Abstract
Key message
TFs involved in drought tolerance in plants may be utilized in future for developing drought tolerant cultivars of wheat and some other crops.
Abstract
Plants have developed a fairly complex stress response system to deal with drought and other abiotic stresses. These response systems often make use of transcription factors (TFs); a gene encoding a specific TF together with -its target genes constitute a regulon, and take part in signal transduction to activate/silence genes involved in response to drought. Since, five specific families of TFs (out of >80 known families of TFs) have gained widespread attention on account of their significant role in drought tolerance in plants, TFs and regulons belonging to these five multi-gene families (AP2/EREBP, bZIP, MYB/MYC, NAC and WRKY) have been described and their role in improving drought tolerance discussed in this brief review. These TFs often undergo reversible phosphorylation to perform their function, and are also involved in complex networks. Therefore, some details about reversible phosphorylation of TFs by different protein kinases/phosphatases and the co-regulatory networks, which involve either only TFs or TFs with miRNAs, have also been discussed. Literature on transgenics involving genes encoding TFs and that on QTLs and markers associated with TF genes involved in drought tolerance has also been reviewed. Throughout the review, there is a major emphasis on wheat as an important crop, although examples from the model cereal rice (sometimes maize also), and the model plant Arabidopsis have also been used. This knowledge base may eventually allow the use of TF genes for development of drought tolerant cultivars, particularly in wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drought stress has an adverse impact on growth and productivity of plants including all major crops worldwide. Genetic variation for adaptation for drought tolerance is available in all plant species, and involves genes that express at multiple levels including physiology, morphology and cellular/molecular levels. In particular, the drought stress initiates transcription factor (TF)-mediated expression of a variety of genes in many plant species (Shinozaki et al. 2003; Bartels and Sunkar 2005; Lata et al. 2015; Singh and Laxmi 2015). It has also been shown that in plant genomes ~6–10 % genes encode TFs (Franco-Zorrilla et al. 2014). DNA-binding domains of these TFs are highly conserved among species, so that the characteristics of these domains have been used to classify TFs into families. The number of families of TFs differ in different plant species, so that in different plant systems the number of TF families ranges from 26 to 83 (Jin et al. 2014). For instance, as many as 34 families containing 1,533 TFs are known in Arabidopsis thaliana; about ~45 % of these TFs belong to families that are specific to plants (Riechmann et al. 2000). Some of the major families of TFs include the following: AP2/EREBP or AP2/ERF, ABI3VP1, ARF, bZIP/HD-ZIP, C2H2, GRAS, MYB/MYC, Zinc fingers, MADS, NAC and WRKY; of these, TFs belonging to five multi-gene families [bZIP (mainly AREB/ABF), DREB (AP2/EREBP), MYB/MYC, NAC and WRKY)] have been shown to be associated with drought tolerance (although there are exceptions; see later). Three of these five families (AP2/EREBP, NAC and WRKY) are also known to be unique to plants, although AP2/EREBP domain was also reported to be present in TFs of protists, cyanobacteria and phages (Wessler 2005). It is known that most TFs belonging to the above five families are key regulators of several developmental processes and are also involved in response to biotic and abiotic stresses including drought (Riechmann and Meyerowitz 1998; Uno et al. 2000; Jakoby et al. 2002; Abe et al. 2003; Fujita et al. 2004; Tran et al. 2004; 2007; Yanhui et al. 2006; Lata and Prasad, 2011; Zhu et al. 2013; Huang et al. 2015). The above five families of TFs have been systematically studied in a number of plant systems including the following: Arabidopsis thaliana, Arachis hypogea, Brassica napus, Cicer aritinum, Gossypium hirsutum, Gylcine max, Hordeum vulgare, Oryza sativa, Pennisetum glaucum, Poncirus trifoliata, Solanum tuberosum, Sorghum bicolor, Triticum aestivum, Vitis vinifera and Zea mays (Table 1; Table S1).
Since drought tolerance may require regulation of the expression of a number of genes including TF genes, and because some TFs may each regulate transcription of a number of genes involved in drought tolerance, a study of TFs in all major crops including wheat has been rewarding. A successful example of TFs used in breeding for abiotic stress is the conversion of flood sensitive rice varieties into flood-tolerant genotypes after introgression of Sub1 locus that encodes a TF (ethylene response factor), which has been shown to induce transcription of ~900 genes (Xu et al. 2006; Septiningsih et al. 2009). There are also reports in several other crops including wheat, where the role of TFs in imparting tolerance against abiotic stresses including drought has been systematically examined (Table 1; Table S1), so that TF genes for drought tolerance may be used in future for developing drought tolerant cultivars in some important crops including wheat (Fig. 1).
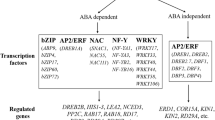
A schematic representation of drought stress signal perception and gene expression via ABA-dependent and ABA-independent pathways in plants; their utilization in the development of drought tolerant wheat cultivars using marker-assisted selection is also shown (Modified from Shinozaki and Yamaguchi-Shinozaki 2000)
A number of wheat genes for TFs (TabZIP1, TabZIP60, TaABRE3, TaDREB1, TaPIMP1, TaNAC29 and TAWRKY44) have been shown to exhibit induced expression during exposure to drought stress, suggesting that these genes may be used for improving stress tolerance in wheat (Egawa et al. 2006; Cao et al. 2012; Zhang L et al. 2012; Zhang Z et al. 2012; Zhang L et al. 2015; Huang et al. 2015; Wang et al. 2015; Wang J et al. 2016). The wheat gene TaGT2L1 encoding a TF belonging to GT family has also been recently shown to play a role in regulation of stomatal density and drought tolerance (Zheng et al. 2016). Genes for TFs involved in stress tolerance have also been studied in two model plant systems, Arabidopsis and rice, the latter often having orthologues in wheat (Hu et al. 2006; Lu et al. 2009; Fang et al. 2015).
In this brief review, we first describe regulons involving TFs of the above five major families and their interaction with cis-regulatory elements (CREs) that are available in the promoters of drought responsive genes; at the whole genome level, these cis-elements in a species constitute, what are described as cistrome and epicistrome for that species (O’Malley et al. 2016). We are aware that some details about TF regulons are also available in earlier recent reviews (Nakashima et al. 2009, 2014; Singh and Laxmi 2015; Joshi et al. 2016), but this review has a focus on TFs involved in drought tolerance and that too in wheat. Other novel features of this review include a brief account of reversible phosphorylation involved in the function of TFs and the co-regulatory networks, which involve either only TFs, or TFs with miRNAs. A brief account of databases of plant TFs and cistrome/epicistrome is also included.
Regulons involved in drought tolerance: TFs and their target genes
A TF generally has two domains, the DNA binding (DB) domain and an activation domain (AD). With the help of DB domain, a TF binds to a cis-acting element (also described as TF binding site or TFBS) available in the promoter of a stress responsive gene; this brings AD in close proximity of the concerned gene thus facilitating activation or repression of the target gene. A number of genes that carry the same cis-acting element (s) and thus induced by the same TF (s) constitute a regulon. A number of such drought responsive regulons along with associated TFs are now known (Table 2). Five classes of such extensively studied regulons that are involved in drought response, along with their TFs will be described in brief (more details are available in some recent reviews, Nakashima et al. 2009, 2014; Singh and Laxmi 2015).
bZIP TFs: the AREB/ABF regulons
Basic leucine zipper (bZIP) TFs represent one of the most diverse and largest TF family (Pérez-Rodríguez et al. 2010). TFs belonging to this family have been classified in ten groups/subfamilies on the basis of sequence similarity of basic region and the presence of additional conserved motifs (Jakoby et al. 2002). An important subfamily of these bZIP TFs, which regulate expression of stress responsive gene via abscisic acid (ABA) belong to group A. An analysis of ABA-inducible genes revealed that each of these genes carry a conserved cis-acting element (ACGTGGC) in their promoters, designated as ABA-responsive element (ABRE), which is a subset of the G-box sequence (CACGTG); therefore, sometimes these response elements are also described as G-ABRE, the corresponding; bZIP TFs are described as ABRE binding protein/factor (AREB/ABF) (Table S1). AREB/ABF TFs harbor a highly conserved bZIP domain composed of two structural features [a basic region and a leucine (Leu) zipper]; the basic region comprises ~16 amino acid residues with the invariant motif N-x7-R/K-x9 and is responsible for nuclear localization and DNA binding, whereas the Leu zipper is composed of heptad repeats of Leu or other bulky hydrophobic amino acids and mediates homo- and/or heterodimerization of bZIP proteins (Jakoby et al. 2002; Fig. 2a). In rice, bZIP TFs have also been shown to be involved in feedback regulation of ABA level during drought stress by targeting the ABA biosynthesis gene OsNCED4 (Zong et al. 2016).
Ribbon structures of TF-DNA complexes involving five families of wheat TFs. a AP2/ERF b bZIP c Myb d NAC e WRKY. In each case, information on the following features is written in the corresponding legend: (1) structural features, (2) recognition sequences, (3) number of amino-acids (aa) (4) Plant TFDB v3 (http://planttfdb.cbi.pku.edu.cn/index.php) entry codes of wheat TF (except bZIP; structure taken from RCBS protein data bank; http://www.rcsb.org/pdb/home/home.do). The figures for molecules were drawn using TFmodeller (http://maya.ccg.unam.mx/~tfmodell/) and PDB file view on SWISS PDB viewer v4 (Guex et al. 2009)
TFs of bZIP family that are involved in AREB/ABF regulons provide a good example of interaction between TFs and the stress responsive genes (which carry cis-acting element, ABRE), particularly in the context of drought tolerance (Choi et al. 2000; Fujita et al. 2005; Nakashima et al. 2009; Wang J et al. 2016). In wheat, the first example of such an AREB/ABF and the corresponding ABRE became available, when ABA-induced transient expression assay in rice protoplasts was used to identify an ABA-responsive promoter in some wheat genes (Marcotte et al. 1988); an ABRE sequence was also identified soon in the wheat gene Em (early maturity; Marcotte et al. 1989). Subsequently, ABREs were detected in a number of ABA-responsive genes (i.e., RD29, COR15, COR47 etc.) in several plant species (for a review, see Kim 2006).
Recently, an AREB wheat gene (TaAREB3) encoding a bZIP TF was also identified and characterized (Wang J et al. 2016). When this gene was overexpressed in Arabidopsis, the transgenic lines were found to be relatively more sensitive to ABA and also more tolerant to drought and freezing. Functional analysis showed that the TF encoded by TaAREB3 can bind promoters of four drought responsive genes (RD29A, RD29B, COR15A and COR47) and activate their expression under stress conditions like drought and freezing (Wang J et al. 2016). These four genes are key ABA signaling regulators and are involved in freezing tolerance. Another wheat bZIP TF gene TabZIP60, when overexpressed in Arabidopsis transgenic plants, imparted improved drought tolerance. These transgenic plants also exhibited higher expression of four key ABA signaling regulators and cold-responsive genes (AtRD29A, AtRD20, AtRD29B and AtCOR47) under drought. These results suggested that TabZIP60 is involved in regulating the expression of some key ABA signaling regulators and cold-responsive genes under drought condition (Zhang et al. 2015).
The promoters of drought responsive genes may also each carry one or more distal or proximal coupling elements (CE), such as CE1 and CE3 to induce expression of drought responsive genes. For instance, CEs in barley have been shown to form an abscisic acid response complex (ABRC), which might be necessary and sufficient to confer ABA response or to trigger ABA-mediated gene expression (Shen et al. 1996). Binding sites have also been identified in CEs, where ABFs bind (Choi et al. 2000; Niu et al. 2002). TFs belonging to AP2/ERF proteins could also interact with CE1 (Lee et al. 2010a).
DREB (AP2/ERF) TFs and DREB/CBF regulons
The dehydration-responsive element-binding (DREB) proteins (TFs) play an important role in response to drought stress. They represent a large subfamily of TFs belonging to the family AP2/EREBP or AP2/ERF (APETALA2/ethylene-responsive element-binding protein/factor). Several members of this subfamily have been identified in different plant species (details are given in Table S1). Each DREB TF contains one AP2/ERF DNA-binding domain (highly conserved region of about 60 to 70 amino acids) that binds to the cis-acting element DRE consisting of 9-bp core sequence (TACCGACAT) that is available in promoters of drought responsive genes (Fig. 2b). The presence of this DRE has been reported in several drought responsive genes (e.g., RD29A and RD29B in Arabidopsis; Yamaguchi-Shinozaki and Shinozaki 1994). Similar cis-acting elements named C-repeat (CRT), each containing A/GCCGAC motif (found in DRE) have also been reported in the promoters of cold inducible genes in Arabidopsis, where CRT is used for binding of the TF CBF (CRT binding factor) under cold stress (Saleh et al. 2005).
A DREB TF and the corresponding one or more cis-acting elements (e.g., DRE) constitute a regulon, thus providing another important example of well characterized interaction between a TF and a cis-element. In wheat, DREB1/CBF and DREB2 genes were identified and named TaDREB1A and WDREB2 (Vagujfalvi et al. 2005; Egawa et al. 2006; Nakashima et al. 2009; Hassan et al. 2015). Overexpression of these two wheat genes in transgenic Arabidopsis or tobacco plants resulted in overexpression of downstream stress-inducible genes (RD29A, Wdhn13, Wrab17, Wrab18 and Wrab19) having DRE in their promoter region; these transgenic lines also exhibited tolerance to abiotic stresses such as drought and freezing (Pellegrineschi et al. 2004; Egawa et al. 2006). Similar, results were also obtained using Arabidopsis genes for DREB TFs. For instance, transgenic Arabidopsis plants exhibiting overexpression of its own gene AtDREB1/CBF were found to exhibit tolerance to drought, high salinity and cold stress (Jaglo-Ottosen et al. 1998; Liu et al. 1998; Kasuga et al. 1999) suggesting that DREBs/CBFs target multiple genes. The target genes for DREB/CBF include genes encoding LEA (late embryogenesis abundant) proteins, KIN (cold-inducible) proteins, osmoprotectant biosynthesis proteins, and protease inhibitors, which are known to function in response to abiotic stresses. More than 40 such target genes containing either DRE/CRT or other core motifs in their promoters have so far been identified (Seki et al. 2001; Fowler and Thomashow 2002; Maruyama et al. 2004).
MYB/MYC TFs and MYB/MYC regulons
The TFs belonging to the MYB (myeloblastosis)/MYC (myelocytomatosis) family of proteins are found in both plants and animals and have varied functions (Abe et al. 2003). Each TF of MYB family contains a MYB domain with 1–3 imperfect repeats, each with about 52 amino acid residues, which acquire a helix-turn-helix (HTH) conformation and intercalates in the major groove of DNA (Yanhui et al. 2006; Fig. 2c). Similarly, each MYC TF harbors a highly conserved basic helix-loop-helix (bHLH) domain (Kazan and Manners 2013). MYB and MYC TFs are often together involved in constituting, common regulons described as MYB/MYC regulons.
MYB/MYC regulons participate in some important transcriptional pathways that are involved in drought stress responses via ABA-dependent signaling systems (Abe et al. 1997; Baldoni et al. 2015). In wheat, MYB TF gene TaMYB30-B was found to be involved in drought stress responses. Arabidopsis transgenic plants, overexpressing TaMYB30-B gene also exhibited altered expression levels of some drought stress-responsive genes (RD29A, ERD1); they also carried improved drought stress tolerance during germination and seedling stages (Zhang L et al. 2012). Another wheat MYB TF gene TaPIMP1 is also involved in regulation of genes involved in drought response; its own expression level is also positively correlated with drought tolerance (as shown in transgenic plants). Electrophoretic mobility shift assay and yeast-one-hybrid assays suggested that the TF TaPIMP1 can bind to MYB-binding site, and activate the expression of the genes with the Myb cis-element. The TF TaPIMP1 was also shown to upregulate the expression of a subset of defense- and stress-related genes (e.g., RD22, TLP4 and PR1a) as revealed through microarray analysis. (Zhang Z et al. 2012). Another wheat MYB TF gene TaMYB3R1 was also found to be involved in drought stress response. In transgenic Arabidopsis lines overexpressing TaMYB3R1, enhanced mRNA levels were observed for several dehydration inducible genes including both, ABA-dependent genes (RD29A, RD29B, and ABF3) and ABA-independent genes (COR15A, ADH1 and CBF4). These results suggested that TaMYB3R1 also affects expression of dehydration-responsive genes in both ABA-dependent and ABA-independent pathways (Cai et al. 2015).
In the model plant Arabidopsis also, induced expression of RD22 gene has been shown to be mediated by abscisic acid (ABA), although it lacks ABRE cis-element in its promoter region; its expression is regulated by two closely located putative recognition sites, namely MYCR, CNNTG for the MYC TF (AtMYC2) and MYBR, TAACNA/G for MYB TF (AtMYB2) (Abe et al. 1997, 2003). This represents an example of a regulon, where same gene may carry two binding sites for two TFs. Overexpression of two Arabidopsis TF genes (AtMYC2 and AtMYB2), which overexpressed in transgenic Arabidopsis plants (studied through microarray analysis) suggested that their target genes include many ABA- or jasmonic-acid (JA)-inducible genes (e.g., RD22, AtADH1). These transgenic plants were also found to be ABA responsive and carried improved osmotic tolerance (Abe et al. 2003).
NAC TFs and NAC regulons
The TFs of NAC family (NAM, ATAF and CUC) have also been shown to be involved in activation of drought responsive genes. These TFs are unique to plants and have not been reported in animal systems. NAC TFs share a conserved N-terminal with ~150–160 amino-acids long DNA binding region (NAC domain) carrying five sub-domains (A–E) and a variable C-terminal (Ooka et al. 2003) that is responsible for the observed differences in the regulatory function of NAC proteins (Jensen et al. 2010). Some NAC proteins, described as NTL or transmembrane motif (TMs) 1-like, also contain α-helical transmembrane motifs at their C-terminals (Kim et al. 2010) (Fig. 2d), which allow activation or repression of a variety of downstream genes thus regulating multiple cellular or molecular processes (Nakashima et al. 2012; Puranik et al. 2012).
NAC TF genes and their corresponding cis-acting elements (NACRS) constitute NAC regulons, and provide another important example of well characterized interaction between a TF and one or more cis-elements that work during drought stress. Recently it was shown that Arabidopsis transgenic plants overexpressing three wheat NAC TF genes (TaNAC47, TaNAC67 and TaNAC29) exhibited an improved tolerance to drought stress. In transgenic plants carrying TaNAC47, the expression levels of six stress-responsive genes (RD29A, RD29B, COR47, RD20, GSTF6 and P5CS1) increased. Similarly, in transgenic plants carrying TaNAC67, the expression of five stress-responsive genes (DERB1A, RD29B, RD29A, RAB18 and ABI5) increased. The promoters of these genes carried NAC-binding cis-elements, suggesting that these genes might be transcriptionally activated by TaNAC47/TaNAC67 and constitute a NAC regulon (Mao et al. 2014; Zhang et al. 2016). In contrast, in the transgenic plants carrying TaNAC29, the expression level of some key ABA signaling regulators and senescence-associated genes (RD29B, SAG13, SAG113, AIB1, ERD11 and ABI5) was significantly reduced (Huang et al. 2015), thus suggesting that wheat NAC TFs may be either involved in activation or repression of downstream genes.
NAC TFs have also been studied in two model plant systems including Arabidopsis and rice. For instance JUB1 (JUNGBRUNNEN1) regulates the Gibberellins (GAs)/brassinosteroids (BRs)-DELLA signaling in Arabidopsis; in doing so, JUB1 causes accumulation of DELLA proteins in two different ways: (1) suppression of the activity of genes GA3ox1 and DWF4, thus negatively regulating the synthesis of GA and BR biosynthesis; (2) direct activation of genes, GAI (GA INSENSITIVE) and RGL1 (RGA-LIKE). DELLAs, in turn, cause inhibition of cell elongation, reduction of intracellular H2O2 levels, and promotion of stress tolerance (Shahnejat-Bushehri et al. 2016). Another Arabidopsis gene ERD1 (early dehydration stress 1) is also induced by several NAC TFs including ANAC019, ANAC055, ANAC072 (Tran et al. 2004, 2007). Recently in rice, OsNAM (a NAC TF) was shown to regulate the expression of five genes (OsCESA, OsGDP, OsMtN3, OsAH and OsGdpD) under drought stress (Dixit et al. 2015). Examples of such NAC TFs, which use NACRS motif for binding include ANAC019, ANAC055, and ANAC072 in Arabidopsis (Tran et al. 2004) and SNAC1, ENAC1 and SNAC2 in rice (Hu et al. 2008; Sun et al. 2012).
WRKY TFs and WRKY regulons
WRKY is the largest superfamily of TFs that are unique to plants. These TFs not only impart tolerance against abiotic stresses, but also regulate plant growth and development (Wu et al. 2008; Tripathi et al. 2012, 2014; Zhu et al. 2013). Several members of this family have been identified in various plant species (details were given in Table S1; Rushton et al. 2010; Banerjee and Roychoudhury 2015; Xu et al. 2016). TFs of this family are characterized by the presence of WRKY domain (~60 aa), which is composed of highly conserved WRKYGQK sequence followed by a zinc-finger motif. This WRKY domain exhibits high binding affinity for the cis-acting element called W-box (TTGACC/T), which occurs in several drought responsive genes (Ulker and Somssich 2004; Rushton et al. 2010; Fig. 2e).
The genes encoding TFs of WRKY family and the cis-acting element W-box constitute another class of regulons, which are involved in drought stress response and signaling. In wheat also, WRKY regulons have been reported to be involved in drought stress. Transgenic Arabidopsis plants overexpressing wheat genes TaWRKY2 and TaWRKY19 exhibited increased tolerance to drought stress. TaWRKY2 and TaWRKY19 are two nuclear proteins (TFs), which displayed specific binding to W box; TaWRKY2 binding the promoter of downstream gene RD29B, and TaWRKY19 binding the promoters of three genes (RD29A, RD29B and COR6.6). In both cases, TF binding leads to increase in expressions of downstream genes during drought stress (Niu et al. 2012).
In another study, Wang et al. (2013) identified ten (10) TaWRKY genes, designated TaWRKY1–TaWRKY10. Among these genes, TaWRKY10 was examined in more detail, and was shown to confer drought tolerance by regulating osmosis and reducing ROS accumulation. Overexpression of TaWRKY10 in tobacco transgenic plants significantly activated the expression of following three stress-related genes, which can be classified into the following two groups: (1) NtERD10C (encoding early response to drought 10C) and NtSPSA (encoding sucrose–phosphate synthase activity) both dealing with osmotic stress, and (2) NtGPX (encoding glutathioneperoxidase) involved in scavenging ROS.
Another wheat WRKY TF gene (TaWRKY44) that was overexpressed in tobacco transgenic plants, imparted an improved tolerance to drought stress with increased relative water content (RWC), proline and soluble sugar accumulation, improved antioxidant system (Wang et al. 2015). The 14 ROS-related and stress-responsive genes that, were upregulated in these transgenic plants, could be classified in the following four groups: (1) genes encoding enzymes for ROS detoxification (NtSOD, NtAPX, NtCAT, NtPOX, and NtGST); (2) genes encoding enzymes involved in the biosynthesis of polyamine (NtADC1 and NtSAMDC), sucrose (NtSPSA) or ABA (NtNCED1); (3) stress-defensive protein gene (NtERD10C); and (4) lipid-transfer protein genes (NtLTP1 and TobLTP1). These results suggested that TaWRKYs are involved in regulating the expression of some key ROS-related and stress responsive genes under drought.
WRKY regulons were also reported in the model plant Arabidopsis. Transgenic Arabidopsis lines exhibiting overexperssion of its own genes WRKY18, WRKY40 and WRKY60 had altered plant sensitivity to ABA, salt and osmotic stress. During drought stress, TFs WRKY18 and WRKY60 were found to be positive regulators of ABA signaling, and the TF WRKY40 was found to be a negative regulator. They bind to the promoters of multiple genes including some stress-inducible downstream genes (RD29A and COR47) and some TF genes (e.g., ABI5, DREB1A/CBF3, DREB2A and MYB2), and influence their expression (Chen et al. 2010; Shang et al. 2010). Further, a transgenic Arabidopsis line exhibiting overexpression of ABO3 gene that encodes a WRKY TF (AtWRKY63) was also found to exhibit hypersensitive response for ABA in the seedling stage, associated with reduced drought tolerance. It was also found that a mutant ABO3 downregulates the expression of two downstream genes (RD29A and COR47) on ABA treatment. A study involving gel-shift assay revealed that ABO3 protein binds to the W-box localized in the ABF2 promoter (Ren et al. 2010).
Kinases and phosphorylation of TFs for drought tolerance
The products of TF genes in the form of transcripts and proteins (TFs) have been shown to undergo post-transcriptional and post-translational modifications respectively. The major modifications at the post-transcriptional level include alternate splicing, while modifications at the post-translational level include phosphorylation and protein–protein interactions. Of these modifications, however, phosphorylation seems to be the most common and widespread modification that has been witnessed in almost all TFs. Phosphorylation in TFs has also been shown to be induced as a response to abiotic stresses (Luan 2003; You et al. 2014). During phosphorylation, a protein kinase transfers the terminal phosphate group of an ATP molecule to the hydroxyl group on a Ser, Thr, or Tyr side chain of the protein. Reversible phosphorylation allows TFs to switch rapidly from dephosphorylated state to phosphorylated state and vice versa, thus permitting plants to respond to stress stimuli rapidly and accurately.
Variations have been observed in the kinases that cause phosphorylation and also in the specific amino acid residues of TFs that are involved in phosphorylation (i.e., Ser, Thr and Tyr). Serine-threonine kinases (e.g., SnRKs) are generally used in phosphorylation of bZIP, NAC and DREB TFs, mitogen-activated protein kinases (MAPKs) are used for phosphorylation of MYB/MYC and WRKY TFs, and calcium-dependent protein kinases (CDPKs) are involved in phosphorylation of some bZIP TFs (e.g., ABF1 and ABF4). In the following section, some details about kinases and phosphatases involved in reversible phosphorylation of TFs, involved in response to drought stress are presented. Modifications of transcripts and TFs other than phosphorylation will not be described, since either they don’t appear to be so widespread, or else these have not been worked out as extensively as the kinases and phosphatases and their involvement in reversible phosphorylation.
The members of SnRK2 family are plant-specific serine/threonine kinases involved in phosphorylation that is involved in plant response to abiotic stresses (Kulik et al. 2011). It has been shown that the AREB/ABF TFs activate expression of genes only after phosphorylation of their conserved regions (Ser/Thr residues of R-XX-S/T sites) (Furihata et al. 2006; Fujita et al. 2011). This phosphorylation is catalyzed by phosphorylated form of a SnRK2 kinase itself that is negatively regulated by a phosphatase PP2C (protein phosphatase 2C), which in turn is negatively regulated by a complex between ABA and its receptors including PYR/PYL/RCARs (pyrabactin resistance/pyrabactin resistance 1-like/regulatory component of ABA receptors). This complex inhibits phosphatase activity of PP2C, so that SnRK2 remains phosphorylated and active, and brings about phosphorylation of AREB/ABF. It is only in its phosphorylated form that AREB/ABF induces expression of downstream genes (Umezawa et al. 2010). Later, the phosphatase activity of PP2C is restored due to dissociation of PYR/PYL/RCAR, so that now it dephosphorylates and thereby inactivates SnRK2. This inactivated and released dephosphorylated SnRK2 gets accumulated in the cell, where it gets phosphorylated again to initiate another cycle of AREB/ABF phosphorylation (Fig. 3a). The SnRK2 kinases are also involved in phosphorylation of NAC TFs, which get phosphorylated at Thr-142 due to the kinase SnRK2.8 under drought conditions in Arabidopsis (Kim et al. 2012).
Models showing phosphorylation: a ABF TFs using SnRK2 pathway b WRKY TFs using MAPK pathway and c ABF TFs using CDPK pathway in plant response to drought stress. (Modified from Zhu S-Y et al. 2007; Kulik et al. 2011; Banerjee and Roychoudhury 2015). ABA abscisic acid, ABF ABRE binding protein/factors, CPK calcium-dependent protein kinase, PYR/PYL/RCARs pyrabactin resistance/pyrabactin resistance 1-like/regulatory component of ABA receptors, MAPK mitogen-activated protein kinase, MAPKK MAP kinase kinase, MAPKKK MAP kinase kinase kinase
In wheat, both kinases (e.g., SnRK2) and phosphatases (TaPP2Ac-1) and the corresponding genes for reversible phosphorylation have been reported and their involvement in response to abiotic stress tolerance has been demonstrated (Mao et al. 2010). In some cases, it has also been shown that the TFs have sites for phosphorylation, suggesting that the TaSnRK2 genes may be involved in phosphorylation of TFs. For instance, an SnRK2 ortholog named PKABA1 was initially reported, which phosphorylates wheat AREB1 ortholog, TaABF (Johnson et al. 2002, 2008; Fig. S1). Another TF, TaAREB3 was shown to contain a conserved N-terminal of 145 amino acids (aa) with Ser/Thr-like protein kinase phosphorylation sites, a variable M region (146-257 aa) and a conserved C-terminal region with a bZIP domain (258–311 aa). Alignment and phylogenetic analysis also revealed that TaAREB3 is similar to AtAREB3 (a member of AREB subgroup) that belongs to A group of bZIP family (Wang J et al. 2016).
More recently, 10 different genes encoding wheat SnRK2s have been isolated and characterized (Zhang et al. 2016). Based on their kinase domains and the C-terminus, the 10 SnRK2s were classified into three subclasses (I, II and III) (Fig. S2). Expression pattern analysis revealed that all TaSnRK2s were involved in responses to PEG, NaCl, and cold stress. TaSnRK2s in subclass III were strongly induced by ABA, those in subclass II responded weakly to ABA, whereas those in subclass I were not activated by ABA treatment. Physical and functional interaction between TaSnRK2s and a typical group A PP2C (TaABI1) was also examined, suggesting that PP2C interacted physically with subclass III TaSnRK2s, while having no interaction with TaSnRK2s from subclasses I and II. It was concluded that subclass III TaSnRK2s were involved in ABA regulated stress responses, whereas subclasses I and II TaSnRK2s responded to other abiotic stressors in an ABA-independent manner. Rice SnRK2 orthologs, SAPK8, SAPK9 and SAPK10 have also been shown to phosphorylate the AREB1 ortholog TRAB1, as demonstrated through experiments conducted in vitro (Johnson et al. 2002; Kagaya et al. 2002; Kobayashi et al. 2005).
MAPKs represent another class of important kinases that are generally involved in phosphorylation of WRKY and MYB/MYC TFs, thus playing a crucial role in signal transduction as a response to external stresses (Sheikh et al. 2016). It has been shown that the WRKY TFs activate expression of their targets genes (containing W-box cis-element) only after phosphorylation of their conserved regions by MAPKs; these cascades are also involved in transduction of downstream signals in ABA-dependent manner during stress response (Banerjee and Roychoudhury 2015; Fig. 3b). For instance, a MAPK in Arabidopsis (AtMPK3) phosphorylates AtWRKY46 TF (Sheikh et al. 2016), and in rice, OsMPK3 phosphorylates OsWRKY30, which is known to enhance drought tolerance (Shen et al. 2012). An example of MAPK mediated phosphorylation of MYB TFs is also available in pine (Pinus taeda). In this study, a MAPK protein (PtMAPK6) phosphorylates a Ser-236 residue, located in the C-terminal activation domain of MYB TF (PtMYB4) during early stages of xylem development (Morse et al. 2009).
In wheat also, 19 MAPK genes (including six MAP kinase kinase kinase genes, two MAP kinase kinase genes, and 11 MAP kinase genes) were recently characterized (Wen et al. 2015). Apparently these kinases are involved in phosphorylation in response to deprivation of N/P, salinity and drought. Temporal expression profiles of these MAPKs indicated that these genes were each regulated by stress and exhibited typical recovery responses, when these genes were exposed again to normal growth conditions.
Other kinases namely Calcium-dependent protein kinases (CDPKs), OPEN STOMATA 1 (OST1) and Calcium/calmodulin-dependent protein kinase (CCaMK) are also involved in phosphorylation of TFs. CDPKs, (also written as CPKs; e.g., Arabidopsis AtCPK1–AtCPK34), are exclusive to plants (and certain protists) and have been reported to be involved in Ca2+-responsive kinase activity in plants during abiotic stress response and abscisic acid (ABA) signaling (Cheng et al. 2002; Wei et al. 2014). For instance, Arabidopsis CDPKs (CPK4 and CPK11) are involved in ABA signaling by phosphorylating bZIP TFs (ABF1 and ABF4) (Zhu S-Y et al. 2007) (Fig. 3c). Also, in in vitro kinase assays of CPK3 protein with a suite of substrates demonstrated that CPK3 phosphorylates some TFs (including ERF1, HsfB2a and CZF1/ZFAR1) in the presence of Ca2+ (Kanchiswamy et al. 2010). OST1 was initially identified for its role in stomatal closure in response to drought in Arabidopsis (Mustilli et al. 2002). In maize, ZmOST1 was shown to cause phosphorylation of the ZmSNAC1 (Vilela et al. 2013) and ZmCCaMK was shown to phosphorylate ZmNAC84 (Zhu et al. 2016) leading to enhanced drought tolerance in both the cases.
The above examples suggest that the system of regulation of TFs due to reversible phosphorylation by different protein kinases/phosphatases should be conserved among plant species, which needs further investigation for a better understanding of the mechanism involved in activation of stress responsive genes under drought.
Database for TFs in plants
Efforts to develop databases for eukaryotic TFs started more than 20 years ago (Wingender 1988) and resulted in a number of publicly available plant TF databases (Table 3). PlnTFDB was the first comprehensive database for plant TFs and initially contained TFs for the following five species: A. thaliana, P. trichocarpa, O. sativa, Chlamydomonas reinhardtii and Ostreococcus tauri (Riano-Pachon et al. 2007). Current version of PlnTFDB (v3.0) contains a total of 26,184 distinct proteins representing 84 families of different TFs and transcriptional regulators from 19 plant species (Pérez-Rodríguez et al. 2010). Another important comprehensive database for plant TFs is PlantTFDB 1.0 (Guo et al. 2008) with 26,402 TFs from 22 species; its latest updated version (PlantTFDB 3.0) contains information for 129,288 TFs representing 58 families of TFs from 83 species (Jin et al. 2014); genome sequences are also available for 67 of these 83 species.
Two wheat specific TF databases (wDBTF; Romeuf et al. 2010 and wheatTFDB; Chen et al. 2015) are also publicly available. The wDBTF contains 3,820 wheat TF genes belonging to 40 families and 84 subfamilies (Romeuf et al. 2010). This includes 295 AP2/EREBP, 167 bZIP, 116 MYB, 269 NAC and 187 WRKY TFs, which are actively involved in drought tolerance. Similarly, wheatTFDB contains 2,407 putative TFs belonging to 63 families. This database also includes 226 AP2/EREBP, 110 bZIP, 127 MYB, 193 NAC and 135 WRKY TFs, which are actively involved in drought tolerance (Chen et al. 2015). This database also includes 1,257 developmental stage-specific TFs and 1104 tissue-specific TFs, which were identified based on publicly available gene expression databases. The above two wheat databases will be useful in identifying target TFs involved in response to drought stress at a specific stage of development.
Databases for TFs and corresponding cis-elements in the promoters of drought responsive genes belonging to individual plant species other than wheat are also available. For instance, TRANSFAC is an important database of cis-acting elements and trans-acting TFs (Matys et al. 2006), which mainly includes TFs from A. thaliana. Other TF databases focusing on individual plant species include the following: (1) AtTFDB (Davuluri et al. 2003), DATF (Guo A et al. 2005), RARTF (Iida et al. 2005) and STIFDB (Shameer et al. 2009) for A. thaliana; (2) DRTF (Gao et al. 2006) and RiceSRTFDB (Priya and Jain 2013) for rice (O. sativa), (3) DPTF (Zhu QH et al. 2007) for P. tricharpa, (4) TOBFAC (Rushton et al. 2008) for N. tabacum, (5) SoyTFKB (http://www.igece.org/Soybean_TF) and SoyDB (Wang et al. 2010) for soybean (G. max), and (6) DMTR (Mochida et al. 2010) for M. truncatula.
A ‘plant cistrome database (PlantCistromeDB)’ giving details of genome-wide transcription factor binding sites, the TFBSs (cis-elements) including cistrome and epicistrome has also been recently developed (O’Malley et al. 2016). In this database, TFBS for 529 Arabidopsis TFs (of 1,812 TFs comprising 80 families, surveyed) and details of methylated sites, which inhibit or promote TF binding are available (http://neomorph.salk.edu/ PlantCistromeDB). In future, data on cistromes and epicistromes of other plant species will certainly be added to this PlantCistromeDB.
Cross-talks and networking involving TFs
There is evidence that individual TFs generally do not function in isolation and that often there are cross-talks between the signaling pathways involving two or more TFs. At least two examples of such cross-talks of TFs from the family ABF (a subfamily) of bZIP are available, one involving crosstalk with DREB, and the other involving cross-talk with NAC. However, such interactions are yet to be discovered in wheat.
Interaction between ABF and DREB
The TFs belonging to ABF and DREB families are often involved in a network as shown in case of Arabidopsis drought-responsive gene RD29A, which carries with it two cis-elements, ABRE and DRE/CRT-core motifs that are used for binding of two different TFs. A detailed study of the involvement of ABRE and DRE in ABA-dependent gene expression suggested that ABRE and DRE are interdependent and are synergistic in action for induction of the stress-responsive expression of RD29A gene. DRE motif sometimes also functions as a coupling element (CE) for ABRE (Narusaka et al. 2003).
Using yeast two hybrid system, ABF TFs (ABF2, ABF3 and ABF4) have also been shown to interact physically with DREB/CBF TFs (DREB1A/CBF3, DREB2A and DREB2C) (Lee et al. 2010b). In another study, it was found that DREB2A promoter contains ABRE and coupling element 3 (CE3)-like sequences that are necessary for promoter activity under osmotic stress conditions, such as dehydration. Using yeast one-hybrid and chromatin immunoprecipitation (ChIP) assays, it was also found that the AREB/ABF bZIP TFs can bind DREB2A promoter and activate the expression of DREB2A gene in an ABRE-dependent manner (Kim et al. 2011).
Interaction between ABF and NAC
Interactions between the AREB/ABF and NAC TFs have also been demonstrated in some studies. For instance, ATAF1 (an Arabidopsis NAC TF) has been shown to regulate the ABA biosynthetic gene 9-cis-epoxycarotenoid dioxygenases 3 (NCED3) in Arabidopsis, suggesting that SNACs may regulate the expression of genes from AREB/ABF regulons (Jensen et al. 2013). Cooperation between NAC TF and TFs AREB/ABF of the bZIP family during dehydration stress was also reported in two other studies. In one of these studies, ANAC096 was shown to interact synergistically with ABF2 and ABF4 for inducing activation of RD29A (Xu et al. 2013), while in the other study, a NAC TF (ANAC016) and the protein encoded by its target gene NAP, were both shown to bind to a site in the promoter of AREB1, leading to repression of AREB1 transcription. This was described as a trifurcate feed-forward pathway involving functions of NAC016, NAP, and AREB1 in the drought stress response (Sakuraba et al. 2015).
MicroRNA and TF co-regulatory networks for drought tolerance
Generally, TFs regulate expression of protein-coding genes at the transcription level (Fire et al. 1998; Stefani and Slack 2008). However, during last more than a decade, non-coding RNA mediated riboregulation of gene expression at the post-transcriptional level has attracted attention of the scientific community. A major component of this riboregulation is a class of small non-coding RNA represented by microRNAs (miRNAs), which are ~19–24 bp long and repress expression of a large number of genes in all known eukaryotic systems (Chen 2004; Cobb et al. 2005). Interestingly, about 50 % of the miRNA targets that have been identified are genes encoding TFs, which are involved in regulation of abiotic stress responses in plants (Bartel 2004; Kidner and Martienssen 2005; Zhang et al. 2006; Zhang, 2015; Zhang and Wang 2015).
In bread wheat and emmer wheat also, expression of miRNAs under drought stress was examined. For instance, a number of miRNAs (miR1867, miR474, miR398, miR1450, miR1881, miR894, miR156 and miR1432) are induced by drought in wild emmer wheat (Triticum dicoccoides) (Kantar et al. 2011). Similarly, in bread wheat genotype C306, following five miRNAs were found to be differentially expressed under dehydration stress (Gupta et al. 2014): miR159, miR164, miR393, miR529 and miR1029. In another study, 367 miRNAs were found to be differentially expressed, when a drought tolerant genotype (Hanxuan10) and a drought sensitive genotype (Zhengyin1) were grown under dehydration stress, and expression analysis was conducted using a deep-sequencing method (Ma et al. 2015). The gene targets of the differentially expressed miRNAs encoded proteins that were involved in metabolic processes, response to stress, and regulation of transcription (including TFs). For instance, gene target of miR159a encodes the MYB3 TF, which plays a role in drought-stress. The target of miR164b is the NAC TF family, which is also having functions related to various abiotic stresses including drought (Ma et al. 2015). Majority of miRNAs regulate plant growth and development involving plant architecture, and are therefore also involved in imparting tolerance to biotic and abiotic stresses. Sometimes, they do so by controlling the level of TFs (Zhang et al. 2006; Rubio-Somoza and Weigel 2011; Ma et al. 2015). In such cases, there is a complex networking between TFs and miRNAs, which ultimately regulate gene expression (Fig. 4). The function of one TF may be affected by one or more miRNAs, and one miRNA may affect more than one TF (Zhang et al. 2006). It is also known that the synthesis of a specific miRNA itself may also be affected by one or more TFs and that one TF may affect synthesis of more than one miRNAs (Gutierrez et al. 2009; Meng et al. 2011). Because the expression of miRNAs often depends on the regulation of TF synthesis and vice versa, it is not surprising that both families of regulators are tightly related to each other in gene regulatory networks. The coordinated transcriptional regulation of miRNAs and their target genes by TFs also involves a network, which will be illustrated using some specific examples of coregulation by miRNAs and TFs in wheat and in two model plants (Arabidopsis and rice). A summary of miRNAs and their target TFs is given in Table 4.
miR164 and miR169 and NAC/NFY TFs co-regulate drought tolerance
The miRNAs form miRNA164 family are crucial players in regulation of response to drought stress; they target and regulate the activity of a number of TFs of the NAC family, which in turn regulate signal transduction pathways involved in a variety of developmental processes (lateral root emergence, formation of vegetative and floral organs and age-dependent cell death) thus imparting tolerance against abiotic stresses (Guo HS et al. 2005; Kim et al. 2009). In wheat, it has been shown that miR164 accumulates in the seedlings of the cultivar C-306 during drought stress (Gupta et al. 2014), suggesting that, miR164 might target NAC TFs and influence response to drought stress (including root development) as earlier reported in rice and Arabidopsis (Kim et al. 2009; Fang et al. 2014). In rice, six miR164-targeted NAC genes (OMTN1–OMTN6) were characterized for drought response. Over-expression of four of these six genes (OMTN2, OMTN3, OMTN4 and OMTN6) significantly reduced drought tolerance at the reproductive stage, suggesting that the OMTNs function as negative regulators of drought tolerance in rice, in addition to their role in some developmental processes (Fang et al. 2014).
Another important miRNA family that is involved in response to drought stress is miR169, which targets CAAT-box NFY TFs (Li et al. 2008), which are responsive to drought stress in wheat (Stephenson et al. 2007). In another study in wheat, it was shown that, miR169d was repressed in the drought-tolerant cultivar after dehydration stress, which might target the CAAT-box TFs, and also influence ABA-responsive transcription resulting in enhanced drought tolerance (Ma et al. 2015). Studies have also been conducted in model plant Arabidopsis, where miRNA169 targets NFY TFs and regulates drought tolerance (Zhao et al. 2009).
miR160, miR167 and ARF TFs co-regulate adventitious rooting
In Arabidopsis, miR160 and miR167 are known to regulate ARF TFs at post-transcription level (ARF10, ARF16 and ARF17 in case of miR160; ARF6 and ARF8 in case of miR167) and are involved in development of adventitious roots (Rhoades et al. 2002; Mallory et al. 2005; Gutierrez et al. 2009). ARF17 is a negative regulator, whereas ARF6 and ARF8 are positive regulator of adventitious rooting, suggesting their involvement in a response to drought through root development. The above three ARFs also regulate each other’s expression at both transcriptional and posttranscriptional levels by modulating the availability of miR160 and miR167. This feedback regulation of microRNA homeostasis by their target TFs causes adventitious rooting in Arabidopsis (Gutierrez et al. 2009).
miR156, miR172 and SPL/AP2-like TFs co-regulate developmental timing
In Arabidopsis, the regulatory network for developmental timing involves two miRNAs (miR156 and miR172) and some TFs from SPL/AP2 family (Wu et al. 2009). The miR156 targets 10 members of the SPL TF family (SPL2, SPL3, SPL4, SPL5, SPL6, SPL9, SPL10, SPL11, SPL13 and SPL15), while miR172 targets six AP2-like TFs (AP2, TOE1, TOE2, TOE3, SMZ and SNZ). However, since, five of the above 10 SPL TFs (SPL3, SPL4, SPL5, SPL9 and SPL10) are involved in regulation of developmental timing, it is evident that miR156 also regulates developmental timing by repressing the expression of these five SPL TFs. Two of the above five SPLs (SPL9 and SPL10) also promote the transcription of miR172 indicating the involvement of miR156 in the regulation of expression of miR172 via SPLs 9 and 10. The same two SPLs are in turn involved in regulation of miR156 and miR172 thus forming a negative feedback loop. These interlinking connections ultimately form a regulatory cascade miR156-SPL-miR172-AP2 which is involved in developmental timing in Arabidopsis (Wu et al. 2009), thus suggesting their indirect involvement in drought tolerance.
The above results suggest that the TF–miRNA regulation is one of the most important aspects of the study of both miRNAs and TFs for their role in drought stress. Recent interest in TF–miRNA regulation led to the creation of a manually curated database called ‘TransmiR’ (Wang et al. 2009), which is a valuable resource for the study of TF–miRNA regulation. This database would be helpful for understanding not only the mechanisms by which TFs regulate miRNAs, but also about their contribution to plant development and tolerance to biotic/abiotic stress.
TF genes versus major QTL for yield under drought stress
A number of QTLs for drought tolerance have also been reported in wheat and several other crops. Since genes encoding TFs involved in drought tolerance are also known, it was considered fruitful to examine if at least some of the QTLs may actually represent the genes encoding TFs involved in drought tolerance. For this purpose, BLAST search was conducted against the mapped wheat sequences available at EnsemblPlants database (Yates et al. 2016). Interestingly, out of 42 wheat TF genes, nine TF genes (TaNAC29, TaABP1, TaWRKY10, TaWRKY12, TaWRKY16, TaWRKY19, TaWRKY20, TaWRKY44 and TaWRKY82) were mapped on five wheat chromosomes/arms that were already known to harbor major QTLs for drought tolerance (Table 5; Table S2; Fig. S3; Quarrie et al. 2006; Kirigwi et al. 2007; Pinto et al. 2010; Alexander et al. 2012; Kadam et al. 2012; Nezhad et al. 2012; Maphosa et al. 2014; Shukla et al. 2014). Phenotypic variance explained (PVE; R 2) for these QTLs ranged from 14.0 to 23.9 %.
Presence of both drought TFs and major yield QTL under drought stress on the same chromosome indicated that the above-mentioned five wheat chromosomes are important in imparting drought tolerance. However, it was not possible to identify specific regions of these chromosomes, because at least in some cases, the positions of TF genes and QTLs did not seem to overlap. Further studies to find out if some of the QTLs represent genes encoding TFs may be rewarding.
Meta-QTLs (MQTLs) with precise and narrow confidence intervals prove useful in deducing the candidate genes for the concerned trait. MQTL analyses for grain yield and its related traits under drought stress have been conducted in rice (Swamy et al. 2011), barley (Li et al. 2013) and wheat (Acuna-Galindo et al. 2015). In these studies several candidate genes located in the regions of MQTLs were reported, and were found to be stress-inducible, and were known to be involved in growth, development, signal transduction (including TFs) and sugar transport. The important TF genes includes NAC, MADS box, Zinc finger and CCAAT box. The candidacy of above genes including TF genes for their role in drought stress has already been proved in rice and other crops (Swamy et al. 2011).
Transgenic plants for drought tolerance
A number of reports are now available on development of transgenic plants for drought tolerance using TF genes (Yang et al. 2010); these reports are listed in a recent review (Wang H et al. 2016). However, no drought-tolerant cultivar has been released using these TFs, although a transgenic wheat developed for DREB1A gene at Japan International Research Center for Agricultural Sciences is under field testing (Blum 2014; Waltz 2014). One of the limitations in such studies is lack of precision in phenotyping for drought tolerance.
For developing drought-tolerant transgenics, phenotyping is generally done under controlled environments and the results generally are not reproducible under field conditions. This led to the failure in utilizing the results of these studies in plant breeding programs. In wheat, 14 transgenic wheat lines (DREB1A-wheat) that were selected under greenhouse conditions for survival to severe drought (SURV) and for high water use efficiency (WUE) failed to outperform the controls in terms of grain yield under water deficit; however, the events selected for WUE were identified as lines that combine an acceptable yield under well irrigated conditions (Pierre et al. 2012). Most of the studies for drought tolerance in transgenic plants suggested improved performance under laboratory and greenhouse conditions only (Dubouzet et al. 2003; Yang et al. 2010; Wang H et al. 2016), except few studies that confirmed improved drought resistance in transgenics under field conditions [SNAC1 (Hu et al. 2006), NFYB1 (Nelson et al. 2007), and DREB1A/CBF3 (Xiao et al. 2009)].
From the above account, it is obvious that phenotyping for drought tolerance is very crucial and should be conducted under suboptimal field conditions (Yang et al. 2010). This will enable us to draw correct conclusions about the real function of the discovered genes towards drought tolerance and their utilizations in plant breeding. Since drought tolerance is a complex trait, and several phenotypic parameters are available to measures it, a combination of the following phenotypic parameters may prove useful for improving yield under water-limited conditions- (1) water use efficiency, (2) root architecture, (3) carbon isotope discrimination, (4) stomatal conductance, (5) osmotic adjustment, (6) remobilization of water-soluble carbohydrates, (7) chlorophyll concentration, (8) stay-green, and (9) delayed leaf senescence (Gupta et al. 2012; Tuberosa 2012).
Molecular markers for TF genes and their possible use in MAS
Marker-assisted selection (MAS) involving markers associated with TF genes can also be used for the development of drought tolerant genotypes (Lata et al. 2011; Budak et al. 2013, 2015). As many as 16 gene specific markers have been listed by Budak et al. (2015), of which 10 markers are associated with genes for DREB and WRKY TFs (Wei et al. 2009; Huseynova and Rustamova 2010; Mondini et al. 2012, 2015; Edae et al. 2013). In one of these studies (Wei et al. 2009), five primer pairs based on the available sequences of DREB1 genes in common wheat and related species were designed. Two of these primer pairs (P21F/P21R and P25F/PRa) were A genome specific (located on 3A), one primer pair (P18F/P18R) was B genome specific (located on 3B), and two primer pairs (P20F/P20R and P22F/PRa) were D genome specific (located on 3D). The DREB1 gene that was amplified using primer pair specific to B genome carried two SNPs (S646 and S770), which showed polymorphism between the two parents of a mapping population (Opata 85 and W7984). However, no polymorphism was observed between the orthologues from A and D genomes. One of these SNPs (S770) also helped in mapping the DREB-B1 gene between the markers Xmwg818 and Xfbb117 on chromosome arm 3BL (Wei et al. 2009). In another study, four SNPs were identified within DREB1A, which were found to be associated with four traits, including final biomass, normalized vegetation index (NDVI), days to heading, and spikelet number. The percentage of phenotypic variation explained by those SNPs ranged from 6.4 % for heading date to 9.7 % for NDVI (Edae et al. 2013). Also, HRM (high resolution melting) technology was used to identify and characterize SNPs in EREBP/AP2 domain of genes DREB2, DREB3 and DREB4 in some durum wheat (T. turgidum L. var durum) cultivars, which differed in the level of drought tolerance (Mondini et al. 2015).
In addition to the above studies, microsatellite markers derived from genome-wide TF genes were also identified in two legume species. These included a large number of microsatellite markers associated with TF genes that are involved in drought tolerance in chickpea (Kujur et al. 2013) and in Medicago truncatula (Liu et al. 2015). Some of these TFs included the following: AP2/ERF, bZIP, WRKY, NAC, NAM, bHLH and MYB. However, to our knowledge, no such SSRs associated with drought-related TFs have been reported in wheat. Therefore, one may like to search such markers in wheat, so that the same along with those discussed above may be used for the development of drought tolerant wheat cultivars using MAS (Fig. 1).
Conclusions and perspectives
The research area of TF-mediated gene regulation in plants has shown rapid progress in recent years. This resulted in a better understanding of the role of TFs in imparting tolerance against different abiotic stresses including drought (Bartels and Sunkar 2005; Yoshida et al. 2014). Progress has also been made in understanding the complex mechanisms underlying various aspects of plant responses to drought and other abiotic stresses. Nevertheless, keeping in view the fact that as much as 5 % of the genome in a crop plant encodes TFs, further studies on the function of TFs and their use in imparting drought tolerance in crop plants should prove useful.
To achieve a better understanding of the role of TFs in providing tolerance against drought stress, it is critical to identify not only the interaction of TFs with cis-regulatory elements (CREs), but also to identify the target-downstream genes for TFs. Post-transcriptional/post-translational modifications (including phosphorylation) of TFs and miRNA-TF co-regulatory networks under drought stress are other areas of research, which will receive increased attention in future. While conducting these future studies, we also need to keep in mind that the regulation of gene expression by TFs is also highly context-dependent. Human ENCODE project (The ENCODE Project Consortium 2012) provides strong evidence to support the assumption that the binding of a TF to its TF specific binding site (TFBS) is also dependent on its chromatin state, DNA methylation and numerous additional factors (Wang H et al. 2012; Wang J et al. 2012; Yanez-Cuna et al. 2012). This has led to the concept of Epicistrome; the genome wide information on epicistrome has been collected recently in Arabidopsis (O’Malley et al. 2016), and in future, similar information should become available in crop plants (including wheat) also. These features of the function of TFs are difficult to measure and predict, and constitute the so-called ‘‘cellular context’’ that influences the expression level of genes. In future, an understanding of the complex mechanisms of signaling and transcriptional regulation operated by TFs under drought stress will certainly improve. This will help in developing improved drought tolerant cultivars in wheat and in other important crops for increasing yield and quality under drought stress, thus contributing to sustainable agriculture.
Author contribution statement
PKG conceived, outlined, edited and wrote a part of the review. VG prepared outline and wrote the first draft of the manuscript and assisted PKG in finalising the manuscript. VJ wrote MicroRNA and TF co-regulatory networks section and AK wrote the TF database section.
References
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9:1859–1868
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 5:63–78
Achard P, Herr A, Baulcombe DC, Harberd NP (2004) Modulation of floral development by a gibberellin-regulated microRNA. Development 131:3357–3365
Acuna-Galindo MA, Mason RE, Subramanian NK, Hays DB (2015) Meta-analysis of wheat QTL regions associated with adaptation to drought and heat stress. Crop Sci 55:477–492
Alexander LM, Kirigwi FM, Fritz AK, Fellers JP (2012) Mapping and quantitative trait loci analysis of drought tolerance in a spring wheat population using amplified fragment length polymorphism and diversity array technology markers. Crop Sci 52:253–261
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15:2730–2741
Baldoni E, Genga A, Cominelli E (2015) Plant MYB transcription factors: their role in drought response mechanisms. Int J Mol Sci 16:15811–15851
Baloglu MC, MT O¨z, O¨ktem HA, Yu¨cel M (2012) Expression analysis of TaNAC69-1 and TtNAMB-2, wheat NAC family transcription factor genes under abiotic stress conditions in durum wheat (Triticum turgidum). Plant Mol Biol Rep 30:1246–1252
Banerjee A, Roychoudhury A (2015) WRKY proteins: signaling and regulation of expression during abiotic stress responses. Sci World J 2015:807560
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24:23–58
Bennett D, Reynolds M, Mullan D, Izanloo A, Kuchel H, Langridge P, Schnurbusch T (2012) Detection of two major grain yield QTL in bread wheat (Triticum aestivum L.) under heat, drought and high yield potential environments. Theor Appl Genet 125:1473–1485
Blum A (2014) Genomics for drought resistance—getting down to earth. Funct Plant Biol 41:1191–1198
Budak H, Kantar M, Kurtoglu KY (2013) Drought tolerance in modern and wild wheat. Sci World J 548246:1–16
Budak H, Hussain B, Khan Z, Ozturk NZ, Ullah N (2015) From genetics to functional genomics: improvement in drought signaling and tolerance in wheat. Front Plant Sci 6:1012
Cai H, Tian S, Dong H, Guo C (2015) Pleiotropic effects of TaMYB3R1 on plant development and response to osmotic stress in transgenic Arabidopsis. Gene 558:227–234
Cao X, Chen M, Xu Z Chen, Li Y, Yu L, Liu Y, Ma Y (2012) Isolation and functional analysis of the bZIP transcription factor gene TaABP1 from a Chinese wheat landrace. J Integr Agril 11:1580–1591
Chen X (2004) A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303:2022–2025
Chen H, Lai Z, Shi J, Xiao Y, Chen Z, Xu X (2010) Roles of Arabidopsis WRKY18, WRKY40 and WRKY60 transcription factors in plant responses to abscisic acid and abiotic stress. BMC Plant Biol 10:281
Chen Z-Y, Guo X-J, Chen Z-X, Chen W-Y, Liu D-C, Zheng Y-L, Liu Y-X, Wei Y-M, Wang J-R (2015) Genome-wide characterization of developmental stage- and tissue-specific transcription factors in wheat. BMC Genom 16:1–15
Cheng S-H, Willmann MR, Chen H-C, Sheen J (2002) Calcium signaling through protein kinases. The Arabidopsis calcium-dependent protein kinase gene family. Plant Physiol 129:469–485
Choi H, Hong JH, Ha JO, Kang JY, Kim SY (2000) ABFs, a family of ABA-responsive element binding factors. J Biol Chem 275:1723–1730
Cobb BS, Nesterova TB, Thompson E, Hertweck A, O’Connor E, Godwin J, Wilson CB, Brockdorff N, Fisher AG, Smale ST, Merkenschlager M (2005) T cell lineage choice and differentiation in the absence of the RNase III enzyme Dicer. J Exp Med 201:1367–1373
Davuluri RV, Sun H, Palaniswamy SK, Matthews N, Molina C, Kurtz M, Grotewold E (2003) AGRIS: Arabidopsis gene regulatory information server, an information resource of Arabidopsis cis-regulatory elements and transcription factors. BMC Bioinform 4:25
Dixit S, Biswal AK, Min A, Henry A, Oane RH, Raorane ML, Longkumer T, Pabuayon IM, Mutte SK, Vardarajan AR, Miro B, Govindan G, Albano-Enriquez B, Pueffeld M, Sreenivasulu N, Slamet-Loedin I, Sundarvelpandian K, Tsai Y-C, Raghuvanshi S, Hsing Y-IC, Kumar A, Kohli A (2015) Action of multiple intra-QTL genes concerted around a co-localized transcription factor underpins a large effect QTL. Sci Rep 5:15183
Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33:751–763
Edae E, Byrne PF, Manmathan H, Haley SD, Moragues M, Lopes MS, Reynolds MP (2013) Association mapping and nucleotide sequence variation in five drought tolerance candidate genes in spring wheat. Plant Genom 6:13
Egawa C, Kobayashi F, Ishibashi M, Nakamura T, Nakamura C, Takumi S (2006) Differential regulation of transcript accumulation and alternative splicing of a DREB2 homolog under abiotic stress conditions in common wheat. Genes Genet Syst 81:77–91
Fang Y, Xie K, Xiong L (2014) Conserved miR164-targeted NAC genes negatively regulate drought resistance in rice. J Exp Bot 65:2119–21356
Fang Y, Liao K, Du H, Xu Y, Song H, Li X, Xiong L (2015) A stress-responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice. J Exp Bot. doi:10.1093/jxb/erv386
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Floyd SK, Bowman JL (2004) Ancient microRNA target sequences in plants. Nature 428:485–486
Fowler S, Thomashow F (2002) Arabidopsis transcriptome profiling indicates that multiple regulatory pathways are activated during cold acclimation in addition to the CBF cold response pathway. Plant Cell 14:1675–1690
Franco-Zorrilla JM, López-Vidriero I, Carrasco JL, Godoy M, Vera P, Solano R (2014) DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc Natl Acad Sci USA 111:2367–2372
Fredslund J (2008) DATFAP: a database of primers and homology alignments for transcription factors from 13 plant species. BMC Genom 9:140
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran LSP, Yamaguchi-Shinozaki K, Shinozaki K (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39:863–876
Fujita Y, Fujita M, Satoh R, Maruyama K, Parvez MM, Seki M, Hiratsu K, Ohme-Takagi M, Shinozaki K, Yamaguchi-Shinozaki K (2005) AREB1 is a transcription activator of novel AREB dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 17:3470–3488
Fujita Y, Fujita M, Shinozaki K, Yamaguchi-Shinozaki K (2011) ABA-mediated transcriptional regulation in response to osmotic stress in plants. J Plant Res 124:509–525
Furihata T, Maruyama K, Fujita Y, Umezawa T, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2006) Abscisic acid-dependent multisite phosphorylation regulates the activity of a transcription activator AREB1. Proc Natl Acad Sci USA 103:1988–1993
Gao G, Zhong Y, Guo A, Zhu Q, Tang W, Zheng W, Gu X, Wei L, Luo J (2006) DRTF: a database of rice transcription factors. Bioinformatics 22:1286–1287
Guex N, Peitsch MC, Schwede T (2009) Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: A historical perspective. Electrophoresis 30(S1):S162–S173
Guo A, He K, Liu D, Bai S, Gu X, Wei L, Luo J (2005) DATF: a database of Arabidopsis transcription factors. Bioinformatics 21:2568–2569
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to down-regulate auxin signals for Arabidopsis lateral root development. Plant Cell 17:1376–1386
Guo AY, Chen X, Gao G, Zhang H, Zhu QH, Liu XC, Zhong YF, Gu X, He K, Luo J (2008) PlantTFDB: a comprehensive plant transcription factor database. Nucl Acids Res 36:D966–D969
Gupta PK, Balyan HS, Gahlaut V, Kulwal P (2012) Phenotyping, genetic dissection and breeding for tolerance to drought and heat in common wheat: present status and future prospects. Plant Breed Rev 36:85–168
Gupta OP, Meena NL, Sharma I, Sharma P (2014) Differential regulation of microRNAs in response to osmotic, salt and cold stresses in wheat. Mol Biol Rep 41:4623–4629
Gutierrez L, Bussell JD, Pacurar DI, Schwambach J, Pacurar M, Bellini C (2009) Phenotypic plasticity of adventitious rooting in Arabidopsis is controlled by complex regulation of AUXIN RESPONSE FACTOR transcripts and microRNA abundance. Plant Cell 21:3119–3132
Hassan NM, El-Bastawisy ZM, El-Sayed AK, Ebeed HT, Nemat Alla MM (2015) Roles of dehydrin genes in wheat tolerance to drought stress. J Adv Res 6:179–188
He G-H, Xu J-Y, Wang Y-X, Liu J-M, Li P-S, Chen M, Ma Y-Z, Xu Z-S (2016) Drought-responsive WRKY transcription factor genes TaWRKY1 and TaWRKY33 from wheat confer drought and/or heat resistance in Arabidopsis. BMC Plant Biol 16:116
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 35:12987–12992
Hu H, You J, Fang J, Zhu X, Qi Z, Xiong L (2008) Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice. Plant Mol Biol 67:169–181
Huang Q, Yan Wang Y, Li B, Chang J, Chen M, Li K, Yang G, He G (2015) TaNAC29, a NAC transcription factor from wheat, enhances salt and drought tolerance in transgenic Arabidopsis. BMC Plant Biol 15:268
Huseynova I, Rustamova S (2010) Screening for drought stress tolerance in wheat genotypes using molecular markers. Biol Sci 65:132–139
Iida K, Seki M, Sakurai T, Satou M, Akiyama K, Toyoda T, Konagaya A, Shinozaki K (2005) RARTF: database and tools for complete sets of Arabidopsis transcription factors. DNA Res 12:247–256
Jaglo-Ottosen KR, Gilmour SJ, Zarka DG, Schabenberger O, Thomashow MF (1998) Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280:104–106
Jakoby M, Weisshaar B, Dröge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7:106–111
Jensen MK, Kjaersgaard T, Nielsen MM, Galberg P, Petersen K, O’Shea C, Skriver K (2010) The Arabidopsis thaliana NAC transcription factor family: structure-function relationships and determinants of ANAC019 stress signaling. Biochem J 426:183–196
Jensen MK, Lindemose S, De Masi F, Reimer JJ, Nielsen M, Perera V, Workman CT, Turck F, Grant MR, Mundy J, Petersen M, Skriver K (2013) ATAF1 transcription factor directly regulates abscisic acid biosynthetic gene NCED3 in Arabidopsis thaliana. FEBS Open Bio 3:321–327
Jin J, Zhang H, Kong L, Gao G, Luo J (2014) PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucl Acids Res 42:D1182–D1187
Johnson RR, Wagner RL, SD Verhey, Walker-Simmons MK (2002) The abscisic acid-responsive kinase PKABA1 interacts with a seed-specific abscisic acid response element-binding factor, TaABF, and phosphorylates TaABF peptide sequences. Plant Physiol 130:837–846
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Joshi R, Wani SH, Singh B, Bohra A, Dar ZA, Lone AA, Pareek A, Singla-Pareek SL (2016) Transcription factors and plants response to drought stress: Current understanding and future directions. Front Plant Sci 7:1029
Juarez MT, Kui JS, Thomas J, Heller BA, Timmermans MC (2004) microRNA-mediated repression of rolled leaf1 specifies maize leaf polarity. Nature 428:84–88
Kadam S, Singh K, Shukla S, Goel S, Vikram P, Pawar V, Gaikwad K, Khanna-Chopra R, Singh N (2012) Genomic associations for drought tolerance on the short arm of wheat chromosome 4B. Funct Integr Genom 12:447–464
Kagaya Y, Hobo T, Murata M, Ban A, Hattori T (2002) Abscisic acid-induced transcription is mediated by phosphorylation of an abscisic acid response element binding factor, TRAB1. Plant Cell 14:3177–3189
Kanchiswamy CN, Takahashi H, Quadro S, Maffei ME, Bossi S, Bertea C, Zebelo SA, Muroi A, Ishihama N, Yoshioka H, Boland W, Takabayashi J, Endo Y, Sawasaki T, Arimura G (2010) Regulation of Arabidopsis defense responses against Spodoptera littoralis by CPK-mediated calcium signaling. BMC Plant Biol 10:97
Kantar M, Lucas SJ, Budak H (2011) miRNA expression patterns of Triticum dicoccoides in response to shock drought stress. Planta 233:471–484
Kasschau KD, Xie Z, Allen E, Llave C, Chapman EJ, Krizan KA, Carrington JC (2003) P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA unction. Dev Cell 4:205–217
Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1999) Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17:287–291
Kazan K, Manners JM (2013) MYC2: the master in action. Mol Plant 6:686–703
Kidner CA, Martienssen RA (2005) The developmental role of microRNA in plants. Curr Opin Plant Biol 8:38–44
Kim SY (2006) The role of ABF family bZIP class transcription factors in stress response. Physiol Plant 126:519–527
Kim J, Jung JH, