Abstract
Anaerobic digestion is a very complex process that is mediated by various microorganisms, and the understanding of the microbial community assembly and its corresponding function is critical in order to better control the anaerobic process. The present study investigated the effect of different inocula on the microbial community assembly in biogas reactors treating cellulose with various inocula, and three parallel biogas reactors with the same inoculum were also operated in order to reveal the reproducibility of both microbial communities and functions of the biogas reactors. The results showed that the biogas production, volatile fatty acid (VFA) concentrations, and pH were different for the biogas reactors with different inocula, and different steady-state microbial community patterns were also obtained in different biogas reactors as reflected by Bray-Curtis similarity matrices and taxonomic classification. It indicated that inoculum played an important role in shaping the microbial communities of biogas reactor in the present study, and the microbial community assembly in biogas reactor did not follow the niche-based ecology theory. Furthermore, it was found that the microbial communities and reactor performances of parallel biogas reactors with the same inoculum were different, which could be explained by the neutral-based ecology theory and stochastic factors should played important roles in the microbial community assembly in the biogas reactors. The Bray-Curtis similarity matrices analysis suggested that inoculum affected more on the microbial community assembly compared to stochastic factors, since the samples with different inocula had lower similarity (10–20 %) compared to the samples from the parallel biogas reactors (30 %).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Anaerobic digestion for biogas production has been widely used in the treatment of organic wastes, and it is a very complex microbial process involving different microorganisms (Werner et al. 2011). The degradation of organic wastes includes four sequential steps: hydrolysis (conversion of organic wastes into oligomers and monomers), fermentation (conversion of oligomers and monomers into acetate, CO2, and H2), acetogenesis (conversion of CO2 and H2 to acetate), and methanogenesis (conversion of CO2, H2, and acetate into methane) (Schluter et al. 2008). The first three steps are mediated by bacteria, while the last step is mediated by archaea (also known as methanogens). The syntrophic relationship between bacteria and archaea is essential for the efficiency and stability of the biogas process (Luo et al. 2015).
The understanding of microbial community assembly in biogas reactors is crucial for effective biodiversity management and maintenance. Ecologists have studied the microbial community assembly in different ecosystems, and now it is known that deterministic factors (such as environmental selection and interspecies interactions) and stochastic factors (stochastic processes of birth, death, colonization, extinction, and speciation) affect microbial community assembly (Pagaling et al. 2014; Zhou et al. 2013). However, the relative importance of these factors is still unclear. There are controversial views on the microbial community assembly. The traditional niche-based theory supports the view that the community is shaped mainly by deterministic factors and thereby asserts that community composition should converge toward a single pattern under similar environmental conditions from diverse starting species sets (Fargione et al. 2003). The view has been demonstrated in several ecosystems including activated sludge bioreactors treating wastewater (Ayarza and Erijman 2011) and sediment slurry microcosms (Kurtz et al. 1998). On the contrary, the neutral-based theory assumes that many natural different community patterns can be generated under similar environmental conditions by stochastic factors, which was demonstrated in several ecosystems including microbiomes from microbial fuel cells, wetlands et al. (Baptista et al. 2008; Chave 2004; Zhou et al. 2013). The stochastic factors are impossible to manipulate or reconstruct in the ecosystems (Chase 2003).
The microbial community in biogas reactors has been studied for several decades, and our understanding of the microbiomes in biogas reactors has been increased greatly with the establishment of culture-independent molecular methods including PCR-DGGE, PCR-cloning, and the recently developed PCR-high-throughput sequencing (Luo and Angelidaki 2014; Luo et al. 2013; Riviere et al. 2009; Sundberg et al. 2013). The roles of predictable factors including temperature, reactor configuration, feedstock et al., on the microbial communities in biogas reactors have been studied previously (Nielsen et al. 2004; Rincón et al. 2008). However, the microbial community assembly in biogas reactors is still not well exploited based on the ecological theories. The importance of inoculum sources on the biogas production has been demonstrated previously in batch experiments treating different organic wastes, and the suitable inoculum sources are essential for efficient biogas production (Dhamodharan et al. 2015; Gu et al. 2014; Lopes et al. 2004). However, inoculum sources should not be important for biogas production based on the niche-based theory, where similar microbial communities should be shaped under similar environmental conditions from diverse inoculum sources. It should be noted that all the abovementioned studies were conducted under batch conditions (lasting for around 1 month), and stable microbial communities were not achieved. Therefore, it is not appropriate to evaluate the importance of inoculum source for biogas production only based on the results from batch experiments, and the role of inoculum in shaping the microbial communities in biogas reactors remains to be elucidated.
