Abstract
The red macroalga Asparagopsis taxiformis has been shown to significantly decrease methane production by rumen microbial communities. This has been attributed to the bioaccumulation of halogenated methane analogues produced as algal secondary metabolites. The objective of this study was to evaluate the impact of A. taxiformis supplementation on the relative abundance of methanogens and microbial community structure during in vitro batch fermentation. Addition of A. taxiformis (2% organic matter) or the halogenated methane analogue bromoform (5 μM) reduced methane production by over 99% compared to a basal substrate-only control. Quantitative PCR confirmed that the decrease in methane production was correlated with a decrease in the relative abundance of methanogens. High-throughput 16S ribosomal RNA gene amplicon sequencing showed that both treatments reduced the abundance of the three main orders of methanogens present in ruminants (Methanobacteriales, Methanomassiliicoccales and Methanomicrobiales). Shifts in bacterial community structure due to the addition of A. taxiformis and 5 μM bromoform were similar and concomitant with increases in hydrogen concentration in the headspace of the fermenters. With high potency and broad-spectrum activity against rumen methanogens, A. taxiformis represents a promising natural strategy for reducing enteric methane emissions from ruminant livestock.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The rumen supports a diverse range of microorganisms including bacteria, archaea, protozoa, fungi and viruses [1, 2]. Essential energy and nutrients are provided to the host animal from the microbial degradation and fermentation of ingested plant biomass [3]. The archaeal component of the rumen microbiome is almost exclusively composed of methanogenic archaea [4]. These phylogenetically diverse microbes predominantly use hydrogen to generate methane via the reduction of different end products (carbon dioxide, methanol or methylated amines [5, 6]) produced during anaerobic fermentation. Methane is the predominant sink for hydrogen in the rumen and is a potent greenhouse gas, with a global warming potential 25 times greater than carbon dioxide over a 100-year period [7]. Enteric fermentation processes in domesticated ruminants are responsible for 6.4% of the total anthropogenic methane released into the atmosphere each year [8]. With the global demand for meat and milk predicted to rise, practical and targeted approaches for the mitigation of methane emissions are increasingly important.
Mechanisms for the biosynthesis of methane by methanogenic archaea have been described in detail [5]. Despite differences in the mechanisms used to generate methyl-coenzyme M [9], all known methanogens use the same enzymatic reaction to reduce this intermediate compound with coenzyme B to form methane [10,11,12]. Halogenated methane analogues (HMAs) such as bromochloromethane (BCM; CH2BrCl [13,14,15,16]), bromoform (BF; CHBr3 [17]), chloroform (CHCl3 [18]) and dichloromethane (CH2Cl2 [19]) have been shown to have antimethanogenic properties. This has been attributed to their ability to bind with reduced vitamin B12, blocking the cobamide-dependent methyltransferase reaction required for formation of methyl-coenzyme M [17]. Given their potential for broad-spectrum activity against the seven validly described orders of methanogens [9, 20], HMAs have been investigated for their ability to inhibit methane production in ruminants. Bromochloromethane can be formulated within a stabilised cyclodextrin matrix for delivery into the rumen, and methane emissions from cattle were reported to be reduced by 59% within 2 h of dosing [13]. The response suggests that BCM acts as a direct inhibitor of methanogenesis in vivo. Subsequent studies have shown that administration of BCM does not hinder animal performance [15] or rumen microbial fermentation [21], demonstrating the potential for reducing ruminant methane emissions using HMAs.
Despite the demonstrated efficacy at low concentrations, the use of artificially formulated HMAs in livestock production systems is prohibited because of their ozone-depleting effect [22]. Naturally derived sources of HMAs may provide a practical alternative method for delivery into the rumen. The red macroalga (or seaweed) Asparagopsis taxiformis produces high concentrations of BF as a secondary metabolite, which accumulates within vacuoles of gland cells [23]. The concentration of BF in this macroalga can vary from 0.17% [24] to 1.45% of the total biomass dry weight [25]. The impact of A. taxiformis supplementation on rumen microbiota in batch fermentation has been thoroughly evaluated [24, 26,27,28,29]. At an inclusion rate of 2% organic matter (OM), near-complete abatement of methane production is accompanied by a 12–25% reduction in total volatile fatty acid (VFA) yield with minimal impact on pH or substrate degradability. The aim of the present study was to build upon these previous observations by characterising the effect of A. taxiformis on rumen microbiota using DNA-based molecular techniques.
