Abstract
l-Amino acid oxidases (LAAOs), which catalyze the stereospecific oxidative deamination of l-amino acids to α-keto acids and ammonia, are flavin adenine dinucleotide-containing homodimeric proteins. l-Amino acid oxidases are widely distributed in diverse organisms and have a range of properties. Because expressing LAAOs as recombinant proteins in heterologous hosts is difficult, their biotechnological applications have not been thoroughly advanced. LAAOs are thought to contribute to amino acid catabolism, enhance iron acquisition, display antimicrobial activity, and catalyze keto acid production, among other roles. Here, we review the types, properties, structures, biological functions, heterologous expression, and applications of LAAOs obtained from microbial sources. We expect this review to increase interest in LAAO studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
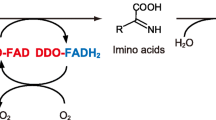
l-Amino acid oxidases or deaminases (LAAOs; EC 1.4.3.2) are widely distributed in mammals (Puiffe et al. 2013; Nakano and Danowski 1966), snake venom (Du and Clemetson 2002), insect venoms (Sakurai et al. 2001), sea hare (Yang et al. 2005), fungi (Davis et al. 2005), bacteria (Huang et al. 2011), and algae (Vallon et al. 1993). As a result, LAAOs form a family of proteins with various enzymatic properties, similar overall structure, and a wide range of biological functions and applications. First described by Zeller and Maritz in (1944), LAAOs are predominantly dimeric flavoproteins with some monomeric types (Yang et al. 2012). They contain non-covalently bound flavin adenine dinucleotide (FAD) as a cofactor (Pawelek et al. 2000). LAAOs catalyze the stereospecific oxidative deamination of l-amino acids to α-keto acids that produces ammonia and H2O2 via an imino acid intermediate (Macheroux et al. 2001). Other LAAOs produce water instead of H2O2 and are associated with a respiratory electron transport chain, donating electrons to the quinone pool that are then transferred to a terminal cytochrome oxidase to reduce molecular oxygen to H2O. Specifically, membrane-bound amino acid oxidases are associated with the respiratory electron transport chain (Anraku and Gennis 1987; Franklin and Venables 1976; Jones and Venables 1983; Olsiewski et al. 1980; Yu and DeVoe 1981). The membrane-associated LAAOs in Proteus mirabilis (Pelmont et al. 1972) and Proteus rettgeri (Duerre and Chakrabarty 1975) are suggested to interact with the respiratory electron transport chain.
Until recently, LAAOs in snake venom had been the most thoroughly investigated member of this enzyme family with respect to toxicology, biochemistry, physiology, and medicine. LAAOs are attracting increasing attention owing to their key biological roles, such as keto acid production, anti-tumor and antimicrobial activity, amino acid consumption, and anti-insect properties. Some of the biological roles of LAAOs are associated with the H2O2 produced (Ehara et al. 2002).
In this review, we provide an overview of the types of LAAOs, their structural, enzymatic, and functional properties, and recent advances in microbial LAAO production and application. Future research directions for the development of cost-effective fermentation processes using LAAOs on an industrial scale are also discussed.
Types of LAAOs from microbial sources
From a biochemical point of view, LAAOs from various sources are distinguished by substrate specificity, molecular mass, regulation, and post-translational modification. This assortment suggests that LAAOs have undergone enormous evolutionary alterations since their split-up from a putative ancestral protein (Macheroux et al. 2001). To date, different types of LAAOs have been reported from various microbes, including bacteria and fungi. Bacteria such as Streptomyces sp. X-119-6, Rhodococcus opacus, lactic acid bacteria, Proteus mirabilis, Proteus vulgaris, Proteus rettgeri, Pseudomonas sp. P-501, Pseudoalteromonas tunicata, Pseudoalteromonas luteoviolacea, Pseudoalteromonas flavipulchra, Marinomonas mediterranea, Rheinheimera sp., Streptococcus oligofermentans, Corynebacterium, Cellulomonas cellulans, Bacillus carotarum, Providencia, Lechevalieria aerocolonigenes, and Synechocystis sp. PCC 6803 produce different types of LAAOs. The fungal LAAOs from Trichoderma viride, Trichoderma harzianum (Cheng et al. 2011), Aspergillus oryzae, Aspergillus flavipes, Aspergillus nidulans, Aspergillus fumigatus (Singh et al. 2009), Neurospora crassa (Sikora and Marzluf 1982), Hebeloma cylindrosporum (Nuutinen et al. 2012), and Laccaria bicolor (Nuutinen and Timoneni 2008) produce various types of oxidases.
Functional, enzymatic, structural properties, and cellular localization of LAAOs
Some functional properties and cellular localization of the different types of LAAOs are shown in Tables 1 and 2, respectively. One of the main biological functions of LAAOs is antimicrobial activity. The Pseudoalteromonas flavipulchra LAAO has broad substrate specificity against methicillin-resistant Staphylococcus aureus that occurs during the stationary phase of the strain growth curve (Chen et al. 2010). The antimicrobial activity of LAAO is likely associated with H2O2 formation generated by enzyme activity because its antimicrobial effect is significantly decreased in the presence of H2O2 scavengers such as catalase and peroxidase. LAAOs can also be intracellular, membrane bound, or extracellular (Table 2) which depend on their sources as well as their corresponding functions.
LAAO of Streptococcus oligofermentans seems to have a role in competition between species (Tong et al. 2008). A function of LAAO from the cyanobacteria Aspergillus nidulans is in electron transfer on the thylakoid membrane (Pistorius and Voss 1982). The LAAOs found in insects are cytotoxic, killing tumor cells via H2O2 generation (Ahn et al. 2000). Similarly, the l-tryptophan oxidase isolated from Chromobacterium violaceum is implicated in the biosynthesis of violacein, which is believed to have bactericidal and tumoricidal activities (Genet et al. 1995). As an immunosuppressive enzyme expressed in a series of 315 human malignancies, l-phenylalanine oxidase could be involved in a general mechanism for the relationship between tumor cells and the immune system (Carbonnelle-Puscian et al. 2009). Moreover, α-keto acids, which are the products of LAAO reactions, have roles as siderophores, as seen in Proteus mirabilis (Massad et al. 1995).