Based on the above considerations, the present study aims to reveal the role of inoculum sources in shaping the microbial communities in biogas reactors. Five different inocula were obtained from different sources and inoculated in continuously running biogas reactors with the same operational conditions. In addition, one of the above inocula was inoculated into three parallel biogas reactors in order to see the reproducibility of both microbial communities and functions of the biogas reactors. The steady-state microbial communities in the biogas reactors were analyzed by 16S rRNA genes high-throughput sequencing.
Material and methods
Inoculum and feed composition
The inocula used in the present study were obtained from five different sources, including the digested stillage from mesophilic continuously stirred tank reactor (CSTR) treating stillage in an ethanol plant (pH 7.1, total solid (TS) 23.2 g/L, volatile solid (VS) 16.5 g/L), the anaerobic granular sludge from a mesophilic up-flow anaerobic sludge blanket (UASB) reactor treating paper mill wastewater (pH 7.2, TS 72.8 g/L, VS 43.5 g/L), the digested manure from mesophilic CSTR reactor treating cattle manure (pH 7.8, TS 35.2 g/L, VS 20.6 g/L), the raw cattle manure (pH 7.5, TS 65.4 g/L, VS 44.3 g/L), and the digested sewage sludge from mesophilic CSTR reactor in a wastewater treatment plant (pH 7.2, TS 24.5 g/L, VS 13.7 g/L). The feed was composed of cellulose and basic anaerobic (BA) medium. The cellulose concentration in the feed was 7.5 g/L. The BA medium was prepared based on a previous literature (Angelidaki and Sanders 2004). The NaHCO3 concentration in the feed was 5 g/L. The feed was sterilized in an autoclave to reduce the influence of microorganisms that might be grown in the non-sterilized feed.
Experimental setup
Seven identical 2 L CSTR reactors with working volume of 1.8 L were used in the present study, and all the reactors were continuously stirred by magnetic stirrer at a stirring speed of 200 rpm to achieve well mixing of the reactors. All the reactors were equipped with thermal jackets in order to maintain a temperature of 37 °C. The produced biogas was collected by gas bags. The HRT of the reactors was 15 days. Each inoculum was added to one CSTR reactor except for digested manure, which was added to three parallel CSTR reactors. All the reactors were initially filled with the inoculum, BA medium, and feed (200 mL), and the VS concentration of inoculum in all the reactors was 5 g/L. In the beginning, all the reactors were run in batch condition without feeding, and the continuous feeding was started once the biogas production from all the reactors achieved maximum values. The evolved biogas was collected with gas bag, and the amount of biogas was determined periodically using syringe. The parameters such as biogas production, volatile fatty acid (VFA) concentration, and pH were monitored during the whole operational phase. The steady state in this study was defined as a stable biogas production with daily variation of lower than 10 % for a period at least 10 days. Liquid samples from all the reactors were collected once the steady-state conditions were reached in all the reactors. The reactors were named as 1 (inoculated by digested stillage), 2 (inoculated by anaerobic granular sludge), 3a (inoculated by digested manure), 3b (inoculated by digested manure), 3c (inoculated by digested manure), 4(inoculated by raw cattle manure), and 5 (inoculated by digested sewage sludge).