Materials and Methods
Experimental Design and Treatments
Rumen fluid was collected and pooled from four rumen-cannulated Brahman steers (Bos indicus) fed Rhodes grass ray (Chloris gayana) ad libitum. Experimental protocols were approved by the CSIRO Animal Ethics Committee (A5/2011) under the Australian Code of Practice for Care and Use of Animals for Scientific Purposes [30]. The rumen fluid sampling and batch fermentations were carried out as previously described [24, 28].
The treatments consisted of A. taxiformis biomass at a dose equivalent to 2% total OM (24.7 mg dry weight), two concentrations of BF (1 μM and 5 μM) and one concentration of BCM (5 μM), included as a positive control. Selection of the BF concentrations was based on a previous study demonstrating that inclusion of A. taxiformis at 2% OM delivers a final concentration of 1.3 μM BF [24]. Subsamples of the same algal biomass were used in both studies. The biomass was prepared by rinsing, freeze-drying and grinding through a 1-mm sieve. Bromoform and BCM treatments were prepared in dimethyl sulfoxide (DMSO) as previously described [24]. Control incubations contained the basal substrate (Rhodes grass hay) and DMSO equivalent to that added to the BF and BCM treatments (20 μL 125 mL−1). Six biological replicates for each treatment were prepared, with triplicates harvested after 48 and 72 h. Total gas production was recorded, and the concentrations of CH4 and H2 within the headspace were analysed by gas chromatography ([24], Table S1).
DNA Extraction and Illumina Sequencing
Samples from each replicate were filtered through a 130-μm mesh, and two 5-mL aliquots were immediately frozen on dry ice and stored at − 80 °C prior to subsequent analysis. Samples were thawed on wet ice and gently mixed by inversion. A 1-mL aliquot was transferred to a sterile 1.5-mL tube and centrifuged at 10000×g for 5 min at 4 °C to generate a cell-rich pellet. DNA extraction was performed using a PowerSoil® DNA Kit (Mo Bio Laboratories, Carlsbad, CA, USA), with the protocol modified to replicate the repeated bead-beating plus column (RBB+C) purification [31]. The DNA concentration in all samples was determined with a Qubit® 2.0 Fluorometer and dsDNA BR Assay Kit (Invitrogen, Carlsbad, CA, USA). All samples were diluted to 2 ng μL−1 in molecular-grade water prior to DNA amplification. DNA extraction was performed from duplicate frozen samples (technical replicates) for each batch fermentation (biological replicate), giving a total of six DNA extractions for each treatment/time point combination.
Preparation of 16S Ribosomal RNA Gene Amplicons for Next-Generation Sequencing (NGS)
Primers for amplification of 16S ribosomal RNA (rRNA) genes (515F and 806R) have been described by the Earth Microbiome Project (http://www.earthmicrobiome.org/emp-standard-protocols/16s/, [32]). A total of 5 μL DNA was added to a 96-well plate containing barcoded PCR primers (final concentration of 2 μM each; universal 515F primer and barcoded 806R primer) and 10 μL of MyTaq™ Hot Start DNA Polymerase mix (Bioline, London, UK). Polymerase chain reactions (PCRs) were performed using a MyCycler™ Thermal System (Bio-Rad, Hercules, CA, USA). The PCR reaction started at 98 °C for 3 min to denature DNA samples, followed by 25 cycles of 95 °C for 15 s, 55 °C for 15 s and 72 °C for 1 min and a final cycle of 72 °C for 3 min. Duplicate PCR products for each DNA extraction were pooled and visualised by electrophoresis on a 2% (wt/vol) agarose gel stained with SYBR Safe (Invitrogen, Carlsbad, CA, USA). Equimolar amounts of each PCR product were pooled in a single tube and further purified using a QIAEX II Gel Extraction Kit (Qiagen, Melbourne, Australia) followed by Agencourt AMPure® XP purification (Beckman Coulter, Brea, CA, USA). The purified DNA was sequenced using the Illumina MiSeq v2 2 × 250 bp chemistry (Ramaciotti Centre for Functional Genomics, UNSW Sydney, Australia).
