Abstract
Clostridium beijerinckii is identified as a promising Clostridium strain for industrialization of acetone and butanol (AB) fermentation. It has been reported that high reducing power levels are associated with high butanol yield. In this study, we regulated reducing power by blocking NAD(P)H consumption in C. beijerinckii NCIMB 8052. Gene Cbei_4110, encoding NADH-quinone oxidoreductase (nuoG), is a subunit of the electron transport chain complex I. After inactivation of gene Cbei_4110, the generated mutant strain exhibited a remarkable increase in glucose utilization ratio and enhanced butanol production to 9.5 g/L in P2 medium containing 30 g/L of glucose. NAD(P)H and ATP levels were also increased by one to two times and three to five times, respectively. Furthermore, a comparative transcriptome analysis was carried out in order to determine the mechanism involved in the enhanced activity of the Cbei_4110-inactivated mutant strain. This strategy may be extended for making industrial bio-butanol more economically attractive.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acetone and butanol (AB) fermentation was one of the largest industrial fermentation processes in the early twentieth century (Jones and Woods 1986; Lee et al. 2008). It has recently attracted the interest of scientists as a potential solution to decreasing petroleum resources and the ever-deteriorating environment (Green 2011; Tracy 2012). Before new energy resources (such as nuclear energy and solar energy) are commonly utilized, it is predicted that biomass power will be the first alternative energy source used, which will complement fossil feedstock (Lin and Tanaka, 2006). However, high fermentation substrate prices, low solvent production, and other drawbacks limit industrialization of bio-butanol (Ezeji et al. 2007; Jones and Woods 1986 and Qureshi et al. 2008).
Clostridium beijerinckii, an anaerobic Gram-positive Clostridium species, is capable of utilizing a broader spectrum of substrates with no glucose repression effect and has better toxin resistance to lignocellulosic hydrolysate than another main solvent-forming strain: Clostridium acetobutylicum (Ezeji et al. 2007). Since C. beijerinckii is identified as a promising Clostridium strain which can realize the economic benefit by using lignocellulosic sugars, improving butanol yield of AB fermentation by C. beijerinckii has attracted the attention of many researchers.
Energy and reducing power (NADH/NADPH) has been reported to have a significant influence in cell growth, metabolism, and production (Jones and Woods 1986). It was reported that high ATP concentration could increase solvent production in continuous cultures despite its effect on cell growth and metabolism (Meyer and Papoutsakis 1989). Reducing power participates in multiple bio-reactions in Clostridia and plays a crucial role in solvent formation. Along with formation of acids, NAD(P)H accumulates in excess and is oxidized in formation of hydrogen which is delivered by ferredoxins and catalyzed by hydrogenases (Jones and Woods 1986). Previous studies demonstrated that regulating Clostridia intracellular reducing power levels would produce a significant increase in solvent yield (Liu et al. 2013; Meyer and Papoutsakis 1989; Ventura et al. 2013).
In this study, we focused on genetic modification of C. beijerinckii NCIMB 8052 to improve its intracellular reducing power levels. Two genes were identified and mutated using ClosTron technology (Cooksley et al. 2012; Heap et al. 2009). Consequently, gene Cbei_4110 encoding NADH-quinone oxidoreductase (nuoG) in electron transport chain complex I was inactivated and the mutant strain showed a significant increase in the glucose conversion ratio and butanol yield than that of the wild-type strain. However, inactivation of the gene Cbei_3796 encoding hydrogenase (hydA) could not be achieved. In order to determine correlation between the phenotypic changes of growth and solvent production with the intracellular energy and reducing power in C. beijerinckii, the NAD(P)+, NAD(P)H, and ATP levels of the mutant and the wild-type strains were tested. Furthermore, we carried out a transcriptome analysis and determined transcriptional differences between these two strains.
Materials and methods
Strains, plasmids, media, and cultivation conditions
The bacterial strains and plasmids used in the study were listed in Table 1. Escherichia coli DH5α and TOP10 cells were grown at 37 °C in Luria-Bertani broth or agar medium supplemented with 15 mg/L tetracycline and 50 mg/L ampicillin as required. C. beijerinckii cells were grown anaerobically in liquid or solid YPS medium at 37 °C (1-L medium contains 3.0 g yeast extract, 5.0 g peptone, 10.0 g soluble starch, 2.0 g ammonium acetate, 2.0 g NaCl, 3.0 g MgSO4·7H2O, 1.0 g KH2PO4, 1.0 g K2HPO4, and 0.1 g FeSO4·7H2O, adjusted to pH 6.0 with 1 mol/L HCl) with addition of 10 mg/L erythromycin as required (Guo et al. 2012). P2 medium (Baer et al. 1987) (contained the following components: 30 g/L glucose, 0.5 g/L KH2PO4, 0.5 g/L K2HPO4, ammonium acetate 2.2 g/L, 1 mg/L para-amino-benzoic acid, 1 mg/L thiamine, 0.01 mg/L biotin, 0.01 g/L MnSO4·H2O, 0.01 g/L NaCl, 0.2 g/L MgSO4·7H2O, and 0.01 g/L FeSO4·7H2O) was used for C. beijerinckii fermentation. Escherichia coli and C. beijerinckii strains were maintained in 30 % (v/v) glycerol and stored at −80 °C.
