Abstract
Clostridium tyrobutyricum is a promising organism for butyrate and n-butanol production, but cannot grow on sucrose. Three genes (scrA, scrB, and scrK) involved in the sucrose catabolic pathway, along with an aldehyde/alcohol dehydrogenase gene, were cloned from Clostridium acetobutylicum and introduced into C. tyrobutyricum (Δack) with acetate kinase knockout. In batch fermentation, the engineered strain Ct(Δack)-pscrBAK produced 14.8–18.8 g/L butanol, with a high butanol/total solvent ratio of ∼0.94 (w/w), from sucrose and sugarcane juice. Moreover, stable high butanol production with a high butanol yield of 0.25 g/g and productivity of 0.28 g/L∙h was obtained in batch fermentation without using antibiotics for selection pressure, suggesting that Ct(Δack)-pscrBAK is genetically stable. Furthermore, sucrose utilization by Ct(Δack)-pscrBAK was not inhibited by glucose, which would usually cause carbon catabolite repression on solventogenic clostridia. Ct(Δack)-pscrBAK is thus advantageous for use in biobutanol production from sugarcane juice and other sucrose-rich feedstocks.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Butanol is a promising advanced biofuel with superior fuel properties, including a 30% higher energy content and lower hygroscopicity and volatility compared to ethanol (Jang et al. 2012; Zhao et al. 2013). Currently, butanol is mainly manufactured via petrochemical processes. Butanol can also be produced from renewable carbon sources in acetone–butanol–ethanol (ABE) fermentation by solventogenic clostridia, including Clostridium acetobutylicum and Clostridium beijerinckii (Wang et al. 2014; Xue et al. 2013; Zheng et al. 2015). However, conventional ABE fermentation suffers from low butanol titer, yield, and productivity because of butanol cytotoxicity and the coproduction of acetone (Nicolaou et al. 2010; Zhao et al. 2013). Consequently, biobutanol with a high production cost cannot compete with petroleum-based butanol (Green 2011). To improve ABE fermentation, there have been extensive research efforts in metabolic and process engineering of solventogenic clostridia (Cho et al. 2015; Du et al. 2015; Lee et al. 2008; Lütke-Eversloh 2014; Papoutsakis 2008; Wang et al. 2014; Xu et al. 2015; Zheng et al. 2015). In addition, as the conventional ABE fermentation feedstock, mainly corn and cassava, usually accounts for more than 50% of the product cost (Green 2011), alternative low-cost feedstocks including lignocellulosic biomass (Jang et al. 2012; Yang et al. 2015) and sugarcane juice and molasses (Bellido et al. 2015; Jiang et al. 2014; Mariano et al. 2013; Ni et al. 2012) have been a focus of recent studies of ABE fermentation.
The goal of this study was to engineer Clostridium tyrobutyricum, a Gram-positive anaerobe naturally producing butyric and acetic acids as two main products (Zhu and Yang 2003), for n-butanol production from sucrose present in sugarcane juice, which is abundant and inexpensive in South China and Brazil. Previously, C. tyrobutyricum with acetate kinase (ack) knockout (Liu et al. 2006) was engineered to overexpress aldehyde/alcohol dehydrogenase gene (adhE2) encoding a bifunctional aldehyde/alcohol dehydrogenase (Yu et al. 2011, 2012), and the engineered strain, Ct(Δack)-pM2, produced n-butanol as the main product, and some ethanol but no acetone, from glucose (Du et al. 2015). However, C. tyrobutyricum is unable to utilize sucrose or ferment sucrose-rich substrates, such as sugarcane and sweet sorghum juices and beet molasses.
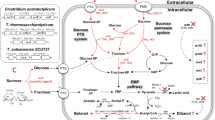
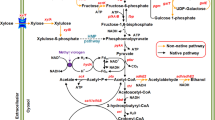
In bacteria, sucrose is transported into cells as sucrose 6-phosphate via sucrose-specific phosphoenolpyruvate (PEP)-dependent phosphotransferase system (PTS); sucrose 6-phosphate hydrolase (or sucrase) then cleaves sucrose 6-phosphate to glucose 6-phosphate and fructose, which is phosphorylated to fructose-6-phosphate by fructokinase; and finally, glucose-6-phosphate and fructose-6-phosphate enter the Embden–Meyerhof–Parnas (EMP) pathway (see Fig. 1). In C. acetobutylicum, three genes (scrA, scrK, and scrB), encoding for the proteins involved in the sucrose catabolic pathway, are present in the scr operon (Tangney and Mitchell 2000). However, these genes are not found in the genome of C. tyrobutyricum (Bassi et al. 2013; Jiang et al. 2013). Therefore, we cloned scrA, scrK, and scrB from C. acetobutylicum and overexpressed them along with adhE2 in C. tyrobutyricum (Δack). The engineered strain, Ct(Δack)-pscrBAK, was able to use sucrose for growth and n-butanol production. Moreover, the engineered strain can simultaneously use sucrose, glucose, and fructose present in sugarcane juice without the glucose-induced carbon catabolite repression (CCR) observed with C. acetobutylicum and C. beijerinckii. By applying an artificial electron carrier, methyl viologen (MV), Ct(Δack)-pscrBAK gave good butanol production with high yield and productivity from sucrose and sugarcane juice, demonstrating its potential industrial application for n-butanol production from low-cost sugarcane juice and molasses.
Metabolic pathways in Ct(∆ack)-pscrBAK. The pathways for sucrose utilization and butanol and ethanol formation shown in dotted lines are absent in the wild-type Clostridium tyrobutyricum. CoA transferase (cat1) catalyzes the conversion of butyryl-CoA to butyrate by transferring CoA to acetate, instead of using ptb and buk, which are not present in the annotated genome of C. tyrobutyricum ATCC 25755 (Lee et al. 2016). It is noted that adhE2 can also function as adh and bdh, which are present in the genome of wild-type C. tyrobutyricum. Key enzymes and genes in the pathways: scrA sucrose-specific PTS; scrB sucrose-6-phosphate hydrolase or sucrase; scrK fructokinase; pta phosphotransacetylase; ack acetate kinase; thl thiolase; hbd β-hydroxybutyryl-CoA dehydrogenase; crt crotonase; bcd butyryl-CoA dehydrogenase; etf electron transferring flavoprotein; ptb phosphotransbutyrylase; buk butyrate kinase; cat1 butyryl-CoA/acetate CoA transferase; adh alcohol dehydrogenase; bdh butanol dehydrogenase; adhE2 aldehyde–alcohol dehydrogenase
Materials and methods
Bacterial strains and culture media
All bacterial strains, recombinant plasmids, and PCR primers used in this study are listed in Table 1. Ct(Δack), a mutant strain of C. tyrobutyricum ATCC 25755 with ack knockout (Liu et al. 2006), was used as the host for metabolic engineering. C. acetobutylicum ATCC 824 (Cac 824) was used to extract genes from its genome. All Clostridium strains were cultured in clostridial growth medium (CGM) and on CGM agar plates supplemented with glucose or sucrose as carbon source at 37 °C under strict anaerobic conditions. The CGM contained (g/L): 1 K2HPO4·3H2O, 0.5 KH2PO4, 2 (NH4)2SO4, 0.1 MgSO4·7H2O, 2 yeast extract, 4 tryptone, and trace minerals (Zhu and Yang 2003). Escherichia coli DH5α (Invitrogen, Carlsbad, CA) used in the preparation of recombinant plasmids and E. coli CA434 (Williams et al. 1990) as the donor strain in conjugation were cultured in liquid Luria–Bertani (LB) and on LB agar plates at 37 °C. After autoclaving at 121 °C for 30 min, media were supplemented with 25 μg/mL chloramphenicol, 30–45 μg/mL thiamphenicol, or 250 μg/mL cycloserine as needed.
DNA isolation and plasmid construction
Two recombinant plasmids expressing Cac 824 scr operon genes were constructed using pMTL82151-adhE2 (pM2), which contained the native C. tyrobutyricum thiolase (thl) promoter driving the constitutive expression of Cac 824 adhE2 gene (Yu et al. 2011). The first plasmid pscrTAKB was constructed by inserting the entire scr operon, including its promoter (P), amplified from the Cac 824 genomic DNA by PCR using the primers shown in Table 1, into pM2 at the SbfI site (see Fig. S1 in supplemental materials) using Clontech In-Fusion Cloning Kit (Mountain View, CA). The second plasmid pscrBAK carrying only scrB (CA_C0425), scrA (CA_C0423), and scrK (CA_C0424) was constructed from pM2 in two steps (see Fig. S1). The scrB gene was first amplified from the Cac 824 genomic DNA by PCR and inserted at the SacII site downstream of the adhE2 gene. The original ribosome binding site of scrB was replaced with a consensus sequence “AGGAGG” to optimize the gene expression. Then, scrA and scrK genes were amplified together by PCR from the Cac 824 genomic DNA and inserted after the scrB gene at the SacI site. Both scrA and scrK genes carried native A/G rich ribosome binding sites. The final constructed plasmids were confirmed by restriction digestion and DNA sequencing.
Construction of C. tyrobutyricum Ct(Δack)-pscrBAK and Ct(Δack)-pscrTAKB
Unless otherwise noted, the constructed plasmids were transformed into Ct(Δack) by conjugation under anaerobic conditions following the previously described method (Yu et al. 2015b). The plasmid was first transformed into E. coli CA434 cells by heat shock under aerobic conditions. The E. coli CA434 cells carrying the plasmid were cultured in LB medium containing 25 μg/mL chloramphenicol and 50 μg/mL kanamycin overnight at 37 °C and 200 rpm to reach an optical density at 600 nm (OD600) of 1.5–2.0. Cells were collected by centrifugation at 4000×g for 2 min, washed once with 1 mL sterile phosphate-buffered saline (PBS, pH 7.0), and then mixed with 200 μL of Ct(Δack) cells pre-cultured at 37 °C overnight. The mixture was pipetted onto CGM agar plates supplemented with 20 g/L glucose and incubated anaerobically at 37 °C for 8–24 h for mating. The cells were then collected and resuspended in 1 mL PBS and spread onto CGM agar plates containing 45 μg/mL thiamphenicol and 250 μg/mL cycloserine for 2–3 days or until colonies were apparent. Positive transformants were confirmed by colony PCR screening and plasmid extraction. The selected transformants were stored at −80 °C.
