Abstract
Strains of Bacillus thuringiensis (Bt) are commonly commercialized as bioinoculants for insect pest control, but their benefits go beyond their insecticidal property: they can act as plant growth-promoters. Auxins play a major role in the plant growth promotion. However, the mechanism of auxin production by the Bacilli group, and more specifically by Bt strains, is unclear. In previous work, the plant growth-promoting rhizobacterium (PGPR) B. thuringiensis strain RZ2MS9 increased the corn roots. This drew our attention to the strain’s auxin production trait, earlier detected in vitro. Here, we demonstrate that in its genome, RZ2MS9 harbours the complete set of genes required in two pathways that are used for Indole acetic acid (IAA) production. We also detected that the strain produces almost five times more IAA during the stationary phase. The bacterial application increased the shoot dry weight of the Micro-Tom (MT) tomato by 24%. The application also modified MT root architecture, with an increase of 26% in the average lateral root length and inhibition of the axial root. At the cellular level, RZ2MS9-treated MT plants presented elongated root cortical cells with intensified mitotic activity. Altogether, these are the best characterized auxin-associated phenotypes. Besides that, no growth alteration was detected in the auxin-insensitive diageotropic (dgt) plants either with or without the RZ2MS9 inoculation. Our results suggest that auxins play an important role in the ability of B. thuringiensis RZ2MS9 to promote MT growth and provide a better understanding of the auxin production mechanism by a Bt strain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Auxins, such as the Indole-3-acetic acid (IAA), are major regulators of plant growth, development, and stress response (Liu et al. 2014). At the cellular level, these phytohormones can control many aspects of cell division, differentiation, and elongation by stimulating cell wall synthesis and/or increasing membrane plasticity (Bergfeld et al. 1988; Overvoorde et al. 2010). The biosynthesis of auxins by plant-associated bacteria is well-described and probably one of the most important causes of improved growth and yield in various crops when they are applied with plant growth-promoting rhizobacteria (PGPR) (Steenhoudt and Vanderleyden 2000; Bloemberg and Lugtenberg 2001).
A large proportion (around 80%) of bacteria from the rhizosphere has been reported as producing IAA, and the amino acid l-tryptophan (Trp) has been identified as the main precursor for its biosynthesis (Loper and Schroth 1986; Patten and Glick 1996). Four major Trp-dependent IAA synthesis pathways are known in bacteria, which are classified according to their intermediates: indole-3-acetonitrile (IAN), indole-3-acetamide (IAM), indole-3-pyruvate (IPA), and tryptamine (TPM) (Zhang et al. 2019) (Fig. 1). Briefly, in the IAN pathway, Trp is converted into indole-3-acetaldoxime (IAOx) by an oxidoreductase. Subsequently, IAOx is converted into IAN by an IAOx dehydratase. Once IAN is formed, it can be converted to IAA by the action of nitrilases. In the IAM pathway, the enzyme tryptophan-2-monooxygenase converts Trp into IAM, which is then converted to IAA by an IAM hydrolase. In the IPA pathway, Trp is firstly transformed into IPA by an aromatic amino acid transferase, and then IPA can be converted into indole-3-acetaldehyde (IAAld) by the well-documented enzyme indolepyruvate decarboxylase (IPDC). IAAld is then transformed into IAA by dehydrogenases. Finally, in the TPM pathway, Trp is first converted into TPM by a Trp decarboxylase and TPM is subsequently converted to IAAld by an amine oxidase. As in the last step of the IPA pathway, the IAAld can be transformed to IAA by dehydrogenases (Spaepen and Vanderleyden 2011; Zhang et al. 2019). Although several experiments have been performed to understand the above named mechanisms responsible for the bacterial IAA synthesis, the biosynthetic pathways present in Gram-positive bacteria, in particular in the Bacilli group, remain vague (Idris et al. 2007; Ali et al. 2009; Shao et al. 2015a).

adapted from Zhang et al. (2019). Arrows with solid lines indicate the enzymes which genes were detected in the B. thuringiensis RZ2MS9 genome. Arrows with dashed lines indicate the enzymes which genes were not detected in RZ2MS9 genome. ORD, oxidoreductase; IAOxD, indole-3-acetaldoxime dehydratase; TMO, tryptophan-2-monooxygenase; IAH, indole-3-acetamide hydrolase; AAT, aromatic amino transferase; IPDC, indole-3-pyruvate decarboxylase; ALDH, aldehyde dehydrogenase; TDC, tryptophan decarboxylase; MAO, monoamine oxidase
Main L-tryptophan-dependent metabolic pathways involved in the bacterial production of indole 3-acetic acid (IAA)—
Bacillus thuringiensis RZ2MS9 is a PGPR, previously isolated from guarana (Paullinia cupana) plants cultivated in the Brazilian Amazon, which successfully promotes the growth of corn and soybean plants. In prior studies, we detected that the bacterium produces auxins and siderophores in vitro and it can solubilize phosphate and grow in a N-free medium (Batista et al. 2018). However, the highly beneficial effect of this strain on corn’s roots growth drew attention to its previously detected auxin production trait. RZ2MS9 application increased corn’s root dry weight by almost 250% in greenhouse conditions (against the 136% increase observed with the application of the strain RZ2MS16, an auxin-producer Burkholderia ambifaria from the same bacterial collection) (Batista et al. 2018).
To assess whether the plant growth-promotion caused by the inoculation of B. thuringiensis RZ2MS9 may be explained by its auxin production, we utilized a plant host that has a mutant insensitive to the effect of exogenous auxins as study model: the dwarf tomato Micro-Tom (MT) (Solanum lycopersicum cv. Micro-Tom) and its mutant diageotropic (dgt). Tomato is one of the best-studied organisms in genetic research and has been increasingly proposed as a model of studies due to its agronomic relevance (Kimura and Sinha 2008; Campos et al. 2010). Originally developed as an ornamental cultivar, the MT presents reduced size (less than 15 cm in height) and a rapid life cycle (about 10 weeks). MT was used as a genetic background in which mutations were introduced, creating a collection of mutants to study a wide range of processes stimulated by bio-stimulants (Carvalho et al. 2011; Zandonadi et al. 2016). The dgt is a spontaneous single gene recessive mutant that is insensitive to exogenously applied auxin (Kelly and Bradford 1986; Daniel et al. 1989; Hicks et al. 1989). This is due to a defect in the Dgt gene which encodes a cyclophilin, a component of auxin signaling and plant development (Oh et al. 2006).
The main aims of this study were to (1) investigate the molecular basis of auxin production by the PGPR B. thuringiensis strain RZ2MS9, (2) optimize the cultivation conditions of RZ2MS9 to enhance its auxin production, and to (3) test the effect of B. thuringiensis RZ2MS9 on the growth of MT and dgt tomato plants.
Materials and methods
Biological materials
Bacillus thuringiensis RZ2MS9, a plant growth-promoting rhizobacterium (PGPR) previously isolated from the guarana rhizosphere (Batista et al. 2018) is deposited in the Brazilian Collection of Microorganisms from the Environment and Industry (Coleção Brasileira de Micro-organismos de Ambiente e Indústria—CBMAI), registered number 823, strain access number CBMAI1824. The strain RZ2MS9 was selected for the present study due to its superior performance in promoting corn growth in greenhouse conditions when compared to another IAA-producer PGPR tested as well as its harmless to human healthy.
In our laboratory, RZ2MS9 is kept stored in a Luria–Bertani (LB) medium (Sambrook and Russell 2001) containing 15% glycerol at − 80 °C. Fresh cultures are routinely started from frozen stocks by culturing the strain on the agar LB and incubating the plates at 28 °C for 24 h.
Tomato seeds (Solanum lycopersicum) of the cultivar Micro-Tom (MT) and its diageotropic mutant (dgt) were obtained from the HCPD laboratory collection of Micro-Tom mutants (http://www.esalq.usp.br/tomato/) maintained at the Luiz de Queiroz College of Agriculture (ESALQ), University of São Paulo (USP), Piracicaba—SP, Brazil.
RZ2MS9 taxonomic analysis
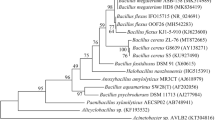
The 16S rDNA gene partial sequencing of the strain RZ2MS9 done previously (Batista et al. 2018) revealed that the strain belongs to the Bacillus cereus sensu lato group, which comprises notable members such as the B. cereus (stricto sensu) and the B. thuringiensis. Thus, we retrieved all complete genomes from B. cereus sensu lato group (n = 231) available in the National Center for Biotechnology Information (NCBI) Assembly Database in February 2021 (Supplementary Table 1). The complete genome of B. thuringiensis RZ2MS9 (NCBI Reference Sequence: NZ_CP049978.1) (Bonatelli et al. 2020) was also included in the following analyses.
FastANI version 1.32 (Jain et al. 2018) generated the average nucleotide identity (ANI) between RZ2MS9 and all the B. cereus sensu lato group genomes. Pangenome analysis was performed with Roary version 3.13 (Page et al. 2015). Only the genomes from B. cereus and B. thuringiensis species were considered in this analysis. All genomes were annotated with Prokka version 1.14.5 (Seemann 2014), considering the flag-genus Bacillus. Default parameters were used in both FastANI and Roary. With the core genome generated (n = 1823), FastTree version 2.1.10 (Price et al. 2010) was used to generate the approximately-maximum-likelihood phylogenetic tree using the generalized time-reversible (GTR) model of nucleotide evolution. iTOL (Letunic and Bork 2019) was used for tree visualization.
RZ2MS9 genome mining for putative IAA-related genes
The complete genome sequence of B. thuringiensis RZ2MS9 (Bonatelli et al. 2020) was also used for mining IAA-related genes. The functional annotation of the genome was performed against the KEGG database (http://www.genome.jp/kegg/), using the BlastKOALA tool. The annotated genes were manually inspected and identified based on information of genes/enzymes already reported to function in IAA metabolism. The detected IAA-related genes were confirmed using tBLASTn.
Kinetics of IAA biosynthesis by Bacillus sp. RZ2MS9
To evaluate the IAA production by RZ2MS9 over the time, the strain was grown in 100 ml Erlenmeyer flasks containing 30 ml of LB medium supplemented with Trp (1 g/l). Each flask, 3 replicates, was inoculated with the same initial bacterial load (optical density at 600nm [OD600] = 0.05, approximately 1 × 108 CFU/ml). The flasks were incubated in the dark at 28 °C under shaking conditions (180 rpm). We measured the IAA production and bacterial growth (OD600) 3, 12, 15, 18, 21, and 30 h after inoculation (HAI).
To evaluate the influence of Trp concentration on IAA production by RZ2MS9, the strain was grown in 100 ml Erlenmeyer flasks containing 30 ml of LB medium supplemented with different amounts of Trp (0, 0.1, 0.5, 1.0 and 2.0 g/l). Each flask, 3 replicates per treatment, was inoculated with the same initial bacterial load (OD600 = 0.05, approximately 1 × 108 CFU/ml). The flasks were incubated in the dark for 21 h at 28 °C under shaking conditions (180 rpm).
For IAA measurement, we centrifuged (10,000 × g) the bacterial cultures for 5 min and mixed 1 ml of the supernatant with 2 ml of the Salkowski’s reagent (1 ml of 0.5 M FeCl3 in 50 ml of 35% HClO4 solution). The development of a pink color indicated auxin production, which was detected by OD measurement, at 530 nm (OD530), using a spectrophotometer (Ultrospec 3000 Amersham Pharmacia Biotech) (Gordon and Weber 1951). IAA production per ml was then calculated by interpolation of the obtained OD530 with an IAA calibration curve.
Plant growth-promotion assay
To treat the tomato seeds with the B. thuringiensis RZ2MS9, the strain was grown in 250 ml Erlenmeyer flasks containing 100 ml of LB medium. The flasks were initially inoculated with 1 ml of the strain cell suspension and incubated under shaking conditions (180 rpm) at 28 °C for approximately 21 h. The cell density was adjusted to 1 × 108 CFU/ml. Micro-Tom (MT) and the diageotropic mutant (dgt) seeds were immersed in the bacterial suspension and incubated at 28 °C under shaking conditions (180 rpm) for 30 min. Controls consisted of MT and dgt seeds immersed in the LB culture medium without bacterial growth and incubated under the same conditions as described above. After, the seeds were carefully sown into pre-filled tray cells containing a 1:1 mixture of a commercial mix (Bioplant®) with expanded vermiculite and kept in a greenhouse at 28 °C. 10 days after germination, plants were transferred to 400 ml pots containing the soil mix, which were kept in the greenhouse under the same conditions as described previously. Both when in the trays and when in the pots, the plants were irrigated daily and the Peters® general-purpose water-soluble fertilizer (20–20-20 NPK + micronutrients) was applied once a week.
The 36-pot experiment, in a randomized experimental design, consisted of a factorial scheme (2 plant genotypes × 2 bacterial treatments). The treatments were: Micro-Tom (MT) plants which seeds had the application of RZ2MS9 cells suspension (MT_RZ2MS9) and the control without bacterial application (MT_Control); and dgt plants which seeds were treated with RZ2MS9 (dgt_RZ2MS9) and the non-inoculated control (dgt_Control). Nine replicates were used per treatment in the bioassay.
Plants were collected at the flowering stage, sixty days after transferring the plants to individual pots. The roots were separated from the shoots, washed and kept in pots containing alcohol (70%). The shoots were relocated in paper bags and the shoot dry weight (SDW) was measured in an analytical balance after oven-drying at 70 °C for 5 days.
The roots were scattered in a clear layer of water on a tray (30 cm × 20 cm), and their image was captured at 400 dpi with an Epson® Expression 11000XL professional scanner system. The images were analyzed for total root length (TRL), lateral root length (LRL), axial root length (ARL), total root surface area (RSA) and total root diameter (TRD) using WinRHIZO™ Arabidopsis software (Regent Instruments Inc., Quebec, Canada).
The ten root diameter classes provided by the WinRHIZO™ Arabidopsis software were simplified into only two (LRL and ARL), according to the methodology proposed by Trachsel et al. (2009). Thus, we considered the root portions with a diameter of less than or equal to 0.5 mm as LRL, and the root portions with a diameter greater than 0.5 mm as ARL.
Root anatomical analysis
Root tips of the MT plants, with and without B. thuringiensis RZ2MS9 treatment, were fixed in the Karnovsky solution for 24 h (Karnovsky 1965), dehydrated through a graded alcohol series (10, 20, 30, 50, 70, 80, 90 and 100%) and embedded in plastic resin (Tecnovit® 7100). The samples were sectioned in rotational microtome Leica RM2235 at 5–7 μm thickness. The sections were displayed on glass slides and subsequently stained with toluidine blue (O’Brien et al. 1964) and mounted with the synthetic resin Entellan® (Merck, Darmstadt, Germany). The slides were analyzed under a Leica DM 500, and the images documented with an ICC50W camera.
Statistical analysis
Data were submitted to analysis of variance and Tukey’s test (p-value < 0.05) with bacterial inoculation and plant genotype as sources of variation using the package “ExpDes.pt” in software R (R Core Team 2014). Pearson's correlation coefficients between IAA production and cell density, as well as between IAA production and the levels of Trp added to the culture medium were also calculated using the software R.
Results
RZ2MS9 taxonomic analysis
FastANI analysis showed that the closest species to the strain RZ2MS9 was the B. thuringiensis subsp. jinghongiensis strain YGd22-03, with 99.05% ANI (Supplementary Table 1). Interestingly, the phylogenetic tree based on the B. cereus and B. thuringiensis species core genomes (n = 1823) clustered the RZ2MS9 and the B. thuringiensis strain subsp. jinghongiensis YGd22-03 together in a clade mainly composed by B. thuringiensis representatives (Supplementary Fig. 1). Therefore, the strain RZ2MS9 will be referred as B. thuringiensis RZ2MS9 hereafter.
RZ2MS9 genome mining for genes likely involved in IAA biosynthesis
A graphic overview of the main pathways proposed for the synthesis of bacterial IAA is presented in Fig. 1. IAA synthesis from Trp can occur via four main alternative pathways, named according to its main intermediate: indole-3-acetonitrile (IAN), indole-3-acetamide (IAM), indole-3-pyruvate (IPA), and tryptamine (TPM). The complete B. thuringiensis RZ2MS9 genome was mined for genes putatively involved in each step of these pathways. No genes related to the IAN and IAM pathways were detected. However, the whole set of genes involved in the IPA and in the TPM pathways were identified in the RZ2MS9 genome (Table 1). In the IPA pathway, we detected two genes that can putatively encode aminotransferases involved in the conversion of tryptophan into IPA. We also identified the well-documented ipdC gene presumably responsible for the production of the indole-3-pyruvate decarboxylase (IPDC) enzyme, which converts IPA into IAAld. In the TPM pathway, a Trp decarboxylase gene may be involved in the tryptophan decarboxylation reaction to TPM, while monoamine oxidases (MAO) may convert TPM to IAAld. In both IPA and TPM pathways, the final step is the conversion of IAAld into IAA, which is catalyzed by aldehyde dehydrogenase (ALDH). Three aldH genes detected in RZ2MS9 may encode this enzyme.
IAA production is influenced by the incubation period of B. thuringiensis RZ2MS9 and Trp concentration
To investigate the effect of RZ2MS9 incubation period on the IAA production, samples were collected from the culture media at 3, 12, 15, 18, 21, and 30 h after inoculation (HAI). Data analysis suggested that, overall, the bacterial cell density and the amount of auxin detected were positively and significantly correlated (r = 0.72, p-value < 0.01). In other words, the more RZ2MS9 cells present in the medium, the higher the amount of IAA detected.
At 3 HAI, approximately 0.06 μg/ml of auxin was detected in the medium (Fig. 2a). This amount remained constant during the log phase of bacterial growth (3–12 HAI). The bacterium reached the stationary phase of growth approximately 15 HAI, and maximum IAA (0.20 μg/ml) was produced by RZ2MS9 at 21 HAI, still during the stationary phase. Subsequently, IAA production declined, reaching around 0.10 μg/ml at 30 HAI.
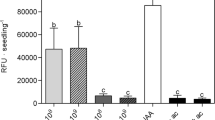
Factors that influence the production of IAA by B. thuringiensis RZ2MS9. a The influence of the incubation period on IAA production (μg/ml) and growth (optical density—OD at 600 nm) of RZ2MS9. The bacterium was grown in LB medium supplemented with 1 g/l of l-tryptophan (Trp). The results shown are representative of 3 replicates of the experiment. b Effect of Trp concentration on IAA production (μg/ml) and growth (OD at 600 nm) of RZ2MS9. The bacterium was grown in LB medium supplemented with different amounts of Trp (0, 0.1, 0.5, 1.0 and 2.0 g/l). The results shown are representative of 3 replicates of the experiment. We measured both the IAA production and bacterial growth 21 h after the beginning of the incubation period
We observed that B. thuringiensis RZ2MS9 growth was not affected by different concentrations of the Trp (Fig. 2B). The bacterial IAA production increased when the concentration of Trp in the medium was increased from 0.1 to 1 g/l. RZ2MS9 showed maximal IAA production (0.20 μg/ml) when 1 g/l of Trp was added to the medium. At this concentration of Trp, bacterial IAA production was around five times higher than when the bacterium was grown in the medium without Trp. On the other hand, IAA production by RZ2MS9 slightly declined to around 0.18 μg/ml in the medium containing 2 g/l of Trp. Data analysis indicated that the auxin production and Trp concentration in the medium were positively and significantly correlated (r = 0.76, p-value < 0.01).
Effect of B. thuringiensis RZ2MS9 inoculation on MT and dgt growth
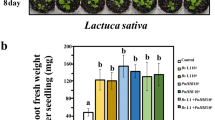
The inoculation with the auxin-producing B. thuringiensis RZ2MS9 showed significant differences in several parameters of MT tomato growth (Figs. 3 and 4). We observed a significant increase of 24% in average shoot dry weight (SDW) of MT plants inoculated with RZ2MS9 (MT_RZ2MS9) when compared to MT plants without the bacterial treatment (MT_Control). However, the SDW of dgt plants inoculated (dgt_RZ2MS9) and non-inoculated (dgt_Control) with RZ2MS9 did not differ from each other. It is also worth mentioning that the application of RZ2MS9 seems to have accelerated the flowering in MT plants (Fig. 3).
Effect on the shoot dry weight (SDW) of 60-day old dgt and MT plants either with or without the application of B. thuringiensis RZ2MS9. Mean values (9 replicates per treatment) followed by different capital letters represent significant differences between the plant genotypes (dgt × MT). Means followed by lowercase letters represent significant differences among all the treatments according to Tukey’s test (p < 0.05)
Effect on several root parameters of 60-day old dgt and MT plants either with or without the application of B. thuringiensis RZ2MS9 (a–e), and effect of bacterial application on the MT root architecture (f and g). a Total root length (TRL), b lateral root length (LRL), c axial root length (ARL), d root superficial area (RSA), e total root diameter (TRD). Mean values (9 replicates per treatment) followed by different capital letters represent significant differences between the plant genotypes (dgt × MT). Means values followed by lowercase letters represent significant differences among all the treatments according to Tukey’s test (p < 0.05). f Roots of MT plants with RZ2MS9 application and g roots of MT plants without bacterial application. Ctrl , non-inoculated control
The root architecture of MT plants inoculated with B. thuringiensis RZ2MS9 was modified when compared to MT_Control plants (Fig. 4f and g). The bacterial inoculation led to an average significant increase of 25, 26 and 17% in total root length (TRL) (Fig. 4a), lateral root length (LRL) (Fig. 4b) and total root surface area (RSA) (Fig. 4d), respectively, and a significant decrease of 7% in the total root diameter (TRD) (Fig. 4e) of MT_RZ2MS9 plants when compared to MT_Control plants. No statistically significant effects on these parameters were observed when comparing dgt_RZ2MS9 plants and dgt_Control. Besides that, the axial root length (ARL) did not differ between the treatments MT_RZ2MS9 and MT_Control as well as between dgt_RZ2MS9 and dgt_Control (Fig. 4c).
