Abstract
Key message
GWAS on multi-environment data identified genomic regions associated with trade-offs for grain weight and grain number.
Abstract
Grain yield (GY) can be dissected into its components thousand grain weight (TGW) and grain number (GN), but little has been achieved in assessing the trade-off between them in spring wheat. In the present study, the Wheat Association Mapping Initiative (WAMI) panel of 287 elite spring bread wheat lines was phenotyped for GY, GN, and TGW in ten environments across different wheat growing regions in Mexico, South Asia, and North Africa. The panel genotyped with the 90 K Illumina Infinitum SNP array resulted in 26,814 SNPs for genome-wide association study (GWAS). Statistical analysis of the multi-environmental data for GY, GN, and TGW observed repeatability estimates of 0.76, 0.62, and 0.95, respectively. GWAS on BLUPs of combined environment analysis identified 38 loci associated with the traits. Among them four loci—6A (85 cM), 5A (98 cM), 3B (99 cM), and 2B (96 cM)—were associated with multiple traits. The study identified two loci that showed positive association between GY and TGW, with allelic substitution effects of 4% (GY) and 1.7% (TGW) for 6A locus and 0.2% (GY) and 7.2% (TGW) for 2B locus. The locus in chromosome 6A (79–85 cM) harbored a gene TaGW2-6A. We also identified that a combination of markers associated with GY, TGW, and GN together explained higher variation for GY (32%), than the markers associated with GY alone (27%). The marker-trait associations from the present study can be used for marker-assisted selection (MAS) and to discover the underlying genes for these traits in spring wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum L.) provides 20% of the total calories and 20% of plant-derived protein to the world population (Food and Agricultural Organization of the United Nations, 2010). However, the production levels need to be increased by 70% to meet the projected food requirements by 2050 (Ray et al. 2012). Even though progress has been made through conventional breeding approaches in increasing genetic gains of grain yield (GY) of spring bread wheat, it is less than 1% per year (Sharma et al. 2012; Aisawi et al. 2015; Crespo-Herrera et al. 2017). To meet the predicted demand, it is important to complement the conventional approaches through molecular breeding for complex traits (Reynolds and Langridge 2016). Even though GY is the most important trait in any plant breeding program, there still exists a large gap in understanding the genetic and molecular mechanism of the trait and its components (Valluru et al. 2014).
Linkage mapping and genome-wide association studies (GWAS) are two methods widely used to identify and understand the genetic basis of complex traits (Zhu et al. 2008). Grain yield is a complex trait determined by multiple quantitative trait loci (QTL) that interact with each other and with the environment (Sehgal et al. 2017). Genetic analysis to identify the genomic region for GY is often subjected to genotype-by-environment interaction (G × E) due to the complex nature of the trait. A recent study has shown the phenotypic plasticity for GY in the US Great Plains ranged from 1.3 to 5.3 Mg ha−1 (Grogan et al. 2016). To discover the genetic basis of grain yield it is important to have multi-environmental experiments conducted. In those studies, the identified QTL are constitutive—when consistently detected across most environments—or adaptive—when only detected in specific environmental conditions (Vargas et al. 1998).
Until a decade ago, QTL mapping was the first choice for genetic analysis to understand complex traits like GY (Quarrie et al. 1994, 2006; Kato et al. 1999; Kirigwi et al. 2007) but the trend has moved to GWAS due to less time required for population development and higher mapping resolutions (Zhang et al. 2010; Huang et al. 2010; Sukumaran et al. 2012). GWAS was used to identify genomic regions associated with GY and related traits in several populations (Sonah et al. 2015; Maccaferri et al. 2016; Valluru et al. 2017). Genomic regions related to plant development—phenology and plant height—were identified in chromosomes 1AL, 1BS, 2AL, 2BS, 2BL, 4BL, 5BL, and 6AL, with constitutive QTL on chromosomes 5BL and 6AL (Ain et al. 2015). Our previous study in temperate irrigated environments in Mexico identified genomic regions related to GY and yield components in chromosomes 5A (98 cM) and 6A (77–85 cM) (Sukumaran et al. 2015a).
Genes and QTLs associated with thousand grain weight (TGW) have been studied in bread wheat (Ramya et al. 2010; Zanke et al. 2014; Kumar et al. 2015; Simmonds et al. 2016; Brinton et al. 2017a). A region responsible for grain weight was fine mapped in chromosome 7D using introgression lines in winter wheat (Röder et al. 2008). However, the most studied gene affecting TGW in wheat is TaGW2-6A an orthologue of the OsGW2 gene, a RING-type E3 ubiquitin ligase in rice that influences grain width and weight (Su et al. 2011; Yang et al. 2012; Zhang et al. 2013; Jaiswal et al. 2015). This gene has two haplotypes—Hap-6A-A and Hap-6A-G—detected in the promotor region (Su et al. 2011). However, the superior effect of both haplotypes in different germplasms were reported. For instance, Hap-6A-A is superior allele in Chinese germplasm, increasing 3 g of TGW (Su et al. 2011) but another report suggested Hap-6A-G is the superior allele (Zhang et al. 2013). When this gene has an insertion in the eighth exon it reduces the protein sequence from 424 to 328 amino acids, it is responsible for large kernel in a European wheat variety (Yang et al. 2012). QTLs have been identified for grain number (GN), but none of them was studied in detail in spring wheat (Börner et al. 2002; Kuchel et al. 2007). A QTL mapping approach was used to identify a locus in chromosome 1B, which controls GN, and a locus in 7B associated with TGW without significant reduction in grain number (Griffiths et al. 2015). Overall, there has been little effort on dissecting the TGW and GN trade-off in spring wheat.
In general, GY is positively associated with GN, the positive association with TGW is less profound, and the two components themselves are usually negatively correlated. The aims of this study were (1) to identify genomic regions associated with GY, GN, and TGW in an elite spring wheat panel in diverse environments; (2) to identify sets of markers associated with GY and components that maximize the probability of identifying high GY wheat lines; and (3) identify specific markers for TGW and GN independent of GY, those can be used to maximize TGW and GN (e.g. increasing TGW while increasing GY and GN).
