Abstract
Plant growth promoting rhizobacteria (PGPR) are a group of bacteria that promote plants growth in the rhizosphere. PGPRs are involved in various mechanisms that reinforce plant development. In this study, we screened for PGPRs that were effective in early growth of Arabidopsis thaliana when added to the media and one Bacillus subtilis strain L1 (Bs L1) was selected for further study. When Bs L1 was placed near the roots, seedlings showed notably stronger growth than that in the control, particularly in biomass and root hair. Quantitative reverse transcription polymerase chain reaction analysis revealed a high level of expression of the high affinity nitrate transporter gene, NRT2.1 in A. thaliana treated with Bs L1. After considering how Bs L1 could promote plant growth, we focused on nitrate, which is essential to plant growth. The nitrate content was lower in A. thaliana treated with Bs L1. However, examination of the activity of nitrate reductase revealed higher activity in plants treated with PGPR than in the control. Bs L1 had pronounced effects in representative crops (wheat and lettuce). These results suggest that Bs L1 promotes the assimilation and use of nitrate and plant growth.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant growth promoting rhizobacteria (PGPR) include beneficial bacteria located in the rhizosphere that can enhance plant growth performance. The mechanisms responsible for PGPR activities include enhancement of abiotic stress tolerance, pathogen control, secretion of growth-related phytohormones, augmentation of plant nutrient uptake, and production of siderophores and other beneficial organic compounds (García-Fraile et al. 2015; Siddikee et al. 2015; Vejan et al. 2016).
Plant growth requires a variety of nutrients. Among them, nitrogen is one of the most important nutrients in plant growth and development and is the critical element in the biosynthesis of many fundamental molecules in plant, such as nucleic acids, chlorophyll, and ATP (O’Brien et al. 2016; Vejan et al. 2016). Since plants cannot directly fix atmospheric nitrogen. Instead they absorb nitrate or ammonium ions from the soil and convert them into the necessary metabolic materials (Miller et al. 2007). As nitrate can be artificially provided in fertilizer, PGPR application would enhance nitrate absorption, permitting plants to make full use of the nitrogen source. Plants have various genes encoding transport proteins, that allow the efficient absorption of nitrate. Of these genes, the NRT1 and NRT2 gene families are crucial (Cerezo et al. 2001). The NRT1.1 gene encodes a protein that has dual roles as a nitrogen transporter and a sensor of available nitrate (Tsay et al. 1993). The transport protein NRT2, plays a major role in nitrogen transport. The NRT2 family comprises high affinity, nitrate specific transporters in many plant systems, including Arabidopsis thaliana (L) Hevnh. (Krapp et al. 2014). The NRT2 family in A. thaliana consists of seven proteins (NRT2.1 to NRT2.7). NRT2.1 is key for the high affinity transport system (HATS) of root nitrate uptake under most conditions (Cerezo et al. 2001; Filleur et al. 2001). The stimulation of NRT2 expression of host plants by exogenous PGPR is involved in the PGPR growth promotion effect (Kechid et al. 2013). NRT2 genes have roles in the biotic defense system in plants. NRT2.1 is involved in resistance against Pseudomonas syringae pv. tomato (Camanes et al. 2012) and NRT2.6 in the resistance against Erwinia amylovora (Dechorgnat et al. 2012). Those studies indicate that the induction of NRT2 genes may be part of an induced systemic response (ISR) in recognition of microbial infection. Moreover, the induction of NRT2.1, NRT2.5, NRT2.6 accompanied by the morphological changes of the root architecture in the presence of PGPR could imply the potential improvement to nitrate absorption by plant roots (Kechid et al. 2013). Thus, PGPR that also trigger the induction of NRT2 genes in the host plant may correlate with either the PGPR mechanism of inducing ISR in plants, improving plant nutrient uptake, or both.
In this study, an evaluation of several soil bacteria led to a focus on the Bacillus subtilis strain L1 (Bs L1). B. subtilis is involved in various reactions and contributes to the availability of nitrogen and phosphorus in the soil (Idriss and Borriss 2002). In addition, the bacterium contributes to the improvement of stress resistance of plants (Pliego and Ramos 2010). Our co-cultivation experiment results clearly indicated that Bs L1 enhanced the growth performance of the tested A. thaliana, Lactuca sativa, and Triticum aestivum plants. In addition, morphological improvements in co-cultivated L. sativa and wild type, transgenic Arabidopsis seedlings were closely observed and measured. Quantitative real time polymerase chain reaction (qRT-PCR) analysis determined the molecular mechanism underlying the PGPR growth promoting effect in the co-cultivated Arabidopsis plants. The results demonstrate the plant growth promoting effect of Bs L1.
