Abstract
To optimize the expression of cry genes in a Bacillus thuringiensis sigK mutant failing in crystal releasing, the transcriptional activity of the cry promoters cry1A, cry3A, cry4A, and cry8E was compared using lacZ gene fusions. A beta-galactosidase assay indicated that the cry8E promoter showed the highest transcriptional activity. A novel Escherichia coli-B. thuringiensis shuttle vector pHT315-8E21b was constructed for cry gene expression using the cry8E promoter and the multiple cloning sites from vector pET21b, based on vector pHT315. SDS-PAGE analysis showed that the expression of the cry1Ac gene directed by the cry8E promoter was increased by approximately 2.4-fold over the expression directed by the cry3A promoter. The cry1Ba gene was expressed in the sigK mutant with the constructed vector pHT315-8E21b. Normal bipyramidal crystals encapsulated in mother cell were observed by transmission electron microscopy (TEM). The encapsulated Cry1Ba protein expressed in the sigK mutant showed activity against Ostrinia furnacalis and Plutella xylostella similar to that of the released Cry1Ba protein expressed in the acrystalliferous strain HD73 and can be protected from inactivation by UV light. All these results suggest that the cry8E promoter can be an efficient transcriptional element for cry gene expression in sigK mutants and can be utilized for the construction of a genetically engineered strain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacillus thuringiensis (Bt) is a Gram-positive bacterium that produces highly specific insecticidal crystal proteins encoded by the cry or cyt genes, also called δ-endotoxin, during sporulation. These insecticidal proteins are specifically toxic to the larvae of Lepidoptera, Coleoptera, Diptera, and nematodes and are more environmentally friendly than conventional pesticides. Thus, Bt-based biopesticides have become the most used microbial insecticides worldwide. Nonetheless, to date, Bt-based biopesticides have captured only ~2 % of the pesticide market (Bravo et al. 2011; Huang et al. 2007; Bradley et al. 1995). The low persistence of Bt agents after application due to environmental stresses, such as UV radiation, temperature, and rain, particularly the sunlight-mediated inactivation of the toxins, has become a conspicuous influencing factor for the further development of Bt-based biopesticides (Leong et al. 1980; Marianne et al. 1991). Some methods have been developed to increase the stability of toxins in the environment, such as microencapsulation (Yang et al. 2012b), melanin (Ruan et al. 2004), and genetic modification (Sanchis et al. 1999; Bravo et al. 1996). Of these, the genetic modification of the sigK mutant has proved to be an efficient way to prevent the inactivation of the crystal by sunlight due to the failure to release the crystal in the mother cell (Sanchis et al. 1999; Yang et al. 2013). However, the expression of some cry genes and the comparison of transcriptional activity among cry genes have not been clarified for the sigK mutant.
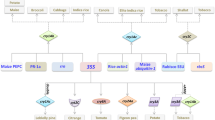
The products of B. thuringiensis cry genes are largely synthesized and accumulate as crystalline inclusions in the mother cell, processes that are dependent on highly ordered programs for the expression of the corresponding genes, which are regulated by a series of sigma factors in a signaling cascade (Kroos et al. 1999; Agaisse and Lereclus 1995). Usually, these cry genes are classified as sporulation dependent or sporulation independent based on their transcriptional mechanisms (Kroos et al. 1999; Schnepf et al. 1998). The cry3A gene, a typical example of a sporulation-independent cry gene, is controlled by σA and is expressed during vegetative growth (Agaisse and Lereclus 1994a). In contrast, most cry genes (e.g., cry1A, cry1B, cry1C, cry2A, cry4A, cry4B, cry11A, cry18Aa, cry34, and cry40) are sporulation-dependent genes controlled by σE, σK, or both (Bravo et al. 1996; Kroos et al. 1999; Brown 1993; Brizzard et al. 1991; Yoshisue et al. 1993; Zhang et al. 1998; Dervyn et al. 1995). Although the cry8Ea1 gene also belongs to the sporulation-dependent type, its transcription is controlled by two promoters, P orf1 and P cry8E , which are located upstream of the orf1 gene in the intergenic region mapping between orf1 and cry8Ea1 and are controlled by σE and σH, respectively (Du et al. 2012).
