Abstract
A putative glycoside hydrolase family 43 β-xylosidase/α-arabinofuranosidase (CoXyl43) that promotes plant biomass saccharification was isolated via functional screening of a compost microbial metagenomic library and characterized. CoXyl43 promoted the saccharification of plant biomasses, including xylans (xylan and arabinoxylan), rice straw, and Erianthus, by degrading xylooligosaccharide residues to monosaccharide residues. The recombinant CoXyl43 protein exhibited both β-xylosidase and α-arabinofuranosidase activities for chromogenic substrates, with optimal activity at pH 7.5 and 55 °C. Both of these activities were inactivated by ethanol, dimethylsulfoxide, and zinc and copper ions but were activated by manganese ions. Only the β-xylosidase activity of recombinant CoXyl43 was enhanced in the presence of calcium ions. These results indicate that CoXyl43 exhibits unique enzymatic properties useful for biomass saccharification.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The replacement of fossil fuels on a global scale has the potential to solve multiple environmental problems in addition to fossil fuel depletion. Lignocellulosic biomass materials, such as various agricultural residues (e.g., rice straw, corn stover, and timber from forest thinning), can be converted into renewable biofuels and biochemicals. Lignocellulosic biomass is composed of cellulose, hemicelluloses, and lignin (Lynd et al. 2002) and is available in large quantities. Cellulose consists of linear chains of β-1,4-linked glucose and is the most abundant component of lignocellulosic biomass. Xylans are hemicelluloses and are the most common hetero-polysaccharides in lignocellulosic biomass (Saha 2003). Xylans are composed of homopolymeric backbone chains of β-1,4-linked xylopyranose units and may contain arabinose, glucuronic acid, and other small sugars (Table 1) (Kormelink and Voragen 1993; Saha 2003). In natural environments, lignocellulosic biomass is degraded by enzymes such as glycoside hydrolase, which is produced by environmental microorganisms. For example, cellulose is hydrolyzed by cellobiohydrolase (Teeri et al. 1983, 1987) and endo-β-1,4-glucanase (Penttilä et al. 1986; Okada et al. 1998; Saloheimo et al. 1988). Xylans are hydrolyzed by xylanase (Tenkanen et al. 1992; Xu et al. 1998) and xylosidase (Shallom et al. 2005). These glycoside hydrolases are vital for the saccharification and degradation of plant biomass.
The filamentous fungus, Trichoderma reesei (also known as Hypocrea jecorina), produces cellulases and hemicellulases in large quantities and efficiently degrades cellulosic biomass materials. Many mutants that produce large amounts of cellulase have been generated from wild-type T. reesei (Peterson and Nevalainen 2012). T. reesei PC-3-7, isolated from the QM9414 strain, is a cellulose hyperproducing mutant (Nogawa et al. 2001). Although T. reesei produces large amounts of cellulases and hemicellulases, these enzymes are too costly for use in commercial cellulosic biomass saccharification. Therefore, the discovery of new cellulases is important to facilitate biomass saccharification.
Although environmental microorganisms can be used to produce various glycoside hydrolases for the commercial degradation of lignocellulosic biomass, more than 99 % of environmental microorganisms are difficult to culture and have not been fully characterized (Torsvik and Øvreås 2002; Kimura et al. 2010). This indicates that the vast majority of microbial resources have not been accessed for biotechnology (Handelsman et al. 1998). This limitation can be overcome to a large extent by metagenomics, which is the culture-independent genomic analysis of microorganisms. Metagenomics has been used to screen for novel microbial enzymes in forest soil (Lee et al. 2008), activated sludge (Suenaga et al. 2007), marine environments (Okamura et al. 2010), and mammalian rumen (Beloqui et al. 2006; Ferrer et al. 2012).
In this study, a metagenomic approach was used to identify an enzyme that acts synergistically with T. reesei cellulase enzymes to improve the efficiency of lignocellulose saccharification. Genes encoding glycoside hydrolases were isolated from a compost metagenomic library using a chromogenic (p-nitrophenyl (pNP)) substrate mixture (pNP-β-D-lactopyranoside, pNP-β-D-xylopyranoside, pNP-β-D-mannopyranoside, and pNP-β-D-galactopyranoside). Among the isolated hydrolases was a putative glycoside hydrolase family 43 (GH43) protein (named CoXyl43) that enhanced the saccharification of lignocellulosic biomass and xylans using T. reesei cellulase enzymes. CoXyl43 had bifunctional β-xylosidase/α-arabinofuranosidase activity and was able to hydrolyze xylooligosaccharide intermediates produced by T. reesei cellulases and hemicellulases into xylose.
Materials and methods
Materials
Xylan from birch wood, pNP-α-L-arabinofuranoside, pNP-α-L-arabinopyranoside, pNP-β-L-arabinopyranoside, pNP-β-D-cellobioside, pNP-α-L-fucopyranoside, pNP-β-D-fucopyranoside, pNP-β-L-fucopyranoside, pNP-β-D-galactopyranoside, pNP-β-D-glucopyranoside, pNP-β-D-mannopyranoside, pNP-α-L-rhamnopyranoside, and pNP-α-D-xylopyranoside were purchased from Sigma-Aldrich (St. Louis, MO, USA). pNP-α-D-galactopyranoside, pNP-α-D-glucopyranoside, and pNP-β-D-xylopyranoside were purchased from Nacalai Tesque (Kyoto, Japan). pNP-α-D-mannopyranoside was purchased from Wako Pure Chemical Industries (Osaka, Japan). pNP-β-D-xylopyranoside, pNP-α-L-arabinofuranoside, pNP-β-D-cellobioside, pNP-β-D-fucopyranoside, pNP-β-L-fucopyranoside, pNP-β-D-galactopyranoside, pNP-α-D-galactopyranoside, and pNP-β-D-glucopyranoside were dissolved in water at a final concentration of 20 mM. pNP-α-L-arabinopyranoside, pNP-β-L-arabinopyranoside, pNP-α-L-fucopyranoside, pNP-β-D-mannopyranoside, pNP-α-D-mannopyranoside, pNP-α-L-rhamnopyranoside, pNP-α-D-xylopyranoside, and pNP-α-D-glucopyranoside were dissolved in 25 % dimethyl sulfoxide (DMSO) at a final concentration of 20 mM. Arabinoxylan from wheat flour (low viscosity) was obtained from Megazyme (Wicklow, Ireland). Arabinofuranosyl-xylooligosaccharides, O-α-L-arabinofuranosyl-(1→3)-O-β-xylopyranosyl-(1→4)-D-xylopyranose (Araf-X2) and O-β-D-xylopyranosyl-(1→4)-[O-α-L-arabinofuranosyl-(1→3)]-O-β-xylopyranosyl-(1→4)-D-xylopyranose (Araf-X3) were prepared as reported previously (Fujimoto et al. 2004).
