Abstract
Endophytic microorganisms can metabolize organic contaminants and assist in plant growth, thus facilitating the phytoremediation of polluted environments. An endophytic bacterium capable of decoloring malachite green (MG) was isolated from the leaves of the wetland plant Suaeda salsa and was identified as Klebsiella aerogenes S27. Complete decolorization of MG (100 mg/l) was achieved in 8 h at 30 °C and pH 7.0. Ultraviolet-visible spectroscopy and Fourier-transform infrared spectroscopy analyses indicated the degradation of MG by the isolate. The enzymic assays of the strain showed the triphenylmethane reductase (TMR) activity. A gene encoding putative TMR-like protein (named as KaTMR) was cloned and heterologously expressed in Escherichia coli. KaTMR showed only 42.6–43.3% identities in amino acids compared with well-studied TMRs, and it phylogenetically formed a new branch in the family of TMRs. The degraded metabolites by recombinant KaTMR were detected by liquid chromatography-mass spectrometry, showing differences from the products of reported TMRs. The biotransformation pathway of MG was proposed. Phytotoxicity studies revealed the less-toxic nature of the degraded metabolites compared to the dye. This study presented the first report of an endophyte on the degradation and detoxification of triphenylmethane dye via a novel oxidoreductase, thus facilitating the study of the plant-endophyte symbiosis in the bioremediation processes.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Malachite green (4-[(4-dimethylaminophenyl)-phenyl-methyl]-N,N-dimethyl aniline; MG) is a triphenylmethane dye that had been extensively used in dye industries or in aquaculture as an antifungal agent before the year 1993 when it was nominated by the United States Food and Drug Administration (FDA) as a priority chemical for carcinogenicity testing (Zhou et al. 2018; Culp and Beland 1996). Nevertheless, MG is still illegally used because of its low cost and high efficacy (Bañuelos et al. 2016) and has been detected with an accumulation in fish tissues and the aquaculture water (Li et al. 2017; Wu et al. 2017). Among the physical, biological, and chemical methods that have been applied for the elimination of these toxic dyes, phytoremediation is generally considered to be the most promising approach as to its long-term applicabilities using living organisms for the removal of contaminants. So far, a diversity of fungi and algae have been reported with the ability of adsorption and biodegradation of triphenylmethane dyes (Shedbalkar et al. 2008; Daneshvar et al. 2007; Khan et al. 2013), whereas only a few bacteria, such as Citrobacter, Aeromonas, and Pseudomonas, have also been characterized with the capacity to decolorize these dyes (An et al. 2002; Ayed et al. 2010; Du et al. 2012; Wu et al. 2013; Jang et al. 2005; Schlüter et al. 2007; Huan et al. 2011).
In the last decade, different mechanisms have been proposed for these bacteria in biodegradation of malachite green, involving enzymes of triphenylmethane reductase (TMR) (Wang et al. 2012), manganese peroxidase (Yang et al. 2016), laccase (Casas et al. 2009), and tyrosinase (Shedbalkar et al. 2008). The latter three have wild broad substrates, whereas TMR is an enzyme specifically degrading triphenylmethane dyes to their corresponding colorless leucoderivatives (Wang et al. 2012). The tmr genes have been characterized from Gram-negative bacteria such as Pseudomonas (GenBank No. HQ11652), Comamonas (GenBank No. AFJ11835), Exiguobacterium (GenBank No. X436462), Citrobacter (GenBank No. AY756172), and Aeromonas (GenBank No. EF010984). Genetic studies showed the tmr genes might locate in both the genomic DNA and plasmids from these bacteria, and it appears to be a component of a composite transposon associated with a multisubstrate resistance region (Dutta et al. 2014). Although there is no report of plants harboring the tmr genes, evidence showed effective biodegradation of triphenylmethane dye by a transgenic Arabidopsis with the overexpressing of a Citrobacter sp. triphenylmethane reductase (Fu et al. 2013). However, most of the TMRs are intracellular, facing restrictions in biodegradation of MG due to the transport barrier imposed by the cell membranes (Zeng et al. 2013). Moreover, the metabolite leucomalachite green (LMG) produced by TMR is also a toxic inorganic contaminant that is hazardous to the health of humans and other organisms (Zhou et al. 2018). Thus, these drawbacks limited the application of TMRs.
Bacteria of genus Klebsiella have been isolated as plant growth-promoting endophytes or rhizobacteria associated with various plants contributing in the phytoremediation of pollutants such as petroleum-contaminated soil and metalliferous soil (Wei et al. 2014; Rosenblueth et al. 2004; Liu et al. 2015; Ahemad 2015). The species Klebsiella aerogenes (formerly Enterobacter aerogenes) is widely distributed in water, soil, air, and polluted sediment (Iyer et al. 2017). It has been reported to be capable of removing heavy metals (Rudd et al. 1983). Despite the genomic data analysis showing the potential adaptation of this strain to the abiotic stress (Iyer et al. 2017), limited information was currently available for K. aerogenes on its ecological role in phytoremediation or the involved molecular mechanisms.
Here, we isolated an endophytic bacterium, K. aerogenes S27, from a wetland plant Suaeda salsa propagating vegetatively in high-salinity soils in the littoral zone of Yellow River Delta in China (Song and Wang 2015; Liu et al. 2018). The strain S27 was evaluated for its ability to decolorize malachite green at different pH, temperature, and salinity. The degradation products of S27 were analyzed by ultraviolet-visible spectrophotometry (UV-Vis), Fourier-transform infrared spectroscopy (FTIR), and liquid chromatography-mass spectrometry (LC-MS), and showed a less-toxic nature by the phytotoxicity study. Extracellularly triphenylmethane reductase activity was detected from K. aerogenes S27, and a putative tmr-like gene was identified by genomic data mining. The gene was cloned and expressed in Escherichia coli. The detoxification activities and the other biochemical characterizations of this recombinant TMR-like protein were also elucidated. This work may facilitate a better understanding of the role of endophytic bacteria in the phytoremediation of triphenylmethane dyes.
Materials and methods
Isolation and identification of an endophytic bacterium from S. salsa Pall.
Leaves from the plant S. salsa Pall. living along the seaside saline soil of Yellow River Delta in Dongying City of China (GPS coordinates: N 37° 51′, E 118° 46′) were collected and washed thoroughly with running water and sterile water. Approximately 5 g of leaves were immersed in 0.1% Tween 80 for 3 min, followed by 0.1% HgCl2 for 1 min, and then immersing in 75% (v/v) ethanol for 3 min, before being washed with sterilized H2O for five times. The leaves were grounded with a mortar and pestle, added to 5 ml sterile water, and incubated in LB culture at 30 °C for 48 h. The enriched culture was subsequently incubated for a week on LB agar.