DNA extraction, 16S rRNA sequencing, and analysis
Twelve microbial samples were obtained including five inocula and seven liquid samples from all the biogas reactors in steady states. All the samples were centrifuged at 10,000 rpm for 10 min, and the resulting supernatant was discarded. The solid residues were used for DNA extraction. Total genomic DNA of the collected samples was extracted using QIAamp DNA Stool Mini Kit (QIAGEN 51504) according to the manufacturer’s instructions. The extracted DNA was PCR amplified with the universal primers 515f and 806r targeting the V4 region of the 16S rRNA gene according to previous studies (Bates et al. 2011; Regueiro et al. 2015), and the PCR products were purified using the QIAquick spin columns (QIAGEN) to remove the excess primer dimers and dNTPs. The concentration of PCR amplicons was measured by NanoDrop spectrophotometer. The PCR products were then sent out for sequencing on an Ion Torrent PGM machine at Shanghai Shenggong. The raw sequences were quality-checked by removing the low-quality sequences without exact matches to the forward and reverse primers, with length shorter than 100 bp, and containing any ambiguous base calls by RDP tools (Cole et al. 2009). Chimeras were removed from the data by using the Find Chimeras web tool. The numbers of high-quality sequences are shown in Table 1 with an average length around 265 bp. The sequences were then normalized to the same sequencing depths (20,000 sequences) by MOTHUR program to facilitate the comparison. The normalized sequences were phylogenetically assigned to taxonomic classifications by RDP Classifier with a confidence threshold of 50 %. The sequences were clustered into operational taxonomic units (OTU) by setting a 0.03 distance limit by MOTHUR program. Rarefaction curve, Shannon diversity index, species richness estimator of Chao1, and dendrogram based on Bray-Curtis similarity matrix were also generated by MOTHUR program. All the sequence data were submitted to NCBI (Bioproject PRJNA293546).
Analytical methods
The concentrations of acetate, propionate, isobutyrate, butyrate, iso-valerate, and valerate were determined by HPLC, and they were separated with a 7.8 × 300 Aminex HPX-87-H column (Bio-Rad) at 55 °C with a refractive index detector at 50 °C. The mobile phase was 5 mmol H2SO4, at a flow rate of 0.4 mL/min. The biogas composition was analyzed by GC-TCD equipped with a 2-m stainless column packed with Porapak T (50/80mesh). Helium was used as carrier gas. The operational temperatures of the injection port, the oven, and the detector were 120, 120, and 110 °C, respectively. TS, VS, and COD were analyzed according to APHA (APHA 1995).
Results
Reactor performances
All the biogas reactors were operated for 8 months, and steady-state reactor performances were obtained in all the reactors. The steady-state reactor performances were summarized in Table 2. The biogas production from the seven reactors was different. Reactors 1 and 5 had relatively lower biogas production (below 320 mL/day) compared to the other reactors (all above 450 mL/day). The lower biogas production in reactors 1 and 5 also corresponded to the lower methane content, pH, and COD removal efficiencies. Acetate and propionate were the main constituents in volatile fatty acids (VFA). The higher acetate and propionate concentrations in reactors 1 and 5 showed that the microorganisms in reactors 1 and 5 could not effectively convert the organics into methane. Reactors 3a, 3b, and 3c were all inoculated with the same digested manure; however, the steady-state reactor performances were not the same. The biogas production in reactors 3a and 3c was significantly higher than that in reactor 3b. Lower methane content, pH, and COD removal efficiency were also observed in reactor 3b compared to reactors 3a and 3c. The methane yields from all the reactors were shown in Fig. 1. The theoretical methane yield was calculated to be 413 mL CH4/g cellulose, and the higher methane yields were obtained from reactors 3a, 3c, and 4, which were around 86 % of the theoretical value.
Diversity of the microbial communities
Both the inocula and the steady-state microbial communities of all the reactors were analyzed by the high-throughput sequencing of the 16S rRNA genes. The obtained sequences were quality checked and normalized to 20,000 sequences for the following analysis. The parameters related to the diversity of the microbial communities are shown in Table 3. All the inocula had higher species richness compared to the samples from steady-state reactors as reflected by the higher numbers of OTUs and CHAO1. The Shannon diversity index, which provides both species richness and evenness of the species among all the species in the community (Lu et al. 2012), also showed that the diversity of microbial communities in the inocula was higher than the samples from the biogas reactors. The rarefaction curves of all the samples at 0.03 distance are shown in Fig. 2. For samples 2I and 5I, the numbers of OTUs tended to increase faster compared to the other samples, which suggested that the sequencing depth of 20,000 was still not enough to cover the whole diversity. However, the higher coverage values (80 %) for all the samples clearly indicated that the dominant OTUs were detected in the present study.