In this paper, we analyzed the influence of the sigK gene deletion on the activity of the cry1Ac, cry3A, cry4A, and cry8E gene promoters in B. thuringiensis and compared the transcriptional activity among these promoters in the sigK mutant (HD∆sigK). A high-level expression vector, pHT315-8E21b, and a sigK mutant engineered strain of B. thuringiensis, ∆sigK−-8E1Ba, were constructed using the cry8E promoter. We demonstrated that the cry8E promoter is a high-efficiency transcriptional element for cry gene expression in the sigK mutant and can be utilized for the construction of genetically engineered strains.
Material and methods
Bacterial strains, plasmids, and media. The bacterial strains and plasmids used in this study are listed in Table 1. Escherichia coli JM109 was used for the cloning experiments, and SCS110 was used to produce nonmethylated plasmid DNA for B. thuringiensis transformations (Macaluso and Mettus 1991). B. thuringiensis subsp. kurstaki HD73 and the acrystalliferous sigK mutant strain HDΔsigK− were used as the recipient strains to measure promoter activities (Du and Nickerson 1996). All the B. thuringiensis strains were grown with 220-rpm rotary agitation at 30 °C in Schaeffer’s sporulation medium (SSM); E. coli strains were grown at 37 °C in Luria-Bertani (LB) medium (1 % NaCl, 1 % tryptone, and 0.5 % yeast extract) (Schaeffer et al. 1965). The antibiotic concentrations used for bacterial selection were the following: 100 μg/ml ampicillin (for E. coli), 5 μg/ml erythromycin (for B. thuringiensis), and 200 μg/ml kanamycin (for B. thuringiensis).
DNA manipulation and transformation. Plasmid DNA was extracted from E. coli by the standard alkaline lysis procedure with a Plasmid Miniprep Kit (Axygen, Hangzhou, China). Restriction enzymes and T4 DNA ligase were used according to the manufacturer’s instructions (Takara Biotechnology Corporation, Dalian, China). PCR was performed with high-fidelity DNA polymerase (TOYOBO). The DNA fragments were separated on 0.7 % agarose gels after digestion and extracted from the gels using a DNA gel extraction kit (Axygen, Hangzhou, China). Standard procedures were used for E. coli transformation (Sambrook and Russell 2001), and Bt cells were transformed by electroporation as previously described (Lereclus et al. 1989).
Screening of acrystalliferous mutant strain HDΔsigK−. To obtain a acrystalliferous sigK mutant HDΔsigK− as receipt strain, we cure the plasmid pHT73 harboring cry1Ac gene from a sigK mutant strain HDΔsigK, which was obtained from strain HD73 with sigK gene deletion (Du et al. 2011). The sigK mutant strain HDΔsigK was grown in BP medium (1 % tryptone, 0.5 % NaCl, 0.36 % Na2HPO4, and 0.15 % KH2PO4) with 200 μg/ml kanamycin at 42 °C for 48 h with 220 rpm. The 2-ml culture was inoculated into 50-ml fresh BP medium with kanamycin at 45 °C for 12 h with 220-rpm rotate agitation. Then, the cells were plated on LB agar medium without antibiotics and grown at 30 °C for 48 h. The resulting single colonies were screened and identified by PCR amplification with primers cry1Ac-5/cry1Ac-3 and pHT73-5/pHT73-3 (Du and Nickerson 1996). Further identification of the acrystalliferous mutant was performed by SDS-PAGE and optical microscopy.
Construction of cry gene promoters with lacZ fusions. For determining the transcription activity of cry gene promoters, a high-copy-number plasmid pHTlac was constructed. The lacZ fragment was amplified by using the plasmid pHT304-18Z (Agaisse and Lereclus 1994a) as template, and the fragment then was digested and ligated into the HindIII and PstI sites of the B. thuringiensis-E. coli shuttle vector pHT315 (Arantes and Lereclus 1991) to produce the recombinant plasmid pHTlac harboring the promoterless lacZ gene. The promoter fragments of cry1Ac, cry3A, cry4A, and cry8E were amplified from B. thuringiensis HD73, Bt22, Bti, and Bt185 genomic DNA using the specific primer pairs according to previously studies LP1Ac-5/LP1Ac-3, LP3A-5/LP3A-3, LP4A-5/LP4A-3, and LPorf1-5/LP8E-3 (with 5′-KpnI and 3′-XbaI) (Table 2), respectively. P cry1Ac is a 382-bp fragment upstream of the cry1Ac translational start codon (access number AAB46989.1) (Wong et al. 1983). P cry3A is a 570 bp upstream of the cry3A translational start codon, and the access number is CAB41411.1 (Agaisse and Lereclus 1994b). P cry4A is a 826 bp upstream of the cry4A translational start codon, and the access number is CAD30148.1 (Yoshisue et al. 1993). Pcry8E is a 1,352-bp fragment upstream of the cry8E translational start codon, and the access number is AY329081.1 (Du et al. 2012). The PCR products were digested by restriction endonuclease KpnI and XbaI and then ligated into the vector pHTlac. The recombinants, pHTPcry1Ac, pHTPcry3A, pHTPcry4A, and pHTPorf1-cry8E, were separately introduced into HD73 and the sigK mutant.