To characterize the isolates, the 16S rRNA genes were amplified using primers 27F (AGAGTTTGATCCTGGCTCAG) and 1492R (AGAGTTTGATCCTGGCTCAG) and were sequenced by BioSune Biotechnology (Shanghai, China). For the phylogenetic analysis, the 16S rRNA sequences of the related strains were retrieved from the GenBank database, followed by the alignment by CLUSTAL_X 2.0 program (Larkin et al. 2007). The neighbor-joining method was used to construct the phylogenetic tree by using the MEGA 6 software (Tamura et al. 2013).
Assay of MG decolorization
The decolorization percentage of malachite green was calculated by the decrease in absorbance at 620 nm analyzed by a UV-Vis scanning spectrophotometer (TU -1810, Persee, Beijing, China). Briefly, a single colony of the strain was picked and aerobically cultured overnight in 5 ml LB medium at 30 °C with shaking at 150 rpm. Then, 1% of the pre-cultured cells were transferred to 100 ml fresh LB medium and further incubated at 30 °C to reach the log-status at OD600nm = 1.0. Totally, 10 ml of cells were centrifuged at 4000 rpm for 10 min and resuspended by adding 1 ml MG solution. The MG solution was prepared by solving MG powder in the sterile water to a concentration of 100 mg/l. After incubation for 12–24 h at 30 °C, bacterial cells were removed by centrifugation at 10,000 rpm for 5 min, leaving the supernatant for further measurement. The percentage decolorization was calculated as:
in which A represented initial absorbance and B represented observed absorbance after treatment.
Analyses of degradation products of MG
The final products of malachite green catalyzed by the strain were characterized by UV-Vis and FTIR. MG (100 mg/l) was incubated with the collected K. aerogenes S27 cells as described above at 30 °C for 12 h until the dye turned to be colorless. UV-Vis spectrophotometer (NanoDrop 2000c, Thermo Fisher Scientific, Waltham, USA) was applied to measure the absorption spectrum of 2 μl degraded metabolites with a wavelength range of 190–840 nm. The FTIR spectra of the samples were recorded on a Fourier-transform infrared spectrometer (model Tensor II, Bruker, Karlsruh, Germany) in the mid-IR region of 400–4000 cm−1.
The LC-MS analysis was also carried out to detect the MG metabolites. The S27 cells (100 ml) were collected at their log-status at OD600nm = 1.0 and were incubated at 30 °C for 12 h with 10 ml MG solution at the final concentration of 100 mg/l. The degraded MG metabolites were extracted using 10 ml ethyl acetate, dried in a rotary evaporator, and further dissolved into 1 ml methanol for analysis and filtered using a 0.45-mm membrane filter. For the LC-MS analysis of the metabolites degraded by the enzyme, the assay was carried out in a 5 ml solution containing 50 mM phosphate buffer (pH = 7.0), 1 mg/ml MG, 0.1 mM NADH, and 0.01 mg/ml of the purified recombinant enzyme. The reaction mixture was incubated at 30 °C for 8 h until the dye was removed completely, and was centrifugated at 10,000 rpm for 30 min to remove any possible precipitations. Then, the solution was extracted using an equal volume of ethyl acetate, the extract dried and dissolved in 1 ml methanol. The LC-MS analysis was carried out by using the liquid chromatography-mass spectrometry (Agilent 6530 Accurate-Mass Q-TOF, Agilent Technologies, Palo Alto, California, USA) equipped with a reversed-phase C-18 analytical column of 150 mm × 2.1 mm and 5-μm particle size (XAqua-C18, Acchrom, Beijing, China) with a mobile phase of acetonitrile and 20 mM ammonium acetate. The acetonitrile proportion was initially 5% with ramping gradually to 60% in 10 min and was kept for another 10 min at 60%. Finally, the proportion reached 95% in 5 min and was held for another 5 min. The flow rate was 0.3 ml/min. The analysis was carried out using ESI in positive ion mode with the injection volume of 0.2 μl. The reference wavelength was 360 nm. Other parameters of the mass spectrometer include 345 °C of sheath gas temperature, 11 l/min of sheath gas flow rate, 50 psig of the nebulizer, 4000 V of Vcap, and 500 V of nozzle voltage. The LC-MS analysis was carried out by Agilent Mass Hunter Qualitative Analysis B.04.00 software (Agilent Technologies, Santa Clara, CA, USA).
Effects of different parameters on decolorization assay
To evaluate the effect of salt, pH, and temperature on the decolorization rate of the strain, the decolorization assays were performed under different salt concentrations (2, 3, 4, 5, 6, 7, 8, 9, and 10%), pH values (4.0, 5.0, 6.0, 7.0, 8.0, 9.0, and 10.0) and temperatures (25, 30, 35, and 42 °C) with the same processes as described above. To study the effects of initial dye concentration on decolorization, decolorization experiment was carried out at different initial concentrations of 100, 200, 300, 400, and 500 mg/l at 30 °C. The effect of MG (100–500 mg/l) on the growth of K. aerogenes S27 was measured by counting cell numbers using a hemacytometer. To determine the decolorization specificities of the strain to different dyes, azo magenta, methyl orange, or congo red at 100 mg/l was incubated with the cell culture for the measurement of absorbance using the UV-Vis spectrometer. All experiments in this study were conducted in triplicate.