RT-PCR analysis of gene expression
RT-PCR was performed to confirm scrA, scrB, scrK, and adhE2 expressions in Ct(Δack)-pscrBAK. For RNA isolation and purification, cells were cultured in CGM supplemented with 30 μg/mL thiamphenicol for 2 days. RNAs were then extracted and purified from the collected cells using an RNeasy Mini Kit (Qiagen, cat. no. 74104). RNA quality and quantity were measured by using NanoDrop ND Spectrophotometer (NanoDrop Technologies, Wilmington, DE). Reverse transcription reactions were performed using SuperScript III One-Step RT-PCR System with Platinum Taq DNA Polymerase (Invitrogen, cat. no. 12574-018), and the primers shown in Table 1 under the following conditions: 30 min at 55 °C for cDNA synthesis, 2 min at 94 °C as a hot start, 35 thermal cycles (15 s at 95 °C for denaturation, 1 min at 55 °C for annealing, 1 min at 68 °C for extension), and 5 min at 68 °C for final extension. A minus RT negative control was performed to ensure no trace of DNA existed in all RNA samples (data not shown). RNAs extracted from Ct(Δack) cells were used as negative control, and RNAs extracted from Cac 824 cells were used as positive control. RT-PCR results were analyzed on a 2.5% agarose gel.
Fermentation kinetics
Unless otherwise noted, batch fermentations were carried out in 1-L stirred-tank bioreactors, each containing 600 mL CGM with 120 g/L sucrose or 218 g/L concentrated sugarcane juice, which was obtained from a sugarcane mill in Brazil and contained 44% (w/w) sucrose, 6% (w/w) glucose, and 5% (w/w) fructose at 37 °C. After autoclaving at 121 °C for 30 min, the bioreactor was sparged with nitrogen for ∼30 min to reach anaerobiosis and then inoculated with an overnight culture at a volume ratio of 5%. Unless otherwise noted, thiamphenicol was added (final concentration 30 μg/mL) at the time of inoculation to ensure plasmid stability, and medium pH was controlled at 6.0 by adding 40% ammonium hydroxide throughout the fermentation. Methyl viologen as an artificial electron carrier was also added to a final concentration of 350 μM in the CGM to study its effect on enhancing butanol production. In addition, glucose-induced CCR was also studied with media containing 30 g/L sucrose and 30 g/L glucose in serum bottles. Samples were taken twice a day to monitor cell growth, substrate consumption and butanol, butyrate, acetate, and ethanol production.
Segregational stability
Segregational stability of the plasmid pscrBAK in Ct(Δack)-pscrBAK was evaluated in a series of subcultures in CGM containing glucose or sucrose without antibiotics following the method described previously (Yu et al. 2012). Cells from a colony on the CGM agar plate were picked and cultured in 1 mL CGM supplemented with 20 g/L glucose and 30 μg/mL thiamphenicol at 37 °C for 24 h. To start the segregational stability evaluation, cells were collected and washed with PBS and then inoculated at 1% (v/v) in 1 mL CGM containing glucose or sucrose (without antibiotics) every 24 for 96 h. After dilution by 104-fold with PBS, cells were plated on CGM plates (20 g/L glucose; with or without antibiotics) and incubated at 37 °C for 5 days to determine the number of colony-forming units (CFU). The segregational stability P was estimated using the formula: P = \( \sqrt[\ n]{R} \), where R is the fraction of cells carrying the plasmid and n is the number of generation in 96 h (Yu et al. 2015c), which was ∼21 based on the specific growth rate of ∼0.15/h (generation time = 4.62 h) in the absence of antibiotic observed in this study.
Analytical methods
Cell density was analyzed by measuring optical density (OD) at 600 nm using a spectrophotometer (Shimadzu, Columbia, MD, UV-16-1). The specific growth rate was calculated based on the OD data in the exponential growth phase in batch fermentation. Sucrose, glucose, and fructose were determined by high-performance liquid chromatography (HPLC) with a refractive index detector (Shimadzu RID-10A) and a carbohydrate analysis column (CARBOSep CHO-882, Transgenomic) operated at 80 °C with water as the eluant at 0.5 mL/min. Butanol, butyrate, acetate, and ethanol were determined with a gas chromatograph (GC) (GC-2014 Shimadzu Columbia, MD) equipped with a flame ionization detector (FID) and a 30-m fused silica column (Stabilwax-DA, 0.25 mm film thickness and 0.25 mm ID, Restek, Bellefonte, PA), operated initially at 80 °C for 3 min, increased linearly to 150 at 30 °C per min, and held at 150 °C for 3.7 min. Fermentation samples were diluted 20-fold with a phosphoric acid (1%) solution containing isobutanol (0.5 g/L) and isobutyric acid (0.1 g/L) as internal standards and injected (1 μL each) with an autoinjector (AOC-20i, Shimadzu). Both injector and detector were at 250 °C.
Statistical analysis
All fermentations were repeated at least once and representative data with averages and standard errors are reported. Student’s t test analysis was performed with the significance level of P < 0.05.
Results
Reverse transcription analysis
The presence of messenger RNAs (mRNAs) for scrA, scrB, scrK, and adhE2 was analyzed by using One-Step RT-PCR (see Fig. S2 for agarose gel results in supplemental materials). As expected, the mRNAs of the four genes were undetectable in Ct(Δack) total RNA. In contrast, they were detected in Cac 824 (positive control) and Ct(Δack)-pscrBAK, confirming that these four genes on the plasmid pscrBAK were transcribed in Ct(Δack)-pscrBAK.