Materials and methods
Plant material
The plant material used in the study was the Wheat Association Mapping Initiative (WAMI) panel, which consisted of 287 spring bread wheat lines assembled from several of the CIMMYT’s wheat international nurseries distributed around the world; Elite Spring Wheat Yield Trials (ESWYT); Semi-arid Wheat Yield Trials (SAWYT), and High Temperature Wheat Yield Trials (HTWYT) (Lopes et al. 2012). In the WAMI panel, several studies were conducted; GWAS for GY and yield components in temperate irrigated environments (Sukumaran et al. 2015a), genomic regions for adaptation to density (Sukumaran et al. 2015b), identification of earliness per se (eps) locus (Sukumaran et al. 2016), markers for grain yield in moisture stress environments (Edae et al. 2014), QTLs for spike ethylene (Valluru et al. 2017), and candidate gene association mapping for drought tolerance (Edae et al. 2013). The population structure of the panel is loosely based on 1B.1R translocation as well as pedigree of lines, e.g. lines crossed with and derived from elite lines like Pastor, Weebil, and Baviacora are in high frequency in this panel (Lopes et al. 2015; Sukumaran et al. 2015a). The WAMI population has a low range of phenology; range of 9 days for heading and 35 cm for plant height, when grown under temperate irrigated conditions in Mexico (Lopes et al. 2015; Sukumaran et al. 2015a).
Phenotyping
The WAMI population was phenotyped in 31 major wheat growing areas in Bangladesh, India, Pakistan, Nepal, Sudan, and Mexico in 2009–2010 and 2010–2011 growing seasons (Lopes et al. 2012, 2015; Sukumaran et al. 2015b). We used a subset of the data—ten environments—from this, based on heritability estimates (Table 1); Bangladesh Agricultural Research Institute (BARI), Joydebpur, Indian Agricultural Research Institute (IARI), Indore (India I), National Wheat Research Program (NWRP) Bhairahawa, Rupandehi (Nepal B), National Agricultural Research Centre (NARC), Islamabad, (PAK I), Sudan Wad Medani (Sudan W), and CIMMYT’s experimental station, Campo Experimental Normal E. Borlaug (CENEB), Sonora Mexico (Mex), where I, D, H, and HD denotes, irrigated, drought, heat, and heat + drought conditions, respectively. These environments were diverse in terms of rainfall, heat stress, drought stress, and solar radiation patterns (Sukumaran et al. 2017). Minimum and maximum temperatures and coordinates of the environments were described in earlier publications (Sukumaran et al. 2016). The experimental design at each site was an α-lattice with two replications. Blocks were arranged based on the heading date of the materials. Checks were included as entry number 50, 100, 150, and 200 to verify any planting errors.
Several traits were recorded and among them three traits were analyzed in the present study: GY, GN, and TGW using standard protocols (Sayre et al. 1997; see Pask et al. 2012). At most locations, plants were sown on flat beds generally of 2.5–3.0 m length with 4–6 rows (20–25 cm between rows) providing total harvestable area > 3 m2 (Reynolds et al. 2017). Grain number was estimated from GY and TGW. Days to heading (DH) was recorded when 50% of the spikes in a plot had emerged from the boot leaf (Zadoks stage 59) from days of emergence and was used as a co-variate in the analyses (Zadoks et al. 1974). The phenotypic data for each environment and trait, as well as the genomic data, is available from the link http://hdl.handle.net/11529/10714.
Genotyping and statistical analysis
Genotyping of the panel with the 90 K Illumina Infinitum SNP array and SNP processing are described in an earlier publication (Sukumaran et al. 2015a). In short, genetic data of 38K SNPs was processed for monomorphic markers, missing values (< 5%), and minor allele frequency (> 10%) that resulted in 28K SNPs for GWAS. Several markers related to the genes for vernalization, photoperiod, plant height, and 1B/1R translocation were scored in the WAMI panel (Sukumaran et al. 2017). The genes with known position in the panel were: Vrn-A1 (5A) (Yan et al. 2003), Ppd-B1 (2B), and Rht-B1 (4B) (Ellis et al. 2002), earliness per se (Eps-D1) (Zikhali et al. 2014; Sukumaran et al. 2016), and TaGW2-6A (Simmonds et al. 2016) identified through blind association analysis, where marker score was used as a phenotype.
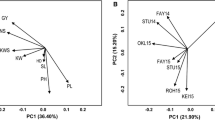
Analysis of variance (ANOVA) was conducted in SAS using the PROC MIXED commands. Genotypes, environments, and genotype-by-environment interaction (G × E) were considered as random factors in the model in estimating the Best Linear Unbiased predictions (BLUPs) for each environment and in combined analysis using META-R (Vargas et al. 2013). Principal component analysis of the data was performed using the R package “FactoMineR” (Lê et al. 2008) and correlations between the environments were estimated using the R package “corrplot” (Wei and Simko 2017). For the individual environment analyses, the following model was used:
where Y is the trait of interest, μ is the mean effect, R i is the effect of the ith replicate, B j (R i ) is the effect of the jth incomplete block within the ith replicate, G k is the effect of the kth genotype, Cov is the effect of the covariate, and ε ijk is the error associated with the ith replication, jth incomplete block, and kth genotype, which is assumed to be normally and independently distributed, with mean zero and variance σ 2. Broad sense repeatability (H 2) was estimated as
where \(\sigma_{\text{g}}^{2}\) and \(\sigma_{\text{e}}^{2}\) are genotype and environment variance and r is the number of replications.
For a combined analysis across all environments, the following model used for the estimation of BLUPs;
where the new terms L i and L i × G l are the effects of the ith environment and the G × E, respectively. For the combined analysis the H 2 was estimated as,
where \(\sigma_{\text{ge}}^{2}\) is the genotype by environment interaction variance and l is the number of environments.