Construction of plasmid pWJ1-392a, pWJ1-1146s, pWJ1-148a, pWJ1-771s, and pWD-4110
The introns were designed by a publically available computational method (Perutka et al. 2004). We chose the 392a and 1146s positions for Cbei_4110 insertion and chose 148a and 771s for Cbei_3796 insertion using the Clostron system (http://www.clostron.com), then generated 350 bp group II intron insertion fragments. Subsequently, we synthesized group II intron fragments and constructed plasmids pWJ-392a/1146s/148a/771s, by inserting these two 350 bp group II intron fragments into Xho I and BsrG I restriction sites of the pWJ1 plasmid. All primers involved in this process were listed in Table 2.
Construction of the expression plasmid was performed as follows: we amplified the full-length CDS of the Cbei_4110 gene using prime pIMP1-4110-up/dn and the C. beijerinckii NCIMB 8052 genome DNA as a template. The carrier vector was digested by the Nde I restriction enzyme. After DNA purification, gene Cbei_4110 was cloned into pWD using the infusion one-step clone kit (Vazyme Biotech Inc., Nanjing, China).
Transformation of C. beijerinckii and identification of mutants
All plasmids were initially methylated in E.coli TOP10 (pAN2) then transformed via electroporation. The specific process was performed as follows: First, 200 μL of frozen stock was inoculated into 10 mL of liquid YPS medium and incubated in an anaerobic environment at 37 °C for 12 h. Then 2 % (v/v) of the above inoculum was transferred into 50 mL YPS medium for secondary seed followed by anaerobic incubation at 37 °C for approximately 4 h when the optical density of 600 nm (OD600) of the cell reached 1.0. Cells were then harvested by centrifugation at 4000g for 10 min at 4 °C. The cell pellet was washed with 25 mL of ice-cold ETM buffer (270 mM sucrose, 0.6 mM Na2HPO4, 4.4 mM NaH2PO4, and 10 mM MgCl2; pH = 6.0) and resuspended gently, then repeated once more. The resulting cell pellet was resuspended in 1 mL ice-cold ET buffer (270 mM sucrose, 0.6 mM Na2HPO4, and 4.4 mM NaH2PO4; pH = 6.0) and used immediately for electroporation. For each electroporation, 190 μL of C. beijerinckii cells suspended in ET buffer were mixed with 10 μL methylated plasmid (no more than 1 μg) and added into a 0.2 cm precooled electroporation cuvette. Cells were then electrically pulsed by a Gene Pulser Xcell System (Bio-Rad) under conditions of 1800 V, 25 uF, and 200 Ω. Then, the competent cells were added to 5 mL of YPS medium and incubated for 5 h for recovery. Recovered cells were plated on YPS agar containing 10 mg/L erythromycin and incubated anaerobically at 37 °C for 48 h (Xiao et al. 2012).
Colony PCR was used for screening inactivated mutants to identify the positive transformation and isolate the mutant strains. The electrophoretic band of the PCR product of intron-inserted strains was about 1000 bp longer than the wild-type strain. For the next genetic operation, we needed to derive a strain that had lost the replicating plasmid. The intron-inserted strains were streak inoculated on YPS agar to isolate individual colonies and then incubated in YPS agar with or without erythromycin. The strains that grew in YPS medium but could not survive in erythromycin selection agar were considered the losing strain pWJ1 plasmids. For gene expression in C. beijerinckii, the methylated plasmids were introduced into cells by electroporation and identified via colony PCR as described above (Xiao et al. 2012).
Fermentations and analysis methods
A two-step level of seed cell activation was carried out before fermentation of C. beijerinckii. P2 medium (30 g/L glucose) was used for C. beijerinckii fermentation. Secondary seed cells (5 mL; OD600 = 2) were added to 45 mL of P2 medium in 100 mL serum bottles (Scott, Landshut, Germany) and flushed into sterile nitrogen gas for 3 min to eliminate oxygen poisoning. Bottle fermentation was then carried out in an anaerobic condition at 37 °C and continued for 72 h (Guo et al. 2012).
Batch fermentation was carried out in a 3-L fermenter (Bioflo110, NBS, CT, USA) with an initial broth volume of 2 L containing P2 (30 g/L glucose) medium, sterilized in an autoclave (115 °C for 20 min), and nitrogen gas was then bubbled through the medium to remove oxygen before and after inoculation; the temperature was maintained at 37 °C without agitation or pH control for 72 h. During the course of fermentation, 10 mL samples were collected every 6 h for measurement of OD600, sugar, and analysis of productions.
Glucose, acetate, and butyrate concentrations were determined by HPLC analysis (Agilent 1100 series, Wilmington, DE, USA), using an Aminex HPX-87H ion exclusion column (300 × 7.8 mm; Bio-Rad Laboratories, Hercules, CA, USA), with a 5.0-mM H2SO4 solution used as the mobile phase (0.6 mL/min) at 50 °C and a refractive index detector. Acetone, ethanol, and butanol were analyzed using gas chromatography (7890A, Agilent, Wilmington, DE, USA) equipped with a flame ionization detector (FID) and an Agilent HP-INNOWAX column (0.25 mm × 60 m). To determine hydrogen production, all produced gases were collected in gasbags and the hydrogen ratio was measured by a gas analyzer (Tandem Multiplex, Magellan Instruments, Herts, UK).