The enzymatic properties of LAAOs depend on their microbial sources, and these properties are usually dissimilar. Generally, the optimum pH of LAAOs depends on substrate, and enzyme activity is relatively stable within the range from 25 to 45 °C (Arinbasarova et al. 2012; Leese et al. 2012; Liu et al. 2013). LAAOs can be inactivated by decreasing the pH and vice versa (Wellner 1966), and inactivation by freezing is more pronounced between −20 and −30 °C (Curti et al. 1968). These characteristics suggest that enzyme inactivation is likely due to conformational changes in the protein structure (Curti et al. 1968). Most LAAOs exhibit a broad range of substrate spectra. They display a noticeable preference for hydrophobic amino acid substrates such as phenylalanine, leucine, tyrosine, and tryptophan (Du and Clemetson 2002; Mandal and Bhattacharyya 2008). This substrate partiality perhaps comes from the binding of hydrophobic side chains of amino acids with the enzyme. The catalysis is proposed to follow one of two diverse mechanisms: in the first pathway, the proton is transferred to the FAD cofactor from the α-carbon atom of the substrate, leaving a negative charge followed by a two-electron transfer which is also called a carbanion pathway; and in the second mechanism, an α hydrogen atom is transferred as a hydride ion carrying two electrons simultaneously by a hydride transfer pathway (Fitzpatrick 2004).
The LAAO isolated from Rhodococcus opacus oxidizes 39 l-amino acids, including not only all of the 20 l-amino acids but also some derivatives (Geueke and Hummel 2002). In addition, LAAOs of Proteus rettgeri (Duerre and Chakrabarty 1975), Mytilus edulis (Blaschko and Hope 1956), Proteus vulgaris (Stumpf and Green 1944), Cellulomonas cellulans (Braun et al. 1992), and Corynebacterium (Coudert and Vandecasteele 1975) exhibit broad substrate specificity. Although a few LAAOs accept d-amino acids as substrates, the reaction rates are relatively low compared with those using l-amino acids. The LAAO from B. carotarum 2Pfa accepts 10 l-amino acids and 7 d-amino acids as substrates (Brearley et al. 1994), among which, 3 aromatic l-amino acids phenyl alanine, tyrosine, and tryptophan are preferred. Some LAAOs, such as those from the fresh water cyanobacteria Synechococcus elongatus PCC 6301, Synechococcus elongatus PCC 7942, Synechococcus cedrorum PCC 6908 (Gau et al. 2007), and Anacystis nidulans (Pistorius and Voss 1980), have relatively narrower substrate specificities, with high preference for only basic l-amino acids. By contrast, several LAAOs have very firm preferences for specific substrates and are named according to their favored substrate—for example, l-glycine oxidase from Bacillus subtilis (Nishiya and Imanaka 1998), l-cysteine oxidase from Neisseria meningitidis (Yu and DeVoe 1981), l-glutamate oxidases from Streptomyces endus (Böhmer et al. 1989) and Streptomyces sp. (Arima et al. 2009), and l-lysine oxidases from T. viride (Kusakabe et al. 1980), Pseudoalteromonas tunicata (James et al. 1996), and Marinomonas mediterranea (Lucas-Elio et al. 2005).
Structural knowledge about microbial LAAOs is extremely limited. LAAO from Rhodococcus opacus has been crystallized and examined with preliminary x-ray analysis (Faust et al. 2006, 2007). LAAOs have a range of isoelectric points varying widely from pH 4.0 to 9.4 and molecular masses between approximately 50 and 300 kDa. In general, LAAOs are first synthesized as precursors carrying a signal peptide and then form mature proteins via post-translational modification through limited proteolysis, possibly owing to their potential toxicity (Ahn et al. 2000). The LAAO activity that catalyzes the stereospecific oxidation of l-amino acids along with the release of ammonia and H2O2 is speculated to be toxic to or have negative effects on the growth of host cells. Therefore, LAAO exists in cells as a precursor with low initial activity and then is secreted with high activity from cells after digestion by an endopeptidase. For example, the LAAO from Neurospora crassa consists of 695 amino acids as a precursor, a length of 129 amino acids longer than that of the mature enzyme (Neidermann and Lerch 1990). Similarly, the mature form of l-phenylalanine oxidase from Pseudomonas sp. P-501 is produced and triggered by the proteolytic cleavage of a noncatalytic proenzyme (Ida et al. 2008). The Rhodococcus opacus LAAO gene shows an open reading frame coding for 534 amino acids, including a signal peptide of 45 amino acids (Geueke and Hummel 2002). Along with these structures, l-glutamate oxidase (LGOX) from Rhodococcus opacus, LAAO from Streptomyces sp. X-119-6 (Arima et al. 2009), l-aspartate oxidase from Escherichia coli (Bossi et al. 2002), and l-phenyl alanine oxidase from Pseudomonas sp. P501 (Ida et al. 2008; 2011) have been reported. A topological view of those structures shows that each LAAO subunit consists of three domains: one for FAD binding, one for substrate binding, and a helical domain. For example, the structure of l-aspartate oxidase includes a FAD-binding domain with the classic dinucleotide-binding fold observed in many flavoproteins, a capping domain with an irregular α + β topology, and a helical C-terminal domain. Although the structures of these enzymes are similar to a certain extent, they have several differences that make them unique. In the LGOX structure, two funnel-shaped entrances with distinct functions lead from the surface to the active site (Arima et al. 2009). The structure of l-phenyl alanine oxidase suggests that the pro-sequence peptide occupies the funnel of the substrate amino acid from the outside of the protein to the interior flavin ring (Ida et al. 2008). However, no funnel orientating and directing the substrates to the active site is present in the LAAO from Rhodococcus opacus. This funnel conformation and the structure of the enzyme are probably associated with substrate specificity, and the absence of a funnel may explain the broader substrate specificity of the Rhodococcus opacus LAAO (Faust et al. 2006). Moreover, LGOX has a hexameric structure, α2β2γ2, and by recombinant expression, it has been shown to have a homodimeric structure of its precursor (Arima et al. 2003).
