Abstract
SpoIIID is a small, sequence-specific DNA-binding protein which can direct many genes’ transcription and has an effect on spore formation in Bacillus subtilis. We investigated the role of SpoIIID in mother cell lysis in Bacillus thuringiensis. A β-galactosidase assay based on the promoter fusions with lacZ indicated that the sigK gene was positively regulated by SpoIIID and σK negatively regulated the expression of sigE. The spoIIID mutant strain exhibited no mother cell lysis in Schaeffer’s sporulation medium (SSM) but did in ½ Luria-Bertani (LB) medium. cwlC is an essential hydrolase gene for mother cell lysis. Moreover, the expression of a PcwlC-lacZ fusion in spoIIID mutant was proved to be higher in ½ LB medium than in SSM. HD (ΔspoIIID)(ΔcwlC) mutant was obtained by knocking out the cwlC gene in HD(ΔspoIIID) and displayed no mother cell lysis in both SSM and ½ LB mediums. The deletion of spoIIID decreased the crystal protein production in HD73. The expression of Porf1cry8E and P5014 promoter fusions with lacZ gene in the acrystalliferous HD−(ΔspoIIID) mutant showed similar activity to that in the acrystalliferous HD73− strain before T7 and slightly higher than that in the acrystalliferous HD73− after T7. Sodium dodecyl sulfate polyacrylamide gel electrophoresis showed that Cry1Ac production in HD−(ΔspoIIID) directed by the Porf1cry8E and P5014 promoters was at a similar level as that in HD73 wild strain. Altogether, these results suggested that the spoIIID mutant with Porf1cry8E or P5014 promoters could be an alternative delivery system for cry gene expression with no mature spore formation and medium-dependent mother cell lysis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
SpoIIID is a pivotal transcriptional regulator that governs gene expression with σE, σK, and GerE during sporulation in Bacillus subtilis (Zheng and Losick 1990). The alternative sigma factor, σE, becomes active in the mother cell to regulate transcription of the spoIIID gene, and the sigma factor for later-acting mother cells, σK, can be upregulated by SpoIIID (Halberg and Kroos 1994; Kroos 2007). As a small and sequence-specific DNA-binding protein, SpoIIID can regulate more than 100 genes including many genes in σE or σK regulon and particularly the former. Moreover, it can also negatively regulate genes in the σK regulon (Eichenberger et al. 2004; Halberg and Kroos 1994; Ichikawa and Kroos 2000; Zhang et al. 1997).
Bacillus thuringiensis is a gram-positive bacterium that belongs to the Bacillus cereus group and is one of the most successful insect pathogens currently used for insect control, characterized by the formation of parasporal crystal proteins and spores during the stationary phase of its growth cycle (Bravo et al. 2011; Schnepf et al. 1998). In B. thuringiensis, spoIIID is also a crucial gene for spore formation (Zhang et al. 2009). However, the transcriptional regulation among σE, SpoIIID, and σK in B. thuringiensis remains unknown.
The released Cry proteins are rapidly inactivated by UV light when B. thuringiensis is applied in the field (Myasnik et al. 2001). A previous study showed that sigK mutation blocked mother cell lysis, which promoted the encapsulation of crystals with cell envelop to prevent UV inactivation (Sanchis et al. 1999; Zhou et al. 2014). However, the expression of some cry genes was obviously decreased during the late sporulation phase because of deletion of the sigK gene (Sanchis et al. 1999; Zhou et al. 2014). Cell wall hydrolases, forming a large and highly diverse group of enzymes by cleaving bonds in polymeric peptidoglycans to hydrolyze bacterial cell wall, maybe an efficient alternative, which does not affect the production of Cry proteins. Two hydrolase genes, cwlB and cwlC, controlled by σK, have been found to be involved in mother cell lysis in B. thuringiensis subsp. kurstaki strain HD73. The deletion of the cwlB gene in B. thuringiensis caused a delay in mother cell lysis and disrupting the cwlC gene completely blocked mother cell lysis without impacting sporulation, crystal protein production, and insecticidal activity (Chen et al. 2018; Yang et al. 2013). SpoIIID can positively regulate the transcription of sigK in B. subtilis. Thus, investigating the effect of spoIIID on mother cell lysis in B. thuringiensis may provide insight into the transcriptional mechanism.
In this study, we analyzed the regulatory relationship between SpoIIID, σE, and σK in B. thuringiensis, which is similar to that in B. subtilis. HD(ΔspoIIID) mutant lacking the ability to form spores did not lyse in Schaeffer’s sporulation medium (SSM) but did in ½ Luria-Bertani (LB) medium. HD(ΔspoIIID)(ΔcwlC) mutant was constructed and exhibited no mother cell lysis. We demonstrated that Porf1cry8E and P5014 are strong promoters for cry1Ac gene expression in the spoIIID mutant, indicating a potential delivery system for Cry protein expression.