Measurement of the intracellular NAD(P)+, NAD(P)H, and ATP concentrations
Intracellular concentrations of NAD(P)H were determined by the enzyme cycling method of Bernofsky and Swan (1973) with modifications. As described previously (Liu et al. 2013), 5 mL of sample were centrifuged, the supernatant was discarded and the remaining pellet was dissolved in 0.5 mL of 0.2 M NaOH and HCl for cell lysate and decomposing NAD(P)H and NAD(P)+, respectively. The cell lysate was heated at 60 °C for 30 min, then cooled to 0 °C and centrifuged. The following reagents were sequentially added to the test tube and kept at 37 °C for 5 min in the dark: 350 μL of ddH2O, 200 μL of 0.1 M HCl, 100 μL of Tris-HCl (1 M, pH 7.8), 200 μL of the alkaline extract, 100 μL of 4.2 mM MTT, 150 μL of 16.6 mM PES, and 100 μL of ethanol for the determination of NADH or 100 μL of 60 mM glucose 6-phosphate for the determination of NADPH. The reaction was started by adding 10 μL of alcohol dehydrogenase (660 units/mL for NADH) or glucose 6-phosphate dehydrogenase (70 units/mL for NADPH) and incubated at 37 °C for 30 min in the dark. Absorbance at 570 nm was then determined. The same procedure was followed for NAD(P)H standards.
Intracellular concentrations of ATP were measured by fluorescent detection. To measure the intracellular ATP concentration, 1 mL of cold 30 % (v/v) trichloroacetic acid was added to the samples (4 mL) and mixed thoroughly. The ATP concentrations were then measured using the BacTiter-Glo™ Microbial Cell Viability Assay kit on the GloMax-Multi+ Detection System (Promega, Madison, WI, USA)
Total RNA and protein extraction, cDNA preparation, and RNA-seq analysis
Total RNA was extracted using TRIzol reagent according to the manufacturer’s protocol (Takara Bio Inc., Otsu, Japan). The integrity and purity of the RNA were determined using Trizol™ (Invitrogen, Carlsbad, CA, USA) and electrophoresis using a 1 % agarose gel. Illumina sequencing was performed at the Beijing Genomics Institute (BGI)-Shenzhen, Shenzhen, China (http://www.genomics.cn/index.php) according to the manufacturer’s instructions (Illumina, San Diego, CA, USA).
Mapping of RNA-Seq reads and quantitative analysis of gene expression
Clean reads were obtained by removing raw reads that contained the adaptor or unknown or low-quality sequences. These reads were then used for mapping to the C. beijerinckii NCIMB 8052 reference genome using the short oligonucleotide alignment program (SOAP), allowing up to five base mismatches (Gao et al. 2011; Li et al. 2009). For quantification, gene expression levels were calculated using the reads per kilobase per million reads (RPKM) method (Mortazavi et al. 2008), thereby limiting the effects of different gene lengths and sequencing levels. Rigorous algorithms were applied to identify differentially expressed genes (DEGs) between control and heat-treated samples at the BGI based on previously described methods (Audic and Claverie 1997). DEGs were identified using a false discovery rate (FDR) ≤0.001 and an absolute value of the log2 ratio ≥1 as the threshold.
DEGs functional annotation and pathway enrichment analysis
To further characterize the biological functions and metabolic pathways of DEGs, the DEGs were subjected to a gene ontology (GO) functional analysis (http://www.geneontology.org/) and Encyclopedia of Genes and Genome (KEGG) pathway enrichment analysis (Kanehisa et al. 2008). Both the annotation and pathway enrichment analyses utilized a corrected p value ≤0.05 as a threshold.
Illumina sequencing data
The Illumina sequencing data of the present study was deposited into the NCBI database with an accession number of SRP060345.
Results
Inactivation of gene Cbei_4110 promoted C. beijerinckii fermentation
Gene Cbei_4110 was inactivated at sites 392a and 1146s using TargeTron technology. Colony PCR and sequencing were used for screening and confirming the mutant strains 8052-4110392a and 8052-41101146s. As expected, both of the mutant strains 8052-MU (the two mutant strains demonstrated similar characteristics and were therefore collectively referred to as 8052-MU) showed significant increases in growth, glucose conversion ratio, and butanol yield than the 8052-wild-type (WT) strain. As seen in Fig. 1, the fermentation period of the 8052-MU was remarkably shorter than the wild-type strain, which were 38 and 60 h, respectively. Similar results were also seen with the glucose consumption rate. At the end of batch fermentation, the mutant strains 8052-MU yielded 9.5 g/L butanol and 12.1 g/L total solvent while that of the wild strain were 7.8 g/L and 10.2 g/L, respectively. Butanol and total solvent production were therefore enhanced by 21.8 and 18.6 %, respectively, and the butanol ratio rose to 78.5 %.
To confirm that the phenotypic changes observed in the mutant strains 8052-MU were caused by inactivation of gene Cbei_4110, a complementation mutant (named 8052-CP) was constructed. The expression plasmid pWD-4110 was introduced into 8052-MU, and the empty vector pWD was also introduced into 8052-MU; these mutants were confirmed by colony PCR. Compared to the mutant strains 8052-MU, the Cbei_4110 complement mutant strain 8052-CP showed obvious decreases in growth rate, glucose consumption, and ABE production, which was even lower than its counterpart 8052-WT. This result confirmed that the changes of the mutant strain 8052-MU was caused by inactivation of gene Cbei_4110. The excessive decrease in activity observed may be due to the fact that the ptb promoter of the pWD expression plasmid is a strong promoter, which results in increased Cbei_4110 expression (Xiao et al. 2012).
