Abstract
A halotolerant bacterial strain isolated and identified as Bacillus gibsonii was used for extracellular lipase production. The bacterial strain was able to grow up to 1200 mM salt concentration and showed maximum growth at 600 mM NaCl concentration. The present study includes production of extracellular lipase enzyme and characterization of partially purified lipase with respect to its kinetic and thermodynamic behaviour. Maximum lipase activity was observed at 60 °C under alkaline pH (9.0) condition. The kinetic parameters such as Vmax, Km and Kcat were calculated as 158.73 U/mL, 0.539 mM and 483.93 min−1 at 60 °C, respectively, suggested thermostable nature of the enzyme. The thermal inactivation energy [Ea(d)] was calculated as 66.98 kJ/mol. The values of Gibb’s free energy (86.31 kJ/mol), enthalpy (64.26 kJ/mol) and entropy (− 66.21 × 10–3 kJ/mol/K) for the enzyme inactivation obtained at 60 °C corroborated the assumption that 60 °C was the optimum temperature. Further, the deactivation rate constant (kd) values calculated at 60 °C and 80 °C were found to be 0.0907 and 0.182 min−1, respectively, which suggested that enzyme was more stable at 60 °C and it was partly inactivated at 80 °C.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Lipases are defined as triacylglycerol acylhydrolases (EC 3.1.1.3) which naturally hydrolyze triglycerides into diglycerides, monoglycerides, glycerol and fatty acids (Houde et al. 2004). Microbial lipases have some additional properties, which make them potent enzyme for esterification, transesterification, aminolysis, alcoholysis, and deacetylation reactions (Sarmah et al. 2018). Microbial lipases, not only hydrolyze ester bonds of triglycerides, but also resolve racemic mixture and synthesize ester bonds in non-aqueous media (Hiol et al. 1999). Lipolytic activity in bacteria is dependent on the presence of carbon source as well as inducers such as oil, fatty acids, bile salts and triacyl glycerols (Bora and Mohan 2008). Microbial lipases produced by mesophilic microorganisms are active over a wide range of alkaline pH conditions, but are generally unstable at temperatures above 70 °C. Microbial lipases produced by mesophilic bacteria are generally more active between 30 and 65 °C (Royter et al. 2009). The thermostable microbial lipases have been purified and characterized from Bacillus sp. (Sugihara et al. 1991), Bacillus thermocatenulatus (Schmidt-Dannert et al. 1996), B. stearothermophilus (Kim et al. 2000), Bacillus sp. J33 (Nawani and Kaur 2000), B. strain A30-1 (Wang et al. 1995), Bacillus sp. RSJ-1 (Sharma et al. 2002) and B. thermoleovorans ID-1 (Lee et al. 1999).
Since microbial lipases are mostly used in industries at elevated temperatures, it is pertinent to study their thermal behaviour. Discovery of new thermo-stable microbial lipases have attracted the attention of many researchers with respect to developing strategies to enhance stability of lipases at elevated temperature. A major constraint in achieving the thermal stability of enzyme is the kinetic limitations imposed by high temperature, they require more rigid material, which are unsupportive to product formation (Li and Zhang 2005). The importance of thermostable lipases has been growing as they are more stable than the mesophilic lipase, which make them more potent for biotechnological applications. Thermostable lipase are endowed with some unusual characteristics such as higher stability with higher rate of catalysis, higher rate of diffusion, enhanced solubility of reactants in water and positive equilibrium displacement in endothermic reactions with reduced risk of microbial contamination (Hasan et al. 2006). Due to these advantages, the thermostable lipases are still highly attractive and are in high demand for industrial application. Thermostable lipase are not only produced by thermophilic or mesophilic microorganisms, but are also produced by psychrophilic microorganism (Griffiths et al. 1981).
Alkalophilic thermostable lipase enzymes are often considered to be more efficient and useful for industrial applications. Microbial lipase retain adequate functional characteristics with some restrictive factor. For example, lipase added to detergent powders and pre-wash soaking agents require to be stable against alkaline pH condition, i.e., above pH 7.0 (Treves et al. 1984). Furthermore, when tallow is the feedstock for hydrolysis, the lipase should be erratically active and thermostable between temperature ranges of 45 and 55 °C (Wang et al. 1995). The need of thermostable alkaline lipase emerged as majority of the lipases were not stable above pH 7.0 and 42 °C temperature. Therefore, a thermostable alkaline lipase is needed to meet the industrial and commercial demand (Bora et al. 2008).
Alkaline thermostable lipases have great potential for inclusion in the formulations of detergent and leather industries. At present, market of microbial lipases in detergent industry has increased by more than 60% (Hasan et al. 2006). Thermostable microbial lipase have found a place in the production of biofuels, cosmetics products, detergent formulation, synthesis of organic compounds, pharmaceuticals and medical industries, food and feed industry (Gandhi 1997). Microbial production of extracellular lipase is considered to be highly advantageous due to its low cost of production, ease of biotechnological manipulation of microbial system at high temperature and hence, bacterial lipases are better suited for industrial processes under harsh conditions.
The present study aimed to screen lipase producing halophilic bacterium and characterization of catalytic characteristics of lipase enzyme. Further characterization of enzyme was carried out with respect to kinetic and thermodynamic behaviour of enzyme under varying temperature. To the best of our knowledge, this is the first report on catalytic, kinetic and thermodynamic behaviour of extracellular lipase produced by halophilic bacterium B. gibsonii.
