Abstract
Key message
Rsc15, a novel locus underlying soybean resistance to SMV, was fine mapped to a 95-kb region on chromosome 6. The Rsc15- mediated resistance is likely attributed to the gene GmPEX14 , the relative expression of which was highly correlated with the accumulation of H 2 O 2 along with the activities of POD and CAT during the early stages of SMV infection in RN-9.
Abstract
Soybean mosaic virus (SMV) causes severe yield losses and seed quality deterioration in soybean [Glycine max (L.) Merr.] worldwide. A series of single dominant SMV resistance genes have been identified on respective soybean chromosomes 2, 13 and 14, while one novel locus, Rsc15, underlying resistance to the virulent SMV strain SC15 from soybean cultivar RN-9 has been recently mapped to a 14.6-cM region on chromosome 6. However, candidate gene has not yet been identified within this region. In the present study, we aimed to fine map the Rsc15 region and identify candidate gene(s) for this invaluable locus. High-resolution fine-mapping revealed that the Rsc15 gene was located in a 95-kb genomic region which was flanked by the two simple sequence repeat (SSR) markers SSR_06_17 and BARCSOYSSR_06_0835. Allelic sequence comparison and expression profile analysis of candidate genes inferred that the gene Glyma.06g182600 (designated as GmPEX14) was the best candidate gene attributing for the resistance of Rsc15, and that genes encoding receptor-like kinase (RLK) (i.e., Glyma.06g175100 and Glyma.06g184400) and serine/threonine kinase (STK) (i.e., Glyma.06g182900 and Glyma.06g183500) were also potential candidates. High correlations were established between the relative expression level of GmPEX14 and the hydrogen peroxide (H2O2) concentration and activities of catalase (CAT) and peroxidase (POD) during the early stages of SMV-SC15 infection in RN-9. The results of the present study will be useful in marker-assisted breeding for SMV resistance and will lead to further understanding of the molecular mechanisms of host resistance against SMV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soybean [Glycine max (L.) Merr] is the most important legume crop in the world due to its high protein and oil contents. Soybean mosaic virus (SMV; Potyvirus), a seed-borne and aphid-transmitted virus, generally infects soybean varieties, causing a typical mosaic pattern in the leaves, top-necrosis, stunted plant growth and seed mottling. Widespread SMV leads to seed quality deterioration and serious yield losses (up to ~50%) in almost all soybean production regions worldwide (Hill et al. 1987; Wrather et al. 2001a, b; Yang et al. 2014).
Based on the expression of disease symptoms in different soybean cultivars (Conover 1948), SMV isolates have been grouped into different strains. Seven pathotypic strains (G1–G7) have been identified in the United States and Korea (Cho and Goodman 1979; Cho et al. 1983), while 22 strains (SC1–SC22) have been identified in China (Yang 2002; Wang et al. 2003, 2004; Guo et al. 2005; Zhan et al. 2006; Li et al. 2010; Wang et al. 2014). However, the pathotypic relationships among these groups have not yet been fully established. Due to host–pathogen co-evolution, resistance-breaking virulent strains have been identified in South Korea, China, Canada and Iran (Choi et al. 2005; Guo et al. 2005; Viel et al. 2009; Ahangaran et al. 2013). In China, SC15 is the most virulent strain among the 22 strains infecting all ten soybean differentials. This strain was first identified from the SMV isolates collected from Shandong and Beijing (Guo et al. 2005). Furthermore, the following identification of the isolates from soybean fields in Southern and Northeastern China have revealed that SC15 is one of the most virulent and widespread strains (Li et al. 2010, 2014). The distribution of the SMV SC15 strain in China is summarized in Fig. 1.
Because breeding soybean cultivars with broad-spectrum and durable resistance is the most cost-effective and eco-friendly approaches for SMV management in soybean production, and many attempts have been made to identify and explore diverse SMV resistance genes in soybeans. Marker-assisted selection (MAS) is one of the key methods assisting both traditional breeding practices as well as resistance gene-mediated transgenic breeding approaches.
SMV resistance in soybean has been known for many years, and the inheritance of resistance has been the subject of several studies. The SMV resistance in soybean is controlled mostly by single dominant genes. Thus far, multiple independent resistance genes with different SMV strain specificities have been identified. Three distinct SMV resistance loci (‘Rsv’) were first identified from PI 96983 (Rsv1), L29 (Rsv3) and V94-5152 (Rsv4) in the United States. These three loci, which underlie extreme resistance (ER) to SMV strains G1–G4, G5–G7 and G1–G7, were identified on chromosomes 2, 13 and 14, respectively (Kiihl and Hartwig 1979; Buzzell and Tu 1989; Yu et al. 1994; Hayes et al. 2000; Jeong et al. 2002; Gore et al. 2002). In addition, many resistance loci involving the resistance to certain Chinese SMV strains (‘Rsc’) from soybean cultivars Kefeng No. 1 (Rsc7 and Rsc8, etc.), Qihuang No. 1 (Rsc3 and Rsc14) and Dabaima (Rsc4) were also positioned on chromosome 2, 13 and 14, respectively (Yan et al. 2015; Li et al. 2014; Ma et al. 2011; Zheng et al. 2014; Ma et al. 2010; Li et al. 2012; Wang et al. 2011a, b; Zhao et al. 2016). In a previous study, we screened 957 simple sequence repeat (SSR) markers throughout the soybean genome (Song et al. 2004) using bulked segregant analysis (BSA) of 200 F7:10 recombinant inbred lines (RILs). We found that novel resistance locus Rsc15 was located within a 14.6-cM (~1.6-Mb) genomic interval flanked by Sat_213 and Satt286 on chromosome 6 (Yang and Gai 2011) (Figs. 2a, 3b, c). Although the ‘Rsv’ and ‘Rsc’ genes are located in the vicinity of each other on the genome, the allelic relationships between these genes remain unclear.