Gene cloning and heterologous expression of LAAOs
The first bacterial heterologous expression of an LAAO of Rhodococcus opacus was reported by Geueke and Hummel (2003). The lao gene encoding LAAO was cloned into E. coli, B. subtilis, and Streptomyces lividans expression vectors. Expression in E. coli results in the accumulation of insoluble protein, but Streptomyces lividans is a relatively suitable host for the heterologous production of LAAO (Geueke and Hummel 2003; Liu et al. 2013). Comparison of native and recombinant LAAOs has shown the same specific activities (Liu et al. 2013), similar spectral properties, and the same molecular mass. The LAAO isolated from sea hare, which is not a microbial source, was first functionally expressed in E. coli, but the expression level for soluble recombinant LAAO is relatively low—approximately 0.2 mg/L culture medium (Yang et al. 2005). This phenomenon has several explanations: (1) much of the LAAO is present in insoluble inclusion bodies, and (2) LAAO inhibits the growth of E. coli and other bacteria at doses below 1 mg/L, which also likely inhibits the level of bacterial expression. Other LAAO genes from Streptococcus oligofermentans and Proteus mirabilis have been successfully cloned and overexpressed in E. coli BL21(DE3) strain (Liu et al. 2013; Tong et al. 2008). The functional expression of LAAO in heterologous hosts remains an enormous challenge.
To date, a wide variety of LAAO-encoding sequences have been published, revealing that LAAO family members have flavin as a common coenzyme, with the exception of the lysine oxidase of Marinomonas mediterranea (Lucas-Elio et al. 2006) and two characteristic sequence motifs. One is a dinucleotide-binding motif comprising a β-strand/α-helix/β-strand of the secondary structure; the other is a GG motif (x-G-G-R-x-x) positioning shortly after the dinucleotide-binding motif (Vallon 2000). In 1990, the Neurospora crassa LAAO gene was cloned and sequenced (Neidermann and Lerch 1990). The Synechococcus PCC6301 LAAO nucleotide sequence (Bockholt et al. 1995) infers that the protein consists of 355 amino acid residues showing no homology with the LAAO of Neurospora crassa (Neidermann and Lerch 1990), in which the expression of LAAO can be induced via addition of l-amino acids to nitrogen-starved cultures as well as addition of protein synthesis inhibitors or d-amino acids (DeBusk and Ogilvie 1984a, 1984b; Neidermann and Lerch 1991; Sikora and Marzluf 1982). Its gene expression is controlled by NIT2 and the nmr gene product, and the LAAO gene is regulated at the transcriptional level. Likely because of the post-translational modification requirement of LAAO, only some heterologous expression systems have been reported.
Methods for measuring LAAO activity and high-throughput screening
A variety of methods can be used to measure LAAO activity, including capacity of production of compounds such as ammonia (Timmer et al. 2005), α-keto acid (Singh et al. 2009), and H2O2 (Okubo et al. 2012); measurement of oxygen consumption using a classic Warburg manometer or an oxygen-sensitive electrode (Ralph et al. 2006); and measurement of amino acid substrate change (Roth 1971). All of these methods have both benefits and drawbacks. Although detection methods for the measurement of H2O2 production have been most widely used for LAAOs assays in which horseradish peroxidase (HRP) acts as an H2O2-sensitive probe, in-gel detection of l-amino acid oxidases based on the visualization of H2O2 production is a reasonable and reproducible method (Rau and Fischer 2011; Yu et al. 2013). However, most HRP substrates, including 2,2′-azinobis (3-ethylbenzthiazoline-6) sulfonic acid, o-phenylenediamine, and o-dianisidine are tremendously toxic or carcinogenic, and even HRP is easily inactivated and costly. Moreover, high-throughput screening (HTS) is essential for the production of mutant LAAOs with improved activity and specificity for certain l-amino acids. Although mutagenesis techniques such as directed evolution and gene mosaics have become prevalent, the sensitivity and selectivity of direct enzyme assay has not advanced. In some cases, a peroxidase/o-dianisidine-based assay has been used to screen the library of mutant clones on microplates, but the method is hazardous, labor-intensive, time-consuming, and requires multiple handling of each clone—namely, cultivation in microplate wells, cell transfer to assay microplates, cell lysis, cell separation, and activity determination. Therefore, a suitable method must be developed for the HTS assay of LAAO that can detect micro-quantities of the enzyme in biological fluids and intact cell colonies and is also fast and cost-effective.