Genome-wide association analysis
GWAS were performed on the BLUPs for each trait from the combined analyses of ten environments. We followed the unified mixed model approach (Yu et al. 2006; Zhang et al. 2010) as well as the generalized linear models incorporated in the TASSEL 5.0 software (Bradbury et al. 2007) to test for marker trait associations (MTAs). To account for population structure, principal components (PC1–5) from the principal component analysis of the genotypic data, Q matrix (Q1–5) derived from the STRUCTURE software (Pritchard and Rosenberg 1999; Falush et al. 2003, 2007), and non-metric multi-dimensional scaling (nMDS1–5) (Zhu and Yu 2009) were used. In addition, kinship matrix (K) calculated using SPARGeDi (Hardy and Vekemans 2002) and coefficient of parentage matrix (COP) was used as random factor in the mixed model. We fitted several different models—including generalized linear models—accounted only fixed effects—and mixed linear models—accounted population structure and family relatedness—(i.e. simple model, Q1–5, K, COP, Q1–5 + K, Q1–5 + COP, PC1–5 + K, PC1–5 + COP, nMDS1–5 + K, and nMDS1–5 + COP) using the analysis options (Yu et al. 2006) in TASSEL. Best model to estimate the marker effects and marker traits associations were decided based on the quantile–quantile plots of p values from each model fitting; the model that is close to the 1:1 ratio as the best model (Yu et al. 2006). The threshold to call marker-trait associations significant was based on the p value where a drastic deviation of the expected p values from the observed p values was observed (Sukumaran et al. 2015a). To define a genomic region as QTL, linkage disequilibrium (LD) was estimated for the region with several significant MTAs.
Candidate-gene association mapping
We also performed candidate gene based association analysis with all the known genes in this panel. Random 600 SNPs at least 5 cM apart and distributed in the genome were used as background markers. The candidate genes along with these markers were analyzed to identify significant association with the phenotypes. Similar to GWAS, several models—linear and mixed—were fitted and the best model—determined by the Q–Q plots—was used to identify the MTAs. In addition, the effect of haplotype TaGW2-6A, which was identified to be associated with TGW from earlier studies, was tested on the BLUPs of TGW from combined analysis of the environments. We also tested the effect of this haplotype on TGW in ten individual environments using t test and results were reported.
Marker effects and best combinations of significant markers for grain yield
We used step-wise forward and backward regression to identify the best marker combinations for each trait (Schulthess et al. 2017). Multiple regression was performed with the Q matrix and significant markers for each trait to estimate the likelihood-ratio-based R 2 (LRR 2) (Sun et al. 2010) for each marker. LRR 2 estimation avoids the effect of intercept that potentially have confounding effect on the variation explained by each marker. In addition, all significant markers for GY, TGW, and GN were fitted together in the step-wise regression models and multiple regression models to identify the optimum combination of makers and to estimate the marker effects for GY. These analyses were performed using the custom-made scripts in R software.
Results
Agronomic variation and repeatability estimates
Among the 31 environments phenotyped for grain yield, ten environments with moderate to high H 2 values were used for the analyses. The average yield of the WAMI panel from all environments was 3.81 t/ha with a range of 2.51 t/ha in Nepal B11 to 6.48 t/ha in Mex I10. The highest heritability estimate for GY was observed in Mex H10 and Mex I10 (0.75) with a mean H 2 of 0.67 among all environments. The average TGW was 33.45 g and varied from 29.36 g in Mex HD10 to 43.58 g in Mex I10. The highest H 2 estimate for TGW (0.96) was observed in Mex I10 and the lowest was in Pak I10 (0.72). GN was estimated from GY and TGW and it varied from 767.88 grains per m−2 in Nepal B11 to 17,554 grains per m−2 in Ind I11. The H 2 estimate for GN ranged from 0.48 (Nepal B11) to 0.85 (Mex h10) (Table 1). The H 2 values across environments for GY, TGW, and GN were 0.68, 0.95, and 0.42, respectively. The mean values across environments were 3.81 t/ha, 33.4 g, 3739.1, for GY, TGW, and GN (Table 2).
Correlations between the traits and environments
Among environments, the highest correlation coefficients (r) was observed for TGW, followed by GN, and GY. The highest correlation for GY was observed between Mex H10 and Mex HD10 (r = 0.65). The Mexican environments had moderate to high r with most other environments. The lowest correlations among environments were observed with Sudan W10 and Pak I10 for GY. In Sudan W10, GY showed negative correlations with BGLD J11 and Mex HD10. The correlations for GN between the environments followed a pattern similar to GY with highest correlation between Mex I10 and Mex D10 (r = 0.68). Seven environments (BGLD J11, Nepal B11, Sudan W10, Mea H10, Mex HD10, Mex D10, and Mex I10) showed correlations (r > 0.30). TGW, the highest heritability trait showed the highest r (0.86) between Mex HD10 and Mex H10. Most environments were highly correlated for TGW and the lowest r value being 0.36 between Pak I10 and BGLD J11 (Fig. 1). The r value between GY and TGW was (r = 0.21), between GY and GN was (r = 0.56), and between GN and TGW was (r = − 0.50). In general, GN was positively correlated with GY than TGW in all environments.
GWAS results
GWAS was conducted in TASSEL using GLM and MLM models (Fig. 2). For most of the traits, PC3 + K matrix was the best model and showed less deviation of the expected values from the observed values in the Q–Q plots.