Energy and reduction energy changes in 8052-MU and 8052-WT
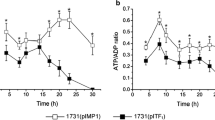
In order to investigate the energy and reduction energy changes of C. beijerinckii after Cbei_4110 inactivation, samples of the 8052-MU and 8052-WT strains were analyzed at both the mid-acidogenic (time of fastest pH drop rate) and mid-solventogenic (time of fastest butanol formation rate) phases. Since 8052-MU strains demonstrated higher fermentation efficiency than the 8052-WT strain, the sampling times were 16 and 25 h for 8052-MU whereas 8052-WT was 18 and 30 h after onset of fermentation. As shown in Table 3, ATP levels of the 8052-MU were significantly increased, which resulted in a five times increase in the acid formation period and three times increase in the solvent formation period. The reduction energy levels of 8052-MU were similar to 8052-WT in the acid formation period. When fermentation reached solvent formation period, the reduction energy levels of the mutant strains were twice than that of the parent strain. These results indicate that energy supply in 8052-MU is more abundant than 8052-WT in the whole fermentation period. Reduction energy in C. beijerinckii has reached the peak point in acidogenic phase and which is insufficient during the solventogenic phase. High concentration of NAD(P)H links with high concentration solvent.
Comparative transcriptome analysis of wild-type C. beijerinckii NCIMB 8052 and mutant strains
To explore the effects of Cbei_4110 gene inactivation on cellular metabolism, RNA sequencing was used to carry out a comparative transcriptome analysis between the wild-type and the Cbei_4110 inactivated mutant strain. RNA samples were extracted from fermentation of both the mid-acidogenic and the mid-solventogenic phase and then sequenced using the Illumina HiSeq 2000 system. A total of 5003 C. beijerinckii NCIMB 8052 genes capturing about 98.0 % of the genome were examined. Approximately 1538 genes were differentially expressed (up- or downregulated) by 8052-MU strains compared with the 8052-WT strain in the acidogenic growth phase, while there were 929 differentially expressed genes (DEGs) identified in the solventogenic growth phase (Fig. 2). DEGs were relative to redox cofactors, membrane transporters, and nucleotide metabolisms. In addition, expression patterns between the acidogenic and solventogenic growth phases were dramatically different.
Expression of sugar transporter and glycolysis genes
The phosphoenolpyruvate (PEP)-dependent phosphotransferase system (PTS) plays an important role in sugar uptake by Clostridia and consists of multiprotein complexes (enzyme I, HPr, and enzyme II) that mediate the uptake and phosphorylation of carbohydrates (Lee and Blaschek 2001). Among the PTS systems, the mannose family PTS (Man-PTS) transporters are deemed the major sugar transporters exhibiting broad substrate specificity toward glucose, mannose, sorbose, fructose, and a variety of other sugars (Barabote and Saier 2005). In the acidogenic phase of batch fermentation, a consistent fourfold upregulation in expression of the 8052-MU genes (Cbei_0711, Cbei_0712, and Cbei_0713) encoding Man-PTS transporters (manII AB, manII C, and manII D, respectively) was observed, which is consistent with the work of Wang et al. (2012). Interestingly, the trend of solventogenesis was contrary to that of acidogenesis which abovementioned genes expressed twofold lower (Fig. 3a). The expression level of enzyme I encoded by Cbei_0196 increased one time than that in the acidogenic phase. These results support previously reported findings demonstrating that ATP-dependent glucose phosphorylation was predominant and phosphoenolpyruvate-dependent glucose phosphorylation was repressed during the solventogenic stage (Wang et al. 2012). The gene expression patterns of other PTS systems were similar to that of the Man-PTS transporters. As shown in Fig. 3b, genes involved in glycolysis including gck (Cbei_4895), pfk (Cbei_4852), tpi (Cbei_0599), pgk (Cbei_0598), gap (Cbei_0597), gpmA (Cbei_1987 and Cbei_1759), and pyk (Cbei_4851 and Cbei_1412) were significantly upregulated (two- to sixfold) in the acidogenic phase. In the solventogenic phase, however, glycolysis gene expression patterns were consistent with the 8052-WT strain. These results indicate that the 8052-MU strains had faster glucose uptake and glycolytic capacity at the beginning of batch fermentation than the 8052-WT strain.
Expression of acidogenesis and solventogenesis genes
During acid formation, butyryl-CoA formation genes: crt (Cbei_0321 and Cbei_2034), bcd (Cbei_0322), etfA (Cbei_0323), etfB (Cbei_0323), and thlA (Cbei_0411), were highly expressed with twofold increase; butyrate formation genes: ptb (Cbei_0203), buk (Cbei_0204, Cbei_4006, and Cbei_4609), were slightly increased; however, acetate formation genes: pta (Cbei_1164) and ack (Cbei_1165) expression was downregulated by a fold in the acidogenic phase. Sovlentogenesis genes encoding ald (Cbei_3832), ctfA (Cbei_3833), ctfB (Cbei_3834), adc (Cbei_3835), and adh (Cbei_1722 and Cbei_2181) demonstrated an expression pattern very similar to 8052-WT. Samples extracted in the mid-solventogenic phase of 8052-MU showed generally a decline in expression by twofold of both acidogenesis and solventogenesis genes (shown in Fig. 3c).