Materials and methods
Bacterial strains, plasmids, and growth conditions
All the bacterial strains and plasmids used in this study are summarized in Table 1. Escherichia coli TG1 was used as a host for molecular cloning, and E. coli SCS110 (also called ET12567) was used for the production of non-methylated plasmid DNA for B. thuringiensis transformation (Hoffmann et al. 2015; Wang et al. 2006). E. coli strains were grown at 37 °C in LB medium (1% tryptone, 0.5% yeast extract, and 0.5% NaCl) with ampicillin (Amp, 100 μg/ml) when required. B. thuringiensis strains were grown at 30 °C in either LB or ½ LB medium (0.5% tryptone, 0.25% yeast extract, and 0.25% NaCl) or SSM (0.8% nutrient broth, 1 mM MgSO4, 13.4 mM KCl, and 0.5 mM NaOH with 1 mM Ca(NO3)2, 0.01 μM MnCl2, and 1 μM FeSO4 added before use) (Schaeffer et al. 1965). When required, erythromycin (Ery, 5 μg/ml) or kanamycin (Kan, 100 μg/ml) was added for the growth of B. thuringiensis. The standard strain B. thuringiensis subsp. kurstaki HD73 (BGSC strain number BGSC 4D4, hereafter abbreviated as HD73, GenBank Accession Number: NC_020238.1) served as the recipient for monitoring gene transcriptional activity and genetic manipulation of B. thuringiensis (Du and Nickerson 1996; Liu et al. 2013).
DNA amplification, purification, and transformation
Routine handling of nucleic acids was performed according to the standard protocols. Polymerase chain reaction (PCR) was performed with either PrimeSTAR HS DNA polymerase (TaKaRa Biotechnology Corporation, Beijing, China) or Taq DNA polymerase (BioMed, Beijing, China). The bacterial lysate or the plasmid extracted from E. coli cells with the Plasmid Miniprep Kit (Axygen, Beijing, China) was used as the template DNA for PCR. Restriction enzymes and T4 DNA ligase (TaKaRa Biotechnology Corporation) were used in accordance with the manufacturer’s instructions. After agarose gel electrophoresis, all DNA fragments were isolated and purified with the AxyPrep DNA Gel Extraction Kit (Axygen).
Standard procedures were followed for E. coli transformation (Kaiser et al. 1998). B. thuringiensis cells were transformed by electroporation as described previously (Lereclus et al. 1989).
Construction of lacZ fusions and cry1Ac expression strains
To compare the transcriptional activities of the different genes in the wild-type and mutant strains, the promoter regions of sigK (PsigK, 1048 bp) and sigE (PsigE, 264 bp) were amplified by PCR using B. thuringiensis genomic DNA as the template. All the oligonucleotides are listed in Table 2. The amplified fragments of PsigK and PsigE were digested by BamHI/HindIII and PstI/HindIII, respectively. After gel purification, the fragments were inserted into the linearized pHT304-18Z plasmid (Agaisse and Lereclus 1994), which harbors a promoterless lacZ gene, by ligation reaction. The resulting plasmids, pHT304-PsigK and pHT304-PsigE, were transformed into B. thuringiensis HD73 and the mutant HD(ΔspoIIID), respectively. The plasmid pHT304-PcwlC of PcwlC (PcwlC, 717 bp) with the lacZ gene was transformed into B. thuringiensis HD73 and the mutant HD(ΔspoIIID), respectively. The plasmids pHT304-Porf1cry8E and pHT304-P5014 of Porf1cry8E (Porf1cry8E, 1352 bp) and P5014 (P5014, 710 bp) with the lacZ gene were also transformed into B. thuringiensis HD73− (acrystalliferous HD73 mutant strain) and the mutant HD−(ΔspoIIID) (acrystalliferous spoIIID mutant strain), respectively (Chen et al. 2018; Du et al. 2012; Zhang et al. 2018).
The constructed plasmids pHT315-P8E-1Ac and pHT315-P5014-1Ac containing the Porf1cry8E-cry1Ac and P5014-cry1Ac fusion (Zhang et al. 2018; Zhou et al. 2014) were introduced into HD−(ΔspoIIID) (Wang et al. 2012) mutants by electroporation (Lereclus et al. 1989). All the transformants were cultured at 30 °C and selected on Ery agar plates.
Construction of the cwlC and spoIIID disruption mutant
To knock out the cwlC gene in the HD(ΔspoIIID) strain, the pRN5101_cwlC plasmid carrying a Kan resistance gene (Kan) fragment (Chen et al. 2018) was transformed into the HD(ΔspoIIID) cells by electroporation. The transformants were selected on LB agar plates containing Ery and Kan at 30 °C and verified by PCR with the pRN5101-F and pRN5101-R primers. Allelic replacement of the pRN5101_cwlC plasmid in the HD(ΔspoIIID) cells was achieved by homologous recombination as reported previously (Yang et al. 2013). The mutant was verified by PCR and sequencing with the primers cwlC-a and cwlC-b.
Microscopic analysis
The wild-type strain (HD73), spoIIID deletion mutant [HD(ΔspoIIID)], sigK deletion mutation [HD(ΔsigK)], cwlC deletion mutant [HD(ΔcwlC)], and spoIIID and cwlC double-deletion mutant [HD(ΔspoIIID)(ΔcwlC)] were cultured in 50 ml of SSM and ½ LB at 30 °C with shaking at 220 rpm, respectively. At the designated time point (T18, T24, T30, and day 15; T0 is the end of the exponential growth phase, and Tn is the n hours after T0), 1 ml of each sample was collected by centrifugation (12,000 rpm, 2 min). The pellets were resuspended in a final volume of 100 μl of deionized water. Each cell sample (1.5 μl) was spotted onto the center of a glass slide and covered with a coverslip. Cell samples were analyzed with an optical microscope (BX61, Olympus, Tokyo, Japan).
Assays of β-galactosidase activities
B. thuringiensis cells were grown in SSM at 30 °C with shaking at 220 rpm. Two milliliters of culture was collected at each time point (for PsigK-lacZ: T2–T12; for PsigE-lacZ: T1–T12; for PcwlC-lacZ: T8–T19; for Porf1cry8E-lacZ: T0–T12; for P5014-lacZ: T0–T12; T0 indicates the end of the exponential growth phase, and Tn indicates the n hours after T0) and centrifuged at 12,000 rpm for 1 min, and the pellets were stored at − 70 °C ready for use. β-Galactosidase activities in the pelleted cells were measured as described previously (Miller 1972). β-Galactosidase activity is expressed as Miller units (Miller 1972). The data shown are the mean values of at least three independent experiments.
Sodium dodecyl sulfate polyacrylamide gel electrophoresis analysis of Cry protein production
Tested strains were cultured in 100 ml of SSM at 30 °C with shaking (220 rpm) to T24, and the cells were centrifuged at 4 °C at 10,000 rpm, followed by freeze-drying for 72 h until the pellets became lyophilized powders. The same amount of experimental powder by weight was dissolved in an equal volume of double-distilled water in a 2-ml centrifuge tube to obtain a suitable concentration, followed by disruption using the BeadBeater (Biospec Products, Inc., Bartlesville, OK, USA). The supernatant was transferred to a new centrifuge tube after centrifuging at 12,000×g. Then, 20 μl of cell lysates was mixed with 5 μl of 5× loading buffer and boiled for 15 min. Total protein quantitation and sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) analysis were performed as described previously (Miller 1972; Yang et al. 2013).
Results
The expression of PsigK-lacZ and PsigE-lacZ fusions in spoIIID mutant
We separately constructed transcriptional gene fusions between the 5′ regions of sigK and sigE and a promoterless lacZ gene harbored in the pHT304-18Z plasmid. The recombinant plasmids PHT304-PsigK and PHT304-PsigE were introduced into HD73 and HD(ΔspoIIID) cells by electroporation, respectively. β-Galactosidase assay suggested that the transcriptional activity of PsigK in HD(ΔspoIIID) was much lower than that in the HD73 wild type (Fig. 1a). It suggested that SpoIIID positively regulates sigK gene. The activity of sigE in both HD73 and HD(ΔspoIIID) was similar before T5 but higher in HD(ΔspoIIID) than in the HD73 strain after T5 (Fig. 1b). The previous studies showed that SpoIIID activated the transcription of sigK and σK repressed the transcription of sigE in B. subtilis (Halberg and Kroos 1994; Zhang and Kroos 1997; Zhang et al. 1999; Zheng and Losick 1990). The increasing of PsigE expression in HD(ΔspoIIID) after T5 might have resulted from the similar feedback regulation of σK in B. thuringiensis.
The effect of spoIIID mutation on the expression of sigK and sigE promoters. a Expression of PsigK-lacZ in HD73 and HD(ΔspoIIID). b Expression of PsigE-lacZ in HD73 and HD(ΔspoIIID). β-Galactosidase activity during sporulation after resuspension in SSM was measured for B. thuringiensis containing the lacZ gene fused with the sigK and sigE promoters in wild-type HD73 (black line) and HD(ΔspoIIID) mutant (red line). Each point represents the average of three determinations, and error bars show one standard deviation of the data
spoIIID mutation affects mother cell lysis
Knocking out the cwlC gene, which is regulated by σK, blocks mother cell lysis in HD73 (Chen et al. 2018). Disruption of sigK also blocks mother cell lysis (Sanchis et al. 1999; Zhou et al. 2014). The results described in the previous section showed that SpoIIID positively regulates the sigK gene. Thus, we investigated the role of SpoIIID in mother cell lysis. We first examined the morphology of the B. thuringiensis strains at different growth phases by optical microscopy. HD73 mother cells could naturally lyse in three different media, including SSM, ½ LB, and LB (Fig. 2, data for LB not shown). On the other hand, HD(ΔsigK) mutant did not lyse in any media (Fig. 2, data for LB not shown). HD(ΔspoIIID) mutant showed little mother cell lysis even cultured to 15 days in SSM and LB mediums (Fig. 2, data for LB not shown). Surprisingly, the majority of HD(ΔspoIIID) mother cells autolyzed in ½ LB medium at T30 (Fig. 2b). Altogether, these results indicated that mother cell lysis in spoIIID mutant depends on the growth medium.
Observation of mother cell lysis of B. thuringiensis HD73 (wild-type strain) cells, the HD(ΔspoIIID) mutant cells, and the HD(ΔsigK) mutant cells. a Three strains cultivated in SSM were observed by optical microscopy at T24, T30, and 15 days. b Three strains cultivated in ½ LB were observed by optical microscopy at T18, T30, and 15 days. The black arrows pointing to the sporangium at 15 days and the red arrows pointing to the free crystals
Deletion of the spoIIID gene decreases PcwlC-lacZ expression
CwlC has been proven to be essential for mother cell lysis in B. thuringiensis subsp. kurstaki strain HD73 (Chen et al. 2018). Therefore, we detected PcwlC expression in spoIIID deletion mutant. The PcwlC-lacZ recombinant plasmid was introduced into HD73 and HD(ΔspoIIID) cells by electroporation, respectively. The expression of PcwlC-lacZ in HD(ΔspoIIID) was considerably lower than that in the HD73 wild-type strain in both SSM and ½ LB medium (Fig. 3). Besides, the expression of PcwlC-lacZ in spoIIID mutant was higher in ½ LB than in SSM (Fig. 3). This result indicated that the lower expression of PcwlC in spoIIID mutant might result in no mother cell lysis in SSM medium.
The expression of PcwlC-lacZ in SSM and ½ LB mediums. a Expression of PcwlC-lacZ in HD73 and HD(ΔspoIIID) cultured in SSM medium. b Expression of PcwlC-lacZ in HD73 and HD(ΔspoIIID) cultured in ½ LB medium. β-Galactosidase activity during sporulation after resuspension in SSM or ½ LB mediums was measured for B. thuringiensis containing lacZ fused to the cwlC promoter in wild-type HD73 (black line) and HD(ΔspoIIID) mutant (red line). Each point represents the average of three determinations, and error bars show one standard deviation of the data
spoIIID and cwlC double mutant exhibit no mother cell lysis
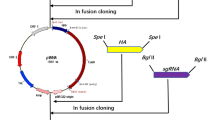
Mother cells of spoIIID mutant still lysed if cultured in ½ LB. Thus, double-gene mutant HD(ΔspoIIID)(ΔcwlC) was constructed by replacing the cwlC coding sequence with the Kan resistance gene through homologous recombination in the HD(ΔspoIIID) mutant (Fig. 4a). The morphology of the HD(ΔspoIIID)(ΔcwlC) and HD(ΔcwlC) strain cells grown in SSM and ½ LB medium was observed through optical microscopy. Both cell types did not autolyze even after 15 days of growth, and most of the cells contained crystals. These results indicated that knocking out the cwlC gene in spoIIID mutant completely blocked mother cell lysis in both SSM and ½ LB medium (Fig. 4b, c).
Mother cell lysis of the double mutant. a The construction of plasmid and mutants. b HD(ΔspoIIID)(ΔcwlC) and HD(ΔcwlC) mutant cells cultivated in SSM were observed at T24, T30, and 15 days. c HD(ΔspoIIID)(ΔcwlC) and HD(ΔcwlC) mutant cells cultivated in ½ LB medium were observed at T18, T30, and 15 days. Observation of mother cell lysis by optical microscopy. The black arrows pointing to a sporangium with a mature spore for the HD(ΔcwlC) mutant and red arrows pointing to a sporangium with an immature spore for the HD(ΔspoIIID)(ΔcwlC) mutant at 15 days
cry1Ac gene expression in spoIIID mutant directed by two promoters
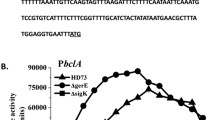
Porf1cry8E and P5014 are two strong promoters that can direct high cry1Ac gene expression in HD73 and HD(ΔsigK) during the sporulation stages (Zhang et al. 2018; Zhou et al. 2014). The sequences of Porf1cry8E and P5014 were cloned from HD73 genomic DNA and integrated into the promoterless vector pHT304-18Z (Du et al. 2012; Zhang et al. 2018). The resulting plasmids, PHT304-Porf1cry8E and PHT304-P5014, were introduced into the spoIIID mutant and subjected to β-galactosidase assays. The promoters’ expression of orf1cry8E and 5014 in HD−(ΔspoIIID) were almost equal to those in HD73−, which revealed that the deletion of spoIIID did not decrease the activity of the two promoters (Fig. 5a, b). The pHT315-P8E-1Ac and pHT315-P5014-1Ac plasmids separately containing the Porf1cry8E-1Ac and P5014-1Ac fusions were introduced into the acrystalliferous spoIIID mutant HD−(ΔspoIIID) to express Cry1Ac protein. SDS-PAGE revealed that both Porf1cry8E and P5014 can direct Cry1Ac protein expression with a similar yield in HD−(ΔspoIIID) as that of the control strain HD73. Much less Cry1Ac protein was expressed in the HD(ΔspoIIID) strain which was directed by native cry1Ac promoter. The recipient strain HD−(ΔspoIIID) also produced rarely Cry1Ac protein (Fig. 5c). Taken together, these results suggested that Porf1cry8E and P5014 are suitable candidates to direct cry gene expression in spoIIID mutant.
Cry1Ac expression in the B. thuringiensis strain. a Expression of Porf1cry8E-lacZ in HD73− and HD−(ΔspoIIID). b Expression of P5014-lacZ in HD73− and HD−(ΔspoIIID). β-Galactosidase activity during sporulation after resuspension in SSM was measured for B. thuringiensis containing lacZ fused to the orf1cry8E and 5014 promoters in wild-type HD73− (black line) and HD−(ΔspoIIID) mutant (red line). Each point represents the average of three determinations, and error bars show one standard deviation of the data. c SDS-PAGE analysis of Cry1Ac production in HD73, HD−(ΔspoIIID)-P8E-1Ac (Porf1cry8E in the figure), HD(ΔspoIIID) (Cry1Ac directed by native promoter), HD−(ΔspoIIID)-P5014-1Ac (P5014 in the figure), and HD−(ΔspoIIID) strains cultured in SSM medium after T24. The numbers below indicate the Cry1Ac yield in the five strains calculated by Image J software
Discussion
In this study, we found that SpoIIID positively regulates the sigK gene. Also, σK directly or indirectly represses transcription of sigE in B. thuringiensis, based on our finding that PsigE-lacZ expression was higher in spoIIID mutation than in wild type after T5. Hence, transcriptional regulation among σE, SpoIIID, and σK in B. thuringiensis appears to be similar to that in B. subtilis (Halberg and Kroos 1994; Zhang and Kroos 1997; Zhang et al. 1999; Zheng and Losick 1990). In agreement, we found that a spoIIID mutation impairs mother cell lysis in B. thuringiensis. Prior work had shown that σK controls the expression of PcwlC-lacZ, and CwlC is essential for mother cell lysis in B. thuringiensis (Chen et al. 2018; Myasnik et al. 2001; Sanchis et al. 1999). Thus, it is reasonable to expect spoIIID mutation to result in no mother cell lysis. However, we found that the effect of spoIIID mutation depended on the culture medium; HD(ΔspoIIID) mutant did not lyse in SSM and LB (data for LB not shown) but did in ½ LB (Fig. 2).
CwlC is a critical cell wall hydrolase for mother cell lysis that does not decrease the production of Cry protein, insecticidal activity, and mature spore formation (Chen et al. 2018). We further compared the expression of PcwlC-lacZ in cells grown in ½ LB and SSM. Expression from PcwlC-lacZ in the spoIIID mutant was higher in ½ LB than in SSM, but not as high as in wild-type HD73 in either medium (Fig. 3), and correspondingly, lysis of the spoIIID mutant was caused in ½ LB compared with that of wild type (Fig. 2b). A previous study showed that the expression of the PcwlC-lacZ began at T8 in SSM medium (Chen et al. 2018). The difference in mother cell lysis of HD(ΔspoIIID) between ½ LB and SSM likely partly resulted from CwlC accumulation. It seems that the accumulation of CwlC is important for the initiation of mother cell lysis. Although three genes have been found to be involved in mother cell lysis in B. subtilis, the initiation mechanism of mother cell lysis remains unknown (Foster 1992, 1994; Nugroho et al. 1999; Vollmer et al. 2010). Our findings provide a clue for further investigation of this cellular process.