Materials and methods
Screening and isolation of lipase producing microorganism
The halotolerant bacterium was isolated from saline water collected from the coastal regions of Gujarat, India. For screening of halo-tolerant bacterial species, the water sample was enriched with nutrients of culture media by inoculating 1 mL of water sample into 50 mL growth medium containing peptone 0.5%; yeast extract 0.3%; tributyrin 0.1%; pH 7.0 in a 250 mL Erlenmeyer flask (Griebeler et al. 2011). The inoculated culture medium was kept on orbital shaker at 30 °C for 48–72 h under shaking condition at 150 rpm. The sample culture was streaked on tributyrin agar plate (TBA) medium, prepared by mixing 2% (w/v) agar in culture medium. The large transparent bacterial colonies on the plate forming halo zones were picked up and were further purified by repeated streaking on tributyrin-agar-plates (TBA).
Effect of salt on growth of isolated bacterial strain
The growth of bacterial isolate obtained from saline water was optimized under different growth conditions. Effect of salt (NaCl) on the growth of bacterial isolate was studied by inoculating the bacterial culture (1% inoculum) in the medium containing varying NaCl concentrations (50–1200 mM). The flasks were incubated at 30 °C under shaking condition at 150 rpm for 120 h. The sample was collected every 8 h up to 120 h for recording the optical density of the culture at 600 nm, using double beam UV–Vis spectrophotometer (UV-1601, Shimadzu, Japan).
Molecular identification of isolated bacterial strain
The monoclonal culture of extracellular lipase producing strain named as HYL was sent to CSIR-NCL (National Chemical Laboratories), Pune (India) for molecular identification. The 16S rRNA sequence of the isolated strain was amplified using bacteria specific universal primer 27F (5ˊ- CCAGAGTTTGATCCTGGCTCAG-3ˊ) and 1492R (5ˊ- CGGTTACCTTGTTACGACTT- 3ˊ). The sequence was compared with 16S rRNA gene database using BLAST. The rRNA sequence data were submitted to NCBI. The phylogenetic tree of the isolated strain was constructed using MEGA-X with the neighbour joining method. The isolated strain was given an accession number MK835660 and the sequence has been submitted to the Genbank Data Library.
Production of extracellular lipase enzyme
The isolated halotolerant bacterial strain was used to study the production of extracellular lipase enzyme. The growth medium containing 100 mL of medium (g/L): Beef extract 1.5, Yeast extract 1.5, Peptone 5.0, NaCl 600 mM with 1% (v/v) olive oil with pH 7.2 ± 0.2 was autoclaved at 121 °C for 15 min. The sterilized medium was inoculated with isolated bacterial strain (2%, v/v) and was incubated at 30 °C under shaking condition (150 rpm) for 72 h. At the end of incubation period (72 h.), the culture broth was centrifuged at 12,000xg for 15 min. The cell free supernatant exhibiting lipolytic activity was used as a source of extracellular lipase enzyme and was subjected for partial purification and characterization.
Enzyme assay and protein determination
The enzyme activity was assayed under different temperatures using p-nitrophenyl palmitate (pNPP) as the substrate (Ertuğrul et al. 2007). The reaction mixture was freshly prepared by mixing solution A [30 mg of substrate (pNPP) in 10 mL of isopropanol] with solution B (0.1 g of gum Arabic and 0.4 mL Triton X-100 in 90 mL of 0.05 M Tris–HCl buffer at pH 8.0, by stirring until the mixture was dissolved. The final reaction mixture for enzyme assay was prepared by mixing 9 μL of substrate solution and 1 μL of enzyme solution. The reaction mixture was incubated at 30 ± 0.1 °C for 15 min and absorbance of the reaction mixture was measured at 410 nm in a double beam UV–visible spectrophotometer. The rate of enzyme activity was expressed in terms of U/mL. Dissolved protein concentration was determined by the method of Lowry (Lowry and Tinsley 1976) using bovine serum albumin (BSA) as a standard.
Ammonium sulphate precipitation
The enzyme was partially purified from a fresh culture broth of halotolerant bacterium. The cell free culture broth obtained after 72 h of incubation was used for purification of the lipase enzyme. The partial purification involved an appropriate amount of ammonium sulphate at a different concentration range from 40 to 70% (w/v) saturations. The ammonium sulphate precipitate obtained after centrifugation was dissolved in 0.2 M phosphate buffer (pH 7.0 ± 0.2). After dissolving the precipitate in phosphate buffer, the protein solution was dialyzed against the same phosphate buffer. The phosphate buffer was changed regularly after every 1 hour till the release of ammonium sulphate salt stopped coming from the dialysis tube. The partially purified lipase was stored at − 4 °C for further use.
Characterization of extracellular lipase enzyme
Effect of pH on lipase enzyme
Effect of different pH conditions (pH 4.0–11) was studied on the enzyme catalysis (30 °C). As described by Sinchaikul et al. (2001), different buffers were used to maintain various pH conditions, such as 50 mM sodium acetate buffer was used for pH range from pH 4.0–6.0, 50 mM phosphate buffer was used for pH range from pH 6.0–7.5, 50 mM Tris–HCl buffer for pH range from pH 8.0–10 and 50 mM Tris–glycine buffer for pH 9.0–11.
