/1574-6976.12025
Enzmann F, Mayer F, Rother M, Holtmann D (2018) Methanogens: biochemical background and biotechnological applications. AMB Express 8
Fernandez L, Langa S, Martin V, Maldonado A, Jimenez E, Martin R, Rodriguez JM (2013) The human milk microbiota: origin and potential roles in health and disease. Pharmacol Res 69(1):1–10. https://doi.org/10.1016/j.phrs.2012.09.001
Ferrer M, Martinez-Martinez M, Bargiela R, Streit WR, Golyshina OV, Golyshin PN (2016) Estimating the success of enzyme bioprospecting through metagenomics: current status and future trends. Microb Biotechnol 9(1):22–34
Fonty G, Gouet P, Jouany JP, Senaud J (1987) Establishment of the microflora and anaerobic fungi in the rumen of lambs. J Gen Microbiol 133:1835–1843
Fonty G, Joblin K, Chavarot M, Roux R, Naylor G, Michallon F (2007) Establishment and development of ruminal hydrogenotrophs in methanogen-free lambs. Appl Environ Microbiol 73(20):6391–6403. https://doi.org/10.1128/Aem.00181-07
Fotidis IA, Treu L, Angelidaki I (2017) Enriched ammonia-tolerant methanogenic cultures as bioaugmentation inocula in continuous biomethanation processes. J Clean Prod 166:1305–1313
Fotidis IA, Wang H, Fiedel NR, Luo G, Karakashev DB, Angelidaki I (2014) Bioaugmentation as a solution to increase methane production from an ammonia-rich substrate. Environ Sci Technol 48(13):7669–7676
Fukatsu T, Hosokawa T (2002) Capsule-transmitted gut symbiotic bacterium of the Japanese common plataspid stinkbug, Megacopta punctatissima. Appl Environ Microbiol 68(1):389–396. https://doi.org/10.1128/Aem.68.1.389-396.2002
Gagen EJ, Padmanabha J, Denman SE, McSweeney CS (2015) Hydrogenotrophic culture enrichment reveals rumen Lachnospiraceae and Ruminococcaceae acetogens and hydrogen-responsive Bacteroidetes from pasture-fed cattle. FEMS Microbiol Lett 362(14):fnv104. https://doi.org/10.1093/femsle/fnv104
Gharechahi J, Salekdeh GH (2018) A metagenomic analysis of the camel rumen's microbiome identifies the major microbes responsible for lignocellulose degradation and fermentation. Biotechnol Biofuels 11
Gijzen HJ, Zwart KB, Verhagen FJ, Vogels GP (1988) High-Rate two-phase process for the anaerobic degradation of cellulose, employing rumen microorganisms for an efficient acidogenesis. Biotechnol Bioeng 31(5):418–425. https://doi.org/10.1002/bit.260310505
Godon JJ, Arcemisbehere L, Escudie R, Harmand J, Miambi E, Steyer JP (2013) Overview of the oldest existing set of substrate-optimized anaerobic processes: digestive tracts. Bioenerg Res 6(3):1063–1081. https://doi.org/10.1007/s12155-013-9339-y
Godon JJ, Arulazhagan P, Steyer JP, Hamelin J (2016) Vertebrate bacterial gut diversity: size also matters. BMC Ecol 16:12. https://doi.org/10.1186/s12898-016-0071-2
Güllert S, Fischer MA, Turaev D, Noebauer B, Ilmberger N, Wemheuer B, Alawi M, Rattei T, Daniel R, Schmitz RA, Grundhoff A, Streit WR (2016) Deep metagenome and metatranscriptome analyses of microbial communities affiliated with an industrial biogas fermenter, a cow rumen, and elephant feces reveal major differences in carbohydrate hydrolysis strategies. Biotechnol Biofuels 9:121
Guzman CE, Bereza-Malcolm LT, De Groef B, Franks AE (2015) Presence of selected methanogens, fibrolytic bacteria, and proteobacteria in the gastrointestinal tract of neonatal dairy calves from birth to 72 hours. PLoS One 10(7):ARTN e0133048). https://doi.org/10.1371/journal.pone.0133048
Harrison MD, Geijskes J, Coleman HD, Shand K, Kinkema M, Palupe A, Hassall R, Sainz M, Lloyd R, Miles S, Dale JL (2011) Accumulation of recombinant cellobiohydrolase and endoglucanase in the leaves of mature transgenic sugar cane. Plant Biotechnol J 9(8):884–896
Harrison MD, Zhang ZY, Shand K, Chong BF, Nichols J, Oeller P, O'Hara IM, Doherty WOS, Dale JL (2014) The combination of plant-expressed cellobiohydrolase and low dosages of cellulases for the hydrolysis of sugar cane bagasse. Biotechnol Biofuels 7
Hart KJ, Yanez-Ruiz DR, Duval SM, McEwan NR, Newbold CJ (2008) Plant extracts to manipulate rumen fermentation. Anim Feed Sci Technol 147(1–3):8–35. https://doi.org/10.1016/j.anifeedsci.2007.09.007
He SM, Ivanova N, Kirton E, Allgaier M, Bergin C, Scheffrahn RH, Kyrpides NC, Warnecke F, Tringe SG, Hugenholtz P (2013) Comparative metagenomic and metatranscriptomic analysis of hindgut paunch microbiota in wood- and dung-feeding higher termites. PLoS One 8(4):e61126. https://doi.org/10.1371/journal.pone.0061126e61126