Genetic linkage map of Rsc15 with the F2, RIL population derived from RN-9×7605. a Linkage map of Rsc15 constructed by Yang et al. with the RIL population. b, c Linkage maps of Rsc15 constructed with the F2 and RIL populations in this study. d Linkage maps of Rsc15 using the RIL population genotyped with the additional 17 SSR markers. Genetic distances between the neighboring markers are on the left side of linkage groups and the marker names are shown on the right side
Genetic and physical maps of SMV resistance locus (Rsc15) and transcriptions of candidate resistance genes. a Putative position of SMV resistance locus (Rsc15) and quantitative trait loci (QTLs) of other traits on chromosome 6. Markers are arranged according to the linkage map of soybean chromosome 6 constructed by Song et al. (2004). The blue short bars indicate the putative position of QTLs for seed size traits (Xu et al. 2011), flowering time (qFT6) (Zhang et al. 2013) and maturity (T and E1) (Molnar et al. 2003). The red short bars indicate the putative location of QTLs for the resistance of soybean sudden death syndrome (SDS) (qFDS 003,004) (Njiti et al. 1998; Abdelmajid et al. 2012) and Asian soybean rust (ASR) [Rpp3 and Rpp? (Hyuuga)] (Hyten et al. 2009; Monteros et al. 2007; Hossain et al. 2015). b Genetic maps of SMV resistance locus Rsc15. SSR markers were arranged above the bar according to the linkage map of the RIL population (Fig. 2d). Numbers under the line indicate the recombinants detected between the corresponding markers and the resistance gene. c Physical map position of Rsc15. Numbers at the foot of the markers under the line indicate the physical start position of corresponding markers on chromosome 6 according to the Williams 82 reference whole genomic sequence (WGS) (Glyma Wm 82.a2.v1 accessible at http://soybase.org/). d Transcription models of Rsc15 candidate genes. Blue and green arrows indicate the positions of RLK and STK-encoding genes, respectively. Totally, 5 putative genes existed in the target genomic region of Rsc15 flanked by the two SSR markers SSR_06_17 and BARCSOYSSR_06_0835, according to the soybean reference genome annotation (Glycine max Wm82.a2.v1, accessible at http://soybase.org/, December 2016) (color figure online)
The availability of soybean reference whole genome sequence (WGS) has laid the basis for the development of new markers and identification of candidate genes. With high-density markers developed by Song et al. (2010), many SMV resistance loci have been delimited into meticulous regions, revealing the likely involvement of disease-resistant genes. Researchers discovered that the ER to SMV mediated by Rsv1, Rsv3 and Rsc4 is likely attributed to one or more members of the NBS-LRR class of disease-resistant genes (Hayes et al. 2004; Shi et al. 2008; Zhang et al. 2012; Yang et al. 2013; Ma et al. 2016; Suh et al. 2011; Wang et al. 2011a; Li et al. 2016). However, Rsv4, Rsc7 and Rsc8 are not similar to other resistance genes and belong to an original class of disease R-genes (Maroof et al. 2010; Ilut et al. 2016; Yan et al. 2015; Wang et al. 2011b; Zhao et al. 2016). Despite studies focussing primarily on resistance gene identification for decades, none of SMV resistance gene has been cloned.
Cloned plant disease-resistance genes share conservative architectural features, and majority of these genes, such as tobacco mosaic virus (TMV) resistance gene N in tobacco (Whitham et al. 1994) and a strong candidate gene 3Gg2 for Rsv1 (Hayes et al. 2004), belong to the NBS-LRR classification. Following NBS-LRR, receptor-like kinases (RLKs) and serine/threonine kinases (STKs) are the second most abundant type of resistance gene classes, such as Cladosporium fulvum resistance genes Cf-9 and Cf-2 in tomato (Jones et al. 1994; Dixon et al. 1996); Xanthomonas oryzae resistance genes Xa21 and Xa26 in rice (Song et al. 1995; Sun et al. 2004), Pseudomonas syringae resistance gene Pto in tomato (Martin et al. 1993), Puccinia triticina resistance gene Lrk10 in wheat (Feuillet et al. 1997) and Puccinia graminis resistance gene Rpg1 in barley (Brueggeman et al. 2002). These conservative architectural features of resistance genes would provide references for the identification of SMV resistance genes.
Virus infection of plants is often accompanied by a burst of reactive oxygen species and changes in the levels of endogenous phytohormones. The levels of hydrogen peroxide (H2O2) and superoxide (O2 −) are increased with the spontaneous cell death and they activate defense responses in plants (Apel and Hirt 2004; Gadjev et al. 2006; Hernández et al. 2016; Liu et al. 2011). In addition, stress and defense-related proteins, such as pathogenesis-related proteins, peroxidase (POD) and catalase (CAT), are involved in host response to SMV infection (Yang et al. 2011; Liu et al. 2011). The phytohormone salicylic acid (SA) is a signaling molecule of the systemic acquired resistance pathway involved in host response to SMV infection (Sandhu et al. 2009; Hajimorad and Hill 2001), and abscisic acid (ABA) can also induce GmPP2C3a to function as a key regulator of Rsv3-mediated ER against SMV (Seo et al. 2014). Exploring the involvement of H2O2, POD, CAT and phytohormones in SMV infection would facilitate the clarifying of the molecular mechanisms of host resistance against the SMV in soybean.
Based on the previous study, the objectives of the present study were to: (1) fine map the Rsc15 gene region on chromosome 6; (2) identify candidate genes for Rsc15 using sequencing and expression analyses in the presence of SMV; and (3) explore the likely involvement of resistance mediated by Rsc15.
Materials and methods
Plant materials and mapping populations
Soybean cultivar RN-9 contains the Rsc15 gene conferring resistance to SMV strain SC15 (Yang and Gai 2011). The popular cultivars 7605 and Nannong 1138-2 are susceptible to all SMV strains (SC1-SC22). The F1, F2 (315 F2 individuals) and RIL population (comprising 216 F7:11 lines) derived from the cross of RN-9×7605 were used to verify the previous results (Table 1), and the RILs were used for the fine-mapping of Rsc15. A detailed account of the RIL population is provided in Yang and Gai (2011). The soybean seeds of the two parents along with the F1, F2 and RIL populations were planted in plastic pots (20 cm × 20 cm) in aphid-free greenhouses. For each line, a total of 15–20 seeds were planted. All seed materials were obtained from the National Center for Soybean Improvement (NCSI; Nanjing Agricultural University, China).
Inoculation and resistance evaluation
SMV strain SC15, which was previously used by Yang and Gai (2011) to map Rsc15, was used in the present study. The SC15 strain was obtained by the NCSI and maintained in leaves of the susceptible cultivar Nannong 1138-2. Young symptomatic leaves were ground using a sterilized mortar and pestle in sodium phosphate-buffered saline (PBS, 0.01 M, pH 7.4) at a ratio of 1:2 (w/v) mixed with a small amount of carborundum powder (600-mesh). Fully expanded primary leaves of soybean plants were mechanically inoculated by gently rubbing with the inoculum using a paintbrush (Li et al. 2010). Immediately after inoculation, tap water was sprayed to the inoculated leaves. Pesticides were regularly sprayed onto the plants to prevent cross-infection via aphids.
All plants of the parents, and the F1, F2 and RIL populations were evaluated for virus resistance through weekly visual assessment with detailed scoring in 40 days post-inoculation (dpi). Any inoculated plants that showed a mosaic pattern or chlorotic symptoms in upper leaves were considered as susceptible (S), while those that were symptomless and necrotic were classified as resistant (R) to the virus (Buzzell and Tu 1984; Chen et al. 1991).
DNA isolation, new SSR marker development and genotyping
Young trifoliate leaves from parents and the mapping populations were collected and ground using a mortar and pestle in liquid nitrogen. Total genomic DNA was isolated using a modified cetyltrimethyl ammonium bromide method (Doyle 1990).
A total of 15 SSR markers (i.e., Satt640, Satt227, Sat_153, Satt322, Sat_213, Sat_246, Satt286, Satt277, Satt557, Satt658, Satt100, Satt708, Sat_238, Staga001 and Satt433) on soybean chromosome 6 were selected and used to verify the previous mapping results. For this purpose, the F2 and RIL populations were used. After verifying the position of Rsc15, fine-mapping was performed in the RIL population.
A total of 132 SSR markers flanking the target region, including 108 available SSR markers (Song et al. 2010) and 24 newly developed SSR markers, were used to shorten the Rsc15 flanking interval. Briefly, for SSR development, the targeted region of the DNA sequence was obtained from the WGS (http://www.phytozome.net/soybean), and SSRs were identified using SSRHunter 1.3 software (Li and Wan 2005). The primer pairs were designed using Primer Premier 5.0 software (Premier, Palo Alto, CA, USA) (Supplement S1). SSR markers were screened using polymerase chain reaction (PCR), polyacrylamide gel electrophoresis and silver staining following the method used by Wang et al. (2011a).
The phenotypic data (the resistance gene served as a morphological marker) and genotypic data of the F2 and RIL populations were used to construct linkage maps. Genetic distances between SMV resistance loci and markers were calculated using ‘Join Map 4.0′ linkage analysis software (Van Ooijen 2006). MapChart 2.2 (Voorrips 2002) was used for map construction and modification. The markers were assigned to linkage groups (LGs) using a minimum logarithm of likelihood ratio (LOD) score of 3.0 as a criterion to test the linkage.
Collection of various tissues and stress treatments
The roots, stems and leaves were collected from RN-9 plants at the V2 stage; flowers were collected at the R2 stage; and immature pods were collected at the R5 stage. Plant stress treatments were performed at the VC stage. Mock-inoculated (inoculated with PBS) and SMV-infected RN-9 and 7605 leaves were collected at 0, 1, 2, 4, 8, 12 and 24 h post-inoculation (hpi). RN-9 seedlings were used for SA, jasmonic acid (JA), ABA, ethylene (ET) and cold-stress treatments. The seedlings were sprayed with SA (2 mM SA) and JA (100 μM methyl-jasmonate dissolved in 0.01% ethanol), ABA (200 μM ABA) and ET (100 μM 1-aminocyclopropane-1-carboxylic acid). For chilling treatments, the seedlings were placed in a 4 °C growth chamber, and the leaves were harvested at 0, 1, 2, 3, 6, 12, 24, 36, 48, 72 h post-treatment (hpt). Each sample was independently collected with three biological replicates and stored at −80 °C immediately after freezing in liquid nitrogen. Total RNA was extracted from various tissues, and quantitative real-time PCR was conducted.
Candidate gene prediction and quantitative real-time PCR (qRT-PCR)
Putative candidate gene models in the identified flanking region were predicted according to the WGS (https://soybase.org/gb2/gbrowse/gmax2.0/; Glycine max Wm82.a2.v1; verified December, 2016). Gene-specific primers for qRT-PCR were designed from the respective gene transcript sequences (http://www.phytozome.net/soybean) using Primer-BLAST software (https://www.ncbi.nlm.nih.gov/tools/primer-blast/) (Ye et al. 2012). The gene Tubulin (accession No. AY907703) was used as an internal reference control to normalize the total amount of cDNA in each reaction (Supplement S1). PCR was conducted, and only primers that amplified a single product were selected for qRT-PCR.
Tissues with different treatments were collected, and total RNA was extracted using an RNA Simple Total RNA Kit (Tiangen, Beijing, China). Subsequently, the DNA-free RNA was used for first-strand cDNA synthetization using Oligo (dT) primers and a PrimeScript™ И 1st strand cDNA Synthesis Kit 6210A (Takara, Dalian, China) following the manufacturer’s instructions. The qRT-PCR was performed in a 20-μL reaction volume comprising 2.0 μL (approximately 50 ng) of 10× diluted first-strand cDNA, 0.8 μL of each primers (10.0 µM), 10.0 μL of 2× SYBR Green I Master Mix and 6.4 μL of sterile distilled H2O (Takara, Dalian, China). All reactions were performed in 96-well reaction plates using a LightCycler 480® Real-Time PCR System (Roche, Germany) with three technical replicates. The following PCR conditions were used: 95 °C for 5 min, followed by 40 cycles at 95 °C for 15 s, 60 °C for 30 s and 72 °C for 30 s, and then at 68 °C for 5 min.
The expression of candidate genes was quantified using the relative quantification (2−∆∆CT) method (Wang et al. 2011b). The ratios of relative expression levels between samples infected with SMV and samples inoculated with PBS were regarded as a calibration of the relative expression levels induced by SMV infection alone at different time points.
Sequence alignments of candidate resistance genes
For sequence alignments of candidate genes, gene-specific primers were designed according to the promoter, genomic and transcript sequences of each gene models using Primer Premier 5.0 software (Premier, Palo Alto, CA, USA) (Supplement S1). The fragments for each gene were amplified using the DNA or cDNA of RN-9 and 7605. Purified PCR products were cloned into the pMD19-T vector (Takara, Dalian, China) according to the manufacturer’s instructions, and sequencing was subsequently performed at Invitrogen Biotechnology Co., Ltd (Shanghai, China). Sequence alignments and polymorphism evaluation were performed using DNAMAN™ (LynnonBiosoft version 8.0, Pointe-Claire, Canada). The sequences were analyzed for open reading frames using GENESCAN (http://genes.mit.edu/GENSCAN.html) and protein domains were annotated using SMART (http://smart.embl-heidelberg.de) and InterProScan (http://www.ebi.ac.uk/InterProScan).
Detection of H2O2 and enzyme activity assays
The inoculated leaves of RN-9 and Nannong 1138-2 (inoculated with SC15 and PBS) were collected at 0, 1, 2, 3, 6, 12, 24, 36, 48 and 72 hpi. Subsequently, the samples were ground and suspended in 0.9 mL PBS (pH 7.4) and centrifuged at 4 °C, 2500 rpm for 10 min. The resulting supernatants were collected for H2O2 determination and enzyme activity assays. The H2O2 concentration was detected by monitoring the absorbance of the titanium-peroxide complex at 405 nm using the determination kit [A064-1; Nanjing Jiancheng Bioengineering Institute (NJBI), Nanjing], and the CAT and POD activities were assayed using the determination kit (A007-1 and A084-3; NJBI, Nanjing) according to the manufacturer’s instructions (Li et al. 2013; Yong et al. 2008).
Statistical analyses
The observed R:S ratios were determined based on the goodness-of-fit for the expected ratios tested using Chi-square (χ 2) analysis with a P value greater than 0.05. The Chi-square (χ2) test and correlation analyses were both conducted using SAS (SAS Institute v. 9.2).
Results
Response of parents, F1, F2 and RILs populations to SC15
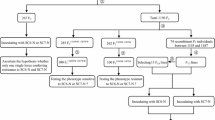
Responses of the parents and different populations to SC15 are summarized in Table 1. The results confirmed that RN-9 was R, but that Nannong 1138-2 and 7605 were S to SC15. The F1 plants generated with RN-9 and 7605 showed the same response to SC15 with RN-9, suggesting the dominant nature of SC15 resistance in RN-9. Through self-pollination of F1 plants, we generated 315 F2 plants including 247 R and 68 S plants. The Chi-squared text indicated that this segregation pattern was consistent with a Mendelian segregation ratio of 3:1 (R:S). Thus, SC15 resistance derived from RN-9 is controlled by a single dominant gene. Additionally, the results for the RIL population were also consistent with an expected Mendelian inheritance ratio of 1:1 (R:S). This finding further demonstrated that SC15 resistance was controlled by one dominant gene, consistent with the report of Yang and Gai (2011).
Fine-mapping of Rsc15 with the RILs
The objective of the present study was to fine map Rsc15, the locus previously described by Yang and Gai (2011). The Rsc15 locus was mapped within a 14.6-cM (~1.6-Mb) genomic region flanked by Sat_213 and Satt286 on chromosome 6, using 200 RILs and a genetic map comprising ten SSRs. To verify the map location of the Rsc15 gene, 4 and 15 SSR markers were screened in the F2 and the RIL population (216 lines), respectively. The genotypic data were used for linkage analysis, and the results revealed that the resistance gene was located between Sat_246 and Satt286 (Fig. 2b, c). These results were consistent with those of Yang and Gai (2011).
To fine map Rsc15, new markers were added around and within the original map interval using RILs. A total of 108 SSR markers flanking Rsc15, developed by Song et al. (2010), were used to screen the parental lines. Among the 108 SSRs tested, 16 markers exhibited polymorphisms between the parents. These markers were used to genotype the RILs, and Rsc15 was delimited within the interval flanked by BARCSOYSSR_06_0830 and BARCSOYSSR_06_0835. To further narrow down the region, a total of 24 new SSR markers in the targeted region were developed, and two of the markers (SSR_06_15 and SSR_06_17) were polymorphic. RILs were subsequently used for genotyping analysis. The result revealed that Rsc15 was located within the 95-kb region in the Williams 82 reference genome (Glyma Wm 82.a2.v1), and it was flanked by the two markers SSR_06_17 and BARCSOYSSR_06_0835 at genetic distances of 2.38 and 1.66 cM, respectively (Figs. 2d, 3b, c).
Identification of candidate resistance genes through allelic sequence comparison
To identify the likely candidate genes of Rsc15, the target region was examined for predicted genes according to the soybean reference genome annotation database (Glycine max Wm82.a2.v1 accessible at https://soybase.org/gb2/gbrowse/gmax2.0/; December 2016). A total of 136 genes were identified between Sat_246 (physical start position Gm06:14487532) and Satt286 (Gm06: 16221141), but 23 genes were not annotated for the loci (Supplement S2; Fig. 3c). None of the remaining 113 genes was found to belong to NBS-LRR family, while two genes encoded RLK (i.e., Glyma.06g175100 and Glyma.06g184400) and five genes encoded STK (i.e., Glyma.06g176800, Glyma.06g177700, Glyma.06g182300, Glyma.06g182900 and Glyma.06g183500) (Fig. 3c).
In the 95-kb Rsc15 region, five gene models were predicted (Fig. 3d; Table 2). Gene Glyma.06g182500 encodes a protein of unknown function (DUF1645); Glyma.06g182600 (designated as GmPEX14) encodes a peroxisomal membrane anchor protein; Glyma.06g182700 encodes a predicted carbonic anhydrase involved in protection against oxidative damage; Glyma.06g182800 encodes a tetratricopeptide repeat (TPR)-like superfamily protein; and Glyma.06g182900 encodes an STK (Table 2). Allelic sequence comparison was conducted using the DNA and cDNA sequences from the two parents. The results showed that all of these five candidate genes were conserved in the coding sequence (CDS) with their allelic counterparts, and the three genes (Glyma.06g182700, Glyma.06g182800 and Glyma.06g182900) were also conserved in promoter sequences, while a nucleotide insertion/deletion mutation (InDel; 8 bp) and a single nucleotide polymorphism (SNP) were detected in the promoter sequence of Glyma.06g182600 (Supplement S3). Moreover, the CDS of all three STK-encoding genes (Glyma.06g176800, Glyma.06g177700 and Glyma.06g183300) were conserved between the two parents (Supplement S3). In contrast, 3 and 6 SNPs were identified in the CDS or promoter of Glyma.06g175100 and Glyma.06g183500 between the two parents, respectively. The gene model Glyma.06g175100 encodes an RLK protein containing one signal peptide, one LRR domain, one low complexity region, one TM region and one STK domain, and only one of the 3 SNP resulted in an R203H AA substitution between the two parents (Supplement S3). The gene Glyma.06g183500 encodes an STK protein containing one S_TKc domain, and 6 SNPs resulted in a V371A AA substitution between the two parents (Supplement S3). For another RLK-encoding gene, Glyma.06g184400, a single base-pair deletion close to the initiation codon delayed the transcription in 7605 (Supplement S3). Full-length PCR products of Glyma.06g182800 could not be obtained from the two parents using four pairs of specific primers.
Taken together, among the five genes predicted in the identified interval, GmPEX14 (Glyma.06g182600) was the only gene possessing polymorphisms in the aligned DNA or cDNA sequences. Moreover, the RLK and STK genes (namely, Glyma.06g175100, Glyma.06g183500 and Glyma.06g184400), which exhibited different polymorphisms were also potential candidate genes.
Expression analysis of candidate resistance genes using qRT-PCR
Based on architectural features, these five genes predicted in the resistance locus, along with 12 other genes near Rsc15 (genes encoding RLK, STK or disease-resistance-related protein), were further selected for expression analysis. The results showed that most of these genes could be induced by SMV infection with different magnitudes of relative expression levels (Fig. 4; Supplement S5a). For the five genes located in the Rsc15 flanking region, only GmPEX14 (Glyma.06g182600) was up-regulated in RN-9 from 1 to 12 hpi, while its expression was significantly suppressed in 7605. The remaining four genes displayed almost the same expression profile in the two parents after inoculation. Other plant immunity-related genes near Rsc15 could, to some extent, be induced by SMV infection. For instance, Glyma.06g175700 (E3 ubiquitin ligase) was significantly up-regulated in RN-9 from 4 to 12 hpi; Glyma.06g181300 (F-box family protein) and Glyma.06g183300 (zinc finger protein, C3HC4 type) were both significantly up-regulated from 8 to 24 hpi, but there was no significant difference in the expression level of Glyma.06g185400 (U-box domain containing protein). In summary, the results revealed that 8 of the 17 candidate genes (Glyma.06g175100, Glyma.06g176800, Glyma.06g177700, Glyma.06g181300, Glyma.06g182300, Glyma.06g182600, Glyma.06g183300 and Glyma.06g183500) showed significant differences in expression profiles between the two parents in response to SMV infection.
Expression analyses of 11 of 17 selected candidates by qRT-PCR. a, b The two RLK-encoding genes (namely Glyma.06g175100 and Glyma.06g175100). c–g the five genes predicted in the identified interval (Glyma.06g182500, Glyma.06g182600, Glyma.06g182700, Glyma.06g182800 and Glyma.06g182900). h–k The four STK-encoding genes (Glyma.06g177700, Glyma.06g182300, Glyma.06g183500, Glyma.06g176800). Y-axes indicate the ratios of relative expression levels between samples infected with SMV and samples inoculated with PBS (phosphate buffer saline); X-axes indicate the time points post-inoculation (hpi; 0–24 hpi); data are expressed as means of three biological replicates with error bars indicating the standard deviation (SD)
Phytohormones, such as SA and ABA, play important roles as signaling molecules of the resistance pathway against SMV (Hajimorad and Hill 2001; Sandhu et al. 2009; Seo et al. 2014; Liu et al. 2011). The expression of candidate genes, including RLK- and STK-encoding genes, and Glyma.06g182600, was further examined under various stress treatments, and all eight genes showed significant differences on average (Fig. 5). For example, Glyma.06g175100 was up-regulated by approximately 20-fold at 12 hpt with SA and ACC, while Glyma.06g182600 was up-regulated by approximately 2.5-fold from 3 to 48 hpt with ABA. The results showed that these genes responded to the different phytohormones at different time intervals. Further, the expression profiles of these genes were examined in specific tissues (root, stem, leaf, flower and immature pod) (Fig. 6). The results revealed that the expression of these genes in the leaves was generally higher than that in other tissues and that the relative expression of Glyma.06g182300 was much lower compared with that of the other tested genes.
Expression analyses of the four selected candidates under various stress treatments (SA, JA, ABA, ET and cold) by qRT-PCR. Y-axes indicate the ratios of relative expression levels between samples under various stress treatments (SA, JA, ABA, ET and cold) and control at different time points (0–48 hpi); data are expressed as mean of three biological replicates with error bars indicating the SD
Expression analysis of the eight selected candidates in different tissues (root, stem, leaf, flower and immature pod) by qRT-PCR. Y-axes indicate the relative expression levels in different tissues (root, stem, leaf, flower and immature pod) at different time points (0–9 days); data are expressed as the mean of three biological replicates, with error bars indicating the SD
Overall, expression analysis of candidate resistance genes revealed that 8 of the 17 candidate genes (namely Glyma.06g175100, Glyma.06g176800, Glyma.06g177700, Glyma.06g181300, Glyma.06g182300, Glyma.06g182600, Glyma.06g183300 and Glyma.06g183500) showed significant differences in expression profiles between the two parents under various stress treatments, including SMV infection.
Effects of H2O2, CAT and POD on the expression of GmPEX14 in SMV infection
H2O2 detection and enzyme activity assays were conducted, and the results revealed that H2O2 concentration and activities of CAT and POD were all involved in SMV infection. Overall, the levels of endogenous H2O2 in RN-9 were higher than those in Nannong 1138-2, and they were significantly increased from 2 to 12 hpi in RN-9 infected with SC15 compared with the mock control (Fig. 7a). The CAT activity, an antioxidant enzyme that specifically catalyzes the degradation of H2O2, was negatively correlated with the levels of endogenous H2O2. The CAT activity in Nannong 1138-2 was higher than that of in RN-9, and two peaks were observed in Nannong 1138-2 infected with SC15 at 2 and 24 hpi, while the levels remained relatively stable in the mock control (Fig. 7b). POD is an antioxidant enzyme that promotes the removal of peroxides, including H2O2. However, no significant difference in POD activity was detected between Nannong 1138-2 and RN-9 infected with SC15 (Fig. 7c). These results indicated that H2O2, CAT and POD might be involved in SMV infection during the early stages (approximately 2–12 hpi).
Because GmPEX14 was significantly up-regulated from 1 to 12 hpi, accompanied by a burst of H2O2 in the SMV-resistant genotype RN-9 (Figs. 5d, 7a), we further investigated the relationship between the expression level of GmPEX14 and the involvement of the H2O2 concentration along with CAT and POD activities in SMV infection (Table 3). High correlations were established between the relative expression of GmPEX14 and the H2O2 concentration along with activities of CAT and POD at the early stages of SMV infection in RN-9 (0–12 hpi) (Fig. 8). However, it was difficult to establish any correlation either during the other time intervals in RN-9 and during the same time interval in the susceptible genotype Nannong 1138-2 (data not shown).
Correlations between the concentration of H2O2, the activities of CAT and POD and the relative expression of PEX14 in RN-9 at different hours post-inoculation (0–12 hpi). “E” represents the relative expression of PEX14; “C” indicates the concentration of H2O2; “CAT” and “POD” stands for the activities of CAT and POD, respectively
Discussion
Among the 22 SMV strains have been characterized in China, SC15 is the most virulent and widely distributed strain (Fig. 1). The popular soybean cultivars Kefeng No. 1 and Qihuang No. 1 are resistant to most of the Chinese strains, but are susceptible to SC15 (Guo et al. 2005; Li et al. 2010, 2014). Yang and Gai (2011) conducted a large-scale screening for SC15 resistance sources in 205 diverse soybean accessions originating from China and other countries. This screening test showed that ten accessions, including RN9 with ER to SC15, were obtained. Further inheritance analyses and mapping studies indicated that one single dominant gene controls the resistance to SC15 in RN-9, and that this gene was located within the 14.6-cM (~1.6-Mb) genomic region flanked by Sat_213 and Satt286 on chromosome 6 (Yang and Gai 2011). The present study also confirmed these results, consistent with the report by Yang and Gai (2011). Previously, many ‘Rsv’ and ‘Rsc’ genes for certain SMV strains have been identified on chromosomes 2, 13 and 14, respectively. The Rsc15 gene is an interesting novel locus underlying resistance gene to the virulent strain SC15 and is the first SMV resistance gene identified on chromosome 6.
In the present study, we further fine mapped the Rsc15 gene and identified the candidate resistance genes. Using SSR markers to perform high-resolution fine-mapping in the modified RIL population, we narrowed the genomic region from 1.6 Mb to 95 kb, which was flanked by the SSR markers SSR_06_17 and BARCSOYSSR_06_0835 (Figs. 2d, 3b, c). Here, we also identified numerous SSR markers closely linked to Rsc15. These markers could be used as effective and consistent tool that enable traditional breeders to tag the genetic loci linked to SMV resistance.
To our knowledge, there are many QTLs for seed size traits (i.e., qSWT, qSLW, qSW and qST) (Xu et al. 2011), and flowering time (qFT6) (Zhang et al. 2013), and maturity genes (T and E1) (Molnar et al. 2003) distributed in different regions on chromosome 6. In addition, many QTLs for resistance to sudden death syndrome (SDS) (qFDS 003, 004) (Njiti et al. 1998; Abdelmajid et al. 2012), Asian soybean rust (ASR) [Rpp3 and Rpp? (Hyuuga)] (Monteros et al. 2007; Hyten et al. 2009; Hossain et al. 2015), soybean cyst nematode (SCN) (Concibido et al. 2004) and White mold (Huynh et al. 2010) were also located in different regions close to Rsc15 on chromosome 6 (Fig. 3a). Previously, Kang et al. (2012) reported that 23 NBS-LRR genes are located on this chromosome, and the resistance of Rpp? (Hyuuga) is likely attributed to a cluster of NBS-LRR genes (Hossain et al. 2015). Therefore, we examined the genomic region around the putative resistance locus (Rsc15), flanked by Sat_246 (Gm06:14487532) and Satt277 (Gm06:17218711), for predicted genes. The results revealed that no genes belonged to the NBS-LRR family, but that three RLK- and six STK-encoding genes shared conservative architectural features with other cloned R-genes (Supplement S2). Moreover, these RLK- and STK-encoding genes (namely, Glyma.06g175100, Glyma.06g183500 and Glyma.06g184400) showed polymorphisms in the DNA or cDNA sequences, while the other RLK- and STK-encoding genes were all conserved in the CDS and promoter sequences (Table 2; Supplement S3). In addition, two genes (Glyma.06g175100 and Glyma.06g183500) both showed significant differences in expression profiles between the two parents during SMV infection and under various stress treatments. Therefore, these RLK and STK genes were potential candidate genes. These data indicated that Rsc15 was located in a disease-resistance gene-poor region, and that the resistance associated with this locus might be attributed to an original class of disease-resistant genes, similar to that of Rsv4, Rsc7 and Rsc8 (Maroof et al. 2010; Ilut et al. 2016; Yan et al. 2015; Wang et al. 2011b; Zhao et al. 2016).
Despite none of candidate genes was annotated to the NBS-LRR resistance gene family (Table 2), Glyma.06g182600 (GmPEX14) was inferred as a strong candidate gene governing the Rsc15 resistance. Among the five genes predicted in the identified interval, only GmPEX14 possessed polymorphisms in the DNA/cDNA sequence alignments (1 SNP and 1 InDel in the promoter sequence) (Supplement S3). Meanwhile, GmPEX14 showed significant differences in expression profiles in response to SMV infection between the two parents (Fig. 4d), and could be induced by JA, ABA and cold treatments (Fig. 5). However, whether the variations in cis-regulatory elements in the promoter sequence (~2000 bp) lead to differences in plant gene expression and regulation should be further explored. To date, no SMV resistance gene has been cloned, and a greater concentration in the delimitation of mapping intervals is required to verify the likely involvement and perform cloning of SMV resistance genes.
The strong candidate gene Glyma.06g182600 was annotated to encode the peroxisomal membrane protein PEX14. PEX14 is the sole peroxin that has a unique dual function in peroxisome formation and selective degradation (van Zutphen et al. 2008). The formation and selective degradation of peroxisome play a key role in the maintenance of cellular oxidative balance (Nordgren and Fransen 2014). In our study, the relative expression level of GmPEX14 was highly correlated with the concentration of H2O2 and the activities of CAT and POD during the early stages of SMV infection in RN-9 (Fig. 8). As known, H2O2 could trigger programmed plant cell death during the hypersensitive response mediated by R-genes (Chen et al. 2017), acting as a signal in the induction of SAR and inducing defense-related genes (Apel and Hirt 2004; Gadjev et al. 2006; Hernández et al. 2016). As well, plant cells contain oxygen radical-detoxifying enzymes, such as CAT, POD and peroxin, that protect them from oxidative damage. Taken together, these results suggest that PEX14 might confer the resistance to SC15 in RN-9 through the involvement of cellular oxidative balance, in which CAT and POD might also be involved. However, more efforts are necessary to clarify the relationship between the involvement of peroxide and the resistance.
In summary, our research presents evidence that the SSR markers SSR_06_17 and BARCSOYSSR_06_0835 are two markers that delimit a 95-kb region in the reference Williams 82 genome, flanking the Rsc15 locus. In such a region, the gene GmPEX14 (Glyma.06g182600) is the best candidate gene attributed for the resistance of Rsc15, and the four RLK and STK genes (i.e., Glyma.06g175100, Glyma.06g182900, Glyma.06g183500 and Glyma.06g184400) are also potential candidates. Peroxide, particularly H2O2, might be involved in the resistance mediated by Rsc15 during the early stages of SMV infection. Moreover, the markers that are closely linked to SMV resistance locus Rsc15, as determined in this study, will be potentially used for MAS in soybean breeding programs and the pyramiding of resistance genes.
Author contribution statement
HZ, KL, QY and RR designed the methods and experiments. RR, SL, JY, TW, HN, AK, YY, and LW conducted SMV inoculations and resistance evaluation. RR, SL and TW performed genotype analysis. RR, SL, and JY performed qRT-PCR and sequencing. RR, SL and YY conducted the H2O2 detection and enzymes activity assays. RR and KL analyzed the data. HZ, KL, RR and AK drafted the manuscript. All authors critically reviewed and approved the final manuscript.
References
Abdelmajid KM, Ramos L, Leandro L, Mbofung G, Hyten DL, Kantartzi SK, Grier I, Robert L, Njiti VN, Meksem K (2012) The ‘PI 438489B’by ‘Hamilton’ SNP-based genetic linkage map of soybean [Glycine max (L.) Merr.] identified quantitative trait loci that underlie seedling SDS resistance. J Plant Genome Sci 1:18–30
Ahangaran A, Habibi MK, Mohammadi G-HM, Winter S, García-Arenal F (2013) Analysis of Soybean mosaic virus genetic diversity in Iran allows the characterization of a new mutation resulting in overcoming Rsv4-resistance. J Gen Virol 94:2557–2568
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Brueggeman R, Rostoks N, Kudrna D, Kilian A, Han F, Chen J, Druka A, Steffenson B, Kleinhofs A (2002) The barley stem rust-resistance gene Rpg1 is a novel disease-resistance gene with homology to receptor kinases. Proc Natl Acad Sci 99:9328–9333
Buzzell R, Tu J (1984) Inheritance of soybean resistance to soybean mosaic virus. J Hered 75:82
Buzzell R, Tu J (1989) Inheritance of a soybean stem-tip necrosis reaction to soybean mosaic virus. J Hered 80:400–401
Chen P, Buss G, Roane C, Tolin S (1991) Allelism among genes for resistance to soybean mosaic virus in strain-differential soybean cultivars. Crop Sci 31:305–309
Chen H, Adam Arsovski A, Yu K, Wang A (2017) Deep sequencing leads to the identification of eukaryotic translation initiation factor 5a as a key element in Rsv1-mediated lethal systemic hypersensitive response to Soybean mosaic virus infection in Soybean. Mol Plant Pathol 18:391–404
Cho E-K, Goodman RM (1979) Strains of soybean mosaic virus: classification based on virulence in resistant soybean cultivars. Phytopathology 69:467–470
Cho E, Choi S, Cho W (1983) Newly recognized soybean mosaic virus mutants and sources of resistance in soybeans. The research report of the Office of Rural Development Soil fertilizer, crop protection, mycology and farm products utilization
Choi B, Koo J, Ahn H, Yum H, Choi C, Ryu K, Chen P, Tolin S (2005) Emergence of Rsv-resistance breaking Soybean mosaic virus isolates from Korean soybean cultivars. Virus Res 112:42–51
Concibido VC, Diers BW, Arelli PR (2004) A decade of QTL mapping for cyst nematode resistance in soybean. Crop Sci 44:1121–1131
Conover RA (1948) Studies of two viruses causing mosaic diseases in Soybean. Phytopathology 38:724–735
Dixon MS, Jones DA, Keddie JS, Thomas CM, Harrison K, Jones JD (1996) The tomato Cf-2 disease resistance locus comprises two functional genes encoding leucine-rich repeat proteins. Cell 84:451–459
Doyle JJ (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Feuillet C, Schachermayr G, Keller B (1997) Molecular cloning of a new receptor-like kinase gene encoded at the Lr10 disease resistance locus of wheat. Plant J 11:45–52
Gadjev I, Vanderauwera S, Gechev TS, Laloi C, Minkov IN, Shulaev V, Apel K, Inzé D, Mittler R, Van Breusegem F (2006) Transcriptomic footprints disclose specificity of reactive oxygen species signaling in Arabidopsis. Plant Physiol 141:436–445
Gore M, Hayes A, Jeong S, Yue Y, Buss G, Maroof MS (2002) Mapping tightly linked genes controlling potyvirus infection at the Rsv1 and Rpv1 region in soybean. Genome 45:592–599
Guo D, Zhi H, Wang Y (2005) Identification and distribution of soybean mosaic virus strains in Middle and Northern Huang Huai Region of China. Chin J Oil Crop Sci 27:64–68
Hajimorad M, Hill J (2001) Rsv1-mediated resistance against Soybean mosaic virus-N is hypersensitive response-independent at inoculation site, but has the potential to initiate a hypersensitive response-like mechanism. Mol Plant Microbe Interact 14:587–598
Hayes AJ, Ma G, Buss GR, Maroof M (2000) Molecular marker mapping of Rsv4, a gene conferring resistance to all known strains of Soybean Mosaic Virus. Crop Sci 40:1434–1437
Hayes A, Jeong S, Gore M, Yu Y, Buss G, Tolin S, Maroof MS (2004) Recombination within a nucleotide-binding-site/leucine-rich-repeat gene cluster produces new variants conditioning resistance to soybean mosaic virus in soybeans. Genetics 166:493–503
Hernández JA, Gullner G, Clemente-Moreno MJ, Künstler A, Juhász C, Díaz-Vivancos P, Király L (2016) Oxidative stress and antioxidative responses in plant–virus interactions. Physiol Mol Plant Pathol 94:134–148
Hill J, Bailey T, Benner H, Tachibana H, Durand D (1987) Soybean mosaic virus: effects of primary disease incidence on yield and seed quality. Plant Dis 71:237–239
Hossain M, Akamatsu H, Morishita M, Mori T, Yamaoka Y, Suenaga K, Soares R, Bogado A, Ivancovich A, Yamanaka N (2015) Molecular mapping of Asian soybean rust resistance in soybean landraces PI 594767A, PI 587905 and PI 416764. Plant Pathol 64:147–156
Huynh T, Bastien M, Iquira E, Turcotte P, Belzile F (2010) Identification of QTLs associated with partial resistance to white mold in soybean using field-based inoculation. Crop Sci 50:969–979
Hyten DL, Smith JR, Frederick RD, Tucker ML, Song Q, Cregan PB (2009) Bulked segregant analysis using the GoldenGate assay to locate the locus that confers resistance to soybean rust in soybean. Crop Sci 49:265–271
Ilut DC, Lipka AE, Jeong N, Bae DN, Kim DH, Kim JH, Redekar N, Yang K, Park W, Kang S-T (2016) Identification of haplotypes at the Rsv4 genomic region in soybean associated with durable resistance to soybean mosaic virus. Theor Appl Genet 129:453–468
Jeong S, Kristipati S, Hayes A, Maughan P, Noffsinger S, Gunduz I, Buss G, Maroof M (2002) Genetic and sequence analysis of markers tightly linked to the resistance gene, Rsv3. Crop Sci 42:265–270
Jones DA, Thomas CM, Hammond-Kosack KE, Balint-Kurti PJ, Jones JD (1994) Isolation of the tomato Cf-9 gene for resistance to Cladosporium fulvum by transposon tagging. Science 266:789–793
Kang YJ, Kim KH, Shim S, Yoon MY, Sun S, Kim MY, Van K, Lee S-H (2012) Genome-wide mapping of NBS-LRR genes and their association with disease resistance in soybean. BMC Plant Biol 12:139
Kiihl RA, Hartwig E (1979) Inheritance of reaction to soybean mosaic virus in soybeans. Crop Sci 19:372–375
Li Q, Wan J (2005) SSRHunter: development of a local searching software for SSR sites. Hereditas 27:808–810
Li K, Yang Q, Zhi H, Gai J (2010) Identification and distribution of soybean mosaic virus strains in southern China. Plant Dis 94:351–357
Li C, Yang Y, Wang D, Li H, Zheng G, Wang T, Zhi H (2012) Studies on mapping and inheritance of resistance genes to SMV strain SC10 in soybean. Scientia Agricultura Sinica 45:4335–4342
Li H, Xiao Y, Cao L, Yan X, Li C, Shi H, Wang J, Ye Y (2013) Cerebroside C increases tolerance to chilling injury and alters lipid composition in wheat roots. PLoS One 8:e73380
Li K, Xia Y, Wang D, Yang Y, Ren R, Gao L, Zhang K, Zhi H (2014) Analysis of dynamic change of soybean mosaic virus strains in Heilongjiang province of China. Soybean Sci 33:880–884
Li N, Yin J, Li C, Wang D, Yang Y, Karthikeyan A, Luan H, Zhi H (2016) NB-LRR gene family required for Rsc4-mediated resistance to Soybean mosaic virus. Crop Pasture Sci 67:541–552
Liu J, Horstman HD, Braun E, Graham MA, Zhang C, Navarre D, Qiu W, Lee Y, Nettleton D, Hill JH (2011) Soybean homologs of MPK4 negatively regulate defense responses and positively regulate growth and development. Plant Physiol 157:1363–1378
Ma Y, Li H, Wang D, Liu N, Zhi H (2010) Molecular mapping and marker assisted selection of soybean mosaic virus resistance gene RSC12 in soybean. Legume Genom Genet 1:41–46
Ma Y, Wang D, Li H, Zheng G, Yang Y, Li H, Zhi H (2011) Fine mapping of the RSC14Q locus for resistance to soybean mosaic virus in soybean. Euphytica 181:127–135
Ma F, Wu X, Chen Y, Liu Y, Shao Z, Wu P, Wu M, Liu C, Wu W, Yang J (2016) Fine mapping of the Rsv1-h gene in the soybean cultivar Suweon 97 that confers resistance to two Chinese strains of the soybean mosaic virus. Theor Appl Genet 129:2227–2236
Maroof M, Tucker DM, Skoneczka JA, Bowman BC, Tripathy S, Tolin SA (2010) Fine mapping and candidate gene discovery of the soybean mosaic virus resistance gene, Rsv4. Plant Genome 3:14–22
Martin GB, Brommonschenkel SH, Chunwongse J, Frary A, Ganal MW, Spivey R, Wu T, Earle ED, Tanksley SD (1993) Map-based cloning of a protein kinase gene conferring disease resistance in tomato. Science 262:1432–1436
Molnar SJ, Rai S, Charette M, Cober ER (2003) Simple sequence repeat (SSR) markers linked to E1, E3, E4, and E7 maturity genes in soybean. Genome 46:1024–1036
Monteros MJ, Missaoui AM, Phillips DV, Walker DR, Boerma HR (2007) Mapping and confirmation of the ‘Hyuuga’ red-brown lesion resistance gene for Asian soybean rust. Crop Sci 47:829–834
Njiti V, Doubler T, Suttner RJ, Gray L, Gibson P, Lightfoot D (1998) Resistance to soybean sudden death syndrome and root colonization by Fusarium solani f. sp. glycine in near-isogenic lines. Crop Sci 38:472–477
Nordgren M, Fransen M (2014) Peroxisomal metabolism and oxidative stress. Biochimie 98:56–62
Sandhu D, Tasma IM, Frasch R, Bhattacharyya MK (2009) Systemic acquired resistance in soybean is regulated by two proteins, orthologous to Arabidopsis NPR1. BMC Plant Biol 9:105
Seo J-K, Kwon S-J, Cho WK, Choi H-S, Kim K-H (2014) Type 2C protein phosphatase is a key regulator of antiviral extreme resistance limiting virus spread. Sci Rep 4:5905
Shi A, Chen P, Zheng C, Hou A, Zhang B (2008) A PCR-based marker for the locus conferring resistance to Soybean Mosaic Virus. Crop Sci 48:262–268
Song W-Y, Wang G-L, Chen L-L, Kim H-S (1995) A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 270:1804–1806
Song Q, Marek L, Shoemaker R, Lark KG, Concibido V, Delannay X, Specht JE, Cregan P (2004) A new integrated genetic linkage map of the soybean. Theor Appl Genet 109:122–128