Materials and methods
Plant materials and growth conditions
A. thaliana (Col-0 ecotype) seeds were sterilized with sodium chlorate and rinsed four times with sterilized water. The seeds were kept at 4 °C for 3 days and planted perpendicularly in Murashige and Skoog (MS) agar containing 2% sucrose. Then, they were incubated in a culture chamber at 23 °C with a 16 h light/8 h dark cycle and a relative humidity of 60%. After 4 days, the seedlings were moved to the test board. L. sativa and T. aestivum seeds were sterilized for 15 min with sodium chlorite and rinsed five times with sterilized water. After 3 days of stratification at 4 °C, the seeds were cultivated in MS agar containing 2% sucrose in a culture chamber at 23 °C with the aforementioned light/dark cycle and relative humidity. After 6 days, the seedlings were moved to the soil for joint cultivation (Lee et al. 2017). Col-0 ecotype was obtained from TAIR site (arabodopsis.org) and Lactuca sativa L was obtained from Danong market (www.danong.co.kr). The cultivar of Lactuca sativa we purchased is Lactuca sativa L. var. Cheongchima. KeumKang (K2) indicates Korean wheat cultivar of T.aestivum. Seeds of CycB1pro::GUS (Colón-Carmona et al. 1999) and DR5::GUS (Ulmasov et al. 1997) transgenic Arabidopsis plants were provided by Dr. Peter Doerner and Dr. Tom J. Guilfoyle, respectively.
Bacterial isolation, identification, and inoculation
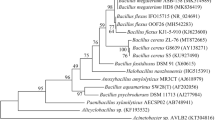
Various types of bacteria were isolated from soil samples collected from local farms (Namyangju, Republic of Korea). First, lettuce plants growing at the farm were rooted and transported to the laboratory, and the large lumps of soil attached to the roots were removed. Next, the roots were gently shaken in sterilized distilled water to collect the remaining soil. Bacteria were then isolated from soil samples using the plate streaking method (Sander 2012; Trinh et al. 2018). In this study, lysogeny broth (LB, Miller) of Sigma-Aldrich (L5322) was used and 1 g of soil was put into a 15 mL conical tube containing 10 mL of sterilized distilled water. After 5 min of vortex, 100 μL suspension was transferred to microtubes containing 900 μL of LB to obtain 10−1 dilution. The culture was further diluted to 10−2, 10−3, 10−4 before incubation on the nutrient agar at 37 °C for 24 h. Among these bacteria, Bs L1 (Genbank; Accession number: MN542211.1) was identified by 16S rDNA sequencing (Ha 2017; Lei et al. 2018). The bacterial cell stock was stored at − 80 °C in a 40% glycerol solution. The day before co-cultivation with A. thaliana, Bs L1 was grown in the shaker chamber at 37 °C for 18 h in LB media. For the co-cultivation experiment, 200 µL of bacterial suspension (109 colony forming units (CFU) mL−1) was applied in a line 5 cm under the seedling roots.
Co-cultivation condition
Four-day old, uniform A. thaliana seedlings were moved to MS agar and a 200 µL bacterial suspension (109 CFU mL−1) in 2% sucrose was added 5 cm under the roots (Wintermans et al. 2016). The intermediate plate treated with 200 µL of bacteria-free LB was used as the control. Plants were grown vertically with Bs L1 for 8 days in the growth chamber at 23 °C using the aforementioned light/dark cycle and relative humidity. For β-glucuronidase (GUS) staining, plants were cultivated for 4 days. The seedlings were collected and stored at − 80 °C. To create a nitrogen (N) deficient state, N-free Hoagland agar (Hoagland Nitrogen free, 30630038; bioPLUS) containing 2% sucrose was used as a test plate. For L. sativa and T. aestivum, 4-day old seedlings of the same size were moved to a container with soil mixed with 70 mL distilled water. One mL volume of LB suspension of Bs L1 (103 or 105 CFU mL−1) was used to directly inoculate the plant roots. Soil containers in which 1 mL of bacteria-free LB was applied to plant roots were used as controls. L. sativa and Bs L1 were co-cultivated in growth chambers at 23 °C using the aforementioned light/dark cycle and relative humidity. Growth performance was observed from 0 to 9 days after inoculation.