β-Galactosidase assay. B. thuringiensis strains containing lacZ fusions were grown in SSM at 30 °C with 220 rpm. Samples of 2 ml were removed at 1-h intervals from T 1 to T 12 (T 0 indicates the end of the exponential growth phase; T n indicates n hours of the postexponential phase). The cells were harvested by centrifugation, and the pellets were stored at −20 °C. The β-galactosidase-specific activities were determined as previously described (Yang et al. 2012a) and are expressed as Miller units (Miller 1972). The values reported are the means of at least three independent assays.
Construction of an E. coli-B. thuringiensis shuttle vector. A B. thuringiensis-E. coli shuttle vector was constructed to express the cry gene under the control of the cry8E gene promoter. The 1,352-bp promoter fragment of the cry8E gene was obtained as described previously (Du et al. 2012), and the 213-bp expression sequence from plasmid pET-21b was amplified using pET21b-5/pET21b-3 as primers. The 1,565-bp overlapping fragment was amplified using the mixture of both the above fragments as a template and LPorf1-5 and pET21b-3 as primers. The overlapping PCR product was digested with SmaI and ligated into the EcoRI and HindIII sites of the B. thuringiensis-E. coli shuttle vector pHT315 by blunt-end ligation to produce the recombinant plasmid pHT315-8E21b, which carries the promoter of the cry8E gene, a multiple cloning site (MCS), and the T7 terminator (Fig. 3a).
Construction of the expression plasmid of cry1Ba with the cry8E promoter. The cry1Ba fragment was amplified from B. thuringiensis UV17 chromosomal DNA (Wang et al. 2006) using the primers 1Ba-5/1Ba-3 (with 5′-BamHI and 3′-SalI). The BamHI-SalI fragment of cry1Ba was then ligated into the B. thuringiensis-E. coli shuttle vector pHT315-8E21b (Fig. 3a), which harbors the cry8E gene promoter. The resulting plasmid was introduced into the HD73− and HDΔsigK− strains to produce the corresponding strains HD−-8E1Ba and ΔsigK−-8E1Ba.
SDS-PAGE analysis of Cry protein production. Different B. thuringiensis strains were grown at 30 °C with 220 rpm in SSM medium. After complete autolysis, 2.0-ml samples were centrifuged at 12,000 × g for 10 min, and the cells were suspended in 0.5 ml Tris–HCl (50 mM, pH 8.0). The bacterial cells were then ruptured with a Mini-Beadbeater. Samples of 100 μl were mixed with 5× loading buffer and boiled for 10 min for subsequent total protein quantitation and SDS-PAGE. The total protein quantitation was determined using Pierce 660 nm Protein Assay Reagent (Thermo Scientific). ImageJ software (National Institutes of Health) was used to determine the intensity of protein bands.
Irradiation of B. thuringiensis samples with a solar simulator. B. thuringiensis samples were irradiated with a solar analyzer (XT5409-XPC80) from Xutemp, which delivered a spectrum equivalent to that of sunlight passing through the Earth’s atmosphere. The products (60 μg of toxins) were sprayed onto 2.5- by 7.5-cm glass plates and air-dried (Sanchis et al. 1999). The coated plates were irradiated for 4 and 8 h using a solar analyzer. The analyzer uses a xenon lamp emitting from 280 to 800 nm at 100 klx. During irradiation, the glass plates were maintained at 25 °C. The irradiated toxins were recovered and then bioassayed against Plutella xylostella and Ostrinia furnacalis, as described below. For each irradiated sample, a control sample from the same batch was sprayed onto glass plates, air-dried, and recovered under the same conditions as for the irradiated plates.
Bioassay of insecticidal activity. Biological assays were performed by using free ingestion techniques and neonates. The protein concentration in the samples was estimated using a Total Protein Quantity Kit (Thermo Scientific, Beijing, China).