Many previous studies focused on cry gene expression in acrystalliferous B. thuringiensis strains (Amadio et al. 2013; Gómez et al. 2014). Higher-yield Cry3A can be obtained in Bacillus thuringiensis subsp. morrisoni using the sporulation-dependent cyt1Aa promoter to drive the expression of cry3Aa; in Bacillus thuringiensis subsp. jegathesan, increased yield of Cry19A was achieved by applying a recombinant promoter (cyt1A-p/STAB-SD) (Barboza-Corona et al. 2012; Park et al. 1998). The Cry64Ba/Cry64Ca complex and Cry8Kb3 and Cry8Pa3 ovoid crystal proteins as well as Cry30Ca, Cry60Aa, and Cry60Ba can also be expressed in an acrystalliferous B. thuringiensis strain (Liu et al. 2018; Navas et al. 2014; Sun et al. 2013). However, very few studies have investigated cry gene expression in a sporulation mutant. Although many cry genes are sporulation-dependent, such as cry1A, cry1B, cry1C, cry2A, cry4A, cry4B, cry11A, cry18Aa, cry34, and cry40 controlled by σE, σK, or both (Bravo et al. 1996; Dervyn et al. 1995; Huang et al. 2007; Sanchis et al. 1999; Zhang et al. 1998), the strong promoters can be screened to express cry genes in sporulation-deficient mutants. The orf1cry8E promoter and the promoter of the non-cry gene HD73_5014 are the two strongest promoters that can direct cry1Ac expression with high-level transcriptional activity in acrystalliferous sigK mutant (Du et al. 2012; Sanchis et al. 1999; Zhang et al. 2018; Zhou et al. 2014). We confirmed that Porf1cry8E and P5014 in HD(ΔspoIIID) mutant showed the similar expression level as in HD73 strain, and our acrystalliferous and non-spore-forming recipient bacteria HD−(ΔspoIIID) yielded a similar amount of Cry1Ac protein to that of HD73 wild-type strain.
Based on the spoIIID mutant, we further knocked out the gene cwlC, whose deletion completely blocked mother cell lysis (Fig. 4), revealing that spoIIID is an influential factor but cwlC is a key gene for mother cell lysis. Although spoIIID mutation would decrease the expression of some σK-dependent cry genes, the strong σE-dependent promoters can be introduced into spoIIID mutant with special features such as non-spore-forming and medium-dependent mother cell lysis, to drive the expression of σK-dependent cry genes whose production significantly decreased in the spoIIID mutant (Fig. 5c). Thus, spoIIID mutant can be developed as an alternative delivery system for cry gene expression.
References
Agaisse H, Lereclus D (1994) Structural and functional analysis of the promoter region involved in full expression of the cryIIIA toxin gene of Bacillus thuringiensis. Mol Microbiol 13(1):97–107. https://doi.org/10.1111/j.1365-2958.1994.tb00405.x
Amadio AF, Navas LE, Sauka DH, Berretta MF, Benintende GB, Zandomeni RO (2013) Identification, cloning and expression of an insecticide cry8 gene from Bacillus thuringiensis INTA Fr7-4. J Mol Microbiol Biotechnol 23(23):401–409. https://doi.org/10.1159/000353206
Arantes O, Lereclus D (1991) Construction of cloning vectors for Bacillus thuringiensis. Gene 108(1):115–119. https://doi.org/10.1016/0378-1119(91)90495-W
Barboza-Corona JE, Park HW, Bideshi DK, Federici BA (2012) The 60-kilodalton protein encoded by orf2 in the cry19A operon of Bacillus thuringiensis subsp. jegathesan functions like a C-terminal crystallization domain. Appl Environ Microbiol 78(6):2005–2012. https://doi.org/10.1128/AEM.06750-11
Bravo A, Agaisse H, Salamitou S, Lereclus D (1996) Analysis of cryIAa expression in sigE and sigK mutants of Bacillus thuringiensis. Mol Gen Genet MGG 250(6):734–741. https://doi.org/10.1007/BF02172985
Bravo A, Likitvivatanavong S, Gill SS, Soberón M (2011) Bacillus thuringiensis: a story of a successful bioinsecticide. Insect Biochem Mol Biol 41(7):423–431. https://doi.org/10.1016/j.ibmb.2011.02.006
Chen X, Gao T, Peng Q, Zhang J, Chai Y, Song F (2018) The novel cell wall hydrolase CwlC from Bacillus thuringiensis is essential for mother cell lysis. Appl Environ Microbiol 84(7):e02640–e02617. https://doi.org/10.1128/AEM.02640-17
Dervyn E, Poncet S, Klier A, Rapoport G (1995) Transcriptional regulation of the cryIVD gene operon from Bacillus thuringiensis subsp. israelensis. J Bacteriol 177(9):2283–2291. https://doi.org/10.1128/jb.177.9.2283-2291
Du C, Nickerson KW (1996) Bacillus thuringiensis HD-73 spores have surface-localized Cry1Ac toxin: physiological and pathogenic consequences. Appl Environ Microbiol 62(10):3722–3726