Grain yield
GWAS of the combined environment BLUPs detected 27 MTAs in six chromosomes—2A, 3B, 4A, 4B, 5A, and 7A—associated with GY that explained 4–7% of the variation of the trait with p values < 0.001 (Supplementary Table 1). The MTAs could be localized into ten genomic regions based on LD; 2A (106 cM), 3B (86 cM), 3B (91 cM), 3B (95 cM), 3B (115 cM), 4A (151 cM), 4B (66–68 cM), 6A (77–85 cM), and 7A (35 cM) (Table 3). We further explored the 3B and 6A regions and found that, the markers in chromosome 3B at 86, 91, 95, and 119 cM are in high LD, with r 2 = 1, indicating a large QTL for GY in chromosome 3B from 86 to 119 cM (Supplementary Fig. 1). LD estimates in the chromosome 6A region from 77 to 85 cM indicated the presence of an LD block in 77–81 cM region, but on individual environment analysis markers at 85 cM were also associated (Supplementary Fig. 2). A blind association analysis of the TaGW2-6A haplotype indicted the possible location of the TaGW2 gene in chromosome 6A (77–78 cM).
Thousand grain weight
We identified 34 MTAs for TGW from combined analysis of environment BLUPs in chromosomes; 1B, 2A, 2B, 3B, 3D, 5A, 6A, 6B, and 7D (Supplementary Table 2). Based on LD analysis, these MTAs corresponded to 15 QTLs—1B (141–148, and 164 cM), 2A (143 cM), 2B (20–26 cM), 2B (96–99 cM and 145 cM, 3B (51 cM), 3D (48 cM), 5A (26, 60, and 98 cM), 6A (77–85 cM), 6B (113 cM), and 7D (55 and 78 cM), located in nine chromosomes (Table 3). The Vrn-A1 gene in chromosome 5A (90 cM) had significant effect on flowering time in an earlier study (Sukumaran et al. 2015a), but the present locus associated with TGW is in chromosome 5A at 98 cM. LD analysis indicated these loci are not in high LD, which suggests that these are separate loci (Supplementary Fig. 3).
Grain number
For GN, we identified 31 significant MTAs in twelve chromosomes; 1A, 1D, 2B, 3A, 3B, 4B, 5A, 5B, 5D, 6A, 6D, and 7A (Supplementary Table 3). LD analysis of the significant MTAs narrowed the significant loci into 13 QTLs on chromosomes 1A (130 cM), 1D (51 cM) 2B (96–99 cM), 3A (15 cM), 3B (99 cM), 4B (81 cM), 5A (98 cM), 5B (3–4 cM), 5D (203 cM), 6A (141 cM), 6D (77 cM), and 7A (120 and 135 cM) (Table 4).
Common markers and trade-off for grain weight and grain number
A comparison of the GWAS results for all traits identified four common regions associated with multiple traits. A common locus for GY and TGW was located in chromosome 6A (77–85 cM). A common QTL for GY and GN was detected in chromosome 3B (99 cM). Common loci for GN and TGW were in chromosome 5A (98 cM) and 2B (96 cM) (Fig. 3).
Venn diagram illustrating the common genomic regions for grain yield, grain weight, and grain number for the combined data from 10 environments. Subscript numbers indicate the centi morgan (cM) position of marker-trait associations in a chromosome based on the 90 K consensus map. The arrows indicates positive (↑) or negative (↓) additive effects of minor alleles
We also estimated the effect of these loci on GY, TGW, and GN. The locus in 6A (85 cM) had positive effect on GY, TGW, and GN with an allele substitution effect of 4.7, 1.7, and 2%, respectively. The locus on chromosome 3B (99 cM) had an allelic substitute effect of 3.0, − 4.5, and 3.1% for GY, TGW, and GN, respectively. The locus on chromosome 2B (96 cM) had a positive allelic substitution effect on GY (0.2%) and TGW (7.16%), but negative effect on GN (− 2.2%). The locus in chromosome 5A (98 cM) had positive effect on GY (0.18%) and GN (1.54%), but negative effect on TGW (− 2.85%) (Fig. 4).
Marker combinations and effects
We also identified the optimum marker combination that explained highest variation for GY, TGW, and GN using step-wise regression (Supplementary Tables 4, 5, and 6) and multiple regression analyses. Five marker combinations—IWA2963 (4B), IWB2774 (3B), IWB52628 (6A), IWB51659 (4B), and IWB72516 (2A)—explained 27% of the variation for GY based on multiple regression analysis, where as the population structure (Q3) explained 7% (Table 5). The marker IWA2963 (4B) explained 12% of the variation in GY based on LRR 2 and the other three markers explained 15% of the variance. The combination of 10 markers—IWB65271 (1B), IWB18267 (7D), IWB40900 (3B), IWA6949 (5A), IWB65783 (6A), IWB23810 (3D), IWB686 (5A), IWB32380 (2B), IWB2414 (2B), and IWB42660 (2B)—explained 31% of the variation for TGW, where as Q3 explained 28%. Multiple regression analysis identified the combination of 11 markers—IWB24502 (6D), IWB6365 (3A), IWB53601 (6A), IWB50952 (1A), IWB63867 (7A), IWB53861 (5D), IWB15488 (1D), IWB686 (5A), IWB68969 (7A), IWB9095 (3B), and IWB49242 (2B)—that explained 37% of the variation for GN. The population structure (Q3) explained 3% of the variation for GN.
We also fitted all significant MTAs for GY, TGW, and GN in step-wise regression to identify the most effective marker combination for GY (Supplementary Table 6). Multiple regression analysis identified a combination of 12 markers—IWB2774 (3B), IWB52628 (6A), IWB50952 (1A), IWA6949 (5A), IWB53601 (6A), IWB65783 (6A), IWB63656 (3A), IWB51659 (4B), IWB15488 (1D), IWB18267 (7D), IWB49242 (2B), and IWB686 (5A)—that explained 35% of the variation for GY, Q3 explaining 9% (Table 6).