GUS activity assay
GUS activity was determined by tissue chemical dyeing using GUS solution. Four day old DR5:GUS (Ulmasov et al. 1997) and CycB1pro:GUS transgenic seedlings (Colón-Carmona et al. 1999) were co-cultivated with Bs L1 under normal conditions. After PGPR treatment, seedlings were collected and dyed with GUS solution in a 37 °C incubator for 2 h and then at 70 °C in 70% ethanol until all chlorophyll was removed (Li 2011). Ecotype of the DR5-GUS and CycB1pro:GUS transgenic seedlings is Col-0 (Table 1).
qRT PCR analysis
RNA extracted from a 50 mg sample by TRIzol Reagent was converted to cDNA using the RevertAid First Strand cDNA Synthetic Kit (H1622; Thermo Fisher Scientific) as described previously (Jeong et al. 2018; Joo et al. 2017; Nguyen et al. 2016). qRT-PCR was performed in a heat cycler (Bio-Rad Laboratories). The transcript of Actin2 gene was used as an internal standard. Three independent replications were performed for each sample. Error bar = standard deviation. The statistical analyses were performed by the one-way ANOVA, followed by Tukey’s test for comparison of means at the 95% confidence level. Different letters indicate significant differences at P < 0.05 according to one-way ANOVA and Tukey’s test.
Nitrate content and nitrate reductase (NR) activity
Total nitrate content was quantified using the salicylic acid method (Cataldo et al. 1975; Xu et al. 2016; Vendrell and Zupancic 1990). A. thaliana seedlings of 50 mg were boiled for 20 min at 100 °C in 1 mL distilled water. The boiled sample was centrifuged at 15,871g for 10 min and the 0.1 mL of supernatant was transferred to a new 15 mL tube. For the assay, 0.4 mL of salicylic-sulfuric acid solution (5 g of salicylic acid in 100 mL of sulfuric acid) was added to the mixture for 20 min at room temperature. Next, an 8.5 mL solution of 8% NaOH was added for 5 min at room temperature and the absorption was measured using a multi detection micro plate reader at OD410. Nitrate content is calculated using the following formula: nitrate content = C × V/W (C, nitrate concentration calculated using the regression equation in the OD410; V, volume of the extracted sample; and W, sample weight). In an empty sample, 0.1 mL distilled water was used instead of 0.1 mL of the supernatant sample. A standard curve was created using KNO3 at a concentration of 0.5–2 mg L−1, and the regression equation (R2 = 0.99) was calculated. Nitrate reducing enzyme activity is measured primarily by the previously described method, (Smarrelli and Campbell, 1983) and about 0.1 g of fresh samples were frozen by liquid nitrogen and ground into powder. A 1 mL volume of Buffer A (25 mM phosphate buffer, pH 7.5, 10 mM KNO3, 0.05 mM EDTA) was added to the sample which was then centrifuged for 10 min at 4 °C and 28,486 g. The 50 µL supernatant was transferred to a new 1.5 mL tube and added to 900 µL of buffer A, 2 mM β NADH solution, 58 mM sulfanilamide 3 M HCl solution, 0.77 mM N-(1-Naphthyl) ethylenediamine dihydrochloride (NED) solution. For standard curves, we used 1.45 mM sodium nitrate instead of sample supernatant at 25 °C for 30 min. The OD540 value was measured. A = C × V1/V2/W/t (A, nitrate reducing enzyme activity; C, the amount of nitrite derived from regression equations; V1, volume of extracts added to the reaction; V2, volume added to the reaction; W, sample weight; t, reaction time). The absorption intensity was measured using the Multi Detection Microplate Reader (HIDEX).
Luminescence imaging
As previously described (Yu and Lee 2017), the C24 line expressing a chimeric gene consisting of the luciferase reporter gene fused to the RD29A promoter was used. The seeds were germinated and grown for 4 days in MS agar containing 2% sucrose. Ten similar seedlings were selected and transferred to the PGPR plate for 8 days. The samples were then lightly sprayed with luciferin and kept in the dark for 5 min. Images of the samples were taken for 10 min using a model 7513-002 luminescence imaging device (Roper Scientific) and Lumazone 1300B CCD camera (Roper Scientific). Liquid nitrogen was used to maintain the low temperature of the device before use.
Statistical analyses
The statistical analyses were performed by the one-way ANOVA, followed by Tukey’s test for comparison of means at the 95% confidence level. Different letters indicate significant differences at P < 0.05 according to one-way ANOVA and Tukey’s test.
Results and discussion
Bs L1 promotes growth of lettuce
Co-cultivation experiments were conducted using soil microbes with Col-0 accession of Arabidopsis. Rhizobacteria with a noticeable growth enhancement effect on the host plants were selected for further investigation. We previously isolated Pseudomonas nitroreducens strain IHB B 13561 (PnIHB) as PGPR by using A. thaliana (Trinh et al. 2018). The growth promotion effect observed for Arabidopsis grown in MS medium was not always observed for vegetables, such as lettuce. Thus, as shown in Fig. 1, we screened soilborne microorganisms and isolated Bs L1, which promoted the growth of lettuce in sterile soil instead of MS medium using the method we previously established (Trinh et al. 2018). Blast analysis showed that partial sequences of 16S rDNA of Bs L1 had more than 99% similarity with 16S rDNA sequences of Bs Y16 (Khan et al. 2017). The authors isolated rhizobacteria with ACC-deaminase activity in the wheat rhizosphere and tested their plant growth promotion activities. However, there were few physiological studies on how Bs Y16 promotes plant growth. Therefore, in this study we decided to study the mechanism by which Bs L1 promotes plant growth.