Henderson G, Cox F, Ganesh S, Jonker A, Young W, Global Rumen Census C, Janssen PH (2015) Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range. Sci Rep 5:14567. https://doi.org/10.1038/srep14567
Henderson G, Naylor GE, Leahy SC, Janssen PH (2010) Presence of novel, potentially homoacetogenic bacteria in the rumen as determined by analysis of formyltetrahydrofolate synthetase sequences from ruminants. Appl Environ Microbiol 76(7):2058–2066. https://doi.org/10.1128/Aem.02580-09
Herbel Z, Rakhely G, Bagi Z, Ivanova G, Acs N, Kovacs E, Kovacs KL (2010) Exploitation of the extremely thermophilic Caldicellulosiruptor saccharolyticus in hydrogen and biogas production from biomasses. Environ Technol 31(8–9):1017–1024
Hess M, Sczyrba A, Egan R, Kim TW, Chokhawala H, Schroth G, Luo SJ, Clark DS, Chen F, Zhang T, Mackie RI, Pennacchio LA, Tringe SG, Visel A, Woyke T, Wang Z, Rubin EM (2011) Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331(6016):463–467
Hill C, Guarner F, Reid G, Gibson GR, Merenstein DJ, Pot B, Morelli L, Canani RB, Flint HJ, Salminen S, Calder PC, Sanders ME (2014) The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 11(8):506–514. https://doi.org/10.1038/nrgastro.2014.66
Hongoh Y, Deevong P, Inoue T, Moriya S, Trakulnaleamsai S, Ohkuma M, Vongkaluang C, Noparatnaraporn N, Kudol T (2005) Intra- and interspecific comparisons of bacterial diversity and community structure support coevolution of gut microbiota and termite host. Appl Environ Microbiol 71(11):6590–6599. https://doi.org/10.1128/Aem.71.11.6590-6599.2005
Hongoh Y, Ohkuma M, Kudo T (2003) Molecular analysis of bacterial microbiota in the gut of the termite Reticulitermes speratus (Isoptera; Rhinotermitidae). FEMS Microbiol Ecol 44(2):231–242
Hook SE, Wright ADG, McBride BW (2010) Methanogens: methane producers of the rumen and mitigation strategies. Archaea:Artn 945785. https://doi.org/10.1155/2010/945785
Hosokawa T, Kikuchi Y, Nikoh N, Shimada M, Fukatsu T (2006) Strict host-symbiont cospeciation and reductive genome evolution in insect gut bacteria. PLoS Biol 4(10):1841–1851. https://doi.org/10.1371/journal.pbio.0040337
Huang SW, Zhang HY, Marshall S, Jackson TA (2010) The scarab gut: a potential bioreactor for bio-fuel production. Insect Sci 17(3):175–183. https://doi.org/10.1111/j.1744-7917.2010.01320.x
Inward D, Beccaloni G, Eggleton P (2007) Death of an order: a comprehensive molecular phylogenetic study confirms that termites are eusocial cockroaches. Biol Lett 3(3):331–335. https://doi.org/10.1098/rsbl.2007.0102
Jami E, Israel A, Kotser A, Mizrahi I (2013) Exploring the bovine rumen bacterial community from birth to adulthood. ISME J 7(6):1069–1079. https://doi.org/10.1038/ismej.2013.2
Janke L, Leite A, Batista K, Weinrich S, Strauber H, Nikolausz M, Nelles M, Stinner W (2016) Optimization of hydrolysis and volatile fatty acids production from sugarcane filter cake: effects of urea supplementation and sodium hydroxide pretreatment. Bioresour Technol 199:235–244. https://doi.org/10.1016/j.biortech.2015.07.117
Janke L, Weinrich S, Leite AF, Terzariol FK, Nikolausz M, Nelles M, Stinner W (2017) Improving anaerobic digestion of sugarcane straw for methane production: combined benefits of mechanical and sodium hydroxide pretreatment for process designing. Energ Convers Manag 141:378–389
Janssen PH, Kirs M (2008) Structure of the archaeal community of the rumen. Appl Environ Microbiol 74(12):3619–3625. https://doi.org/10.1128/AEM.02812-07
Jeyanathan J, Martin C, Morgavi DP (2014) The use of direct-fed microbials for mitigation of ruminant methane emissions: a review. Animal 8(2):250–261. https://doi.org/10.1017/S1751731113002085
Jiang XR, Zhou XY, Jiang WY, Gao XR, Li WL (2011) Expressions of thermostable bacterial cellulases in tobacco plant. Biotechnol Lett 33(9):1797–1803
Jiao JZ, Li XP, Beauchemin KA, Tan ZL, Tang SX, Zhou CS (2015) Rumen development process in goats as affected by supplemental feeding v. grazing: age-related anatomic development, functional achievement and microbial colonisation. Brit J Nutr 113(6):888–900. https://doi.org/10.1017/S0007114514004413
Jin W, Cheng YF, Zhu WY (2017) The community structure of Methanomassiliicoccales in the rumen of Chinese goats and its response to a high-grain diet. J Anim Sci Biotechno 1(8):47. https://doi.org/10.1186/s40104-017-0178-0