Expression of cofactor genes
Energy is the fundamental demand of microorganism growth and metabolism. ATP, a common currency of energy, directly participates in all kinds of bioactivities. As shown in Fig. 3d, in the acidogenic phase, expression of ATP synthase genes (Cbei_0412–Cbei_0419) in 8052-MU were highly upregulated by twofold than 8052-WT, which may indicate that the mutant strain had a faster ATP synthesis speed rate than the parent strain. ATP-dependent enzymes and other ATPase-related genes exhibited remarkable differences in expression. The gene ontology (GO) analysis shows that these ATP-relevant genes are involved in ATP-binding cassette transport systems (sugar, amino acid, peptide, inorganic salts, and metal ions), DNA helicase (Cbei_1824), and DNA replication (Cbei_4288). Furthermore, Cbei_1224–Cbei_1227 and Cbei_1206, which are involved in the riboflavin biosynthesis pathway, were expressed sixfold higher in the acidogenic phase and sixfold lower in the solventogenic phase than the parent strain. Genes involved in the folate biosynthesis pathway (Cbei_1784, Cbei_2234, and Cbei_3249) were downregulated about 1.5 times in the acidogenic phase, and Cbei_0206 was downregulated twofold in the solventogenic phase. Genes (Cbei_3795–3798) encoding hydA expression were 1.5–4 times lower in the solventogenic phase while slightly higher (<1 time) in the acidogenic phase.
Other differentially expressed genes
Besides the differences in gene expression demonstrated above, genes in many other biological processes, cellular components, and molecular functions also demonstrated significant changes in expression. In the acid formation period, genes involved in amino acid synthesis, fatty acid metabolism, purine metabolism, and ribosomal proteins were universally upregulated. Genes involved in the two-component signal transduction system were alternatively upregulated or downregulated. Expression of chemotaxis and cell motility genes and genes expressed in folate synthesis and some degradation processes were downregulated. Other differentially expressed genes are shown in the supplementary material (Table S1 and Table S2).
Discussion
C. beijerinckii NCIMB 8052, one kind of anaerobic heterotrophic microorganism, maintains energy metabolism mainly through substrate level phosphorylation (SLP) by conversion of carbohydrates to hydrogen, carbon dioxide, acids, and solvents (Thauer et al. 1977). In the initial stages of fermentation, ATP is the desired energy source for biosynthesis of cell materials in glycolysis accompanied by excess generation of reduced cofactors (NADH + H+). For keeping up the redox balance, these reduced cofactors must be regenerated by means of hydrogenases with release of H2 or by reduction of electron accepters with a final formation of end products such as ethanol and butanol. It was previously reported that regulation of reducing power has a significant influence on butanol yield. This finding created a research hotspot centered around improving reducing power endogenously by limiting hydrogenases (hydA) (Meyer and Papoutsakis 1989).
Hydrogenases (H2ases) are metalloenzymes that contain iron-sulfur clusters and catalyze the reversible oxidation of molecular hydrogen (2 H+ + 2e− ↔ H2). They also play a critical role in microbial energy metabolism (Vignais et al. 2001) and are classified into three types: [NiFe]-H2ases, [FeFe]-H2ases, and [Fe]-H2ases. The classification and structural and functional relationships of H2ases and their subunits has been discussed in detail by Vignais (2008). [FeFe]-hydrogenases are found in anaerobic prokaryotes, such as clostridia and sulfate reducers, and are generally associated with the formation of H2. In C. beijerinckii, several H2ases have been found using genomic sequencing and have been annotated differently by various researchers, which mostly named and classified them by sequence alignment. Indeed, these H2ases have similar components: 350 residues containing the active site (H-cluster), iron hydrogenase small subunit (Fe_hyd_SSU), iron-only hydrogenase large subunit in the C-terminal domain (Fe_hyd_lg_C), NADH-ubiquinone oxidoreductase-G iron-sulfur binding region (NADH-G_4Fe-4S_3), etc. (Nicolet et al. 2002; Nicolet et al. 2000; Peters 1999; Peters et al. 1998). However, the functions of these H2ases are entirely different.
In this study, we regulated the reducing power by inactivation of two kinds of H2ases. Cbei_4110, according to the KEGG pathways, belongs to oxidative phosphorylation of energy metabolism and encodes NADH-quinone oxidoreductase subunit G (nuoG). NADH-ubiquinone oxidoreductase is a membrane-bound enzyme that is relevant to energy conversion and is called complex I in mitochondria or NDH-1 in bacteria (Yagi 1993; Yagi et al. 1998). Complex I (or NDH-1) is L-shaped with a membrane domain and a peripheral arm extending into the cytosol in which oxidate NADH to NAD+, proton transfer to periplasm occurs, forming a proton gradient between the periplasm and the cytoplasm (Thauer et al. 1977) In mitochondria, complex I couples with ATP synthesis by a proton pump or ATPase (Abrahams et al. 1994; Blair et al. 1996; Boyer 1993). This is described as the chemiosmotic hypothesis, but whether this coupling exists in C. beijerinckii is unclear.
Studies on decreasing hydA activities in order to improve the reducing power level have been carried out (Cooksley et al. 2012; Grupe and Gottschalk 1992; Jang et al. 2014; Liu et al. 2015; Meyer et al. 1986). However, most of them failed to disrupt hydA or lower cell activity and solvent yield after limited hydA activities. It is controversial in C. beijerinckii that which H2ase controls H2 formation. Gene Cbei_4110 was once recognized as the hydA gene (Zhao et al. 2012) which was proved incorrect in this study. Another gene, Cbei_3796 was identified by gene blast and functional domain analysis, was considered encoding the hydA which directly catalyzes H2 formation in C. beijerinckii. Genetic inactivation of gene Cbei_3796 in C. beijerinckii NCIMB 8052 was performed in our laboratory by using ClosTron technology and no mutants were obtained, either (data not shown). In conclusion, we believe that gene Cbei_3796 encodes the hydA and it seems to be an essential gene in C. beijerinckii NCIMB 8052.