Analysis of Microbial Community Profiles
Paired DNA sequence reads were demultiplexed and merged using a custom Python script before processing using default parameters in USEARCH 6.0 [33]. After removal of the forward and reverse primer sequences, reads were trimmed to 250 bp. The trimmed pairs were dereplicated and clustered at a similarity threshold of 97%. Chimeras were removed using the GOLD reference database before sequences were assigned to operational taxonomic units (OTUs). Results were normalised using a custom Python script to perform 100 independent rarefactions without replacement to a depth of 70,000 sequences per sample. OTUs with an average number of sequences lower than 35 (representing ≤ 0.05% of any given sample) were removed prior to computation of principal coordinate analysis (PCoA) using the weighted UniFrac parameters in QIIME 1.7.0 [34]. The initial UniFrac tree and PCoA plot were inspected to confirm that data from each pair of technical replicates were most similar to each other (OTU table is provided as Table S2). Counts from technical replicates were combined and the PCoA recomputed to represent data from biological replicates (n = 3 batch fermentations per treatment and time combination). Data from the rumen fluid inoculum was not included in the PCoA computation. A representative sequence from each OTU was classified to a taxonomic hierarchy using a minimal bootstrap of 50% in RDP Release 11.2 [35]. Taxonomic data presented in Fig. 2 represents the average of biological replicates.
Quantitative PCR to Enumerate Methanogen DNA
A single plasmid (p-STANDARD-1) containing a single copy of the 16S rRNA gene PCR product (from Escherichia coli XL1-Blue strain; for bacteria) and mcrA gene PCR product (from Methanobrevibacter ruminantium M1 strain; for methanogenic archaea) was used as a qPCR standard. PCR assays were performed in a Roche LightCycler® 480 II System (Roche). Each 10-μL reaction consisted of 5 μL SensiFAST™ SYBR® No-Rox reagent (Bioline, London, UK) and 1 μM each of forward and reverse primer (qmcrA-F and qmcrA-R for methanogens; or BacF and BacR for bacteria [13]). The cycle conditions of the qPCR assays were as follows: a single activation step at 95 °C for 5 s, and 45 amplification cycles at 95 °C for 5 s, 60 °C for 10 s and 72 °C for 20 s followed by an amplicon melting profile step. The qPCR assays showed consistent amplification efficiencies of greater than 1.9 and an error rate lower than 0.02 based on analysis of triplicate standards ranging from 3 × 102 to 3 × 107 copies of pSTANDARD-1. Target quantification was carried out using 2 μL of each DNA sample diluted to 100 pg μL−1. Each sample was amplified in triplicate providing three quantifications of each sample. Results are presented as a relative percentage of the methanogen mcrA gene to bacterial 16S rRNA gene for each treatment. Analysis of relative abundance was based on this ratio, and the copy number of targeted genes was not assumed [36].
Statistical Analysis
Total gas production was blank corrected and fitted to a modified sigmoidal Gompertz model [27] in JMP® 10 (SAS Institute, Cary, NC, USA). The effect of each treatment on gas production after 72 h of incubation was determined using one-way analysis of variance (ANOVA) and Tukey’s post hoc test (JMP® 10, Table S1). The effect of incubation time (48 and 72 h) and each treatment on the relative abundance of methanogens (qPCR) was determined through a two-factor permutational analysis of variance (PERMANOVA [37]) using Primer v6 software [38]. Pairwise comparisons were used to determine the significance of differences between the control and treatments.
Nucleotide Sequence Accession Numbers
A representative sequence for each OTU has been deposited in NCBI GenBank under accession numbers KT168398–KT174433. FASTA sequences and taxonomic lineages are included in Table S2.
Results
A. taxiformis and Halogenated Methane Analogues Decrease Methane Production and the Relative Abundance of Methanogens
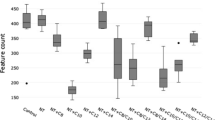
Total gas production decreased by 29–34% for all treatments (2% A. taxiformis, 1 μM BF, 5 μM BF and 5 μM BCM) compared with the control at 48 and 72 h (Fig. 1a, Table S1). The amount of methane formed was reduced by more than 77% with the addition of 1 μM BF and by more than 99% with the addition of 5 μM BCM, 5 μM BF or A. taxiformis at both time points (Fig. 1b). All treatments increased hydrogen accumulation after 48 h, at a rate of 1.8 mL g−1 OM for 1 μM BF and 5 μM BCM and 4.7 mL g−1 OM for 5 μM BF and A. taxiformis (Fig. 1c). After 72 h, hydrogen accumulation was still evident in the 5 μM BF (6 mL g−1 OM) and A. taxiformis (3.9 mL g−1 OM) treatments. The rate of hydrogen accumulation was comparable to the control for the 1 μM BF and 5 μM BCM treatments after 72 h (Fig. 1c).