Enzyme assays of the strain
Activities of laccase, manganese peroxidase, tyrosinase, and malachite green reductase were determined for the cell lysate of the strain. Briefly, pre-cultured cells were harvested by centrifugation at 10,000 rpm for 15 min and were washed for three times with phosphate buffer (100 mM, pH 7.2). Then, cells were suspended in the phosphate buffer and sonicated for 20 min, keeping the sonifier output at 130 W, 20 HZ and giving strokes of 10 s each with a 1-min interval at 4 °C. The suspension was centrifuged at 10,000 rpm for 15 min at 4 °C, and 0.1 ml of supernatant was used as a crude enzyme solution for enzyme analysis. Laccase activity was determined by measuring the oxidation of 2,2’-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) diammonium salt (ABTS) as the substrate at 420 nm in 2 ml mixture containing 10% ABTS in 0.1 M acetate buffer (pH = 5.0) (Du et al. 2012). The laccase activity was calculated using the extinction coefficient of 3.6 × 104 M−1 cm−1. Tyrosinase activity was determined by measuring the liberated catechol quinone at 410 nm with a 3 ml reaction of 0.01% catechol in 100 mM phosphate buffer (pH = 7.4) (Novotny et al. 2001). The tyrosinase activity was calculated using the extinction coefficient of 1.22 × 104 M−1 cm−1. Manganese peroxidase activity was determined by measuring the oxidation product of MnSO4 at 468 nm in a reaction mixture of 3 ml containing 1 mM MnSO4 and 0.1 mM H2O2 in acetate buffer (pH = 5.0). The MG reductase activity was performed by monitoring the decreased intensity at 620 nm with a reaction mixture of 5 ml containing 323 μM MG, 50 μM NADH in 50 mM phosphate buffers (pH 7.4), and 0.1 ml of cell lysate. A unit of laccase or manganese peroxidase activity is defined as an absorbance change in units of min/ml. One unit of MG reductase activity was expressed as the reduction of 1 μg of MG per minute per ml cell lysate or cell culture supernatant.
Cloning, sequencing, and expression of the tmr gene in E. coli
The genomic DNA of strain S27 was extracted by using the DNA extraction kit (Tiangen, Beijing, China). The tmr gene was cloned from the genomic DNA using primers 5′-CGCGGATCCGTCGGCGATTACCGGCGCCAC-3′ (BamHI restriction site underlined) and 5′-CGGCTCGAGTGTGATGCCTTTCACGCTATCG-3′ (XhoI restriction site underlined). The PCR product was purified and inserted between the BamHI and XhoI cutting sites of the pET-22b(+) vector which contains the T7 promoter and the sequence for six histidine residues at the 3′ end of the insertion site (Novagen, Madison, WI, USA). Recombinant vectors were transformed into E. coli BL21 competent cells (Vazyme, Nanjing, China). To induce the expression of the gene, the transformed bacteria were grown in LB medium containing 100 μg/ml ampicillin at 37 °C and 200 rpm to reached an OD600nm of 0.6 and were further induced with 1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) for further incubation at 16 °C for 10 h. To purify the recombinant protein, cells were suspended in 50 mM sodium phosphate buffer (pH 7.0) and disrupted for 20-min sonication giving five strokes each of 10 s at 1-min intervals at 20 Hz on ice. After centrifugation, the supernatant was applied to Ni-NTA agarose (GE Healthcare, Beijing, China), washed with phosphate buffer (pH 7.2) containing 0.3 M NaCl and 20 mM imidazole, and eluted with 500 mM imidazole. All purification procedures were carried out at 4 °C. The purified protein was analyzed by SDS-PAGE, and the protein concentration was determined by Bio-Rad protein assay kit (Beyotime, Shanghai, China) according to the instructions.
Kinetic analysis
To determine the kinetic constant of recombinant TMR for MG, a series of assay solutions dissolved in 1 ml of 50 mM sodium phosphate buffer (pH 7.0) containing various amount of MG (4 to 20 μM) and a fixed concentration of the cofactor NADH (100 μM) and the purified enzyme KaTMR (0.05 mg protein/ml) were incubated at 30 °C (Jang et al. 2005). Each reaction was initiated by adding the enzyme, and the initial reaction rate was determined by monitoring the decrease in absorbance at 620 nm in the first 2 min. Enzyme activity was a linear function of both incubation time and protein concentration. The reciprocals of the substrate concentrations and the reciprocals of the corresponding initial velocities were then used to generate a Lineweaver-Burk plot to determine Km and Vmax values (Jang et al. 2005).
Toxicity studies
The toxicity of MG and its degraded products on the germination of Chinese cabbage seeds was determined according to Du et al. (2015). The seeds of Chinese cabbage were sterilized with 3–5% H2O2 solution for 5 min and washed with sterile H2O for 3–5 times. Twenty seeds of Chinese cabbage were put into individual plates with double sterile filter paper immersed with 7 ml of MG (1000 mg/l) solution or the MG-degraded products. To give the MG-degraded products, 7 ml MG (1000 mg/l) were incubated at 30 °C for 24 h with collected cells from 100 ml of cell suspension cultures with an OD600nm of 1.0 or 0.2 mg recombinant TMR. For the control experiments, sterile H2O was used instead of the MG solutions. All plates were further cultivated at 22 °C with 65% humidity. The germination percentage of the seeds were determined after a week, and the root length of the seeds was also measured.
Accession number
Strain K. aerogenes S27 has been deposited at CCTCC under the accession no. M2017452. The 16S rRNA sequence of K. aerogenes S27 has been deposited at DDBJ/ENA/GenBank under the accession no. MF076897. The gene coding KaTMR has been deposited at DDBJ/ENA/GenBank under the accession no. MH377117.
Results
Isolation of an endophytic bacterium capable of decolorization of dyes
Among 12 bacterial strains isolated from the leaves of plant S. salsa Pall. and screened for decolorization of malachite green in LB medium, a cream-colored bacterium named S27 was selected based on the complete decolorization (above 99%) of MG (100 mg/l) at 30 °C, pH 7.0 in 8 h. The 16S rRNA gene sequence (GenBank No. MF076897) for strain S27 showed high identity (99%) to the type strain K. aerogenes KCTC 2190T (NR_102493) and other K. aerogenes strains such as strain JP27 (MF086659), strain J42 (MF141727), and strain B02 (MF141719). The neighbor-joining phylogenetic tree also revealed that strain S27 was clustered within the genus Klebsiella and most related to the species of K. aerogenes (Supplemental Fig. S1). Thus, this strain was identified as K. aerogenes S27.
The decoloration potential of strain S27 to different dyes was assessed. At 25 °C, the decoloring rate of 94.7% was achieved for MG (100 mg/l), whereas less than 10% decoloring was detected for azo magenta (100 mg/l) and congo red (100 mg/l), and none for the methyl orange (100 mg/l) (Fig. 1). With the increased temperature at 30–42 °C, the complete decolorization (above 99%) for MG has been detected, compared to a slightly increased decoloring rate below 40% for azo magenta and congo red (Fig. 1). Compared with the formerly reported MG-degrading bacteria, strain S27 showed a relative high decolorization rate (2.289 μmol min−1 g−1 DWC) against MG at 274 μM (Supplemental Table S1). These findings suggested that K. aerogenes S27 is capable of decoloring different types of dyes and showed the highest specificity and efficiency to the decoloring of the triphenylmethane dye.
The dye decolorization ability by strain K. aerogenes S27 under different temperatures. Dyes of MG (black bars), azo magenta (dark gray bars), methyl orange (white bars), or congo red (light gray bars) at 100 mg/l were independently incubated for 12 h with collected cells from 10 ml of cell suspension cultures with an OD600nm of 1.0. The absorbances of the solutions were measured by using a UV-Vis spectrometer. Each bar represents the mean ± SE of triplicate cultures
The biodegradation products of MG by K. aerogenes S27
To further clarify the decoloration process of MG by K. aerogenes S27, UV-Vis, FTIR, and LC-MS analyses were performed. UV-Vis absorption spectra showed a prominent peak at 619 nm for MG (100 mg/ml). When incubated with the K. aerogenes S27 cells in the sterile water, this absorption peak gradually decreased and disappeared after 24 h (Fig. 2a), indicating a complete decoloration of MG by the strain. Moreover, the FTIR analysis revealed a significant difference between MG and decolorized products. For MG, the FTIR spectra showed a fingerprint region for the benzene rings from 1500 to 500 cm−1, including a peak at 828 cm−1 corresponding to the aromatic ring structure (Ayed et al. 2008). The vibration bands at 1167 cm−1 and 1365 cm−1 were characteristic peaks of C–N stretching and CH2. Besides, a peak at 1585 cm−1 for the C=C stretching of the benzene rings and the one at 2870 cm−1 for the C–H asymmetric structure showed an amide antisymmetric stretching vibration for MG (Parshetti et al. 2006). Comparably, the degraded MG metabolites by strain S27 gave a peak at 1091 cm−1 for C–N stretch, and a peak at 1660 cm−1 for aromatic ketones, as well as the wide peak around 3000–3500 cm−1 for N–H stretch, represents the formation of primary and secondary amines. In addition, the sharp peak at 2852 cm−1 for C–H stretching by asymmetric CH2 groups indicates the product of methylene-substituted metabolites, whereas the absence of the peak at 828 cm−1 indicates the loss of the aromaticity or benzene ring (Ayed et al. 2008) (Fig. 2a). As to the LC-MS analyses, the significant peak of MG appeared at a retention time of 25 min with the molecular ion peak at m/z 329.2, thereby confirming the chemical structure of MG (Supplemental Fig. S2). The LC-MS analysis of the MG-degraded metabolites showed the absence of the peak of MG but a new peak at a retention time of 1.306 min with the molecular ion peak at m/z 121, corresponding to the structure of N,N-dimethylaniline (Supplemental Fig. S3). The chromatogram of the degraded sample also showed a peak at a retention time of 17.215 min which represents the structure of desmethyl leucomalachite green (m/z 317) (Supplemental Fig. S3).
Analyses of the catabolic intermediates of MG by strain K. aerogenes S27. A, UV-Vis spectral analysis of MG (0.1 μg) before (solid curve) and after (dashed curve) being treated by K. aerogenes S27 cells for 24 h. B, FTIR spectra of MG solution before (curve a) and after (curve b) being treated by the K. aerogenes S27 cells for 24 h
Effect of various conditions on the degradation potential of K. aerogenes S27
To detect the degradation capacity of K. aerogenes S27 to malachite green, 10 ml of the cells at the log-phase (OD600nm = 1.0) was collected and incubated in 1 ml sterile water containing MG at 100, 200, 300, 400, and 500 mg/l. The results showed that 100 mg/l of MG was completely decolorized in 8 h at 30 °C. At the concentration ranging from 100–300 mg/l, above 60% of decolorization of MG was achieved for 100–300 mg/l MG in the initial 5 h (Supplemental Fig. S4). As a control, the untreated MG showed little self-decolorization at 30 °C for 5 h (Supplemental Fig. S5). By comparison, the time for decoloring MG at 400–500 mg/l by the strain was delayed and required 5–7 h (Supplemental Fig. S4), indicating the negative effect of high concentrations of MG on the growth or metabolic activity of strain S27. Thus, we measured the growth rate of K. aerogenes S27 in LB media containing MG at 100–500 mg/l. However, results showed that addition of MG did not significantly inhibit the survival of the strain. With the treatment of increased concentration of MG, the cells delayed to enter their exponential phase, but the cell numbers reached to the same order of magnitude (1010) after 24-h incubation at 30 °C at the stabilized phase (Supplemental Fig. S6). Moreover, a decolorization rate of 500 mg/l MG above 90% was achieved in 16 h (Supplemental Fig. S4), showing the high degrading capacity of K. aerogenes S27 to MG.
Considering the application of high amounts of salts in the textile dyeing, the degradation potential of strain S27 was evaluated at different NaCl concentrations from 2.0 to 10%. As shown in Fig. 3, 90% decolorization was observed within 8 h for salt concentration ranging from 2.0–5.0%. Complete decolorization was observed within 24 h for salt concentration less than 8%. When the concentration increased to 9.0%, the degradation rate of MG reached 97% in 32 h (Fig. 3). However, only 49% of MG was degraded at 32 h when 10% NaCl was supplemented (Fig. 3), which may be due to the negative effect on the growth of strain S27 by such high salt concentration. The effect of pH for decolorization of MG (100 mg/l) was tested at 30 °C for 24 h. Results showed an increased decolorization rate by strain S27 when the pH was increased, with an optimal pH of 8.0 exhibiting a decolorization of MG of 96% (Supplemental Fig. S7). The rate was 94% and 93% at pH 9.0 and pH 10 respectively, but only 53–65% at pH 4.0–6.0 (Supplemental Fig. S7), suggesting that the alkaline conditions are preferable for the decolorization of MG by strain S27.
The effect of NaCl concentration on the decolorization of MG by strain K. aerogenes S27. The decolorization assays were carried out by collecting 10 ml cultured cells at their log-status (OD600nm = 1.0) and incubating them with 100 mg/l MG at 30 °C for 0–32 h under different NaCl concentrations from 2 to 10%. Results are expressed as the means ± SE of triplicate cultures
The candidate enzymes involved in MG degradation
To further investigate the mechanism of strain S27 in MG degradation, the enzymic activities of laccase, tyrosinase, manganese peroxidase, and triphenylmethane reductase were assessed both in the presence and absence of MG. As shown in Table 1, while activities of laccase, tyrosinase, and manganese peroxidase have not been detected, the intracellular TMR activity was 11.6 μg MG reduced/min/ml in the absence of MG and raised to 22.3 μg MG reduced/min/ml in the presence of 100 mg/ml MG. Extracellular TMR activity was detected at 78.9 μg MG reduced/min/ml in the absence of MG and raised to 184.2 μg MG reduced/min/ml in the presence of 100 mg/ml MG, indicating the TMR could be secreted into the cell culture supernatant.
