Abstract
Acidogenic clostridia naturally producing acetic and butyric acids has attracted high interest as a novel host for butyrate and n-butanol production. Among them, Clostridium tyrobutyricum is a hyper butyrate-producing bacterium, which re-assimilates acetate for butyrate biosynthesis by butyryl-CoA/acetate CoA transferase (CoAT), rather than the phosphotransbutyrylase-butyrate kinase (PTB-BK) pathway widely found in clostridia and other microbial species. To date, C. tyrobutyricum has been engineered to overexpress a heterologous alcohol/aldehyde dehydrogenase, which converts butyryl-CoA to n-butanol. Compared to conventional solventogenic clostridia, which produce acetone, ethanol, and butanol in a biphasic fermentation process, the engineered C. tyrobutyricum with a high metabolic flux toward butyryl-CoA produced n-butanol at a high yield of > 0.30 g/g and titer of > 20 g/L in glucose fermentation. With no acetone production and a high C4/C2 ratio, butanol was the only major fermentation product by the recombinant C. tyrobutyricum, allowing simplified downstream processing for product purification. In this review, novel metabolic engineering strategies to improve n-butanol and butyrate production by C. tyrobutyricum from various substrates, including glucose, xylose, galactose, sucrose, and cellulosic hydrolysates containing the mixture of glucose and xylose, are discussed. Compared to other recombinant hosts such as Clostridium acetobutylicum and Escherichia coli, the engineered C. tyrobutyricum strains with higher butyrate and butanol titers, yields and productivities are the most promising hosts for potential industrial applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
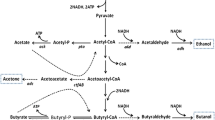
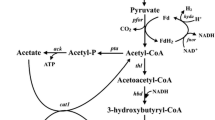
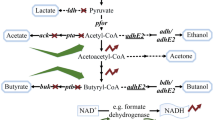
Butyric acid, a short-chain volatile fatty acid with broad applications in the chemical industry, is currently produced primarily by chemical synthesis from petroleum-based feedstocks (Wang et al. 2016; Zigova and Sturdik 2000). However, there is a high demand for biobased butyric acid as a natural ingredient for use in animal feeds, cosmetics, foods, and pharmaceuticals (Dwidar et al. 2012; Jha et al. 2014). Therefore, there is an urgent need in developing bacterial strains for butyric acid production from sugars and renewable feedstocks in fermentation (Jiang et al. 2018). Clostridium tyrobutyricum, a Gram-positive, strictly anaerobic acidogen, produces acetic and butyric acids as the main products from glucose (see Fig. 1). In the dairy industry, C. tyrobutyricum is recognized as the main microbial contaminant affecting cheese quality due to the off flavor from butyrate (D'Incecco et al. 2015; Morandi et al. 2015). On the other hand, C. tyrobutyricum has been considered as the most promising microbial cell factory for butyric acid production because of its high metabolic flux toward butyryl-CoA and high butyric acid tolerance (Jiang et al. 2018; Yang et al. 2013a, b). Compared to other butyric acid producing bacteria including native Clostridium butyricum (Cummins and Johnson 1971; Sushkova et al. 2013; Zigova et al. 1999) and engineered Clostridium acetobutylicum (Jang et al. 2014; Siller et al. 2008) and Escherichia coli (Jawed et al. 2016; Kataoka et al. 2017; Saini et al. 2014), C. tyrobutyricum can produce more butyrate at a higher titer with a higher product yield and purity (Jiang et al. 2018). Figure 1 shows the metabolic pathways involved in butyric acid biosynthesis from glucose and other carbon sources.
Metabolic pathways in engineered C. tyrobutyricum for butyrate and n-butanol production from glucose, xylose, galactose, and sucrose. (Gene name and abbreviation: ack: acetate kinase; adhE2: aldehyde/alcohol dehydrogenase; adh alcohol dehydrogenase; bcd: butyryl-CoA dehydrogenase; buk: butyrate kinase; cat1: butyryl-CoA/acetate CoA transferase; ctfAB: CoA transferase; crt: crotonase; etf: electron transferring flavoprotein; fba: fructose-1,6-diphosphatase; galK: galactokinase; galE: UDP-galactose 4-epimerase; galT: galactose-1-phosphate uridylyltransferase; gpi: phosphoglucose isomerase; hbd: β-hydroxybutyryl-CoA dehydrogenase; hydA: hydrogenase; pta: phosphotransacetylase; ptb: phosphotransbutylase; pfkA: phosphofructokinase; pykA: pyruvate kinase; pgm: phosphoglyceromutase; scrB: sucrose-6-phosphate hydrolase; scrK: fructokinase; scrA: sucrose-specific PTS; thl: thiolase; xylA: xylose isomerase; xylB: xylulokinase; xylT: D-xylose-proton symporter; HMP pathway: hexose monophosphate pathway)
In addition to butyric acid, engineered C. tyrobutyricum has been affirmed as a superior host for n-butanol production, leveraging its high metabolic flux toward butyryl-CoA and tolerance to butanol at high concentrations (> 15 g/L) (Yu et al. 2011). As illustrated in Fig. 1, n-butanol can be produced from butyryl-CoA via a bifunctional aldehyde/alcohol dehydrogenase (adhE2) from C. acetobutylicum. Unlike traditional acetone-butanol-ethanol (ABE) fermentation with solventogenic clostridia such as Clostridium beijerinckii and C. acetobutylicum, butanol fermentation with engineered C. tyrobutyricum overexpressing adhE2 does not produce acetone. In addition, the heterologous n-butanol biosynthesis pathway in C. tyrobutyricum is easier to control as the fermentation does not involve a phase transition from acidogenesis to solventogenesis in the ABE fermentation, which is also suppressed by sporulation under butanol stress (Xu et al. 2017). In addition, industrial ABE fermentation is susceptible to bacteriophage infection (Jones et al. 2000), which has rarely been observed or reported for C. tyrobutyricum (Mayer et al. 2010). Therefore, the engineered C. tyrobutyricum-adhE2 with a simple and un-regulated butanol biosynthesis pathway has outperformed any known microbes in n-butanol production and achieved the highest titer (> 25 g/L) and yield (> 0.3 g/g from glucose) reported so far (Yu et al. 2011; Zhang et al. 2018).
While n-butanol has long been produced from sugars and starch in industrial ABE fermentation, currently butanol is almost exclusively produced via petrochemical routes and used mainly in industrial solvents and in the manufacturing of acrylate esters, amino resins, and butylamines, with a worldwide market of ~ 1.5 billion gallons (4.5 million metric tons) (Zhao et al. 2013). Butanol is also an attractive drop-in biofuel with superior fuel properties (high energy density, lower volatility, etc.). It has lower water miscibility, flammability, and corrosiveness than ethanol and is compatible with existing fuel infrastructures and can directly replace gasoline in engines without modification. Increasing demands for biobutanol as a green solvent in food, pharmaceutical, cosmetic and biofuel industries have prompted the search and development of novel bacterial strains for butanol production from renewable feedstocks (Cheng et al. 2019a; Wang et al. 2014; Xue et al. 2017).
This mini-review provides an overview of recent advances in metabolic engineering of C. tyrobutyricum for butyrate and n-butanol production from sugars and low-cost biomass feedstocks, highlighting strategies and challenges to enhance fermentation efficiency in order to reduce production cost for industrial applications. We also give a brief introduction about C. tyrobutyricum and genetic engineering tools, including replicative plasmids for heterologous gene expression and CRISPR-Cas systems for genome editing, available for rational metabolic engineering of clostridia. We conclude with a brief discussion on perspectives for future research and development.