Biomass pretreatment
Pretreated plant biomass for enzymatic saccharification was prepared by the alkaline treatment of rice straw or Erianthus as described previously with some modifications (Kawai et al. 2012). Alkaline-pretreated rice straw was treated with 0.5 % sodium hydroxide at 100 °C for 5 min. Alkaline-pretreated Erianthus was treated with 1 % sodium hydroxide at 120 °C for 5 min. The compositions of alkaline-pretreated rice straw and Erianthus were determined by acid hydrolysis and high-performance liquid chromatography (HPLC) according to the procedure published by NREL (http://www.nrel.gov/biomass/analytical_procedures.html). The monosaccharide compositions of alkaline-pretreated rice straw and Erianthus are listed in Table 1.
Construction of metagenomic library and screening of glycoside hydrolases with a positive effect on biomass saccharification
Metagenomic DNA extracted from composts was integrated into p18GFP plasmid vectors as described previously (Uchiyama et al. 2013).
Escherichia coli DH10B (Life Technologies Corporation, Carlsbad, CA, USA) harboring the metagenomic library were selected on Luria-Bertani (LB) agar plates containing ampicillin (LB + Amp), and 20 colonies were inoculated together into the same well of 96-well plates containing 900 μl LB + Amp medium with 10 μM isopropyl thiogalactoside (IPTG). After overnight cultivation at 37 °C, cells were collected by centrifugation (4000 rpm, 5 min) and resuspended in 220 μl Milli-Q water. Ten microliters of each cell suspension was mixed with 10 μl of a pNP substrate mixture (1 mM pNP-β-D-lactopyranoside, 1 mM pNP-β-D-xylopyranoside, 1 mM pNP-β-D-mannopyranoside, and 1 mM pNP-β-D-galactopyranoside) and incubated overnight at room temperature. After incubation, wells that had developed a yellow color, derived from pNP, were selected (pNP substrate screening). To isolate individual clones from the selected wells, a 20-clone mixture was diluted and cultured on LB + Amp plates. Clones that degraded the pNP substrate were selected (pNP substrate positive clones).
pNP substrate positive clones were then inoculated into 900 μl LB + Amp medium containing 10 μM IPTG. After overnight cultivation at 37 °C, cells were collected and resuspended in 145 μl Milli-Q water and 10 μl of 10× BugBuster (Novagen, Madison, WI, USA). The cell suspensions were incubated for 30 min at room temperature for protein extraction, and soluble fractions (4000 rpm, 20 min, 4 °C) of cell lysates were collected. Ten-microliter aliquots of the soluble fractions were mixed with 100 μl biomass premix (100 mM sodium acetate buffer (pH 5.0), 42.2 % (w/v) alkaline-treated rice straw, 0.02 % sodium azide), 70 μl 100 mM sodium acetate buffer (pH 5.0), and 20 μl 10 μg/ml cellulase CTec2 (Novozyme, Bagsværd, Denmark) and incubated overnight at 37 °C. To determine the polysaccharide degradation activity, reducing sugars were measured using the 3,5-dinitrosalicylic acid (DNS) reagent method (Miller 1959). Inserted DNA fragments encoding putative glycoside hydrolase, termed coxyl43, that had a positive effect on biomass saccharification were sequenced. The nucleotide sequence of coxyl43 was deposited in DDBJ/EMBL/GenBank under accession number LC025936.
Cloning, expression, and purification of glycoside hydrolase
To express the mature region of CoXyl43 in E. coli, the coxyl43 gene was synthesized in its codon-optimized form using DNA2.0 Inc. (Menlo Park, CA, USA) (DDBJ/EMBL/GenBank accession number LC027446) and cloned into a pET-28b vector (Novagen) digested with NdeI and XhoI. E. coli BL21(DE3) (NIPPON GENE, Tokyo, Japan) harboring the pET28b-putative glycoside hydrolase gene was cultured overnight in 100 ml Overnight Expression Instant LB medium (Novagen) containing 20 μg/ml kanamycin at 37 °C. After cultivation, the cells were harvested by centrifugation (5000 rpm, 3 min). The cell pellet was resuspended in BugBuster (Novagen) with Benzonase (Novagen) and incubated for 40 min at room temperature. Cell debris was removed by centrifugation (10,000 rpm, 20 min) at 4 °C. The supernatant was applied to a HisTrap HP Ni2+-affinity column (GE Healthcare, Buckinghamshire, England) to purify the recombinant enzyme. The recombinant protein was eluted with elution buffer (20 mM sodium phosphate, 500 mM NaCl, 500 mM imidazole, pH 7.4) and concentrated using an ultrafiltration membrane (Amicon Ultra 10K cutoff, Millipore, Darmstadt, Germany). The concentrated recombinant enzyme was then applied to a HiLoad 16/60 Superdex 200 prep-grade gel-filtration column (GE Healthcare) and eluted with PBS buffer (137 mM NaCl, 2.7 mM KCl, 10 mM disodium hydrogen phosphate, 1.8 mM potassium dihydrogen phosphate: pH 7.4). The enzyme fraction was concentrated using an ultrafiltration membrane (Amicon Ultra 10K cutoff, Millipore).
Biomass saccharification
Xylans (xylan from birch wood or arabinoxylan from wheat) were dissolved in Milli-Q water to a final concentration of 2 % and incubated at 98 °C for 5 min with agitation (1000 rpm) using a Thermomixer comfort (Eppendorf, Hamburg, Germany). After centrifugation (8000 rpm for 5 min), the supernatant was used as a source of xylans or arabinoxylans.