Karakashev D, Batstone DJ, Angelidaki I (2005) Influence of environmental conditions on methanogenic compositions in anaerobic biogas reactors. Appl Environ Microbiol 71(1):331–338
Ke J, Chen SL (2013) Selective lignin modification process in termites for effective lignocellulose deconstruction. Abstr Pap Am Chem S 245
Ke J, Laskar DD, Singh D, Chen SL (2011) In situ lignocellulosic unlocking mechanism for carbohydrate hydrolysis in termites: crucial lignin modification. Biotechnology for Biofuels 4
Ke J, Sun JZ, Nguyen HD, Singh D, Lee KC, Beyenal H, Chen SL (2010) In-situ oxygen profiling and lignin modification in guts of wood-feeding termites. Insect Sci 17(3):277–290
Kim JS, Lee YY, Kim TH (2016) A review on alkaline pretreatment technology for bioconversion of lignocellulosic biomass. Bioresour Technol 199:42–48. https://doi.org/10.1016/j.biortech.2015.08.085
Kinet R, Destain J, Hiligsmann S, Thonart P, Delhalle L, Taminiau B, Daube G, Delvigne F (2015) Thermophilic and cellulolytic consortium isolated from composting plants improves anaerobic digestion of cellulosic biomass: toward a microbial resource management approach. Bioresour Technol 189:138–144
Kitade O (2004) Comparison of symbiotic flagellate faunae between termites and a wood-feeding cockroach of the genus Cryptocercus. Microbes Environ 19(3):215–220. https://doi.org/10.1264/jsme2.19.215
Klieve AV, Hennessy D, Ouwerkerk D, Forster RJ, Mackie RI, Attwood GT (2003) Establishing populations of Megasphaera elsdenii YE 34 and Butyrivibrio fibrisolvens YE 44 in the rumen of cattle fed high grain diets. J Appl Microbiol 95(3):621–630. https://doi.org/10.1046/j.1365-2672.2003.02024.x
Kmet V, Flint HJ, Wallace RJ (1993) Probiotics and manipulation of rumen development and function. Arch Tierernahr 44(1):1–10
Kohler T, Dietrich C, Scheffrahn RH, Brune A (2012) High-resolution analysis of gut environment and bacterial microbiota reveals functional compartmentation of the gut in wood-feeding hgher termites (Nasutitermes spp.). Appl Environ Microbiol 78(13):4691–4701. https://doi.org/10.1128/Aem.00683-12
Kolsch G, Pedersen BV (2010) Can the tight co-speciation between reed beetles (Col., Chrysomelidae, Donaciinae) and their bacterial endosymbionts, which provide cocoon material, clarify the deeper phylogeny of the hosts? Mol Phylogenet Evol 54(3):810–821. https://doi.org/10.1016/j.ympev.2009.10.038
Kovacs KL, Acs N, Kovacs E, Wirth R, Rakhely G, Strang O, Herbel Z, Bagi Z (2013) Improvement of biogas production by bioaugmentation. Biomed Res Int:Article ID 482653. https://doi.org/10.1155/2013/482653
Kröber M, Bekel T, Diaz NN, Goesmann A, Jaenicke S, Krause L, Miller D, Runte KJ, Viehover P, Puhler A, Schluter A (2009) Phylogenetic characterization of a biogas plant microbial community integrating clone library 16S-rDNA sequences and metagenome sequence data obtained by 454-pyrosequencing. J Biotechnol 142(1):38–49
Lang K, Schuldes J, Klingl A, Poehlein A, Daniel R, Brune A (2015) New mode of energy metabolism in the seventh order of methanogens as revealed by comparative genome analysis of “Candidatus Methanoplasma termitum”. Appl Environ Microbiol 81(4):1338–1352. https://doi.org/10.1128/aem.03389-14
Leite AF, Janke L, Harms H, Zang JW, Fonseca-Zang WA, Stinner W, Nikolausz M (2015) Assessment of the variations in characteristics and methane potential of major waste products from the Brazilian bioethanol industry along an operating season. Energy Fuel 29(7):4022–4029
Lemke T, Stingl U, Egert M, Friedrich MW, Brune A (2003) Physicochemical conditions and microbial activities in the highly alkaline gut of the humus-feeding larva of Pachnoda ephippiata (Coleoptera: Scarabaeidae). Appl Environ Microbiol 69(11):6650–6658
Levy B, Jami E (2018) Exploring the prokaryotic community associated with the rumen ciliate protozoa population. Front Microbiol 9:2526. https://doi.org/10.3389/fmicb.2018.02526
Ley RE, Lozupone CA, Hamady M, Knight R, Gordon JI (2008) Worlds within worlds: evolution of the vertebrate gut microbiota. Nat Rev Microbiol 6(10):776–788. https://doi.org/10.1038/nrmicro1978
Letunic I, Bork P (2019). Interactive Tree Of Life (iTOL) v4: recent updates and new developments."Nucleic Acids Res 47(W1): W256-W259.
Li RW, Connor EE, Li CJ, Baldwin RL, Sparks ME (2012) Characterization of the rumen microbiota of pre-ruminant calves using metagenomic tools. Environ Microbiol 14(1):129–139. https://doi.org/10.1111/j.1462-2920.2011.02543.x
Lim S, Seo J, Choi H, Yoon D, Nam J, Kim H, Cho S, Chang J (2013) Metagenome analysis of protein domain collocation within cellulase genes of goat rumen microbes. Asian Aust J Anim 26(8):1144–1151
Lindner J, Zielonka S, Oechsner H, Lemmer A (2016) Is the continuous two-stage anaerobic digestion process well suited for all substrates? Bioresour Technol 200:470–476
Lü F, Ji JQ, Shao LM, He PJ (2013) Bacterial bioaugmentation for improving methane and hydrogen production from microalgae. Biotechnol Biofuels 6:92. https://doi.org/10.1186/1754-6834-6-92
Lucas R, Kuchenbuch A, Fetzer I, Harms H, Kleinsteuber S (2015) Long-term monitoring reveals stable and remarkably similar microbial communities in parallel full-scale biogas reactors digesting energy crops. FEMS Microbiol Ecol 91(3). https://doi.org/10.1093/femsec/fiv004
Lv Z, Leite AF, Harms H, Glaser K, Liebetrau J, Kleinsteuber S, Nikolausz M (2019) Microbial community shifts in biogas reactors upon complete or partial ammonia inhibition. Appl Microbiol Biotechnol 103(1):519–533
Martin-Ryals A, Schideman L, Li P, Wilkinson H, Wagner R (2015) Improving anaerobic digestion of a cellulosic waste via routine bioaugmentation with cellulolytic microorganisms. Bioresour Technol 189:62–70
Martin SA, Nisbet DJ (1992) Effect of direct-fed microbials on rumen microbial fermentation. J Dairy Sci 75(6):1736–1744. https://doi.org/10.3168/jds.S0022-0302(92)77932-6
Mason PM, Stuckey DC (2016) Biofilms, bubbles and boundary layers—a new approach to understanding cellulolysis in anaerobic and ruminant digestion. Water Res 104:93–100. https://doi.org/10.1016/j.watres.2016.07.063
Mathai PP, Zitomer DH, Maki JS (2015) Quantitative detection of syntrophic fatty acid-degrading bacterial communities in methanogenic environments. Microbiol-Sgm 161:1189–1197. https://doi.org/10.1099/mic.0.000085
Minkley N, Fujita A, Brune A, Kirchner WH (2006) Nest specificity of the bacterial community in termite guts (Hodotermes mossambicus). Insect Soc 53(3):339–344
Mir BA, Myburg AA, Mizrachi E, Cowan DA (2017) In planta expression of hyperthermophilic enzymes as a strategy for accelerated lignocellulosic digestion. Sci Rep Uk 7(1):11462. https://doi.org/10.1038/s41598-017-11026-1
Mulat DG, Huerta SG, Kalyani D, Horn SJ (2018) Enhancing methane production from lignocellulosic biomass by combined steam-explosion pretreatment and bioaugmentation with cellulolytic bacterium Caldicellulosiruptor bescii. Biotechnol Biofuels 11
Musa HH, Wu SL, Zhu CH, Seri HI, Zhu GQ (2009) The potential benefits of probiotics in animal production and health. J Anim Vet Adv 8(2):313–321
Nalepa CA (2015) Origin of termite eusociality: trophallaxis integrates the social, nutritional, and microbial environments. Ecol Entomol 40(4):323–335. https://doi.org/10.1111/een.12197
Nettmann E, Bergmann I, Mundt K, Linke B, Klocke M (2008) Archaea diversity within a commercial biogas plant utilizing herbal biomass determined by 16S rDNA and mcrA analysis. J Appl Microbiol 105(6):1835–1850
Nettmann E, Bergmann I, Pramschufer S, Mundt K, Plogsties V, Herrmann C, Klocke M (2010) Polyphasic analyses of methanogenic archaeal communities in agricultural biogas plants. Appl Environ Microbiol 76(8):2540–2548
Nkemka VN, Gilroyed B, Yanke J, Gruninger R, Vedres D, McAllister T, Hao XY (2015) Bioaugmentation with an anaerobic fungus in a two-stage process for biohydrogen and biogas production using corn silage and cattail. Bioresour Technol 185:79–88. https://doi.org/10.1016/j.biortech.2015.02.100
Noel SJ, Attwood GT, Rakonjac J, Moon CD, Waghorn GC, Janssen PH (2017) Seasonal changes in the digesta-adherent rumen bacterial communities of dairy cattle grazing pasture. PLoS One 12(3):ARTN e0173819). https://doi.org/10.1371/journal.pone.0173819
Nyonyo T, Shinkai T, Mitsumori M (2014) Improved culturability of cellulolytic rumen bacteria and phylogenetic diversity of culturable cellulolytic and xylanolytic bacteria newly isolated from the bovine rumen. FEMS Microbiol Ecol 88(3):528–537