Anatomical changes
We analyzed the anatomy of the tips of MT plants’ roots with and without application of RZ2MS9. In this region, we observed a single-epidermal cell layer, 6–7 layers of cortical cells, and the vascular cylinder where the procambium is differentiated into protoxylem and phloem cells (Fig. 5a–h). Two main alterations were detected in the cortical cells of MT_RZ2MS9 root tips: (1) the are more elongated (Fig. 5c, d) than MT_Control plants’ roots (Fig. 5a, b); (2) and exhibited more mitotic activity (Fig. 5g, h) when compared to non-inoculated plants (Fig. 5e, f).
Longitudinal sections of the Micro-tom MT root tips either with (MT_RZ2MS9) (c, d; g, h) or without (MT_Control) (a, b, e, f) the B. thuringiensis RZ2MS9 application. b and f are details of squares in a and e, respectively, while d and h are details of squares in c and g, respectively. All detailed regions are associated to the first developed protoxylem cells (arrows in b, f, d, and h). c and d presented larger cortical cells than the control. g and h presented higher mitotic activity of cortical cells. CT, Cortex; EP, epidermis; PH, phloem; XY, xylem; PR, protoxylem
Discussion
RZ2MS9 is currently classified as Bacillus thuringiensis
The vast majority of bacteria present in the rhizosphere produce auxins such as the indole-3-acetic acid (IAA) (Patten and Glick 1996). From a mutualistic point of view, rhizobacteria synthesize auxins to enlarge the superficial area of the roots, thus increasing plant exudation and nutrient availability in the rhizosphere. In exchange, plant development is favored by greater access to water and soil nutrients (Etesami et al. 2015). This mechanism is very likely the main cause of plant growth promotion by PGPR.
Strains belonging to the genus Bacillus are among the most commonly reported PGPR (Kloepper et al. 2004; Idris et al. 2007). In particular B. thuringiensis, which is well-known by its insecticidal activities, has not received the deserved attention regarding its potential to promote plant growth, even with many studies reporting that it enhances plant development (Selvakumar et al. 2008; Mishra et al. 2009; Armada et al. 2015). B. thuringiensis belongs to the B. cereus sensu lato group which comprises several species, but the most-well studied are B. cereus, B. thuringiensis and B. anthracis (Liu et al. 2015; Ehling-Schulz et al. 2019). The taxonomy of B. cereus group is complex and recent studies have used ANI (Carroll et al. 2020; Torres Mano et al. 2020) and pan- and core-genome analyses (Bazinet 2017; Baek et al. 2019) to clarify species definition and investigate the group phylogeny without relying on the analysis of plasmids, which can be easily transferred or lost in the evolutionary history (Liu et al. 2015), but still there is no consensus. Here, we used both ANI and core-genome analyses to classify RZ2MS9 as B. thuringiensis. Interestingly, in both analyses we found a close similarity between the B. thuringiensis subsp. jinghongiensis strain YGd22-03 (Wu et al. 2017) and the RZ2MS9, both originally isolated from soil.
RZ2MS9 genome mining for genes likely involved in IAA biosynthesis
Despite the importance of Bacillus as PGPR, most of the studies portraying bacterial IAA production are based on Gram-negative strains (Wahyudi et al. 2011; Ortiz-Castro et al. 2020). However, some IAA-producing Gram-positive Bacillus sp. strains have started to receive more attention in the last few decades (Idris et al 2004). For instance, the role of IAA produced by B. amyloliquefaciens FZB42 in plant growth promotion was validated by the use of mutant strains deficient in a gene responsible for IAA biosynthesis, suggesting that the main route of IAA biosynthesis in this bacterium was dependent on Trp (Idris et al. 2007). The Trp, naturally present in root exudates of plants (Olanrewaju et al. 2017) can be converted to IAA through several pathways. More than one pathway could be present in bacteria (Patten and Glick 1996). Here, two complete Trp-dependent IAA biosynthesis pathways, the indole-3-pyruvate (IPA) and the tryptamine (TPM), were identified in the genome of B. thuringiensis RZ2MS9.
Six of the detected genes might be involved in the IPA pathway: two amino acid transferases (or aminotransferases), the indolepyruvate decarboxylase (ipdC) and three aldehyde dehydrogenases (aldH). Aminotransferases catalyze the reversible transamination of Trp to IPA. Bacteria can possess several isoforms of aminotransferases which are involved in multiple pathways (Whitaker et al. 1982). The activity of these enzymes in IAA-producing bacteria has been demonstrated mainly in Gram-negative bacteria such as Pseudomonas sp. and Azospirillum sp. (Baca et al. 1994; Pedraza et al. 2004). The decarboxylation of IPA to indole-3-acetaldehyde (IAAld) by the action of the enzyme indole-3-pyruvate decarboxylase (IPDC), encoded by the ipdC gene, appears to be the rate-limiting step of the IPA pathway. This gene is well-documented, and its disruption significantly decreased the IAA production of Gram-negative strains such as Pseudomonas putida (Patten and Glick 2002), Erwinia herbicola (Brandl and Lindow 1998), Azospirillium brasilense (Dobbelaere et al. 1999; Malhotra and Srivastava 2008) and Klebsiella sp. (Gang et al. 2018). However, the presence of this gene in the genome of IAA-producing Gram-positive bacteria, especially in Bacillus sp. strains, is less common. For instance, the gene was not detected in the genome of the well-studied auxin-producing B. amyloliquefaciens strains FZB42 (Idris et al. 2007), and SQR9 (Shao et al. 2015a). Interestingly, some isolates of B. thuringiensis were reported to possess ipdC gene in their genomes (Raddadi et al. 2008).
In addition, six of the detected genes in the RZ2MS9 are possibly involved in the IAM pathway: one Trp decarboxylase (tdc), two monoamine oxidases (mao), and the three aldehyde dehydrogenases (aldH) which are shared with the IPA pathway. Recently, a large-scale genomic study revealed that rhizobacteria predominantly synthesize IAA via the indole-3-acetamide (IAM) and TPM pathways. More precisely, the Firmicutes, the taxa group to which Bacillus sp. belongs, likely mainly use the TPM pathway for IAA production since this is the main pathway detected among the hundreds of genomes studied (Zhang et al. 2019). The TPM pathway is present in plants (Mano and Nemoto 2012) and fungi (Bunsangiam et al. 2019); however, it is much less well-documented in bacteria (Cassán et al. 2014; Williams et al. 2014). This pathway was suggested for Bacillus cereus (Perley and Stowe 1966) and Azospirillum brasilense (Hartmann et al. 1983) based on their ability to produce IAA from TPM in chemically defined culture medium (Cassán et al. 2014). Genomes of Pseudomonas putida strains were reported to encode enzymes involved in the TPM pathway, including a putative tryptophan decarboxylase, a putative amine oxidase, and several candidate aldehyde dehydrogenases (ALDH) (Wu et al. 2011; Kunkel and Harper 2018). In the RZ2MS9 genome, we identified three aldH genes, which are predicted to encode the ALDH involved in the final step of both IPA and TPM pathways. A gene encoding a potential ALDH was also detected in the PGPR B. amyloliquefaciens SQR9 genome and in response to the addition of Ltrp to the medium its transcription increased 2.5-fold (Shao et al. 2015a). All genes involved in the IAA pathways detected in the RZ2MS9 genome will serve as a foundation for future functional studies.
