Abstract
Bacillus amyloliquefaciens SQR9, isolated from the rhizosphere of cucumber, can control Fusarium wilt of cucumber and directly stimulate plant growth. To evaluate its potential agricultural use, the plant growth promotion of B. amyloliquefaciens SQR9 was evaluated, and the relative mechanisms, especially the production of the phytohormone indole-3-acetic acid (IAA), were investigated. The related plant-growth-promoting factors were genetically and chemically analyzed, and a mutant library was constructed for selecting strains with different IAA production. B. amyloliquefaciens SQR9 showed a growth-promoting activity in greenhouse experiments. Plant-growth-promoting factors like extracellular phytase, volatile components including acetoin, 2,3-butanediol, and phytohormone IAA were detected in B. amyloliquefaciens SQR9 cultures grown under laboratory conditions. Three IAA production mutant strains showed variation in plant-growth-promoting effect. IAA production in B. amyloliquefaciens SQR9 was related to its plant-growth-promoting effect, but IAA alone could not account for the overall observed plant-growth-promoting effect. The promoted plant growth by the rhizospheric strain B. amyloliquefaciens SQR9 can be attributed to multiple factors, including production of phytohormones, volatile compounds, and extracellular enzymes. Therefore, the strain B. amyloliquefaciens SQR9 may be used as a plant-growth-promoting agent to increase crop yield.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant-growth-promoting rhizobacteria (PGPR) are a wide range of microorganisms, colonizing the root surface and enhancing seed emergence, plant biomass, and crop yield (Kloepper et al. 1989; Frommel et al. 1991; Babalola 2010). PGPR can directly stimulate plant growth through various mechanisms, including production of phytohormones (Bottini et al. 2004; Chaiharn and Lumyong 2011; Kochar et al. 2011), increasing the availability of plant nutrients in the rhizosphere (Idriss et al. 2002; Adesemoye et al. 2008; Palacios et al. 2014), suppressing ethylene production by root cells (Penrose and Glick 2001; Madhaiyan et al. 2006), and emitting plant-growth-promoting volatile compounds (Ryu et al. 2003). Today, most of the commercial PGPR are Bacillus strains because they form heat- and desiccation-resistant endospores, which facilitate their survival during production and storage period (Emmert and Handelsman 1999). The use of Bacillus strains as a bioorganic fertilizer also depends on the production of various antibiotics (Liu et al. 2013; Wang et al. 2013). It is very likely that plant-growth-promoting effect of rhizosphere bacilli results from the complex combined action of several factors mentioned above, though little is known about the basic molecular mechanisms responsible for this beneficial action.
Low-molecular-weight C6-volatile compounds such as methyl jasmonate, jasmonate, and terpenes, have been identified as signal molecules for plant growth (Farmer 2001; Farag and Paré 2002). Airborne chemicals released by specific PGPR can regulate auxin homeostasis are implicated in cell expansion, trigger growth promotion, and induce systemic resistance (ISR) in Arabidopsis thaliana seedlings (Ping and Boland 2004; Ryu et al. 2004; Farag et al. 2006; Zhang et al. 2007). Studies on Bacillus subtilis GB03 and Bacillus amyloliquefaciens IN937a revealed that 3-hydroxy-2-butanone (acetoin) and 2,3-butanediol could be the volatile chemical signals enhancing the growth of A. thaliana (Ryu et al. 2003, 2004).
Improving P nutrition is achievable by releasing P from insoluble inorganic form such as polyphosphates and/or phytate (myo-inositol hexakiphosphate), which accounts for 20–50 % of the total organic P in soil (Richardson et al. 2001). The possible role of microbial phytase produced by PGPR in supporting plant growth has been investigated in B. amyloliquefaciens FZB45, which can promote maize growth under P limiting condition (Idriss et al. 2002).
Auxin (indole-3-acetic acid (IAA)), which can promote plant growth (Idris et al. 2007; Spaepen et al. 2007), was the first hormone identified in plants. This phytohormone can change cell wall in plants, increase cell permeability with swelling of exposed plant cells, and induce the production of another hormone, ethylene (Woodward and Bartel 2005; Teale et al. 2006). IAA can be produced by 80 % of rhizosphere bacteria (Idris et al. 2004). Studies using IAA-deficient bacterial mutants showed that microbial IAA production increases lateral root length and number as well as root hair density (Barbieri and Galli 1993). By increasing the physical size of the root system, plant mineral uptake, and root exudation, it also increases bacterial colonization on plant roots. B. amyloliquefaciens FZB42 IAA mutant strains were less efficient in promoting plant growth, indicating that IAA and plant growth promotion are functionally related in the bacterial strain (Idris et al. 2007). However, the relative importance of IAA in the plant-growth-promoting effect of Bacillus is not well understood.
B. amyloliquefaciens SQR9 isolated from cucumber rhizosphere showed an efficient root colonization (Cao et al. 2011; Qiu et al. 2012). The antifungal antibiotic, bacillomycin D, is involved in the biofilm formation and root colonization by B. amyloliquefaciens SQR9 (Xu et al. 2013), whereas the global transcription regulator AbrB inhibited its root colonization (Weng et al. 2013). B. amyloliquefaciens SQR9 was an IAA production strain, and we hypothesized that IAA produced by SQR9 was important for its plant-growth-promoting activity. To determine the effects of IAA production by B. amyloliquefaciens SQR9 on plant growth promotion, three IAA mutant strains were selected from a random insertion mutant library, and changes in plant-growth-promoting effect of these mutants were monitored and compared to the effects of the wild-type strain in greenhouse experiments.
Materials and methods
Strains and growth conditions
B. amyloliquefaciens SQR9 (China General Microbiology Culture Collection Center, CGMCC accession No. 5808), green fluorescent protein (GFP)-labeled B. amyloliquefaciens SQR9 (SQR9-gfp) (Cao et al. 2011), and Escherichia coli DH5α (DH5α) were grown at 37 °C in Luria-Bertani (LB) medium, when necessary with the appropriate antibiotics (kanamycin at 10 μg ml−1 and erythromycin at 3 μg ml−1), solidified with 1.5 % agar. Strains were stored at −80 °C in LB containing 30 % glycerol. The production of phytase was determined by fermentating B. amyloliquefaciens SQR9 in 500-ml Erlenmeyer flasks with 100 ml of artificial sea water (ASW) medium (Idriss et al. 2002) in a rotary shaker at 37 °C, at 200 r min−1. Both acetoin and 2,3-butanediol production were determined by incubating B. amyloliquefaciens SQR9 cells in the LB medium containing glucose (1 % [w/v] final concentration) at 37 °C, at 90 r min−1. For IAA production, B. amyloliquefaciens SQR9 was cultured in liquid Landy medium (Landy et al. 1948) with or without l-tryptophan (3 mM) at 25 °C at 90 r min−1 shaking in the dark.
Growth-promoting assays
Cucumber (Jinchun 4) seeds were surface-sterilized (2-min, 70 % ethanol soaking followed by a 5-min, 10 % sodium hypochlorite soaking) and rinsed (four times) with sterile distilled water, and then grown in the nursery cups. They were transplanted to pots filled with 400 g soil-less growth media (Klasmann-Deilmann Base Substrate, Recipe-No. 422, blended with sterile vermiculite, 1:1) at two true leaves age. The experiment includes four treatments: CK1, plants were treated with 5 ml of inactivated (moist heat sterilized for 20 min) B. amyloliquefaciens SQR9 suspensions (108 cells ml−1); CK2, plants treated with 10 ml of inactivated B. amyloliquefaciens SQR9 suspensions (108 cells ml−1); T1, plants treated with 5 ml B. amyloliquefaciens SQR9 suspensions (108 cells ml−1); and T2, plants treated with 10 ml B. amyloliquefaciens SQR9 suspensions (108 cells ml−1). Suspensions of B. amyloliquefaciens SQR9 were prepared by shaking cells for 6 h in the liquid LB medium, and then cells were collected by centrifugation for 10 min at 8000×g and suspended in sterile distilled water (washed twice by sterile distilled water). Each treatment was replicated 30 times, and the experimental plan included three blocks in a completely randomized design (10 plants for each block). All treatments were incubated in a greenhouse at 70 % humidity, with 27 ± 2 °C at day and 22 ± 2 °C at night, natural light. Plants were irrigated regularly during the growing period, and pots were fertilized with 1 % (w/w) commercial fertilizer (alkali-hydrolyzed N, 6.27 %; available P, 4.71 %; available K 10.01 %). Ten randomly selected plants of each treatment were harvested after 55 days of transplanting, and plant height, root length and surface area, and shoot dry weight were measured. Data were analyzed using JMP software (SAS Institute Inc., Cary, NC).
Seeds of A. thaliana (ecotypes Columbia) were surface-sterilized as described above for cucumber seeds and then inoculated with B. amyloliquefaciens SQR9. Seeds were placed on Petri dishes containing half-strength modified Murashige and Skoog (MS) medium with 1.5 % (w/v) sucrose. The seeds were vernalized for 2 days at 4 °C in the dark. Then, the seedlings were placed in a growth chamber (14-h-light/10-h-dark cycles under 40-W fluorescent lights; the temperature was maintained at 25 ± 1 °C with a 50–60 % relative humidity). After 3 days, germinated seedlings (6–10 seedlings per plate) were transferred to one side of a divided Petri dish prepared with modified MS solid medium; then, 20 μl of B. amyloliquefaciens SQR9 (108 cells ml−1) in phosphorous buffer (PBS buffer, pH 7.0) and E. coli DH5α suspensions (108 cells ml−1) in PBS buffer (pH 7.0) (control) or sterilized PBS (control) were spotted onto the other side. The Petri dishes were sealed with parafilm. Co-cultivation was performed for 14 days in a growth chamber as described above.
Investigation of plant-growth-promoting factors
The phytase activity of B. amyloliquefaciens SQR9 was measured as described by Shimizu (1992). Detection and quantification of acetoin and 2,3-butanediol produced by B. amyloliquefaciens SQR9 were performed by the method of Nicholson (2008). HPLC and ELISA analysis were used for determining IAA produced by B. amyloliquefaciens SQR9. After incubation for 65 h, the bacterial abundance of each culture was measured at 600 nm, and the cells of B. amyloliquefaciens SQR9 and mutant strains were separated by centrifugation (5000×g, 20 min, 4 °C). Next, 50 μl culture supernatant of each strain were subjected to ELISA analysis with the IAA ELISA KIT (Cloud-Clone Corp., USA), and the IAA content quantified. Supernatants were adjusted to pH 2.5 with 1.0 M HCl and extracted three times with ethyl acetate (1:3, v/v). The organic solvents were vacuum-dried at 37 °C and then dissolved in 3 ml methanol. The extracted samples were filtered through a 0.45-μm membrane before HPLC (1200 series, Agilent, USA) detection at 220 nm using a UV detector. Mobile phase was methanol 0.1 % acetic acid (60/40) at a flow rate of 0.4 ml min−1 for 20 min. IAA was determined and quantified by integrating the areas of peaks with the help of standard samples (supplementary material).
Transposon mutagenesis
A shuttle plasmid pMarA (Breton et al. 2006), which contained a mariner-based transposon, was used to construct a random insertion library for screening mutant strains with different IAA production. The pMarA plasmids (kindly supplied by Plant Protection College, Nanjing Agricultural University) were transformed into competent cells of B. amyloliquefaciens SQR9 (Cao et al. 2011) selecting for Kanr and Emr at 30 °C. Plasmid DNA was then extracted from the transformants, subjected to KpnI digestion, and HimarI transposon fragments were amplified by PCR primers (oAtnpFwd and oAtnpRev, Table 1) to verify if these clones contained the original intact plasmid. A representative plasmid-containing colony was cultured overnight in liquid LB at 30 °C, then diluted (10−4) and plated on LB agar plates containing kanamycin and grown at 50 °C to select transposants.
Screening of IAA production variation mutants
Bacteria were cultivated for 12 h in Landy medium without l-tryptophan, and then a 20-μl aliquot was transferred into 5 ml of fresh Landy medium supplemented with 3 mM l-tryptophan. After 48 h (at 25 °C), the abundance of bacteria cells was measured at 600 nm; then, cells were separated from culture medium by centrifugation (5000×g, 20 min). One hundred microliters of the supernatant were mixed with 100 μl Salkowski’s reagent (0.5 M FeCl3 35 % HClO4, 2:100) in a 96-well plate. After 30 min at room temperature in the dark, the absorbance was measured at 535 nm. The concentration of IAA in the culture was determined by a standard IAA (Sigma) curve. IAA levels produced by SQR9 and its mutants were calculated by the OD535/OD600 ratio. IAA production of mutants was also quantified by HPLC analysis and ELISA analysis as described above with the growth curve being determined by OD600. Measurements were replicated three times.
The transposon insertion sites were analyzed by inverse-PCR (IPCR) as reported by Breton et al. (2006). The oIPCR1 and oIPCR2 (Table 1) primers, which face outward from the transposon sequence, were used in IPCR, whose products were purified by the quick PCR purification kit and sequenced after amplification by the oIPCR3 primer (Table 1).
Investigation of phytase, acetoin, and 2,3-butanediol produced by the mutant strains
SQR9 strains were cultured in the ASW and LB (containing 1 % glucose [w/v]) media for phytase, and acetoin and 2,3-butanediol productions, respectively. The detection methods were described above and shown in details in the supplementary material; the abundance of bacteria cells was continuously measured at 600 nm.
Colonization assay of B. amyloliquefaciens SQR9 strains
To study root colonization, green fluorescent protein (GFP)-labeled B. amyloliquefaciens SQR9 (Cao et al. 2011) was used instead of SQR9. Cultures of the SQR9 strains were grown in 50 ml LB media with kanamycin at 37 °C until reaching the stationary phase. The cells were washed twice in sterile PBS buffer (pH 7.0) and resuspended in PBS buffer (OD600 = 0.5) prior to use. Axenic-prepared cucumber seedlings were incubated in bacterial suspensions for 1 h at room temperature. Then, the seedlings were transplanted to containers with 200 ml of sterile 1/2 MS medium. The plants were grown in a growth chamber at 27 °C with a 16-h-light regimen. Each treatment was replicated 10 times. After 5 days, bacteria were extracted from 0.2-g roots randomly selected from each treatment by briefly rinsing the roots in sterile water; then, roots were homogenized in 1.8 ml PBS buffer using a mortar and pestle. The homogenates were serially diluted and plated onto LB plates containing kanamycin. SQR9 strains on cucumber seedling roots were determined.
Greenhouse experiment of B. amyloliquefaciens SQR9 and its derived strains
Pot experiments were performed for comparing the plant growth promotion effect of the wild-type B. amyloliquefaciens SQR9 and its strains with different IAA production (SQR9-IAA1, SQR9-IAA2, and SQR9-IAA3). Cucumber (Jinchun 4) seeds were surface-sterilized as described above. Seedlings were treated with 10-ml suspensions (108 CFU ml−1) of SQR9 strains, equal amounts of sterile distilled water served as control. Suspensions of SQR9 strains were prepared by shaking cells for 6 h in liquid LB media, and cells were collected by 10-min centrifugation at 5000×g; then, cells were suspended in sterile distilled water (washed twice by sterile distilled water). Each treatment was replicated 30 times and the experimental plan included three blocks in a completely randomized design (10 plants for each block). Pots were incubated in a greenhouse at 70 % humidity with a natural light and 27 ± 2 °C at day and 22 ± 2 °C at night. Plants were irrigated regularly during the growing period. Ten randomly selected plants of each treatment were harvested after 55 days. Plant height and shoot dry weight were measured. Data were analyzed using JMP software (SAS Institute Inc., Cary, NC).
Results
B. amyloliquefaciens SQR9 significantly stimulated growth of cucumber and A. thaliana
Equal amounts of inactivated B. amyloliquefaciens SQR9 cells were used as control (CK). Obvious differences among CK replicates were not observed, and this revealed that the presence of bacterial cells is not a factor of plant growth promotion (Table 2). When 5-ml B. amyloliquefaciens SQR9 suspensions (108 cells ml−1) were applied to each pot (T1 treatment), the cucumber yield, shoot height, root length, and root surface area increased by 60.1, 45.7, 29.3, and 30.7 %, respectively (Table 2). When 10-ml cell suspensions (108 cells ml−1) were applied (T2 treatment), increases amounted to 90.0, 71.6, 56.3, and 65.6 %, respectively (Table 2).
The effects of volatile chemicals produced by B. amyloliquefaciens SQR9 on plant growth were tested with divided Petri dishes so that only airborne signals could be transmitted between bacteria and the plant seedlings. After 14 days of incubation, B. amyloliquefaciens SQR9 significantly stimulated A. thaliana growth compared with the PBS and E. coli DH5α controls (Fig. 1a).
Acetoin and 2,3-butanediol produced by B. amyloliquefaciens SQR9. a Growth promotion of A. thaliana after exposure to airborne compounds released from B. amyloliquefaciens SQR9 compared with a nongrowth-promoting E. coli strain DH5α and PBS treatments. b The profile of acetoin and 2,3-butanediol productions and growth curve of B. amyloliquefaciens SQR9. Bars represent standard deviations of three biological replicates. c TLC assay of 2,3-butanediol in culture filtrates of B. amyloliquefaciens SQR9 in a TLC plate after spraying with modified Seebach solution and heat treatment
B. amyloliquefaciens SQR9 production of potentially plant-growth-promoting factors
Acetoin production by B. amyloliquefaciens SQR9 was low during exponential growth, increased during stationary phase, and peaked at 60 h after inoculation, reaching a concentration of 15.87 mM (Fig. 1b). Thin-layer chromatography (TLC) analysis showed that B. amyloliquefaciens SQR9 could produce 2,3-butanediol (Fig. 1c), which peaked at 22 h after inoculation (Fig. 1b). B. amyloliquefaciens SQR9 was incubated in phosphate-free ASW, and phytase activity increased markedly during the late exponential growth and peaked at 100 h after inoculation (0.28 U ml−1) (Fig. 2).
B. amyloliquefaciens SQR9 cultured in Landy medium with or without l-tryptophan (3 mM) showed positive color reaction by using Salkowski reagent, indicating the presence of an IAA-like substance in the supernatant. Both HPLC and ELISA analysis of supernatants confirmed the production of IAA by B. amyloliquefaciens SQR9 (Fig. S1) and the produced IAA reached a concentration of 9.46 mg l−1 when l-tryptophan was added to Landy medium (Fig. 3), which was almost 3.6-fold compared to the no-l-tryptophan treatment (data not shown). This observation indicated that IAA production in B. amyloliquefaciens SQR9 is tryptophan dependent.
Screening of SQR9 strains differing in the IAA production
Above results showed that SQR9 is an efficient plant-growth-promoting strain producing IAA. To evaluate the contribution of IAA production to plant growth promotion, a random transposon mutant library of SQR9 was constructed to select different IAA production strains. About 3000 mutants were initially screened by colorimetric assays. Three mutants were selected, one mutant (designated as SQR9-IAA1) was almost completely deficient in IAA production, another mutant (designated as SQR9-IAA2) showed decreased IAA production, and the third mutant (designated as SQR9-IAA3) showed increased IAA production. In mutant SQR9-IAA1, SQR9-IAA2, and SQR9-IAA3 strains, the TnYLB-1 transposon was inserted at qoxB, ylmD, and yngF genes, respectively.
IAA production and bacterial growth were measured in culture supernatants after 65 h (Fig. 4). Both SQR9-IAA2 and SQR9-IAA3 strains exhibited almost identical growth curves with the wild-type strain, while SQR9-IAA1 strain grew slower than the wild-type strain. Less than 10 % of the IAA produced by the wild type was detected in the culture supernatant of SQR9-IAA1 strain, and IAA production in SQR9-IAA2 strain was nearly 50 % of the wild-type strain, while in SQR9-IAA3 strain, the IAA production was nearly 240 % compared with that of the wild-type strain (Fig. 4).