Du L, Qiu L, Peng Q, Lereclus D, Zhang J, Song F, Huang D (2012) Identification of the promoter in the intergenic region between orf1 and cry8Ea1 controlled by SigmaH factor. Appl Environ Microbiol 78(12):4164–4168. https://doi.org/10.1128/AEM.00622-12
Eichenberger P, Fujita M, Jensen ST, Conlon EM, Rudner DZ, Wang ST, Ferguson C, Haga K, Sato T, Liu JS, Losick R (2004) The program of gene transcription for a single differentiating cell type during sporulation in Bacillus subtilis. PLoS Biol 2(10):e328. https://doi.org/10.1371/journal.pbio.0020328
Foster SJ (1992) Analysis of the autolysins of Bacillus subtilis 168 during vegetative growth and differentiation by using renaturing polyacrylamide gel electrophoresis. J Bacteriol 174(2):464–470. https://doi.org/10.1128/jb.174.2.464-470.1992
Foster SJ (1994) The role and regulation of cell wall structural dynamics during differentiation of endospore-forming bacteria. J Appl Microbiol 76(S23):25S–39S. https://doi.org/10.1111/j.1365-2672.1994.tb04355.x
Gómez I, Sánchez J, Muñoz-Garay C, Matus V, Gill SS, Soberón M, Bravo A (2014) Bacillus thuringiensis Cry1A toxins are versatile proteins with multiple modes of action: Two distinct pre-pores are involved in toxicity. Biochem J 459(2):383–396. https://doi.org/10.1042/BJ20131408
Halberg R, Kroos L (1994) Sporulation regulatory protein SpoIIID from Bacillus subtilis activates and represses transcription by both mother-cell-specific forms of RNA polymerase. J Mol Biol 243(3):425–436. https://doi.org/10.1006/jmbi.1994.1670
Hoffmann F, Schmidt M, Rinas U (2015) Simple technique for simultaneous on-line estimation of biomass and acetate from base consumption and conductivity measurements in high-cell density cultures of Escherichia coli. Biotechnol Bioeng 70(3):358–361. https://doi.org/10.1002/1097-0290(20001105)70:3<358::AID-BIT14>3.0.CO;2-T
Huang D, Zhang J, Song F, Lang Z (2007) Microbial control and biotechnology research on Bacillus thuringiensis in China. J Invertebr Pathol 95(3):175–180. https://doi.org/10.1016/j.jip.2007.02.016
Ichikawa H, Kroos L (2000) Combined action of two transcription factors regulates genes encoding spore coat proteins of Bacillus subtilis. J Biol Chem 275(18):13849–13,855. https://doi.org/10.1074/jbc.275.18.13849
Kaiser C, Michaelis S, Mitchell A (1998) Methods in yeast genetics: a cold spring harbor laboratory course manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Korz DJ, Rinas U, Hellmuth K, Sanders EA, Deckwer WD (1995) Simple fed-batch technique for high cell density cultivation of Escherichia coli. J Biotechnol 39(1):59–65. https://doi.org/10.1016/0168-1656(94)00143-Z
Kroos L (2007) The Bacillus and Myxococcus developmental networks and their transcriptional regulators. Annu Rev Genet 41(41):13–39. https://doi.org/10.1146/annurev.genet.41.110306.130400
Lereclus D, Arantés O, Chaufaux J, Lecadet MM (1989) Transformation and expression of a cloned endotoxin gene in Bacillus thuringiensis. FEMS Microbiol Lett 51(1):211–217. https://doi.org/10.1111/j.1574-6968.1989.tb03448.x
Liu G, Song L, Shu C, Wang P, Deng C, Peng Q, Lereclus D, Wang X, Huang D, Zhang J, Song F (2013) Complete genome sequence of Bacillus thuringiensis subsp. kurstaki strain HD73. Genome Announc 1(2):e0008013. https://doi.org/10.1128/genomeA.00080-13
Liu Y, Wang Y, Shu C, Lin K, Song F, Bravo A, Soberón M, Zhang J (2018) Cry64Ba and Cry64Ca, two ETX/MTX2-type Bacillus thuringiensis insecticidal proteins active against hemipteran pests. Appl Environ Microbiol 84(3):e01996–e01917. https://doi.org/10.1128/AEM.01996-17
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Myasnik M, Manasherob R, Ben-Dov E, Zaritsky A, Margalith Y, Barak ZE (2001) Comparative sensitivity to UV-B radiation of two Bacillus thuringiensis subspecies and other Bacillus sp. Curr Microbiol 43(2):140–143. https://doi.org/10.1007/s002840010276
Navas LE, Berretta MF, Pérez MP, Amadio AF, Ortiz EM, Sauka DH, Benintende GB, Zandomeni RO (2014) Sequence and expression of two cry8 genes from Bacillus thuringiensis INTA Fr7-4, a native strain from Argentina. J Mol Microbiol Biotechnol 24(4):241–248. https://doi.org/10.1159/000365929
Nugroho FA, Yamamoto H, Kobayashi Y, Sekiguchi J (1999) Characterization of a new SigmaK-dependent peptidoglycan hydrolase gene that plays a role in Bacillus subtilis mother cell lysis. J Bacteriol 181(20):6230–6237