Genomics and metabolic pathway engineering of C. tyrobutyricum
The genomes of three C. tyrobutyricum strains have been fully sequenced and annotated. For the most widely studied strain ATCC 25755, it has a chromosome of 3,071,606 bp in size, a plasmid of 62,831 bp, and totally 3,220 genes (Lee et al. 2016). The strain CCTCC W428 has a chromosome of 3,011,209 bp, a similar plasmid of 62,833 bp, and totally 3,038 genes (Wu et al. 2017a). The third strain Cirm BIA 2237 has a slightly larger chromosome of 3,159,003 bp but no plasmid and 3,182 genes (Munier et al. 2019). In addition, several draft genome sequences have also been reported for different strains (Bassi et al. 2013; Soggiu et al. 2015; Storari et al. 2015a, b; Wasels et al. 2016). These genome sequences can provide genome-scale information about genes involved in various metabolic and regulatory pathways, which are not fully elucidated yet as many genes have not been annotated or their functions remained unspecified. Nevertheless, the available genomic information has given us a better understanding of the metabolism of C. tyrobutyricum and facilitated rational metabolic engineering for creating mutant strains with superior fermentation ability to produce butyric acid and other chemicals such as n-butanol. For example, a recent genomic analysis has uncovered that C. tyrobutyricum re-assimilates acetic acid for butyric acid biosynthesis through the CoA transferase (CoAT encoded by cat1) pathway, rather than the phosphotransbutyrylase-butyrate kinase (PTB-BK) pathway as in other clostridia including C. butyricum and solventogenic Clostridium species (Lee et al. 2016).
Among the butyric acid producing clostridia, C. tyrobutyricum has the smallest genome and a narrower substrate spectrum, but the highest butyric acid production potential based on the final titer and yield attained in fermentation (see Table 1). Since acetate, a major byproduct in butyrate fermentation, can be re-assimilated to generate butyrate via the CoAT pathway in C. tyrobutyricum, acetate production can be reduced to minimum with butyrate as the main or only fermentation product (Fu et al. 2017b; 2020), which not only increase butyrate yield but also ease downstream processing for product purification. Recent genomic and proteomic studies of C. tyrobutyricum have also suggested that the carbon distribution and energy conservation in C. tyrobutyricum favored the biosynthesis of C4 (butyrate) over C2 (acetate) metabolites (Lee et al. 2016; Ma et al. 2015). Although C. tyrobutyricum is the leading candidate for fermentation production of biobased butyric acid and n-butanol, its prospects for industrial application can be improved by further increasing product yield, productivity, and titer.
Several replicative plasmids with G(+) replicons have been developed for clostridia, including pSOS94 (with pIM13 replicon from Bacillus subtilis), pJIR (with pIP404 replicon from Clostridium perfringens), pMTL007 (with pCB102 replicon from C. butyricum), and pMTL80000 series with replicons pIM13, pCB102, pCD6 (from Clostridium difficile), and pBP1 (from Clostridium botulinum), respectively (Heap et al. 2007, 2009). These plasmids have been successfully used for gene expression in various Clostridium species including C. tyrobutyricum (Yu et al. 2011; 2012). Among them, pMTL82151 with pBP1 replicon gave the highest gene expression, plasmid stability, and transformation efficiency (Yu et al. 2012), which are critical to the development of a stable and robust recombinant strain for industrial fermentation. The PMTL plasmids can also be used with the retargeted Group II intron (ClosTron) for gene knockout on the chromosome (Heap et al. 2010). More recently, an endogenous type I-B CRISPR-Cas system with significantly decreased toxicity was developed for genome editing in C. tyrobutyricum (Zhang et al. 2018). Furthermore, eliminating native plasmid and type-I restriction endonuclease in C. tyrobutyricum increased the transformation efficiency and facilitated genome editing using the CRISPR-Cas9/Cpf1 system which was not applicable in wild-type C. tyrobutyricum (Zhang et al. 2020). The ability to perform efficient genome editing with CRISPR-Cas systems in C. tyrobutyricum is a major breakthrough that can facilitate multiple gene modifications and create stable strains without requiring a selection pressure (such as antibiotic resistance) suitable for use in industrial fermentation.
Empowered with newly available genomics data and genetic engineering tools, scientists constructed various C. tyrobutyricum mutant strains with desirable properties like increased butyrate/butanol titer, yield, productivity, tolerance, and substrate variety and utilization efficiency, which are discussed in the following sections.
Engineering strategies for enhancing butyrate production from glucose
Clostridium tyrobutyricum has been profoundly studied for improving butyrate production from various substrates through rational metabolic engineering strategies (Jiang et al. 2018). Table 2 summarizes metabolic engineering strategies applied to date with notable fermentation performance boosts in final product titer, yield, productivity and purity or selectivity as indicated by the butyric acid to acetic acid ratio (BA/AA). So far most metabolic engineering studies have been focusing on eliminating acetic acid accumulation and overexpressing genes in the butyrate biosynthesis pathway. An earlier attempt to knock out the pta and ack genes in the acetic acid biosynthesis pathway resulted in mutants (PTA-Em and ACK-Em) with a ~ 14% decrease in acetic acid production and ~ 30% higher butyrate production in fermentation (Liu et al. 2006b; Zhu et al. 2005). For butyrate biosynthesis, C. tyrobutyricum uses butyryl-CoA/acetate CoA transferase (cat1) to convert butyryl-CoA to butyrate, instead of the phosphotransacetylase and butyrate kinase (PTB-BUK) pathway commonly utilized in other clostridia such as C. butyricum and C. acetobutylicum (Lee et al. 2016). Overexpressing cat1 and crotonase (crt) in C. tyrobutyricum thus enhanced the flux from acetyl-CoA to butyrate and significantly reduced acetic acid production, which resulted in a 2.24-fold increase in the butyric acid to acetic acid ratio (BA/AA) to 15.76 g/g (Suo et al. 2018a). Meanwhile, overexpressing a [FeFe]-hydrogenase in C. tyrobutyricum increased hydrogen and butyrate production (Jo et al. 2010). Since hydrogen production has significant effects on electron balance and product distribution, adding artificial electron carriers, such as benzyl viologen (BV) and methyl viologen (MV), in the fermentation medium was found to inhibit hydrogen production and shift the metabolism from acetic acid production to reassimilation for butyrate production, which resulted in a high BA/AA of 58 g/g or a product purity of 98.3% in batch fermentation (Choi et al. 2012; Fu et al. 2017b).
To increase butyrate productivity from glucose, 6-phosphofructokinase (pfkA) and pyruvate kinase (pykA) in the EMP pathway were overexpressed individually or simultaneously in C. tyrobutyricum to enhance glucose catabolism (Suo et al. 2018b). Then, genes involved in butyric acid biosynthetic pathway, including thiolase (thl), crotonase (crt), and butyryl-CoA/acetate CoA transferase (cat1), were further investigated for enhancing the butyrate titer and yield (Suo et al. 2018c). Finally, C. tyrobutyricum mutant strain co-expressing crt, cat1, pfkA, and pykA was shown to produce the highest level of butyric acid of 46.8 g/L with a productivity of 0.83 g/L·h and butyrate/acetate ratio of 13.22 g/g in batch fermentation, which were 33.7%, 69.4% and 83.1% increase, respectively, as compared to the wild-type C. tyrobutyricum (Suo et al. 2018c).