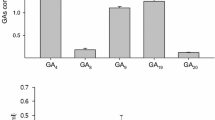
Growth curve and IAA production (column diagram) of B. amyloliquefaciens SQR9 strains grown in Landy medium supplemented with tryptophan. IAA production was quantified at stationary phase (65 h) and the IAA concentration produced by SQR9 was defined as 1. Bars represent standard deviations of three biological replicates
Phytase, acetoin, and 2,3-butanediol produced by IAA production variation mutant strains
As these mutants could have effects unrelated to IAA synthesis, we investigated their abilities to produce phytase, acetoin, and 2,3-butanediol. The three mutants showed no difference in growth curve in ASW medium and LB medium containing glucose. In addition, phytase, acetoin, and 2,3-butanediol produced by the three mutants were not affected by the transposon mutagenesis (Fig. 5).
Root colonization
The population size attained by the mutant strains on the surface 5 days after inoculation was similar to that attained by the SQR9-gfp (about 106 CFU g−1 root) suggesting that the reduced and increased IAA production in SQR9 strains did not affect the ability to colonize cucumber seedling roots (Fig. 6).
Effect of SQR9 and IAA production on cucumber growth
All bacterial treatments showed different plant-growth-promoting effects compared with the non-treated control. Strain SQR9-IAA1, nearly deficient in IAA production, decreased the biomass weight and shoot height by 24 and 18.9 % compared to the wild-type strain (Fig. 7). SQR9-IAA3 strain, which produced approximately 240 % IAA of the wild-type strain, also significantly (P < 0.05) promoted cucumber biomass over the other strains. Although SQR9-IAA2 strain produced about 50 % IAA of the wild-type strain, it showed no reduction in plant growth promotion compared with the wild-type strain (Fig. 7).
Effect of B. amyloliquefaciens SQR9 strains on growth of cucumber transplant plugs. Different letters indicate significantly differences at P < 0.05, least significant difference. CK, seedlings inoculated with 10 ml sterile distilled water; SQR9-IAA1, seedlings inoculated with 10-ml cell suspensions of SQR9-IAA1 strain (l08 CFU ml−1); SQR9, seedlings inoculated with 10-ml cell suspensions of the wild-type SQR9 (l08 CFU ml−1); SQR9-IAA2, seedlings inoculated with10-ml cell suspensions of SQR9-IAA2 strain (l08 CFU ml−1); SQR9-IAA3, seedlings inoculated with 10-ml cell suspensions of SQR9-IAA3 strain (l08 CFU ml−1). Plants were grown in greenhouse for 55 days. Bars represent standard deviations of three biological replicates
Discussion
B. amyloliquefaciens SQR9 is a rhizobacterium isolated from the rhizosphere of cucumber (Cao et al. 2011). We have demonstrated that B. amyloliquefaciens SQR9 can directly promote plant growth by various mechanisms. The prerequisite of disease suppression and plant growth promotion is the efficient colonization of plant roots by the PGPR strain (Qiu et al. 2014). Indeed B. amyloliquefaciens SQR9 is a root colonizer since it can maintain the population at 106 CFU g−1 root after 23 days of inoculation (Cao et al. 2011).
Genomic analysis of SQR9 (NCBI accession no. CP006890) revealed several genes involved in the plant-growth-promoting effects. A whole set of genes alsRSD (V529_35850, V529_35860, and V529_35870) responsible for acetoin synthesis were identified. In this cluster, alsS gene potentially encodes the enzyme α-acetolactate synthase, which convert pyruvate to α-acetolactate; alsD gene encodes α-acetolactate decarboxylase catalyzing the conversion of α-acetolactate to acetoin; alsR gene, which encodes a positive transcriptional regulator of alsSD, is located downstream of alsSD (Renna et al. 1993). YwrO was found upstream of alsRSD gene cluster, which encodes a putative NAD (P)-dependent oxidoreductase in Bacillus (Renna et al. 1993). However, the bdhA gene of B. subtilis (Nicholson 2008), which encodes 2,3-butanediol dehydrogenase responsible for the conversation of acetoin to 2,3-butanediol, was not found in B. amyloliquefaciens SQR9 genome. The phy gene (V529_21880), which encodes phytase in Bacillus spp. (Kerovuo et al. 1998; Kim et al. 1998; Idriss et al. 2002), was also identified in B. amyloliquefaciens SQR9. Moreover, acetoin, 2,3-butanediol and phytase activity were detected in the culture supernatant of B. amyloliquefaciens SQR9. The volatiles acetoin and 2,3-butanediol released by Bacillus strains can enhance plant growth (Ryu et al. 2003). Also, extracellular bacteria phytase released by B. amyloliquefaciens FZB strains can contribute to the plant-growth-promoting activity under P limiting condition (Idriss et al. 2002; Makarewicz et al. 2006).