Butanol production from sucrose and sugarcane juice
Figure 2 shows typical batch fermentation profiles for Ct(Δack)-pscrBAK with sucrose and sugarcane juice as substrate. As expected, Ct(Δack)-pscrBAK was able to grow on sucrose after a short lag phase of ∼10 h. About 27.2 g/L butyrate, 16.9 g/L acetate, 6.7 g/L butanol, and 1.0 g/L ethanol were produced from ∼104 g/L sucrose consumed in ∼60 h (Fig. 2a). Similarly, about 17.9 g/L butyrate, 16.2 g/L acetate, 8.7 g/L butanol, and 1.0 g/L ethanol were produced from sugarcane juice containing 109 g/L sucrose, 9.0 g/L glucose, and 11.6 g/L fructose (Fig. 2b). The results confirmed that the heterologous sucrose PTS was functional in C. tyrobutyricum, facilitating the utilization of sucrose as a carbon source. Furthermore, all three sugars in sugarcane juice were simultaneously consumed by Ct(Δack)-pscrBAK, indicating that sucrose metabolism was not repressed or inhibited by glucose or fructose, which was further verified with fermentation using medium containing both sucrose and glucose.
Batch fermentation of C. tyrobutyricum Ct(Δack)-pscrBAK with sucrose (a) and sugarcane juice (b) as substrate and 30 μg/mL thiamphenicol. No growth or sucrose consumption was observed with the wild-type C. tyrobutyricum in similar fermentation because of a lack of the sucrose catabolic pathway genes
Although Ct(Δack)-pscrBAK can use all three sugars in sugarcane juice, a slightly longer lag phase and slower growth were observed for fermentation with sugarcane juice as compared to the fermentation with sucrose in CGM. Nevertheless, the specific growth rate in sugarcane juice (0.11 ± 0.01/h) was comparable to that for sucrose (0.12 ± 0.01/h) (Table 2). Butyric acid production was significantly lower while butanol production was higher with sugarcane juice than with sucrose in CGM. Butanol yields and productivities were 0.08 ± 0.01 g/g and 0.12 ± 0.01 g/L/h, respectively, from sugarcane juice, and 0.06 ± 0.01 g/g and 0.09 ± 0.01 g/L/h, respectively, from sucrose (Table 2). The higher butanol production in sugarcane juice could be attributed to the lower butyrate production as both butanol and butyrate had the same precursor, butyryl-CoA (see Fig. 1).
Glucose-induced catabolite repression
The glucose present in sugarcane juice did not inhibit sucrose utilization by Ct(Δack)-pscrBAK. However, glucose as the preferred carbon source would usually inhibit the utilization of other carbon sources like xylose (Fu et al. 2016; Wei et al. 2016) and sucrose (Reid et al. 1999; Sabri et al. 2013), which is known as glucose-mediated CCR (Görke and Stülke 2008; Yao and Shimizu 2013). The effect of glucose on sucrose uptake by cells was thus studied for Ct(Δack)-pscrBAK, C. acetobutylicum JB200, and C. beijerinckii BA101 in serum bottles with CGM or P2 medium (Xue et al. 2012) containing ∼30 g/L sucrose and ∼30 g/L glucose as carbon source and 40 g/L CaCO3 for pH buffering. As can be seen in Fig. 3, sucrose and glucose were utilized simultaneously by Ct(Δack)-pscrBAK (Fig. 3a). The sucrose uptake rate was 0.79 g/L/h, which was even higher than the glucose uptake rate of ∼0.54 g/L/h. In contrast, only 34% of sucrose in the medium was consumed by C. beijerinckii BA101 with a much lower sucrose uptake rate of ∼0.17 g/L/h before glucose was totally consumed (Fig. 3b). Moreover, sucrose in the medium was almost not consumed in the presence of glucose by C. acetobutylicum JB200 (Fig. 3c), which is consistent with the results from a previous study with C. acetobutylicum ATCC 824 (Tangney and Mitchell 2000).
Effects of MV on fermentation
Adding an artificial electron carrier, such as MV, in the fermentation medium can increase NADH availability and thus increase butanol production (Du et al. 2015; Yu et al. 2015a). The effect of MV at 350 μM on butanol production by Ct(Δack)-pscrBAK was thus studied, and the results are shown in Fig. 4. As expected, butanol production from sucrose was significantly improved to reach a higher titer of 18.8 g/L (vs. 6.7 g/L), with a higher butanol yield of 0.21 ± 0.01 g/g (vs. 0.06 ± 0.01 g/g) and productivity of 0.20 ± 0.01 g/L/h (vs. 0.09 ± 0.01 g/L/h), as compared to the fermentation without MV (see Table 2). Meanwhile, acid production was reduced by 79–92%, with only 5.8 g/L butyrate and 1.4 g/L acetate produced, while ethanol production was not significantly affected by the addition of MV (Fig. 4a). Consequently, the ratio of butanol to total products (w/w) increased to 0.68 (from 0.13), a clear indication of metabolic flux shift from acid biosynthesis to butanol biosynthesis in the presence of MV. Similarly, MV also increased butanol production in the fermentation with sugarcane juice as substrate (Fig. 4b). The final butanol titer increased to 14.8 g/L (vs. 8.7 g/L), yield increased to 0.21 ± 0.01 g/g sugar (vs. 0.08 ± 0.01 g/g), and productivity increased to 0.15 ± 0.01 g/L/h (vs. 0.12 ± 0.01 g/L/h), while acid production decreased 49% for butyrate and 91% for acetate, compared to the fermentation without MV (see Table 2). Efficient co-utilization of glucose, fructose, and sucrose was also observed in the fermentation with MV, suggesting that MV did not impose any adverse effect on sugar uptake. However, MV did significantly inhibit cell growth, resulting in a lower cell density and specific growth rate (0.08–0.09 vs. 0.11–0.12/h) (see Table 2). Also, the amount of sucrose utilized in the fermentation with MV was reduced due to increased butanol production, which inhibited the fermentation. For complete sucrose utilization, in situ butanol separation with gas stripping can be used to alleviate butanol toxicity and facilitate higher butanol production (Du et al. 2015).