IAA production is influenced by the incubation period of B. thuringiensis RZ2MS9 and Trp concentration
It has been reported that IAA production by bacteria is influenced by the growth stage and Trp concentration added to the medium (Myo et al. 2019). We tested both conditions in the present study. Firstly, we detected that maximum IAA was produced by RZ2MS9 at 21 HAI, during the stationary phase. IAA is produced by some bacterial strains, including Bacillus sp. strains, when they are in the stationary phase of growth (Patten et al. 2013; Ali 2015; Shim et al. 2015). It was suggested that IAA production can be induced in response to cell density-dependent signal (Quorum Sensing—QS). The inactivation of N-acylhomoserine lactones (AHLs), QS-related signalling compounds, reduced IAA production by the Gram-negative Azospirillum lipoferum during the stationary phase (Boyer et al. 2008). Besides that, increased bacterial auxin production during the stationary phase may be related to stress factors (such as carbon and nitrogen depletion in the medium, reduction in growth rate, and oxygen deprivation) triggering the expression of IAA-related genes (Ona et al. 2005). The regulation of genes such as the ipdC in some Gram-negative bacteria is performed by the rpoS gene, the stress response sigma factor (Patten and Glick 2002). Moreover, bacteria may be able to obtain the maximum Trp from the dead bacterial mass during the stationary phase, which could result in increased IAA production during this phase (Swain et al. 2007).
Subsequently, we studied the effect of Trp concentration on RZ2MS9 IAA production. The application of 1 g/l of Trp to the medium resulted in an almost fivefold increase in IAA secretion by the bacterium. Similarly, Shao et al. (2015b) reported an increase of almost 3.6-fold in IAA produced by B. amyloliquefaciens SQR9 when Trp was added to the medium. Furthermore, our experiments showed that the auxin production by RZ2MS9 is positively and significantly correlated with the amount of Trp added to the medium. Likewise, IAA production by Paenibacillus polymyxa SK1was strongly correlated with the application of exogenous tryptophan concentrations in the growth medium (Khan et al. 2020).
Therefore, we found that growing the RZ2MS9 in a medium supplemented with 1 g/l of Trp until the stationary phase enhanced its auxin production, which reached the maximal value of 0.20 μg/ml in our studies. Shreya et al. (2020) also reported that the plant growth promoting and chromium-tolerant B. thuringiensis strain S3VKR2 produced around 0.15 μg/ml of IAA when grown for 5 days in a medium supplemented with Trp. In this sense, longer term analysis (during the late stationary phase) will be carried out in the future to reveal whether the auxin amount produced by RZ2MS9 remains constant, decreases or increases.
Effect of B. thuringiensis RZ2MS9 inoculation on MT and dgt root development and anatomical changes
The Micro-Tom MT (Solanum lycorpersicum cv Micro-Tom) is a miniature tomato with various advantageous properties to serve as a model system to study plant-microbes interaction (Park et al. 2007; Campos et al. 2010). This plant has been used in plant growth promotion studies with a range of microbes, including fungi and actinomycetes (Palaniyandi et al. 2014; Fiorini et al. 2016). MT and dgt have been used in combination to confirm auxin-like activity in the humic matter (Dobbss et al. 2007; Canellas et al. 2011) and to investigate the role of phytohormones in mediating the effect of cadmium on growth and metabolism of tomato (Chu et al. 2020). However, to the best of our knowledge, this is the first study using this study model to investigate the role of auxin produced by a PGPR.
The inoculation of B. thuringiensis RZ2MS9 significantly increased the lateral roots length (LRL) by 26% while it inhibited the growth of axial roots length (ARL) in MT plants when compared to non-inoculated MT plants. It is generally accepted that auxin inhibits root elongation and promotes the development of lateral roots (Sukumar et al. 2013). Besides that, the reduction by 7% of total mean root diameter (TRD) by application of the bacterium is more evidence of the increased development of lateral roots, which have a smaller diameter when compared to axial roots. Although thicker roots have a greater capacity to transport water and nutrients and are less vulnerable to adverse edaphic conditions, fine roots allow a much larger soil exploration (Richardson et al. 2009). Altogether, the modifications in the root architecture of MT plants caused by RZ2MS9 certainly had a great influence in the improvement of water and nutrients acquisition by the plants, which was reflected in the significant increase of 24% in MT shoot dry weight. Interestingly, despite not being quantified, it seems that the bacterial application caused an acceleration of flowering time in MT plants. The effect caused on flowering, as well as on the final fruit production, will be further investigated. On the other hand, the inoculation of the auxin-insensitive dgt plants with RZ2MS9 did not affect any of the roots and shoots parameters evaluated. This is an additional indication of the relevance of the auxin produced by RZ2MS9 in the plant response.
Regarding the anatomical alterations, herein we observed that, for MT_RZ2MS9 plants, at the differentiation zone of the root where the first protoxylem starts to differentiate, cortical cells are expanded, and their mitotic activity seems to be accelerated when comparing to roots from non-inoculated MT plants. This data suggests a modification at cellular level possibly induced by phytohormones. Plant responses commonly related to auxins include cell enlargement, cell division, increased protein, RNA synthesis, and RNA polymerase activity (Ahmed and Hasnain 2014). Kollarova et al. (2004), investigating the effect of auxins on Karwinskia humboldtiana roots, observed that IAA promotes the mitotic activity and can also induce the plant ethylene production, which can lead to cell enlargement. Taken together, our results provide evidence that plant growth-promotion elicited by the PGPR B. thuringiensis RZ2MS9 likely results from its auxin production trait. However, it is possible that the bacterium can interfere in other phytohormone pathways that will lead to increased plant growth. This indirect effect needs further studies.
Deciphering which PGP trait is relevant and how to optimize the cultivation of the bacterium to obtain the best expression of this trait has key importance for the consistent use of PGPR in agricultural systems (Martínez-Viveros et al. 2010). Here, we demonstrate that B. thuringiensis RZ2MS9 not only harbours genes involved in IAA production from Trp but also produces around five times more IAA during the stationary phase in a medium supplemented with 1 g/l of Trp. The anatomical analysis showed that MT plants applied with RZ2MS9 had the root cortical cells either expanded or with high mitotic activity when compared with MT plants without bacterial application. Moreover, we have brought to light a study model (MT + dgt) that seems to be useful in elucidating the auxin production mechanism used by PGPR to promote plant growth.