Tryptophan has been postulated as a main precursor for IAA synthesis in bacteria (Spaepen et al. 2007; Duca et al. 2013); we also showed that the primary route of IAA biosynthesis in B. amyloliquefaciens SQR9 depends on the presence of this amino acid since lower IAA concentration was detected in the supernatant without tryptophan supply. Genes with high homology to ysnE (putative IAA transacetylase) and yhcX (putative nitrilase) in B. amyloliquefaciens FZB42 (Idris et al. 2007) were found in the genome of B. amyloliquefaciens SQR9 (V529_38080 and V529_08860), and these two genes were suggested to be involved in tryptophan-dependent IAA synthesis pathways in B. amyloliquefaciens FZB42. Three strains showing variation in IAA production were obtained by construction of a mutant library with plasmid pMarA. The transposon was inserted into an ORF coding for quinol oxidase polypeptide I (V529_38120) in strain SQR9-IAA1. The menaquinol oxidase (aa3-600) catalyzes oxidation of quinols with release of energy during vegetative growth; the relation between growth rate and qox expression reflects the need of energy of the cell (Santana et al. 1992). In glucose minimal medium, the qoxB mutant strain grows slower than the B. subtilis wild-type strain (Santana et al. 1992), as also found by us with the Landy medium containing 20 g l−1 glucose and 1 g l−1 yeast extract. We suggest that the quinol oxidase was involved in the electron transport chains of tryptophan metabolism, since the IAA biosynthesis contains oxidation-reduction reactions like indole-3-acetaldehyde dehydrogenase converting indole-3-acetaldehyde to IAA. In strain SQR9-IAA2, the transposon was inserted into ylmD ORF (V529_14750), encoding the multi-copper polyphenol oxidoreductase laccase, which may be involved in cytochrom c maturation system (Kuras et al. 2007). In strain SQR9-IAA3, the transposon was inserted into yngF ORF (V529_18040), which is located in yng operon and is expected to be required for degradation of fatty acids to acetyl-CoA (Koburger et al. 2005). 3-Methylbutyryl-CoA, produced by the oxidation of fatty acids, can be decarboxylated to acetyl-CoA and acetoacetyl-CoA by enzyme encoded by yng operon; this intermediate can also be produced by the degradation of branched chain amino acids (Kanehisa et al. 2002). However, the role of qoxB, ylmD and yngF in tryptophan metabolism and IAA synthesis in B. amyloliquefaciens SQR9 remains to be illustrated.
Greenhouse experiment of SQR9 strains showed that IAA production was directly related to its plant-growth-promoting effect. However, the lowest IAA production mutant SQR9-IAA1 still promoted growth of cucumber compared to the control, and SQR9-IAA2 strain, with decreased synthesis of IAA, showed similar plant growth promotion effect of the wild-type SQR9 strain. The reduced or enhanced growth promoting effects were not attributed to the root colonization since all mutants showed similar root colonization ability to the SQR9-gfp strain. This confirms the report by Patten and Glick (2002), who showed that the Pseudomonas putida GR12-2 IAA mutant strain was able to colonize canola roots as the wild-type strain, while IAA mutant strain of Erwinia herbicola 299R was less competitive than the wild-type strain for root colonization (Brandl and Lindow 1997). Plant-growth-promoting factors like phytase, acetoin, and 2,3-butanediol produced by the three IAA production variation mutants were not affected by the transposon mutagenesis. These results suggest that IAA production is involved in the plant-growth-promoting effect but IAA biosynthesis alone cannot account for the overall effects of B. amyloliquefaciens SQR9. In our study, seedlings were grown in sterile soil, and this avoided the effects of soil native microflora. PGPR can interact with other microbes in soil, especially mycorrhizal fungi, to give a net growth response (Meyer and Linderman 1986). Another review by Bashan et al. (2004) on Azospirillum indicated the potential role of multiple factors in plant growth enhancement including co-inoculation of Azospirillum with other microbial species.
In conclusion, our results indicated that in B. amyloliquefaciens SQR9, IAA production alone cannot account for the overall observed effects, since its stimulation of plant growth can be attributed to multiple factors, including generation of phytohormone, production of volatile compounds, and release of extracellular enzymes. Our study not only has improved our understanding of the plant growth promotion mechanisms but also has provided some indications for the isolation of plant-growth-promoting rhizosphere bacteria for agricultural application.