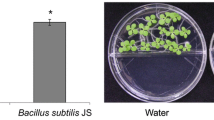
Growth stimulation of Pn IHB and Bs L1 in L. sativa. a Germinating lettuce seeds were stratified for 4 days in MS agar containing 2% sucrose. During germination of lettuce seedlings, the seedlings are transferred to containers containing sterilized soil treated with 70 mL of water. Immediately after that, the soil was treated with 1 mL of LB containing Pn IHB and Bs L1 at different concentrations (103 or 105 CFU mL−1) and the lettuce phenotypes were observed. b On day 8 after the transfer, weight of lettuce shoots and roots was determined. Different letters indicate significant differences at P < 0.05 according to one-way ANOVA and Tukey’s test
To observe how plant growth promotion differed between Pn IHB and Bs L1, we used Pn IHB, Bs L1, or both to inoculate the sterile soil medium of 5-day old lettuce. As shown in Fig. 1, both Pn IHB and Bs L1 started to show distinct growth promotion effects 6 d after inoculation. We hypothesized that plant growth would be accelerated if Pn IHB and Bs L1 were co-inoculated. However, that was not the case (Fig. 1a). When root and shoot weights were determined, it was found that growth was similarly promoted by Pn IHB and Bs L1. This indicated that these PGPRs contributed to the increase of biomass throughout the plant. This seemed to lead directly to the increase in the fresh weight of A. thaliana (Fig. 1b). In addition to these plants, a similar growth promotion effect of Bs L1 was evident for T. aestivum (Fig. S2). Thus, either these two PGPRs work through similar pathways to promote growth or growth is already maximized by a sufficient amount of PGPR bacteria, resulting in no further growth promotion.
Bs L1 increases the number of root hairs and lateral roots of Arabidopsis
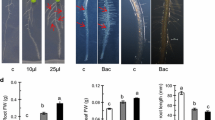
After confirming that Bs L1 had a growth promotion effect in lettuce, the mechanism of growth promotion was investigated using A. thaliana. In addition to the Bs L1 induced growth of the shoots, promotion of root growth was also observed as the increase in the number of lateral roots and density of root hairs (Fig. 2a). This seemed to lead directly to the increase in the fresh weight of Arabidopsis (Fig. 2b). To evaluate whether Bs L1 can promote plant root cell division, we used transgenic plants containing the GUS reporter gene fused with the promoter of CYCB1 encoding a cell cycle marker protein in the root meristem (Day and Reddy 1998). Although slight, the GUS staining was darker in the Bs L1 treated group compared to the controls (Fig. 3a). In addition, optical microscopy revealed considerably more root hair development in Bs L1 treated A. thaliana (Fig. 3b).
Phenotypic improvements in Arabidopsis co-cultivated with Bs L1. Col-0 seedlings germinated in MS agar containing 2% sucrose were transferred to PGPR plate (200 µL of bacteria suspension (109 CFU mL−1) in MS agar containing 2% sucrose). a Physical changes for Col-0 seedlings co-cultivated with Bs L1 were observed for 16 days. b Comparison of fresh weights
Histochemical GUS assay and optical microscopy observation of root ends. The 4 days CycB1pro:GUS seedlings grown in MS agar containing 2% sucrose were transferred to PGPR plates containing MS agar with 2% sucrose and 200 µL of bacteria suspension (109 CFU/mL). aCycB1pro:GUS analysis of GUS seedlings. No significant difference was evident after 4 days of co-cultivation. b Optical microscopy examination of root tip (× 100) revealed greater development of root hair
Next, to determine if auxin is involved in the growth difference caused by the PGPR, DR5:GUS plants were used. These plants have enhanced GUS activity in response to auxin (Béziat et al. 2017). The root tips of plants treated with Bs L1 showed more GUS activity than the control plants (Fig. 4a). However, there was no difference in the GUS activity in other tissues such as the leaves or stems (Fig. 4a). The characteristics of PGPR that promote plant growth include the direct production of plant hormones and the control of homeostasis in plants (Lugtenberg and Kamilova 2009). The effects of various bacterial strains that produce plant hormones have been reported. In the case of rhizobium, approximately 80% of the bacteria can produce indole acetic acid (Ahmed and Hasnain 2014; Duca et al. 2014; Swain et al. 2007). Therefore, Bs L1 may also synthesize auxin for plant growth. To determine whether Bs L1 induces a more active auxin signaling pathway, we evaluated the mRNA expression level of the auxin signaling genes ARF19, ARF7, and AXR4 using qRT-PCR. No significant difference was observed between the expression levels of the control and the Bs L1 treated plants (Fig. 4b). In addition, only a small difference was evident in the expression of ammonia-related genes including ammonium transporter 1, 1.2, 2 (AMT1, AMT1.1, AMT2). Since ARF7 and ARF19 were not enhanced in their transcript levels in response to the Bs L1, it seems that increased lateral root growth is not associated with ARF7 and ARF19 dependent auxin signaling in lateral root formation, as reported elsewhere (Okushima et al. 2007).