Analysis of marker by environment interaction using step-wise regression of the 12 significant markers on the BLUPs for grain yield in each environment indicated variation in the marker effects. The markers directly associated with GY—IWB2774 (3.35%) and IWB52628 (3.02%)—explained highest variation for GY on average than markers indirectly associated for GY, i.e. markers identified for TGW and GN (Table 7). The highest LRR 2 values for a marker was in MexI10 environment, IWB2774 explaining 12.01% variation for GY. Total variation explained by the twelve markers were highest in MexI10 (24.08%) followed by MexHD10 (21.24%).
Association of known genes with the traits
We did candidate gene-based association mapping of several known genes. Results indicated that Rht-B1 and Vrn-D1a were associated with grain yield with a p value of < 0.001. Other genes were not significantly associated with any of the traits when a p value threshold of 0.001 was used. A blind association mapping—using marker score as trait—for the TaGW2-6A polymorphism in the WAMI population identified the location of the gene in chromosome 6A at 77–78 cM. However, in the combined analysis of BLUPs for TGW, the locus we identified was at 77 cM but individual analysis the locus was at 85 cM. A t-test of the TaGW2-6A haplotypes—Hap-6A-A and Hap-6A-G—with the phenotypic data of all individual environments indicated that the SNP is significantly associated with TGW in only two environments (Supplementary Table 9). The p value from t test was significant in Mex H10 (p = 0.002) and Nepal B10 (p = 0.02) at significant level of α = 0.05. In addition, the TaGW2-6A Hap 6A-G haplotype was superior in Mexico D10 but TaGW2-6A Hap-6A-A haplotype was superior in Nepal B10. In all other environments, the marker effect was not significant as denoted by standard error in bar plot (Supplementary Fig. 4).
Discussion
In the present study, we identified several MTAs for GY, TGW, and GN using large-scale multi-environment data in spring wheat. We observed that the combined analysis of environments had high heritability estimates and increased the power to detect causative loci. Large-scale multi-environment studies are associated with high G × E, which prompted us to reduce the environments from 31 to 10 based on heritability estimates of the traits. In our previous study, we successfully followed the same approach for flowering time where we used 19 environments with high heritability estimates and identified earliness per se (eps-D1) locus in CIMMYT spring wheat germplasm (Sukumaran et al. 2016). Phenotypic correlations between the environments for each trait were high, indicating the importance of research conducted at Mexican environments that is applicable several countries (Braun et al. 1996). TGW showed the highest correlation between the environments similar to our earlier study (Sukumaran et al. 2017).
The locus in chromosome 6A (77–85 cM) was associated with GY and TGW. We have detected the same locus important for GY, GN, and GW in temperate irrigated conditions in Cd. Obregon, Mexico in an earlier study (Sukumaran et al. 2015a). A QTL in chromosome 6A associated with TGW, GY, and green canopy duration was identified earlier in winter wheat and might be similar in spring wheat (Simmonds et al. 2014). Candidate gene based association mapping of the TaGW2-6A haplotype did not show significant association between the gene and TGW on BLUPs from combined analysis of environments. We observed the significant effect of this marker on TGW in two out of ten environments, and it was opposite in those environments. In Mex HD 10, the TaGW2-6A Hap-6A-G was superior whereas in Nepal the TaGW2-6A Hap-6A-A was superior. Taken together, it indicates that the effect of the gene is dependent on the environment and probably there could be another gene controlling the expression TaGW2-6A. Our study indicated that this gene does not show consistent and significant association with grain weight across different environments; however, it is close to the causative loci for TGW variation.
In addition to the 6A locus for GY and TGW, a common MTA for TGW and GN was detected in 5A at 98 cM. This locus was also associated with GN and TGW in temperate irrigated environments (Sukumaran et al. 2015a). A GWAS study using the 90 K SNP array in spring wheat also identified a similar genomic region associated with GY (Ain et al. 2015). Recently, a stable QTL on chromosome 5A associated with 6.9% increase in grain weight was identified which lead to 4% longer grains and 1.5% wider grains (Brinton et al. 2017b). The QTL contributes to increased pericarp cell length, thus contributing to grain weight We believe this might be the same locus in spring wheat and winter wheat but need to be further explored.
A chromosome 3B genomic region associated with grain yield was also reported in multiple studies (Bonneau et al. 2013; Lopes et al. 2013; Edae et al. 2014; Sukumaran et al. 2015a). The locus identified in chromosomes 3B (86–99 cM) was associated with multiple traits (GY, maturity, and chlorophyll content at vegetative state) in the temperate irrigated environments (Sukumaran et al. 2015a). An earlier study had identified MTAs on chromosome 3B at 70 cM for adaptation to density in the same WAMI panel (Sukumaran et al. 2015b). A QTL with large effect for GY was also identified in chromosome 3B using a different genetic map (Bonneau et al. 2013, 2017).
We will propose these four candidate regions in chromosomes 2B (96 cM), 3B (99 cM), 5A (98 cM), and 6A (77–85 cM) for further gene discovery and validation that is associated with the trade-off for grain weight and grain number. In general, it is possible to increase GY by increases in GN, but increases in GN is associated with decreased TGW. From our study, we found loci that are associated with a positive association between GY, TGW and GN.
Stepwise regression and multiple regression analysis identified the optimum combination significant markers for the traits. The additive effect of markers decreases as more number of markers are added to the model. We used population structure in the multiple regression model to avoid the overestimation of the marker effects from regression analysis. Likelihood ratio based R2 was estimated for each marker based on the marker substitution effect from multiple regression analysis (Sun et al. 2010). The best marker combinations for traits explained 27, 31, and 33% for GY, TGW, and GN, respectively. However, our analysis to find the optimum combination of markers found a marker based model that could explain 32% of GY, indicating a combination of makers for GY, TGW, and GN is important to select for GY in a breeding program, instead of marker for GY alone (27%). This indicated that the markers that are not directly related to the trait also has an effect on the trait and much of the variation is not explained when grain yield is dissected directly (Reynolds and Langridge 2016).