References
Adesemoye AO, Torbert HA, Kloepper JW (2008) Enhanced plant nutrient use efficiency with PGPR and AMF in an integrated nutrient management system. Can J Microbiol 54:876–886
Babalola OO (2010) Beneficial bacteria of agricultural importance. Biotechnol Lett 32:1559–1570
Barbieri P, Galli E (1993) Effect on wheat root development of inoculation with an Azospirillum brasilense mutant with altered indole-3-acetic acid production. Res Microbiol 144:69–75
Bashan Y, Holguin G, De-Bashan LE (2004) Azospirillum-plant relationships: physiological, molecular, agricultural, and environmental advances (1997–2003). Can J Microbiol 50:521–577
Bottini R, Cassán F, Piccoli P (2004) Gibberellin production by bacteria and its involvement in plant growth promotion and yield increase. Appl Microbiol Biotechnol 65:497–503
Brandl MT, Lindow SE (1997) Environmental signals modulate the expression of an indole-3-acetic acid biosynthetic gene in Erwinia herbicola. Mol Plant Microbe Interact 10:499–505
Breton YL, Mohapatra NP, Haldenwang WG (2006) In vivo random mutagenesis of Bacillus subtilis by use of TnYLB-1, a mariner-based transposon. Appl Environ Microbiol 72:327–333
Cao Y, Zhang Z, Ling N, Yuan Y, Zheng X, Shen B, Shen Q (2011) Bacillus subtilis SQR 9 can control Fusarium wilt in cucumber by colonizing plant roots. Biol Fertil Soils 47:495–506
Chaiharn M, Lumyong S (2011) Screening and optimization of indole-3-acetic acid production and phosphate solubilization from rhizobacteria aimed at improving plant growth. Curr Microbiol 62:173–181
Duca D, Lorv J, Patten CL, Rose D, Glick BR (2013) Indole-3-acetic acid in plant–microbe interactions. Antonie Van Leeuwenhoek 1–41
Emmert EA, Handelsman J (1999) Biocontrol of plant disease: a gram-positive perspective. FEMS Microbiol Lett 171:1–9
Farag MA, Paré PW (2002) C6-Green leaf volatiles trigger local and systemic VOC emissions in tomato. Phytochemistry 61:545–554
Farag MA, Ryu CM, Sumner LW, Paré PW (2006) GC–MS SPME profiling of rhizobacterial volatiles reveals prospective inducers of growth promotion and induced systemic resistance in plants. Phytochemistry 67:2262–2268
Farmer EE (2001) Surface-to-air signals. Nature 411:854–856
Frommel MI, Nowak J, Lazarovits G (1991) Growth enhancement and developmental modifications of in vitro grown potato (solanumtuberosum spp. tuberosum) as affected by a nonfluorescent pseudomonas sp. Plant Physiol 96:928–936
Idris EE, Bochow H, Ross H, Borriss R (2004) Use of Bacillus subtilis as biocontrol agent. VI. Phytohormone-like action of culture filtrates prepared from plant growth-promoting Bacillus amyloliquefaciens FZB24, FZB42, FZB45 and Bacillus subtilis FZB37. J Plant Dis Prot 111:583–597
Idris EE, Iglesias DJ, Talon M, Borriss R (2007) Tryptophan-dependent production of indole-3-acetic acid (IAA) affects level of plant growth promotion by Bacillus amyloliquefaciens FZB42. Mol Plant Microbe Interact 20:619–626
Idriss EE, Makarewicz O, Farouk A, Rosner K, Greiner R, Bochow H, Richter T, Borriss R (2002) Extracellular phytase activity of Bacillus amyloliquefaciens FZB45 contributes to its plant-growth-promoting effect. Microbiology 148:2097–2109
Kanehisa M, Goto S, Kawashima S, Nakaya A (2002) The KEGG databases at GenomeNet. Nucleic Acids Res 30:42–46
Kerovuo J, Lauraeus M, Nurminen P, Kalkkinen N, Apajalahti J (1998) Isolation, characterization, molecular gene cloning, and sequencing of a novel phytase from Bacillus subtilis. Appl Environ Microbiol 64:2079–2085
Kim YO, Lee JK, Kim HK, Yu JH, Oh TK (1998) Cloning of the thermostable phytase gene (phy) from Bacillus sp. DS11 and its overexpression in Escherichia coli. FEMS Microbiol Lett 162:185–191
Kloepper JW, Lifshitz R, Zablotowicz RM (1989) Free-living bacterial inocula for enhancing crop productivity. Trends Biotechnol 7:39–44
Koburger T, Weibezahn J, Bernhardt J, Homuth G, Hecker M (2005) Genome-wide mRNA profiling in glucose starved Bacillus subtilis cells. Mol Genet Genomics 274:1–12
Kochar M, Upadhyay A, Srivastava S (2011) Indole-3-acetic acid biosynthesis in the biocontrol strain Pseudomonas fluorescens Psd and plant growth regulation by hormone overexpression. Res Microbiol 162:426–435
Kuras R, Saint-Marcoux D, Wollman FA, Vitry CD (2007) A specific c-type cytochrome maturation system is required for oxygenic photosynthesis. Proc Natl Acad Sci 104:9906–9910
Landy M, Warren GH, Rosenman SB, Colio LG (1948) Bacillomycin an antibiotic from Bacillus subtilis active against pathogenic Fungi. Exp Biol Med 67:539–541
Liu Y, Shi J, Feng Y, Yang X, Li X, Shen Q (2013) Tobacco bacterial wilt can be biologically controlled by the application of antagonistic strains in combination with organic fertilizer. Biol Fertil Soils 49:447–464
Madhaiyan M, Poonguzhali S, Ryu J, Sa T (2006) Regulation of ethylene levels in canola (Brassica campestris) by 1-aminocyclopropane-1-carboxylate deaminase-containing Methylobacterium fujisawaense. Planta 224:268–278