Availability of data and material
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Ahmed A, Hasnain S (2014) Auxins as one of the factors of plant growth improvement by plant growth promoting rhizobacteria. Pol J Microbiol 63:261–266. https://doi.org/10.33073/pjm-2014-035
Ali B (2015) Bacterial auxin signaling: comparative study of growth induction in Arabidopsis thaliana and Triticum aestivum. Turk J Bot 39:1–9. https://doi.org/10.3906/bot-1401-31
Ali B, Sabri AN, Ljung K, Hasnain S (2009) Quantification of indole-3-acetic acid from plant associated Bacillus spp. and their phytostimulatory effect on Vigna radiata (L.). World J Microb Biot 25:519–526. https://doi.org/10.1007/s11274-008-9918-9
Armada E, Azcón R, López-Castillo OM, Calvo-Polanco M, Ruiz-Lozano JM (2015) Autochthonous arbuscular mycorrhizal fungi and Bacillus thuringiensis from a degraded Mediterranean area can be used to improve physiological traits and performance of a plant of agronomic interest under drought conditions. Plant Physiol Biochem 90:64–74. https://doi.org/10.1016/j.plaphy.2015.03.004
Baca BE, Soto-Urzua L, Xochihua-Corona YG, Cuervo-Garcia A (1994) Characterization of two aromatic amino acid aminotransferases and production of indoleacetic acid in Azospirillum strains. Soil Biol Biochem 26:57–63. https://doi.org/10.1016/0038-0717(94)90195-3
Baek I, Lee K, Goodfellow M, Chun J (2019) Comparative genomic and phylogenomic analyses clarify relationships within and between Bacillus cereus and Bacillus thuringiensis: proposal for the recognition of two Bacillus thuringiensis genomovars. Front Microbiol 10:1978. https://doi.org/10.3389/fmicb.2019.01978
Batista BD, Lacava PT, Ferrari A, Teixeira-Silva NS, Bonatelli ML, Tsui S, Mondin M, Kitajima EW, Pereira JO, Azevedo JL, Quecine MC (2018) Screening of tropically derived, multi-trait plant growth-promoting rhizobacteria and evaluation of corn and soybean colonization ability. Microbiol Res 206:33–42. https://doi.org/10.1016/j.micres.2017.09.007
Bazinet A (2017) Pan-genome and phylogeny of Bacillus cereus sensu lato. BMC Evol Biol 17:176. https://doi.org/10.1186/s12862-017-1020-1
Bergfeld R, Speth V, Schopfer P (1988) Reorientation of microfibrils and microtubules at the outer epidermal wall of maize coleoptiles during auxin-mediated growth. Bot Acta 101:57–67. https://doi.org/10.1111/j.1438-8677.1988.tb00012.x
Bloemberg GV, Lugtenberg BJ (2001) Molecular basis of plant growth promotion and biocontrol by rhizobacteria. Curr Opin Plant Biol 4:343–350. https://doi.org/10.1016/S1369-5266(00)00183-7
Bonatelli ML, Li J, Dumaresq A, Quecine MC, Settles Matthew L (2020) Complete genome sequence of Bacillus sp. strain RZ2MS9, a multi-trait plant growth promoter. Microbiol Resour Announc 9:e00623-e720. https://doi.org/10.1128/MRA.00623-20
Boyer M, Bally R, Perrotto S, Chaintreuil C, Wisniewski-Dyé F (2008) A quorum-quenching approach to identify quorum-sensing-regulated functions in Azospirillum lipoferum. Res Microbiol 159:699–708. https://doi.org/10.1016/j.resmic.2008.08.003
Brandl MT, Lindow SE (1998) Contribution of indole-3-acetic acid production to the epiphytic fitness of Erwinia herbicola. Appl Environ Microbiol 64:3256–3263. https://doi.org/10.1128/AEM.64.9.3256-3263.1998
Bunsangiam S, Sakpuntoon V, Srisuk N, Ohashi T, Fujiyama K, Limtong S (2019) Biosynthetic pathway of indole-3-acetic acid in basidiomycetous yeast Rhodosporidiobolus fluvialis. Mycobiol 47:292–300. https://doi.org/10.1080/12298093.2019.1638672
Campos ML, Carvalho RF, Benedito VA, Peres LEP (2010) Small and remarkable: the micro-Tom model system as a tool to discover novel hormonal functions and interactions. Plant Signal Behav 5:267–270. https://doi.org/10.4161/psb.5.3.10622
Canellas LP, Dantas DJ, Aguiar NO et al (2011) Probing the hormonal activity of fractionated molecular humic components in tomato auxin mutants. Ann Appl Biol 159:202–211. https://doi.org/10.1111/j.1744-7348.2011.00487.x
Carroll LM, Wiedmann M, Kovac J (2020) Proposal of a taxonomic nomenclature for the Bacillus cereus group which reconciles genomic definitions of bacterial species with clinical and industrial phenotypes. Mbio 11:e00034-e120. https://doi.org/10.1128/mBio.00034-20
Carvalho RF, Campos ML, Pino LE et al (2011) Convergence of developmental mutants into a single tomato model system: “Micro-Tom” as an effective toolkit for plant development research. Plant Methods 7:18. https://doi.org/10.1186/1746-4811-7-18
Cassán F, Vanderleyden J, Spaepen S (2014) Physiological and agronomical aspects of phytohormone production by model Plant-growth-promoting rhizobacteria (PGPR) belonging to the genus Azospirillum. J Plant Growth Regul 33:440–459. https://doi.org/10.1007/s00344-013-9362-4
Chu Z, Munir S, Zhao G et al (2020) Linking phytohormones with growth, transport activity and metabolic responses to cadmium in tomato. Plant Growth Regul 90:557–569. https://doi.org/10.1007/s10725-020-00580-w
Daniel SG, Rayle DL, Cleland RE (1989) Auxin physiology of the tomato mutant diageotropica. Plant Physiol 91:804–807. https://doi.org/10.1104/pp.91.3.804
Dobbelaere S, Croonenborghs A, Thys A, Broek AV, Vanderleyden J (1999) Phytostimulatory effect of Azospirillum brasilense wild type and mutant strains altered in IAA production on wheat. Plant Soil 212:153–162. https://doi.org/10.1023/A:1004658000815
Dobbss LB, Medici LO, Peres LEP, Pino-Nunes LE, Rumjanek VM, Façanha AR, Canellas LP (2007) Changes in root development of Arabidopsis promoted by organic matter from oxisols. Ann Appl Biol 151:199–211. https://doi.org/10.1111/j.1744-7348.2007.00166.x
Ehling-Schulz M, Lereclus D, Koehler TM (2019) The Bacillus cereus group: Bacillus species with pathogenic potential. In: Fischetti VA, Novick RP, Ferretti JJ, Portnoy DA, Braunstein M, Rood JI (eds) Gram-positive pathogens, 3rd edn. ASM Press, Washington, pp 875–902. https://doi.org/10.1128/9781683670131.ch55
Etesami H, Alikhani HA, Hosseini HM (2015) Indole-3-acetic acid (IAA) production trait, a useful screening to select endophytic and rhizosphere competent bacteria for rice growth promoting agents. MethodsX 2:72–78. https://doi.org/10.1016/j.mex.2015.02.008
Fiorini L, Guglielminetti L, Mariotti L et al (2016) Trichoderma harzianum T6776 modulates a complex metabolic network to stimulate tomato cv. Micro-Tom Growth Plant Soil 400:351–366. https://doi.org/10.1007/s11104-015-2736-6
Gang S, Saraf M, Waite CJ, Buck M, Schumacher J (2018) Mutualism between Klebsiella SGM 81 and Dianthus caryophyllus in modulating root plasticity and rhizospheric bacterial density. Plant Soil 424:273–288. https://doi.org/10.1007/s11104-017-3440-5
Gordon SA, Weber RP (1951) Colorimetric estimation of indoleacetic acid. Plant Physiol 26:192–195. https://doi.org/10.1104/pp.26.1.192