Ohkuma M (2003) Termite symbiotic systems: efficient bio-recycling of lignocellulose. Appl Microbiol Biotechnol 61(1):1–9. https://doi.org/10.1007/s00253-002-1189-z
Okwakol MJN (1980) Estimation of soil and organic-matter consumption by termites of the genus Cubitermes. Afr J Ecol 18(1):127–131. https://doi.org/10.1111/j.1365-2028.1980.tb00276.x
Omar B, Abou-Shanab R, El-Gammal M, Fotidis IA, Kougias PG, Zhang YF, Angelidaki I (2018) Simultaneous biogas upgrading and biochemicals production using anaerobic bacterial mixed cultures. Water Res 142:86–95. https://doi.org/10.1016/j.watres.2018.05.049
Öner BE, Akyol C, Bozan M, Ince O, Aydin S, Ince B (2018) Bioaugmentation with Clostridium thermocellum to enhance the anaerobic biodegradation of lignocellulosic agricultural residues. Bioresour Technol 249:620–625
Oren A (2014) The Family Methanotrichaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes: other major lineages of bacteria and the Archaea. Springer, Berlin Heidelberg, pp 297–306
Ozbayram EG, Akyol C, Ince B, Karakoc C, Ince O (2018a) Rumen bacteria at work: bioaugmentation strategies to enhance biogas production from cow manure. J Appl Microbiol 124(2):491–502. https://doi.org/10.1111/jam.13668
Ozbayram EG, Ince O, Ince B, Harms H, Kleinsteuber S (2018b) Comparison of rumen and manure microbiomes and implications for the inoculation of anaerobic digesters. Microorganisms 6:15. https://doi.org/10.3390/microorganisms6010015
Ozbayram EG, Kleinsteuber S, Nikolausz M, Ince B, Ince O (2017) Effect of bioaugmentation by cellulolytic bacteria enriched from sheep rumen on methane production from wheat straw. Anaerobe 46:122–130. https://doi.org/10.1016/j.anaerobe.2017.03.013
Ozbayram EG, Kleinsteuber S, Nikolausz M, Ince B, Ince O (2018c) Bioaugmentation of anaerobic digesters treating lignocellulosic feedstock by enriched microbial consortia. Eng Life Sci 18(7):440–446. https://doi.org/10.1002/elsc.201700199
Ozbayram EG, Kleinsteuber S, Nikolausz M, Ince B, Ince O (2018d) Enrichment of lignocellulose-degrading microbial communities from natural and engineered methanogenic environments. Appl Microbiol Biotechnol 102(2):1035–1043. https://doi.org/10.1007/s00253-017-8632-7
Paul SS, Dey A, Baro D, Punia BS (2017) Comparative community structure of archaea in rumen of buffaloes and cattle. J Sci Food Agric 97(10):3284–3293. https://doi.org/10.1002/jsfa.8177
Peng XW, Borner RA, Nges IA, Liu J (2014) Impact of bioaugmentation on biochemical methane potential for wheat straw with addition of Clostridium cellulolyticum. Bioresour Technol 152:567–571
Pitta DW, Kumar S, Veiccharelli B, Parmar N, Reddy B, Joshi CG (2014) Bacterial diversity associated with feeding dry forage at different dietary concentrations in the rumen contents of Mehshana buffalo (Bubalus bubalis) using 16S pyrotags. Anaerobe 25:31–41
Prochazka J, Mrazek J, Strosova L, Fliegerova K, Zabranska J, Dohanyos M (2012) Enhanced biogas yield from energy crops with rumen anaerobic fungi. Eng Life Sci 12(3):343–351
Quintero M, Castro L, Ortiz C, Guzman C, Escalante H (2012) Enhancement of starting up anaerobic digestion of lignocellulosic substrate: fique's bagasse as an example. Bioresour Technol 108:8–13
Rahman NA, Parks DH, Vanwonterghem I, Morrison M, Tyson GW, Hugenholtz P (2016) A Phylogenomic analysis of the bacterial phylum fibrobacteres. Front Microbiol 7(6):1469. https://doi.org/10.3389/fmicb.2015.01469
Ransom-Jones E, Jones DL, McCarthy AJ, McDonald JE (2012) The Fibrobacteres: an important phylum of cellulose-degrading bacteria. Microb Ecol 63(2):267–281
Rastogi G, Ranade DR, Yeole TY, Patole MS, Shouche YS (2008) Investigation of methanogen population structure in biogas reactor by molecular characterization of methyl-coenzyme M reductase A (mcrA) genes. Bioresour Technol 99(13):5317–5326
Rey M, Enjalbert F, Combes S, Cauquil L, Bouchez O, Monteils V (2014) Establishment of ruminal bacterial community in dairy calves from birth to weaning is sequential. J Appl Microbiol 116(2):245–257. https://doi.org/10.1111/jam.12405
Rodriguez JM, Murphy K, Stanton C, Ross RP, Kober OI, Juge N, Avershina E, Rudi K, Narbad A, Jenmalm MC, Marchesi JR, Collado MC (2015) The composition of the gut microbiota throughout life, with an emphasis on early life. Microb Ecol Health D 26:ARTN 26050. https://doi.org/10.3402/mehd.v26.26050