Culture supernatant of T. reesei PC-3-7 (ATCC 66589) (T. reesei PC-3-7 was cultured in the liquid medium containing Avicel) was used as a source of cellulases (Kawai et al. 2012; Kawamori et al. 1986). Saccharification of xylans by cellulases was performed in 200-μl tubes in 50 mM sodium phosphate buffer (pH 6.0) with or without CoXyl43. For xylan saccharification, 20 μl xylan solution, 10 μl 50 μg/ml cellulases (or 10 μl sterile water), and 10 μl 50 μg/ml CoXyl43 in 200 mM sodium phosphate buffer (pH 6.0) (or 10 μl 200 mM sodium phosphate buffer) were mixed and incubated at 40 °C for 24 h. For arabinoxylan saccharification, 20 μl arabinoxylan solution, 10 μl 50 μg/ml cellulases (or 10 μl sterile water), and 10 μl 50 μg/ml CoXyl43 in 200 mM sodium phosphate buffer (pH 6.0) (or 10 μl 200 mM sodium phosphate buffer) were mixed and incubated at 40 °C for 24 h. After incubation, the produced sugars were measured by DNS and analyzed using a high-performance ion chromatography system (HPIC) as described previously (Nakazawa et al. 2012). The degree of saccharification was calculated relative to the complete saccharification of xylans by phenol-sulfuric acid reaction (Dubois et al. 1956). Briefly, 200 μl 5 % phenol and 200 μl diluted xylan or arabinoxylan solutions were mixed, and 1 ml concentrated sulfuric acid was then added. After incubation at room temperature for 20 min, the absorbance was measured at 490 nm. A standard curve was constructed using D-xylose.
Biomass (alkaline-treated rice straw and Erianthus) saccharification by cellulases (T. reesei PC-3-7 strain) and CoXyl43 was performed in 20-ml plastic bottles. The total reaction volume was 2 ml. The reaction mixture contained 0.54 g alkaline-treated rice straw (water content: 81.4 %; final concentration 5 % (w/v)) or 0.56 g alkaline-treated Erianthus (water content: 82.1 %, final concentration 5 % (w/v)), 500 μl of 400 mM sodium phosphate buffer (pH 6.0), 10 μl 0.5 mg/ml CoXyl43 (or 10 μl sterile water), 117.6 μl 1.7 mg/ml cellulases (culture supernatant of T. reesei PC-3-7), 20 μl 2 % sodium azide, and sterile water. The reaction was performed at 40 °C with shaking at 150 rpm for 72 h. To inactivate the enzymes, 200 μl supernatant were incubated at 100 °C for 5 min. The produced sugars were measured by DNS and analyzed using an HPIC system as described previously (Nakazawa et al. 2012). A standard curve for DNS was constructed using D-glucose.
Substrate specificity
The total reaction volume was 20 μl and contained 0.15 μg of the recombinant enzyme, 50 mM sodium phosphate buffer (pH 7.5), and 5 mM pNP-substrate (pNP-β-D-xylopyranoside, pNP-α-D-xylopyranoside, pNP-α-L-arabinofuranoside, pNP-α-L-arabinopyranoside, pNP-β-L-arabinopyranoside, pNP-β-D-cellobioside, pNP-α-L-fucopyranoside, pNP-β-D-fucopyranoside, pNP-β-L-fucopyranoside, pNP-α-D-galactopyranoside, pNP-β-D-galactopyranoside, pNP-α-D-glucopyranoside, pNP-β-D-glucopyranoside, pNP-α-D-mannopyranoside, pNP-β-D-mannopyranoside, or pNP-α-L-rhamnopyranoside). The reaction mixture was incubated at 50 °C for 5 min. To stop the reaction, 50 μl of a 1.0-M sodium bicarbonate solution were added. The concentration of released pNP was determined by measuring the solution absorbance at 405 nm (Infinite M200 PRO, Tecan (Zurich, Switzerland)).
The substrate specificity of CoXyl43 for xylan and arabinoxylan was analyzed using xylan and arabinoxylan solutions (described above) in a final reaction volume of 20 μl. The reaction mixture contained 0.2 μg of the recombinant enzyme, 10 μl of the xylan or arabinoxylan solution, and 50 mM sodium phosphate buffer (pH 7.5). The reaction mixture was incubated at 50 °C for 10 min. The produced sugars were measured by DNS.
Arabinofuranosyl-xylooligosaccharide hydrolysis
Each reaction (100 μl final volume) contained 25 μg of the recombinant enzyme, 0.3 % Araf-X2 or Araf-X3, and 50 mM sodium phosphate buffer (pH 7.5). Reaction mixtures were incubated at 50 °C for 24 h. The produced sugars were analyzed by HPIC as described previously (Nakazawa et al. 2012).
Effects of pH and temperature on hydrolysis activity
The optimum pH for recombinant glycoside hydrolase activity with 3 mM pNP-β-D-xylopyranoside was evaluated at 40 °C for 5 min in McIlvaine’s buffer ranging from pH 3.0 to pH 9.0 (McIlvaine 1921). The optimum temperature for recombinant glycoside hydrolase activity with 2.86 mM pNP-β-D-xylopyranoside in McIlvaine buffer (pH 7.5) was evaluated from 20 to 80 °C for 5 min.
Kinetic analyses
The kinetic parameters of CoXyl43 for chromogenic substrates were determined using 0.025 μg recombinant CoXyl43 at pNP-β-D-xylopyranoside or pNP-α-L-arabinofuranoside concentrations from 0.125 to 8 mM in 50 mM sodium phosphate buffer (pH 7.5) at 50 °C for 5 min. The total reaction volume was 20 μl. To stop the reaction, 50 μl of a 1.0-M sodium bicarbonate solution were added. The kinetic parameters of CoXyl43 for xylobiose were determined using 0.5 μg recombinant CoXyl43 at xylobiose concentrations from 0.125 to 50 mM in 50 mM sodium phosphate buffer (pH 7.5) at 50 °C for 5 min. The reaction was stopped by heating the reaction mixture to 98 °C for 10 min. The xylose concentration was determined using a D-Xylose assay kit obtained from Megazyme (Wicklow, Ireland). Kinetic constants (K m and k cat) were calculated using a nonlinear regression of the Michaelis-Menten equation using GraphPad Prism version 5.0 (GraphPad Software, La Jolla, CA, USA).