Hartmann A, Singh M, Klingmüller W (1983) Isolation and characterization of Azospirillum mutants excreting high amounts of indoleacetic acid. Can J Microbiol 29:916–923. https://doi.org/10.1139/m83-147
Hicks GR, Rayle DL, Lomax TL (1989) The diageotropica mutant of tomato lacks high specific activity auxin binding sites. Science 245:52–54. https://doi.org/10.1126/science.245.4913.52
Idris EE, Bochow H, Ross H, Borriss R (2004) Use of Bacillus subtilis as biocontrol agent. VI. Phytohormone-like action of culture filtrates prepared from plant growth-promoting Bacillus amyloliquefaciens FZB24, FZB42, FZB45 and Bacillus subtilis FZB37. J Plant Dis Protec 111:583–597
Idris EE, Iglesias DJ, Talon M, Borriss R (2007) Tryptophan-dependent production of indole-3-acetic acid (IAA) affects level of plant growth promotion by Bacillus amyloliquefaciens FZB42. Mol Plant Microbe Interact 20:619–626. https://doi.org/10.1094/MPMI-20-6-0619
Jain C, Rodriguez-RLM PAM, Konstantinidis KT, Aluru S (2018) High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun 9:5114. https://doi.org/10.1038/s41467-018-07641-9
Karnovsky JM (1965) A formaldehyde glutaraldehyde fixative of high osmolality for use in electron microscopy. J Cell Biol 27:137-138A
Kelly MO, Bradford KJ (1986) Insensitivity of the diageotropica tomato mutant to auxin. Plant Physiol 82:713–717. https://doi.org/10.1104/pp.82.3.713
Khan MS, Gao J, Chen X et al (2020) Isolation and characterization of plant growth-promoting endophytic bacteria Paenibacillus polymyxa SK1 from Lilium lancifolium. Biomed Res Int. https://doi.org/10.1155/2020/8650957
Kimura S, Sinha N (2008) Tomato (Solanum lycopersicum): a model fruit-bearing crop. Cold Spring Harb Protoc 11:1pdb-emo105. https://doi.org/10.1101/pdb.emo105
Kloepper JW, Ryu CM, Zhang S (2004) Induced systemic resistance and promotion of plant growth by Bacillus spp. Phytopathol 94:1259–1266. https://doi.org/10.1094/PHYTO.2004.94.11.1259
Kollarova K, Liskova D, Kakoniova D, Lux A (2004) Effect of auxins on Karwinskia humboldtiana root cultures. Plant Cell Tiss Organ 79:213–221. https://doi.org/10.1007/s11240-004-0662-z
Kunkel BN, Harper CP (2018) The roles of auxin during interactions between bacterial plant pathogens and their hosts. J Exp Bot 69:245–254. https://doi.org/10.1093/jxb/erx447
Letunic I, Bork P (2019) Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:W256–W259. https://doi.org/10.1093/nar/gkz239
Liu L, Guo G, Wang Z, Ji H, Mu F, Li X (2014) Auxin in plant growth and stress responses. In: Tran LSP, Pal S (eds) Phytohormones: a window to metabolism, signaling and biotechnological applications. Springer, Berlin. https://doi.org/10.1007/978-1-4939-0491-4_1
Liu Y, Lai Q, Göker M, Meier-Kolthoff JP, Wang M, Sun Y, Wang L, Shao Z (2015) Genomic insights into the taxonomic status of the Bacillus cereus group. Sci Rep 5:14082. https://doi.org/10.1038/srep14082
Loper JE, Schroth MN (1986) Influence of bacterial sources of indole-3-acetic acid on root elongation of sugar beet. Phytopathology 76:386–389. https://doi.org/10.1094/Phyto-76-386
Malhotra M, Srivastava S (2008) An ipdC gene knock-out of Azospirillum brasilense strain SM and its implications on indole-3-acetic acid biosynthesis and plant growth promotion. Antonie Van Leeuwenhoek 93:425–433. https://doi.org/10.1007/s10482-007-9207-x
Mano Y, Nemoto K (2012) The pathway of auxin biosynthesis in plants. J Exp Bot 63:2853–2872. https://doi.org/10.1093/jxb/ers091
Martínez-Viveros O, Jorquera MA, Crowley DE, Gajardo GMLM, Mora ML (2010) Mechanisms and practical considerations involved in plant growth promotion by rhizobacteria. J Soil Sci Plant Nutr 10:293–319. https://doi.org/10.4067/S0718-95162010000100006
Mishra PK, Mishra S, Selvakumar G, Kundu S, Gupta HS (2009) Enhanced soybean (Glycine max L.) plant growth and nodulation by Bradyrhizobium japonicum-SB1 in presence of Bacillus thuringiensis-KR1. Acta Agr Scand B-SP 59:189–196. https://doi.org/10.1080/09064710802040558
Myo EM, Ge B, Ma J et al (2019) Indole-3-acetic acid production by Streptomyces fradiae NKZ-259 and its formulation to enhance plant growth. BMC Microbiol 19:155. https://doi.org/10.1186/s12866-019-1528-1
O’Brien T, Feder N, McCully ME (1964) Polychromatic staining of plant cell walls by toluidine blue O. Protoplasma 59:368–373. https://doi.org/10.1007/BF01248568
Oh K, Ivanchenko MG, White TJ, Lomax TL (2006) The diageotropica gene of tomato encodes a cyclophilin: a novel player in auxin signaling. Planta 224:133–144. https://doi.org/10.1007/s00425-005-0202-z
Olanrewaju OS, Glick BR, Babalola OO (2017) Mechanisms of action of plant growth promoting bacteria. World J Microbiol Biotechnol 33:197. https://doi.org/10.1007/s11274-017-2364-9
Ona O, Van Impe J, Prinsen E, Vanderleyden J (2005) Growth and indole-3-acetic acid biosynthesis of Azospirillum brasilense Sp245 is environmentally controlled. FEMS Microbiol Lett 246:125–132. https://doi.org/10.1016/j.femsle.2005.03.048
Ortiz-Castro R, Campos-García J, López-Bucio J (2020) Pseudomonas putida and Pseudomonas fluorescens influence Arabidopsis root system architecture through an auxin response mediated by bioactive cyclodipeptides. J Plant Growth Regul 39:254–265. https://doi.org/10.1007/s00344-019-09979-w
Overvoorde P, Fukaki H, Beeckman T (2010) Auxin control of root development. Cold Spring Harb Perspect Biol 2:a001537. https://doi.org/10.1101/cshperspect.a001537
Page AJ, Cummins CA, Hunt M, Wong VK, Reuter S, Holden MT, Fookes M, Falush D, Keane JA, Parkhill J (2015) Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31:3691–3693. https://doi.org/10.1093/bioinformatics/btv421
Palaniyandi SA, Damodharan K, Yang SH, Suh JW (2014) Streptomyces sp. strain PGPA39 alleviates salt stress and promotes growth of ‘Micro Tom’ tomato plants. J Appl Microbiol 117:766–773. https://doi.org/10.1111/jam.12563
Park EJ, Lee SD, Chung EJ et al (2007) Micro Tom—a model plant system to study bacterial wilt by Ralstonia solanacearum. Plant Pathol J 23:239–244. https://doi.org/10.5423/PPJ.2007.23.4.239
Patten CL, Glick BR (1996) Bacterial biosynthesis of indole-3-acetic acid. Can J Microbiol 42:207–220. https://doi.org/10.1139/m96-032
Patten CL, Glick BR (2002) Role of Pseudomonas putida indoleacetic acid in development of the host plant root system. Appl Environ Microbiol 68:3795–3801. https://doi.org/10.1128/aem.68.8.3795-3801.2002
Patten CL, Blakney AJ, Coulson TJ (2013) Activity, distribution and function of indole-3-acetic acid biosynthetic pathways in bacteria. Crit Rev Microbiol 39:395–415. https://doi.org/10.3109/1040841X.2012.716819
Pedraza RO, Ramírez-Mata A, Xiqui ML, Baca BE (2004) Aromatic amino acid aminotransferase activity and indole-3-acetic acid production by associative nitrogen-fixing bacteria. FEMS Microbiol Lett 233:15–21. https://doi.org/10.1016/j.femsle.2004.01.047