Butyrate production in fermentation is strongly inhibited by butyric acid, which at > 10 g/L would reduce cell growth and metabolic activities by more than 80% (Wu and Yang 2003; Zhu and Yang 2003). Several Class I heat shock proteins (including dnaJ, dnaK, grpE, groES, groEL, and htpG) known to play important roles in resisting environmental stress were investigated for their effects on butyrate tolerance in C. tyrobutyricum. Among them, the overexpression of groESL significantly improved the butyrate tolerance and the mutant gave a high level of butyric acid production of up to 52.2 g/L, which was a 15.2% increase compared to the wild type strain (Suo et al. 2017). Wu et al. reported that overexpressing trehalose synthase (TreS), which converted maltose to trehalose, in C. tyrobutyricum increased the host’s oxidative resistance and robustness under hypoxic and aerobic conditions (Wu et al. 2017b). Interestingly, compared to the wild type strain the mutant also produced significantly more butyrate in batch fermentations under acidic conditions (pH 4.0 and 5.0). In addition, many earlier studies have focused on enhancing cell butyrate tolerance through adaptation in immobilized-cell bioreactor such as the fibrous bed bioreactor (FFB) (Jiang et al. 2011; Zhu and Yang 2003). While the original strain was unable to grow in the presence of 40 g/L butyric acid, cells immobilized in a FBB were able to produce up to 86.9 g/L butyric acid from glucose in a repeated fed-batch fermentation process (Jiang et al. 2011). The adapted cells in the FBB had an elongated rod morphology and significantly elevated intracellular pH, which might have contributed to the higher butyric acid tolerance.
Engineering strategies for enhancing n-butanol production from glucose
Yu et al. first introduced adhE2 into various strains of C. tyrobutyricum for n-butanol production from glucose, achieving a high butanol yield of 0.27 g/g (Yu et al. 2011). After optimizing the conjugative plasmid expression system, C. tyrobutyricum Δack-adhE2 produced 20.5 g/L of n-butanol with a high yield of 0.33 g/g with mannitol as the substrate (Yu et al. 2012). However, large amounts of acids (acetate and butyrate) were also produced. To overcome this problem, CoA transferase (encoded by ctfAB) from C. acetobutylicum was co-overexpressed with adhE2 in C. tyrobutyricum to facilitate the reassimilation of butyrate for n-butanol production, leading to over twofold increase in butanol productivity and yield (Yu et al. 2015a). However, acetone was also produced in the fermentation. More recently, using the native CRISPR-Cas system, Zhang et al. successfully knocked out cat1 with adhE2 insertion on the genome of C. tyrobutyricum and the mutant strain Δcat1::adhE2 produced 26.2 g/L n-butanol with a yield of 0.23 g/g and very little butyrate production (Zhang et al. 2018). However, large amounts of acetate and ethanol were also produced by this mutant in the fermentation. Additional metabolic and process engineering efforts are thus required to direct more carbon flux toward C4 compounds in order to further enhance n-butanol production, which might also be limited by NADH availability. These metabolic engineering strategies along with additional studies described in the following sections are summarized in Table 3.
Compared to butyric acid, each mole of butanol produced from butyryl-CoA requires additional two moles of NADH (see Fig. 1), which may cause redox imbalance. To increase NADH availability for butanol biosynthesis from glucose, Nguyen et al. (2018) knocked out the redox-sensing transcriptional repressor gene (rexA) and replaced NAD+-dependent 3-hydroxybutyryl-CoA dehydrogenase (hbd) with a heterologous NADP+-dependent 3-hydroxybutyryl-CoA dehydrogenase (hbd1) in C. acetobutylicum. They also replaced the native thiolase (thlA) with a heterologous acetoacetyl-CoA thiolase/synthase (atoB) to increase the flux from C2 (acetyl-CoA) to C4 (butyryl-CoA) and knocked out CoA transferase (ctfAB), butyrate kinase (buk) and phosphotransbutyrylase (ptb). The resulting mutant produced n-butanol as the main metabolic product at a high yield of 0.34 g/g glucose. Replacing NAD+-dependent 3-hydroxybutyryl-CoA dehydrogenase with NADP+-dependent one thus should have a positive effect on NADH availability for n-butanol biosynthesis in C. tyrobutyricum, which remains to be verified.
Some process engineering strategies have also been applied to improve n-butanol production. For example, adding MV as an artificial electron carrier in the fermentation by C. tyrobutyricum Δack-adhE2 reduced acetate and butyrate production by more than 80–90% and increased n-butanol production to 14.5 g/L with a high yield of > 0.3 g/g (Du et al. 2015). The MV effect on increased butanol production can be attributed to its effects on inhibiting hydrogen production and thus increasing available NADH for butanol biosynthesis. In addition, FBB was applied to immobilize C. tyrobutyricum adhE2, which not only dramatically increased cell density, but also improved butanol titer, yield, and productivity with reduced acid production (Huang et al. 2019).
Engineering C. tyrobutyricum for butyrate/butanol production from low-cost feedstocks
Although high-titer n-butanol and butyrate can be produced from glucose with a high yield because few byproducts are coproduced in the fermentation, especially with the addition of MV (Du et al. 2015), C. tyrobutyricum has a narrow substrate spectrum and can use only a few monosaccharides (glucose, xylose, and fructose), mannitol, and lactate for growth (Dwidar et al. 2012). This can be attributed to the fact that C. tyrobutyricum’s relatively small genome, compared to C. butyricum and C. acetobutylicum, is lacking genes for starch and disaccharides, such as maltose and sucrose, transport and catabolism (Jiang et al. 2018). In order to expand the substrate spectrum of C. tyrobutyricum, heterologous sucrose, maltose, and galactose catabolism pathways have been successfully introduced into C. tyrobutyricum.
For sucrose catabolism, sucrose-specific PTS (scrA), sucrose-6-phosphate hydrolase (scrB), and fructokinase (scrK) from C. acetobutylicum were co-expressed in C. tyrobutyricum (Guo et al. 2020). The mutant strain was able to utilize cane molasses as both nitrogen and carbon sources and produced 45.7 g/L butyric acid with a yield of 0.39 g/g in fed-batch fermentation. Similarly, C. tyrobutyricum was engineered to co-express adhE2 with scrK, scrB, and scrA for n-butanol production from sucrose (Zhang et al. 2017a). The mutant produced 16 g/L n-butanol with a yield of 0.24 g/g sugars from sugarcane juice supplemented with corn steep liquor (CSL) (Zhang et al. 2017b). Compared to glucose as the substrate, the feedstock cost was reduced by ~ 50% when cane molasses or sugarcane juice was used in the fermentation.
Metabolic engineering of C. tyrobutyricum for n-butanol production from maltose and soluble starch was also studied (Yu et al. 2015b). Two α-glucosidase genes, agluI and agluII, from C. acetobutylicum were cloned and co-expressed with adhE2 in C. tyrobutyricum Δack. The mutant expressing agluI demonstrated robust activity in breaking down the α-1,4-glycosidic bonds in maltose and starch and produced 17.2 g/L butanol from maltose with a yield of 0.20 g/g and productivity of 0.29 g/L·h in batch fermentation. With soluble starch, 16.2 g/L butanol was produced with a yield of 0.17 g/g and productivity of 0.20 g/L·h. Because of the inherent higher butanol tolerance, the mutant was able to produce more butanol at a remarkably higher productivity as compared to C. acetobutylicum ATCC 824 (11.2 g/L at 0.10 g/L·h from maltose and 8.8 g/L at 0.10 g/L·h from soluble starch).
For galactose catabolism, the recombinant C. tyrobutyricum kept was constructed by co-expressing UDP-galactose 4-epimerase (galE), galactokinase (galK), phosphoglucomutase (galP), and galactose-1-phosphate uridylyltransferase (galT) genes from C. acetobutylicum, which utilized glucose and galactose simultaneously without glucose-mediated carbon catabolite repression (CCR) (He et al. 2020). When using hydrolyzed coffee ground (rich in galactose) as the substrate, the mutant strain produced 34.3 g/L butyric acid with a yield of 0.37 g/g, which were 78.6% and 56.5%, respectively, higher than those from the wild-type strain.