Effects of organic solvents, metal ions, and chelating agent
The effects of additives including organic solvents (ethanol and DMSO), metal ions (Ca2+, Mg2+, Mn2+, Zn2+, Cu2+, and Fe2+), and ethylenediaminetetraacetic acid (EDTA) were evaluated by measuring enzyme activity in the presence of these additives with 5 mM pNP-β-D-xylopyranoside or pNP-α-L-arabinofuranoside and 50 mM sodium phosphate buffer (pH 7.5) at 50 °C for 5 min following the measurement of pNP release.
Results
Screening of clones with glycoside hydrolase activity from the metagenomic library
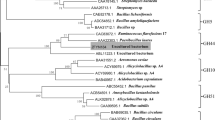
About 40 clones exhibiting glycoside hydrolase activity in a mixture of pNP substrates (mixture of pNP-β-D-lactopyranoside, pNP-β-D-xylopyranoside, pNP-β-D-mannopyranoside, and pNP-β-D-galactopyranoside) were screened from the metagenomic library (approximately 30,000 colonies). Among these pNP substrate positive clones, one showing a positive effect on biomass saccharification was isolated. Plasmid DNA was extracted from this clone, and the inserted DNA was sequenced. A gene encoding a putative glycoside hydrolase, named CoXyl43, was isolated from the sequencing data. CoXyl43 contains an open reading frame of 1107 bp encoding a putative protein of 369 amino acids. The N-terminus of the mature protein may begin at the 47th residue from the first Met, because the first 46 amino acids from the N-terminus are predicted to be a signal sequence. N-terminal proteolytic cleavage resulted in a protein of 323 amino acid residues with a predicted molecular mass of 36,186 Da. CoXyl43 was predicted to belong to the glycoside hydrolase family 43 (GH43), which contains β-xylosidase (EC 3.2.1.37) (Shallom et al. 2005; Brüx et al. 2006), α-arabinofuranosidase (EC 3.2.1.55) (Flipphi et al. 1993b), and endo-α-arabinase (EC 3.2.1.99) (Flipphi et al. 1993a). Comparisons against the BLAST database (NCBI BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi, nonredundant UniProtKB/SwissProt sequences) revealed that CoXyl43 is similar to Bacteroides ovatus Xsa (identity: 63 %, similarity: 75 %) (Whitehead 1995) and Prevotella ruminicola XynB (identity: 57 %, similarity: 69 %) (Gasparic et al. 1995), both of which belong to the GH43 class of carbohydrate-active enzymes in the CAZy database (Fig. 1). Both B. ovatus Xsa and P. ruminicola XynB are bifunctional β-xylosidases/α-arabinofuranosidases (EC 3.2.1.37, EC 3.2.1.55). CoXyl43 had a relatively low degree of similarity with the characterized GH43 arabinanases (EC 3.2.1.99) (Fig. 1).
Phylogenetic tree of GH43 enzymes. The amino acid sequences of GH43 enzymes sharing a high sequence similarity with CoXyl43 were obtained from UniProtKB (www.uniprot.org). Sequence alignment was performed using ClustalW (http://clustalw.ddbj.nig.ac.jp/), and the phylogenetic tree was constructed using FigTree v1.4.2 (http://tree.bio.ed.ac.uk/software/figtree/). UniProtKB accession numbers are as follows: Bacteroides ovatus Xsa, P49943; Prevotella ruminicola XynB, P48791; Bacillus subtilis XynD, Q45071; Paenibacillus polymyxa XynD, P45796; Clostridium stercorarium XylA, P48790; Streptomyces chartreusis Afase, P82594; Butyrivibrio fibrisolvens XylB, P45982; Escherichia coli YagH, P77713; B. subtilis XynB, P94489; Prevotella ruminicola β-xylosidase, Q9WXE8; B. subtilis Abn2, P42293; Thermotoga petrophila AbnA, A5IKD4; B. subtilis AbnA, P94522; Geobacillus stearothermophilus AbnB, B3EYM8; Geobacillus thermodenitrificans abn-ts, Q93HT9; Aspergillus nidulans AbnA, Q5BA96; Aspergillus terreus AbnA, Q0CS14; Aspergillus fumigatus AbnA, Q4WYX7; Neosartorya fischeri AbnA, A1D5W1; Aspergillus clavatus AbnA, A1CLG4; Aspergillus niger AbnA, A2QT85; A. niger AbnC, A5AAG2; Aspergillus oryzae AbnC, Q2U1X8; N. fischeri AbnC, A1DKY5; A. fumigatus AbnC, Q4W930; A. clavatus AbnC, A1CN18; A. fumigatus AbnB, B0XTS5; N. fischeri AbnB, A1DHW8; A. oryzae AbnB, Q2UI74; Aspergillus flavus AbnB, B8N803; A. nidulans AbnB, Q5AZC8; A. terreus AbnB, Q0CY27. EC numbers: 3.2.1.37, β-xylosidase; 3.2.1.55, α-L-arabinofuranosidase; 3.2.1.99, endo-1,5-α-L-arabinanase
The recombinant CoXyl43 enzyme was expressed in E. coli for characterization. His-tagged recombinant CoXyl43 was purified using Ni2+-affinity chromatography and gel filtration.
Synergism of cellulases and CoXyl43 on biomass saccharification
CoXyl43 and PC-3-7 cellulases (alone or combined) were incubated with birch wood xylan (Fig. 2a, c) or wheat flour arabinoxylan (Fig. 2b, d) to evaluate any synergistic effects on saccharification. Some reducing sugar (xylose) was released from both xylan and arabinoxylan in the presence of CoXyl43 alone, suggesting that CoXyl43 has a slight exo-xylanase activity with xylan and arabinoxylan (left panels of Fig. 2c, d). With T. reesei cellulases (PC-3-7), saccharification yields of xylan and arabinoxylan were approximately 36 and 43 %, respectively, and xylooligosaccharide (such as xylobiose and xylotriose) accumulation was detected (middle panels of Fig. 2c, d). When CoXyl43 was added to a xylan saccharification solution containing PC-3-7 cellulases, the accumulation of xylooligosaccharides was negligible (right panels of Fig. 2c, d), resulting in a significant increase in saccharification yields (Fig. 2a, b).