Perley JE, Stowe BB (1966) On the ability of Taphrina deformans to produce indoleacetic acid from tryptophan by way of tryptamine. Plant Physiol 41:234–237. https://doi.org/10.1104/pp.41.2.234
Price MN, Dehal PS, Arkin AP (2010) FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5:e9490. https://doi.org/10.1371/journal.pone.0009490
R Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Raddadi N, Cherif A, Boudabous A, Daffonchio D (2008) Screening of plant growth promoting traits of Bacillus thuringiensis. Ann Microbiol 58:47–52. https://doi.org/10.1007/BF03179444
Richardson AE, Barea JM, McNeill AM, Prigent-Combaret C (2009) Acquisition of phosphorus and nitrogen in the rhizosphere and plant growth promotion by microorganisms. Plant Soil 321:305–339. https://doi.org/10.1007/s11104-009-9895-2
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, p A2.2
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Selvakumar G, Kundu S, Gupta AD, Shouche YS, Gupta HS (2008) Isolation and characterization of nonrhizobial plant growth promoting bacteria from nodules of kudzu (Pueraria thunbergiana) and their effect on wheat seedling growth. Curr Microbiol 56:134–139. https://doi.org/10.1007/s00284-007-9062-z
Shao J, Li S, Zhang N et al (2015a) Analysis and cloning of the synthetic pathway of the phytohormone indole-3-acetic acid in the plant-beneficial Bacillus amyloliquefaciens SQR9. Microb Cell Fact 14:130. https://doi.org/10.1186/s12934-015-0323-4
Shao J, Xu Z, Zhang N, Shen Q, Zhang R (2015b) Contribution of indole-3-acetic acid in the plant growth promotion by the rhizospheric strain Bacillus amyloliquefaciens SQR9. Biol Fertil Soils 51:321–330. https://doi.org/10.1007/s00374-014-0978-8
Shim J, Kim JW, Shea PJ, Oh BT (2015) IAA production by Bacillus sp. JH 2–2 promotes Indian mustard growth in the presence of hexavalent chromium. J Basic Microbiol 55:652–658. https://doi.org/10.1002/jobm.201400311
Shreya D, Jinal HN, Kartik VP et al (2020) Amelioration effect of chromium-tolerant bacteria on growth, physiological properties and chromium mobilization in chickpea (Cicer arietinum) under chromium stress. Arch Microbiol 202:887–894. https://doi.org/10.1007/s00203-019-01801-1
Spaepen S, Vanderleyden J (2011) Auxin and plant-microbe interactions. Cold Spring Harb Perspect Biol 3:a001438. https://doi.org/10.1101/cshperspect.a001438
Steenhoudt O, Vanderleyden J (2000) Azospirillum, a free-living nitrogen-fixing bacterium closely associated with grasses: genetic, biochemical and ecological aspects. FEMS Microbiol Rev 24:487–506. https://doi.org/10.1111/j.1574-6976.2000.tb00552.x
Sukumar P, Legue V, Vayssieres A, Martin F, Tuskan GA, Kalluri UC (2013) Involvement of auxin pathways in modulating root architecture during beneficial plant–microorganism interactions. Plant Cell Environ 36:909–919. https://doi.org/10.1111/pce.12036
Swain MR, Naskar SK, Ray RC (2007) Indole-3-acetic acid production and effect on sprouting of yam (Dioscorea rotundata L.) minisetts by Bacillus subtilis isolated from culturable cowdung microflora. Pol J Microbiol 56:103–110
Torres Manno MA, Repizo GD, Magni C, Dunlap CA, Espariz M (2020) The assessment of leading traits in the taxonomy of the Bacillus cereus group. Antonie Van Leeuwenhoek 113:2223–2242. https://doi.org/10.1007/s10482-020-01494-3
Trachsel S, Messmer R, Stamp P, Hund A (2009) Mapping of QTLs for lateral and axile root growth of tropical maize. Theor Appl Genet 119:1413–1424. https://doi.org/10.1007/s00122-009-1144-9
Wahyudi AT, Astuti RP, Widyawati A, Mery A, Nawangsih AA (2011) Characterization of Bacillus sp. strains isolated from rhizosphere of soybean plants for their use as potential plant growth for promoting rhizobacteria. J Microbiol Antimicrob 3:34–40
Whitaker RJ, Gaines CG, Jensen RA (1982) A multispecific quintet of aromatic aminotransferases that overlap different biochemical pathways in Pseudomonas aeruginosa. J Biol Chem 257:13550–13556
Williams BB, Van Benschoten AH, Cimermancic P et al (2014) Discovery and characterization of gut microbiota decarboxylases that can produce the neurotransmitter tryptamine. Cell Host Microbe 16:495–503. https://doi.org/10.1016/j.chom.2014.09.001
Wu X, Monchy S, Taghavi S, Zhu W, Ramos J, van der Lelie D (2011) Comparative genomics and functional analysis of niche-specific adaptation in Pseudomonas putida. FEMS Microbiol Rev 35:299–323. https://doi.org/10.1111/j.1574-6976.2010.00249.x
Wu Y, Fu Y, Yuan Y, Gao M (2017) Complete genome sequence of Bacillus thuringiensis subsp. jinghongiensis reference strain YGd22-03. Genome Announc 5:e00740-e817. https://doi.org/10.1128/genomeA.00740-17
Zandonadi DB, Santos MP, Caixeta LS, Marinho EB, Peres LEP, Façanha AR (2016) Plant proton pumps as markers of biostimulant action. Sci Agr 73:24–28. https://doi.org/10.1590/0103-9016-2015-0076
Zhang P, Jin T, Kumar Sahu S, Xu J, Shi Q, Liu H, Wang Y (2019) The distribution of tryptophan-dependent indole-3-acetic acid synthesis pathways in bacteria unravelled by large-scale genomic analysis. Molecules 24:1411. https://doi.org/10.3390/molecules24071411
Acknowledgements
We are grateful for the technical assistance provided by the Laboratory of Electron Microscopy “Elliot W. Kitajima”—ESALQ/USP.
Funding
This work was supported by grants from the “São Paulo Research Foundation” (FAPESP) (Proc. No. 2015/01188-9) and “National Council for Scientific and Technological Development” (CNPq) (Proc. No. 427669/2018-9). We thank CNPq for the fellowships to BDB, MND and ROH. We would like to thank FAPESP for the fellowship to EFF (Proc. No. 2017/11026-1). We also thank “Coordination for the Improvement of Higher Education Personnel” (CAPES) for the fellowship to JPRM and MLB.
Author information
Authors and Affiliations
Contributions
BDB and MCQ conceived the research plan. BDB performed the in-silico analysis for genome mining. BDB, ROH, FAP, MND carried out the experiments (in vitro and in vivo—greenhouse). JPRM analysed the anatomical changes in plant roots using microscopy. MLB and MLS performed the taxonomic analysis. BDB and MCQ wrote the manuscript with the support of EFF, JLA and MLB. All authors provided critical feedback and helped to shape the research, analysis and final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest in the publication.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Batista, B.D., Dourado, M.N., Figueredo, E.F. et al. The auxin-producing Bacillus thuringiensis RZ2MS9 promotes the growth and modifies the root architecture of tomato (Solanum lycopersicum cv. Micro-Tom). Arch Microbiol 203, 3869–3882 (2021). https://doi.org/10.1007/s00203-021-02361-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02361-z