Structure of the microbial communities
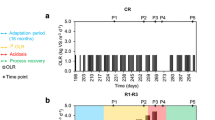
The dendrograms from Bray-Curtis similarity matrices that took into account the abundance of sequences in each OTU were generated to assess the differences among all the samples, and the result is shown in Fig. 3. It was obvious that the microbial community of the sample from each biogas reactor and its corresponding inoculum were clustered together (except 5 and 5I), although the similarity (between 10 and 20 %) was not very high. The phylogenetic classification of the sequences was conducted by Ribosomal Database Project classifier, and both bacteria and archaea classifications were obtained since universal primers for PCR were used in the present study. Figure 4 shows the phylogenetic classification of sequences assigned to bacteria. Different distributions of the sequences at phylum level were observed for the five different inocula. For example, Proteobacteria were found to be dominant in 2I and 4I, while Firmicutes were dominant in 1I, 3I, and 5I. The different phylum level distributions of the inocula resulted in the different similarity as shown in Fig. 3. For the samples obtained in the present study from the biogas reactors, the dominant phyla were Firmicutes and Bacteroidetes. The higher abundance of Firmicutes in 3a compared to 3b and 3c resulted in the different distribution of microorganism in the samples from parallel biogas reactors, which also supported the observation in Fig. 4. Standing in the class level, it was further found that the relative abundance of Clostridia, belonging to Firmicutes, were usually higher in the samples from biogas reactors compared to the inocula.
The order level identification of the sequences belonging to archaea is shown in Fig. 5. The sequences were mainly assigned to three orders (Methanobacteriales, Methanomirobiales, and Methanosarcinales) mediating hydrogenotrophic and aceticlastic methanogenesis. Different order level distributions of the sequences obtained from all the inocula were observed. Only Methanobacteriales and Methanomirobiales were detected in 1I, while the three orders were all detected in 2I, 3I, 4I, and 5I though their relative abundances were different in each sample. The samples from the biogas reactors also had different order level distributions of the sequences, and Methanosarcinales belonging to aceticlastic methanogens were found in all the samples. Methanomicrobiales were absent in sample 4 but present in all the other samples from the biogas reactors. Samples 3a, 3b, and 3c had the same dominant orders (Methanobacteriales, Methanomirobiales, and Methanosarcinales) and genus (Methanosarcina, Methanoculleus, and Methanobacterium) but with different relative abundances, which also might contribute to their difference in Fig. 3.
Discussion
The results obtained from the performances of the long-term operated continuous reactors indicated that the inoculum had significant effects on the biogas production from cellulose, and different reactor performances can be obtained with the same inoculum and operational conditions. The differences between actual and theoretical methane yields (Fig. 1) were due to the organics left in the effluent, which included unconverted VFA, residual cellulose, and also the produced biomass.
The microbial community analysis showed that the inocula generally had higher diversity than the samples from the biogas reactors, and it could be due to that the microbial communities accumulated in the biogas reactors are special for the anaerobic degradation of pure cellulose, while the inocula had more diverse microorganisms to degrade complex substrates including stillage, manure, paper milling wastewater, et al. (Sundberg et al. 2013).
The Bray-Curtis similarity analysis showed that inoculum played an important role in shaping the microbial communities in biogas reactors treating cellulose since the sample from each biogas reactor was clustered together with its corresponding inoculum. Samples 3a, 3b, and 3c were obtained from parallel biogas reactors; however, they only had around 30 % similarity, and it could be related to the different reactor performances as mentioned before. The results also showed that different microbial community patterns could be generated under similar environmental conditions. Although reactors 2, 3a, 3b, 3c, and 4 had relatively higher methane yields compared to the other biogas reactors, the microbial communities only had 5 % similarity. It suggested that the efficient biogas production from cellulose could be obtained with different microbial community patterns. Sample 5I (raw cattle manure) had the lowest similarity to all the other samples, which was expected since all the other samples were obtained from biogas reactors treating different feedstock.