Histochemical GUS assay and relative gene expression level analysis. DR5pro:GUS seedlings grown for 4 days in MS agar containing 2% sucrose were transferred to PGPR plates with MS agar containing 2% sucrose and 200 µL bacteria suspension (109 CFU mL−1). a There was no difference between control and DR5pro:GUS, but there was a slight difference at the end of the roots. b Relative expression analysis of auxin and ammonia-related genes in inoculation and non-inoculated seedlings. Different letters indicate significant differences at P < 0.05 according to one-way ANOVA and Tukey’s test
Bs L1 has a high growth promotion effect, especially in a nitrogen-rich medium
Nitrogen is a very important factor in the growth of plants. Thus, we measured the expression of nitrogen transport protein genes. The NRT1 and NRT2 proteins have important roles in nitrogen transport. Bs L1 had little effect on the expression of NRT1.1, but it increased expression of NRT2 genes including NRT2.1, 2.2, 2.5 and 2.6 (Fig. 5a). Arabidopsis NRT1.1, also known as CHL1 and NPF6.3, is one of the most studied nitrogen transport genes (Tsay et al. 1993). Since the discovery of NRT1.1 in 1993, numerous studies have revealed the principles of nitrogen absorption and transport mechanisms in plants (Wang et al. 2012). One study showed that NRT1.1 is involved in nitrogen transport activities at both high concentration and low concentration nitrate conditions (Liu et al. 1999). Absorption of nitrate (NO3−) by root cells in soil solutions is a very important response in most plant species. Thus, both the high concentration and low concentration transport systems (HATS and LATS) enable efficient nitrogen binding to various external nitrate concentrations, and are crucial for plant growth (Miller et al. 2007). The NRT1.1, NRT2.1 and NRT2.2 transport proteins have major roles in nitrate transport and are present in Arabidopsis. Most HATS activities seem to be carried out by NRT2.1 (Miller et al. 2007). Indeed, a NRT2.1 null mutant was shown to lose a significant portion (up to 75%) of the HATS activity (Cerezo et al. 2001; Filleur et al. 2001; Li et al. 2007).
Relative gene expression level analysis of NRT family, nitrate contents, and evaluation of nitrate reductase activity. Seedlings grown for 4 days in MS agar containing 2% sucrose were transferred to PGPR plates with MS agar containing 2% sucrose and 200 µL bacteria suspension (109 CFU mL−1) a Relative expression analysis of NRT genes in Bs L1 treated seedlings and untreated seedlings. We collected the seedlings treated with Bs L1 8 days after transfer. We collected seedlings at the same weight of 100 mg for the nitrate content and nitrate reductase assays. b The amount of nitrate was less than that of the treated and non-treated samples. c Nitrate reductase activity was higher in the treated samples. d Relative expression analysis of genes encoding nitrate reductase (NIA1, NIA2) in Bs L1 treated seedlings and untreated seedlings. Different letters indicate significant differences at P < 0.05 according to one-way ANOVA and Tukey’s test
Since the expression of NRT2, encoding the main protein of nitrate transport, has increased significantly in Bs L1 treated plants, we examined the nitrate contents of the control as well as the Bs L1 treated plants. The nitrate content was slightly lower in the plants treated with Bs L1 (Fig. 5b). To determine if the results were due to the more efficient metabolism of nitrate in Bs L1 treated plants, we then investigated the activity of NR in the plants. As shown in Fig. 5c, the activity of NR in the Bs L1 treated plants was approximately four times higher than that of the control. Although an increase in nitrate reducing enzyme activity in response to Bs L1 was observed (Fig. 5c), the level of expression of genes encoding nitrate reductase (NIA1, NIA2) is decreased (Fig. 5d). Nitrate-reducing enzyme activity is very important for plant adaptation to challenging environments, and the NR activity is strictly controlled. Therefore, since Bs L1 displayed increased NR activity, its transcript level could be obviated by negative regulation. Given the high activity of NR in the Bs L1 treated plants, we hypothesized that these plants might grow better under low nitrate conditions. To test this, we examined the growth promotion effect of Bs L1 under low nitrate conditions (Fig. 6a). Even under a very low concentration of nitrate (0.5 mM), Bs L1 clearly promoted plant growth. Furthermore, growth was doubled at a nitrate concentration of 10 mM (Fig. 6a). We also determined the relative expression levels of nitrate-responsive genes at different nitrate concentrations. In the case of NIR1, encoding a nitrite reductase (Konishi and Yanagisawa 2010) even if the nitrate concentration increased, there was no difference regardless of the treatment (Fig. 6e). NRT2.1 displayed a very high relative expression in the Bs L1 treatment group, as the nitrate concentration increased while the NRT1.1 and NLP6, encoding transcriptional regulator (Konishi and Yanagisawa 2013) exhibited similar expressions (Fig. 6). Phyllobacterium brassicacearum STM196 reduces the negative effects of high nitrate on lateral root development (Mantelin et al. 2006). The authors found that STM196 strongly upregulated the nitrate transporter genes, NRT2.5 and NRT2.6, which altered the shoot:root biomass allocation for optimal root development in the conditions used. Based on the present results and the previous data, although STM196 and Bs L1 similarly promote the growth of plant roots, their molecular mechanisms of action differ.