C. tyrobutyricum can also use xylose as the sole carbon source in fermentation (Liu and Yang 2006). The product (butyrate and butanol) yields from xylose were comparable to those from glucose although xylose utilization involves the hexose monophosphate pathway that would give slightly less ATP but more NADH and thus somewhat different product profiles under different pH conditions (Zhu and Yang 2004). However, in the presence of glucose, xylose utilization was greatly inhibited by glucose-mediated CCR, which could be alleviated by overexpressing three xylose catabolism genes (xylB: xylulokinase, xylT: D-xylose-proton symporter, and xylA: xylose isomerase) from C. acetobutylicum (Fu et al. 2017a; Yu et al. 2015c).
Compared to C. butyricum and C. acetobutylicum, engineered C. tyrobutyricum strains with heterologous galactose, maltose, and sucrose catabolism genes gave better fermentation performance due to their higher tolerance to butyrate and butanol. The engineered C. tyrobutyricum strains thus can provide more robust and cost-effective processes for industrial butyrate and butanol production from food processing wastes such as sugarcane molasses and spent coffee ground.
Engineering strategies for using lignocellulosic biomass hydrolysates
Lignocellulosic biomass (LCB) is the most abundant renewable resource on the planet (Kumar et al. 2013). The feasibility of using LCB hydrolysates as low-cost feedstock has thus also been explored for butyric acid and butanol production by C. tyrobutyricum (Baroi et al. 2015; Cao et al. 2020; Chen et al. 2017; Huang et al. 2011; 2016a; 2016b; 2019; Liu et al. 2013; Oh et al. 2019; Sjoblom et al. 2016; Song et al. 2011; Wei et al. 2013; Xiao et al. 2018; Zheng et al. 2018; Zhu et al. 2002). The application of LCB in fermentation requires relatively harsh chemical, physical, and/or thermal pretreatments before enzymatic hydrolysis of cellulose. The pretreatment process usually generates chemical inhibitors derived from the degradation of lignin and sugars (Amiri and Karimi 2018; Sharma et al. 2019). In general, immobilized cells had better resistance to the hydrolysate inhibitors, especially after adaptation in bioreactors. Compared to free-cell fermentation, significantly higher butyrate and butanol titers and productivities were obtained from LCB hydrolysates when C. tyrobutyricum cells were immobilized in fibrous bed bioreactor (FBB) (Fu et al. 2017b; Li et al. 2019; Wei et al. 2013; Xiao et al. 2018). Various detoxification approaches (chemical, physical or biological methods) have been developed to remove inhibitors in the hydrolysates prior to fermentation (Jönsson et al. 2013). For example, an in-situ detoxification process using Tween 80 as a surfactant was found to be effective in removing hydrolysate inhibitors in pretreated rice straw hydrolysate, which after detoxification could be directly added in C. tyrobutyricum fermentation broth for butyrate production with comparable performance to that from pristine sugars (Lee et al. 2015).
However, detoxification is not always effective and can be costly (Jönsson et al. 2013). Improving cell tolerance to LCB-derived inhibitors via metabolic engineering was thus investigated. One study showed that the overexpression of Class I heat shock protein genes (groESL) improved the fermentation performance of C. tyrobutyricum with a significantly higher butyrate production from glucose (Suo et al. 2017) as well as LCB (corn straw and rice straw) hydrolysates as compared to the wild type (Suo et al. 2018b). More recently, a short-chain reductase (SDR) and aldo/keto reductases (AKR) from C. beijerinckii were investigated for enhancing the fermentability of undetoxified corncob acid hydrolysate (Suo et al. 2019). SDR and AKR can catalyze the reduction of furfural and 5-hyroxymethyl furfural (HMF) to corresponding alcohols, which are less toxic than the aldehydes (Suo et al. 2019). Compared to the parental strain, butyrate fermentation productivity was improved to 0.29 g/L·h with the butyric acid titer increased by 28.1% when sdr and groESL genes were co-overexpressed in C. tyrobutyricum (Suo et al. 2019).
LCB hydrolysates contain glucose and xylose as two main types of monosaccharide. Although most of clostridia, including C. acetobutylicum and C. tyrobutyricum, can use xylose as the sole carbon source, xylose utilization in the presence of glucose was greatly inhibited by CCR, leading to poor xylose consumption and low fermentation productivity (Xiao et al. 2012). To overcome the CCR in glucose/xylose co-fermentation, three xylose catabolism genes xylB, xylT, and xylA from C. acetobutylicum were expressed in C. tyrobutyricum Δack and Δack:adhE2 for butyrate and butanol production, respectively (Fu et al. 2017a; Yu et al. 2015c). Glucose and xylose co-utilization with significantly reduced residual xylose was achieved in batch fermentations with these mutants. The mutant Ct-pTBA was evaluated with the hydrolysates of sugarcane bagasse, rice straw, corn fiber, wheat straw, and soybean hull. A high butyric acid titer of 42.6 g/L with a yield of 0.36 g/g and productivity of 0.56 g/L·h was obtained from sugarcane bagasse hydrolysate (Fu et al. 2017b), which were significantly higher than those from the wild type (see Table 2). In batch fermentation with C. tyrobutyricum Δack:adhE2-pTBA, 15.7 g/L n-butanol with a yield of 0.24 g/g was produced from soybean hull hydrolysate (Yu et al. 2015c). Clearly, expressing xylA, xylB, and xylT alleviated the CCR bottleneck in C. tyrobutyricum and was effective in enhancing butyrate and butanol production from LCB hydrolysates containing glucose and xylose. Table 3 summarizes notable metabolic engineering strategies applied to C. tyrobutyricum for n-butanol production from various substrates.
Comparison to other bacterial hosts for butyrate and n-butanol production
Compared to the best recombinant microbes engineered to date for butyrate and n-butanol production, engineered C. tyrobutyricum strains generally gave higher product titer, yield, and productivity and thus would have greater potential for industrial application (see Table 4). Native solventogenic C. acetobutylicum produces acetone, butanol, and ethanol as the main products at a mass ratio of 6:3:1 with a relatively low butanol titer (10 − 14 g/L) and yield (~ 0.2 g/g). After multiple gene manipulations (overexpression and deletion of multiple genes) Nguyen et al. (2018), were able to engineer C. acetobutylicum to produce mainly n-butanol without acetone, achieving a high butanol yield of 0.34 g/g. Several mutant strains of C. acetobutylicum ATCC 824 with pta, ctfB, and adhE1 knockouts were able to produce up to 31 g/L butyric acid with a high BA/AA ratio of 31.3 g/g and negligible solvent production when buk was also inactivated (Jang et al. 2014). In addition, E. coli, which has the most well-developed genetic tools and has been extensively studied as a robust host for production of a variety of chemicals, has also been metabolically engineered to produce butyrate and n-butanol. For example, Shen et al. (2011) engineered E. coli to express a chimeric n-butanol biosynthetic pathway with increased NADH availability to achieve a high n-butanol titer of ~ 15 g/L with a yield of 0.28 g/g (~ 70% theoretical). Metabolically engineered E. coli strains were also constructed to produce butyrate at a high yield (0.31 − 0.43 g/g) with minimal acetate production, achieving a high selectivity with the highest BA/AA ratio of 143 g/g obtained from 20 g/L glucose and 8 g/L acetate in an LB medium (Saini et al. 2014). However, E. coli has relatively poor tolerance to butyric acid and butanol, and the highest butyrate and butanol titers produced so far were much lower than those from clostridial fermentations. Although C. tyrobutyricum is more difficult to engineer because of limited genetic engineering tools and its relatively low transformation efficiency, overall it is a better host with superior fermentation performance in product titer, yield, and productivity.
Moreover, compared to C. acetobutylicum and other solventogenic clostridia used in industrial ABE fermentation, C. tyrobutyricum is not as susceptible to sporulation (Xu et al. 2017) and bacteriophage infection (Jones et al. 2000). Although several strains of C. tyrobutyricum (NCIMB 9582, NCIMB 701753 and 701756) were found to be susceptible to the phage φCTP1 isolated from a landfill site (Mayer et al. 2010), no bacteriophage infection of C. tyrobutyricum ATCC 25755 has ever been observed in a continuous or fed-batch fermentation process operated for an extended period (over a month). It is noted that phage-resistant strains can be obtained through screening/isolation (Liu et al. 2017) or genetic engineering to clone and express a potent restriction/modification system (such as using CRISPR/Cas9 technology for double-strand DNA cleavage) targeting selected phage genes (e.g., endolysin) (Baltz et al. 2018).