Romano RT, Zhang RH, Teter S, McGarvey JA (2009) The effect of enzyme addition on anaerobic digestion of Jose tall wheat grass. Bioresour Technol 100(20):4564–4571. https://doi.org/10.1016/j.biortech.2008.12.065
Rosenberg E, Zilber-Rosenberg I (2011) Symbiosis and development: the hologenome concept. Birth Defects Res C 93(1):56–66. https://doi.org/10.1002/bdrc.20196
Rosenberg E, Zilber-Rosenberg I (2016) Microbes drive evolution of animals and plants: the hologenome concept. Mbio 7(2):ARTN e01395-15). https://doi.org/10.1128/mBio.01395-15
Rosenberg E, Zilber-Rosenberg I (2018) The hologenome concept of evolution after 10 years. Microbiome 6:ARTN 78. https://doi.org/10.1186/s40168-018-0457-9
Rubin EM (2008) Genomics of cellulosic biofuels. Nature 454(7206):841–845. https://doi.org/10.1038/nature07190
Sabree ZL, Huang CY, Arakawa G, Tokuda G, Lo N, Watanabe H, Moran NA (2012) Genome shrinkage and loss of nutrient-providing potential in the obligate symbiont of the primitive termite Mastotermes darwiniensis. Appl Environ Microbiol 78(1):204–210. https://doi.org/10.1128/Aem.06540-11
Sambusiti C, Monlau F, Ficara E, Carrere H, Malpei F (2013) A comparison of different pre-treatments to increase methane production from two agricultural substrates. Appl Energy 104:62–70
Saunois M, Bousquet P, Poulter B, Peregon A, Ciais P, Canadell JG, Dlugokencky EJ, Etiope G, Bastviken D, Houweling S, Janssens-Maenhout G, Tubiello FN, Castaldi S, Jackson RB, Alexe M, Arora VK, Beerling DJ, Bergamaschi P, Blake DR, Brailsford G, Brovkin V, Bruhwiler L, Crevoisier C, Crill P, Covey K, Curry C, Frankenberg C, Gedney N, Hoglund-Isaksson L, Ishizawa M, Ito A, Joos F, Kim HS, Kleinen T, Krummel P, Lamarque JF, Langenfelds R, Locatelli R, Machida T, Maksyutov S, McDonald KC, Marshall J, Melton JR, Morino I, Naik V, O'Doherty S, Parmentier FJW, Patra PK, Peng CH, Peng SS, Peters GP, Pison I, Prigent C, Prinn R, Ramonet M, Riley WJ, Saito M, Santini M, Schroeder R, Simpson IJ, Spahni R, Steele P, Takizawa A, Thornton BF, Tian HQ, Tohjima Y, Viovy N, Voulgarakis A, van Weele M, van der Werf GR, Weiss R, Wiedinmyer C, Wilton DJ, Wiltshire A, Worthy D, Wunch D, Xu XY, Yoshida Y, Zhang B, Zhang Z, Zhu Q (2016) The global methane budget 2000–2012. Earth Syst Sci Data 8(2):697–751. https://doi.org/10.5194/essd-8-697-2016
Sawatdeenarunat C, Surendra KC, Takara D, Oechsner H, Khanal SK (2015) Anaerobic digestion of lignocellulosic biomass: challenges and opportunities. Bioresour Technol 178:178–186. https://doi.org/10.1016/j.biortech.2014.09.103
Scharf ME, Tartar A (2008) Termite digestomes as sources for novel lignocellulases. Biofuels Bioprod Biorefin 2(6):540–552. https://doi.org/10.1002/bbb.107
Schink B (1997) Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol R 61(2):262–280
Schnürer A, Zellner G, Svensson BH (1999) Mesophilic syntrophic acetate oxidation during methane formation in biogas reactors. FEMS Microbiol Ecol 29(3):249–261
Shah RK, Patel AK, Davla DM, Parikh IK, Subramanian RB, Patel KC, Jakhesara SJ, Joshi CG (2017) Molecular cloning, heterologous expression, and functional characterization of a cellulolytic enzyme (Cel PRII) from buffalo rumen metagenome. 3 Biotech 7(4):257. https://doi.org/10.1007/s13205-017-0895-2
Shapira M (2016) Gut microbiotas and host evolution: scaling up symbiosis. Trends Ecol Evol 31(7):539–549. https://doi.org/10.1016/j.tree.2016.03.006
Shi Y, Huang Z, Han S, Fan S, Yang H (2015) Phylogenetic diversity of Archaea in the intestinal tract of termites from different lineages. J Basic Microbiol 55(8):1021–1028. https://doi.org/10.1002/jobm.201400678
Silva GGD, Couturier M, Berrin JG, Buleon A, Rouau X (2012) Effects of grinding processes on enzymatic degradation of wheat straw. Bioresour Technol 103(1):192–200
Singh KM, Reddy B, Patel D, Patel AK, Parmar N, Patel A, Patel JB, Joshi CG (2014) High potential source for biomass degradation enzyme discovery and environmental aspects revealed through metagenomics of Indian buffalo rumen. Biomed Res Int. https://doi.org/10.1155/2014/267189
Siriwongrungson V, Zeny RJ, Angelidaki I (2007) Homoacetogenesis as the alternative pathway for H2 sink during thermophilic anaerobic degradation of butyrate under suppressed methanogenesis. Water Res 41(18):4204–4210. https://doi.org/10.1016/j.watres.2007.05.037