Identification of a tmr-like gene and the phylogenetic analysis
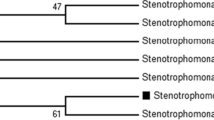
By genomic data mining of the closest related strain of S27, K. aerogenes KCTC 2190T (GenBank No. CP002824.1), also known as the type strain of K. aerogenes (Shin et al. 2012), a gene that encodes a putative TMR-like protein was identified (WP_015704185). Based on the above sequence, the gene named katmr was amplified from strain S27, sequenced and the sequence was deposited in GenBank under the accession number MH377117. Gene katmr encodes a protein consisting of 282 amino acids with a calculated molecular weight of 31.02 kDa. KaTMR shows only 42.6%, 43.3%, and 43.3% amino acid identities with the well-characterized TMRs from Aeromonas hydrophila subsp. decolorationis (ABJ99087, AhTMR) (Ren et al. 2006), Pseudomonas sp. MDB-1 (EF463103, TMR2) (Li et al. 2009), and Citrobacter sp. strain KCTC 18061P (AAW88298, CsTMR) (Jang et al. 2005), respectively, thus indicating a clear difference of KaTMR from the previously characterized TMRs. The HMMSCAN results from Pfam (Finn et al. 2016) showed that KaTMR fell into the NmrA-like family. The phylogenetic analysis showed that KaTMR was clustered in the radiation of the TMRs group but formed a single branch distant from other TMRs (Fig. 4). Taken together, KaTMR represents a novel family of TMR-like proteins with uncharacterized properties.
Phylogenetic tree based on the amino acid sequences of KaTMR characterized in this work (indicated by boldface type), and other reported TMRs and AZRs. Bootstrap values are expressed as percentages of 1000 replications, and only bootstrap values above 50% are shown. GenBank accession numbers are given in parentheses. Bar, 0.2 substitutions per amino acid
Expression, purification, and the enzymic properties of KaTMR
The gene encoding KaTMR was heterologously overexpressed in E. coli BL21. The recombinant KaTMR was purified and detected by SDS-PAGE with a band corresponding to the molecular mass of about 32 kDa (Fig. 5a; Supplemental Fig. S8). The steady-state kinetic constants of the recombinant KaTMR were calculated, and the Lineweaver-Burk plot generated using NADH as a cofactor and MG as the substrate (Fig. 5b). Results showed the values of Km, Vmax values, and kcat/Km are 68.89 μmol/l, 10.76 μmol/l min, and 1.23 s−1 μmol−1, respectively.
Enzymic properties of purified recombinant KaTMR. A, SDS-PAGE analysis of the purified enzyme. Lane 1, the recombinant KaTMR purified from E. coli BL21. Lane M, protein molecular weight marker. B, the Lineweaver-Burk plot of KaTMR activity against MG. The reaction was carried out by incubating 0.05 mg purified KaTMR with 4–20 μM of MG in 50 mM sodium phosphate buffer and using 100 μM of NADH as a cofactor
Identification of the degradation products from MG by KaTMR
To study the probable pathway of the degradation of MG by the recombinant KaTMR, LC-MS analysis was performed after the complete decoloration of MG (1 mg) incubated with 0.01 mg of the purified enzyme. The LC-MS chromatograms of MG-degraded metabolites showed four new peaks at retention times 1.344 min, 13.183 min, 14.82 min, and 20.123 min (Fig. 6) corresponding to N,N-dimethylaniline (m/z 121), (4-dimethylamino-phenyl)-phenyl-methanone (m/z 225), leucomalachite green (m/z 330, LMG), and desmethyl malachite green (m/z 315), respectively. It can be speculated that KaTMR might transform MG into its primary metabolite LMG and then degrades it into the smaller products N,N-dimethylaniline and (4-dimethylamino-phenyl)-phenyl-methanone. Based on these available data, the proposed schematic pathway for the biotransformation of MG is depicted in Fig. 7.
LC-MS profile (A) and extracted ion chromatograms (B–E) for MG metabolites obtained through catalysis by recombinant KaTMR. The analysis was carried out using ESI in positive ion mode. A, LC-MS profile of MG metabolites. B, m/z 122, N,N-dimethylaniline (retention time 1.344 min); C, m/z 226, (4-dimethylamino-phenyl)-phenyl-methanone (retention time 13.183 min); D, m/z 331, leucomalachite green (retention time 14.82 min); E, m/z 315, desmethyl malachite green (retention time 20.123 min)
Toxicity assessment of MG and its degradation products
The phytotoxicity of malachite green solution and its degradation products were evaluated. As shown in Table 2, the germination percentage of Chinese cabbage seeds was significantly inhibited to 80% by treatment with MG, compared with the control treated with water (100%). However, the germination rate raised to 90% when seeds were treated with the degraded MG products by strain S27, and was recovered to 100% when incubated with the degradation products obtained by the recombinant enzyme KaTMR. As to the root length, the average length of the untreated cabbage was 5.2 cm, whereas the length reduced to 1.1 cm when seeds were treated with MG. Comparingly, the roots grew with the products of MG degraded by kaTMR to a length of 1.75 cm while the length significantly increased to a length of 7.53 cm when seeds were incubated with the products obtained by strain S27, indicating the detoxification of MG by both. The amazingly highest length of the enzyme-treated root length was probably achieved by the potential growth-promoting ability of strain S27 as an endophytic bacterium. Besides, we also observed rot and decay in the roots of cabbage incubated with MG solution for ten days, whereas cabbages grown with the degraded MG products kept growing well. Taken together, both strain K. aerogenes S27 and the enzyme kaTMR could detoxify MG in the decolorization process.