Conclusions and prospects for further developments
Clostridium tyrobutyricum has attracted a great deal of interest as a robust host for butyrate and butanol production. To date, impressive progresses in strain and process engineering have been achieved for butyrate and butanol production from low-cost lignocellulosic biomass. However, at the current oil prices of ~ $40/barrel, bio-butyrate and butanol production by fermentation with native or engineered microorganisms including C. tyrobutyricum is not economically competitive with conventional chemical synthesis routes.
There are challenges and opportunities in further engineering C. tyrobutyricum for efficient utilization of lignocellulosic biomass hydrolysates to attain desirable product titer, yield and productivity suitable for industrial application. Genome-scale analyses, including comparative genomics, transcriptomics, and metabolomics analyses, are valuable in guiding rational metabolic engineering at a systems level and have been applied to clostridia (Yoo and Soucaille 2020; Ou et al. 2020) but not C. tyrobutyricum yet. Further strain engineering may also require more sophisticated strategies and approaches such as multivariate modular metabolic engineering (Biggs et al. 2014), which would require a well-characterized “toolbox” including replicon (ori), ribosomal binding sites (RBS), promoters, and reporters (Joseph et al. 2018). Replicon plays a significant role in plasmid copy number and transformation efficiency (Yu et al. 2012). RBS and promoter are important in regulating gene expression and balancing metabolic flux and redox potential, which are critical to optimizing cell growth and metabolic activities. Efficient reporter systems suitable for anaerobes, such as the one based on a flavin mononucleotide (FMN)-dependent fluorescent protein Bs2 (Cheng et al. 2019b), can facilitate the evaluation and screening of promoters with different strengths and thus would be valuable in promoter engineering. These novel genetic engineering toolkits and CRISPR-Cas9 genome-editing systems have rapidly advanced synthetic biology (Kwon et al. 2020; Joseph et al. 2018) and should facilitate the further development of C. tyrobutyricum for butyrate and n-butanol production.
In addition to metabolic engineering, adaptation or evolutionary engineering has also been demonstrated as an efficient strategy to enhance cell tolerance to toxic chemicals such as butyric acid and n-butanol. Cells highly tolerant to butyric acid or n-butanol were obtained after prolonged exposure to the corresponding metabolite produced in fed-batch or repeated batch fermentation in a FBB (Jiang et al. 2011; Yang and Zhao 2013; Zhu and Yang 2003). Comparative genomic analysis revealed that the butanol tolerant mutant strain C. acetobutylicum JB200 had a single-base deletion in a histidine kinase (encoded by cac3319). This finding led to the development of cac3319 knockout mutant with 45% higher butanol production (~ 18.2 g/L vs. ~ 12.6 g/L for the parental strain) and a 90% higher productivity (Xu et al. 2015). Histidine kinase is involved in the phosphorylation or activation of Spo0A, a global regulator in clostridia which is known to control not only sporulation but also stress response and solventogenesis in C. acetobutylicum (Steiner et al. 2011). It has also been reported that inactivating the sporulation transcription factor (spo0A) enhanced the butanol tolerance and production ability of Clostridium cellulovorans after adaptation (Wen et al. 2019). Therefore, we can speculate that knocking out histidine kinase and/or spo0A in C. tyrobutyricum may also enhance its ability to produce more butyric acid and butanol.
Finally, the engineered C. tyrobutyricum with enhanced tolerance can be used in an integrated process with in situ or on-line product separation, such as liquid–liquid extraction for butyric acid (Wu and Yang 2003) and gas stripping for butanol (Du et al. 2015; Lu et al. 2013), to further increase product titer, productivity, and yield, allowing for economical production of these metabolites in fermentation (Yang and Lu 2013).
References
Alam S, Stevens D, Bajpai R (1988) Production of butyric acid by batch fermentation of cheese whey with Clostridium beijerinckii. J Ind Microbiol 2:359–364
Amiri H, Karimi K (2018) Pretreatment and hydrolysis of lignocellulosic wastes for butanol production: challenges and perspectives. Bioresour Technol 270:702–721
Baltz RH (2018) Bacteriophage-resistant industrial fermentation strains: from the cradle to CRISPR/Cas9. J Ind Microbiol Biot 45:1003–1006
Bao G, Wang R, Zhu Y, Dong H, Mao S, Zhang Y, Chen Z, Li Y, Ma Y (2011) Complete genome sequence of Clostridium acetobutylicum DSM 1731, a solvent-producing strain with multireplicon genome architecture. J Bacteriol 193:5007–5008
Baroi GN, Baumann I, Westermann P, Gavala HN (2015) Butyric acid fermentation from pretreated and hydrolysed wheat straw by an adapted Clostridium tyrobutyricum strain. Microbiol Biotechnol 8:874–882
Bassi D, Fontana C, Gazzola S, Pietta E, Puglisi E, Cappa F, Cocconcelli PS (2013) Draft genome sequence of Clostridium tyrobutyricum strain UC7086, isolated from grana padano cheese with late-blowing defect. Genome Announc 1(4):e00614–e713
Biggs B, De Paepe B, Santos CNS, De Mey M, Ajikumar PK (2014) Multivariate modular metabolic engineering for pathway and strain optimization. Curr Opin Biotechnol 29:156–162
Cao X, Chen Z, Liang L, Guo L, Jiang Z, Tang F, Yun Y, Wang Y (2020) Co-valorization of paper mill sludge and corn steep liquor for enhanced n-butanol production with Clostridium tyrobutyricum Δcat1:adhE2. Bioresour Technol 296:122347
Chen T, Zhang L, Luo G, Yuan W (2017) Butyric acid production by Clostridium tyrobutyricum in sugar mixtures and corncob hydrolysate containing arabinose. BioResources 12:7931–7942
Cheng C, Bao T, Yang ST (2019a) Engineering Clostridium for improved solvent production: recent progress and perspective. Appl Microbiol Biotechnol 103:5549–5566
Cheng C, Lin M, Jiang W, Zhao J, Li W, Yang ST (2019b) Development of an in vivo fluorescence based gene expression reporter system for Clostridium tyrobutyricum. J Biotechnol 305:18–22
Choi O, Um Y, Sang B-I (2012) Butyrate production enhancement by Clostridium tyrobutyricum using electron mediators and a cathodic electron donor. Biotechnol Bioeng 109:2494–2502
Cummins CS, Johnson JL (1971) Taxonomy of clostridia: Wall composition and DNA homologies in Clostridium butyricum and other butyric acid-producing clostridia. J Gen Microbiol 67:33–46
D'Incecco P, Faoro F, Silvetti T, Schrader K, Pellegrino L (2015) Mechanisms of Clostridium tyrobutyricum removal through natural creaming of milk: A microscopy study. J Dairy Sci 98:5164–5172