Applications of LAAOs
LAAOs have important potential biotechnological applications as components of biosensors (Mutaguchi et al. 2011; Pollegioni and Molla 2011). The fast and precise determination of amino acid concentrations has become increasingly important for a varied range of applications including medical, biological, and food technological analyses. For example, in case of phenylketonuria (PKU), which is a disorder of phenylalanine metabolism (Huang et al. 1998), abnormal levels of phenylalanine can be used to diagnose. In case of bioprocess optimization, amino acid concentrations can be used to monitor and control the state of fermentation processes (Varadi et al. 1999). In case of animal nutrition, concentrations of essential amino acids have a vital influence on animal weight gain. Although, amino acids can be measured by liquid- or gas-chromatography and spectrometric and colorimetric methods (Sarkar et al. 1999), the complex laboratory procedures, expensive equipment, and a reasonable time for the measurements are drawbacks for all of those methods (Pei and Li 2000). These problems can be overcome by using a simple, rapid and reliable amino acid biosensor based on the LAAOs. l-amino acids can be determined using an LAAO/platinum biosensor (Chauhan et al. 2013). l-Amino acid concentrations have been measured through amperometric measurement of oxygen consumed by LAAO using specific potentials and electrode systems (Kacaniklic et al. 1994; Simonian et al. 1994) or hydrogen peroxide produced (Preuschoff et al. 1993). For rapid determination of the concentration of l-amino acids, a unique amperometric biosensor has been developed using LAAO by immobilizing with poly(carbamoyl) sulfonate hydrogel (Kwan et al. 2002). LAAOs have also been used as biosensors in the food industry to determine the quality of many products by their l-amino acid content (Lee and Huh 1999; Varadi et al. 1999), in particular, l-lys in maize, rice, and wheat. Although, there are some drawbacks of LAAO sensors due to their various sensitivities to different substrates and the variations in their kinetically limited sensitivity to particular amino acids (Scheller and Schubert 1989), their applicability is much simpler and cost-effective compared to other measuring methods. In food processing, LAAOs can be applied in the enzymatic treatment of dough to reduce viscosity and provide better workability (Christiansen and Budolfsen 2002).
LAAOs can be used in textile bleaching and it has been observed that the bleaching effect of hydrogen peroxide, which is produced enzymatically by LAAOs in typical washing, the bleaching and cleaning process can be considerably improved, even in the absence of any activators. A novel LAAO of T. harzianum in combination with its substrate has been used for in situ generation of hydrogen peroxide in the textile bleaching and may be favorably incorporated into detergent compositions that include a peroxidase for preventing the relocation of dye from dyed fabric to other fabrics during washing (Schneider et al. 1998). It also can be mixed with toothpaste or used for cosmetic preservation. Moreover, the enzyme has been mentioned in applications for wastewater treatment, pulp bleaching in the paper industry, water treatment in pulp production, and lignin improvement for particle board production (Schneider et al. 1998).
LAAOs have important potential biotechnological applications as catalysts in biotransformations of l-amino acids (Mutaguchi et al. 2011; Pollegioni and Molla 2011). LAAO from Proteus mirabilis can be used in the production of α-ketoglutarate from l-glutamic acid (Hossain et al. 2013; Liu et al. 2013). The LAAO of Rhodococcus sp. is used in bioconversion to synthesize amino adipic derivatives that are precursors for β-lactam antibiotics (Isobe et al. 2008) and for the biotransformation of l-DOPA to its α-keto acid (Findrik et al. 2006). LAAOs from Providencia alcalifaciens and T. viride have been applied as a catalyst in the biotransformation of Nε-carboxy(CBZ)- l-Lys into the corresponding keto acid (Hanson et al. 1992).
Since optically pure d-amino acids have industrial significance as chiral building blocks for the synthesis of pharmaceuticals, food ingredients, and drug intermediates, LAAOs can be used to separate enantiomers from racemic mixtures of d-/l-amino acids (Jang et al. 2012; Qi et al. 2009; Singh et al. 2009). LAAOs can also be useful as anti-cancer and anti-tumor drugs. For example, a Trichoderma/l-lysine oxidase has been tested in vivo on several tumors and cancers and presented encouraging data on the promising application of this LAAO in clinical oncology for patients with colorectal cancer (Pokrovsky et al. 2013; Treshalina et al. 2000). Studies have also proved the efficacy of sequential pretreatment with LAAO and LAAO antiserum in the modulation of melphalan activity against intracranial glioma (Moynihan et al. 1997).
LAAOs can be used as biofertilizers for nitrogen acquisition and as biocontrol agents. LAAOs that show the capability to release nitrogen from amino acids (Nuutinen and Timoneni 2008) are present in many soil microorganisms with a variety of lifestyles. Therefore, LAAO-producing microorganisms are potential candidates for biological control or biofertilizers. Most LAAOs from microorganisms have broad substrate spectra, making them ideal molecular mechanisms for the acquisition of nitrogen from diverse amino acid sources. For example, an LAAO of Neurospora crassa can be induced by l-Arg and l-Phe in the absence of readily metabolizable nitrogenous compounds (DeBusk and Ogilvie 1984c). Another LAAO of Chlamydomonas reinhardtii is also inducible if no primary nitrogen source is available to produce ammonia (Muñoz-Blanco et al. 1990). In 2005, Davis et al. found that the LAAO of Anacystis nidulans is the primary route of catabolism for amino acids such as l-cysteine, l-histidine, l-leucine, l-lysine, l-methionine, l-α-amino butyrate, and l-citrulline for growth (Davis et al. 2005). LAAOs expressed in Hebeloma spp. and Laccaria bicolor can also catalyze the mineralization of amino acid nitrogen to NH4 + (Nuutinen and Timoneni 2008). The released inorganic nitrogen might then be re-assimilated and distributed to organic compounds, which complements the previously identified amino acid catabolic machinery, e.g., glutamate dehydrogenase, aspartate aminotransferase, and alanine aminotransferase. In general, bacterial LAAOs are assumed to be necessary in nitrogen and l-amino acid metabolism for the plants. The necessary steps and research perspectives to develop the LAAOs for the future applications are summarized in Fig. 1.
Conclusions and perspectives
The microbial production and application of LAAOs is gaining increasing attention, and noteworthy progress has been made to date. In future research, several key topics might be considered. First, a crucial factor for the successful development of biocatalytic processes is the possibility of using an inexpensive production enzyme with suitable properties (i.e., high activity, stability, and selectivity) on a large-scale. Developing whole-cell biocatalysts with evolvable properties is a potentially effective strategy to achieve this objective in industrial processes (Hossain et al. 2013). With whole-cell biocatalysts, the stability of the dimeric LAAOs could be increased, and downstream enzyme processing costs could be decreased significantly. Second, although the production of LAAOs is low in recombinant E. coli, the gram-positive bacteria B. subtilis and Streptomyces lividans may be promising overproducers of LAAOs for industrial use. Finally, to achieve superior LAAO utilization in biotechnological production, structural and protein engineering with mutagenesis techniques should be applied efficiently.