Relative expression level analysis of nitrate signaling-related genes at different nitrate concentrations. The seedlings germinated for 4 days in MS agar containing 2% sucrose were transferred to PGPR plates with different KNO3 concentrations and raised for 8 days. The growth of plants increased as the concentration of KNO3 increased (0, 0.05, 0.5, 10 mM), and the effect of Bs L1 increased. a The response level was higher than the Bs L1 treatment. e The expression of NIR1 decreased. b and d The expressions of NRT1.1 and NLP6 were similar. Different letters indicate significant differences at P < 0.05 according to one-way ANOVA and Tukey’s test
PGPRs ultimately have a positive effect on plant growth by reducing plant sensitivity to abiotic stress (Yang et al. 2009). Thus, we finally investigated whether plants that grew well with Bs L1 treatment were better able to withstand environmental stresses. Bs L1 was effective in Col-0 and other wild type plants, such as C24. As in Col-0, the root length was shorter than the untreated plant, but weighed more (Fig S1). However, Bs L1 did not appear to significantly improve plant resistance to environmental stresses (data not shown). Nevertheless, a study of plants with a construct of a luciferase reporter fused to the stress responsive promoter RD29A found that activity of the abiotic stress responsive promoter was significantly lowered in normal conditions by Bs L1 treatment (Fig. 7). This indicates that plants may be exposed to some degree of environmental stress even in normal conditions. By contrast, the observations with Bs L1 support the assumption that nitrate metabolism of plants is facilitated, which promotes plant growth and production of various metabolites, leading to better performance of plant growth in normal conditions.
Luciferase imaging of RD29A gene activity and MPK gene expression level after Bs L1 treatment. Seedlings germinated from MS agar containing 2% sucrose for 8 days were transferred to PGPR plates and raised for 8 days. a The results with the stress-reaction promoter RD29A fusion with a luciferase reporter indicated less activity for the Bs L1 treated samples. The treatment with 200 mM NaCl was done the day prior to imaging. PnIHB and Bs L1 were treated at the same concentration (109 CFU mL−1). b The relative expression of RD29A was significantly different for Bs L1 treated samples than for the Bs L1 untreated samples. Different letters indicate significant differences at P < 0.05 according to one-way ANOVA and Tukey’s test
In conclusion, Bs L1 promotes plant growth through activation of the nitrogen signaling pathway of plants. Further research is needed to clarify the plant growth promoting mechanisms of Bs L1. Nitrogen is an indispensable material for the growth of plants. Unfortunately, crops are often cultivated in soil that lacks an adequate amount of nitrogen. As a result, a large quantity of nitrogen fertilizer is used every year and, due to the low nitrogen use efficiency in crops, a considerable amount of these fertilizers leaks into the environment, which causes serious environmental harm (Diaz and Rosenberg 2008; Robertson and Vitousek 2009). Therefore, the application of PGPR, which can increase the activity of nitrogen metabolizing enzymes in plants, will lead to the reduction of nitrogen fertilizer use in agriculture and help to address fertilizer-related environmental pollution.