H2ases share not only sequence similarities but also have similar evolutionary links involved in electron transfer subunits and ion-pumping units (Vignais 2008). Therefore, distinction of the function of H2ases should not totally rely on sequence alignment. The nuoG-inactivated strain showed a remarkable increase in reducing power and solvent yield with slightly increased H2 yield instead of H2 releasing decrease. It has been reported that high NAD(P)H levels accompany high butanol yield (Meyer and Papoutsakis 1989; Ventura et al. 2013) since a good deal of reducing power is required in butyry-CoA and butyraldehyde reduction (Fig. 4). In the current study, the reducing power in the acidogenic phase was consistently similar in the two strains investigated. This may be because the redox potential stays at rock bottom where exceeding reducing power would turn to H2 and be released by catalyzing of hydA. In addition, the amount of H2 generated in the fermentation of 8052-MU and 8052-WT was 38 and 35 % of the total gas generated in fermentation, respectively (data not shown). These results indicated that Cbei_4110 is part of NADH-quinone oxidoreductase rather than hydA.
Acetate and butyrate concentrations in batch fermentation by 8052-MU strains were remarkably different than that observed in the control strain (Fig. 5). Apart from the faster accumulation rate (discussed later), the acetate/butyrate ratio in 8052-MU was 0.35:0.85 (mol). The transcriptome data was consistent with the fermentation results in which acetate formation genes (pta and ack) were downregulated while the expression level of butyrate formation genes (etfA/B, bdc, crt, ptb, and buk) were generally upregulated. During the initial growth phase, glucose catabolizes to pyruvate, which turns to acetate and butyrate resulting in generation of ATP. Theoretically, conversion 1 mol of glucose to acetate would yield 4 mol of ATP and 2 mol NADH whereas only 3 mol of ATP would be generated if 1 mol of glucose was fermented completely to butyrate (Jones and Woods 1986). Although formation of acetate is more efficient in energy supply, usually the acetate/butyrate ratio is 0.5:0.75 (mol) because of intracellular redox balance (Jones and Woods 1986; Liu et al. 2015). After inactivation of nuoG, maintaining the redox balance of 8052-MU is now heavily dependent on hydA. Thus, regulation by improving butyrate occupancy seems to be a supplementary role for hydA. Final acid concentrations also showed an obvious decline in 8052-MU strains compared with the 8052-WT strain. Apparently, adequate reducing power plays a crucial role in this phenomenon. The acetone concentration of these two strains was also consistently similar. Although previous studies showed that acetone production was directly proportional to butanol formation, recent evidence seems to indicate that the acetone formation has no direct impact on butanol yield (Scheel and Lutke-Eversloh 2013; Sillers et al. 2009; Wietzke and Bahl 2012). Instead, it is the supply of NAD(P)H that directly affects the acetone/butanol ratio and butanol yield. Carbon flow is directed towards butanol in sufficient NAD(P)H conditions; however, carbon flow goes towards acetone when reducing power is in short supply (Liu et al. 2013). Conversely, the reassimilation of acetate and butyrate is directly coupled to the production of acetone, and disrupting the acetone formation pathway leads to low butanol yield (Jiang et al. 2009; Kuit et al. 2012; Lee et al. 2008).
ATP, as the currency of energy, catalyzes various biosynthesis processes. The ATP levels of the mutant strain was significantly enhanced, which was consistent with the findings of Meyer and Papoutsakis (1989) who carried out glucose-limited and glucose-sufficient steady-state cultures and observed a five to eight times difference in ATP levels. This phenomenon also correlated with the transcriptome analysis results in the acidogenic phase. However, the improvement contradicts the findings in the solventogenic phases (Fig. 3). In C. beijerinckii NCIMB 8052, ATP is generated mainly by substrate phosphorylation in glycolysis (Thauer et al. 1977). After membrane electron transport chain destruction, proton levels inside or outside the cell membrane remain balanced resulting in ATP synthesis being unavailable. Interestingly, the ATP yield generated by only substrate phosphorylation increased instead of being decreased. The fermentation data showed that the mutant strains had faster cell growth and sugar consumption rates (Fig. 1). This result is contradictory to other studies that limited hydrogen spillover in order to improve reduction potential (Grupe and Gottschalk 1992; Liu et al. 2013; Meyer et al. 1986). The transcriptome sequencing data revealed that sugar transporter and glycolytic genes were highly expressed in the acid formation period. Thus, it seemed that 8052-MU supplied more ATP by improving sugar consumption, yet the mechanism of it is still not clear.
In this study, we provided a new strategy for maintaining high levels of reducing power in C. beijerinckii NCIMB 8052. The nuoG inactive mutant strains demonstrated significantly enhanced reducing force, glucose utilization, and butanol yield compared to the wild-type strain. The fermentation results showed that this strategy does not affect cell growth and metabolism but actually significantly enhances butanol production. In conclusion, this strategy may be extended for making industrial bio-butanol more economically attractive.
References
Abrahams JP, Leslie A, Lutter R, Walker JE (1994) Structure at 2.8 A resolution of F1-ATPase from bovine heart mitochondria. Nature: 621–8.