References
Ahn MY, Ryu KS, Lee YW, Kim YS (2000) Cytotoxicity and l-amino acid oxidase activity of crude insect drugs. Arch Pharm Res 23:477–481
Anraku Y, Gennis RB (1987) The aerobic respiratory chain of Escherichia coli. Trends Biochem Sci 12:262–266
Arima J, Tamura T, Kusakabe H, Ashiuchi M, Yagi T, Tanaka H, Inagaki K (2003) Recombinant expression, biochemical characterization and stabilization through proteolysis of an l- glutamate oxidase from Streptomyces sp. X-119-6. J Biochem 134:805–812
Arima J, Sasaki C, Sakaguchi C, Mizuno H, Tamura T, Kashima A, Kusakabe H, Sugio S, Inagaki K (2009) Structural characterization of l-glutamate oxidase from Streptomyces sp. X-119-6. FEBS J 276:3894–3903
Arinbasarova A, Ashin VV, Makrushin KV, Medentsev AG, Lukasheva EV, Berezov TT (2012) Isolation and properties of l-lysine-alpha-oxidase from the fungus Trichoderma cf. aureoviride RIFAI VKM F-4268D tion. Mikrobiologiia 81:594–599
Baek JO, Seo JW, Kwon O, Seong S, Kim IH, Kim CH (2008) Heterologous expression and characterization of l-amino acid deaminase from Proteus mirabilis in Escherichia coli. Chi J Biotech 24:21–29
Baek JO, Seo JW, Kwon O, Seong SI, Kim IH, Kim CH (2011) Expression and characterization of a second l-amino acid deaminase isolated from Proteus mirabilis in Escherichia coli. J Basic Microbiol 51:129–135
Blaschko H, Hope DB (1956) The oxidation of l-amino acids by Mytilus edulis. Biochem J 62:335–339
Bockholt R, Masepohl B, Kruft V, Wittmann-Liebold B, Pistorius EK (1995) Partial amino acid sequence of a l-amino acid oxidase from the cyanobacterium Synechococcus PCC6301, cloning and DNA sequence analysis of the aoxA gene. Biochim Biophys Acta 1264:289–293
Böhmer A, Müller A, Passarge M, Liebs P, Honeck H, Müller HG (1989) A novel l-glutamate oxidase from Streptomyces endus purification and properties. Eur J Biochem 182:327–332
Bossi RT, Negri A, Tedeschi G, Mattevi A (2002) Structure of FAD-bound l-aspartate oxidase: insight into substrate specificity and catalysis. Biochem 41:3018–3024
Braun M, Kim JM, Schmid RD (1992) Purification and some properties of an extracellular l-amino acid oxidase from Cellulomonas cellulans AM8 isolated from soil. Appl Microbiol Biotechnol 37:594–598
Brearley GM, Price CP, Atkinson T, Hammond PM (1994) Purification and partial characterization of a broad-range l-amino acid oxidase from Bacillus carotarum 2Pfa isolated from soil. Appl Microbiol Biotechnol 41:670–676
Carbonnelle-Puscian A, Copie-Bergman C, Baia M, Martin-Garcia N, Allory Y, Haioun C, Crémades A, Abd-Alsamad I, Farcet JP, Gaulard P, Castellano F, Molinier-Frenkel V (2009) The novel immunosuppressive enzyme IL4I1 is expressed by neoplastic cells of several B-cell lymphomas and by tumor associated macrophages. Leukemia 23:952–960
Chauhan N, Narang J, Sunny, Pundir CS (2013) Immobilization of lysine oxidase on a gold-platinum nanoparticles modified Au electrode for detection of lysine. Enzyme Microb Technol 52:265–271
Chen WM, Lin CY, Che CA, Wang JT, Sheu SY (2010) Involvement of an l-amino acidoxidase in the activity of the marine bacterium Pseudoalteromonas flavipulchra against methicillin resistant Staphylococcus aureus. Enzym Microb Tech 47:52–58
Cheng CH, Yang CA, Liu SY, Lo CT, Huang HC, Liao FC, Peng KC (2011) Cloning of a novel l-amino acid oxidase from Trichoderma harzianum ETS 323 and bioactivity analysis of overexpressed l-amino acid oxidase. J Agric Food Chem 59:9142–9149
Christiansen L, Budolfsen G (2002) Preparation of baked product from dough. US patent 6,890,570, 10 May 2005.