Synergism between T. reesei cellulases and CoXyl43 in xylan saccharification. T. reesei PC-3-7 cellulases and CoXyl43, either in combination (+PC3-7 +CoXyl43) or alone (+PC3-7 -CoXyl43, −PC3-7 +CoXyl43), were incubated with birch wood xylan (a and c) or wheat flour arabinoxylan (b and d) at 40 °C for 24 h. Saccharification yields (a and b) and HPIC chromatograms of the hydrolysates (c and d) are shown
Next, the synergistic effects of CoXyl43 with PC-3-7 cellulases in NaOH-pretreated rice straw and Erianthus saccharification were investigated (Fig. 3). As with xylan, some xylose was released from NaOH-pretreated rice straw and Erianthus in the presence of CoXyl43 alone (Fig. 3a, b). Accumulation of xylooligosaccharides and some cellooligosaccharides, including cellobiose, was also observed during the saccharification of NaOH-pretreated rice straw and Erianthus using PC-3-7 cellulases (Fig. 3a, b). The addition of CoXyl43 dramatically reduced the amounts of accumulated xylooligosaccharides (Fig. 3a, b), and the relative saccharification yield increased by approximately 5 % (Fig. 3c). These results indicated that CoXyl43 could almost completely hydrolyze the xylooligosaccharides derived from plant biomasses into xylose.
Synergism between T. reesei cellulases and CoXyl43 in NaOH-pretreated biomass saccharification. PC-3-7 cellulases and CoXyl43, either in combination (+PC3-7 +CoXyl43) or alone (+PC3-7 -CoXyl43, −PC3-7 +CoXyl43) were incubated with NaOH-pretreated rice straw (a and c) and Erianthus (b and c) at 40 °C for 72 h. HPIC chromatograms (a and b) and saccharification yields (c) are shown
Characterization of recombinant CoXyl43
The hydrolytic activity of recombinant CoXyl43 toward various chromogenic substrates was measured as a means of characterizing substrate specificity (Table 2). CoXyl43 exhibited hydrolytic activity with pNP-β-D-xylopyranoside and pNP-α-L-arabinofuranoside but not with other pNP substrates (such as pNP-β-D-mannopyranoside and pNP-β-D-galactopyranoside) (Table 2). The specific activities of recombinant CoXyl43 toward birch wood xylan and wheat arabinoxylan were 22.0 U/mg protein and 4.2 U/mg protein, respectively. One unit was defined as the amount of enzyme that released 1 μmol of xylose equivalents as reducing sugars per minute. CoXyl43 showed substantial activity between pH 5.5 and 9.0 and between 20 and 60 °C (Fig. 4a, b). The CoXyl43 activity with pNP-β-D-xylopyranoside was maximal at pH 7.5 and 55 °C.
We also investigated the hydrolytic activity of CoXyl43 against the arabinofuranosyl-xylooligosaccharides Araf-X2 and Araf-X3, which are degradation products of arabinoxylan (Fujimoto et al. 2004). Although CoXyl43 partially degraded Araf-X2 into xylose and arabinose (Fig. 5a), little activity was observed against Araf-X3 (Fig. 5b). These results indicate that CoXyl43 was able to degrade both xylooligosaccharides and the arabinose side chain of Araf-X2 (Fig. 5c). However, the xylose residue at the nonreducing end of Araf-X3 inhibited the release of an arabinofuranose residue, thereby inhibiting hydrolysis of the xylotriose backbone of Araf-X3 (Fig. 5d).
Arabinofuranosyl-xylooligosaccharide hydrolysis by CoXyl43. (a) O-α-L-arabinofuranosyl-(1→3)-O-βxylopyranosyl-(1→4)-D-xylopyranose (Araf-X2) and (b) O-β-D-xylopyranosyl-(1→4)-[O-α-L-arabinofuranosyl-(1→3)]-O-β-xylopyranosyl-(1→4)-D-xylopyranose (Araf-X3) were hydrolyzed by CoXyl43. The predicted mechanisms of Araf-X2 and Araf-X3 hydrolysis using CoXyl43 are shown in c and d, respectively
Kinetic constants of CoXyl43
The kinetic parameters (Michaelis constant: K m , turnover number: k cat, and k cat/K m ) of CoXyl43 were measured for pNP-β-D-xylopyranoside, pNP-α-L-arabinofuranoside, and xylobiose (Table 3). The K m of CoXyl43 for xylobiose was 2.02, and the k cat for xylobiose was 17.82. The K m of CoXyl43 for pNP-β-D-xylopyranoside and pNP-α-L-arabinofuranoside were 1.43 and 2.60, respectively, suggesting that pNP-β-D-xylopyranoside was the preferred substrate for CoXyl43. However, the k cat of CoXyl43 for pNP-β-D-xylopyranoside was much lower than that for pNP-α-L-arabinofuranoside, and the catalytic efficiency constant k cat/K m for pNP-β-D-xylopyranoside was almost equivalent to that for pNP-α-L-arabinofuranoside. Almost all of the GH43 bifunctional β-xylosidase/α-arabinofuranosidases have preferential activities. Some enzymes show much higher activity toward pNP-β-D-xylopyranoside than toward pNP-α-L-arabinofuranoside (Dougherty et al. 2012; Jordan et al. 2013; Shallom et al. 2005; Viborg et al. 2013; Whitehead and Cotta 2001; Kim and Yoon 2010; Zhou et al. 2012b) while the reverse relationship has been reported for other enzymes of the same family (Sakka et al. 1993; Shao and Wiegel 1992; Utt et al. 1991; Wagschal et al. 2009). Therefore, the almost equivalent catalytic efficiency constants for pNP-β-D-xylopyranoside and pNP-α-L-arabinofuranoside are unique to CoXyl43.
