Abstract
Key message
Four QTLs for adult-plant resistance to powdery mildew were mapped in the Zhou8425B/Chinese Spring population, and a new QTL on chromosome 3B was validated in 103 wheat cultivars derived from Zhou8425B.
Abstract
Zhou8425B is an elite wheat (Triticum aestivum L.) line widely used as a parent in Chinese wheat breeding programs. Identification of genes for adult-plant resistance (APR) to powdery mildew in Zhou8425B is of high importance for continued controlling the disease. In the current study, the high-density Illumina iSelect 90K single-nucleotide polymorphism (SNP) array was used to map quantitative trait loci (QTL) for APR to powdery mildew in 244 recombinant inbred lines derived from the cross Zhou8425B/Chinese Spring. Inclusive composite interval mapping identified QTL on chromosomes 1B, 3B, 4B, and 7D, designated as QPm.caas-1BL.1, QPm.caas-3BS, QPm.caas-4BL.2, and QPm.caas-7DS, respectively. Resistance alleles at the QPm.caas-1BL.1, QPm.caas-3BS, and QPm.caas-4BL.2 loci were contributed by Zhou8425B, whereas that at QPm.caas-7DS was from Chinese Spring. QPm.caas-3BS, likely to be a new APR gene for powdery mildew resistance, was detected in all four environments. One SNP marker closely linked to QPm.caas-3BS was transferred into a semi-thermal asymmetric reverse PCR (STARP) marker and tested on 103 commercial wheat cultivars derived from Zhou8425B. Cultivars with the resistance allele at the QPm.caas-3BS locus had averaged maximum disease severity reduced by 5.3%. This STARP marker can be used for marker-assisted selection in improvement of the level of powdery mildew resistance in wheat breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat is one of the most important staple crops, but its production is constrained by many biotic and abiotic factors. Powdery mildew, caused by Blumeria graminis f. sp. tritici (Bgt), is a devastating rapidly spreading fungal disease that affects all aerial plant parts including stems, leaves, glumes, and awns. Powdery mildew prevails in many wheat growing regions of eastern Asia, southeastern USA, northeastern Africa, and northern Europe (Saari and Wilcoxson 1974; Roelfs 1977; Selter et al. 2014). The severity and prevalence of powdery mildew has increased in recent decades due to increasing applications of nitrogen fertilizer and irrigation (Olesen et al. 2000). Since the 1980s, this disease has become widespread in most wheat growing regions of China (Li and Zeng 2002) where it affects around 8 million ha of wheat annually (Zhao et al. 2013). In comparison to chemical control, the use of resistant cultivars is a more comprehensive, economical, and environmentally friendly approach to control the disease (Petersen et al. 2015).
To date, 85 powdery mildew resistance genes (Pm1–Pm58) at 54 loci have been catalogued in wheat (Hao et al. 2015; Liu et al. 2017; McIntosh et al. 2017; Wiersma et al. 2017). These genes were derived from 20 Triticeae species including Triticum aestivum, T. monococcum, T. dicoccum, T. spelta, and Aegilops speltoides; 21 loci were from T. aestivum. Many of these genes are race-specific and the majority of them have lost effectiveness. However, adult-plant resistance genes, such as Lr34/Yr18/Pm38, Lr46/Yr29/Pm39, and Lr67/Yr46/Pm46, continue to confer race-non-specific resistance to powdery mildew (Lillemo et al. 2008; Herrera-Foessel et al. 2014; Moore et al. 2015), although levels of protection are often less than adequate when such genes are deployed alone and they do not control the disease at early growth stages.
With recent developments in genotyping arrays, such as the high-density wheat 90K SNP chip platform (Wang et al. 2014), SNP markers are being used increasingly in genetic mapping. Due to co-dominance, abundance, and even distribution of SNP across the genome, high-density genome-wide genotyping arrays further improve the construction of high-resolution genetic maps and QTL mapping. In recent years, QTL of different traits were identified from the whole genome using the wheat 90K iSelect array, e.g., for wheat quality (Colasuonno et al. 2014), flag leaf and grain (Wu et al. 2015a, b, 2016), yield and related traits (Gao et al. 2015), and disease resistance (Liu et al. 2016; Zhang et al. 2017). In addition, the SNPs from arrays can be transferred into KASP (Kompetitive allele-specific PCR, Semagn et al. 2014; Thomson 2014) or STARP (Long et al. 2017) markers that can be easily used in marker-assisted selection (MAS). It is expected that the STARP technique, with the major advantages of simple assay design, flexible throughputs, high accuracy, platform compatibility, and low operational costs, will be applied increasingly in MAS and genetic mapping (Long et al. 2017).
Wheat cultivar Zhou8425B developed by the Zhoukou Academy of Agricultural Sciences in Henan province has good resistance to stripe rust, leaf rust, and powdery mildew (http://wheatpedigree.net/sort/show/92642, ZHOUMAI-8425-B in CIMMYT Genebank). It carries all-stage resistance genes YrZH84 (Li et al. 2006) and LrZH84 (Zhao et al. 2008), and 4 leaf rust APR QTL (Zhang et al. 2017). Zhou8425B is an elite wheat cultivar and has been widely used as a parent in breeding programs in the Yellow and Huai Valleys Autumn-sown Wheat Zone since 1984. More than 100 cultivars derived from this line have been grown on an accumulated area of more than 33 million ha in China during the past 20 years. Although stripe rust and leaf rust resistance genes in Zhou8425B were identified previously (Li et al. 2006; Zhao et al. 2008; Yin et al. 2009; Xiao et al. 2011; Zhang et al. 2017), resistance to powdery mildew has not been studied. The objective of the present study was to identify QTL for APR to powdery mildew in Zhou8425B, validate a major QTL on chromosome 3B, and develop a tightly linked STARP marker for MAS in wheat breeding.
Materials and methods
Plant materials
The 244 F8 recombinant inbred lines (RILs) derived from a cross between Zhou8425B and Chinese Spring were used for construction of a whole-genome high-density linkage map and QTL mapping. One hundred and three cultivars derived from Zhou8425B were used for QTL validation. The highly susceptible wheat cultivar Jingshuang 16 was used as the susceptible control. The resistant wheat line Zhou8425B will be available in CIMMYT Genebank from March, 2018 on (http://wheatpedigree.net/sort/show/92642), and it is also available in the Genebank of Chinese Academy of Agricultural Sciences by the accession number ZM29072 (http://www.cgris.net).
Powdery mildew tests in the greenhouse
Seedlings of 244 F8 RILs and parents were tested for powdery mildew response in the greenhouse following inoculation with prevalent Bgt isolates E09 and E20 obtained from the Plant Protection Institute, CAAS. About 15 seeds of each RIL were planted, and the infection types (IT) based on a 0–4 scale (Liu et al. 1999) were scored 10 days after inoculation when the susceptible control Jingshuang16 showed severe symptoms.
Powdery mildew evaluation in the field
The RILs and parents were evaluated for powdery mildew response at the adult–plant stage in the field at Beijing and Zhengzhou during the 2014–2015 and 2015–2016 cropping seasons following artificial inoculation. The field trials were conducted in randomized complete blocks with three replicates. Each plot consisted of a single 1.5 m row with 20 cm between rows. Approximately 50 seeds were uniformly sown in each row. The highly susceptible cultivar Jingshuang 16 was planted at every tenth row as a control, and also perpendicular and adjacent to the test rows as a spreader for even infection.
The highly susceptible cultivar Jingshuang 16 was planted in 9 cm × 9 cm (diameter × height) plastic pots and inoculated with the virulent isolate E20 at the two-leaf stage in the greenhouse; the fully infected Jingshuang 16 seedlings were then transplanted among the spreader lines with one pot every 50 cm in the field at the stem elongation stage by the end of March. Disease severity on each line was scored as the average percentage of leaf area covered by powdery mildew. The first assessment occurred when the susceptible control Jingshuang 16 was severely diseased about 6 weeks after inoculation. The maximum disease severity (MDS) for each line was evaluated when the disease severity on the control reached a maximum level around 1 week later, which was used for subsequent analysis.
To validate the effect of the new QTL on chromosome 3B, 103 cultivars derived from crosses involving Zhou8425B were planted at Beijing, and at Zhengzhou and Xingyang in Henan province during the 2016–2017 cropping season. The field trials and powdery mildew evaluations in the field were similar to those described above.
Statistical analysis
Phenotypic correlation coefficients, analysis of variance (ANOVA), and t tests were conducted using SAS 9.4 software (SAS Institute, Cary, NC). Broad-sense heritability (H2) of powdery mildew response was calculated following Nyquist and Baker (1991).
QTL analysis
Genomic DNA were extracted from healthy seedling leaves of the RILs and parents by the CTAB method (Saghai-Maroof et al. 1984). Molecular genotyping was performed using the wheat 90K iSelect SNP array and genetic linkage maps were constructed. Twenty-one linkage groups corresponding to all 21 hexaploid wheat chromosomes were assembled from 5636 high-quality polymorphic SNP markers (Gao et al. 2015). QTL analysis was performed by the inclusive composite interval mapping with the ICIM-ADD function using the software QTL IciMapping 4.1 (Li et al. 2007; Meng et al. 2015). Phenotypic values of RILs averaged from three replicates in each environment and BLUEs (best linear unbiased estimates) values of the genetic effects from four environments of RILs by R package lme4 (Bates et al. 2015) were used for QTL detection. QTL were mapped at a logarithm of odds (LOD) threshold of 2.5 based on 1000 permutations and a walk speed of 1.0 cM, with P = 0.001 in stepwise regression. QTL effects were estimated as the proportion of phenotypic variance explained (PVE) by the QTL. Normally, there are minor differences in the peaks of LOD contours for a single QTL across different environments. The QTL within one-log support confidence interval (14 cM) were considered to be the same.