References
Ahmed A, Hasnain S (2014) Auxins as one of the factors of plant growth improvement by plant growth promoting rhizobacteria. Pol J Microbiol 63:261–266
Béziat C, Kleine-Vehn J, Feraru E (2017) Histochemical staining of β-glucuronidase and its spatial quantification. Methods Mol Biol 1497:73–80
Camanes G, Pastor V, Cerezo M, Garcıa-Andrade J, Vicedo B, Garcıa-Agustın P, Flors V (2012) A deletion in NRT2.1 attenuates Pseudomonas syringae-induced hormonal perturbation, resulting in primed plant defenses. Plant Physiol 158:1054–1066
Cataldo DA, Haroon M, Schrader LE, Youngs VL (1975) Rapid colorimetric determination of nitrate in plant-tissue by nitration a salicylic-acid. Commun Soil Sci Plant 6:71–80
Cerezo M, Tillard P, Filleur S, Munos S, Daniel-Vedele F, Gojon A (2001) Major alterations of the regulation of root NO3− uptake are associated with the mutation of NRT2.1 and NRT2.2 genes in Arabidopsis. Plant Physiol 127:262–271
Colón-Carmona A, You R, Haimovitch-Gal T, Doerner P (1999) Technical advance: spatio-temporal analysis of mitotic activity with a labile cyclin–GUS fusion protein. Plant J 20:503–508
Day IS, Reddy AS (1998) Isolation and characterization of two cyclin-like cDNAs from Arabidopsis. Plant Mol Biol 36:451–461
Dechorgnat J, Patrit O, Krapp A, Fagard M, Daniel-Vedele F (2012) Characterization of the NRT2.6 gene in Arabidopsis thaliana: a link with plant response to biotic and abiotic stress. PLoS One 7:e42491
Diaz RJ, Rosenberg R (2008) Spreading dead zones and consequences for marine ecosystems. Science 321:926–929
Duca D, Lorv J, Patten CL, Rose D, Glick BR (2014) Indole-3-acetic acid in plant-microbe interactions. Antonie Van Leeuwenhoek 106:85–125
Filleur S, Dorbe MF, Cerezo M, Orsel M, Granier F, Gojon A, Daniel-Vedele F (2001) An Arabidopsis T-DNA mutant affected in NRT2 genes is impaired in nitrate uptake. FEBS Lett 489:220–224
García-Fraile P, Menéndez E, Rivas R (2015) Role of bacterial biofertilizers in agriculture and forestry. AIMS Bioeng 2:183–205
Ha DD (2017) Degradation of chlorotoluenes by Comamonas testosterone KT5. Appl Biol Chem 60:457–465
Idriss EE, Borriss R (2002) Extracellular phytase activity of Bacillus amyloliquefaciens FZB45 contributes to its plant-growth-promoting effect. Microbiology 148:2097–2109
Jeong CY, Lee WJ, Truong HA, Trinh CS, Hong SW, Lee H (2018) AtMybL-O modulates abscisic acid biosynthesis to optimize plant growth and ABA signaling in response to drought stress. Appl Biol Chem 61:473–477
Joo J, Oh N, Nguyen HN, Lee YH, Kim Y, Song IS, Cheong J (2017) Intergenic transformation of AtMYB44 confers drought stress tolerance in rice seedlings. Appl Biol Chem 60:447–455
Kechid M, Desbrosses G, Rokhsi W, Varoquaux F, Djekoun A, Touraine B (2013) The NRT2.5 and NRT2.6 genes are involved in growth promotion of Arabidopsis by the plant growth-promoting rhizobacterium (PGPR) strain Phyllobacterium brassicacearum STM196. New Phytol 198:514–524
Khan MY, Zahir ZA, Asghar HN, Waraich EA (2017) Preliminary investigations on selection of synergistic halotolerant plant growth promoting rhizobacteria for inducing salinity tolerance in wheat. Pak J Bot 49:1541–1551
Konishi M, Yanagisawa S (2010) Identification of a nitrate-responsive cis-element in the Arabidopsis NIR1 promoter defines the presence of multiple cis-regulatory elements for nitrogen response. Plant J 63:269–282
Konishi M, Yanagisawa S (2013) Arabidopsis NIN-like transcription factors have a central role in nitrate signaling. Nat Commun 4:1617
Krapp A, David LC, Chardin C, Girin T, Marmagne A, Leprince A-S, Chaillou S, Ferrario-Méry S, Meyer C, Daniel-Vedele F (2014) Nitrate transport and signaling in Arabidopsis. J Exp Bot 65:789–798
Lee WJ, Kim J, Lee D, Hong SW, Lee H (2017) Arabidopsis UDP-glycosyltransferase 78D1-overexpressing plants accumulate higher levels of kaempferol 3-O-β-d-glucopyranoside than wild-type plants. Appl Biol Chem 60:647–652
Lei Y, Zhao P, Li C, Zhao H, Shan Z, Wu Q (2018) Isolation, identification and characterization of a novel elastase from Chryseobacterium indologenes. Appl Biol Chem 61:365–372
Li X (2011) Histostaining for tissue expression pattern of promoter-driven GUS activity in arabidopsis. Bio-protocol Bio 101:e93