Effects of various organic solvents, metal ions, and chelating agent on CoXyl43 activity
The effects of various organic solvents, metal ions, and EDTA chelating agent on CoXyl43 were investigated (Table 4). Significant inactivation of CoXyl43 was observed in the presence of ethanol, DMSO, ZnCl2, and CuSO4. The CoXyl43 β-xylosidase activity was decreased by about 45 % with the addition of 100 mM EDTA, but no effect on α-arabinofuranosidase activity was observed with the addition of EDTA. The addition of MnCl2 resulted in a slight activity enhancement of both β-xylosidase and α-arabinofuranosidase activities (156.6 and 162.5 %, respectively). Significant activation of the β-xylosidase activity of CoXyl43 was observed upon addition of Ca2+, CaCl2, or CaSO4 (Table 4). The β-xylosidase activity was approximately 3-fold higher in the presence of 0.01 mM CaCl2 or CaSO4 and more than 5-fold higher in the presence of 1 mM CaCl2 or CaSO4. The β-xylosidase activity for 12.5 mM xylobiose at pH 7.5 (50 °C for 5 min) was enhanced by about 2.4-fold by addition of 1 mM CaCl2. In contrast, the α-arabinofuranosidase activity of CoXyl43 was enhanced by only about 25 % in the presence of 1 mM CaCl2 or CaSO4.
Discussion
Metagenomic analyses of environmental microorganisms are a highly effective means of screening for useful genes such as those suitable for biomass utilization and bioremediation. In the present study, we isolated a new β-xylosidase/α-arabinofuranosidase, CoXyl43, from a compost metagenome focusing on biomass saccharification. CoXyl43 displayed both β-D-xylosidase and α-L-arabinofuranosidase activities, similar to other members of the GH43 family of dual-functional enzymes and also showed xylooligosaccharide hydrolysis activity. CoXyl43 degraded xylooligosaccharides derived from plant biomasses, including both xylobiose and xylotriose, into xylose. Although CoXyl43 displayed high α-arabinofuranosidase activity toward pNP-α-L-arabinofuranoside, CoXyl43 removed few arabinofuranosyl units from the backbone chains of arabinoxylan and Araf-X3 (Figs. 2d and 5b). These results suggest that the backbone chains of β-1,4-linked xylopyranose units interfere with α-arabinofuranosidase activity toward the arabinofuranosyl units of arabinoxylan and Araf-X3.
In recent years, many genes encoding glycoside hydrolases have been obtained using metagenomic approaches. For example, Dougherty et al. (2012) isolated and characterized several endo-xylanases, α-fucosidase, and a bifunctional β-xylosidase/α-arabinofuranosidase from a compost metagenome. In addition, various cellulases and hemicellulases, including endo-glucanase (Pang et al. 2009; Alvarez et al. 2013), endo-xylanase (Gong et al. 2013), β-glucosidase/xylosidase (Zhou et al. 2012a; Bao et al. 2012), β-glucosidases (Uchiyama et al. 2013; McAndrew et al. 2013), β-galactosidase (Gupta et al. 2012), and β-xylosidase/α-arabinofuranosidase (Zhou et al. 2012b), were isolated from soil, compost, hot spring, or rumen using metagenomic methods. It was reported previously that a GH family 3 β-glucosidase/xylosidase and GH43 β-D-xylosidase/α-L-arabinofuranosidase from a yak rumen metagenome showed synergisms with endo-xylanase in xylan hydrolysis (Bao et al. 2012; Zhou et al. 2012b). Synergism between CoXyl43 and T. reesei cellulases was observed not only in xylan (from birch wood) and arabinoxylan (from wheat flour) saccharification but also in NaOH-pretreated biomass (rice straw and Erianthus) saccharification, resulting in a significant increase in saccharification efficiency. These results indicate that CoXyl43 would be useful for the saccharification of various cellulosic/hemicellulosic biomasses and that metagenomic approaches to environmental microorganisms have the potential to identify genes that may be valuable for the utilization of plant biomass. The metagenomic isolation of genes encoding non-GH family proteins that improve the efficiency of biomass saccharification will be explored in future studies.
Various chemical reagents affected CoXyl43 activity. Organic solvents, such as ethanol and DMSO, and zinc ions, and copper ions all strongly inhibited both the β-xylosidase and α-arabinofuranosidase activities of CoXyl43. Zinc and copper ions may inhibit the catalytic reaction or hinder substrate binding at the active site. Manganese ions enhanced both the β-xylosidase and α-arabinofuranosidase activities. The inhibition by zinc and copper ions, and activation by manganese ions, has been observed with other GH43 bifunctional β-xylosidase/α-arabinofuranosidases (Gong et al. 2013; Lee et al. 2013; Yang et al. 2014). Interestingly, the β-xylosidase activity of CoXyl43 was dramatically enhanced by the addition of calcium ions, while the α-arabinofuranosidase activity was not. The presence of EDTA inhibited β-xylosidase activity but not α-arabinofuranosidase activity. These results indicate that manganese ions enhance both β-xylosidase and α-arabinofuranosidase activities while calcium ions enhance only β-xylosidase activity. Further studies, incorporating X-ray crystal structure analyses of CoXyl43 with and without calcium ions, will aim to elucidate the mechanism by which calcium ions activate β-xylosidase activity while having no effect on α-arabinofuranosidase activity. Previous studies have reported distinctly different metal ion sensitivities among the GH43 enzymes (Viborg et al. 2013). Our ongoing studies will clarify how metal ions assist the catalytic function of GH43 enzymes.