Fermentation without antibiotics
In the fermentation kinetics studies, thiamphenicol was added in the medium to ensure that the plasmid pscrBAK responsible for sucrose catabolism would be maintained in cells. However, the addition of antibiotics would increase not only the cost of fermentation but also the metabolic burden on cells. Since wild-type C. tyrobutyricum is unable to use sucrose as carbon source, only cells carrying the pscrBAK plasmid can survive in the sucrose medium. Sucrose as the sole carbon source can thus be used as the selection pressure for Ct(Δack)-pscrBAK carrying the heterologous scrBAK genes. To investigate the effects of antibiotics on cell growth and butanol production, batch fermentations were carried out in media with MV but without thiamphenicol; the results are shown in Fig. 5 and compared to those with thiamphenicol (see Table 2). In the sucrose fermentation without the antibiotic, a higher cell density of OD 21.9 was reached in 32 h (vs. 11 in ∼60 h with antibiotic), with a higher specific growth rate of 0.15 ± 0.02/h (vs. 0.08 ± 0.01/h) due to the released metabolic burden. Consequently, the fermentation without antibiotic was faster with a significantly higher butanol productivity (0.28 ± 0.02 vs. 0.20 ± 0.01 g/L/h) and yield (0.25 ± 0.02 vs. 0.21 ± 0.01 g/g). However, the fermentation stopped earlier at a lower butanol titer of 16.1 g/L (vs. 18.8 g/L) without consuming as much sucrose in the medium, in part due to higher acid production (10.2 vs. 5.8 g/L and 2.5 vs. 1.4 g/L for butyrate and acetate, respectively), which, together with butanol, inhibited the fermentation. Similarly, the fermentation with sugarcane juice without antibiotic also gave a higher specific growth rate (0.11 ± 0.02 vs. 0.09 ± 0.01/h) and butanol productivity (0.21 ± 0.01 vs. 0.15 ± 0.01 g/L/h) as compared to the fermentation with antibiotic. However, the higher butyrate (11.2 vs. 9.1 g/L) and acetate (2.6 vs. 1.4 g/L) production resulted in a lower butanol titer (11.9 vs. 14.8 g/L) but had no significant effect on butanol yield from total sugar consumed, which was 0.20–0.21 g/g with or without the antibiotic. The results indicated that Ct(Δack)-pscrBAK was stable for butanol production without the antibiotic, even though glucose and fructose were also present in sugarcane juice.
Segregational stability
The segregational stability of pscrBAK in Ct(Δack)-pscrBAK was further evaluated in sequential cultures for 96 h. The fractions of cells carrying the plasmid were 91.7 and 91.9% after 96 h in glucose and sucrose medium, respectively. The corresponding probabilities for plasmid loss per generation or P values were 99.59 and 99.60%, respectively, comparable to that of PMTL82151 reported in a previous study (Yu et al. 2012). The high P value of 99.6% in both glucose and sucrose media indicated that the plasmid pscrBAK was stable even in the absence of a selection pressure.
Discussion
C. tyrobutyricum naturally does not have the genes encoding for enzymes involved in the sucrose catabolic pathway. In C. acetobutylicum and C. beijerinckii, sucrose transport and catabolism involves three genes: scrA-encoding enzyme II of the sucrose PTS, scrB-encoding sucrose-6-phosphate hydrolase or sucrase, and scrK-encoding fructokinase (Tangney and Mitchell 2000). Sucrose metabolism is modulated by a regulator repressor protein ScrR in C. beijerinckii (Reid et al. 1999) and an antiterminator protein ScrT encoded by scrT, which is preceded by a transcription terminator, in C. acetobutylicum, respectively (Tangney and Mitchell 2000). These genes are located in the same operon with the gene order of scrTAKB in C. acetobutylicum and scrARBK in C. beijerinckii. Transcription of the scr operon in C. acetobutylicum is controlled by a complex termination and antitermination system involving the formation of a terminating structure that prevents the continuation of transcription and the interaction between an antiterminator and a specific mRNA region to relieve the termination caused by the terminator (Tangney and Mitchell 2000). The expression of scr is also subject to the catabolite control protein A (CcpA)-dependent repression system, in which dimerized CcpA, triggered by the intense glycolytic activity, recognizes catabolite responsive elements (CRE) present in the promoter region of scr operon (Tangney and Mitchell 2000), thereby blocking either the recruitment of RNA polymerases or the elongation of RNA synthesis (Görke and Stülke 2008). In addition, PTS-dependent phosphorylation of antiterminators by histidine protein (HPr) kinase is a common mechanism used by bacteria to ensure that operon expression occurs only in the presence of the substrate (e.g., sucrose) and absence of a more favorable carbon source (i.e., glucose) (Görke and Stülke 2008; Stülke et al. 1998).
In this study, we first introduced the plasmid carrying the whole C. acetobutylicum scr operon, including the antiterminator gene (scrT), into C. tyrobutyricum, but the transformant failed to grow on sucrose as the sole carbon source, suggesting that the genes in the scr operon were not expressed, probably because the complex regulatory system was not functional in the heterologous host cells. We then expressed scrB, scrA, and scrK from C. acetobutylicum, along with adhE2 under the native constitutive thiolase promoter to circumvent the CCR directed by the ρ-independent transcription termination and CcpA-dependent repression system, and the mutant Ct(Δack)-pscrBAK was able to use sucrose to produce butanol, along with butyrate and acetate, without being subject to glucose-induced CCR when using sugarcane juice containing sucrose, glucose, and fructose as substrates (Fig. 2b). When fermentations were carried out in serum bottles using sucrose and glucose as co-substrates, sucrose was uptaken by Ct(Δack)-pscrBAK at a much higher rate compared to C. beijerinckii BA101 and C. acetobutylicum JB200 (Fig. 3). Moreover, no sucrose PTS activity was found in C. acetobutylicum cultured on glucose, and sucrose utilization was repressed or inhibited in the presence of glucose (Tangney and Mitchell 2000). In fact, glucose has been widely reported as the preferred substrate over other sugars (e.g., xylose) for Clostridium, including C. tyrobutyricum (Fu et al. 2016; Yu et al. 2015a, 2015c), C. acetobutylicum (Chen et al. 2013; Servinsky et al. 2010), and C. beijerinckii (Jiang et al. 2014; Xiao et al. 2012). No CCR was observed with C. tyrobutyricum overexpressing scrB, scrA, and scrK from C. acetobutylicum, probably because these genes were constitutively expressed under the control of the thiolase promoter and scrT encoding the antiterminator protein ScrT was not introduced. Compared to solventogenic clostridia, Ct(Δack)-pscrBAK not being subject to glucose catabolite repression would be an advantageous host for butanol production from sugarcane juice.
However, unlike C. acetobutylicum, C. tyrobutyricum does not have CoA transferase and is thus unable to reassimilate acids (butyric and acetic) for alcohols (butanol and ethanol) biosynthesis (Grupe and Gottschalk 1992), resulting in overaccumulation of acids and low butanol production (Yu et al. 2015b). To solve this problem, one approach is to overexpress C. acetobutylicum ctfAB to facilitate reassimilation of acetate and butyrate to acetyl-CoA and butyryl-CoA, respectively, thus leading to more ethanol and butanol production (Yu et al. 2015b). However, the expression of ctfAB also resulted in acetone production from acetoacetate through a non-enzymatic decarboxylation. On the other hand, MV can act as a competitive inhibitor of hydrogenase and an artificial electron carrier directing the electron towards NADH generation, instead of hydrogen production, and thus its presence in the fermentation medium favored butanol biosynthesis due to increased NADH availability in cells (Du et al. 2015; Rao and Mutharasan 1987; Yu et al. 2015a). As demonstrated in this study, the addition of MV resulted in higher butanol titer and yield, making C. tyrobutyricum a more attractive host for biobutanol production. It is also possible to increase the internal NADH pool by inhibiting hydrogenase with CO or H2 or by knocking down hydrogenase via genetic manipulations, thus without using MV or an artificial electron carrier. Overexpressing formate dehydrogenase could also increase NADH availability and butanol production (Ma et al. 2016). These methods to increase NADH availability for enhanced butanol biosynthesis may be explored in future studies.
Strains containing heterologous plasmids are usually not genetically stable and may lose their desirable phenotypes without applying a selection pressure during fermentation, making them unsuitable for industrial applications. However, the segregational stability of the plasmid of the mutant in sucrose and glucose media without antibiotic was high (P = 99.6%). Furthermore, as its growth on sucrose depends on carrying the plasmid pscrBAK, Ct(Δack)-pscrBAK was stable in maintaining its phenotype without antibiotic and produced butanol at a high level (11.9–16.1 g/L) comparable to or even higher than those of other solventogenic clostridia, including C. acetobutylicum JB200 (14.9–18.5 g/L) (Jiang et al. 2014), C. beijerinckii NCP P260 (14.2 g/L) (Ni et al. 2012), and C. saccharobutylicum (11.9 g/L) (Shaheen et al. 2000) in fermentation with sucrose, sugarcane juice, or cane molasses (see Table 3). It should be noted that the butanol titer of 18.8 g/L obtained in the presence of MV and antibiotics was higher than most reported to date for butanol fermentation from sucrose and sucrose based substrates. Butanol titer was also higher (14.8 vs. 11.0 g/L), but with a slightly lower yield (0.21 vs. 0.28 g/g) using sugarcane juice as carbon source when compared to Ct(Δack)-adhE2 using glucose as substrate (see Table 3) (Du et al. 2015). The high butanol production and genetic stability of the engineered strain make it a good host for n-butanol production from sucrose, sugarcane juice, and other sucrose-rich feedstocks. It should also be mentioned that Ct(Δack)-pscrBAK produced n-butanol as the main product, with only a small amount of ethanol and no acetone, which would be much easier for butanol to be separated and purified in downstream processing, offering another advantage over conventional ABE fermentation by C. acetobutylicum and C. beijerinckii (Du et al. 2015).
In summary, three genes scrB, scrA, and scrK in the sucrose operon, along with adhE2, of C. acetobutylicum ATCC 824 were cloned and overexpressed in Ct(Δack) for butanol production from sucrose and sugarcane juice. The mutant produced n-butanol from sucrose and sugarcane juice at a high titer with a yield comparable to those of ABE fermentation with solventogenic clostridia. The engineered Ct(Δack)-pscrBAK does not suffer from glucose-induced CCR and produces only n-butanol as the main solvent product with no acetone, making it a good candidate for n-butanol production from sucrose-rich feedstocks.
References
Bassi D, Fontana C, Gazzola S, Pietta E, Puglisi E, Cappa F, Cocconcelli PS (2013) Draft genome sequence of Clostridium tyrobutyricum strain UC7086, isolated from grana padano cheese with late-blowing defect. Genome Announc. 1:e00614–e00613
Bellido C, Infante C, Coca M, Gonzalez-Benito G, Lucas S, Garcia-Cubero MT (2015) Efficient acetone-butanol-ethanol production by Clostridium beijerinckii from sugar beet pulp. Bioresour Technol 190:332–338
Chen Y, Zhou T, Liu D, Li A, Xu SB, Liu QG, Li BB, Ying HJ (2013) Production of butanol from glucose and xylose with immobilized cells of Clostridium acetobutylicum. Biotechnol Bioprocess 18:234–241
Cho C, Jang YS, Moon HG, Lee J, Lee SY (2015) Metabolic engineering of clostridia for the production of chemicals. Biofuels Bioprod Biorefin 9:211–225
Du Y, Jiang W, Yu M, Tang IC, Yang ST (2015) Metabolic process engineering of Clostridium tyrobutyricum ∆ack-adhE2 for enhanced n-butanol production from glucose: effects of methyl viologen on NADH availability, flux distribution, and fermentation kinetics. Biotechnol Bioeng 112:705–715
Fu H, Yu L, Lin M, Wang J, Xiu Z, Yang ST (2016) Metabolic engineering of Clostridium tyrobutyricum for enhanced butyric acid production from glucose and xylose. Metab Eng. doi:10.1016/j.ymben.2016.12.014
Görke B, Stülke J (2008) Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat Rev Microbiol 6:613–624
Green EM (2011) Fermentative production of butanol—the industrial perspective. Curr Opin Biotechnol 22:337–343
Grupe H, Gottschalk G (1992) Physiological events in Clostridium acetobutylicum during the shift from acidogenesis to solventogenesis in continuous culture and presentation of a model for shift induction. Appl Environ Microbiol 58:3896–3902
Jang YS, Malaviya A, Cho C, Lee J, Lee SY (2012) Butanol production from renewable biomass by clostridia. Bioresour Technol 123:653–663
Jiang L, Zhu L, Xu X, Li Y, Li S, Huang H (2013) Genome sequence of Clostridium tyrobutyricum ATCC 25755, a butyric acid-overproducing strain. Genome Announc 1(3):e00308–e00313
Jiang W, Zhao JB, Wang ZQ, Yang ST (2014) Stable high-titer n-butanol production from sucrose and sugarcane juice by Clostridium acetobutylicum JB200 in repeated batch fermentations. Bioresour Technol 163:172–179
Lee J, Jang Y, Han M-J, Kim JY, Lee SY (2016) Deciphering Clostridium tyrobutyricum metabolism based on the whole-genome sequence and proteome analyses. MBio 7(3):e00743–e00716
Lee SY, Park JH, Jang SH, Nielsen LK, Kim J, Jung KS (2008) Fermentative butanol production by Clostridia. Biotechnol Bioeng 101:209–228
Liu X, Zhu Y, Yang ST (2006) Construction and characterization of ack deleted mutant of Clostridium tyrobutyricum for enhanced butyric acid and hydrogen production. Biotechnol Prog 22:1265–1275
Lütke-Eversloh T (2014) Application of new metabolic engineering tools for Clostridium acetobutylicum. Appl Microbiol Biotechnol 98:5823–5837
Ma C, Ou J, Xu N, Fierst J, Yang ST, Liu X (2016) Rebalancing redox to improve biobutanol production by Clostridium tyrobutyricum. Bioengineering 3:2
Mariano AP, Dias MO, Junqueira TL, Cunha MP, Bonomi A, Filho RM (2013) Butanol production in a first-generation Brazilian sugarcane biorefinery: technical aspects and economics of greenfield projects. Bioresour Technol 135:316–323
Ni Y, Wang Y, Sun ZH (2012) Butanol production from cane molasses by Clostridium saccharobutylicum DSM 13864: batch and semicontinuous fermentation. Appl Biochem Biotechnol 166:1896–91907
Nicolaou SA, Gaida SM, Papoutsakis ET (2010) A comparative view of metabolite and substrate stress and tolerance in microbial bioprocessing: from biofuels and chemicals, to biocatalysis and bioremediation. Metab Eng 12:307–331
Papoutsakis ET (2008) Engineering solventogenic clostridia. Curr Opin Biotechnol 19:420–429
Rao G, Mutharasan R (1987) Altered electron flow in continuous cultures of Clostridium acetobutylicum induced by viologen dyes. Appl Environ Microbiol 53:1232–1235
Reid SJ, Rafudeen MS, Leat NG (1999) The genes controlling sucrose utilisation in Clostridium beijerinckii 8052 constitutes an operon. Microbiology 145:1461–1472
Sabri S, Nielsen LK, Ce V (2013) Molecular control of sucrose utilization in Escherichia coli W, an efficient sucrose-utilizing strain. Appl Environ Microbiol 79:478–487
Servinsky MD, Kiel JT, Dupuy NF, Sund CJ (2010) Transcriptional analysis of differential carbohydrate utilization by Clostridium acetobutylicum. Microbiology 156:3478–3491
Shaheen R, Shirley M, Jones DT (2000) Comparative fermentation studies of industrial strains belonging to four species of solvent-producing clostridia. J Mol Microbiol Biotechnol 2:115–124
Stülke J, Arnaud M, Rapoport G, Martin-Verstraete I (1998) PRD—a protein domain involved in PTS-dependent induction and carbon catabolite repression of catabolic operons in bacteria. Mol Microbiol 28:865–874
Tangney M, Mitchell WJ (2000) Analysis of a catabolic operon for sucrose transport and metabolism in Clostridium acetobutylicum ATCC 824. J Mol Microbiol Biotechnol 2:71–80
Wang J, Yang X, Chen CC, Yang ST (2014) Engineering clostridia for butanol production from biorenewable resources: from cells to process integration. Curr Opin Chem Eng 6:43–54
Wei P, Lin M, Wang Z, Fu H, Yang H, Jiang W, Yang ST (2016) Metabolic engineering of Propionibacterium freudenreichii subsp. shermanii for xylose fermentation. Bioresour Technol 219:91–97
Williams DR, Young DI, Young M (1990) Conjugative plasmid transfer from Escherichia coli to Clostridium acetobutylicum. J Gen Microbiol 136:819–826
Xiao H, Li ZL, Jiang Y, Yang YL, Jiang WH, Gu Y, Yang S (2012) Metabolic engineering of D-xylose pathway in Clostridium beijerinckii to optimize solvent production from xylose mother liquid. Metab Eng 14:569–578
Xu M, Zhao J, Yu L, Tang IC, Xue C, Yang ST (2015) Engineering Clostridium acetobutylicum with a histidine kinase knockout for enhanced n-butanol tolerance and production. Appl Microbiol Biotechnol 99:1011–1022
Xue C, Zhao JB, Lu CC, Yang ST, Bai FW, Tang IC (2012) High-titer n-butanol production by Clostridium acetobutylicum JB200 in fed-batch fermentation with intermittent gas stripping. Biotechnol Bioeng 109:2746–2756
Xue C, Zhao JB, Liu F, Lu CC, Yang ST, Bai FW (2013) Two-stage in situ gas stripping for enhanced butanol fermentation and energy-saving product recovery. Bioresour Technol 135:396–402
Yang X, Xu M, Yang ST (2015) Metabolic and process engineering of Clostridium cellulovorans for biofuel production from cellulose. Metab Eng 32:39–48
Yao R, Shimizu K (2013) Recent progress in metabolic engineering for the production of biofuels and biochemicals from renewable sources with particular emphasis on catabolite regulation and its modulation. Process Biochem 48:1409–1417
Yu M, Zhang Y, Tang IC, Yang ST (2011) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production. Metab Eng 13:373–382
Yu M, Du Y, Jiang W, Chang WL, Yang ST, Tang IC (2012) Effects of different replicons in conjugative plasmids on transformation efficiency, plasmid stability, gene expression and n-butanol biosynthesis in Clostridium tyrobutyricum. Appl Microbiol Biotechnol 93:881–889
Yu L, Xu M, Tang IC, Yang ST (2015a) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production through co-utilization of glucose and xylose. Biotechnol Bioeng 112:2134–2141
Yu L, Zhao J, Xu M, Dong J, Varghese S, Yu M, Tang IC, Yang ST (2015b) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production: effects of CoA transferase. Appl Microbiol Biotechnol 99:4917–4930
Yu L, Xu M, Tang IC, Yang ST (2015c) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production from maltose and soluble starch by overexpressing α-glucosidase. Appl Microbiol Biotechnol 99:6155–6165
Zhao J, Lu C, Chen CC, Yang ST (2013) Biological production of butanol and higher alcohols. In: Yang ST, El-Enshasy HA, Thongchul N (eds) Bioprocessing technologies in biorefinery for sustainable production of fuels, chemicals, and polymers, Ch. 13. Wiley, Hoboken, NJ, pp 235–261
Zheng J, Tashiro Y, Wang QH, Sonomoto K (2015) Recent advances to improve fermentative butanol production: genetic engineering and fermentation technology. J Biosci Bioeng 119:1–9
Zhu Y, Yang ST (2003) Adaptation of Clostridium tyrobutyricum for enhanced tolerance to butyric acid in a fibrous-bed bioreactor. Biotechnol Prog 19:365–372
Acknowledgements
This work was supported in part by the National Science Foundation STTR program (IIP-1026648). Financial support from China Scholarship Council to Zhang (201406350172) for research visit at OSU is also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This research does not involve human participants or animals.
Conflict of interest
The authors declare that they have no conflict of interests.
Additional information
Jianzhi Zhang and Le Yu contributed equally to this work.
Electronic supplementary material
ESM 1
(PDF 878 kb).
Rights and permissions
About this article
Cite this article
Zhang, J., Yu, L., Xu, M. et al. Metabolic engineering of Clostridium tyrobutyricum for n-butanol production from sugarcane juice. Appl Microbiol Biotechnol 101, 4327–4337 (2017). https://doi.org/10.1007/s00253-017-8200-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8200-1