Coudert M, Vandecasteele JP (1975) Characterization and physiological function of a soluble l- amino acid oxidase in Corynebacterium. Arch Microbiol 102:151–153
Curti B, Massey V, Zmudka M (1968) Inactivation of snake venom l-amino acid oxidase by freezing. J Biol Chem 243:2306–2314
Davis MA, Askin MC, Hynes MJ (2005) Amino acid catabolism by an area-regulated gene encoding an l-amino acid oxidase with broad substrate specificity in Aspergillus nidulans. Appl Environ Microbiol 71:3551–3555
DeBusk RM, Ogilvie S (1984a) Participation of an extracellular deaminase in amino acid utilization by Neurospora crassa. J Bacteriol 159:583–589
DeBusk RM, Ogilvie S (1984b) Nitrogen regulation of amino acid utilization by Neurospora crassa. J Bacteriol 160:493–498
DeBusk RM, Ogilvie S (1984c) Regulation of amino acid utilization in Neurospora crassa: effect of nmr-1 and ms-5 mutations. J Bacteriol 60:656–661
Du XY, Clemetson KJ (2002) Snake venom L-amino acid oxidases. Toxicon 40:659–665
Duerre JA, Chakrabarty S (1975) l-amino acid oxidases of Proteus rettgeri. J Bacteriol 121:656–663
Ehara T, Kitajima S, Kanzawa N, Tamiya T, Tsuchiya T (2002) Antimicrobial action of achacin is mediated by l-amino acid oxidase activity. FEBS Lett 531:509–512
Faust A, Geueke B, Niefind K, Hummel W, Schomburg D (2006) Crystallization and preliminary x-ray analysis of a bacterial l-amino-acid oxidase from Rhodococcus opacus. Acta Crystallogr Sect F: Struct Biol Cryst Commun 62:279–281
Faust A, Niefind K, Hummel W, Schombur D (2007) The structure of a bacterial l-amino acid oxidase from Rhodococcus opacus gives new evidence for the hydride mechanism for dehydrogenation. J Mol Biol 367:234–248
Findrik Z, Geueke B, Hummel W, Vasić-Rački D (2006) Modelling of LDOPA enzymatic oxidation catalyzed by l-amino acid oxidases from Crotalus adamanteus and Rhodococcus opacus. Biochem Eng J 27:275–286
Fitzpatrick PF (2004) Carbanion versus hydride transfer mechanisms in flavoprotein-catalyzed dehydrogenations. Bioorg Chem 32:125–139
Franklin FC, Venables WA (1976) Biochemical, genetic, and regulatory studies of alanine catabolism in Escherichia coli K12. Mol Gen Genet 149:229–237
Gau AE, Heindl A, Nodop A, Kahmann U, Pistorius EK (2007) l-Amino acid oxidases with specificity for basic l-amino acids in cyanobacteria. Z Naturforsch C 62:273–284
Genet R, Bénetti PH, Hammadi A, Ménez A (1995) l-Tryptophan 2′,3′-oxidase from Chromobacterium violaceum. J Biol Chem 270:23540–23545
Geueke B, Hummel W (2002) A new bacterial l-amino acid oxidase with a broad substrate specificity: purification and characterization. Enzym Microb Technol 31:77–87
Geueke B, Hummel W (2003) Heterologous expression of Rhodococcus opacus l-amino acid oxidase in Streptomyces lividans. Protein Expr Purif 28:303–309
Hanson RL, Bembenek KS, Patel RN, Szarka LJ (1992) Transformation of N epsilon-CBZ- l-lysine to CBZ-l-oxylysine using l-amino acid oxidase from Providencia alcalifaciens and l-2-hydroxy-isocaproate dehydrogenase from Lactobacillus confusus. Appl Microbiol Biotechnol 37:599–603
Hossain GS, Liu L, Shin HD, Du G, Chen J (2013) Bioconversion of α-ketoglutaric acid to l-glutamic acid by an immobilized whole-cell biocatalyst expressing l-amino acid deaminase from Proteus mirabilis. J Biotechnol. doi:10.1016/j.jbiotec.2013.10.026
Huang T, Warsinke A, Kuwana T, Scheller FW (1998) Determination of l-phenylalanine based on an NADH-detecting biosensor. Anal Chem 70:991–997
Huang YL, Li M, Yu Z, Qian PY (2011) Correlation between pigmentation and larval settlement deterrence by Pseudoalteromonas sp. sf57. Biofouling 27:287–293
Ida K, Kurabayashi M, Suguro M, HirumaY HT, Yamomoto M, Suzuki H (2008) Structural basis of proteolytic activation of l-phenylalanine oxidase from Pseudomonas sp. P-501. J Biol Chem 24:16584–16590
Ida K, Suguro M, Suzuki H (2011) High resolution x-ray crystal structures of l-phenylalanine oxidase (deaminating and decarboxylating) from Pseudomonas sp. P-501. Structures of the enzyme-ligand complex and catalytic mechanism. J Biol Chem 150:659–669
Isobe K, Fukuda N, Nagasawa S (2008) Analysis of selective production of N-alpha-benzyloxycarbonyl- l- aminoadipate-delta-semialdehyde and N-alpha-benzyloxycarbonyl- l-aminoadipic acid by Rhodococcus sp. AIU Z-35-1. J Biosci Bioeng 105:152–156
James SG, Holmstrom C, Kjelleberg S (1996) Purification and characterization of a novel antibacterial protein from marine bacterium D2. Appl Environ Microbiol 62:2783–2788
Jang D, Kim K, Park D, Kim G (2012) Efficient optical resolution of amino acid by alanine racemaze chiral analogue supported on mesoporous carbon. IOP Conf Ser: Mater Sci Eng 40:012024
Jones H, Venables WA (1983) Effects of solubilization on some properties of the membrane-bound respiratory enzyme D-amino acid dehydrogenase of Escherichia coli. FEBS Lett 151:189–192
Kacaniklic V, Johansson K, Marko-Varga G, Gorton L, Jonssön- Pettersson G, Csöregi E (1994) Amperometric biosensors for detection of l- and D-amino acids based on coimmobilized peroxidase and l-and D-amino acid oxidases in carbon paste electrodes. Electroanalysis 6:381–390
Koyama H (1982) Purification and characterization of a novel l-phenylalanine oxidase (Deaminating and decarboxylating) from Pseudomonas sp. P-501. J Bio Chem 92:1235–1240