References
Alvarez TM, Paiva JH, Ruiz DM, Cairo JP, Pereira IO, Paixão DA, de Almeida RF, Tonoli CC, Ruller R, Santos CR, Squina FM, Murakami MT (2013) Structure and function of a novel cellulase 5 from sugarcane soil metagenome. PLoS One 8
Bao L, Huang Q, Chang L, Sun Q, Zhou J, Lu H (2012) Cloning and characterization of two β-glucosidase/xylosidase enzymes from yak rumen metagenome. Appl Biochem Biotechnol 166:72–86
Beloqui A, Pita M, Polaina J, Martinez-Arias A, Golyshina OV, Zumárraga M, Yakimov MM, García-Arellano H, Alcalde M, Fernández VM, Elborough K, Andreu JM, Ballesteros A, Plou FJ, Timmis KN, Ferrer M, Golyshin PN (2006) Novel polyphenol oxidase mined from a metagenome expression library of bovine rumen: biochemical properties, structural analysis, and phylogenetic relationships. J Biol Chem 281:22933–22942
Brüx C, Ben-David A, Shallom-Shezifi D, Leon M, Niefind K, Shoham G, Shoham Y, Schomburg D (2006) The structure of an inverting GH43 β-xylosidase from Geobacillus stearothermophilus with its substrate reveals the role of the three catalytic residues. J Mol Biol 359:97–109
Dougherty MJ, D’haeseleer P, Hazen TC, Simmons BA, Adams PD, Hadi MZ (2012) Glycoside hydrolases from a targeted compost metagenome, activity-screening and functional characterization. BMC Biotechnol 12:38
Dubois M, Gilles KA, Hamilton JK, Rebers PA, Smith F (1956) Colorimetric method for determination of sugars and related substances. Anal Chem 28:350–356
Ferrer M, Ghazi A, Beloqui A, Vieites JM, López-Cortés N, Marín-Navarro J, Nechitaylo TY, Guazzaroni ME, Polaina J, Waliczek A, Chernikova TN, Reva ON, Golyshina OV, Golyshina PN (2012) Functional metagenomics unveils a multifunctional glycosyl hydrolase from the family 43 catalysing the breakdown of plant polymers in the calf rumen. PLoS One 7:e38134
Flipphi MJ, Panneman H, van der Veen P, Visser J, de Graaff LH (1993a) Molecular cloning, expression and structure of the endo-1,5-α-L-arabinase gene of Aspergillus niger. Appl Microbiol Biotechnol 40:318–326
Flipphi MJ, Visser J, van der Veen P, de Graaff LH (1993b) Cloning of the Aspergillus niger gene encoding α-L-arabinofuranosidase A. Appl Microbiol Biotechnol 39:335–340
Fujimoto Z, Kaneko S, Kuno A, Kobayashi H, Kusakabe I, Mizuno H (2004) Crystal structures of decorated xylooligosaccharides bound to a family 10 xylanase from Streptomyces olivaceoviridis E-86. J Biol Chem 279:9606–9614
Gasparic A, Martin J, Daniel AS, Flint HJ (1995) A xylan hydrolase gene cluster in Prevotella ruminicola B14: sequence relationships, synergistic interactions, and oxygen sensitivity of a novel enzyme with exoxylanase and β-(1,4)-xylosidase activities. Appl Environ Microbiol 61:2958–2964
Gong X, Gruniniger RJ, Forster RJ, Teather RM, McAllister TA (2013) Biochemical analysis of a highly specific, pH stable xylanase gene identified from a bovine rumen-derived metagenomic library. Appl Microbiol Biotechnol 97:2423–2431
Gruppen H, Hamer RJ, Voragen AGJ (1992) Water-unextractable cell wall material from wheat flour. 2. Fractionation of alkali-extracted polymers and comparison with water-extractable arabinoxylans. J Cereal Sci 16:53–67
Gupta R, Govil T, Capalash N, Sharma P (2012) Characterization of a glycoside hydrolase family 1 β-glucosidase from hot spring metagenome with transglycosylation activity. Appl Biochem Biotechnol 168:1681–1693
Handelsman J, Rondon MR, Brady SF, Clardy J, Goodman RM (1998) Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products. Chem Biol 5:R245–249
Jordan DB, Wagschal K, Grigorescu AA, Braker JD (2013) Highly active β-xylosidase of glycoside hydrolase family 43 operating on natural and artificial substrates. Appl Microbiol Biotechnol 97:4415–4428
Kawai T, Nakazawa H, Ida N, Okada H, Tani S, Sumitani J, Kawaguchi T, Ogasawara W, Morikawa Y, Kobayashi Y (2012) Analysis of the saccharification capability of high-functional cellulase JN11 for various pretreated biomass through a comparison with commercially available counterparts. J Ind Microbiol Biotechnol 39:1741–1749
Kawamori M, Morikawa Y, Takasawa S (1986) Induction and production of cellulases by L-sorbose in Trichoderma reesei. Appl Microbiol Biotechnol 24:449–453
Kimura N, Sakai K, Nakamura K (2010) Isolation and characterization of a 4-nitrotoluene-oxidizing enzyme from activated sludge by a metagenomic approach. Microbes Environ 25:133–139
Kim YA, Yoon KH (2010) Characterization of a Paenibacillus woosongensis β-xylosidase/α-arabinofuranosidase produced by recombinant Escherichia coli. J Microbiol Biotechnol 20:1711–1716
Kormelink FJM, Vorage AGJ (1993) Degradation of different [(glucurono)arabino]xylans by a combination of purified xylan-degradation enzymes. Appl Microbiol Biotechnol 38:688–695
Lee CC, Braker JD, Grigorescu AA, Wagschal K, Jordan DB (2013) Divalent metal activation of a GH43 β-xylosidase. Enzyme Microb Technol 52:84–90
Lee CM, Yeo YS, Lee JH, Kim SL, Kim JB, Han NS, Koo BS, Yoon SH (2008) Identification of a novel 4-hydroxyphenylpyruvate dioxygenase from the soil metagenome. Biochem Biophys Res Commun 370:322–326
Lynd LR, Weimer PJ, van Zyl WH, Pretorius IS (2002) Microbial cellulose utilization: fundamentals and biotechnology. Microbiol Mol Biol Rev 66:506–577
McAndrew RP, Park JI, Heins RA, Reindl W, Friedland GD, D’haeseleer P, Northen T, Sale KL, Simmons BA, Adams PD (2013) From soil to structure, a novel dimeric β-glucosidase belonging to glycoside hydrolase family 3 isolated from compost using metagenomic analysis. J Biol Chem 288:14985–14992
McIlvaine TC (1921) A buffer solution for colorimetric comparison. J Biol Chem 49:183–186
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Nakazawa H, Kawai T, Ida N, Shida Y, Kobayashi Y, Okada H, Tani S, Sumitani J, Kawaguchi T, Morikawa Y, Ogasawara W (2012) Construction of a recombinant Trichoderma reesei strain expressing Aspergillus aculeatus β-glucosidase 1 for efficient biomass conversion. Biotechnol Bioeng 109:92–99