STARP marker design
The STARP marker designed from an SNP tightly linked with the 3B QTL included five primers: two AMAS-primers (asymmetrically modified allele-specific primers) for specifically amplifying two alleles from genomic DNA to provide priming sites for PEA-primers by six touchdown cycles, two universal PEA-primers (priming element-adjustable primers) amplifying with the AMAS amplification products to distinguish two alleles by gel-free fluorescence signals, and their common reverse primer designed from information in the GSP website (http://probes.pw.usda.gov/GSP/index.php). PCR procedures and conditions followed Long et al. (2017). Gel-free fluorescence signals scanning and allele separation were conducted by microplate reader (Multiscan Spectrum BioTek, Synegy/H1) with the Klustercaller 2.24.0.11 software (LGC, Hoddesdon, UK).
Results
Seedling reactions to powdery mildew in the greenhouse
Seedlings of control cultivar Jingshuang 16 were susceptible (IT 3–4) to powdery mildew. Zhou8425B and Chinese Spring were also susceptible to E09 and E20 (IT 3–4). Among the 244 RILs, 243 exhibited susceptible infection types (IT 3–4), whereas one was highly resistant. This line was considered to be a contaminant and was excluded from subsequent analysis.
Powdery mildew scores in the field
The MDS of the 243 RILs ranged from 0 to 77% across four environments, indicating significant differences among genotypes. Zhou8425B and Chinese Spring exhibited averaged MDS scores of 5 and 7%, respectively. Pearson’s correlation coefficients for the population ranged from 0.53 to 0.73 among four environments (P < 0.01) (Table S1). The frequency distribution of powdery mildew MDS in each environment with a coefficient of variation of 63.6% showed a continuous distribution skewed toward resistance (Figure S1), indicating polygenic inheritance and transgressive segregation. Broad-sense heritability of MDS across the four environments was 0.80. ANOVA confirmed significant variation among genotypes, environments, and genotype × environment interactions (Table S2), demonstrating both genotypic and environmental influences on these traits.
The MDS of 103 derivatives of Zhou8425B were 0–70, 0–50, and 1–70% in Beijing, Xingyang, and Zhengzhou, respectively, with correlation coefficients ranging from 0.45 to 0.58 among three environments.
QTL for powdery mildew resistance
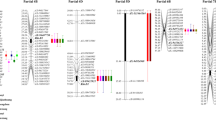
QPm.caas-1BL.1 in marker interval IWB72835-IWB18787 identified at Beijing 2016, Zhengzhou 2016, and the averaged value of four environments explained 5.44, 5.69, and 7.20% of the phenotypic variances, respectively (Fig. 1; Table 1). The additive effects were − 2.33, − 3.22, and − 2.38, respectively. The resistance allele was derived from Zhou8425B.
QPm.caas-3BS in marker interval IWB21064-IWB64002 was stably detected in all environments and averaged values, explaining 4.36–9.05% of the phenotypic variances, with additive effects ranging between − 1.51 and − 3.80. The resistance allele was from Zhou8425B.
QPm.caas-4BL.2 located in the region IWB35851-IWB60096 explained 6.43 and 8.77% of the phenotypic variance in Zhengzhou 2016 and the average for four environments, with additive effects of − 3.43 and − 2.63, respectively. The resistance allele was from Zhou8425B.
QPm.caas-7DS in marker interval IWB41108–IWB53819 was identified in Beijing 2015, Zhengzhou 2015, Beijing 2016, and the average of four environments, explaining 10.37, 5.20, 4.10, and 4.21% of the phenotypic variances, respectively. The additive effects were 4.60, 1.15, 2.05, and 1.84, respectively. The resistance allele was contributed by Chinese Spring.
Validation of QPm.caas-3BS in Zhou8425B derivatives
QPm.caas-3BS with the resistance allele from Zhou8425B was an important APR QTL for powdery mildew resistance, exhibiting a stable effect in all environments. We transferred SNP IWB41105 that was closely linked to QPm.caas-3BS into an STARP marker named Str-IWB41105 (Table 2), and tested 103 Zhou8425B derivatives (Table S3). Student’s t tests indicated that varieties with the resistance allele significantly (P < 0.05) reduced the average MDS of all environments by 5.3% (Table 3).
Discussion
Comparisons with previous reports
QPm.caas-1BL.1
There are several reports of QTL for APR to powdery mildew mapped on chromosome 1BL. These include QPmvt-1BL and QPm.vt-1B detected in the North American winter wheat cultivar Massey and derived line USG 3209, respectively (Liu et al. 2001; Tucker et al. 2007). The pleiotropic APR gene Lr46/Yr29/Pm39 in CIMMYT bread wheat line Saar was at a similar position to the above two QTL (Lillemo et al. 2008), and they all shared the closely linked simple sequence repeat (SSR) locus Xbarc80 (Tucker et al. 2007; Lillemo et al. 2008). Nevertheless, our experiment indicated that parents Zhou8425B and Chinese Spring did not have Lr46/Yr29/Pm39 as tested by the cleaved amplified polymorphic sequences (CAPS) marker csLV46G22 (Figure S2), and this pleiotropic QTL was also not previously detected in the population for leaf rust resistance (Zhang et al. 2017). One of the parents, Zhou8425B (donor of QPm.caas-1BL.1), has winter growth habit which is different from Saar (donor of Pm39) having spring growth habit. Therefore, it is likely that QPm.caas-1BL.1 is different from Pm39.