1. 50 % lethal concentration (LC50). Insecticidal activities were tested by exposing first instar larvae (the Asian corn borer, O. furnacalis) or second instar larvae (diamondback moth, P. xylostella) to an artificial diet incorporating one of seven dilutions of each preparation in water (Xue et al. 2008). Total seven toxin concentrations (0.1, 0.5, 1, 2, 4, 8, and 16 μg/ml) were used to determine LC50 values. The test for each concentration was performed in triplicate. The diet given to O. furnacalis was uniformly distributed into 48-well trays, with 400 mg in each tray. One first instar larva was placed in each of the 48 wells. The 300-mg diet provided to P. xylostella was uniformly distributed into 9- by 9-cm plastic Petri dishes, and 20 second instar larvae were placed in each dish. All tests used water as the control. The number of surviving larvae was recorded after 7 days for O. furnacalis and 3 days for P. xylostella. The LC50 was calculated using probit analysis (Finney 1971).
2. Mortality of samples after solar irradiation. The samples after solar radiation for 8 h, as described above, and control samples were diluted to 16 μg/ml and mixed into the diet of neonate larvae of P. xylostella and O. furnacalis. The concentration (16 μg/ml) which is the highest concentration among the seven dilution concentrations that were assayed to determine LC50 values was used to detect the mortality of samples in consideration of the loss of toxin activity after solar irradiation. The subsequent procedure was identical as the above, and the mortality of each sample was calculated as previously described (Raymond et al. 1993).
Results
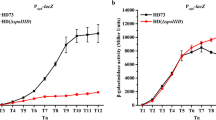
Transcriptional analysis of four cry gene promoters. The promoter activities of the cry1Ac, cry3A, cry4A, and cry8E genes were analyzed by constructing transcriptional fusions with the lacZ gene. β-Galactosidase assays indicated no difference of the activity of the P cry1Ac promoter in HD73 and HDΔsigK from T 0 to T 3. However, the transcriptional activity of P cry1Ac in strain HD73 was higher than that in strain HDΔsigK after T 3, and the highest transcriptional activity was obtained at T 9 in both strains (Fig. 1a). The β-galactosidase activities of P cry3A were detected at T 1, and there was no significant change in transcriptional activity during sporulation (Fig. 1b). The transcriptional activity of P cry4A in strain HDΔsigK was higher than that in strain HD73 after T 9 (Fig. 1c). Before T 9, the transcriptional activity of the cry8E promoter in strain HDΔsigK was similar to that in strain HD73. Thereafter, the β-galactosidase activity in the sigK mutant steadily increased, but it decreased in strain HD73 (Fig. 1d).
The effect of the sigK mutation on the transcriptional activity of the cry promoter. The promoter-directed β-galactosidase synthesis of three clones was determined at the indicated times after growing the cells in SSM at 30 °C. Each value represents the mean of at least three independent replicates. a cry1Ac promoter, b cry3A promoter, c cry4A promoter, d cry8E promoter
The transcriptional activities of the four cry gene promoters were also compared in both HD73 and HDΔsigK. The order of promoter activity, from high to low, was P cry8E , P cry1A , P cry4A , and P cry3A in both strains (Fig. 2a, b).
Comparison of the transcriptional activity among different cry gene promoters. The promoter-directed β-galactosidase synthesis of three clones was determined at the indicated times after growing the cells in SSM at 30 °C. Each value represents the mean of at least three independent replicates. a cry promoters in wild-type strain HD73, b cry promoters in sigK mutant
The comparison of expression efficiency of shuttle vectors with different promoters. To compare the efficiency of the E. coli-B. thuringiensis shuttle vector with respect to the expression of the cry gene, the intact fragment of the cry1Ac gene obtained by PCR amplification was inserted into the BamHI and SalI sites of the recombinant plasmid pHT315-8E21b and the previously used plasmid pSXY-422b, which also derived from pHT315 containing the cry3A gene promoter and pET-21b MCS site (Wang et al. 2006). The two expression plasmids were introduced into the acrystalliferous mutant HD73−, and the resulting strains were designated HD8E-1Ac and HD3A-1Ac. The SDS-PAGE result showed that the expression of the cry1Ac gene was significantly improved in HD8E-1Ac compared to the expression in HD3A-1Ac (Fig. 3b), with an increase of approximately 2.4-fold.