Discussion
In this study, we identified an endophytic bacterium capable of degrading MG from a wetland plant S. salsa Pall., an agricultural plant capable of surviving in a high-salinity region in Europe and Asia (Song and Wang 2015). S. salsa is also an accumulator of toxic substances, removing and recovering the heavy metals from the contaminated soils of wetlands (Wu et al. 2012; Song and Wang 2015). Wetlands are sensitive to be affected by pollution sources of organic dyes (Zhou 2001). MG is a kind of toxic and banned dye, but is still being illegally used in some parts of the world (Zhou et al. 2018; Bañuelos et al. 2016). Results indicated that vegetable plants grown in the MG-polluted environment can take up this toxic compound by roots and accumulate it in the plant tissues (Matpang et al. 2017; Fu et al. 2013), which, when eaten, could be harmful to human health. MG at 2 mg/l or 4 mg/l showed significant inhibition of the leaf length, plant height and leaf width of pak choy, Brassica chinensis (Matpang et al. 2017). In this study, we have also shown a visible toxic effect of MG to the germination and growth of Chinese cabbage seeds. To eliminate the toxic substances, bioremediation has proved to be a promising way (Mosa et al. 2016; Meagher 2000). Former studies showed that cell cultures of the plant Blumea malcolmii Hook can degrade MG into 4-dimethylamino-cyclohexa-2,4 dienone (Kagalkar et al. 2011), while Aloe barbadensis was able to transform MG into nontoxic metabolites (Rai et al. 2014); however, the underlying molecular mechanisms have not been disclosed. Plants usually lack the complete machinery in their catabolic pathways for the efficient degradation of organic contaminants since they do not require to utilize these organic compounds. In contrast, these compounds can be taken up by some microbes as their C, N, and energy sources (Eapen et al. 2007). A combination of plant-associated bacteria showed increased bioremediation abilities of the plant hosts in removing these organic pollutants (Mosa et al. 2016; Meagher 2000). In this work, the isolated endophytic bacterium K. aerogenes S27 showed a remarkable triphenylmethane-detoxifying ability by degrading MG into the nontoxic metabolite N,N-dimethylaniline. This may provide new insight into the study of plant-associated bacterial degradation of triphenylmethane chemicals. Besides, strain K. aerogenes S27 showed plant-growth-promoting properties and tolerance to relative high salt concentrations, which might expand its application in phytoremediation of the contaminated ecosystems.
We also disclosed the molecular mechanism of K. aerogenes S27 in degrading MG and characterized a novel oxidoreductase KaTMR. So far, the enzymes involved in the biotransformation of triphenylmethane dyes have not been fully studied. The reported enzymes can be divided into two main groups according to their specificity towards triphenylmethane. The first group includes manganese peroxidase (Yang et al. 2016) and laccase (Casas et al. 2009) which decolorize dyes in an unspecific manner through the free radical chain reaction (Azmi et al. 1998); however, they can be inhibited by chelating agents (Lorenzo et al. 2005), which limited their application in degradation of complex dye effluent. In the second group, enzymes are supposed to be specific to decoloring triphenylmethane dyes, and only the TMRs have been identified in this group. TMRs belong to the extended NAD(H)-dependent dehydrogenase/reductase (SDR) superfamily (Persson et al. 2003). TMRs from Aeromonas hydrophila, Pseudomonas, and Citrobacter have significantly high amino acid sequence similarities (98.6–100%) (Ren et al. 2006; Jang et al. 2005; Li et al. 2009), and they can only transform MG into LMG, which is still toxic to human and other organisms (Zhou et al. 2018). Genomic data mining revealed some gene products having relative low sequence similarities with TMRs and these genes were therefore annotated as TMR-like genes (refer to the data of GenBank); however, the function of these genes remains obscure. Only the catalytic properties of a TMR-like protein (GtAZR) derived from Geobacillus thermoglucosidasius C56-Y593 have been studied, but this enzyme was finally characterized as an azoreductase (AZR) due to its high specificity for azo dyes rather than triphenylmethane dyes (Gao et al. 2015). Here, we charactered a gene encoding a putative TMR-like protein KaTMR with an amino acid similarity of 42.6–43.3% compared to the well-studied TMRs. The recombinant KaTMR displayed a specific substrate spectrum against the triphenylmethane dye. It was phylogenetically clustered in the family of TMRs, but formed a new branch of the family, apart from the azoreductases. LC-MS analysis showed that recombinant KaTMR could transform MG into the smaller molecules N,N-dimethylaniline and (4-dimethylamino-phenyl)-phenyl-methanone, leading to the degradation of the canonical resonance substructures. Based on the distinct sequence and enzymic property, KaTMR represents a new oxidoreductase distinct from the well-studied TMRs.
Besides, we found that the degraded products N,N-dimethylaniline and (4-dimethylamino-phenyl)-phenyl-methanone are same as the MG-degraded metabolites obtained by Enterobacter asburiae strain XJUHX-4TM (Mukherjee and Das 2014), a species belonging to a genus close to Klebsiella (Supplemental Fig. S1), but the molecular mechanism underlying the degradation of MG by E. asburiae has not been fully disclosed. The characterization of KaTMR may provide insights into the molecular mechanism of the related strains in the phytoremediation of triphenylmethane dyes.
In conclusion, here we isolated an endophytic bacterium K. aerogenes S27 from the wetland plant S. salsa and confirmed its ability in degrading MG into smaller and less-toxic metabolites. We also provided evidence to show that a new oxidoreductase KaTMR is involved in the degradation and detoxification of toxic compounds by Klebsiella. KaTMR can degrade MG into the smaller and nontoxic molecules N,N-dimethylaniline and (4-dimethylamino-phenyl)-phenyl-methanone. These findings indicate potential applications for a novel enzyme as well as of the plant-endophyte system in bioremediation processes of triphenylmethane dyes. Moreover, the heterologous expression and purification of KaTMR achieved in this work will facilitate its potential use in industrial dye effluent purification.