QPm.caas-3BS
Powdery mildew APR QTL QPm.inra-3B linked with Xgwm389 and Xbarc133 and derived from French semi-dwarf cv. Courtot was identified in one environment (Bougot et al. 2006). One major pleiotropic gene Sr2 on chromosome 3BS was tightly linked with pseudoblack chaff (Kota et al. 2006), Lr27 (Singh and McIntosh 1984) and a powdery mildew resistance gene (Mago et al. 2011b). QPm.caas-3BS is different from the powdery mildew gene co-segregating with Sr2, because neither the RILs nor parents have pseudoblack chaff and tests with the CAPS marker csSr2 that co-segregates with Sr2 also indicated the absence of Sr2 (Figure S3) (Mago et al. 2011a). Moreover, there were no evidence that the Hope or H-44 derivatives of Yaroslaw emmer (T. dicoccoides) (McFadden 1930) or a derivative was involved in the pedigree of Zhou8425B. The proximal SNP markers BobWhite_c9711_71 (ID: IWB4653) and Excalibur_c6330_1158 (ID: IWB28189) of QLr.hebau-3BS (Zhang et al. 2017) were located at 53–55 cM (Zhang et al. 2017) in the same region as QPm.caas-3BS. Moreover, QLr.hebau-3BS (in a 14.4 cM interval Xbarc147–Xgwm493) and QPm.inra-3B (16.9 cM from Xgwm389) are in different chromosomal bins (Sourdille et al. 2004). It is thus concluded that QPm.caas-3BS is likely a new APR QTL that is pleiotropic with a QTL for leaf rust resistance (Zhang et al. 2017).
QPm.caas-4BL.2
Five QTL for APR to powdery mildew were identified near the chromosome 4B centromere in previous studies. QPm.ipk-4B and QPm.sfr-4B in the synthetic hexaploid line W7984 and Swiss spelt cv. Oberkulmer were detected in RFLP (restriction fragment length polymorphism) marker intervals Xcdo795–Xbcd1262 and Xpsr593b–Xpsr1112 (Keller et al.1999; Börner et al. 2002). QPm.caas-4BL in Israeli wheat Oligoculm was mapped in SSR interval Xgwm375–Xgwm251 (Liang et al. 2006). QPm.nuls-4BL in wheat line Avocet was between DArT (Diversity arrays technology) marker XwPt1505 and SSR marker Xgwm149 (Lillemo et al. 2008). QPm.caas-4BL.1 located in interval Xgwm149–Xgwm495 in Italian cv. Libellula explained an average PVE of 14.7% (Asad et al. 2012). Xgwm149, Xgwm375, and the SNP marker BS00109813_51 (ID: IWB12434) were adjacent according to the wheat map in Zhang et al. (2017). In the present genetic map, QPm.caas-4BL.2 was flanked by markers IWB35851 and IWB60096 located at 39.97–42.20 cM and closely linked with IWB12434 at position 37.89 cM. Because QPm.ipk-4B and QPm.sfr-4B were mapped only with RFLP markers, it is difficult to compare the locations with our QTL. QPm.caas-4BL.2 was mapped in a similar location to the latter three QTL and, hence, represents a same gene.
QPm.caas-7DS
Chromosome 7DS harbors the multi-pathogen resistance gene Lr34/Yr18/Pm38 that has shown durable resistance for more than 80 years (Krattinger et al. 2009); this gene stimulates senescence-like processes in the flag leaf tips and edges, leading to tip necrosis and functioning resistance in the adult plant (Singh 1992; Kolmer et al. 2008). Six other QTL were also detected in the position of Pm38, including QPm.ipk-7D, QPm.inra-7D.1, Qaprpm.cgb-7D, QPm.caas-7D, and QPm.caas-7DS in Opata 85, Courtot, Hanxuan 10, Opata 85, Fukuho-komugi, and Libellula, respectively (Börner et al. 2002; Huo et al. 2005; Bougot et al. 2006; Liang et al. 2006; Asad et al. 2012). Many previous reports indicated that Chinese Spring had Lr34/Yr18/Pm38 (Dyck 1977; Bossolini et al. 2006; Lagudah et al. 2006, 2009; Krattinger et al. 2009; Wu et al. 2015a, b; Zhang et al. 2017). Thus, QPm.caas-7DS should be Pm38 in Chinese Spring.
Implications of QPm.caas-3BS and QPm.caas-7DS for wheat breeding
The pleiotropic QTL QPm.caas-3BS, significantly reducing MDS to powdery mildew and leaf rust, could be used to develop new cultivars for disease resistance. Aikang 58, containing the resistance allele of QPm.caas-3BS and manifesting prominent resistance, is an excellent powdery mildew resistance source. Zhoumai 22, the most widely grown cultivar in Henan province (Xu et al. 2010; Zou et al. 2017), has resistance to stripe rust, leaf rust, and powdery mildew. Cultivars Zhoumai 26, Zhoumai 28, Zhoumai 36, Xinmai 32, Xinmai 36, and Yude 1, all derived from Zhoumai 22, carry the 3BS resistance allele and can be used as breeding parents.
The pleiotropic APR gene Lr34/Yr18/Pm38 has been successfully used in CIMMYT wheat breeding programs (Singh 1993; Bahl et al. 1997; Singh et al. 2005; Kolmer et al. 2008; Liang et al. 2009; Wu et al. 2010, 2015a, b) and in Canada. It is present at high frequency in Chinese wheat landraces (85.1% of 422 landraces, Yang et al. 2008), and the resistance gene probably originated from Chinese landraces (Dakouri et al. 2014). Nevertheless, due to ease of selection for major resistance genes in Chinese wheat breeding programs, Lr34/Yr18/Pm38 is present in relatively few current wheat cultivars (Yang et al. 2008), and was not found in any of the 103 Zhou8425B derivatives included in the present study. The accumulation of 4–5 slow rusting or slow mildewing resistance genes can achieve high levels of resistance in wheat cultivars (Singh et al. 2000, 2005; Lu et al. 2009). Therefore, it is important to combine genes such as QPm.caas-3BS and Pm38 with other APR genes in the future especially given that at least some of these genes protect against multiple diseases.
Author contribution statement
ALJ performed the experiment and data analysis, and wrote the paper. FMG contributed to construction of the genetic map. YR, JDL, LG, and JZZ participated in the field trials. GHY developed the RIL population and collected 103 Zhou8425B derivative wheat cultivars. ZHH and XCX designed the experiment and wrote the paper. All authors read and approved the final manuscript.
Abbreviations
- ANOVA:
-
Analysis of variance
- APR:
-
Adult-plant resistance
- BLUEs:
-
Best linear unbiased estimates
- CAPS:
-
Cleaved amplified polymorphic sequences
- QTL:
-
Quantitative trait locus (loci)
- ICIM:
-
Inclusive composite interval mapping
- IT:
-
Infection type
- KASP:
-
Kompetitive allele-specific PCR
- LOD:
-
Logarithm of odds
- MAS:
-
Marker-assisted selection
- MDS:
-
Maximum disease severity
- PVE:
-
Phenotypic variance explained
- RFLP:
-
Restriction fragment length polymorphism
- RIL:
-
Recombinant inbred line
- SNP:
-
Single-nucleotide polymorphisms
- STARP:
-
Semi-thermal asymmetric reverse PCR
References
Asad MA, Bai B, Lan CX, Yan J, Xia XC, Zhang Y, He ZH (2012) Molecular mapping of quantitative trait loci for adult-plant resistance to powdery mildew in Italian wheat cultivar Libellula. Crop Pasture Sci 63:539–546
Bahl PN, Salimath PM, Mandal AK (1997) Genetics, cytogenetics and breeding of crop plants. Oxford and IBH Publishing Co. Pvt LTD, New Delhi and Calcutta, pp 75–144
Bates D, Mächler M, Bolker BM, Walker SC (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67:1–48
Börner A, Schumann E, Fürste A, Cöster H, Leithold B, Röder MS, Weber WE (2002) Mapping of quantitative trait loci determining agronomic important characters in hexaploid wheat (Triticum aestivum L.). Theor Appl Genet 105:921–936
Bossolini E, Krattinger SG, Keller B (2006) Development of simple sequence repeat markers specific for the Lr34 resistance region of wheat using sequence information from rice and Aegilops tauschii. Theor Appl Genet 113:1049–1062
Bougot Y, Lemoine J, Pavoine MT, Guyomar’ch H, Gautier V, Muranty H, Barloy D (2006) A major QTL effect controlling resistance to powdery mildew in winter wheat at the adult plant stage. Plant Breed 125:550–556
Colasuonno P, Gadaleta A, Giancaspro A, Nigro D, Giove S, Incerti O, Mangini G, Signorile A, Simeone R, Blanco A (2014) Development of a high-density SNP-based linkage map and detection of yellow pigment content QTLs in durum wheat. Mol Breed 34:1563–1578