The different phylum level distributions of the inocula showed that the inocula chosen in the present study had different microbial community compositions, which ensured the possibility to elucidate the general relationship of microbial communities between various inocula and their final accumulated microbial communities. Firmicutes and Bacteroidetes were dominant in all the samples, and they can degrade a wide range of complex organics including proteins and carbohydrates (Sundberg et al. 2013), and their dominance was also reported in several studies focusing on biogas production (Luo et al. 2015; Riviere et al. 2009). The class Clostridia was found in all the reactors, and it is a highly versatile class that could degrade both carbohydrates and proteins, and their dominance was found in biogas reactors treating both manure and sewage sludge (Lee et al. 2008; Schluter et al. 2008; Sundberg et al. 2013). The enrichment of Clostridia in the present study might be related with their capacity to hydrolyze cellulose substrates and further produce volatile fatty acids and alcohols (Krakat et al. 2011). Thermotogae, which was mainly found in 1I and 1, can utilize different complex carbohydrates to produce hydrogen and volatile fatty acids (Nguyen et al. 2008). They were thought to be thermophilic microorganisms; however, recently, some Thermotogae existing in mesophilic temperatures has been identified (Nesbø et al. 2010). Their dominance in reactor 1 may be related to their existence in the inoculum, which further showed that inoculum had considerable effects in shaping the microbial communities of the biogas reactors.
The results from the analysis of archaea showed that Methanosarcinales were found in sample 1 but not in the inoculum 1I, which indicated different microbial communities could be formed when the cultivation conditions were changed. The lower relative abundances of Methanosarcinales (<30 %) in reactors 1 and 5 could be related with the higher concentrations of VFA and lower pH, and it resulted in the dominance of syntrophic acetic acid oxidation and higher relative abundance of hydrogenotrophic methanogens (Fotidis et al. 2013; Fotidis et al. 2014). Methanobacteriales had a high relative abundance (85 %) in sample 4 that indicated syntrophic acetic acid oxidation occurred in reactor 4, which was not expected since reactor 4 was not running in an extreme conditions such as high ammonia or VFA concentration (Lü et al. 2013). The result might suggest syntrophic acetic acid oxidation could be present in biogas reactors running in smooth conditions, which deserved further investigation.
The present study clearly showed that inoculum had an important role in shaping the microbial community as well as their functions. Previous studies also showed that inoculum affected the biogas production from organic wastes in batch experiments (Dhamodharan et al. 2015; Gu et al. 2014; Lopes et al. 2004), which was consistent with the results from continuous experiments in the present study. However, our study, for the first time, not only investigated the effects of various inocula on the biogas production from cellulose in long-term operated biogas reactors but also made the microbial community analysis. Karakashev et al. (2005) demonstrated that inoculum population appeared to have no influence on the eventual population of the manure-digesting plants. However, we found that inoculum had considerable influence on the microbial community in the biogas reactor treating cellulose since the inocula and the accumulated samples in the biogas reactors were well clustered together as shown in Fig. 3. The difference might be resulted from the feedstock. In the study of Karakashev et al. (2005), manure was the feedstock for the biogas reactors, which were known to be rich in various microorganisms (also demonstrated in our study as shown in Fig. 4), and therefore the influence of the inoculum could be ignored after long-term operation. However, only sterilized cellulose was used in our study and therefore the initial microorganisms in the inoculum were crucial for the biogas production. Some previous studies showed that common environmental selection recruits the same or similar species from diverse starting species sets to produce similar final community structures, which follows the niche-based theory (Ayarza and Erijman 2011; Kurtz et al. 1998). However, all the biogas reactors in our study were operated under identical operational conditions (temperature, substrate, reactor configuration), and different final steady-state microbial communities with different functions (i.e., biogas production, VFA concentration et al.) were generated. It indicated that niche-based theory might not be applicable to the microbial community assembly in biogas reactors in the present study, and inoculum played an important role in the microbial community assembly in the biogas reactors. The results also suggested that for the wastes/wastewater (stillage, wastewater from papermaking plant) that lack microorganisms relating with anaerobic digestion, the selection of proper inoculum is important to get efficient biogas production, while it might not be important to get the proper inoculum for biogas production from the wastes including manure and sewage sludge, which already contained various microorganisms.