Söllinger A, Tveit AT, Poulsen M, Noel SJ, Bengtsson M, Bernhardt J, Hellwing ALF, Lund P, Riedel K, Schleper C, Hojberg O, Urich T (2018) Holistic assessment of rumen microbiome dynamics through quantitative metatranscriptomics reveals multifunctional redundancy during key steps of anaerobic feed degradation. Msystems 3(4):e00038–e00018. https://doi.org/10.1128/mSystems.00038-18
Stewart RD, Auffret MD, Warr A, Wiser AH, Press MO, Langford KW, Liachko I, Snelling TJ, Dewhurst RJ, Walker AW, Roehe R, Watson M (2018) Assembly of 913 microbial genomes from metagenomic sequencing of the cow rumen. Nat Commun 9:870. https://doi.org/10.1038/s41467-018-03317-6
Sträuber H, Bühligen F, Kleinsteuber S, Nikolausz M, Porsch K (2015) Improved anaerobic fermentation of wheat straw by alkaline pre-treatment and addition of alkali-tolerant microorganisms. Bioengineering 2(2):66–93. https://doi.org/10.3390/bioengineering2020066
Svartström O, Alneberg J, Terrapon N, Lombard V, de Bruijn I, Malmsten J, Dalin AM, El Muller E, Shah P, Wilmes P, Henrissat B, Aspeborg H, Andersson AF (2017) Ninety-nine de novo assembled genomes from the moose (Alces alces) rumen microbiome provide new insights into microbial plant biomass degradation. ISME J 11(11):2538–2551
Tale VP, Maki JS, Struble CA, Zitomer DH (2011) Methanogen community structure-activity relationship and bioaugmentation of overloaded anaerobic digesters. Water Res 45(16):5249–5256
Tartar A, Wheeler MM, Zhou X, Coy MR, Boucias DG, Scharf ME (2009) Parallel metatranscriptome analyses of host and symbiont gene expression in the gut of the termite Reticulitermes flavipes. Biotechnol Biofuels 2:25. https://doi.org/10.1186/1754-6834-2-25
Tholen A, Brune A (1999) Localization and in situ activities of homoacetogenic bacteria in the highly compartmentalized hindgut of soil-feeding higher termites (Cubitermes spp.). Appl Environ Microbiol 65(10):4497–4505
Tholen A, Brune A (2000) Impact of oxygen on metabolic fluxes and in situ rates of reductive acetogenesis in the hindgut of the wood-feeding termite Reticulitermes flavipes. Environ Microbiol 2(4):436–449
Tsapekos P, Kougias PG, Vasileiou SA, Treu L, Campanaro S, Lyberatos G, Angelidaki I (2017) Bioaugmentation with hydrolytic microbes to improve the anaerobic biodegradability of lignocellulosic agricultural residues. Bioresour Technol 234:350–359
Urban C, Xu JJ, Sträuber H, Dantas TRD, Muhlenberg J, Hartig C, Angenent LT, Harnisch F (2017) Production of drop-in fuels from biomass at high selectivity by combined microbial and electrochemical conversion. Energy Environ Sci 10(10):2231–2244. https://doi.org/10.1039/c7ee01303e
Vervaeren H, Hostyn K, Ghekiere G, Willems B (2010) Biological ensilage additives as pretreatment for maize to increase the biogas production. Renew Energy 35(9):2089–2093
Vincent JFV, Bogatyreva OA, Bogatyrev NR, Bowyer A, Pahl AK (2006) Biomimetics: its practice and theory. J R Soc Interface 3(9):471–482. https://doi.org/10.1098/rsif.2006.0127
Wahid R, Mulat DG, Gaby JC, Horn SJ (2019) Effects of H2:CO2 ratio and H2 supply fluctuation on methane content and microbial community composition during in-situ biological biogas upgrading. Biotechnol Biofuels 12:ARTN 104. https://doi.org/10.1186/s13068-019-1443-6
Wall DM, Straccialini B, Allen E, Nolan P, Herrmann C, O'Kiely P, Murphy JD (2015) Investigation of effect of particle size and rumen fluid addition on specific methane yields of high lignocellulose grass silage. Bioresour Technol 192:266–271
Warnecke F, Luginbuhl P, Ivanova N, Ghassemian M, Richardson TH, Stege JT, Cayouette M, McHardy AC, Djordjevic G, Aboushadi N, Sorek R, Tringe SG, Podar M, Martin HG, Kunin V, Dalevi D, Madejska J, Kirton E, Platt D, Szeto E, Salamov A, Barry K, Mikhailova N, Kyrpides NC, Matson EG, Ottesen EA, Zhang X, Hernandez M, Murillo C, Acosta LG, Rigoutsos I, Tamayo G, Green BD, Chang C, Rubin EM, Mathur EJ, Robertson DE, Hugenholtz P, Leadbetter JR (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450(7169):560–569. https://doi.org/10.1038/nature06269
Watanabe H, Tokuda G (2010) Cellulolytic systems in insects. Annu Rev Entomol 55:609–632. https://doi.org/10.1146/annurev-ento-112408-085319
Wei SZ (2016) The application of biotechnology on the enhancing of biogas production from lignocellulosic waste. Appl Microbiol Biotechnol 100(23):9821–9836. https://doi.org/10.1007/s00253-016-7926-5