Audic S, Claverie JM (1997) The significance of digital gene expression profiles. Genome Res 7(10):986–995
Baer SH, Hans PB, Smith TL (1987) Effect of butanol challenge and temperature on lipid composition and membrane fluidity of butanol-tolerant Clostridium acetobutylicum. Appl Environ Microb 53(12):2854–2861
Barabote RD, Saier MH (2005) Comparative genomic analyses of the bacterial phosphotransferase system. Microbiol Mol Biol R 69(4):608–634
Bernofsky C, Swan M (1973) An improved cycling assay for nicotinamide adenine dinucleotide. Analytical Biochemistry. 53(2):452–458
Blair A, Ngo L, Park J, Paulsen IT, Saier MH (1996) Phylogenetic analyses of the homologous transmembrane channel-forming proteins of the F0F1-ATPases of bacteria, chloroplasts and mitochondria. Microbiology 142(1):17–32
Boyer PD (1993) The binding change mechanism for ATP synthase—some probabilities and possibilities. BBA-Bioenergetics 1140(3):215–250
Cooksley CM, Zhang Y, Wang H, Redl S, Winzer K, Minton NP (2012) Targeted mutagenesis of the Clostridium acetobutylicum acetone-butanol-ethanol fermentation pathway. Metab Eng 14:630–641
Ezeji TC, Qureshi N, Blaschek HP (2007) Bioproduction of butanol from biomass: from genes to bioreactors. Curr Opin Biotech 18(3):220–227
Gao H, Nielson F, Nielson HR (2011) Analysing protocol stacks for services. In: Wirsing M, Hölzl M (eds) Rigorous software engineering for service-oriented systems. Springer, Berlin Heidelberg, Berlin, Heidelberg, pp. 369–389
Green EM (2011) Fermentative production of butanol-the industrial perspective. Curr Opin Biotech. 22(3):337–343
Grupe H, Gottschalk G (1992) Physiological events in Clostridium acetobutylicum during the shift from acidogenesis to solventogenesis in continuous culture and presentation of a model for shift induction. Appl Environ Microb 58(12):3896–3902
Guo T, Tang Y, Zhang QY, Du TF, Liang DF, Jiang M, Ouyang PK (2012) Clostridium beijerinckii mutant with high inhibitor tolerance obtained by low-energy ion implantation. J Ind Microbiol Biot 39(3):401–407
Heap JT, Pennington OJ, Cartman ST, Carter GP, Minton NP (2007) The ClosTron: a universal gene knock-out system for the genus Clostridium. J Microbiol Meth 70:452–464
Heap JT, Pennington OJ, Cartman ST, Minton NP (2009) A modular system for Clostridium shuttle plasmids. J Microbiol Meth 78(1):79–85
Jang YS, Im JA, Choi SY, Lee JI, Lee SY (2014) Metabolic engineering of Clostridium acetobutylicum for butyric acid production with high butyric acid selectivity. Metab Eng 23:165–174
Jiang Y, Xu C, Dong F, Yang Y, Jiang W, Yang S (2009) Disruption of the acetoacetate decarboxylase gene in solvent-producing Clostridium acetobutylicum increases the butanol ratio. Metab Eng 11:284–291
Jones DT, Woods DR (1986) Acetone-butanol fermentation revisited. Microbiol Rev 50(4):484
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36(suppl 1):D480–D484
Kuit W, Minton NP, López-Contreras AM, Eggink G (2012) Disruption of the acetate kinase (ack) gene of Clostridium acetobutylicum results in delayed acetate production. Appl Microbiol Biotechnol 94(3):729–741
Lee J, Blaschek HP (2001) Glucose uptake in Clostridium beijerinckii NCIMB 8052 and the solvent-hyperproducing mutant BA101. Appl Environ Microb 67(11):5025–5031
Lee SY, Park JH, Jang SH, Nielsen LK, Kim J, Jung KS (2008) Fermentative butanol production by Clostridia. Biotechnol Bioeng 101(2):209–228
Li R, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K, Wang J (2009) SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25(15):1966–1967
Lin Y, Tanaka S (2006) Ethanol fermentation from biomass resources: current state and prospects. Appl Microbiology Biot 69(6):627–642
Liu D, Chen Y, Ding F, Guo T, Xie J, Zhuang W, Niu H, Shi X, Zhu C, Ying H (2015) Simultaneous production of butanol and acetoin by metabolically engineered Clostridium acetobutylicum. Metab Eng 27:107–114
Liu D, Chen Y, Li A, Ding F, Zhou T, He Y, Li B, Niu H, Lin X, Xie J, Chen Y, Wu J, Ying H (2013) Enhanced butanol production by modulation of electron flow in Clostridium acetobutylicum B3 immobilized by surface adsorption. Bioresource Technol 129:321–328
Meyer CL, Papoutsakis ET (1989) Increased levels of ATP and NADH are associated with increased solvent production in continuous cultures of Clostridium acetobutylicum. Appl Microbiol Biotechnol 30(5):450–459
Meyer CL, Roos JW, Papoutsakis ET (1986) Carbon monoxide gasing leads to alcohol production and butyrate uptake without acetone formation in continuous cultures of Clostridium acetobutylicum. Appl Microbiol Biotechnol 24(2):159–167
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7):621–628
Nicolet Y, Cavazza C, Fontecilla-Camps J (2002) Fe-only hydrogenases: structure, function and evolution. J Inorg Biochem 91(1):1–8
Nicolet Y, Lemon BJ, Fontecilla-Camps JC, Peters JW (2000) A novel FeS cluster in Fe-only hydrogenases. Trends Biochem Sci 25(3):138–143
Perutka J, Wang W, Goerlitz D, Lambowitz AM (2004) Use of computer-designed group II introns to disrupt Escherichia coli DExH/D-box protein and DNA helicase genes. J Mol Biol 336(2):421–439
Peters JW (1999) Structure and mechanism of iron-only hydrogenases. Curr Opin Struc Biol 9(6):670–676
Peters JW, Lanzilotta WN, Lemon BJ, Seefeldt LC (1998) X-ray crystal structure of the Fe-only hydrogenase (CpI) from Clostridium pasteurianum to 1.8 angstrom resolution. Science 282(5395):1853–1858
Qureshi N, Ezeji TC, Ebener J, Dien BS, Cotta MA, Blaschek HP (2008) Butanol production by Clostridium beijerinckii. Part I: use of acid and enzyme hydrolyzed corn fiber. Bioresource Technol 99(13): 5915–5922.