Park HW, Ge B, Bauer LS, Federici BA (1998) Optimization of Cry3A yields in Bacillus thuringiensis by use of sporulation-dependent promoters in combination with the STAB-SD mRNA sequence. Appl Environ Microbiol 64(10):3932–3938
Sanchis V, Gohar M, Chaufaux J, Arantes O, Meier A, Agaisse H, Cayley J, Lereclus D (1999) Development and field performance of a broad-spectrum nonviable asporogenic recombinant strain of Bacillus thuringiensis with greater potency and UV resistance. Appl Environ Microbiol 65(9):4032–4039
Schaeffer P, Millet J, Aubert JP (1965) Catabolic repression of bacterial sporulation. Proc Natl Acad Sci U S A 54(3):704–711. https://doi.org/10.1073/pnas.54.3.704
Schnepf E, Crickmore N, Rie JV, Lereclus D, Baum J, Feitelson J, Zeigler DR, Dean DH (1998) Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol Mol Biol Rev 62(3):775–806
Sun Y, Zhao Q, Xia L, Ding X, Hu Q, Federici BA, Park HW (2013) Identification and characterization of three previously undescribed crystal proteins from Bacillus thuringiensis subsp. jegathesan. Appl Environ Microbiol 79(11):3364–3370. https://doi.org/10.1128/AEM.00078-13
Vollmer W, Joris B, Charlier P, Foster S (2010) Bacterial peptidoglycan (murein) hydrolases. FEMS Microbiol Rev 32(2):259–286. https://doi.org/10.1111/j.1574-6976.2007.00099.x
Wang G, Jie Z, Song F, Wu J, Feng S, Huang D (2006) Engineered Bacillus thuringiensis GO33A with broad insecticidal activity against lepidopteran and coleopteran pests. Appl Microbiol Biotechnol 72(5):924–930. https://doi.org/10.1007/s00253-006-0390-x
Wang X, Du L, Peng Q, Liang Y, Li J, Zhang J, Song F (2012) Activity of four cry gene promoters in spoIIID mutant of Bacillus thuringiensis (in Chinese). Acta Microbiol Sin 52(9):1075–1084
Yang J, Peng Q, Chen Z, Deng C, Shu C, Zhang J, Huang D, Song F (2013) Transcriptional regulation and characteristics of a novel N-acetylmuramoyl-L-alanine amidase gene involved in Bacillus thuringiensis mother cell lysis. J Bacteriol 195(12):2887–2897. https://doi.org/10.1128/JB.00112-13
Zhang B, Kroos L (1997) A feedback loop regulates the switch from one sigma factor to the next in the cascade controlling Bacillus subtilis mother cell gene expression. J Bacteriol 179(19):6138–6144. https://doi.org/10.1128/jb.179.19.6138-6144.1997
Zhang B, Daniel RA, Errington J, Kroos L (1997) Bacillus subtilis SpoIIID protein binds to two sites in the spoVD promoter and represses transcription by SigmaE RNA polymerase. J Bacteriol 179(3):972–975. https://doi.org/10.1128/jb.179.3.972-975.1997
Zhang J, Schairer HU, Schnetter W, Lereclus D, Agaisse H (1998) Bacillus popilliae cry18Aa operon is transcribed by SigmaE and SigmaK forms of RNA polymerase from a single initiation site. Nucleic Acids Res 26(5):1288–1293
Zhang B, Struffi P, Kroos L (1999) σK can negatively regulate sigE expression by two different mechanisms during sporulation of Bacillus subtilis. J Bacteriol 181(13):4081–4088
Zhang Q, Shu C, Zhang J, Huang D, Song F (2009) Characterization of Bacillus thuringiensis spoIIID gene mutantation (in Chinese). Acta Microbiol Sin 49(9):1165–1170
Zhang X, Gao T, Peng Q, Song L, Zhang J, Chai Y, Sun D, Song F (2018) A strong promoter of a non- cry gene directs expression of the cry1Ac gene in Bacillus thuringiensis. Appl Microbiol Biotechnol 102(8):1–13. https://doi.org/10.1007/s00253-018-8836-5
Zheng L, Losick R (1990) Cascade regulation of spore coat gene expression in Bacillus subtilis. J Mol Biol 212(4):645–660. https://doi.org/10.1016/0022-2836(90)90227-D
Zhou C, Zheng Q, Peng Q, Du L, Shu C, Zhang J, Song F (2014) Screening of cry-type promoters with strong activity and application in Cry protein encapsulation in a sigK mutant. Appl Microbiol Biotechnol 98(18):7901–7909. https://doi.org/10.1007/s00253-014-5874-5
Acknowledgments
We thank Dr. Didier Lereclus for his critical discussion and suggestions. We also thank Xiaomin Chen, Linghuan Xu, Fan Yang, Na Li, and Jilong Wen for participating in some of the work.
Funding
This work was supported by grants from the National Natural Science Foundation (No. 31530095) and The National Key Research and Development Program of China (No. 2017YFD0200400).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lv, J., Zhang, X., Gao, T. et al. Effect of the spoIIID mutation on mother cell lysis in Bacillus thuringiensis. Appl Microbiol Biotechnol 103, 4103–4112 (2019). https://doi.org/10.1007/s00253-019-09722-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09722-1