References
Ahemad M (2015) Phosphate-solubilizing bacteria-assisted phytoremediation of metalliferous soils: a review. 3 Biotech 5:111–121. https://doi.org/10.1007/s13205-014-0206-0
An SY, Min SK, Cha IH, Choi YL, Cho YS, Kim CH, Lee YC (2002) Decolorization of triphenylmethane and azo dyes by Citrobacter sp. Biotechnol Lett 24:1037–1040. https://doi.org/10.1023/a:1015610018103
Ayed L, Chaieb K, Cheref A, Bakhrouf A (2008) Biodegradation of triphenylmethane dye malachite green by Sphingomonas paucimobilis. World J Microbiol Biotechnol 25:705–711. https://doi.org/10.1007/s11274-008-9941-x
Ayed L, Chaieb K, Cheref A, Bakhrouf A (2010) Biodegradation and decolorization of triphenylmethane dyes by Staphylococcus epidermidis. Desalination 260:137–146. https://doi.org/10.1016/j.desal.2010.04.052
Azmi W, Sani RK, Banerjee UC (1998) Biodegradation of triphenylmethane dyes. Enzym Microb Technol 22:185–191
Bañuelos JA, García-Rodríguez O, El-Ghenymy A, Rodríguez-Valadez FJ, Godínez LA, Brillas E (2016) Advanced oxidation treatment of malachite green dye using a low cost carbon-felt air-diffusion cathode. J Environ Chem Eng 4:2066–2075. https://doi.org/10.1016/j.jece.2016.03.012
Casas N, Parella T, Vicent T, Caminal G, Sarra M (2009) Metabolites from the biodegradation of triphenylmethane dyes by Trametes versicolor or laccase. Chemosphere 75:1344–1349. https://doi.org/10.1016/j.chemosphere.2009.02.029
Culp SJ, Beland FA (1996) Malachite green: a toxicological review. J Am Coll Toxicol 15:219–238
Daneshvar N, Ayazloo M, Khataee AR, Pourhassan M (2007) Biological decolorization of dye solution containing malachite green by microalgae Cosmarium sp. Bioresour Technol 98:1176–1182. https://doi.org/10.1016/j.biortech.2006.05.025
Du LN, Li G, Zhao YH, Xu HK, Wang Y, Zhou Y, Wang L (2015) Efficient metabolism of the azo dye methyl orange by Aeromonas sp. strain DH-6: characteristics and partial mechanism. Int Biodeterior Biodegrad 105:66–72. https://doi.org/10.1016/j.ibiod.2015.08.019
Du LN, Zhao M, Li G, Zhao XP, Zhao YH (2012) Highly efficient decolorization of malachite green by a novel Micrococcus sp. strain BD15. Environ Sci Pollut Res Int 19:2898–2907. https://doi.org/10.1007/s11356-012-0796-1
Dutta V, Elhanafi D, Osborne J, Martinez MR, Kathariou S (2014) Genetic characterization of plasmid-associated triphenylmethane reductase in Listeria monocytogenes. Appl Environ Microbiol 80:5379–5385. https://doi.org/10.1128/aem.01398-14
Eapen S, Singh S, D'Souza SF (2007) Advances in development of transgenic plants for remediation of xenobiotic pollutants. Biotechnol Adv 25:442–451. https://doi.org/10.1016/j.biotechadv.2007.05.001
Finn RD, Coggill P, Eberhardt RY, Eddy SR, Mistry J, Mitchell AL, Potter SC, Punta M, Qureshi M, Sangrador-Vegas A, Salazar GA, Tate J, Bateman A (2016) The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res 44:D279–D285. https://doi.org/10.1093/nar/gkv1344
Fu XY, Zhao W, Xiong AS, Tian YS, Zhu B, Peng RH, Yao QH (2013) Phytoremediation of triphenylmethane dyes by overexpressing a Citrobacter sp. triphenylmethane reductase in transgenic Arabidopsis. Appl Microbiol Biotechnol 97:1799–1806. https://doi.org/10.1007/s00253-012-4106-0
Gao F, Ding H, Shao L, Xu X, Zhao Y (2015) Molecular characterization of a novel thermal stable reductase capable of decoloration of both azo and triphenylmethane dyes. Appl Microbiol Biotechnol 99:255–267. https://doi.org/10.1007/s00253-014-5896-z
Huan M, Lian-Tai L, Cai-Fang Y, Jin-Jin S, Yuan G, Hong Q, Shun-Peng L (2011) Biodegradation of malachite green by strain Pseudomonas sp. K9 and cloning of the tmr2 gene associated with an ISPpu12. World J Microbiol Biotechnol 27:1323–1329. https://doi.org/10.1007/s11274-010-0580-7
Iyer R, Iken B, Damania A (2017) Whole genome of Klebsiella aerogenes PX01 isolated from San Jacinto River sediment west of Baytown, Texas reveals the presence of multiple antibiotic resistance determinants and mobile genetic elements. Genomics Data 14:7–9. https://doi.org/10.1016/j.gdata.2017.07.012
Jang MS, Lee YM, Kim CH, Lee JH, Kang DW, Kim SJ, Lee YC (2005) Triphenylmethane reductase from Citrobacter sp. strain KCTC 18061P: purification, characterization, gene cloning, and overexpression of a functional protein in Escherichia coli. Appl Environ Microbiol 71:7955–7960. https://doi.org/10.1128/aem.71.12.7955-7960.2005
Kagalkar AN, Jadhav MU, Bapat VA, Govindwar SP (2011) Phytodegradation of the triphenylmethane dye malachite green mediated by cell suspension cultures of Blumea malcolmii Hook. Bioresour Technol 102:0312e10318–0312e10318. https://doi.org/10.1016/j.biortech.2011.08.101
Khan R, Bhawana P, Fulekar MH (2013) Microbial decolorization and degradation of synthetic dyes: a review. Rev Environ Sci Biotechnol 12:75–97. https://doi.org/10.1007/s11157-012-9287-6
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Li L, Peng AH, Lin ZZ, Zhong HP, Chen XM, Huang ZY (2017) Biomimetic ELISA detection of malachite green based on molecularly imprinted polymer film. Food Chem 229:403–408. https://doi.org/10.1016/j.foodchem.2017.02.090
Li LT, Hong Q, Yan X, Fang GH, Ali SW, Li SP (2009) Isolation of a malachite green-degrading Pseudomonas sp. MDB-1 strain and cloning of the tmr2 gene. Biodegradation 20:769–776. https://doi.org/10.1007/s10532-009-9265-z
Liu Q, Liu R, Ma Y, Song J (2018) Physiological and molecular evidence for Na+ and Cl− exclusion in the roots of two Suaeda salsa populations. Aquat Bot 146:1–7. https://doi.org/10.1016/j.aquabot.2018.01.001
Liu W, Hou J, Wang Q, Yang H, Luo Y, Christie P (2015) Collection and analysis of root exudates of Festuca arundinacea L. and their role in facilitating the phytoremediation of petroleum-contaminated soil. Plant Soil 389:109–119. https://doi.org/10.1007/s11104-014-2345-9
Lorenzo M, Moldes D, Rodriguez Couto S, Sanroman MA (2005) Inhibition of laccase activity from Trametes versicolor by heavy metals and organic compounds. Chemosphere 60:1124–1128. https://doi.org/10.1016/j.chemosphere.2004.12.051