Weimer PJ, Stevenson DM, Mantovani HC, Man SLC (2010) Host specificity of the ruminal bacterial community in the dairy cow following near-total exchange of ruminal contents. J Dairy Sci 93(12):5902–5912. https://doi.org/10.3168/jds.2010-3500
Weiss S, Zankel A, Lebuhn M, Petrak S, Somitsch W, Guebitz GM (2011) Investigation of mircroorganisms colonising activated zeolites during anaerobic biogas production from grass silage. Bioresour Technol 102(6):4353–4359
Welch JG (1982) Rumination, particle-size and passage from the rumen. J Anim Sci 54(4):885–894
Westerholm M, Dolfing J, Schnurer A (2019) Growth characteristics and thermodynamics of syntrophic acetate oxidizers. Environ Sci Technol 53(9):5512–5520. https://doi.org/10.1021/acs.est.9b00288
Westerholm M, Moestedt J, Schnurer A (2016) Biogas production through syntrophic acetate oxidation and deliberate operating strategies for improved digester performance. Appl Energy 179:124–135. https://doi.org/10.1016/j.apenergy.2016.06.061
Willis JD, Mazarei M, Stewart CN (2016) Transgenic plant-produced hydrolytic enzymes and the potential of insect gut-derived hydrolases for biofuels. Front Plant Sci 7:Article 675. https://doi.org/10.3389/fpls.2016.00675
Yan L, Gao YM, Wang YJ, Liu Q, Sun ZY, Fu BR, Wen X, Cui ZJ, Wang WD (2012) Diversity of a mesophilic lignocellulolytic microbial consortium which is useful for enhancement of biogas production. Bioresour Technol 111:49–54
Yanez-Ruiz DR, Abecia L, Newbold CJ (2015) Manipulating rumen microbiome and fermentation through interventions during early life: a review. Front Microbiol 6:ARTN 1133. https://doi.org/10.3389/fmicb.2015.01133
Zabranska J, Pokorna D (2018) Bioconversion of carbon dioxide to methane using hydrogen and hydrogenotrophic methanogens. Biotechnol Adv 36(3):707–720. https://doi.org/10.1016/j.biotechadv.2017.12.003
Zened A, Combes S, Cauquil L, Mariette J, Klopp C, Bouchez O, Troegeler-Meynadier A, Enjalbert F (2013) Microbial ecology of the rumen evaluated by 454 GS FLX pyrosequencing is affected by starch and oil supplementation of diets. FEMS Microbiol Ecol 83(2):504–514
Zhang J, Guo RB, Qiu YL, Qiao JT, Yuan XZ, Shi XS, Wang CS (2015) Bioaugmentation with an acetate-type fermentation bacterium Acetobacteroides hydrogenigenes improves methane production from corn straw. Bioresour Technol 179:306–313
Zheng MX, Li XJ, Li LQ, Yang XJ, He YF (2009) Enhancing anaerobic biogasification of corn stover through wet state NaOH pretreatment. Bioresour Technol 100(21):5140–5145
Zhong WZ, Zhang ZZ, Luo YJ, Sun SS, Qiao W, Xiao M (2011) Effect of biological pretreatments in enhancing corn straw biogas production. Bioresour Technol 102(24):11177–11182
Ziganshin AM, Schmidt T, Scholwin F, Il'inskaya ON, Harms H, Kleinsteuber S (2011) Bacteria and archaea involved in anaerobic digestion of distillers grains with solubles. Appl Microbiol Biotechnol 89(6):2039–2052. https://doi.org/10.1007/s00253-010-2981-9
Ziganshin AM, Ziganshina EE, Kleinsteuber S, Nikolausz M (2016) Comparative analysis of methanogenic communities in different laboratory-scale anaerobic digesters. Archaea. https://doi.org/10.1155/2016/3401272
Zilber-Rosenberg I, Rosenberg E (2008) Role of microorganisms in the evolution of animals and plants: the hologenome theory of evolution. FEMS Microbiol Rev 32(5):723–735. https://doi.org/10.1111/j.1574-6976.2008.00123.x
Ziolecki A, Briggs CAE (1961) Microflora of rumen of young calf: II. Source, nature and development. J Appl Bacteriol 24(2):148–163. https://doi.org/10.1111/j.1365-2672.1961.tb00247.x
Acknowledgments
The study was funded by the Helmholtz Association, Research Program Renewable Energies. Emine Gozde Ozbayram was supported by the Research Fellowship Program of the Scientific and Technological Research Council of Turkey (grant no. 2214A).The authors would like to acknowledge the financial support from BMBF—German Federal Ministry of Education and Research (# 01DN19018).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ozbayram, E.G., Kleinsteuber, S. & Nikolausz, M. Biotechnological utilization of animal gut microbiota for valorization of lignocellulosic biomass. Appl Microbiol Biotechnol 104, 489–508 (2020). https://doi.org/10.1007/s00253-019-10239-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-10239-w