Total production of a gas, b methane and c hydrogen and d the mean relative abundance of the methanogen mcrA gene to bacterial 16S rRNA gene for each treatment. Error bars represent ± SE (n = 3). BF: bromoform; Asp: A. taxiformis; BCM: bromochloromethane, RF: relative abundance of methanogens in the rumen fluid inoculum (no data of gas production and composition were collected for inoculum as it was being purged with N2 during the experimental setup to maintain the anaerobic conditions)
Quantitative PCR targeting the methanogen mcrA gene and bacterial 16S rRNA gene showed a significant decrease in the relative abundance of methanogens for all treatments compared with the control (PERMANOVA, Pseudo-F = 180.9, p = 0.0001). The proportion of methanogens (Fig. 1d) was lower after 48 h than at 72 h for comparative treatments. Incubations containing A. taxiformis, 5 μM BF or 5 μM BCM showed similar results, with a reduction of 72% after 48 h and 75% after 72 h. The relative abundance of methanogens in incubations containing 1 μM BF was comparable to the other three treatments after 48 h but showed a reduction of only 60% after 72 h. The decreases in methanogen relative abundance were positively correlated with decreases in methane production (Spearman’s correlation coefficient = 0.83, p = 0.0001).
Impact of A. taxiformis and Halogenated Methane Analogues on Microbial Community Composition
The prokaryotic composition of each sample was determined using high-throughput amplicon sequencing of the 16S rRNA gene, rarefied to n = 70,000 per sample. Between 97 and 99% of each sample was composed of reads from the domain Bacteria. All samples contained eight bacterial phyla (> 1% of total reads per phylum), with OTUs belonging to the Firmicutes and Bacteriodetes comprising the majority of sequence reads in all treatments (Fig. 2a). All batch fermentations consistently showed a decrease in Bacteroidetes/Firmicutes ratio compared to the pooled rumen fluid used as the inoculum. This indicates that the experimental conditions shifted the bacterial community diversity compared to the initial composition. Differences between batch fermentations based on treatment or time were minimal at the phylum level.
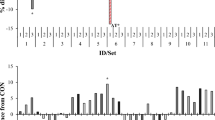
Relative abundance of microbial taxa as determined by 16S rRNA amplicon sequencing. a Relative abundance of microbial taxa at the phylum level. b Relative abundance within the Euryarchaeota phylum displayed at the order level. BF1: 1 μM bromoform; BF5: 5 μM bromoform Asp: 2% A. taxiformis; BCM: 5 μM bromochloromethane
A PCoA plot based on a weighted UniFrac metric was generated to visualise differences in microbial community structure at the OTU level (Fig. 3). Weighted UniFrac is a β diversity measure that accounts for the presence/absence of OTUs and their relative abundance [39]. For comparison, the unweighted PCoA plot (based on the presence/absence of OTUs) is provided as Fig. S1. Note that both measures use phylogenetic information based on alignment of representative sequences to account for the degree of divergence between each OTU [39]. The weighted UniFrac PCoA explains 60.23 and 26.07% of the variation in the batch fermentation data on PC1 and PC2 respectively (Fig. 3). When comparing within experimental conditions, like treatments shifted towards the right on PC1 and towards the bottom on PC2 as the incubation time increased from 48 to 72 h. This indicates that microbial community diversity was affected by the duration of batch fermentation. When comparing within incubation times, all treatments were separated from the controls on both PC1 and PC2. The A. taxiformis and 5 μM BF treatments were grouped at 48 or 72 h, demonstrating a similar impact on rumen microbial communities during batch fermentation. The 5 μM BCM and 1 μM BF treatments also clustered together at each time point. The impact of these treatments was less pronounced than that induced by A. taxiformis and 5 μM BF, with profiles occupying an intermediate position in the plot space.