Effect of different metal salts on lipase activity
Effect of different metal salts on the lipase enzyme was studied by pre-incubating the enzyme solution with 1 mM of each metal salt solution for 30 min. Later on, the reaction mixture was prepared by adding all ingredients of assay mixture including each metal salt solution in 50 mM Tris buffer (pH 8.0) at 30 °C for 30 min. The metal salts used in the study were AlCl3, ZnSO4, BaCl2, MgCl2, MnSO4, CaCl2 and CoCl2, at a final concentration of 1.0 mM, each. The effect of metal salts on lipase activity was expressed as percentage stimulation and inhibition in the enzyme activity as compared to control (without metal).
Effect of different inhibitor and surfactants
The effect of different enzyme inhibitors and surfactants on the lipase enzyme was carried out at 30 °C by pre-incubating the enzyme solution for 30 min at pH 8.0 (0.05 M Tris buffer). The enzyme inhibitors viz dithiothreitol (DTT), 2-mercaptoehanol (BME), phenylmethylsulfonyl fluoride (PMSF), Ethylene diamine tetra acetic acid (EDTA), and ionic surfactants as sodium dodecyl sulfate (SDS) and non-ionic surfactants as Tween 80, Tween 20 and Triton X-100 were used at final concentration of 1 mM each. The enzyme activity was calculated in terms of percentage stimulation and inhibition of the lipase activity as compared to control (without inhibitor).
Kinetic and thermal behaviour of partially purified lipase
Kinetics and thermal properties of lipase
Enzyme activity was measured at different temperatures. The enzyme reaction rate constant (k) for each temperature was calculated. The activation energy (Ea) was determined from the slope of the Arrhenius plot, which was plotted between logarithms of enzyme reaction rate constant versus reciprocal of absolute temperature. The kinetic values were calculated by the following formula (Prajapati et al. 2014).
where, \(E_{a}\) = activation energy, \(R = 8.314J/K/mol\)(gas constant), \(T\) = absolute temperature (K), \(\Delta H^{ * }\) = change in enthalpy, \(\Delta G^{ * }\) = Gibbs free energy of activation. Enthalpy (\(\Delta H^{ * }\)) of activation for each temperature was calculated by following equation
where, \(E_{a}\) = activation energy.
The Gibb’s free energy was calculated from the first- order rate constant:
(Gibb’s free energy of activation)
The activation entropy (\(\Delta S^{ * }\)) for extracellular lipase was calculated from the following equation:
Thermal inactivation of lipase enzyme
Thermal inactivation of the enzyme was performed by pre-incubating the enzyme at various temperatures ranging from 60 to 80 °C (pH 8.0) as described by Whitaker (Whitaker 1994).The residual activity of the enzyme was determined in terms of percentage residual activity at room temperature as mentioned below: (Olusesan et al. 2011).
Residual activity (%) =
where, \(C_{t}\) = enzyme activity at time t (min). \(C_{o}\) = enzyme activity at time 0 (min).
Enzyme inactivation reactions often follow first-order kinetic model, which revealed that enzyme activity decreased log linearly as a function of time. The first-order inactivation rate constant was calculated by the following equation (Ortega et al. 2004).
where, \(C_{t}\) = enzyme activity at time t, \(C_{0}\) = enzyme activity at 0 min, \(t\) = time of enzyme treatment, \(k_{d}\) = inactivation rate constant.
Enzyme inactivation of lipase was determined by incubating the reaction mixture in 50 mM of Tris–HCl buffer for 90 min at various temperatures (60, 70 and 80 °C) without substrate. An aliquot (1 mL) were taken from temperature treatment reaction mixture at a different time interval, cooled for 15 min and enzyme assayed was performed under standard condition. The deactivation rate constant (kd) for each temperature treatment was determined by mentioned below: (Prajapati et al. 2014).
The deactivation energy (\(E_{a(d)}\)) for inactivation reaction was determined by the Arrhenius plot against ln (kd) vs 1/T(K). The half-life (\(t_{1/2}\)) of enzyme was defined as the time at which the residual activity reaches 50% which was calculated by the following expression:
Free energy change (\(\Delta G^{ * }\)) for inactivation of lipase was calculated using following equation:
Decimal reduction time (D value) was describe as the time required by enzyme to reduced 90% of initial activity at a specific temperature was calculated as follows:
Statistical analysis
All the experiments in this study were performed in triplicate and the data presented are mean values ± standard deviation of triplicates experiments.
Results and discussion
Isolation and screening of lipolytic strain
The present study involved the isolation and screening of halotolerant bacterium B. gibsonii for the production of extracellular lipase enzyme. After the serial dilution of water sample the bacterial colonies which grown on nutrient agar plates, were exposed on tributyrin agar plate for confirmation of lipolytic activity. The size of halos zone produced by bacterial strain on tributyrin agar plate were measures, which was found as from 3 mm to 8 mm after 72 h of incubation. Thus, the presence zone indicted the presence of extracellular lipase in the isolated bacterial strain.
Identification of bacterial isolates by 16S r RNA gene sequencing
The extracellular lipase producing bacterial strain (HYL) was identified by 16S r RNA gene sequencing method. The 1335-bp long rRNA segment of isolated strain was purified and amplified by Sanger method 3500xL genetic analyser. The gene sequence was submitted to Genbank database of NCBI and was given unique identification number MK835660. The sequence of rRNA segment was analyzed by BLAST which showed 98.48% homology with B. gibsonii T M3R1 (Accession Number MH298508). A dendrogram was prepared by MEGA-X software by employing Tree-joining method. The dendogram involved 14 nearest neighbours of the relevant strain. (Fig. 1). The percent homology of 16S rRNA of our isolate with different nearest neighbour is mentioned in the parenthesis after the name of selected genus and species.
Effect of salt on growth of isolated bacteria
The growth of lipolytic bacterial strain (Fig. 2) was examined under different salt concentrations (50–1200 mM). The growth was monitored by observing the optical density of culture at different time interval. The time-dependent bacterial growth exhibited the highest growth after 120 h and reached to stationary phase at 120 h of incubation. The result exhibited that the growth of isolated strain was increased with higher salt concentration (400–800 mM), showed optimum growth at 600 mM concentration of NaCl. Further increase in NaCl concentration showed declination in growth.
Production of extracellular lipase
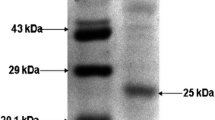
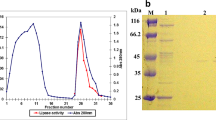
The isolated B. gibsonii produced a high amount of lipase when grown on 1% olive oil as an additional substrate in culture medium incubated at 30 °C at 7.2 pH under shaking condition (150 rpm). The maximum lipase was estimated as 5418.9 U at 30 °C at pH 7.2 from the bacterial culture after 72 h incubation. Further, there was no increase in the lipase production (data not shown). The crude lipase was partially purified by an overnight dialysis process at 4 °C using 70% ammonium sulphate, the obtained lipase showed an increase in the specific activity of enzyme (186.3 U/mg), indicating the enzyme was concentrated by 1.36 fold (Table 1). Kumar et al. (2012), reported that lipase was purified by ammonium sulphate method and specific activity was obtained as 3525.6 U/mg in B. pumilus RK31.
Characterization of the lipase
Effect of pH on lipase activity
Lipase enzyme was assayed under different pH conditions (4.0–11.0). The results revealed that maximum hydrolytic activity of enzyme between pH (8.0–9.0). Whereas the lowest activity was detected at or below pH 7.0 (Fig. 3). An increase in the pH above 9.0 showed a decline in the enzyme activity. Li (Li and Zhang 2005) reported an optimum pH for thermostable lipase from Geobacillus sp. TW1 between pH 7.0–8.0, while Li and Yu (2004) reported that Haloarcula sp. G41 produced extracellular lipase with optimum activity at pH 8.0. Tyndall et al. (2002) have also reported that the purified lipase form B. stearothermophilus P1 exhibited maximum activity at pH 8.5 and at 55 °C temperature. The present investigation revealed that the lipase from B. gobsonii was stable between the pH 8.0 and 9.0, but the enzyme activity registered decline at pH 10.0 and 11.0. However, the enzyme activity in the acidic range below pH 7.0 gradually loses its activity (from 75 to 33%). In the previous report on B. subtilis NS8, the maximum lipase activity was observed at pH 8.0 (Olusesan et al. 2011).
Effect of different metal ions on lipase
The metal ions are considered to be helpful in removing the fatty acids from the oil–water interface. Effect of different metals (1 mM, each) like Ca+2, Ba+2, Mg+2, Al+3, Mn+2, Zn+2, and Co+2 was observed on the lipase activity (Table 2). The result showed that Mg+2, Mn+2, Zn+2 and Co+2 reduced the lipase activity (82.8, 57.9, 86.8 and 61.15%) as compared to control (100%). Surprisingly, addition of Al+3 stimulated the lipase activity by 53.19% over that of control (100%). Sharon et al. (1998) reported that hydrolysis of oil was stimulated in the presence of some metal ions. Goswami and Basu (2012) observed that lipase activity was stimulated by 24%, when the enzyme was treated with Al+3. However, Ba+2 and Ca+2 showed negligible effect (103 and 106%, respectively). Earlier it has been suggested that extracellular lipase activity was uninfluenced in the presence of both Ca+2 and Ba+2 because they form an ion-salts of fatty acids, which are insoluble and prevent product formation (Kambourova et al. 2003). In addition, Ca+2 is known to confer the structural stability to enzymes against denaturation by heat and proteolysis. Rahman et al. (2005) reported that metal ions bind to the enzyme and change its conformational structure, which ultimately leads to effect on the enzyme stability. In previous studied, thermostable lipase from B. stearothermophilus P1 is reported to be partially inhibited by Mg+2, and Mn+2 (Sinchaikul et al. 2001).
Effect of inhibitor and surfactants on enzyme activity
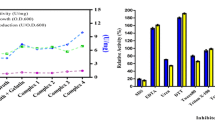
Various enzyme inhibitors and surfactants (1 mM each) were selected to study their effect on partially purified lipase enzyme (Fig. 4). The results showed that extracellular microbial lipase was sensitive to chelating agent EDTA, which inhibited the lipase activity by approx. 55%, whereas the inhibitor-dithiothreitol (DTT) strongly inhibited the enzyme activity (approx. 33%) as compared to control (100%). The Thiol group inhibitor such as 2-mercaptoethanol also inhibited the lipase activity by 40% approx. when compared with control. However, the serine group-specific protein inhibitor PMSF and anionic surfactant SDS worked as weak inducer for the enzyme activity. Both PMSF and SDS stimulated the extracellular lipase activity by approx. 104 and 105%, respectively, as compared to control (100%). However, other non-ionic surfactants Triton X 100, T-20 and T-80 strongly inhibited the lipase activity by approx. 32%, 33% and 37%, respectively, as compared to control. Simil;ar to our findings, Royter et al. (2009) reported that extracellular lipase from Thermoanaerobacter thermohydrosulfuricus SOL1 showed inhibition of the enzyme by EDTA, while surfactants Triton X 100, T-20 and T-80 exhibited inhibitory effect on lipase by approx. 11%, 12% and 22%, respectively.
Effect of temperature on extracellular lipase activity
The enzyme activity was measured under different temperature conditions (Fig. 5). The enzyme solution was pre-incubated at different temperatures (20–80 °C at pH 8.0) for 30 min before the addition of substrate. The result showed temperature dependent increase in the enzyme activity from 20 °C to 60 °C and reached to maximum activity (78.03 U/mL), which was about twofold higher than that recorded at 20 °C. A further increase in the temperature from 60 to 80 °C led to steep decline in the hydrolytic activity of enzyme from 78.03 U/mL to 35.81 U/mL. Similar findings were reported for lipases from Geobacillus sp., Bacillus sp., and Geobacillus thermoleovorans by Li et al. (2005), Nawani and Kaur (2000) and Abdel-Fattah and Gaballa (2008), respectively.
Effect of temperature on kinetic behaviour of lipase enzyme
The kinetic study of extracellular lipase enzyme from B. gibsonii was carried out using at different concentrations of pNPP (0.1–1.5 mM) as substrate under different temperature conditions (10, 30, 60 °C). The Michaelis–Menten saturation kinetics (data not shown), showed a concentration dependent increase in the rate of pNP-palmitate hydrolysis with rate saturation at 1.0 mM concentration under all the temperature conditions. A further increase in the substrate concentration, the rate of pNPP hydrolysis declined under all the temperature conditions (10, 30 and 60 °C). A reciprocal Lineweaver–Burk (Lineweaver and Burk 1934) plot (Fig. 6) of the data, derived from the Michaelis–Menten saturation kinetics, was used to calculate the kinetic constants such as Km, Vmax, and kcat. (Table 3). The Km values calculated for the enzyme activity at 10, 30 and 60 °C were found to be 0.77, 0.65 and 0.53 mM, respectively. The Vmax values of lipase enzyme at 10, 30 and 60 °C were found to be 75.18, 120.48 and 158.73 U/mL, respectively. Noormohamadi et al. (2013) studied Pseudomonas sp. lipase and the Km and Vmax values were reported as 0.77 mM and 49.5 U/ml, respectively.
As evident from the Km and Vmax values of the enzyme, the maximum efficiency of the enzyme activity was observed at 60 °C as evident from lower value of Km and higher Vmax values. A decrease or increase in the temperature beyond the optimum, temperature led to a change in the Km and Vmax values. The kcat (turn over number) values calculated under all the temperature conditions (10, 30, 60 °C) showed that the highest kcat value (483.93 min −1) for the enzyme optimum at 60 °C, indicating higher turnover of the lipase enzyme at this temperature. The catalytic efficiency constant (kcat/Km) at 60 °C was found to be the best (913.07), denoting the highest frequency of enzyme–substrate encounter at 60 °C. The activation energy (Ea) of enzyme is related to that energy which is required to initiate the reaction between enzyme and substrate. It was calculated as 45.28 kJ/mol from Arrhenius plot was (Fig. 7). These kinetic constants of enzyme suggested that the enzyme was thermostable with its best efficiency and activity at 60 °C.
Thermal stability of lipase
The thermal stability of lipase enzyme was presented in terms of percentage residual activity with respect to time of exposure to different temperature (60, 70 and 80 °C). The initial enzyme activity at 0 min was taken as control (100%). During the first 15 min of incubation at 60, 70 and 80 °C, the enzyme retained its activity to the extent of 97, 71 and 52%, respectively, as compared to control (Fig. 8). After 30 min of incubation of enzyme at respective enzyme denaturating temperature, the enzyme activity declined to the level of 95, 60 and 30% at 60.70 and 80 °C, reactively. After 90 min of incubation of the enzyme at 60, 70 and 80 °C, the residual activity declined to the level of 89, 50 and 24%, respectively, as compared to control (100%, 0 min.). The lipase enzyme after 120 min. at 60 °C showed residual activity to the level of 88%, while the residual activity at 70 and 80 °C, was substantially reduced to the level of 41 and 10%, respectively. Olusesan et al. (2011) and Kim (1994) showed that extracellular lipase from B. thermoleovorans ID-1A and Bacillus strain incubated at 60 °C were able to retain about 75% and 50% of activity of the enzymes from the respective strain. The thermostable lipase of G. thermodenitrificans IBRL-nra was able to retain 87.5% of its initial enzyme activity after 15 min of temperature treatment at 70 °C as reported by Balan et al. (2012). The thermal inactivation rate constant (kd) for the lipase from B. gibsonii was calculated by linear regression of the slope obtained by percentage residual against time (min.). The result (Fig. 8) showed that the B. gibsonii lipase followed a first-order reaction kinetic model. The steady inactivation of lipase enzyme occurred with an increase in temperature (60–80 °C). It was observed that the kd value increased from 0.09 min−1 at 60 °C to 0.182 min−1 at 80 °C( Table 3).Whitaker (Whitaker 1994) reported that the inactivation rate constant (kd) is inversely proportional to the stability of the enzyme. Therefore, the lowest kd value obtained at 60 °C further justified the thermostable nature of lipase from B. gibsonii.
The thermal inactivation energy [Ea(d)] constant denotes an amount of minimum energy required to start the enzyme denaturation. The [Ea(d)] calculated as 66.98 kJ/mol, from the slope of the plot of ln kd versus 1/T (temperature) (Fig. 9). The higher value of inactivation energy indicated the larger temperature change needed for enzyme inactivation (Segel 1976). Therefore, the results suggested that thermal inactivation of B. gibsonii lipase required higher inactivation energy. Kumari et al. (2012) have reported that inactivation energy of thermostable lipase LIP8, LIP14 and LIP18, from Yarrowia lipolytica were 64.6, 56.1 and 66.5 kJ/mol, respectively, which suggested that thermostable enzymes required larger change in the temperature for inactivation of the enzyme. Olusesan et al. (2011) also observed that the thermal inactivation energy for lipase from B. subtilis NS 8 was 76.0 kJ/mol.
Hydrophobic interactions are considered to play important role in the stabilization of the native conformation of proteins, but not in the unfolded protein (Kristjansson et al. 1991). During temperature treatment of the enzyme, the enzyme stability is determined by change in Gibbs free energy (ΔG*). The value of ΔG* calculated at 60 °C was 86.31 kJ/mol (Table 3), which was lower than that recorded at 80 °C (1.89 kJ/mol). The closely related result was reported by Owusu (Owusu et al. 1992) in Pseudomonas fluorescens P38 lipase, where Gibb's free energy was obtained in the range of 103–105 kJ/mol between 60 and 80 °C. Adams et al. (1981) reported the Gibb’s free energy (ΔG*) for lipase enzyme from a heat-resistant Pseudomonas sp. to be in the range of 112–121 kJ/mol.
The enthalpy (ΔH*) of a reaction is defined as an amount of heat absorbed or released during an enzyme reaction that occurs at constant pressure, which also represents the minimum energy barrier that must be overcome by the reacting molecules (Ariahu et al. 2000). The values of enthalpy for lipase reactions taking place at from 60 to 80 °C were calculated to be 64.26 and 64.04 kJ/mol, respectively. Similar results on enthalpy (59.56–59.47 kJ/mol) were observed in Cellulomas flavigena UNP3 sp. lipase enzyme (Prajapati et al. 2014) between 60 and 70 °C, which was significantly lower than that obtained for the lipase from B. gibsonii under similar set of temperatures. Entropy (ΔS*) is defined as the structural alteration in the disorderliness of enzyme upon protein folding. There is a direct correlation between enthalpy and entropy in terms of enzyme stability as both the parameters denote the number of non-covalent bonds broken during the enzyme reaction and net change in the enzyme-solvent disorder (Ortega et al. 2004). The entropy values of the lipase from B. gibsonii at 60 and 80 °C were − 66.21 × 10–3 kJ/mol/K and − 78.8 × 10–3 kJ/mol/K, respectively (Table 3). Both the enthalpy and entropy values obtained for lipase of B. gibsonii proved that the enzyme was more stable at 60 °C. A negative value of the entropy for irreversible thermal inactivation of B. gibsonii indicated that the enzyme molecules at the high temperature probably involve an aggregation of partially unfolded enzyme as suggested earlier (Adams et al. 1981).
During thermal inactivation process, the half-life of an enzyme is the time required by the enzyme to operate at half of the initial rate of activity. The half-lives of lipase from B. gibsonii was calculated as 76.40, 58.23 and 38.07 min during temperature treatment at 60, 70 and 80 °C, respectively (Table 4). Sharma et al. (2002) and Kambourova et al. (2003) reported the half-lives of thermostable lipases from Bacillus sp. RSJ-1 and B. stearothermophilus MC 7 as 150 min (60 °C) and 180 min (70 °C), respectively. Kim et al. (1994, 1998) reported the half-lives of lipase from Bacillus sp. strain 398 and B. stearothermophilus was 30 min at 65 and 62 °C, respectively The present result on half-life of the enzyme suggested that the lipase from B. gibsonii was more thermo-tolerant at temperature of 60 °C.
Conclusion
The present investigation relates to the kinetic and thermodynamic evaluation of extracellular lipase produced by halo-tolerant bacterial strain B. gibsonii. The study revealed that maximum production of lipase by the bacterial strain at 60 °C. The enzyme activity showed its optimal activity at 60 °C and pH 9.0. The enzyme activity was stimulated in the presence of surfactant SDS, protein inhibitor PMSF, EDTA and Al+3. Kinetic parameters (Km, Vmax and kcat), thermodynamic parameters and thermal inactivation constants for the enzyme treated under different temperatures (10, 30, 60 °C) exhibited that the activity of lipase enzyme from B. gibsonii was spontaneous and kinetically favourable up to 60 °C. A further increase in temperature led to gradual irreversible inactivation. Therefore, it was concluded that the alkaline extracellular lipase from B. gibsonii was a thermostable enzyme, which exhibits great potential to be used for commercial purpose by various industries.