Makarewicz O, Dubrac S, Msadek T, Borriss R (2006) Dual role of the Pho∼P approximately P response regulator: Bacillus amyloliquefaciens FZB45 phytase gene transcription is directed by positive and negative interactions with the phyC promoter. J Bacteriol 188(19):6953–65
Meyer JR, Linderman RG (1986) Selective influence on populations of rhizosphere or rhizoplane bacteria and actinomycetes by mycorrhizas formed by Glomus fasciculatum. Soil Biol Biochem 18:191–196
Nicholson WL (2008) The Bacillus subtilis ydjL (bdhA) gene encodes acetoin reductase/2,3-butanediol dehydrogenase. Appl Environ Microbiol 74:6832–6838
Palacios OA, Bashan Y, De-Bashan LE (2014) Proven and potential involvement of vitamins in interactions of plants with plant growth-promoting bacteria—an overview. Biol Fertil Soils 50:415–432
Patten CL, Glick BR (2002) Role of Pseudomonas putida indoleacetic acid in development of the host plant root system. Appl Environ Microbiol 68:3795–3801
Penrose DM, Glick BR (2001) Levels of ACC and related compounds in exudate and extracts of canola seeds treated with ACC deaminase-containing plant growth-promoting bacteria. Can J Microbiol 47:368–372
Ping L, Boland W (2004) Signals from the underground: bacterial volatiles promote growth in Arabidopsis. Trends Plant Sci 9:263–266
Qiu M, Li S, Zhou X, Cui X, Vivanco JM, Zhang N, Shen Q, Zhang R (2014) De-coupling of root–microbiome associations followed by antagonist inoculation improves rhizosphere soil suppressiveness. Biol Fertil Soils 50:217–224
Qiu M, Zhang R, Xue C, Zhang S, Li S, Zhang N, Shen Q (2012) Application of bio-organic fertilizer can control Fusarium wilt of cucumber plants by regulating microbial community of rhizosphere soil. Biol Fertil Soils 48:807–816
Renna MC, Najimudin N, Winik LR, Zahler SA (1993) Regulation of the Bacillus subtilis alsS, alsD, and alsR genes involved in post-exponential-phase production of acetoin. J Bacteriol 175:3863–3875
Richardson AE, Hadobas PA, Hayes JE (2001) Extracellular secretion of Aspergillus phytase from Arabidopsis roots enables plants to obtain phosphorus from phytate. Plant J 25:641–649
Ryu CM, Farag MA, Hu CH, Reddy MS, Kloepper JW, Pw P (2004) Bacterial volatiles induce systemic resistance in Arabidopsis. Plant Physiol 134:1017–1026
Ryu CM, Farag MA, Hu CH, Reddy MS, Kloepper JW, Wei HX, Paré PW (2003) Bacterial volatiles promote growth in Arabidopsis. Proc Natl Acad Sci 100:4927–4932
Santana M, Kunst F, Hullo MF, Rapoport G, Danchin A, Glaser P (1992) Molecular cloning, sequencing, and physiological characterization of the qox operon from Bacillus subtilis encoding the aa3-600 quinol oxidase. J Biol Chem 267:10225–10231
Shimizu M (1992) Purification and characterization of phytase from Bacillus subtilis (natto) N-77. Biosci Biotech Bioch 56:1266–1269
Spaepen S, Vanderleyden J, Remans R (2007) Indole-3-acetic acid in microbial and microorganism-plant signaling. FEMS Microbiol Rev 31:425–448
Teale WD, Paponov IA, Palme K (2006) Auxin in action: signalling, transport and the control of plant growth and development. Nat Rev Mol Cell Biol 7:847–859
Wang B, Yuan J, Zhang J, Shen Z, Zhang M, Li R, Ruan Y, Shen Q (2013) Effects of novel bioorganic fertilizer produced by Bacillus amyloliquefaciens W19 on antagonism of Fusarium wilt of banana. Biol Fertil Soils 49:435–446
Weng J, Wang Y, Li J, Shen Q, Zhang R (2013) Enhanced root colonization and biocontrol activity of Bacillus amyloliquefaciens SQR9 by abrB gene disruption. Appl Microbiol Biotechnol 97:8823–8830
Woodward AW, Bartel B (2005) Auxin: regulation, action, and interaction. Ann Bot 95:707–735
Xu Z, Shao J, Li B, Yan X, Shen Q, Zhang R (2013) Contribution of bacillomycin D in Bacillus amyloliquefaciens SQR9 to antifungal activity and biofilm formation. Appl Environ Microbiol 79:808–815
Zhang H, Kim MS, Krishnamachari V, Payton P, Sun Y, Grimson M, Farag MA, Ryu CM, Allen R, Melo IS, Paré PW (2007) Rhizobacterial volatile emissions regulate auxin homeostasis and cell expansion in Arabidopsis. Planta 226:839–851
Acknowledgments
This research was financially supported by the Chinese Ministry of Science and Technology (2013AA102802 and 2011BAD11B03). NZ was supported by the Fundamental Research Funds for the Central Universities (KYZ201408); RZ and QS were also supported by the Priority Academic Program Development (PAPD) of Jiangsu Higher Education Institutions and the 111 Project (B12009).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 198 kb)
Rights and permissions
About this article
Cite this article
Shao, J., Xu, Z., Zhang, N. et al. Contribution of indole-3-acetic acid in the plant growth promotion by the rhizospheric strain Bacillus amyloliquefaciens SQR9. Biol Fertil Soils 51, 321–330 (2015). https://doi.org/10.1007/s00374-014-0978-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-014-0978-8