Dakouri A, McCallum BD, Cloutier S (2014) Haplotype diversity and evolutionary history of the Lr34 locus of wheat. Mol Breed 33:639–655
Dyck PL (1977) Genetics of leaf rust reaction in three introductions of common wheat. Can J Genet Cytol 19:711–716
Gao FM, Wen WE, Liu JD, Rasheed A, Yin GH, Xia XC, Wu XX, He ZH (2015) Genome-wide linkage mapping of QTL for yield components, plant height and yield-related physiological traits in the Chinese wheat cross Zhou 8425B/Chinese Spring. Front Plant Sci 6:1099
Hao YF, Parks R, Cowger C, Chen ZB, Wang YY, Bland D, Murphy JP, Guedira M, Brown-Guedira G, Johnson J (2015) Molecular characterization of a new powdery mildew resistance gene Pm54 in soft red winter wheat. Theor Appl Genet 128:465–476
Herrera-Foessel SA, Singh RP, Lillemo M, Huerta-Espino J, Bhavani S, Singh S, Lan CX, Calvo-Salazar V, Lagudah ES (2014) Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theor Appl Genet 127:781–789
Huo NX, Zhou RH, Zhang FL, Jia JZ (2005) Mapping quantitative trait loci for powdery mildew resistance in wheat. Acta Agron Sin 31:692–696
Keller M, Keller B, Schachermayr G, Winzeler M, Schmid JE, Stamp P, Messmer MM (1999) Quantitative trait loci for resistance against powdery mildew in a segregating wheat × spelt population. Theor Appl Genet 98:903–912
Kolmer JA, Singh RP, Garvin DF, Viccars L, William HM, Huerta-Espino J, Ogbonnaya FC, Raman H, Orford S, Bariana HS, Lagudah ES (2008) Analysis of the Lr34/Yr18 rust resistance region in wheat germplasm. Crop Sci 48:1841–1852
Kota R, Spielmeyer W, McIntosh RA, Lagudah ES (2006) Fine genetic mapping fails to dissociate durable stem rust resistance gene Sr2 from pseudo-black chaff in common wheat (Triticum aestivum L.). Theor Appl Genet 112:492–499
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Lagudah ES, McFadden H, Singh RP, Huerta-Espino J, Bariana HS, Spielmeyer W (2006) Molecular genetic characterization of the Lr34/Yr18 slow rusting resistance gene region in wheat. Theor Appl Genet 114:21–30
Lagudah ES, Krattinger SG, Herrera-Foessel S, Singh RP, Huerta-Espino J, Spielmeyer W, Brown-Guedira G, Selter LL, Keller B (2009) Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor Appl Genet 119:889–898
Li ZQ, Zeng SM (2002) Wheat rust in China. China Agriculture Press, Beijing, pp 180–190
Li ZF, Zheng TC, He ZH, Li GQ, Xu SC, Li XP, Yang GY, Singh RP, Xia XC (2006) Molecular tagging of stripe rust resistance gene YrZH84 in Chinese wheat line Zhou 8425B. Theor Appl Genet 112:1098–1103
Li HH, Ye GY, Wang JK (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175:361–374
Liang SS, Suenaga K, He ZH, Wang ZL, Liu HY, Wang DS, Singh RP, Sourdille P, Xia XC (2006) Quantitative trait loci mapping for adult-plant resistance to powdery mildew in bread wheat. Phytopathology 96:784–789
Liang D, Yang FP, He ZH, Yao DN, Xia XC (2009) Characterization of Lr34/Yr18, Rht-B1b, Rht-D1b genes in CIMMYT wheat cultivars and advanced lines using STS markers. Sci Agric Sin 42:17–27
Lillemo M, Asalf B, Singh RP, Huerta-Espino J, Chen XM, He ZH, Bjørnstad Å (2008) The adult plant rust resistance loci Lr34/Yr18 and Lr46/Yr29 are important determinants of partial resistance to powdery mildew in bread wheat line Saar. Theor Appl Genet 116:1155–1166
Liu ZY, Sun QX, Ni ZF, Yang TM (1999) Development of SCAR markers linked to the Pm21 gene conferring resistance to powdery mildew in common wheat. Plant Breed 118:215–219
Liu SX, Griffey CA, Saghai-Maroof MA (2001) Identification of molecular markers associated with adult plant resistance to powdery mildew in common wheat cultivar Massey. Crop Sci 41:1268–1275
Liu JD, He ZH, Wu L, Bai B, Wen WE, Xie CJ, Xia XC (2016) Genome-wide linkage mapping of QTL for black point reaction in bread wheat (Triticum aestivum L.). Theor Appl Genet 129:2179–2190
Liu WX, Koo DH, Xia Q, Li CX, Bai FQ, Song YL, Friebe B, Gill BS (2017) Homoeologous recombination-based transfer and molecular cytogenetic mapping of powdery mildew-resistant gene Pm57 from Aegilops searsii into wheat. Theor Appl Genet 130:841–848
Long YM, Chao WS, Ma GJ, Xu SS, Qi LL (2017) An innovative SNP genotyping method adapting to multiple platforms and throughputs. Theor Appl Genet 130:597–607