Construction of a highly efficient expression vector, pHT315-8E21b, expressing the cry gene. a Physical map of B. thuringiensis-E. coli shuttle plasmid pHT315-8E21b. The expression region includes the cry8Ea promoter, a T7 tag, an MCS, a His-tag, and a T7 terminator. b SDS-PAGE analysis of Cry1Ac expression in strains HD3A-1Ac and HD8E-1Ac. CK−:recipient strain HD73−. Ratio means the relative value of Cry1Ac production against the Cry1Ac production directed by the Pcry3A promoter, which is defined as 1. c SEM pictures of Bt strains HD3A-1Ac. d SEM pictures of Bt strains HD8E-1Ac. The oval inclusions marked by hollow arrow indicate spores, and the bipyramidal inclusions marked by solid arrow indicate protein crystals
The expression and bioassay of encapsulated Cry1Ba in the sigK mutant. The acrystalliferous sigK mutant strain HDΔsigK− was obtained by culturing the sigK mutant strain HDΔsigK at various temperatures (42, 45 °C) and screening via PCR amplification. To test the expression of the cry gene in the sigK mutant by the pHT315-8E21b vector, a cry1Ba3 gene, which has broad-spectrum insecticidal activity against three orders, Lepidoptera, Coleoptera, and Diptera (Zhong et al. 2000), and no cross-resistance with the Cry1A-resistant and Cry1C-resistant strain of the diamondback moth (P. xylostella) (Liu et al. 2001), was introduced into this vector to generate p8E1Ba. Then, the p8E1Ba vector was transformed into HD73− and HDΔsigK− to obtain the strains HD−-8E1Ba and ΔsigK−-8E1Ba, respectively. The formation of crystal proteins was examined by optical microscopy and electron microscopy (Fig. 4). Large bipyramidal crystals were observed in both the HD−-8E1Ba and ΔsigK−-8E1Ba strains, but the crystals were released from the HD−-8E1Ba strain (Fig. 4a, e); in contrast, those from the sigK mutant strain ΔsigK−-8E1Ba were encapsulated within the cell wall (Fig. 4b, f), and no mature phase refractile spores were observed (Fig. 4f). SDS-PAGE analysis showed that both the ΔsigK−-8E1Ba and HD−-8E1Ba strains produced the 130-kDa protein Cry1Ba and there were no 130-kDa proteins expressed in the recipient strains (Fig. 5). The expression of cry1Ba in the sigK mutant strain ΔsigK−-8E1Ba was similar to that of the control strain HD−-8E1Ba.
CLSM and TEM pictures of Bt strains HD−-8E1Ba and ΔsigK−-8E1Ba. a, b CLSM photos of strains HD−-8E1Ba and ΔsigK−-8E1Ba grown after 24 h in SSM medium at 30 °C, respectively. c–f TEM pictures of different Bt strains grown for 48 h in SSM medium at 30 °C. c HD73−, d HDΔsigK−, e HD−-8E1Ba, f ΔsigK−-8E1Ba
SDS-PAGE analysis of Cry1Ba production in the sigK mutant and HD73−. M protein molecular weight marker, CK1, recipient strain HD73−; CK2, recipient strain HDΔsigK−; Cry1Ba, recipient strains containing the vector p8E1Ba. Ratio means the relative value of Cry1Ba production against the Cry1Ba production from untreated HD−-8E1Ba, which is defined as 1
A bioassay of the activity of the HD−-8E1Ba and ΔsigK−-8E1Ba strains against O. furnacalis and P. xylostella was performed. The results showed that ΔsigK−-8E1Ba had a similar toxicity to P. xylostella and O. furnacalis as the control strain HD−-8E1Ba, in which expressed Cry1Ba proteins are released (Table 3). The LC50 values to P. xylostella and O. furnacalis of strain ΔsigK−-8E1Ba were close to those of strain HD−-8E1Ba.
Encapsulation of Cry proteins in a sigK mutant increases UV resistance. Because the sigK mutation resulted in no spore formation and no crystal release, it has been developed to protect crystal inactivation against UV light (Sanchis et al. 1999). The effect of encapsulation of the Cry1Ba protein in the sigK mutant on UV resistance was also analyzed in this study. Both B. thuringiensis strains HD−-8E1Ba (expressing Cry1Ba protein in the HD73 strain) and ΔsigK−-8E1Ba (expressing Cry1Ba protein in the sigK mutant) were irradiated for 4 and 8 h under a xenon lamp emitting from 280 to 800 nm at 100 klx, and the bioassays against P. xylostella and O. furnacalis were performed. The results indicated that the corrected mortality due to HD−-8E1Ba was significantly lower than that due to ΔsigK−-8E1Ba after irradiation of 8 h, whereas the mortality due to these strains was almost the same before irradiation. The loss percentage of toxicity against P. xylostella was 61.1 % for HD−-8E1Ba and 16.1 % for ΔsigK−-8E1Ba. The loss percentage of toxicity against O. furnacalis was 70.8 % for HD−-8E1Ba and 11.9 % for ΔsigK−-8E1Ba (Table 4).