Scheel M, Lutke-Eversloh T (2013) New options to engineer biofuel microbes: development and application of a high-throughput screening system. Metab Eng 17:51–58
Sillers R, Al-Hinai MA, Papoutsakis ET (2009) Aldehyde-alcohol dehydrogenase and/or thiolase overexpression coupled with CoA transferase downregulation lead to higher alcohol titers and selectivity in Clostridium acetobutylicum fermentations. Biotechnol Bioeng 102(1):38–49
Thauer RK, Jungermann K, Decker K (1977) Energy conservation in chemotrophic anaerobic bacteria. Bacteriol Rev 41(1):100–180
Tracy BP (2012) Improving butanol fermentation to enter the advanced biofuel market. mBio 3(6): e00518–512.
Ventura JR, Hu H, Jahng D (2013) Enhanced butanol production in Clostridium acetobutylicum ATCC 824 by double overexpression of 6-phosphofructokinase and pyruvate kinase genes. Appl Microbiol Biotechnol 97(16):7505–7516
Vignais PM (2008) Hydrogenases and H+-reduction in primary energy conservation. In: Schäfer G, Penefsky H (eds) Bioenergetics. Springer, Berlin Heidelberg, Berlin, Heidelberg, pp. 223–252
Vignais PM, Billoud B, Meyer J (2001) Classification and phylogeny of hydrogenases. FEMS Microbiol Rev 25(4):455–501
Wang Y, Li X, Mao Y, Blaschek HP (2012) Genome-wide dynamic transcriptional profiling in Clostridium beijerinckii NCIMB 8052 using single-nucleotide resolution RNA-Seq. BMC Genomics 13(1):102
Wietzke M, Bahl H (2012) The redox-sensing protein Rex, a transcriptional regulator of solventogenesis in Clostridium acetobutylicum. Appl Microbiology Biot 96(3):749–761
Xiao H, Gu Y, Ning Y, Yang Y, Mitchell WJ, Jiang W, Yang S (2011) Confirmation and elimination of xylose metabolism bottlenecks in glucose phosphoenolpyruvate-dependent phosphotransferase system-deficient Clostridium acetobutylicum for simultaneous utilization of glucose, xylose, and arabinose. Appl Environ Microb 77(22):7886–7895
Xiao H, Li Z, Jiang Y, Yang Y, Jiang W, Gu Y, Yang S (2012) Metabolic engineering of D-xylose pathway in Clostridium beijerinckii to optimize solvent production from xylose mother liquid. Metab Eng 14:569–578
Yagi T (1993) The bacterial energy-transducing NADH-quinone oxidoreductases. BBA-Bioenergetics 1141(1):1–17
Yagi T, Yano T, Di Bernardo S, Matsuno-Yagi A (1998) Procaryotic complex I (NDH-1), an overview. BBA-Bioenergetics 1364(2):125–133
Zhao X, Xing D, Liu B, Lu L, Zhao J, Ren N (2012) The effects of metal ions and L-cysteine on hydA gene expression and hydrogen production by Clostridium beijerinckii RZF-1108. Int J Hydrog Energy 37(18):13711–13717
Acknowledgments
The authors would like to thank Prof. Nigel P. Minton and Michelle L. Kelly from the University of Nottingham, UK, for kindly providing the ClosTron plasmids and Prof. Sheng Yang from the Shanghai Institutes for Biological Sciences for kindly providing the pWJ1 plasmid.
This work was supported by the National Basic Research Program of China (973 Program, 2013CB733602), the restructured institutions innovation capacity of special funds of Ministry of Science and Technology of China (Grant No. 2014EG111227), the National Natural Science Foundation of China (Grant No. 21306032), the National High-Tech Research and Development Program of China (863) (Grant No. 2012AA021200), the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD), Postdoctoral Science Foundation of Jiangsu Province (Grant No. 1302107C), the Major Research Plan of the National Natural Science Foundation of China (21390204), Program for Changjiang Scholars and Innovative Research Team in University (Grant No.: IRT_14R28), and Research Foundation of Guangzhou Sugarcane Industry Research Institute (Grant No. A201302).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
This work is in compliance with ethical standards.
Conflict of Interest
The authors declare that they have no competing interests.
Human and animals studies
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Jun Liu, Ting Guo and Dong Wang contributed equally to this work.
Electronic supplementary materials
ESM 1
(PDF 286 kb)
Rights and permissions
About this article
Cite this article
Liu, J., Guo, T., Wang, D. et al. Enhanced butanol production by increasing NADH and ATP levels in Clostridium beijerinckii NCIMB 8052 by insertional inactivation of Cbei_4110. Appl Microbiol Biotechnol 100, 4985–4996 (2016). https://doi.org/10.1007/s00253-016-7299-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7299-9