A. taxiformis and Halogenated Methane Analogues Are Inhibitory to the Predominant Orders of Ruminal Methanogens
All archaeal sequence reads were taxonomically classified to methanogen lineages. The proportion of reads assigned to methanogen OTUs (Fig. 2b) corresponds to the relative abundance of methanogens as determined using qPCR (Fig. 1c). Methanogen sequences comprised 3.2% of the total reads (Fig. 2a) from the control after 72 h of incubation. This decreased to 0.9% in the A. taxiformis treatment, 0.8% in the 5 μM BF and BCM treatments and 1.2% in the 1 μM BF treatment. The 72-h control samples were dominated by the orders Methanobacteriales (59.5%) and Methanomassiliicoccales (37%). Within the Methanobacteriales, the Methanobrevibacter genus was highly represented (57.8% of total methanogen reads at 72 h), with the Methanosphaera genus comprising a small proportion of the control dataset (2% of total methanogen reads at 72 h). The remaining methanogen reads in the 72-h control samples belonged to the orders Methanomicrobiales (3.4%) and Methanosarcinales (< 1%). All of the treatments applied in this study inhibited the growth of methanogens from the Methanobacteriales, Methanomassiliicoccales and Methanomicrobiales, as shown by an overall decrease in the relative abundance of sequence reads from these orders compared with the control at each time point (Fig. 2b).
Discussion
We have demonstrated that the in vitro antimethanogenic properties of A. taxiformis generate similar results to addition of 5 μM bromoform, the main halogenated methane analogue produced as a secondary metabolite. The BF concentration in the A. taxiformis biomass used for this study was measured at 1.3 μM [24], and we hypothesised that the effect would be similar to 1 μM BF, with 5 μM BF included as a high concentration positive control. At equivalent time points, no significant differences in methane production, methanogen relative abundance or microbial community composition were identified between treatments containing either A. taxiformis or 5 μM BF. This suggests that the A. taxiformis biomass is more potent than anticipated as the effects were more pronounced than what was observed in the 1-μM BF treatments.
A. taxiformis produces a wide range of halogenated metabolites including bromine- and iodine-containing haloforms and other halogenated methanes and ethanes in lesser quantities [40,41,42,43]. The potency of inhibition of methanogenesis varies among halogenated compounds [44], possibly as a reflection of differences in sensitivity between different species of methanogens [45]. In this context, BF could act synergistically with other algal halogenated metabolites to enhance the antimethanogenic activity of the bioactive rich biomass. Given that dietary supplementation with algal biomass would also circumvent the requirement for HMAs to be stabilised for delivery into the rumen [15], this may represent a more holistic approach for the mitigation of enteric methane emissions compared to the use of individual extracts or metabolites.
The in vitro batch fermentation system used in this study supports cultivation of the diversity of methanogens resident in the rumen microbiome [4, 46, 47]. Methanogenic taxa belonging to the orders Methanobacteriales, Methanomassiliicoccales, Methanomicrobiales and Methanosarcinales were present in the control fermentations, although Methanosarcinales were not highly abundant (< 1% of the total reads assigned to all archaeal OTUs). Halogenated methane analogues are predicted to be inhibitory to all known methanogens by preventing synthesis of methyl-coenzyme M [17]. Addition of A. taxiformis, 5 μM BF or 5 μM BCM to the incubations reduced methane production by more than 99% compared with the control. This was concomitant with a decrease in the relative abundance of methanogens (as measured by qPCR). Each treatment was inhibitory towards the three main orders of methanogens found in ruminants, with a statistically significant decrease in the proportion of sequencing reads assigned to OTUs from the orders Methanobacteriales, Methanomassiliicoccales and Methanomicrobiales. This provides further support for the broad-spectrum antimethanogenic properties of HMAs.