Discussion
The transcription of four cry promoters in the sigK mutant was analyzed in this study. The σK factor was previously reported to control the expression of several cry genes, including cry1A, cry1B, cry1C, cry4A, cry11A, and cry18A (Bravo et al. 1996; Brown 1993; Brizzard et al. 1991; Yoshisue et al. 1993; Zhang et al. 1998; Dervyn et al. 1995; Yoshisuea et al. 1997). It is not surprising that the activity of the cry1A promoter decreased in the sigK mutant after T 3 because BtII of the two overlapped cry1Ac promoters is σK dependent, a result that is similar to the result reported by Yang et al. (2012a). Our results also showed that the transcriptional activation of the cry3A promoter in the sigK genetic background was similar to that in the wild type, which was consistent with results reported by Salamitou et al. (1996). The cry4A and cry8E genes have been reported to be primarily controlled by the sporulation factor σE (Du et al. 2012; Yoshisuea et al. 1997). It is very interesting that the activities of both the cry4A and cry8E promoters increased in the sigK mutant after T 9 (Fig. 1c, d). These data suggest that these two promoters were negatively regulated in the same pattern at the late stage of sporulation.
Although transcriptional analyses of many cry genes have been reported (Brown 1993; Brizzard et al. 1991; Yoshisue et al. 1993; Zhang et al. 1998; Dervyn et al. 1995; Yoshisuea et al. 1997; Yang et al. 2012a; Salamitou et al. 1996), it is surprising that little comparison of their promoter activities has been performed. Here, we proved that the cry8E promoter is the strongest of the four tested promoters in both the sigK mutant and the wild-type strain HD73. The cry3A promoter is controlled by σA and is expressed during vegetative growth. The STAB-SD sequence that stabilizes the cry3 transcript-ribosome complex has been used to enhance the yields of the Cry proteins (Park et al. 1998; Park et al. 1999). The vectors based on the cry3A promoter have been widely utilized to express Cry proteins and construct genetically engineered strains (Sanchis et al. 1996; García-Gómez et al. 2013). In this study, a constructed vector, pHT315-8E21b, containing the cry8E promoter proved to be more efficient at expressing the cry gene than a cry3A promoter-related vector. Our results suggest that the pHT315-8E21b vector is an efficient expression system for cry gene expression and the cry8Ea promoter can be a potential expression element for developing genetically engineered Bt strains. The cry8E promoter was also found to be the strongest promoter in the sigK mutant, which cannot release crystals during the late stage of sporulation and has been successfully used to develop a genetically engineered strain against the UV inactivation of crystals by Sanchis et al. (1999). We also confirmed that the encapsulation of the Cry1Ba protein increased its UV resistance in this study. These findings provide an efficient expression system for the future development of more powerful bioinsecticides based on B. thuringiensis.
References
Agaisse H, Lereclus D (1994a) Expression in Bacillus subtilis of the Bacillus thuringiensis cryIIIA toxin gene is not dependent on a sporulation-specific sigma factor and is increased in a spo0A mutant. J Bacteriol 176:4734–4741
Agaisse H, Lereclus D (1994b) Structural and functional analysis of the promoter region involved in full expression of the cryIIIA toxin gene of Bacillus thuringiensis. J Mol Biol 13(1):97–107
Agaisse H, Lereclus D (1995) How does Bacillus thuringiensis produce so much insecticidal crystal protein? J Bacteriol 21:6027–6032
Arantes O, Lereclus D (1991) Construction of cloning vectors for Bacillus thuringiensis. Gene 108:115–119
Bradley D, Harkey MA, Kim MK, Biever KD, Bauer LS (1995) The insecticidal CryIB crystal protein of Bacillus thuringiensis ssp. thuringiensis has dual specificity to coleopteran and lepidopteran larvae. J Invertebr Pathol 65:162–173