We also compared markers associated with GY, TGW, and GN with earlier studies. The MTA on chromosome 4B (66–68 cM) were close to the Rht-B1 locus in chromosome 4B (56 cM) identified by blind association analysis of Rht-B1 score. Candidate gene-based association analysis confirmed the association of Rht-B1 with grain yield (Supplementary Table 8). The MTA in chromosomes 2A (106 cM), 4A (151 cM), and 7A (35 cM) are novel loci associated with GY. Further analysis of the TGW MTAs showed the locus on chromosome 1B at 141–168 cM is close to the 1B/1R translocation from blind association analysis (Lopes et al. 2015). The locus in chromosome 7D (78 cM) was close to Vrn-D3 locus (91 cM) but we expect them to be different based on LD. The MTAs on 2A (143 cM), 2B (20, 26, and 145 cM), 3B (51 cM), 3D (148 cM), 5A (26 and 60 cM), and 6B (113 cM) for TGW are novel. Apart from the common loci for GN and other traits, the MTAs detected in chromosomes 1A (130 cM), 1D (51 cM), 3A (15 cM), 4B (81 cM), 5B (3–4 cM), 5D (203 cM), 6A (141 cM), 6D (77 cM), and 7A (135 cM) are novel in this population.
GWAS is a powerful technique to identify the genomic regions associated with traits of interest, but often with confounding effects of population structure in genetic analyses which was negotiated by the use of models accounting for population structure and familial relatedness (Zhu et al. 2008;Yu et al. 2006; Zhang et al. 2010). We compared the genomic regions detected in the study with previous studies on the same panel and with already known genes/QTLs detected. We identified four loci to be associated with GY and its components in chromosomes 2B, 3B, 5A, and 6A. Among them, the loci in chromosome 6A and 2B showed positive allelic substitution effect for GY and TGW. In many cases, GY is not strongly associated with TGW. Any positive association between GY, TGW, and GN is a perfect scenario for increasing overall GY by increasing GN and TGW.
Conclusions
Genome-wide association analysis identified several key genomic regions associated with grain yield and yield components. Among them, four of them showed a trade-off between thousand grain yield, grain weight, and grain number. A comparison of variation explained by markers associated with trait per se and its components indicated that higher variation is explained by the combination of markers for trait per se and its components. The genomic regions identified in the present study can be used for MAS and need to be further studied to fine map or clone genes.
Change history
16 February 2018
Unfortunately, the Fig. 1 of this original article was incorrectly published. The corrected Fig. 1 is given below.
Abbreviations
- WAMI:
-
The wheat association Mapping Initiative
- BLUPs:
-
Best linear unbiased predictions
- MLM:
-
Mixed linear models
- GLM:
-
Generalized linear models
References
Ain Q-U, Rasheed A, Anwar A et al (2015) Genome-wide association for grain yield under rainfed conditions in historical wheat cultivars from Pakistan. Front Plant Sci 6:743
Aisawi KAB, Reynolds MP, Singh RP, Foulkes MJ (2015) The physiological basis of the genetic progress in yield potential of CIMMYT spring wheat cultivars from 1966–2009. Crop Sci 55:1749–1764
Bonneau J, Taylor J, Parent B et al (2013) Multi-environment analysis and improved mapping of a yield-related QTL on chromosome 3B of wheat. Theor Appl Genet 126:747–761
Börner A, Schumann E, Fürste A, Cöster H, Leithold B, Röder MS, Weber WE (2002) Mapping of quantitative trait loci determining agronomic important characters in hexaploid wheat (Triticum aestivum L.). Theor Appl Genet 105(6–7):921–936. https://doi.org/10.1007/s00122-002-0994-1
Bradbury PJ, Zhang Z, Kroon DE et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Braun H-J, Rajaram S, Ginkel M (1996) CIMMYT’s approach to breeding for wide adaptation. Euphytica 92:175–183
Brinton J, Simmonds J, Minter F et al (2017a) Increased pericarp cell length underlies a major quantitative trait locus for grain weight in hexaploid wheat. New Phytol 6:1–6
Brinton J, Simmonds J, Minter F et al (2017b) Increased pericarp cell length underlies a major QTL for grain weight in hexaploid wheat. bioRxiv 6:1–6
Crespo-Herrera LA, Crossa J, Huerta-Espino J et al (2017) Genetic yield gains in CIMMYT’S international elite spring wheat yield trials by modeling the genotype × environment interaction. Crop Sci 57:789–801
Edae EA, Byrne PF, Manmathan H, Haley SD, Moragues M, Lopes MS, Reynolds MP (2013) Association mapping and nucleotide sequence variation in five drought tolerance candidate genes in spring wheat. Plant Genome 6(2). https://doi.org/10.3835/plantgenome2013.04.0010
Edae EA, Byrne PF, Haley SD et al (2014) Genome-wide association mapping of yield and yield components of spring wheat under contrasting moisture regimes. Theor Appl Genet 127:791–807
Ellis MH, Spielmeyer W, Gale KR et al (2002) “Perfect” markers for the Rht-B1b and Rht-D1b dwarfing genes in wheat. Theor Appl Genet 105:1038–1042
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578
Griffiths S, Wingen L, Pietragalla J et al (2015) Genetic dissection of grain size and grain number trade-offs in CIMMYT wheat germplasm. PLoS ONE 10:1–18
Grogan SM, Anderson J, Stephen Baenziger P et al (2016) Phenotypic plasticity of winter wheat heading date and grain yield across the US great plains. Crop Sci 56:2223–2236