In addition, the present study also found that different microbial communities and reactor performances were obtained from the parallel biogas reactors 3a, 3b, and 3c, which were all operated under similar conditions (temperature, feedstock, reactor configuration, and even the inoculum). It could be explained by the neutral theory, where stochastic processes of birth, death, colonization, extinction, and speciation can lead to alternative communities with distinct functions under similar environmental conditions (Chave 2004). Zhou et al. (2013) demonstrated that the reactor performances (electricity, H2, pH, et al.) in parallel microbial electrolysis cells were different under the same operational conditions (temperature, substrate feeding, applied voltage, reactor structure, and materials), which was similar to our results. Another study found that four parallel biogas reactors with glucose as substrate, which were seeded with the inoculum already accumulated to glucose, had replication of function but without replication of community structures (Fernandez et al. 2000), which could be also related to the stochastic assembly of the microbial communities in the four parallel biogas reactors. Furthermore, they also observed a glucose-fed biogas reactor that experienced a shift in community structure with no detectable change in performance (Fernández et al. 1999). The above results showed that microbial community assembly in biogas process is very complex, and not only the inoculum source but also the stochastic factors played important roles in the microbial ecosystem development. The results had important practical implications for biogas research, and parallel biogas reactors should be operated in order to ensure the reliability of the data obtained, which was already applied in batch experiments for biogas production but not common in continuous biogas reactors. In our study, the inoculum seems to affect more on the microbial community assembly, since the samples with different inocula had lower similarity (10–20 %) compared to the samples (3a, 3b, and 3c) with the same inocula (30 %).
Overall, the present study investigated the microbial community assembly in biogas reactors treating cellulose with various inocula. Different reactor performances (biogas production, VFA concentration, and pH) were obtained from the reactors with different inocula, and it was consistent with the different microbial community patterns as reflected by Bray-Curtis similarity matrices and taxonomic classification. The results also showed that the microbial community assembly in biogas reactors did not follow the niche-based ecology theory, and inoculum played an important role in shaping the microbial communities in the present study. By comparison of the results from parallel reactors with the same inoculum, we further found that stochastic assembly contributed to the different microbial community patterns with distinct functions of the biogas reactors with similar operational conditions.
References
Angelidaki I, Sanders W (2004) Assessment of the anaerobic biodegradability of macropollutants. Reviews in Environ Sci Biotechnol 3(2):117–129. doi:10.1007/s11157-004-2502-3
APHA (1995) Standard methods for the examination of water and wastewater, 19th edn. American Public Health Association, New York, USA
Ayarza J, Erijman L (2011) Balance of neutral and deterministic components in the dynamics of activated sludge floc assembly. Microb Ecol 61(3):486–495. doi:10.1007/s00248-010-9762-y
Baptista JC, Davenport RJ, Donnelly T, Curtis TP (2008) The microbial diversity of laboratory-scale wetlands appears to be randomly assembled. Water Res 42(12):3182–3190. doi:10.1016/j.watres.2008.03.013
Bates ST, Berg-Lyons D, Caporaso JG, Walters WA, Knight R, Fierer N (2011) Examining the global distribution of dominant archaeal populations in soil. ISME J 5(5):908–917. doi:10.1038/ismej.2010.171