Bravo A, Agaisse H, Salamitou S, Lereclus D (1996) Analysis of cry1Aa expression in sigE and sigK mutants of Bacillus thuringiensis. Mol Gen Genet 250:734–741
Bravo A, Likitvivatanavong S, Gill SS, Soberon M (2011) Bacillus thuringiensis: a story of a successful bioinsecticide, insect biochemistry and molecular. Insect Biochem Mol Biol 41:423–431
Brizzard BL, Schnepf HE, Kronstad JW (1991) Expression of the cryIB crystal protein gene of Bacillus thuringiensis. Mol Gen Genet 231:59–64
Brown KL (1993) Transcriptional regulation of the Bacillus thuringiensis subsp. thompsoni crystal protein gene operon. J Bacteriol 175:7951–7957
Dervyn E, Poncet S, Klier A, Rapoport G (1995) Transcriptional regulation of the cryIVD gene operon from Bacillus thuringiensis subsp. israelensis. J Bacteriol 177:2283–2291
Du C, Nickerson KW (1996) Bacillus thuringiensis HD-73 spores have surface-localized Cry1Ac toxin: physiological and pathogenic consequences. Appl Environ Microbiol 62:3722–3726
Du LX, Wei J, Han LL, Chen Z, Zhang J, Song FP, Huang DF (2011) Characterization of Bacillus thuringiensis sigK disruption mutant and its influence on activation of cry3A promoter. Acta Microbiol Sin 51:1177–1184
Du LX, Qiu L, Peng Q, Lereclus D, Zhang J, Song F, Huang D (2012) Identification of the promoter in the intergenic region between orf1 and cry8Ea1 controlled by sigma H factor. Appl Environ Microbiol 78:4164–4168
Finney DJ (1971) Probit analysis. The University Press, Cambridge
García-Gómez BI, Sánchez J, Martínez de Castro DL, Ibarra JE, Bravo A, Soberón M (2013) Efficient production of Bacillus thuringiensis Cry1AMod toxins under regulation of cry3Aa promoter and single cysteine mutations in the protoxin region. Appl Envir Microbiol 79:6969–6973
Huang DF, Zhang J, Song FP, Lang Z (2007) Microbial control and biotechnology research on Bacillus thuringiensis in China. J Invertebr Pathol 95:175–180
Kroos L, Zhang B, Ichikawa H, Yu YT (1999) Control of sigma factor activity during Bacillus subtilis sporulation. Mol Microbiol 31:1285–1294
Leong K, Cano R, Kubinski A (1980) Factors affecting Bacillus thuringiensis total field persistence. Environ Entomol 9:593–599
Lereclus D, Arantes O, Chaufaux J, Lecadet M (1989) Transformation and expression of a cloned delta-endotoxin gene in Bacillus thuringiensis. FEMS Microbiol Lett 51:211–217
Liu YB, Tabashnik B, Meyer S, Crickmore N (2001) Cross-resistance and stability of resistance to Bacillus thuringiensis toxin Cry1C in diamondback moth. Appl Environ Microbiol 67:3216–3219
Macaluso A, Mettus AM (1991) Efficient transformation of Bacillus thuringiensis requires nonmethylated plasmid DNA. J Bacteriol 173(3):1353–1356
Marianne P, Paul F, Larry G, Harvey K, Timothy L, Paul RC (1991) The mechanism of sunlight-mediated inactivation of Bacillicus thuringiensis crystals. Biochem J 273:43–47
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor
Park HW, Ge B, Bauer LS, Federici BA (1998) Optimization of Cry3A yields in Bacillus thuringiensis by use of sporulation-dependent promoters in combination with the STAB-SD mRNA sequence. Appl Environ Microbiol 64:3932–3938
Park HW, Bideshi DK, Johnson JJ, Federici BA (1999) Differential enhancement of Cry2A versus Cry11A yields in Bacillus thuringiensis by use of the cry3A STAB mRNA sequence. Fems Microbiol Lett 181:319–327
Raymond M, Prato G, Ratsira D (1993) Probit analysis of mortality assays displaying quantal response. Version 3.3. Praxeme, Saint Georges d' Orque, France
Ruan L, Yu Z, Fang B, He W, Wang Y, Shen P (2004) Melanin pigment formation and increased UV resistance in Bacillus thuringiensis following high temperature induction. Syst Appl Microbiol 27:286–289
Salamitou S, Agaisse H, Bravo A, Lereclus D (1996) Genetic analysis of cryIIIA gene expression in Bacillus thuringiensis. Microbiol 142:2049–2055
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Sanchis V, Agaisse H, Chaufaux J, Lereclus D (1996) Construction of new insecticidal Bacillus thuringiensis recombinant strains by using the sporulation non-dependent expression system of cryIIIA and a site specific recombination vector. J Biotechnol 48:81–96