Lu YM, Lan CX, Liang SS, Zhou XC, Liu D, Zhou G, Lu LQ, Jing JX, Wang MN, Xia XC, He ZH (2009) QTL mapping for adult-plant resistance to stripe rust in Italian common wheat cultivars Libellula and Strampelli. Theor Appl Genet 119:1349–1359
Mago R, Brown-Guedira G, Dreisigacker S, Breen J, Jin Y, Singh R, Appels R, Lagudah ES, Ellis J, Spielmeyer W (2011a) An accurate DNA marker assay for stem rust resistance gene Sr2 in wheat. Theor Appl Genet 122:735–744
Mago R, Tabe L, McIntosh RA, Pretorius Z, Kota R, Paux E, Wicker T, Breen J, Lagudah ES, Ellis JG, Spielmeyer W (2011b) A multiple resistance locus on chromosome arm 3BS in wheat confers resistance to stem rust (Sr2), leaf rust (Lr27) and powdery mildew. Theor Appl Genet 123:615–623
McFadden ES (1930) A successful transfer of emmer characters to vulgare wheat. J Am Soc Agron 22:1020–1034
McIntosh RA, Dubcovsky J, Rogers WJ, Morris C, Xia XC (2017) Catalogue of gene symbols for wheat: 2017 supplement. In: 12th international wheat genetics symposium, Yokohama
Meng L, Li HH, Zhang LY, Wang JK (2015) QTL IciMapping: integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J 3:269–283
Moore JW, Herrera-Foessel S, Lan CX, Schnippenkoetter W, Ayliffe M, Huerta-Espino J, Lillemo M, Viccars L, Milne R, Periyannan S, Kong XY, Spielmeyer W, Talbot M, Bariana H, Patrick JW, Dodds P, Singh R, Lagudah E (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498
Nyquist WE, Baker RJ (1991) Estimation of heritability and prediction of selection response in plant populations. Crit Rev Plant Sci 10:235–322
Olesen JE, Mortensen JV, Jørgensen LN, Andersen MN (2000) Irrigation strategy, nitrogen application and fungicide control in winter wheat on a sandy soil. I. Yield, yield components and nitrogen uptake. J Agri Sci 134:1–11
Petersen S, Lyerly JH, Worthington ML, Parks WR, Cowger C, Marshall DS, Brown-Guedira G, Murphy JP (2015) Mapping of powdery mildew resistance gene Pm53 introgressed from Aegilops speltoides into soft red winter wheat. Theor Appl Genet 128:303–312
Roelfs AP (1977) Foliar fungal diseases of wheat in the People’s Republic of China. Plant Dis Report 61:836–841
Saari EE, Wilcoxson RD (1974) Plant disease situation of high-yielding dwarf wheats in Asia and Africa. Annu Rev Phytopathol 12:49–68
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal locations and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Selter LL, Shatalina M, Singla J, Keller B (2014) Identification and implementation of resistance: Genomics-assisted use of genetic resources for breeding against powdery mildew and Stagonospora nodorum blotch in wheat. In: Tuberosa R, Graner A, Frison E (eds) Genomics of plant genetic resources, vol 2. Springer, Dordrecht, pp 360–361
Semagn K, Babu R, Hearne S, Olsen M (2014) Single nucleotide polymorphism genotyping using kompetitive allele specific PCR (KASP): overview of the technology and its application in crop improvement. Mol Breed 33:1–14
Singh RP (1992) Association between gene Lr34 for leaf rust resistance and leaf tip necrosis in wheat. Crop Sci 32:874–878
Singh RP (1993) Resistance to leaf rust in 26 Mexican wheat cultivars. Crop Sci 33:633–637
Singh RP, McIntosh RA (1984) Complementary genes for reaction to Puccinia recondita tritici in Triticum aestivum. I. Genetic and linkage studies. Can J Genet Cytol 26:723–735
Singh RP, Huerta-Espino J, Rajaram S (2000) Achieving near-immunity to leaf and stripe rusts in wheat by combining slow rusting resistance genes. Acta Phytopathol Entomol Hung 35:133–139
Singh RP, Huerta-Espino J, William HM (2005) Genetics and breeding for durable resistance to leaf and stripe rusts in wheat. Turk J Agric For 29:121–127
Sourdille P, Singh S, Cadalen T, Brown-Guedira GL, Gay G, Qi LL, Gill BS, Dufour P, Murigneux A, Bernard M (2004) Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Funct Integr Genom 4:12–25
Thomson MJ (2014) High-throughput SNP genotyping to accelerate crop improvement. Plant Breed Biotechnol 2:195–212
Tucker DM, Griffey CA, Liu S, Brown-Guedira G, Marshall DS, Saghai-Maroof MA (2007) Confirmation of three quantitative trait loci conferring adult plant resistance to powdery mildew in two winter wheat populations. Euphytica 155:1–13