The effects of HMAs have been studied in cattle and goats, with positive indications to support their efficacy for mitigating methane emissions in vivo. Bromoform has been shown to inhibit ruminal methanogenesis at a concentration of 25 μM [44]. Bromochloromethane resulted in a reduction in total methane emissions over 12 h of approximately 30% and an increase in propionate/acetate ratio following a 28-day treatment period (0.3 g/100 kg liveweight delivered twice daily). Increases in branched short-chain fatty acids [13] and metagenomic assessments [48] have provided evidence that pathways for utilisation of excess hydrogen are activated in the rumen when methanogens are directly inhibited by HMAs. In a related study, no adverse impacts on liveweight gain or feed conversion ratio for cattle were associated with the use of BCM [15]. In sheep, supplementation of a high-fibre diet with A. taxiformis resulted in significant and persistent decreases in methane output of up to 80% over a 72-day period [49]. This preliminary study noted no significant difference in liveweight gain despite a decrease in total VFAs and acetate with an increase in propionate. The effects of A. taxiformis on productivity, feed intake and digestibility are yet to be addressed in cattle. The microbiota must be able to maintain functional capacity for fermentation of the basal diets. The in vitro degradability of organic matter is not significantly affected by the addition of A. taxiformis at 2% OM [24, 26, 28, 29], despite shifts in the underlying microbial community composition. In conclusion, A. taxiformis shows promise as a potent, specific and natural alternative for the mitigation of methane emissions from ruminant livestock. Further investigations are required to quantify the extent of productivity gains and methane abatement resulting from A. taxiformis supplementation.
References
Kamra D (2005) Rumen microbial ecosystem. Curr Sci 89:124–135
Sirohi SK, Singh N, Dagar SS, Puniya AK (2012) Molecular tools for deciphering the microbial community structure and diversity in rumen ecosystem. Appl Microbiol Biotechnol 95:1135–1154
Wright A-DG, Klieve AV (2011) Does the complexity of the rumen microbial ecology preclude methane mitigation? Anim Feed Sci Technol 166:248–253
Janssen PH, Kirs M (2008) Structure of the archaeal community of the rumen. Appl Environ Microbiol 74:3619–3625
Costa KC, Leigh JA (2014) Metabolic versatility in methanogens. Curr Opin Biotechnol 29:70–75
Liu Y, Whitman WB (2008) Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Ann N Y Acad Sci 1125:171–189
IPCC (2007) Climate change 2007: The physical science basis. In: Solomon S, Qin D, Manning M, Chen Z, Marquis M, Averyt KB, Tignor M, Miller HL (eds) Contribution of working group I to the fourth assessment report of the Intergovernmental Panel on Climate Change. Cambridge University Press, Cambridge, p 1009
Ripple WJ, Smith P, Haberl H, Montzka SA, McAlpine C, Boucher DH (2014) Ruminants, climate change and climate policy. Nat Clim Chang 4:2–5
Borrel G, O’Toole PW, Harris HM, Peyret P, Brugère J-F, Gribaldo S (2013) Phylogenomic data support a seventh order of methylotrophic methanogens and provide insights into the evolution of methanogenesis. Genome Biol Evol 5:1769–1780
Hedderich R, Whitman WB (2006) Physiology and biochemistry of the methane-producing Archaea. The prokaryotes 2:1050–1079
Ellermann J, Hedderich R, Böcher R, Thauer RK (1988) The final step in methane formation. Eur J Biochem 172:669–677
Ermler U, Grabarse W, Shima S, Goubeaud M, Thauer RK (1997) Crystal structure of methyl-coenzyme M reductase: the key enzyme of biological methane formation. Science 278:1457–1462
Denman SE, Tomkins NW, McSweeney CS (2007) Quantitation and diversity analysis of ruminal methanogenic populations in response to the antimethanogenic compound bromochloromethane. FEMS Microbiol Ecol 62:313–322
Goel G, Makkar HP, Becker K (2009) Inhibition of methanogens by bromochloromethane: effects on microbial communities and rumen fermentation using batch and continuous fermentations. Br J Nutr 101:1484–1492