Chase J (2003) Community assembly: when should history matter? Oecologia 136(4):489–498. doi:10.1007/s00442-003-1311-7
Chave J (2004) Neutral theory and community ecology. Ecol Let 7(3):241–253. doi:10.1111/j.1461-0248.2003.00566.x
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucl Acids Res 37:D141–D145. doi:10.1093/nar/gkn879
Dhamodharan K, Kumar V, Kalamdhad AS (2015) Effect of different livestock dungs as inoculum on food waste anaerobic digestion and its kinetics. Bioresour Technol 180(0):237–241. doi:10.1016/j.biortech.2014.12.066
Fargione J, Brown C, Tilman D (2003) Community assembly and invasion: an experimental test of neutral versus niche processes. Proc Nat Acad Sci USA 100:8916–8920
Fernández A, Huang S, Seston S, Xing J, Hickey R, Criddle C, Tiedje J (1999) How stable is stable? Function versus community composition. Appl Environ Microbiol 65(8):3697–3704
Fernandez AS, Hashsham SA, Dollhopf SL, Raskin L, Glagoleva O, Dazzo FB, Hickey RF, Criddle CS, Tiedje JM (2000) Flexible community structure correlates with stable community function in methanogenic bioreactor communities perturbed by glucose. Appl Environ Microbiol 66(9):4058–4067. doi:10.1128/aem.66.9.4058-4067.2000
Fotidis IA, Karakashev D, Kotsopoulos TA, Martzopoulos GG, Angelidaki I (2013) Effect of ammonium and acetate on methanogenic pathway and methanogenic community composition. FEMS Microbiol Ecol 83(1):38–48. doi:10.1111/j.1574-6941.2012.01456.x
Fotidis IA, Wang H, Fiedel NR, Luo G, Karakashev DB, Angelidaki I (2014) Bioaugmentation as a solution to increase methane production from an ammonia-rich substrate. Environ Sci Technol 48(13):7669–7676. doi:10.1021/es5017075
Gu Y, Chen X, Liu Z, Zhou X, Zhang Y (2014) Effect of inoculum sources on the anaerobic digestion of rice straw. Bioresour Technol 158(0):149–155. doi:10.1016/j.biortech.2014.02.011
Karakashev D, Batstone DJ, Angelidaki I (2005) Influence of environmental conditions on methanogenic compositions in anaerobic biogas reactors. Appl Environ Microbiol 71(1):331–338. doi:10.1128/aem.71.1.331-338.2005
Krakat N, Schmidt S, Scherer P (2011) Potential impact of process parameters upon the bacterial diversity in the mesophilic anaerobic digestion of beet silage. Bioresour Technol 102(10):5692–5701. doi:10.1016/j.biortech.2011.02.108
Kurtz JC, Devereux R, Barkay T, Jonas RB (1998) Evaluation of sediment slurry microcosms for modeling microbial communities in estuarine sediments. Environ Toxicol Chem 17(7):1274–1281. doi:10.1002/etc.5620170712
Lü F, Hao L, Guan D, Qi Y, Shao L, He P (2013) Synergetic stress of acids and ammonium on the shift in the methanogenic pathways during thermophilic anaerobic digestion of organics. Water Res 47(7):2297–2306. doi:10.1016/j.watres.2013.01.049
Lee C, Kim J, Shin SG, Hwang S (2008) Monitoring bacterial and archaeal community shifts in a mesophilic anaerobic batch reactor treating a high-strength organic wastewater. FEMS Microbiol Ecol 65(3):544–554. doi:10.1111/j.1574-6941.2008.00530.x
Lopes WS, Leite VD, Prasad S (2004) Influence of inoculum on performance of anaerobic reactors for treating municipal solid waste. Bioresour Technol 94(3):261–266. doi:10.1016/j.biortech.2004.01.006
Lu L, Xing DF, Ren NQ (2012) Pyrosequencing reveals highly diverse microbial communities in microbial electrolysis cells involved in enhanced H-2 production from waste activated sludge. Water Res 46(7):2425–2434. doi:10.1016/j.watres.2012.02.005
Luo G, Angelidaki I (2014) Analysis of bacterial communities and bacterial pathogens in a biogas plant by the combination of ethidium monoazide, PCR and Ion Torrent sequencing. Water Res 60:156–163. doi:10.1016/j.watres.2014.04.047