Du Y, Jiang W, Yu M, Tang IC, Yang ST (2015) Metabolic process engineering of Clostridium tyrobutyricum Δack-adhE2 for enhanced n-butanol production from glucose: effects of methyl viologen on NADH availability, flux distribution, and fermentation kinetics. Biotechnol Bioeng 112:705–715
Dwidar M, Park JY, Mitchell RJ, Sang BI (2012) The future of butyric acid in industry. Sci World J 2012:471417
Fu H, Yu L, Lin M, Wang J, Xiu Z, Yang S-T (2017a) Metabolic engineering of Clostridium tyrobutyricum for enhanced butyric acid production from glucose and xylose. Metab Eng 40:50–58
Fu H, Yang S-T, Wang M, Wang J, Tang IC (2017b) Butyric acid production from lignocellulosic biomass hydrolysates by engineered Clostridium tyrobutyricum overexpressing xylose catabolism genes for glucose and xylose co-utilization. Bioresour Technol 234:389–396
Fu H, Hu J, Guo X, Feng J, Zhang Y, Wang J (2020) High-selectivity butyric acid production from Saccharina japonica hydrolysate by Clostridium tyrobutyricum. Ind Eng Chem Res. https://doi.org/10.1021/acs.iecr.0c01279
Guo X, Fu H, Feng J, Hu J, Wang J (2020) Direct conversion of untreated cane molasses into butyric acid by engineered Clostridium tyrobutyricum. Bioresour Technol 301:122764
He F, Qin S, Yang Z, Bai X, Suo Y, Wang J (2020) Butyric acid production from spent coffee grounds by engineered Clostridium tyrobutyricum overexpressing galactose catabolism genes. Bioresour Technol 304:122977
Heap JT, Pennington OJ, Cartman ST, Carter GP, Minton NP (2007) The ClosTron: a universal gene knock-out system for the genus Clostridium. J Microbiol Meth 70:452–464
Heap JT, Pennington OJ, Cartman ST, Minton NP (2009) A modular system for Clostridium shuttle plasmids. J Microbiol Meth 78:79–85
Heap JT, Kuehne SA, Ehsaan M, Cartman ST, Cooksley CM, Scott JC, Minton NP (2010) The ClosTron: Mutagenesis in Clostridium refined and streamlined. J Microbiol Meth 80:49–55
Huang J, Cai J, Wang J, Zhu XC, Huang L, Yang ST, Xu ZN (2011) Efficient production of butyric acid from Jerusalem artichoke by immobilized Clostridium tyrobutyricum in a fibrous-bed bioreactor. Bioresour Technol 102:3923–3926
Huang J, Dai H, Yan R, Wang P (2016a) Butyric acid production from recycled waste paper by immobilized Clostridium tyrobutyricum in a fibrous-bed bioreactor. J Chem Technol Biotechnol 91:1048–1054
Huang J, Zhu H, Tang W, Wang P, Yang S-T (2016b) Butyric acid production from oilseed rape straw by Clostridium tyrobutyricum immobilized in a fibrous bed bioreactor. Proc Biochem 51:1930–1934
Huang J, Du Y, Bao T, Lin M, Wang J, Yang ST (2019) Production of n-butanol from cassava bagasse hydrolysate by engineered Clostridium tyrobutyricum overexpressing adhE2: Kinetics and cost analysis. Bioresour Technol 292:121969
Jang YS, Im JA, Choi SY, Lee JI, Lee SY (2014) Metabolic engineering of Clostridium acetobutylicum for butyric acid production with high butyric acid selectivity. Metab Eng 23:165–174
Jawed K, Mattam AJ, Fatma Z, Wajid S, Abdin MZ, Yazdani SS (2016) Engineered production of short chain fatty acid in Escherichia coli using fatty acid synthesis pathway. PLoS ONE 11:e0160035
Jha AK, Li J, Yuan Y, Baral N, Ai B (2014) A review on bio-butyric acid production and its optimization. Int J Agric Biol 16:1019–1024
Jiang L, Wang J, Liang S, Cai J, Xu Z, Cen P, Yang ST, Li S (2011) Enhanced butyric acid tolerance and bioproduction by Clostridium tyrobutyricum immobilized in a fibrous bed bioreactor. Biotechnol Bioeng 108:31–40
Jiang L, Zhu L, Xu X, Li YP, Li S, Huang H (2013) Genome sequence of Clostridium tyrobutyricum ATCC 25755, a butyric acid-overproducing strain. Genome Announc 1:e00308–e313
Jiang L, Wu Q, Xu Q, Zhu LY, Huang H (2017) Fermentative hydrogen production from Jerusalem artichoke by Clostridium tyrobutyricum expressing exo-inulinase gene. Sci Rep 7(1):7940
Jiang L, Fu H, Yang HK, Xu W, Wang J, Yang ST (2018) Butyric acid: Applications and recent advances in its bioproduction. Biotechnol Adv 36:2101–2117
Jo JH, Jeon CO, Lee SY, Lee DS, Park JM (2010) Molecular characterization and homologous overexpression of [FeFe]-hydrogenase in Clostridium tyrobutyricum JM1. Int J Hydrogen Energy 35:1065–1073
Jones DT, Shirley M, Wu X, Keis S (2000) Bacteriophage infections in the industrial acetone butanol (AB) fermentation process. J Mol Microbiol Biotechnol 2:21–26
Jönsson LJ, Alriksson B, Nilvebrant N-O (2013) Bioconversion of lignocellulose: inhibitors and detoxification. Biotechnol Biofuels 6:16–25
Joseph RC, Kim NM, Sandoval NR (2018) Recent developments of the synthetic biology toolkit for Clostridium. Front Microbiol 9:154
Kataoka N, Vangnai AS, Pongtharangkul T, Yakushi T, Matsushita K (2017) Butyrate production under aerobic growth conditions by engineered Escherichia coli. J Biosci Bioeng 123:562–568
Kumar M, Goyal Y, Sarkar A, Gayen K (2013) Comparative economic assessment of ABE fermentation based on cellulosic and non-cellulosic feedstocks. Appl Energy 93:193–204
Kwon SW, Paari KA, Malaviya A, Jang YS (2020) Synthetic biology tools for genome and transcriptome engineering of solventogenic Clostridium. Front Bioeng Biotech 8:282
Lee KM, Kim KY, Choi O, Woo HM, Kim Y, Han SO, Sang BI, Um Y (2015) In situ detoxification of lignocellulosic hydrolysate using a surfactant for butyric acid production by Clostridium tyrobutyricum ATCC 25755. Proc Biochem 50:630–635
Lee J, Jang YS, Han MJ, Kim JY, Lee SY (2016) Deciphering Clostridium tyrobutyricum metabolism based on the whole-genome sequence and proteome analyses. mBio 7:e00743-16
Li C, Wang Y, Xie G, Peng B, Zhang B, Chen W, Huang X, Wu H, Zhang B (2016) Complete genome sequence of Clostridium butyricum JKY6D1 isolated from the pit mud of a Chinese flavor liquor-making factory. J Biotechnol 220:23–24
Li J, Du Y, Bao T, Dong J, Lin M, Shim H, Yang ST (2019) n-Butanol production from lignocellulosic biomass hydrolysates without detoxification by Clostridium tyrobutyricum △aack-adhE2 in a fibrous-bed bioreactor. Bioresour Technol 289:121749
Liu X, Yang ST (2006) Kinetics of butyric acid fermentation of glucose and xylose by Clostridium tyrobutyricum wild type and mutant. Process Biochem 41:801–808
Liu S, Bischoff KM, Leathers TD, Qureshi N, Rich JO, Hughes SR (2013) Butyric acid from anaerobic fermentation of lignocellulosic biomass hydrolysates by Clostridium tyrobutyricum strain RPT-4213. Bioresour Technol 143:322–329
Liu X, Turchi B, Mok KC, Taga ME, Miller MJ (2017) HM2-phage resistant solventogenic Clostridium saccharoperbutylacetonicum N1–4 shows increased exopolysaccharide production. FEMS Microbiol Lett 364:fnx191
Liu X, Zhu Y, Yang ST (2006a) Butyric acid and hydrogen production by Clostridium tyrobutyricum ATCC 25755 and mutants. Enzyme Microb Technol 38:521–528
Liu X, Zhu Y, Yang ST (2006b) Construction and characterization of ack deleted mutant of Clostridium tyrobutyricum for enhanced butyric acid and hydrogen production. Biotechnol Progr 22:1265–1275