Tomkins N, Colegate S, Hunter R (2009) A bromochloromethane formulation reduces enteric methanogenesis in cattle fed grain-based diets. Anim Prod Sci 49:1053–1058
Abecia L, Toral PG, Martín-García AI, Martínez G, Tomkins NW, Molina-Alcaide E, Newbold CJ, Yáñez-Ruiz DR (2012) Effect of bromochloromethane on methane emission, rumen fermentation pattern, milk yield, and fatty acid profile in lactating dairy goats. J Dairy Sci 95:2027–2036. https://doi.org/10.3168/jds.2011-4831
Wood J, Kennedy FS, Wolfe R (1968) Reaction of multihalogenated hydrocarbons with free and bound reduced vitamin B12. Biochemistry 7:1707–1713
Martinez-Fernandez G, Denman SE, Yang C, Cheung J, Mitsumori M, McSweeney CS (2016) Methane inhibition alters the microbial community, hydrogen flow, and fermentation response in the rumen of cattle. Front Microbiol 7:1122
Bauchop T (1967) Inhibition of rumen methanogenesis by methane analogues. J Bacteriol 94:171–175
Thauer RK, Kaster A-K, Seedorf H, Buckel W, Hedderich R (2008) Methanogenic archaea: ecologically relevant differences in energy conservation. Nat Rev Micro 6:579–591
Mitsumori M, Shinkai T, Takenaka A, Enishi O, Higuchi K, Kobayashi Y, Nonaka I, Asanuma N, Denman SE, McSweeney CS (2012) Responses in digestion, rumen fermentation and microbial populations to inhibition of methane formation by a halogenated methane analogue. Br J Nutr 108:482–491
Hristov A, Oh J, Meinen R, Montes F, Ott T, Firkins J, Rotz A, Dell C, Adesogan A, Yang W, Tricarico J, Kebreab E, Waghorn G, Dijkstra J, Oosting S (2013) Mitigation of greenhouse gas emissions in livestock production—a review of technical options for non-CO2 emissions. In: Gerber P, Henderson B, Makkar H (eds) FAO Animal Production and Health Paper No 177, Rome, Italy, p 231
Paul NA, Cole L, De Nys R, Steinberg PD (2006) Ultrastructure of the gland cells of the red alga Asparagopsis Armata (bonnemaisoniaceae). J Phycol 42:637–645
Machado L, Magnusson M, Paul NA, Kinley R, de Nys R, Tomkins N (2016) Identification of bioactives from the red seaweed Asparagopsis taxiformis that promote antimethanogenic activity in vitro. J Appl Phycol 28:3117–3126. https://doi.org/10.1007/s10811-016-0830-7
Paul N, de Nys R, Steinberg P (2006) Chemical defence against bacteria in the red alga Asparagopsis armata: linking structure with function. Mar Ecol Prog Ser 306:87–101
Kinley RD, de Nys R, Vucko MJ, Machado L, Tomkins NW (2016) The red macroalgae Asparagopsis taxiformis is a potent natural antimethanogenic that reduces methane production during in vitro fermentation with rumen fluid. Anim Prod Sci 56:282–289. https://doi.org/10.1071/AN15576
Machado L, Magnusson M, Paul NA, de Nys R, Tomkins N (2014) Effects of marine and freshwater macroalgae on in vitro total gas and methane production. PLoS One 9:e85289. https://doi.org/10.1371/journal.pone.0085289
Machado L, Magnusson M, Paul NA, Kinley R, de Nys R, Tomkins N (2016) Dose-response effects of Asparagopsis taxiformis and Oedogonium sp. on in vitro fermentation and methane production. J Appl Phycol 28:1443–1452. https://doi.org/10.1007/s10811-015-0639-9
Kinley RD, Vucko MJ, Machado L, Tomkins NW (2016) In vitro evaluation of the antimethanogenic potency and effects on fermentation of individual and combinations of marine macroalgae. Am J Plant Sci 7:2038
NHMRC (2013) Australian code for the care and use of animals for scientific purposes. National Health and Medical Research Council, Canberra
Yu Z, Morrison M (2004) Improved extraction of PCR-quality community DNA from digesta and fecal samples. BioTechniques 36:808–813
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, Brown CT, Porras-Alfaro A, Kuske CR, Tiedje JM (2013) Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42(D1):D633–D642
Rosewarne CP, Pettigrove V, Stokes HW, Parsons YM (2010) Class 1 integrons in benthic bacterial communities: abundance, association with Tn402-like transposition modules and evidence for coselection with heavy-metal resistance. FEMS Microbiol Ecol 72:35–46
Anderson M, Gorley R, Clarke K (2008) PERMANOVA + for PRIMER: guide to software and statistical methods. PRIMER-E
Clarke KR, Gorley RN (2006) PRIMER v6: user manual/tutorial. PRIMER-E Ltd, Plymouth