Sanchis V, Gohar M, Chaufaux J, Arantes O, Meier A, Agaisse H, Cayley J, Lereclus D (1999) Development and field performance of a broad-spectrum nonviable asporogenic recombinant strain of Bacillus thuringiensis with greater potency and UV resistance. Appl Environ Microbiol 65:4032–4039
Schaeffer P, Millet J, Aubert JP (1965) Catabolic repression of bacterial sporulation. Proc Natl Acad Sci U S A 54:704–711
Schnepf E, Crickmore N, Van Rie J, Lereclus D, Baum J, Feitelson J, Zeigler DR, Dean DH (1998) Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol Mol Biol Rev 62:775–806
Shu C, Yu H, Wang R, Fen S, Su X, Huang DF, Zhang J, Song FP (2009) Characterization of two novel cry8 genes from Bacillus thuringiensis strain BT185. Curr Microbiol 58:389–392
Wang G, Zhang J, Song F, Wu J, Feng S, Huang D (2006) Engineered Bacillus thuringiensis GO33A with broad insecticidal activity against lepidopteran and coleopteran pests. Appl Microbiol Biotechnol 72:924–930
Wang G, Zhang J, Song F, Gu A, Uwais A, Shao T, Huang D (2008) Recombinant Bacillus thuringiensis strain shows high insecticidal activity against Plutella xylostella and Leptinotarsa decemlineata without affecting nontarget species in the field. J Appl Microbiol 105:1365–2672
Wang XM, Du LX, Peng Q, Liang YP, Li J, Zhang J, Song FP (2012) Activity of four cry gene promoters in spoIIID mutant of Bacillus thuringiensis. Acta Microbiol Sin 52:1075–1084
Wong HC, Schnepf HE, Whiteley HR (1983) Transcriptional and translational start sites for the Bacillus thuringiensis crystal protein gene. J Biol Chem 258(3):1960–1967
Xue J, Liang G, Crickmore N, Li H, He K, Song F, Feng X, Huang D, Zhang J (2008) Cloning and characterization of a novel Cry1A toxin from Bacillus thuringiensis with high toxicity to the Asian corn borer and other lepidopteran insects. FEMS Microbiol Lett 280:95–101
Yang H, Wang P, Peng Q, Rong R, Liu C, Lereclus D, Zhang J, Song FP, Huang DF (2012a) Weak transcription of the cry1Ac gene in nonsporulating Bacillus thuringiensis cells. Appl Environ Microbiol 78:6466–6474
Yang W, He K, Zhang J, Guo S (2012b) pH-controlled Bacillus thuringiensis Cry1Ac protoxin loading and release from polyelectrolyte microcapsules. PLoS One 7:1–7
Yang JN, Peng Q, Chen Z, Deng C, Shu CL, Zhang J, Huang DF, Song FP (2013) Transcriptional regulation and characteristics of a novel N-acetylmuramoyl-L- alanineamidase gene involved in Bacillus thuringiensis mother cell lysis. J Bacteriol 195:2887–2897
Yoshisue H, Fukada T, Yoshida K, Sen K, Kurosawa S, Sakai H, Komano T (1993) Transcriptional regulation of Bacillus thuringiensis subsp. israelensis mosquito larvicidal crystal protein gene cryIVA. J Bacteriol 175:7951–7957
Yoshisue H, Sakaia H, Senb K, Yamagiwaa M, Komano T (1997) Identification of a second transcriptional start site for the insecticidal protein gene cryIVA of Bacillus thuringiensis subsp. israelensis. Gene 185:251–255
Zhang J, Schairer HU, Schnetter W, Lereclus D, Agaisse H (1998) Bacillus popilliae cry18Aa operon is transcribed by sigmaE and sigmaK forms of RNA polymerase from a single initiation site. Nucleic Acids Res 26:1288–1293
Zhong C, Ellar DJ, Bishop AH, Johnson C, Lin S, Hart ER (2000) Characterization of a Bacillus thuringiensis δ-endotoxin which is toxic to insects in three orders. J Invertebr Pathol 76:131–139
Acknowledgments
We are grateful to Dr. FengShou Dong for his help about sunlight inactivation experiments. This work was supported by grant from the National Natural Science Foundation (No. 31270111).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, C., Zheng, Q., Peng, Q. et al. Screening of cry-type promoters with strong activity and application in Cry protein encapsulation in a sigK mutant. Appl Microbiol Biotechnol 98, 7901–7909 (2014). https://doi.org/10.1007/s00253-014-5874-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-5874-5