Nogawa M, Goto M, Okada H, Morikawa Y (2001) L-Sorbose induces cellulase gene transcription in the cellulolytic fungus Trichoderma reesei. Curr Genet 38:329–334
Okada H, Tada K, Sekiya T, Yokoyama K, Takahashi A, Tohda H, Kumagai H, Morikawa (1998) Molecular characterization and heterologous expression of the gene encoding a low-molecular mass endoglucanase from Trichoderma reesei QM9414. Appl Environ Microbiol 64:555–563
Okamura Y, Kimura T, Yokouchi H, Meneses-Osorio M, Katoh M, Matsunaga T, Takeyama H (2010) Isolation and characterization of a GDSL esterase from the metagenome of a marine sponge-associated bacteria. Mar Biotechnol (NY) 12:395–402
Pang H, Zhang P, Duan CJ, Mo XC, Tang JL, Feng JX (2009) Identification of cellulase genes from the metagenomes of compost soils and functional characterization of one novel endoglucanase. Curr Microbiol 58:404–408
Penttilä M, Lehtovaara P, Nevalaninen H, Bhikhabhai R, Knowles J (1986) Homology between cellulase genes of Trichoderma reesei: complete nucleotide sequence of the endoglucanase I gene. Gene 45:253–263
Peterson R, Nevalainen H (2012) Trichoderma reesei RUT-C30—thirty years of strain improvement. Microbiology 158:58–68
Saha BC (2003) Hemicellulose bioconversion. J Ind Microbiol Biotechnol 30:279–291
Sakka K, Yoshikawa K, Kojima Y, Karita S, Ohmiya K, Shimada K (1993) Nucleotide sequence of the Clostridium stercorarium xylA gene encoding a bifunctional protein with β-D-xylosidase and α-L-arabinofuranosidase activities, and properties of the translated product. Biosci Biotechnol Biochem 57:268–272
Saloheimo M, Lehtovaara P, Penttilä M, Teeri TT, Ståhlberg J, Johansson G, Claeyssens M, Tomme P, Knowles JK (1988) EGIII, a new endoglucanase from Trichoderma reesei: the characterization of both gene and enzyme. Gene 63:11–22
Shallom D, Leon M, Bravman T, Ben-David A, Zaide G, Belakhov V, Shoham G, Schomburg D, Baasov T, Shoham Y (2005) Biochemical characterization and identification of the catalytic residues of a family 43 β-D-xylosidase from Geobacillus stearothermophilus T-6. Biochemistry 44:387–397
Shao W, Wiegel J (1992) Purification and characterization of a thermostable β-xylosidase from Thermoanaerobacter ethanolicus. J Bacteriol 174:5848–5853
Suenaga H, Ohnuki T, Miyazaki K (2007) Functional screening of a metagenomic library for genes involved in microbial degradation of aromatic compounds. Environ Microbiol 9:2289–2297
Teeri T, Salovuori I, Knowles J (1983) The molecular cloning of the major cellulase gene from Trichoderma reesei. Nat Biotechnol 1:696–699
Teeri TT, Lehtovaara P, Kauppinen S, Salovuori I, Knowles L (1987) Homologous domains in Trichoderma reesei cellulolytic enzymes: gene sequence and expression of cellobiohydrolase II. Gene 51:43–52
Tenkanen M, Puls J, Poutanen K (1992) Two major xylanases of Trichoderma reesei. Enzyme Microb Technol 14:566–574
Torsvik V, Øvreås L (2002) Microbial diversity and function in soil: from genes to ecosystems. Curr Opin Microbiol 5:240–245
Uchiyama T, Miyazaki K, Yaoi K (2013) Characterization of a novel β-glucosidase from a compost microbial metagenome with strong transglycosylation activity. J Bio Chem 288:18325–18334
Utt EA, Eddy CK, Keshav KF, Ingram LO (1991) Sequencing and expression of the Butyrivibrio fibrisolvens xylB gene encoding a novel bifunctional protein with β-D-xylosidase and α-L-arabinofuranosidase activities. Appl Environ Microbiol 57:1227–1234
Viborg AH, Sørensen KI, Gilad O, Steen-Jensen DB, Dilokpimol A, Jacobsen S, Svensson B (2013) Biochemical and kinetic characterization of a novel xylooligosaccharide-upregulated GH43 β-D-xylosidase/α-L-arabinofuranosidase (BXA43) from the probiotic Bifidobacterium animalis subsp. lactis BB-12. AMB Express 3:56
Wagschal K, Heng C, Lee CC, Wong DW (2009) Biochemical characterization of a novel dual-functional arabinofuranosidase/xylosidase isolated from compost starter mixture. Appl Microbiol Biotechnol 81:855–863
Whitehead TR (1995) Nucleotide sequences of xylan-inducible xylanase and xylosidase/arabinosidase genes from Bacteroides ovatus V975. Biochim Biophys Acta 1244:239–241
Whitehead TR, Cotta MA (2001) Identification of a broad-specificity xylosidase/arabinosidase important for xylooligosaccharide fermentation by the ruminal anaerobe Selenomonas ruminantium GA192. Curr Microbiol 43:293–298
Xu J, Takakuwa N, Nogawa M, Okada H, Morikawa Y (1998) A third xylanase from Trichoderma reesei PC-3-7. Appl Microbiol Biotechnol 49:718–724
Yang X, Shi P, Huang H, Luo H, Wang Y, Zhang W, Yao B (2014) Two xylose-tolerant GH43 bifunctional β-xylosidase/α-arabinofuranosidase and one GH11 xylanase from Humicola insolens and their synergy in the degradation of xylan. Food Chem 148:381–387
Zhou J, Bao L, Chang L, Liu Z, You C, Lu H (2012a) β-xylosidase activity of a GH3 glucosidase/xylosidase from yak rumen metagenome promotes the enzymatic degradation of hemicellulosic xylans. Lett Appl Microbiol 54:79–87
Zhou J, Bao L, Chang L, Zhou Y, Lu H (2012b) Biochemical and kinetic characterization of GH43 β-D-xylosidase/α-L-arabinofuranosidase and GH30 α-L-arabinofuranosidase/β-D-xylosidase from metagenome. J Ind Microbiol Biotechnol 39:143–152
Acknowledgments
We thank Dr. Y. Kobayashi (Japan Bioindustry Association) for provision of the biomass materials and helpful discussion. This work was supported in part by grants from the New Energy and Industrial Technology Development Organization (NEDO).
Conflict of interest
The authors declare no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matsuzawa, T., Kaneko, S. & Yaoi, K. Screening, identification, and characterization of a GH43 family β-xylosidase/α-arabinofuranosidase from a compost microbial metagenome. Appl Microbiol Biotechnol 99, 8943–8954 (2015). https://doi.org/10.1007/s00253-015-6647-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6647-5