Lozupone CA, Hamady M, Kelley ST, Knight R (2007) Quantitative and qualitative β diversity measures lead to different insights into factors that structure microbial communities. Appl Environ Microbiol 73:1576–1585
Kladi M, Vagias C, Roussis V (2004) Volatile halogenated metabolites from marine red algae. Phytochem Rev 3:337–366
Burreson B, Moore RE, Roller P (1975) Haloforms in the essential oil of the alga Asparagopsis taxiformis (Rhodophyta). Tetrahedron Lett 16:473–476
Burreson BJ, Moore RE, Roller PP (1976) Volatile halogen compounds in the alga Asparagopsis taxiformis (Rhodophyta). J Agric Food Chem 24:856–861
Woolard FX, Moore RE, Roller PP (1979) Halogenated acetic and acrylic acids from the red alga Asparagopsis taxiformis. Phytochemistry 18:617–620
Lanigan G (1972) Metabolism of pyrrolizidine alkaloids in the ovine rumen. IV. Effects of chloral hydrate and halogenated methanes on rumen methanogenesis and alkaloid metabolism in fistulated sheep. Crop Pasture Sci 23:1085–1091
Ungerfeld E, Rust S, Boone D, Liu Y (2004) Effects of several inhibitors on pure cultures of ruminal methanogens. J Appl Microbiol 97:520–526
Seedorf H, Kittelmann S, Henderson G, Janssen PH (2014) RIM-DB: a taxonomic framework for community structure analysis of methanogenic archaea from the rumen and other intestinal environments. Peer J 2:e494
Kittelmann S, Seedorf H, Walters WA, Clemente JC, Knight R, Gordon JI, Janssen PH (2013) Simultaneous amplicon sequencing to explore co-occurrence patterns of bacterial, archaeal and eukaryotic microorganisms in rumen microbial communities. PLoS One 8:e47879
Denman SE, Fernandez GM, Shinkai T, Mitsumori M, McSweeney CS (2015) Metagenomic analysis of the rumen microbial community following inhibition of methane formation by a halogenated methane analog. Front Microbiol 6:1087. https://doi.org/10.3389/fmicb.2015.01087
Li X, Norman HC, Kinley RD, Laurence M, Wilmot M, Bender H, de Nys R, Tomkins N (2016) Asparagopsis taxiformis decreases enteric methane production from sheep. Anim Prod Sci AN15883. doi: https://doi.org/10.1071/AN15883
Acknowledgements
We thank Dr. Mike Devery for kindly supplying the bromochloromethane used in this study and Dr. Matthew Vucko for assistance with the experiments. We also thank Jeffrey Palpratt from the College of Public Health, Medical & Veterinary Sciences for handling and maintenance of the donor steers, and Dr. Shane Askew from the Advanced Analytical Centre, JCU, for analytical advice.
Funding
This research was supported by the Australian Government Department of Agriculture and Meat & Livestock Australia, funded as part of the National Livestock Methane Program. It was also supported by MBD Energy and the Australian Government through the Advanced Manufacturing Cooperative Research Centre, funded through the Australian Government’s Cooperative Research Centre Scheme.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Experimental protocols were approved by the CSIRO Animal Ethics Committee (A5/2011) under the Australian Code of Practice for Care and Use of Animals for Scientific Purposes
Electronic supplementary material
Table S1
Mean gas production parameters after 72 h incubation (n = 3) (DOCX 13 kb)
Table S2
OTU table with FASTA sequences and taxonomic lineages for experimental and technical replicates. Samples are labelled as treatment_time.Nx, where N is 1, 2 or 3 to denote the experimental replicate; and x is a or b to denote the technical replicate. OTU labelling corresponds to sequences deposited in NCBI Genbank under accession numbers KT168398 – KT174433. (XLSX 1135 kb)
Fig. S1
Principal co-ordinate analysis plot showing relationships between microbial communities under different experimental conditions, based on unweighted UniFrac metric of β diversity. BF1: 1 μM bromoform; BF5: 5 μM bromoform Asp: 2% A. taxiformis; BCM: 5 μM bromochloromethane. (GIF 85 kb)
Rights and permissions
About this article
Cite this article
Machado, L., Tomkins, N., Magnusson, M. et al. In Vitro Response of Rumen Microbiota to the Antimethanogenic Red Macroalga Asparagopsis taxiformis . Microb Ecol 75, 811–818 (2018). https://doi.org/10.1007/s00248-017-1086-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-1086-8