Li W, Wang Y, Okamoto M, Crawford NM, Siddiqi MY, Glass AD (2007) Dissection of the AtNRT2.1: AtNRT2.2 inducible high affinity nitrate transporter gene cluster. Plant Physiol 143:425–433
Liu KH, Huang CY, Tsay YF (1999) CHL1 is a dual-affinity nitrate transporter of Arabidopsis involved in multiple phases of nitrate uptake. Plant Cell 11:865–874
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting rhizobacteria. Annu Rev Microbiol 63:541–556
Mantelin S, Desbrosses G, Larcher M, Tranbarger TJ, Cleyet-Marel JC, Touraine B (2006) Nitrate-dependent control of root architecture and N nutrition are altered by a plant growth-promoting Phyllobacterium sp. Planta 223:591–603
Miller AJ, Fan X, Orsel M, Smith SJ, Wells DM (2007) Nitrate transport and signalling. J Exp Bot 58:2297–2306
Nguyen HD, Jeong CY, Lee WJ, Lee H (2016) Identification of a novel Arabidopsis mutant showing sensitivity to histone deacetylase inhibitors. Appl Biol Chem 59:855–860
O’Brien JA, Vega A, Bouguyon E, Krouk G, Gojon A, Coruzzi G, Gutierrez RA (2016) Nitrate transport, sensing, and response in plants. Mol Plant 9:837–856
Okushima Y, Fukaki H, Onoda M, Theologis A, Tasaka M (2007) ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 19:118–130
Pliego C, Ramos C (2010) Screening for candidate bacterial biocontrol agents against soilborne fungal plant pathogens. Plant Soil 430:505–520
Robertson GP, Vitousek PM (2009) Nitrogen in agriculture: balancing the cost of an essential resource. Annu Rev Environ Resour 34:97–125
Sander ER (2012) Aseptic laboratory techniques: plating methods. J Vis Exp 63:3064
Siddikee MA, Sundaram S, Chandrasekaran M, Kim K, Selvakumar G, Sa T (2015) Halotolerant bacteria with ACC deaminase activity alleviate salt stress effect in canola seed germination. J Korean Soc Appl Biol Chem 58:237–241
Smarrelli J Jr, Campbell WH (1983) Heavy metal inactivation and chelator stimulation of higher plant nitrate reductase. Biochim Biophy Acta 742:435–445
Swain MR, Naskar SK, Ray RC (2007) Indole-3-acetic acid production and effect on sprouting of yam (Dioscorea rotundata L.) minisetts by Bacillus subtilis isolated from culturable cowdung microflora. Pol J Microbiol 56:103–110
Trinh CS, Lee H, Lee WJ, Lee SJ, Chung N, Han J, Kim J, Hong SW, Lee H (2018) Evaluation of the plant growth-promoting activity of Pseudomonas nitroreducens in Arabidopsis thaliana and Lactuca sativa. Plant Cell Rep 37:873–885
Tsay YF, Schroeder JI, Feldmann KA, Crawford NM (1993) The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell 72:705–713
Ulmasov T, Murfett J, Hagen G, Guilfoyle TJ (1997) Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 9:1963–1971
Vejan P, Abdullah R, Khadiran T, Ismail S, Boyce AN (2016) Role of plant growth promoting rhizobacteria in agricultural sustainability—a review. Molecules 21:573
Vendrell PF, Zupancic J (1990) Determination of soil nitrate by transnitration of salicylic acid. Commun Soil Sci Plant Anal 21:1705–1713
Wang YY, Hsu PK, Tsay YF (2012) Uptake, allocation and signaling of nitrate. Trends Plant Sci 17:624–667
Wintermans PCA, Bakker PAHM, Pieterse CMJ (2016) Natural genetic variation in Arabidopsis for responsiveness to plant growth-promoting rhizobacteria. Plant Mol Biol 90:623–634
Xu N, Wang R, Zhao L, Zhang C, Li Z, Lei Z, Liu F, Guan P, Chu Z, Crawford NM, Wanga Y (2016) The Arabidopsis NRG2 protein mediates nitrate signaling and interacts with and regulates key nitrate regulators. Plant Cell 28:485–504
Yang J, Kloepper JW, Ryu CM (2009) Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant Sci 14:1–4
Yu SI, Lee BH (2017) Generation of a stress-inducible luminescent Arabidopsis and its use in genetic screening for stress-responsive gene deregulation mutants. Meth Mol Biol 1631:109–119
Acknowledgements
This work was funded by the National Institute of Crop Science, Rural Development Administration, Republic of Korea [grant number PJ012496022018] and the National Research Foundation of Korea (to Hojoung Lee, 2019; grant NRF-2019R1A2C1088417).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, S., Trịnh, C.S., Lee, W.J. et al. Bacillus subtilis strain L1 promotes nitrate reductase activity in Arabidopsis and elicits enhanced growth performance in Arabidopsis, lettuce, and wheat. J Plant Res 133, 231–244 (2020). https://doi.org/10.1007/s10265-019-01160-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-019-01160-4