Lu C, Dong J, Yang ST (2013) Butanol production from wood pulping hydrolysate in an integrated fermentation-gas stripping process. Bioresour Technol 143:467–475
Luo H, Yang R, Zhao Y, Wang Z, Liu Z, Huang M, Zeng Q (2018) Recent advances and strategies in process and strain engineering for the production of butyric acid by microbial fermentation. Bioresour Technol 253:343–354
Ma C, Kojima K, Xu N, Mobley J, Zhou L, Yang ST, Liu XM (2015) Comparative proteomics analysis of high n-butanol producing metabolically engineered Clostridium tyrobutyricum. J Biotechnol 193:108–119
Mayer MJ, Payne J, Gasson MJ, Narbad A (2010) Genomic sequence and characterization of the virulent bacteriophage ΦCTP1 from Clostridium tyrobutyricum and heterologous expression of its endolysin. Appl Environ Microbiol 76(16):5415–5422
Mo SJ, Kim BS, Yun SJ, Lee JJ, Yoon SH, Oh CH (2015) Genome sequencing of Clostridium butyricum DKU-01, isolated from infant feces. Gut Pathog 7:1–7
Morandi S, Cremonesi P, Silvetti T, Castiglioni B, Brasca M (2015) Development of a triplex real-time PCR assay for the simultaneous detection of Clostridium beijerinckii, Clostridium sporogenes and Clostridium tyrobutyricum in milk. Anaerobe 34:44–49
Munier E, Licandro-Seraut H, Achilleos C, Cachon R, Beuvier E (2019) Whole-genome sequencing and annotation of Clostridium tyrobutyricum strain Cirm BIA 2237, isolated from silage. Microbiol Resour Ann 8:e00492–e519
Nguyen NPT, Raynaud C, Meynial-Salles I, Soucaille P (2018) Reviving the Weizmann process for commercial n-butanol production. Nat Commun 9:3682
Oh HJ, Kim K-Y, Lee KM, Lee S-M, Gong G, Oh M-K, Um Y (2019) Enhanced butyric acid production using mixed biomass of brown algae and rice straw by Clostridium tyrobutyricum ATCC25755. Bioresour Technol 273:446–453
Ou J, Bao T, Ernst P, Si Y, Prabhu SD, Wu H, Zhang JJ, Zhou L, Yang ST, Liu XM (2020) Intracellular metabolism analysis of Clostridium cellulovorans via modeling integrating proteomics, metabolomics and fermentation. Process Biochem 89:9–19
Saini M, Wang ZW, Chiang CJ, Chao YP (2014) Metabolic engineering of Escherichia coli for production of butyric acid. J Agric Food Chem 62:4342–4348
Sharma HK, Xu C, Qin W (2019) Biological pretreatment of lignocellulosic biomass for biofuels and bioproducts: An overview. Waste Biomass Valor 10:235–251
Shen CR, Lan EI, Dekishima Y, Baez A, Cho KM, Liao JC (2011) Driving forces enable high-titer anaerobic 1-butanol synthesis in Escherichia coli. Appl Environ Microbiol 77:2905–2915
Sillers R, Chow A, Tracy B, Papoutsakis ET (2008) Metabolic engineering of the non-sporulating, non-solventogenic Clostridium acetobutylicum strain M5 to produce butanol without acetone demonstrate the robustness of the acid-formation pathways and the importance of the electron balance. Metab Eng 10:321–332
Sjoblom M, Matsakas L, Christakopoulos P, Rova U (2016) Catalytic upgrading of butyric acid towards fine chemicals and biofuels. Fems Microbiol Lett 363:7
Storari M, Wüthrich D, Bruggmann R, Berthoud H, Arias-Roth E (2015a) Draft genome sequences of Clostridium tyrobutyricum strains FAM22552 and FAM22553, isolated from Swiss semihard red-smear cheese. Genome Announc 3:e00078–e115
Soggiu A et al (2015) Draft genome sequence of Clostridium tyrobutyricum strain DIVETGP, isolated from cow's milk for grana padano production. Microbiol Resour Ann 3:e00213–e215
Song J-H, Ventura J-RS, Lee C-H, Jahng D (2011) Butyric acid production from brown algae using Clostridium tyrobutyricum ATCC 25755. Biotechnol Bioprocess Eng 16:42–49
Steiner E, Dago AE, Young DI, Heap JT, Minton NP, Hoch JA, Young M (2011) Multiple orphan histidine kinases interact directly with Spo0A to control the initiation of endospore formation in Clostridium acetobutylicum. Mol Microbiol 80:641–654
Storari M, Wuthrich D, Bruggmann R, Berthoud H, Arias-Roth E (2015b) Draft genome sequences of Clostridium tyrobutyricum strains FAM22552 and FAM22553, isolated from swiss semihard red-smear cheese. Microbiol Resour Ann 3:e00078–e115
Suo YK, Luo S, Zhang YA, Liao ZP, Wang JF (2017) Enhanced butyric acid tolerance and production by Class I heat shock protein-overproducing Clostridium tyrobutyricum ATCC 25755. J Ind Microbiol Biotechnol 44:1145–1156
Suo Y, Ren M, Yang X, Liao Z, Fu H, Wang J (2018a) Metabolic engineering of Clostridium tyrobutyricum for enhanced butyric acid production with high butyrate/acetate ratio. Appl Microbiol Biotechnol 102:4511–4522
Suo Y, Fu H, Ren M, Liao Z, Ma Y, Wang J (2018b) Enhanced butyric acid production in Clostridium tyrobutyricum by overexpression of rate-limiting enzymes in the Embden-Meyerhof-Parnas pathway. J Biotechnol 272:14–21
Suo Y, Fu H, Ren M, Yang X, Liao Z, Wang J (2018c) Butyric acid production from lignocellulosic biomass hydrolysates by engineered Clostridium tyrobutyricum overexpressing Class I heat shock protein GroESL. Bioresour Technol 250:691–698
Suo Y, Liao Z, Qu C, Fu H, Wang J (2019) Metabolic engineering of Clostridium tyrobutyricum for enhanced butyric acid production from undetoxified corncob acid hydrolysate. Bioresour Technol 271:266–273
Sushkova VI, Zhukovskii SV, Berezina OV, Yarotskii SV (2013) Biosynthesis of butyric acid from cabbage stem and molasses by the strain Clostridium butyricum VKPM B-9619. Russ J Bioorg Chem 39:771–776
Tamaru Y, Miyake H, Kuroda K, Nakanishi A, Kawade Y, Yamamoto K, Uemura M, Fujita Y, Doi RH, Ueda M (2010) Genome sequence of the cellulosome-producing mesophilic organism Clostridium cellulovorans 743B. J Bacteriol 192:901–902
Wang JF, Yang XR, Chen CC, Yang ST (2014) Engineering clostridia for butanol production from biorenewable resources: from cells to process integration. Curr Opin Chem Eng 6:43–54
Wang L, Ou MS, Nieves I, Erickson JE, Vermerris W, Ingram LO, Shanmugam KT (2015) Fermentation of sweet sorghum derived sugars to butyric acid at high titer and productivity by a moderate thermophile Clostridium thermobutyricum at 50 °C. Bioresour Technol 198:533–539
Wang J, Lin M, Xu M, Yang ST (2016) Anaerobic fermentation for production of carboxylic acids as bulk chemicals from renewable biomass. Adv Biochem Eng Biotechnol 156:323–361
Wasels F, Clement B, Ferreira NL (2016) Draft genome sequence of the butyric acid producer Clostridium tyrobutyricum strain CIP I-776 (IFP923) Microbiol Resour Ann 4:e00048–16
Wei D, Liu X, Yang ST (2013) Butyric acid production from sugarcane bagasse hydrolysate by Clostridium tyrobutyricum immobilized in a fibrous-bed bioreactor. Bioresour Technol 129:553–560
Wen Z, Ledesma-Amaro R, Lin JP, Jiang Y, Yang S (2019) Improved n-butanol production from Clostridium cellulovorans by integrated metabolic and evolutionary engineering. Appl Environ Microbiol 85:e02560–e2618
Wu ZT, Yang ST (2003) Extractive fermentation for butyric acid production from glucose by Clostridium tyrobutyricum. Biotechnol Bioeng 82:93–102
Wu Q, Liu TT, Zhu LY, Huang H, Jiang L (2017a) Insights from the complete genome sequence of Clostridium tyrobutyricum provide a platform for biotechnological and industrial applications. J Ind Microbiol Biotechnol 44:1245–1260
Wu Q, Zhu L, Xu Q, Huang H, Jiang L, Yang ST (2017b) Tailoring the oxidative stress tolerance of Clostridium tyrobutyricum CCTCC W428 by introducing trehalose biosynthetic capability. J Agric Food Chem 65:8892–8901
Xiao H, Li Z, Jiang Y, Yang Y, Jiang W, Gu Y, Yang S (2012) Metabolic engineering of D-xylose pathway in Clostridium beijerinckii to optimize solvent production from xylose mother liquid. Metab Eng 14:569–578
Xiao Z, Cheng C, Bao T, Liu L, Wang B, Tao W, Pei X, Yang ST, Wang M (2018) Production of butyric acid from acid hydrolysate of corn husk in fermentation by Clostridium tyrobutyricum: kinetics and process economic analysis. Biotechnol Biofuels 11:164
Xu M, Zhao J, Yu L, Tang IC, Xue C, Yang ST (2015) Engineering Clostridium acetobutylicum with a histidine kinase knockout for enhanced n-butanol tolerance and production. Appl Microbiol Biotechnol 99:1011–1022
Xu M, Zhao J, Yu L, Yang ST (2017) Comparative genomic analysis of Clostridium acetobutylicum for understanding the mutations contributing to enhanced butanol tolerance and production. J Biotechnol 263:36–44
Xue C, Zhao JB, Chen LJ, Yang ST, Bai FW (2017) Recent advances and state-of-the-art strategies in strain and process engineering for biobutanol production by Clostridium acetobutylicum. Biotechnol Adv 35:310–322
Yang ST, Lu C (2013) Extraction-fermentation hybrid (extractive fermentation). In: Ramaswamy S, Ramarao BV, Huang H (eds) Separation and purification technologies in biorefineries. Wiley, Chichester, pp 409–437
Yang ST, Zhao J (2013) Adaptive engineering of Clostridium for increased butanol production, US Patent 8450093
Yang ST, Yu M, Chang WL, Tang IC (2013) Anaerobic fermentations for the production of acetic and butyric acids. In: Yang ST, El-Enshasy HA, Thongchul N (eds) Bioprocessing technologies in biorefinery for sustainable production of fuels, chemicals, and polymers. Wiley, Hoboken, pp 351–373
Yang X, Xu M, Yang ST (2015) Metabolic and process engineering of Clostridium cellulovorans for biofuel production from cellulose. Metab Eng 32:39–48
Yoo M, Soucaille P (2020) Trends in systems biology for the analysis and engineering of Clostridium acetobutylicum metabolism. Trends Microbiol 28(2):118–140
Yu L, Zhao J, Xu M, Dong J, Varghese S, Yu M, Tang IC, Yang ST (2015) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production: effects of CoA transferase. Appl Microbiol Biotechnol 99:4917–4930
Yu L, Xu M, Tang IC, Yang ST (2015a) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production from maltose and soluble starch by overexpressing α-glucosidase. Appl Microbiol Biotechnol 99:6155–6165
Yu L, Xu M, Tang IC, Yang ST (2015b) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production through co-utilization of glucose and xylose. Biotechnol Bioeng 112:2134–2141
Yu M, Du Y, Jiang W, Chang WL, Yang ST, Tang IC (2012) Effects of different replicons in conjugative plasmids on transformation efficiency, plasmid stability, gene expression and n-butanol biosynthesis in Clostridium tyrobutyricum. Appl Microbiol Biotechnol 93:881–889
Yu M, Zhang Y, Tang IC, Yang ST (2011) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production. Metab Eng 13:373–382
Zhang J, Hong W, Guo L, Wang Y, Wang Y (2020) Enhancing plasmid transformation efficiency and enabling CRISPR-Cas9/Cpf1-based genome editing in Clostridium tyrobutyricum. Biotechnol Bioeng, in press, https://doi.org/10.1002/bit.27435
Zhang J, Yu L, Xu M, Yang ST, Yan Q, Lin M, Tang IC (2017) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production from sugarcane juice. Appl Microbiol Biotechnol 101:4327–4337
Zhang J, Yu L, Lin M, Yan Q, Yang ST (2017) n-Butanol production from sucrose and sugarcane juice by engineered Clostridium tyrobutyricum overexpressing sucrose catabolism genes and adhE2. Bioresour Technol 233:51–57
Zhang J, Zong W, Hong W, Zhang ZT, Wang Y (2018) Exploiting endogenous CRISPR-Cas system for multiplex genome editing in Clostridium tyrobutyricum and engineer the strain for high-level butanol production. Metab Eng 47:49–59
Zhao J, Lu C, Chen CC, Yang ST (2013) Biological production of butanol and higher alcohols. In: Yang ST, El-Enshasy HA, Thongchul N (eds) Bioprocessing technologies in biorefinery for sustainable production of fuels, chemicals, and polymers. Wiley, New York, pp 235–261
Zheng W, Liu X, Zhu L, Huang H, Wang T, Jiang L (2018) Pretreatment with gamma-valerolactone/mmim dmp and enzymatic hydrolysis on corncob and its application in immobilized butyric acid fermentation. J Agric Food Chem 66:11709–11717
Zhu Y, Yang ST (2003) Adaptation of Clostridium tyrobutyricum for enhanced tolerance to butyric acid in a fibrous-bed bioreactor. Biotechnol Prog 19:365–372
Zhu Y, Yang ST (2004) Effect of pH on metabolic pathway shift in butyric acid fermentation by Clostridium tyrobutyricum. J Biotechnol 110:143–157
Zhu Y, Liu XG, Yang ST (2005) Construction and characterization of pta gene-deleted mutant of Clostridium tyrobutyricum for enhanced butyric acid fermentation. Biotechnol Bioeng 90:154–166
Zhu Y, Wu ZT, Yang ST (2002) Butyric acid production from acid hydrolysate of corn fibre by Clostridium tyrobutyricum in a fibrous-bed bioreactor. Proc Biochem 38:657–666
Zigova J, Sturdik E (2000) Advances in biotechnological production of butyric acid. J Ind Microbiol Biotechnol 24:153–160
Zigova J, Sturdik E, Vandak D, Schlosser S (1999) Butyric acid production by Clostridium butyricum with integrated extraction and pertraction. Process Biochem 34:835–843
Acknowledgments
Some of the authors’ work described in this review was supported by the National Science Foundation STTR program (IIP-0810568, IIP-1026648) and Ohio Department of Development—Third Frontier Advanced Energy Program (Tech 08-036).
Funding
Some of the work described in this review paper was supported by the National Science Foundation STTR program (IIP-0810568, IIP-1026648), Ohio Department of Development—Third Frontier Advanced Energy Program (Tech 08-036).
Author information
Authors and Affiliations
Contributions
STY had the idea for the article. TB, JF, WJ, and HF performed the literature search and data analysis. BT and JF each drafted different sections of the paper. JW and STY critically revised the work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bao, T., Feng, J., Jiang, W. et al. Recent advances in n-butanol and butyrate production using engineered Clostridium tyrobutyricum. World J Microbiol Biotechnol 36, 138 (2020). https://doi.org/10.1007/s11274-020-02914-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02914-2