Luo G, De Francisci D, Kougias P, Laura T, Zhu X, Angelidaki I (2015) New steady-state microbial community compositions and process performances in biogas reactors induced by temperature disturbances. Biotechnol Biofuels 8(1):3
Luo G, Wang W, Angelidaki I (2013) Anaerobic digestion for simultaneous sewage sludge treatment and CO biomethanation: process performance and microbial ecology. Environ Sci Technol 47(18):10685–10693. doi:10.1021/es401018d
Nesbø CL, Kumaraswamy R, Dlutek M, Doolittle WF, Foght J (2010) Searching for mesophilic Thermotogales bacteria: “mesotogas” in the wild. Appl Environ Microbiol 76(14):4896–4900. doi:10.1128/aem.02846-09
Nguyen T-AD, Han SJ, Kim JP, Kim MS, Oh YK, Sim SJ (2008) Hydrogen production by the hyperthermophilic eubacterium, Thermotoga neapolitana, using cellulose pretreated by ionic liquid. Int J Hydrogen Energ 33(19):5161–5168. doi:10.1016/j.ijhydene.2008.05.019
Nielsen HB, Mladenovska Z, Westermann P, Ahring BK (2004) Comparison of two-stage thermophilic (68 °C/55 °C) anaerobic digestion with one-stage thermophilic (55 °C) digestion of cattle manure. Biotechnol Bioeng 86(3):291–300. doi:10.1002/bit.20037
Pagaling E, Strathdee F, Spears BM, Cates ME, Allen RJ, Free A (2014) Community history affects the predictability of microbial ecosystem development. ISME J 8(1):19–30. doi:10.1038/ismej.2013.150
Regueiro L, Spirito CM, Usack JG, Hospodsky D, Werner JJ, Angenent LT (2015) Comparing the inhibitory thresholds of dairy manure co-digesters after prolonged acclimation periods: Part 2—correlations between microbiomes and environment. Water Res. doi:10.1016/j.watres.2015.05.046
Rincón B, Borja R, González JM, Portillo MC, Sáiz-Jiménez C (2008) Influence of organic loading rate and hydraulic retention time on the performance, stability and microbial communities of one-stage anaerobic digestion of two-phase olive mill solid residue. Biochem Eng J 40(2):253–261. doi:10.1016/j.bej.2007.12.019
Riviere D, Desvignes V, Pelletier E, Chaussonnerie S, Guermazi S, Weissenbach J (2009) Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J 3:700–714
Schluter A, Bekel T, Diaz NN, Dondrup M, Eichenlaub R, Gartemann KH, Krahn I, Krause L, Kromeke H, Kruse O, Mussgnug JH, Neuweger H, Niehaus K, Puhler A, Runte KJ, Szczepanowski R, Tauch A, Tilker A, Viehover P, Goesmann A (2008) The metagenome of a biogas-producing microbial community of a production-scale biogas plant fermenter analysed by the 454-pyrosequencing technology. J Biotechnol 136(1–2):77–90. doi:10.1016/j.jbiotec.2008.05.008
Sundberg C, Al-Soud WA, Larsson M, Alm E, Yekta SS, Svensson BH, Sørensen SJ, Karlsson A (2013) 454 pyrosequencing analyses of bacterial and archaeal richness in 21 full-scale biogas digesters. FEMS Microbiol Ecol 85(3):612–626. doi:10.1111/1574-6941.12148
Werner JJ, Knights D, Garcia ML, Scalfone NB, Smith S, Yarasheski K, Cummings TA, Beers AR, Knight R, Angenent LT (2011) Bacterial community structures are unique and resilient in full-scale bioenergy systems. Proc Nat Acad Sci USA 108(10):4158–4163. doi:10.1073/pnas.1015676108
Zhou JZ, Liu WZ, Deng Y, Jiang YH, Xue K, He ZL, Van Nostrand JD, Wu LY, Yang YF, Wang AJ (2013) Stochastic assembly leads to alternative communities with distinct functions in a bioreactor microbial community. Mbio 4(2). doi:10.1128/mBio.00584-12
Acknowledgments
This study was funded by the Yangfan project from Science and Technology Commission of Shanghai Municipality (14YF1400400), National Natural Science Foundation of China (51408133), and SRF for ROCS, SEM.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Han, S., Liu, Y., Zhang, S. et al. Reactor performances and microbial communities of biogas reactors: effects of inoculum sources. Appl Microbiol Biotechnol 100, 987–995 (2016). https://doi.org/10.1007/s00253-015-7062-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-7062-7