Hardy OJ, Vekemans X (2002) SPAGeDI: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Huang X, Wei X, Sang T et al (2010) Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42:961–967
Jaiswal V, Gahlaut V, Mathur S et al (2015) Identification of novel SNP in promoter sequence of TaGW2-6A associated with grain weight and other agronomic traits in wheat (Triticum aestivum L.). PLoS One 10:1–15
Kato K, Miura H, Sawada S (1999) QTL mapping of genes controlling ear emergence time and plant height on chromosome 5A of wheat. TAG Theor Appl Genet 98:472–477
Kirigwi FM, Ginkel VM, Brown-Guedira G, Gill BS (2007) Markers associated with a QTL for grain yield in wheat under drought. Mol Breed 20:401–413
Kuchel H, Williams KJ, Langridge P, Eagles HA, Jefferies SP (2007) Genetic dissection of grain yield in bread wheat. I. QTL analysis. Theor Appl Genet 115(8):1029–1041. https://doi.org/10.1007/s00122-007-0629-7
Kumar U, Laza MR, Soulié JC et al (2015) Analysis and simulation of phenotypic plasticity for traits contributing to yield potential in twelve rice genotypes. Field Crop Res 202:94–107
Lê S, Josse J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25(1):1–18. https://doi.org/10.18637/jss.v025.i01
Lopes MS, Reynolds MP, Jalal-Kamali MR et al (2012) The yield correlations of selectable physiological traits in a population of advanced spring wheat lines grown in warm and drought environments. Field Crop Res 128:129–136
Lopes MS, Reynolds MP, McIntyre CL et al (2013) QTL for yield and associated traits in the Seri/Babax population grown across several environments in Mexico, in the West Asia, North Africa, and South Asia regions. Theor Appl Genet 126:971–984
Lopes MS, Dreisigacker S, Peña RJ et al (2015) Genetic characterization of the wheat association mapping initiative (WAMI) panel for dissection of complex traits in spring wheat. Theor Appl Genet 128:453–464
Maccaferri M, El-Feki W, Nazemi G et al (2016) Prioritizing quantitative trait loci for root system architecture in tetraploid wheat. J Exp Bot 67:1161–1178
Pask AJD, Pietragalla J, Mullan DM, Reynolds MP (2012) Physiological breeding II: a field guide to wheat phenotyping. Cimmyt
Pritchard JK, Rosenberg NA (1999) Use of unlinked genetic markers to detect population stratification in association studies. Am J Hum Genet 65:220–228
Quarrie SA, Gulli M, Calestani C et al (1994) Location of a gene regulating drought-induced abscisic acid production on the long arm of chromosome 5A of wheat. Theor Appl Genet 89:794–800
Quarrie SA, Pekic Quarrie S, Radosevic R et al (2006) Dissecting a wheat QTL for yield present in a range of environments: from the QTL to candidate genes. J Exp Bot 57:2627–2637
Ramya P, Chaubal A, Kulkarni K et al (2010) QTL mapping of 1000-kernel weight, kernel length, and kernel width in bread wheat (Triticum aestivum L.). J Appl Genet 51:421–429
Ray DK, Ramankutty N, Mueller ND et al (2012) Recent patterns of crop yield growth and stagnation. Nat Commun 3:1293
Reynolds M, Langridge P (2016) Physiological breeding. Curr Opin Plant Biol 31:162–171
Reynolds M, Pask A et al (2017) Strategic crossing of biomass and harvest index-source and sink- achieves genetic grain in wheat. Euphytica 213(11):257
Röder MS, Huang XQ, Börner A (2008) Fine mapping of the region on wheat chromosome 7D controlling grain weight. Funct Integr Genom 8:79–86
Sayre KD, Rajaram S, Fischer RA (1997) Yield potential progress in short bread wheats in northwest Mexico. Crop Sci 37:36
Schulthess AW, Reif JC, Ling J, Plieske J, Kollers S, Ebmeyer E, Korzun V, Argillier O, Stiewe G, Ganal MW, Röder (2017) The roles of pleiotropy and close linkage as revealed by association mapping of yield and correlated traits of wheat (Triticum aestivum L.). J Ex Bot 68(15):4089–4101
Sehgal D, Autrique E, Singh R et al (2017) Identification of genomic regions for grain yield and yield stability and their epistatic interactions. Sci Rep 7:41578
Sharma RC, Crossa J, Velu G et al (2012) Genetic gains for grain yield in CIMMYT spring bread wheat across international environments. Crop Sci 52:1522
Simmonds J, Scott P, Leverington-Waite M et al (2014) Identification and independent validation of a stable yield and thousand grain weight QTL on chromosome 6A of hexaploid wheat (Triticum aestivum L.). BMC Plant Biol 14:191
Simmonds J, Scott P, Brinton J et al (2016) A splice acceptor site mutation in TaGW2-A1 increases thousand grain weight in tetraploid and hexaploid wheat through wider and longer grains. Theor Appl Genet 129:1099–1112
Sonah H, O’Donoughue L, Cober E et al (2015) Identification of loci governing eight agronomic traits using a GBS-GWAS approach and validation by QTL mapping in soya bean. Plant Biotechnol J 13:211–221
Su Z, Hao C, Wang L et al (2011) Identification and development of a functional marker of TaGW2 associated with grain weight in bread wheat (Triticum aestivum L.). Theor Appl Genet 122:211–223
Sukumaran S, Xiang W, Bean SR et al (2012) Association mapping for grain quality in a diverse sorghum collection. Plant Genome J 5:126. https://doi.org/10.3835/plantgenome2012.07.0016
Sukumaran S, Dreisigacker S, Lopes M et al (2015a) Genome-wide association study for grain yield and related traits in an elite spring wheat population grown in temperate irrigated environments. Theor Appl Genet 128:353–363
Sukumaran S, Reynolds MP, Lopes MS et al (2015b) Genome-Wide association study for adaptation to agronomic plant density: a component of high yield potential in spring wheat. Crop Sci 55:2609–2619