Kusakabe H, Kodama K, Kuninaka A, Yoshino H (1980) A new antitumor enzyme, l-lysine oxidase from Trichoderma viride: purification and enzymological properties. J Biol Chem 255:976–981
Kwan RCH, Chan CY, Renneberg R (2002) An amperometric biosensor for determining amino acids using a bienzymatic system containing amino acid oxidase and protease. Biotechnol Lett 24:1203–1207
Lee YC, Huh MH (1999) Development of a biosensor with immobilized l-amino acid oxidase for determination of l-amino acids. J Food Bio Chem 23:173–185
Leese C, Fotheringham I, Escalettes F, Speight R, Grogan G (2012) Cloning, expression, characterization and mutational analysis of l-aspartate oxidase from Pseudomonas putida. J Mol Catal B Enzym 85–86:17–22
Liu L, Hossain GS, Shin HD, Li J, Du G, Chen J (2013) One-step production of α-ketoglutaric acid from glutamic acid with an engineered l-amino acid deaminase from Proteus mirabilis. J Biotechnol 164:97–104
Lucas-Elio P, Hernandez P, Sanchez-Amat A, Solano F (2005) Purification and partial characterization of marinocine, a new broad-spectrum antibacterial protein produced by Marinomonas mediterranea. Biochim Biophys Acta 1721:193–203
Lucas-Elio P, Gomez D, Solano F, Sanchez-Amat A (2006) The antimicrobial activity of marinocine, synthesized by Marinomonas mediterranea, is due to hydrogen peroxide generated by its lysine oxidase activity. J Bacteriol 188:2493–2501
Macheroux P, Seth O, Bollschweiler C, Schwarz M, Kurfürst M, Au LC, Ghisla S (2001) l-amino-acid oxidase from the Malayan pit viper Calloselasma rhodostoma. Comparative sequence analysis and characterization of active and inactive forms of the enzyme. Eur J Biochem 268:1679–1686
Mandal S, Bhattacharyya D (2008) Two l-amino acid oxidase isoenzymes from Russell's viper (Daboia russelli russelli) venom with different mechanism of inhibition by substrate analogs. FEBS J 275:2078–2095
Massad G, Zhao H, Mobley HL (1995) Proteus mirabilis amino acid deaminase: cloning, nucleotide sequence, and characterization of aad. J Bacteriol 177:5878–5883
Moynihan K, Elion BG, Pegram C, Reist CJ, Wellner D, Bigner DD, Griffith OW, Friedman HS (1997) l-amino acid oxidase (LOX) modulation of melphalan activity against intracranial glioma. Cancer Chemother Pharmacol 39:179–186
Muñoz-Blanco J, Hidalgo-Martínez J, Cárdenas J (1990) Extracellular deamination of l-amino acids by Chlamydomonas reinhadtii cells. Planta 182:194–198
Mutaguchi Y, Ohmori T, Sakuraba H, Yoneda K, Doi K, Ohshima T (2011) Visible wavelength spectrophotometric assays of l-aspartate and D-aspartate using hyperthermophilic enzyme systems. Anal Chem 409:1–6
Nakano M, Danowski TS (1966) Crystalline mammalian l-amino acid oxidase from rat kidney mitochondria. J Biol Chem 241:2075–2083
Neidermann DM, Lerch K (1990) Molecular cloning of the l-amino-acid oxidase gene from Neurospora crassa. J Biol Chem 265:17246–17251
Neidermann DM, Lerch K (1991) Regulation of biosynthesis of l-amino acid oxidase by Neurospora crassa. FEMS Microbiol Lett 79:309–314
Nishiya Y, Imanaka T (1998) Purification and characterization of a novel glycine oxidase from Bacillus subtilis. FEBS Lett 438:263–266
Nuutinen JT, Timoneni S (2008) Identification of nitrogen mineralization enzymes, l-amino acidoxidases, from the ectomycorrhizal fungi Hebeloma spp. and Laccaria bicolor. Mycol Res 112:1453–1464
Nuutinen JT, Marttinen E, Soliymani R, Hildén K, Timonen S (2012) l-amino acid oxidase of the fungus Hebeloma cylindrosporum displays substrate preference towards glutamate. Microbiol 158:272–283
Okubo BM, Silva ON, Migliolo L, Gomes DG, Porto WF, Batista CL, Ramos CS, Holanda HH, Dias SC, Franco OL, Moreno SE (2012) Evaluation of an antimicrobial l-amino acid oxidase and peptide derivatives from Bothropoides mattogrosensis pit viper venom. PLoS ONE 7:e33639
Olsiewski PJ, Kaczorowski GJ, Walsh C (1980) Purification and properties of D-amino acid dehydrogenase, an inducible membrane bound iron-sulfur flavoenzyme from Escherichia coli. J Biol Chem 255:4487–4494
Pawelek PD, Cheah J, Coulombe R, Macheroux P, Ghisla S, Vrielink A (2000) The structure of l-amino acid oxidase reveals the substrate trajectory into an enantiomerically conserved active site. EMBO J 19:4204–4215
Pei J, Li X (2000) Determination of underivatized amino acids by high-performance liquid chromatography and electrochemical detection at an amino acid oxidase immobilized CuPtCl6 modified electrode. Fresenius J Anal Chem 367:707–713
Pelmont J, Arlaud G, Rossat AM (1972) l-amino acid oxydases des envelopes de Proteus mirabilis: proprie’ te’sge’ nerales. Biochemie 54:1359–1374
Pistorius EK, Voss H (1980) Some properties of a basic l-amino-acid oxidase from Anacystis nidulans. Biochim Biophys Acta 611:227–240
Pistorius EK, Voss H (1982) Presence of an amino acid oxidase in photosystem II of Anacystis nidulans. Eur J Biochem 126:203–209
Pokrovsky VS, Treshalina HM, Lukasheva EV, Sedakova LA, Medentzev AG, Arinbasarova AY, Berezov TT (2013) Enzymatic properties and anticancer activity of l-lysine α-oxidase from Trichoderma cf. aureoviride Rifai BKMF-4268D. Anticancer Drugs 24:846–851
Pollegioni L, Molla G (2011) New biotech applications from evolved D-amino acid oxidases. Trends Biotechnol 29:276–283
Preuschoff F, Spohn U, Weber E, Unverhau K, Mohr KH (1993) Chemiluminometric l-lysine determination with immobilized lysine oxidase by flow-injection analysis. Anal Chimi Acta 280:185–189