Song Q, Jia G, Zhu Y, Grant D, Nelson RT, Hwang E-Y, Hyten DL, Cregan PB (2010) Abundance of SSR motifs and development of candidate polymorphic SSR markers (BARCSOYSSR_1. 0) in soybean. Crop Sci 50:1950–1960
Suh SJ, Bowman BC, Jeong N, Yang K, Kastl C, Tolin SA, Maroof M, Jeong S-C (2011) The Rsv3 locus conferring resistance to soybean mosaic virus is associated with a cluster of coiled-coil nucleotide-binding leucine-rich repeat genes. Plant Genome 4:55–64
Sun X, Cao Y, Yang Z, Xu C, Li X, Wang S, Zhang Q (2004) Xa26, a gene conferring resistance to Xanthomonas oryzae pv. oryzae in rice, encodes an LRR receptor kinase-like protein. Plant J 37:517–527
Van Ooijen J (2006) JoinMap 4: software for the calculation of genetic linkage maps in experimental populations. Wageningen, Kyazma BV
van Zutphen T, Veenhuis M, van der Klei IJ (2008) Pex14 is the sole component of the peroxisomal translocon that is required for pexophagy. Autophagy 4:63–66
Viel C, Ide C, Cui X, Wang A, Farsi M, Michelutti R, Strümvik M (2009) Isolation, partial sequencing, and phylogenetic analyses of Soybean mosaic virus (SMV) in Ontario and Quebec. Can J Plant Pathol 31:108–113
Voorrips R (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang X, Gai J, Pu Z (2003) Classification and distribution of strain groups of Soybean mosaic virus in middle and lower Huang-Huai and Changjiang Valleys. Soybean Sci 22:102–107
Wang Y, Zhi H, Guo D, Gai J, Chen Q, Li K, Li H (2004) Classification and distribution of strain groups of soybean mosaic virus in Northern China spring planting soybean region. Soybean Sci 24:263–268
Wang D, Ma Y, Liu N, Yang Z, Zheng G, Zhi H (2011a) Fine mapping and identification of the soybean RSC4 resistance candidate gene to soybean mosaic virus. Plant Breed 130:653–659
Wang D, Ma Y, Yang Y, Liu N, Li C, Song Y, Zhi H (2011b) Fine mapping and analyses of RSC8 resistance candidate genes to soybean mosaic virus in soybean. Theor Appl Genet 122:555–565
Wang D, Li H, Zhi H, Tian Z, Hu C, Hu G, Huang Z, Zhang L (2014) Identification of strains and screening of resistance resources to soybean mosaic virus in Anhui Province. Chin J Oil Crop Sci 36:374–379
Whitham S, Dinesh-Kumar S, Choi D, Hehl R, Corr C, Baker B (1994) The product of the tobacco mosaic virus resistance gene N: similarity to toll and the interleukin-1 receptor. Cell 78:1101–1115
Wrather J, Anderson T, Arsyad D, Tan Y, Ploper LD, Porta-Puglia A, Ram H, Yorinori J (2001a) Soybean disease loss estimates for the top ten soybean-producing countries in 1998. Can J Plant Pathol 23:115–121
Wrather J, Stienstra W, Koenning S (2001b) Soybean disease loss estimates for the United States from 1996 to 1998. Can J Plant Pathol 23:122–131
Xu Y, Li H-N, Li G-J, Wang X, Cheng L-G, Zhang Y-M (2011) Mapping quantitative trait loci for seed size traits in soybean (Glycine max L. Merr.). Theor Appl Genet 122:581–594
Yan H, Wang H, Cheng H, Hu Z, Chu S, Zhang G, Yu D (2015) Detection and fine-mapping of SC7 resistance genes via linkage and association analysis in soybean. J Integr Plant Biol 57:722–729
Yang Y (2002) Classification and distribution of strains of Soybean mosaic virus in the middle and lower Changjiang River Valleys and the resistance to Soybean mosaic virus in soybeans. Master’s thesis, Nanjing Agricultural University, Nanjing
Yang Q, Gai J (2011) Identification, inheritance and gene mapping of resistance to a virulent Soybean mosaic virus strain SC15 in soybean. Plant Breed 130:128–132
Yang H, Huang Y, Zhi H, Yu D (2011) Proteomics-based analysis of novel genes involved in response toward soybean mosaic virus infection. Mol Biol Rep 38:511–521
Yang Y, Zheng G, Han L, Dagang W, Yang X, Yuan Y, Huang S, Zhi H (2013) Genetic analysis and mapping of genes for resistance to multiple strains of Soybean mosaic virus in a single resistant soybean accession PI 96983. Theor Appl Genet 126:1783–1791
Yang Y, Lin J, Zheng G, Zhang M, Zhi H (2014) Recombinant soybean mosaic virus is prevalent in Chinese soybean fields. Adv Virol 159:1793–1796
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform 13:134
Yong Z, Hao-Ru T, Ya L (2008) Variation in antioxidant enzyme activities of two strawberry cultivars with short-term low temperature stress. World J Agric Sci 4:458–462
Yu Y, Saghai Maroof M, Buss G, Maughan P, Tolin S (1994) RFLP and microsatellite mapping of a gene for soybean mosaic virus resistance. Phytopathology 84:60–64
Zhan Y, Zhi H, Yu D, Gai J (2006) Identification and distribution of SMV strains in Huang-Huai valleys. Sci Agric Sin 39:2009–2015
Zhang C, Grosic S, Whitham SA, Hill JH (2012) The requirement of multiple defense genes in soybean Rsv1–mediated extreme resistance to Soybean mosaic virus. Mol Plant Microbe Interact 25:1307–1313
Zhang D, Cheng H, Hu Z, Wang H, Kan G, Liu C, Yu D (2013) Fine mapping of a major flowering time QTL on soybean chromosome 6 combining linkage and association analysis. Euphytica 191:23–33
Zhao L, Wang D, Zhang H, Shen Y, Yang Y, Li K, Wang L, Yang Y, Zhi H (2016) Fine mapping of the RSC8 locus and expression analysis of candidate SMV resistance genes in soybean. Plant Breed 135:701–706
Zheng G, Yang Y, Ma Y, Yang X, Chen S, Ren R, Wang D, Yang Z, Zhi H (2014) Fine mapping and candidate gene analysis of resistance gene RSC3Q to soybean mosaic virus in Qihuang 1. J Integr Agric 13:2608–2615
Acknowledgements
This work was financially supported through grants from the National Natural Science Foundation of China (Grant Nos. 31671718, 31371646, 31571690), the Fundamental Research Funds for the Central Universities (Y0201600115), the National Soybean Industrial Technology System of China (No. CARS-04), Jiangsu Collaborative Innovation Center for Modern Crop Production (JCIC-MCP), the Fund of Transgenic Breeding for Soybean Resistance to Soybean mosaic virus (No. 2008ZX08004-004) and Changjiang Scholars and Innovative Research Team in University (PCSIRT13073).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The experiments were performed in compliance with the current laws of China.
Additional information
Communicated by Dr. David A Lightfoot.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rui, R., Liu, S., Karthikeyan, A. et al. Fine-mapping and identification of a novel locus Rsc15 underlying soybean resistance to Soybean mosaic virus . Theor Appl Genet 130, 2395–2410 (2017). https://doi.org/10.1007/s00122-017-2966-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-2966-5