Sukumaran S, Lopes MS, Dreisigacker S et al (2016) Identification of earliness per se flowering time locus in spring wheat through a genome-wide association study. Crop Sci 56:2962–2972
Sukumaran S, Crossa J, Jarquin D, Lopes M, Reynolds MP (2017) Genomic prediction with pedigree and genotype × environment interaction in spring wheat grown in South and West Asia, North Africa, and Mexico. G3 (Bethesda) 7(2):481–495. https://doi.org/10.1534/G3.116.036251
Sun G, Zhu C, Kramer MH et al (2010) Variation explained in mixed-model association mapping. Heredity (Edinb) 105:333–340
Valluru R, Reynolds MP, Salse J (2014) Genetic and molecular bases of yield-associated traits: a translational biology approach between rice and wheat. Theor Appl Genet 127:1463–1489
Valluru R, Reynolds MP, Davies WJ, Sukumaran S (2017) Phenotypic and genome-wide association analysis of spike ethylene in diverse wheat genotypes under heat stress. New Phytol 214(1):271–283
Vargas M, Crossa J, Sayre KD et al (1998) Interpreting genotype × environment interaction in wheat by partial least squares regression. Crop Sci 38:679–689
Vargas M, Combs E, Alvarado G, Atlin G, Mathews K, Crossa J (2013) Meta: a suite of sas programs to analyze multienvironment breeding trials. Agron J 105(1):11–19
Wei T, Simko V (2017) R package “corrplot”: Visualization of a correlation matrix (Version 0.84). Available from https://github.com/taiyun/corrplot. Accessed 12 Dec 2017
Yan L, Loukoianov A, Tranquilli G et al (2003) Positional cloning of the wheat vernalization gene VRN1. Proc Natl Acad Sci USA 100:6263–6268
Yang Z, Bai Z, Li X et al (2012) SNP identification and allelic-specific PCR markers development for TaGW2, a gene linked to wheat kernel weight. Theor Appl Genet 125:1057–1068
Yu J, Pressoir G, Briggs WH et al (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zadoks JC, Chang TT, Konzak CF (1974) A decimal code for the growth stages of cereals. Weed Res 14(6):415–421
Zanke CD, Ling J, Plieske J et al (2014) Whole genome association mapping of plant height in winter wheat (Triticum aestivum L). PLoS One 9(11):e113287
Zhang Z, Ersoz E, Lai C-QQ et al (2010) Mixed linear model approach adapted for genome-wide association studies. Nat Genet 42:355–360
Zhang X, Chen J, Shi C et al (2013) Function of TaGW2-6A and its effect on grain weight in wheat (Triticum aestivum L.). Euphytica 192:347–357
Zhu C, Yu J (2009) Nonmetric multidimensional scaling corrects for population structure in association mapping with different sample types. Genetics 182:875–888
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome J 1:5
Zikhali M, Michelle L-W, Fish L et al (2014) Validation of a 1DL earliness per se (eps) flowering QTL in bread wheat (Triticum aestivum). Mol Breed 34:1023–1033
Acknowledgements
This work was implemented by CIMMYT as part of the MasAgro in collaboration with CIMMYT, made possible by the generous support of SAGARPA, IWYP, and ARCADIA Any opinions, findings, conclusion, or recommendations expressed in this publication are those of the author(s) and do not necessarily reflect the view of SAGARPA, IWYP, and ARCADIA.
Author information
Authors and Affiliations
Contributions
SS, MR, ML conceived the study. SS, MR, SD genotyped the panel. SS did the genetic analysis and wrote the manuscript. All authors read, made constructive comments, and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Mark E. Sorrells.
A correction to this article is available online at https://doi.org/10.1007/s00122-018-3066-x.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2017_3037_MOESM1_ESM.tif
Supplementary material 1 (TIFF 1449 kb) Supplementary Fig. 1. Linkage disequilibrium (LD) plot of the chromosome 3B region showing high LD of the markers associated with the traits. Markers used were RAC875_c1997_2590 (85 cM), RAC875_c5427_447 (91 cM), BobWhite_c35398_181 (95 cM), and wsnp_CAP12_c2297_1121142 (119 cM)
122_2017_3037_MOESM2_ESM.tif
Supplementary material 2 (TIFF 1770 kb) Supplementary Fig. 2. Linkage disequilibrium (LD) plot of the chromosome 6A showing the high LD region (77–85 cM). Markers in chromosome 6A were wsnp_Ra_c61979_62215037 (77 cM), wsnp_Ku_rep_c72681_72356010 (78 cM), wsnp_Ra_rep_c100410_86374467 (79 cM), wsnp_Ku_rep_c112734_95776957 (80 cM), wsnp_Ex_c34545_42832894 (81 cM), wsnp_RFL_Contig4424_5193532 (82 cM), wsnp_Ex_c341_667884 (83 cM), wsnp_Ku_c4296_7807837 (84 cM), wsnp_Ra_c11269_18309313 (85 cM) and Excalibur_rep_c111263_307 (86 cM)
122_2017_3037_MOESM3_ESM.tif
Supplementary material 3 (TIFF 1756 kb) Supplementary Fig. 3. Linkage disequilibrium plot (LD) of the 5A region 90–98 cM showing the SNP at 98 cM is not in high LD with the SNPs from 89–98 cM. Markers in chromosome 5A were wsnp_Ra_c12183_19587379 (89 cM), wsnp_Ex_c5998_10513766 (90 cM), wsnp_Ex_rep_c66689_65010988 (91 cM), wsnp_RFL_Contig2265_1693968 (92 cM), wsnp_Ex_rep_c109532_92292121 (93 cM), wsnp_Ra_c3966_7286546 (94 cM), IAAV108 (95 cM), wsnp_BF484028B_Td_2_1 (96 cM), wsnp_Ex_c790_1554988 (97 cM), and wsnp_Ku_c42416_50159250 (98 cM)
Rights and permissions
About this article
Cite this article
Sukumaran, S., Lopes, M., Dreisigacker, S. et al. Genetic analysis of multi-environmental spring wheat trials identifies genomic regions for locus-specific trade-offs for grain weight and grain number. Theor Appl Genet 131, 985–998 (2018). https://doi.org/10.1007/s00122-017-3037-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-3037-7