Matpang P, Sriuttha M, Piwpuan N (2017) Effects of malachite green on growth and tissue accumulation in pak choy (Brassica chinensis Tsen & Lee). Agr Nat Resour 51:96–102
Meagher RB (2000) Phytoremediation of toxic elemental and organic pollutants. Curr Opin Plant Biol 3:153–162. https://doi.org/10.1016/S1369-5266(99)00054-0
Mosa KA, Saadoun I, Kumar K, Helmy M, Dhankher OP (2016) Potential biotechnological strategies for the cleanup of heavy metals and metalloids. Front Plant Sci 7:303. https://doi.org/10.3389/fpls.2016.00303
Mukherjee T, Das M (2014) Degradation of malachite green by Enterobacter asburiae strain XJUHX-4TM. Clean-Soil Air Water 42:849–856. https://doi.org/10.1002/clen.201200246
Novotny C, Rawal B, Bhatt M, Patel M, Sasek V, Molitoris HP (2001) Capacity of Irpex lacteus and Pleurotus ostreatus for decolorization of chemically different dyes. J Biotechnol 89:113–122
Parshetti GK, Kalme SD, Saratale GD, Govindwar SP (2006) Biodegradation of malachite green by Kocuria rosea MTCC 1532. Acta Chim Slov 53:492–498
Persson B, Kallberg Y, Oppermann U, Jornvall H (2003) Coenzyme-based functional assignments of short-chain dehydrogenases/reductases (SDRs). Chem Biol Interact 143-144:271–278
Rai MS, Rama Bhat P, Prajna PS, Jayadev K, Venkatakrishna Rao PS (2014) Degradation of malachite green and congo red using Aloe barabadensis mill. Extract Int J Curr Microbiol App Sc 3:330e340
Ren S, Guo J, Zeng G, Sun G (2006) Decolorization of triphenylmethane, azo, and anthraquinone dyes by a newly isolated Aeromonas hydrophila strain. Appl Microbiol Biotechnol 72:1316–1321. https://doi.org/10.1007/s00253-006-0418-2
Rosenblueth M, Martinez L, Silva J, Martinez-Romero E (2004) Klebsiella variicola, a novel species with clinical and plant-associated isolates. Syst Appl Microbiol 27:27–35. https://doi.org/10.1078/0723-2020-00261
Rudd T, Sterritt RM, Lester JN (1983) Mass balance of heavy metal uptake by encapsulated cultures of Klebsiella aerogenes. Microb Ecol 9:261–272. https://doi.org/10.1007/BF02097741
Schlüter A, Krahn I, Kollin F, Bönemann G, Stiens M, Szczepanowski R, Schneiker S, Puhler A (2007) IncP-1-β plasmid pGNB1 isolated from a bacterial community from a wastewater treatment plant mediates decolorization of triphenylmethane dyes. Appl Environ Microbiol 73:6345–6350. https://doi.org/10.1128/aem.01177-07
Shedbalkar U, Dhanve R, Jadhav J (2008) Biodegradation of triphenylmethane dye cotton blue by Penicillium ochrochloron MTCC 517. J Hazard Mater 157:472–479. https://doi.org/10.1016/j.jhazmat.2008.01.023
Shin SH, Kim S, Kim JY, Lee S, Um Y, Oh MK, Kim YR, Lee J, Yang KS (2012) Complete genome sequence of Enterobacter aerogenes KCTC 2190. J Bacteriol 194:2373–2374. https://doi.org/10.1128/JB.00028-12
Song J, Wang B (2015) Using euhalophytes to understand salt tolerance and to develop saline agriculture: Suaeda salsa as a promising model. Ann Bot.115:541-553. https://doi.org/10.1093/aob/mcu194
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Wang J, Gao F, Liu Z, Qiao M, Niu X, Zhang KQ, Huang X (2012) Pathway and molecular mechanisms for malachite green biodegradation in Exiguobacterium sp. MG2. PLoS One 7:e51808. https://doi.org/10.1371/journal.pone.0051808
Wei CY, Lin L, Luo LJ, Xing YX, Hu CJ, Yang LT, Li YR, An Q (2014) Endophytic nitrogen-fixing Klebsiella variicola strain DX120E promotes sugarcane growth. Biol Fertil Soils 50:657–666. https://doi.org/10.1007/s00374-013-0878-3
Wu HF, Liu XL, Zhao JM, Yu JB (2012) Toxicological responses in halophyte Suaeda salsa to mercury under environmentally relevant salinity. Ecotox Environ Safe 85:64–71. https://doi.org/10.1016/j.ecoenv.2012.03.016
Wu L, Lin ZZ, Zhong HP, Chen XM, Huang ZY (2017) Rapid determination of malachite green in water and fish using a fluorescent probe based on CdTe quantum dots coated with molecularly imprinted polymer. Sensors Actuators B Chem 239:69–75. https://doi.org/10.1016/j.snb.2016.07.166
Wu Y, Xiao X, Xu C, Cao D, Du D (2013) Decolorization and detoxification of a sulfonated triphenylmethane dye aniline blue by Shewanella oneidensis MR-1 under anaerobic conditions. Appl Microbiol Biotechnol 97:439–7446. https://doi.org/10.1007/s00253-012-4476-3
Yang X, Zheng J, Lu Y, Jia R (2016) Degradation and detoxification of the triphenylmethane dye malachite green catalyzed by crude manganese peroxidase from Irpex lacteus F17. Environ Sci Pollut Res Int 23:9585–9597. https://doi.org/10.1007/s11356-016-6164-9
Zeng J, Eckenrode Heather M, Dounce Susan M, Dai HL (2013) Time-resolved molecular transport across living cell membranes. Biophys J 104:139–145. https://doi.org/10.1016/j.bpj.2012.11.3814
Zhou Q (2001) Chemical pollution and transport of organic dyes in water-soil-crop systems of the Chinese Coast. Bull Environ Contam Toxicol 66:784–793. https://doi.org/10.1007/s00128-001-0077-z
Zhou X, Zhang J, Pan Z, Li D (2018) Review of methods for the detection and determination of malachite green and leuco-malachite green in aquaculture. Crit Rev Anal Chem 14:1–20. https://doi.org/10.1080/10408347.2018.1456314
Funding
This study was funded by National Natural Science Foundation of China (Grant No. 31640002), the Natural Science Foundation of Shandong Province (Grant No. ZR2015JL013), the China Postdoctoral Science Foundation (Grant No. 2016 M600551), and the International Postdoctoral Exchange Fellowship, China (Grant No. 20170058).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Research involving human participants and/or animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 399 kb)
Rights and permissions
About this article
Cite this article
Shang, N., Ding, M., Dai, M. et al. Biodegradation of malachite green by an endophytic bacterium Klebsiella aerogenes S27 involving a novel oxidoreductase. Appl Microbiol Biotechnol 103, 2141–2153 (2019). https://doi.org/10.1007/s00253-018-09583-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-09583-0