Puiffe ML, Lachaise I, Molinier-Frenkel V, Castellano F (2013) Antibacterial properties of the mammalian l-amino acid oxidase IL4I1. PLoS ONE 8:e54589
Qi L, Qiao J, Yang G, Chen Y (2009) Chiral ligand-exchange CE assays for separation of amino acid enantiomers and determination of enzyme kinetic constant. Electrophoresis 30:2266–2272
Ralph EC, Anderson MA, Cleland WW, Fitzpatrick PF (2006) Mechanistic studies of the flavoenzyme tryptophan 2-monooxygenase: deuterium and 15 N kinetic isotope effects on alanine oxidation by a l-amino acid oxidase. Biochem 45:15844–15852
Rau JE, Fischer U (2011) In-gel detection of l-amino acid oxidases based on the visualization of hydrogen peroxide production. J Microbiol Methods 85:228–229
Roth M (1971) Fluorescence reaction for amino acids. Anal Chem 43:880–882
Sakurai Y, Takatsuka H, Yoshioka A, Matsui T, Suzuki M, Titani K, Fujimura Y (2001) Inhibition of human platelet aggregation by l-amino acid oxidase purified from Naja naja kaouthia venom. Toxicon 39:1827–1833
Sarkar P, Tothill IE, Setford SJ, Turner AP (1999) Screen-printed amperometric biosensors for the rapid measurement of l-and D-amino acids. Analyst 124:865–870
Scheller FW, Schubert F (1989) Biosensors, 2nd edn. Amsterdam: Elsevier, pp. 157–159
Schneider P, Pedersen AH, Hansen SA (1998) l-amino acid oxidase. US Patent 5,801,035, 1 Sept 1998
Sikora L, Marzluf GA (1982) Regulation of l-amino acid oxidase and D-amino acid oxidase in Neurospora crassa. Mol Gen Genet 186:33–39
Simonian AL, Badalian IE, Berezov TT, Smirnova IP, Khaduev SH (1994) Flow-injection amperometric biosensor based on immobilized l-lysine-a-oxidase for l-lysine determination. Anal Lett 27:2849–2860
Singh S, Gogoi BK, Bezbaruah RL (2009) Optimization of medium and cultivation conditions for l-amino acid oxidase production by Aspergillus fumigatus. Can J Microbiol 55:1096–1102
Stumpf PK, Green DE (1944) l-amino acid oxidase of Proteus vulgaris. J Biol Chem 153:387–399
Takahashi E, Ito K, Yoshimoto T (1999) Cloning of l-amino acid deaminase gene from Proteus vulgaris. Biosci Biotechnol Biochem 63:2244–2247
Timmer B, Olthuis W, van den Berg A (2005) Ammonia sensors and their applications—a review. Sensor Actuat B 107:666–677
Tong H, Chen W, Shi W, Qi F, Dong X (2008) SO-LAAO, a novel l-amino acid oxidase that enables Streptococcus oligofermentans to outcompete Streptococcus mutants by generating H2O2 from peptone. J Bacteriol 190:4716–4721
Treshalina HM, Lukasheva EV, Sedakova LA, Firsova GA, Guerassimova GK, Gogichaeva NV, Berezov TT (2000) Anticancer enzyme l-lysine α-oxidase: properties, application and perspectives. Appl Biochem Biotechnol 88:267–273
Vallon O (2000) New sequence motifs in flavoproteins: evidence for common ancestry and tools to predict structure. Proteins 38:95–114
Vallon O, Bulte L, Kuras R, Olive J, Wollman FA (1993) Extensive accumulation of an extracellular l-amino-acid oxidase during gametogenesis of Chlumydomonas reinhardtii. Eur J Biochem 215:351–360
Varadi M, Adányi N, Szabó EE, Trummer N (1999) Determination of the ratio of D- and l-amino acids in brewing by an immobilized amino acid oxidase enzyme reactor coupled to amperometric detection. Biosens Bioelectron 14:335–340
Wellner D (1966) Evidence for conformational changes in l-amino acid oxidase associated with reversible inactivation. Biochemical 5:1585–1591
Yang H, Johnson PM, Ko KC, Kamio M, Germann MW, Derby CD, Tai PC (2005) Cloning, characterization and expression of escapin, a broadly antimicrobial FAD-containing l-amino acid oxidase from ink of the sea hare Aplysia californica. J Exp Biol 208:609–622
Yang CA, Cheng CH, Lo CT, Liu SY, Lee JW, Peng KC (2011) A novel l-amino acid oxidase from Trichoderma harzianum ETS 323 associated with antagonism of Rhizoctonia solani. J Agric Food Chem 59:4519–4526
Yang CA, Cheng CH, Lee JW, Lo CT, Liu SY, Peng KC (2012) Monomeric l-amino acid oxidase-induced mitochondrial dysfunction in Rhizoctonia solani reveals a novel antagonistic mechanism of Trichoderma harzianum ETS 323. J Agric Food Chem 60:2464–2471
Yu EK, DeVoe IW (1981) l-Cysteine oxidase activity in the membrane of Neisseria meningitidis. J Bacteriol 145:280–287
Yu Z, Zhou N, Zhao C, Qiu J (2013) In-gel determination of l-amino acid oxidase activity based on the visualization of Prussian blue-forming reaction. PLoS One 8:e55548
Zeller A, Maritz A (1944) Ubereineneue l-aminosaure-oxidase. Helv Chim Acta 27:1888–1902
Acknowledgments
This work was financially supported by the Open Funding Project of the State Key Laboratory of Bioreactor Engineering, 863 program (2012AA022202), the Enterprise University research prospective program, Jiangsu Province (BY2013015-3), the Priority Academic Program Development of Jiangsu Higher Education Institutions, and 111 Project (111-2-06).
Author information
Authors and Affiliations
Corresponding author
Additional information
Dr. Xuedong Wang contributed equally to this paper.
Rights and permissions
About this article
Cite this article
Hossain, G.S., Li, J., Shin, Hd. et al. l-Amino acid oxidases from microbial sources: types, properties, functions, and applications. Appl Microbiol Biotechnol 98, 1507–1515 (2014). https://doi.org/10.1007/s00253-013-5444-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-5444-2