Wang SC, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Whan A, Stephen S, Barker G, Wieseke R, Plieske J, International Wheat Genome Sequencing Consortium, Lillemo M, Mather D, Appels R, Dolferus R, Brown-Guedira G, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo MC, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden M, Akhunov E (2014) Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796
Wiersma AT, Pulman JA, Brown LK, Cowger C, Olson EL (2017) Identification of Pm58 from Aegilops tauschii. Theor Appl Genet 130:1123–1133
Wu L, Xia XC, Zhu HZ, Li SZ, Zheng YL, He ZH (2010) Molecular characterization of Lr34/Yr18/Pm38 in 273 CIMMYT wheat cultivars and lines using functional markers. Sci Agric Sin 43:4553–4561
Wu L, Xia XC, Rosewarne GM, Zhu HZ, Li SZ, Zhang ZY, He ZH (2015a) Stripe rust resistance gene Yr18 and its suppressor gene in Chinese wheat landraces. Plant Breed 134:634–640
Wu QH, Chen YX, Zhou SH, Fu L, Chen JJ, Xiao Y, Zhang D, Ouyang SH, Zhao XJ, Cui Y, Zhang DY, Liang Y, Wang ZZ, Xie JZ, Qin JX, Wang GX, Li DL, Huang YL, Yu MH, Lu P, Wang LL, Wang L, Wang H, Dang C, Li J, Zhang Y, Peng HR, Yuan CG, You MS, Sun QX, Wang JR, Wang LX, Luo MC, Han J, Liu ZY (2015b) High-density genetic linkage map construction and QTL mapping of grain shape and size in the wheat population Yanda 1817 × Beinong6. PLoS One 10:e0118144
Wu QH, Chen YG, Fu L, Zhou SH, Chen JJ, Zhao XJ, Zhang D, Ouyang SH, Wang ZZ, Li D, Wang GX, Zhang DY, Yuan CG, Wang LX, You MS, Han J, Liu ZY (2016) QTL mapping of flag leaf traits in common wheat using an integrated high-density SSR and SNP genetic linkage map. Euphytica 208:337–351
Xiao YG, Yin GH, Li HH, Xia XC, Yan J, Zheng TC, Ji WQ, He ZH (2011) Genetic diversity and genome-wide association analysis of stripe rust resistance among the core wheat parent Zhou 8425B and its derivatives. Sci Agric Sin 44:3919–3929
Xu WG, Li CX, Hu L, Zhang L, Zhang JZ, Dong HB, Wang GS (2010) Molecular mapping of powdery mildew resistance gene PmHNK in winter wheat (Triticum aestivum L.) cultivar Zhoumai 22. Mol Breed 26:31–38
Yang WX, Yang FP, Liang D, He ZH, Shang XW, Xia XC (2008) Molecular characterization of slow-rusting genes Lr34/Yr18 in Chinese wheat cultivars. Acta Agron Sin 34:1109–1113
Yin GH, Wang JW, Wen WE, He ZH, Li ZF, Wang H, Xia XC (2009) Mapping of wheat stripe rust resistance gene YrZH84 with RGAP markers and its application. Acta Agron Sin 35:1274–1281
Zhang PP, Yin GH, Zhou Y, Qi AY, Gao FM, Xia XC, He ZH, Li ZF, Liu DQ (2017) QTL mapping of adult-plant resistance to leaf rust in the wheat cross Zhou8425B/Chinese Spring using high-density SNP markers. Front Plant Sci 8:793
Zhao XL, Zheng TC, Xia XC, He ZH, Liu DQ, Yang WX, Yin GH, Li ZF (2008) Molecular mapping of leaf rust resistance gene LrZH84 in Chinese wheat line Zhou 8425B. Theor Appl Genet 117:1069–1075
Zhao ZH, Sun HG, Song W, Lu M, Huang J, Wu LF, Wang XM, Li HJ (2013) Genetic analysis and detection of the gene MlLX99 on chromosome 2BL conferring resistance to powdery mildew in the wheat cultivar Liangxing 99. Theor Appl Genet 126:3081–3089
Zou SK, Yin GH, Tang JW, Han YL, Li SC, Li NN, Huang F, Wang LN, Zhang Q, Gao Y (2017) Molecular and genetic basis of wheat variety Zhoumai 22 and specific primers screening. J Triticeae Crops 37:472–482
Acknowledgements
The authors are grateful to Prof. R. A. McIntosh, Plant Breeding Institute, University of Sydney, for review of this manuscript. This study was supported by the National Key Research and Development Program of China (2016YFD0101802), National Natural Science Foundation of China (31461143021), Natural Science Foundation of Henan province (162300410348), and CAAS Science and Technology Innovation Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical standards
We declare that these experiments complied with the ethical standards in China.
Additional information
Communicated by Hermann Buerstmayr.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jia, A., Ren, Y., Gao, F. et al. Mapping and validation of a new QTL for adult-plant resistance to powdery mildew in Chinese elite bread wheat line Zhou8425B. Theor Appl Genet 131, 1063–1071 (2018). https://doi.org/10.1007/s00122-018-3058-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-018-3058-x