References
Abdel-Fattah YR, Gaballa AA (2008) Identification and over-expression of a thermostable lipase from Geobacillus thermoleovorans Toshki in Escherichia coli. Microbio res 163:13–20. https://doi.org/10.1016/j.micres.2006.02.004
Adams DM, Brawley TG (1981) Factors influencing the heat resistance of a heat-resistant lipase of Pseudomonas. J Food Sci 46:673–676. https://doi.org/10.1111/j.1365-2621.1981.tb15321.x
Ariahu CC, Ogunsua AO (2000) Thermal degradation kinetics of thiamine in periwinkle based formulated low acidity foods. Int J Food Sci Technol 35:315–321. https://doi.org/10.1046/j.1365-2621.2000.00366.x
Balan A, Ibrahim D, Abdul Rahim R, Ahmad Rashid FA (2012) Purification and characterization of a thermostable lipase from Geobacillus thermodenitrificans IBRL-nra. Enzyme Res. https://doi.org/10.1155/2012/987523
Bora L, Kalita MC (2008) Production of thermostable alkaline lipase on vegetable oils from a thermophilic Bacillus sp. DH4, characterization and its potential applications as detergent additive. J Chem Technol Biotechnol 83:688–693. https://doi.org/10.1002/jctb.1853
Ertuğrul S, Dönmez G, Takaç S (2007) Isolation of lipase producing Bacillus sp. from olive mill wastewater and improving its enzyme activity. J Hazard Mater 149:720–724. https://doi.org/10.1016/j.jhazmat.2007.04.034
Gandhi NN (1997) Applications of lipase. J Am Oil Chem Soc 74:621–634. https://doi.org/10.1007/s11746-997-0194-x
Goswami D, De S, Basu JK (2012) Effects of process variables and additives on mustard oil hydrolysis by porcine pancreas lipase. Braz J Chem Eng 29:449–460. https://doi.org/10.1590/S0104-66322012000300002
Griebeler N, Polloni AE, Remonatto D, Arbter F, Vardanega R, Cechet JL, Di Luccio M, de Oliveira D, Treichel H, Cansian RL, Rigo E (2011) Isolation and screening of lipase-producing fungi with hydrolytic activity. Food Bioproc Tech 4:578–586. https://doi.org/10.1007/s11947-008-0176-5
Griffiths MW, Phillips JD, Muir DD (1981) Thermostability of proteases and lipases from a number of species of psychrotrophic bacteria of dairy origin. J Appl Microbiol 50:289–303. https://doi.org/10.1111/j.1365-2672.1981.tb00894.x
Hasan F, Shah AA, Hameed A (2006) Industrial applications of microbial lipases. Enzyme Microb Technol 39:235–251. https://doi.org/10.1016/j.enzmictec.2005.10.016
Hiol A, Jonzo MD, Druet D, Comeau L (1999) Production, purification and characterization of an extracellular lipase from Mucor hiemalis f. hiemalis. Enzyme Microb Technol 25:80–87. https://doi.org/10.1016/S0141-0229(99)00009-5
Houde A, Kademi A, Leblanc D (2004) Lipases and their industrial applications. Appl Biochem Biotechnol 118:155–170
Jaenicke R (1991) Protein stability and molecular adaptation to extreme conditions. In: Christen P, Hofmann E (eds) EJB Reviews, vol 1991. Springer, Heidelberg
Kambourova M, Kirilova N, Mandeva R, Derekova A (2003) Purification and properties of thermostable lipase from a thermophilic Bacillus stearothermophilus MC 7. J. Mol. Catal. B Enzym 22:307–313. https://doi.org/10.1016/S1381-1177(03)00045-6
Kim HK, Sung MH, Kim HM, Oh TK (1994) Occurrence of thermostable lipase in thermophilic Bacillus sp. strain 398. Biosci Biotechnol Biochem 58:961–962. https://doi.org/10.1271/bbb.58.961
Kim HK, Park SY, Lee JK, Oh TK (1998) Gene cloning and characterization of thermostable lipase from Bacillus stearothermophilus L1. Biosci Biotechnol Biochem 62:66–71. https://doi.org/10.1271/bbb.62.66
Kim MH, Kim HK, Lee JK, Park SY, Oh TK (2000) Thermostable lipase of Bacillus stearothermophilus: high-level production, purification, and calcium-dependent thermostability. Biosci Biotechnol Biochem 64:280–286. https://doi.org/10.1271/bbb.64.280
Kristjansson MM, Kinsella JE (1991) Protein and enzyme stability: structural, thermodynamic, and experimental aspects. Adv Food Nutr Res 35:237–316. https://doi.org/10.1016/S1043-4526(08)60066-2
Kumar R, Sharma A, Kumar A, Singh D (2012) Lipase from Bacillus pumilus RK31: production, purification and some properties. World Appl Sci J 16:940–948
Kumari A, Gupta R (2012) Extracellular expression and characterization of thermostable lipases, LIP8, LIP14 and LIP18, from Yarrowia lipolytica. Biotechnol Lett 34:1733–1739. https://doi.org/10.1007/s10529-012-0958-8
Lee DW, Koh YS, Kim KJ, Kim BC, Choi HJ, Kim DS, Suhartono MT, Pyun YR (1999) Isolation and characterization of a thermophilic lipase from Bacillus thermoleovorans ID-1. FEMS Microbiol Lett. 179:393–400. https://doi.org/10.1111/j.1574-6968.1999.tb08754.x
Li X, Yu HY (2014) Characterization of an organic solvent-tolerant lipase from Haloarcula sp. G41 and its application for biodiesel production. Folia Microbiol 59:455–463. https://doi.org/10.1007/s12223-014-0320-8
Li H, Zhang X (2005) Characterization of thermostable lipase from thermophilic Geobacillus sp. TW1. Protein Expr Purif 42:153–159. https://doi.org/10.1016/j.pep.2005.03.011
Lineweaver H, Burk D (1934) The determination of enzyme dissociation constants. J Am Chem Soc 56(3):658–666. https://doi.org/10.1021/ja01318a036
Lowry RR, Tinsley IJ (1976) Rapid colorimetric determination of free fatty acids. J Am Oil Chem Soc 53:470–472. https://doi.org/10.1007/BF02636814
Nawani N, Kaur J (2000) Purification, characterization and thermostability of lipase from a thermophilic Bacillus sp. J33. Mol Cell Biochem 206:91–96. https://doi.org/10.1023/A:1007047328301
Noormohamadi R, Tabandeh F, Shariati P, Otadi M (2013) Characterization of a lipase from a newly isolated Pseudomonas sp. Iran J Microbiol 5:422
Olusesan AT, Azura LK, Forghani B, Bakar FA, Mohamed AK, Radu S, Manap MY, Saari N (2011) Purification, characterization and thermal inactivation kinetics of a non-regioselective thermostable lipase from a genotypically identified extremophilic Bacillus subtilis NS 8. N Biotechnol 28:738–745. https://doi.org/10.1016/j.nbt.2011.01.002
Ortega N, De Diego S, Perez-Mateos M, Busto MD (2004) Kinetic properties and thermal behaviour of polygalacturonase used in fruit juice clarification. Food chem 88:209–217. https://doi.org/10.1016/j.foodchem.2004.01.035
Owusu RK, Makhzoum A, Knapp JS (1992) Heat inactivation of lipase from psychrotrophic Pseudomonas fluorescens P38: activation parameters and enzyme stability at low or ultra-high temperatures. Food chem 44:261–268. https://doi.org/10.1016/0308-8146(92)90048-7
Prajapati V, Patel H, Trivedi U, Patel K (2014) Kinetic and thermodynamic characterization of lipase produced by Cellulomonas flavigena UNP3. J Basic Microbiol 54:976–983. https://doi.org/10.1002/jobm.201300065
Rahman RN, Baharum SN, Basri M, Salleh AB (2005) High-yield purification of an organic solvent-tolerant lipase from Pseudomonas sp. strain S5. Anal Biochem 341:267–274. https://doi.org/10.1016/j.ab.2005.03.006
Royter M, Schmidt M, Elend C, Höbenreich H, Schäfer T, Bornscheuer UT, Antranikian G (2009) Thermostable lipases from the extreme thermophilic anaerobic bacteria Thermoanaerobacter thermohydrosulfuricus SOL1 and Caldanaerobacter subterraneus subsp. tengcongensis. Extremophiles 1:769–783. https://doi.org/10.1007/s00792-009-0265-z
Sarmah N, Revathi D, Sheelu G, Yamuna Rani K, Sridhar S, Mehtab V, Sumana C (2018) Recent advances on sources and industrial applications of lipases. Biotechnol Prog 34:5–28. https://doi.org/10.1002/btpr.2581
Schmidt-Dannert C, Rua ML, Atomi H, Schmid RD (1996) Thermoalkalophilic lipase of Bacillus thermocatenulatus. I. Molecular cloning nucleotide sequence purification and some properties. BBA 1301:105–114. https://doi.org/10.1016/0005-2760(96)00027-6
Segel IH (1976) Enzymes. Biochemical calculations 2nd 447 edn. John Wiley and Sons, pp 208–323.
Sharma R, Soni SK, Vohra RM, Gupta LK, Gupta JK (2002) Purification and characterisation of a thermostable alkaline lipase from a new thermophilic Bacillus sp. RSJ-1. Process Biochem 37:1075–1084. https://doi.org/10.1016/S0032-9592(01)00316-8
Sharon C, Nakazato M, Ogawa HI, Kato Y (1998) Lipase-induced hydrolysis of castor oil: effect of various metals. J Ind Microbiol Biotechnol 21:292–295. https://doi.org/10.1038/sj.jim.2900586
Sinchaikul S, Sookkheo B, Phutrakul S, Pan FM, Chen ST (2001) Optimization of a thermostable lipase from Bacillus stearothermophilus P1: overexpression, purification, and characterization. Protein Expr Purif 22:388–398. https://doi.org/10.1006/prep.2001.1456
Sugihara A, Tani T, Tominaga Y (1991) Purification and characterization of a novel thermostable lipase from Bacillus sp. J Biochem 109:211–216. https://doi.org/10.1093/oxfordjournals.jbchem.a123363
Treves C, Vincenzini MT, Favilli F, Vanni P, Baccari V (1984) On the interaction between synthetic detergents and enzymatic proteins. Canadian J Biochem Cell Biol 62:55–59. https://doi.org/10.1139/o84-009
Tyndall JD, Sinchaikul S, Fothergill-Gilmore LA, Taylor P, Walkinshaw MD (2002) Crystal structure of a thermostable lipase from Bacillus stearothermophilus P1. J Mol Biol 323:859–869. https://doi.org/10.1016/S0022-2836(02)01004-5
Wang Y, Srivastava KC, Shen GJ, Wang HY (1995) Thermostable alkaline lipase from a newly isolated thermophilic Bacillus strain A30–1 (ATCC 53841). J Ferment Bioeng 79:433–438. https://doi.org/10.1016/0922-338X(95)91257-6
Whitaker JR (1993) Principles of enzymology for the food sciences. CRC press
Acknowledgements
The authors are thankful to University Grant Commission (UGC), New Delhi, Government of India for providing financial support for the research work. Author is obliged to The Head, Department of Environmental Science, Babasaheb Bhimrao Ambedkar University, Lucknow (India) for providing the laboratory facilities during the investigation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that this manuscript does not have any conflicts of interest regarding the financial, scientific contents and otherwise.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sonkar, K., Singh, D.P. Kinetic and thermodynamic characterization of novel alkaline lipase from halotolerant Bacillus gibsonii. Arch Microbiol 203, 2199–2209 (2021). https://doi.org/10.1007/s00203-021-02197-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02197-7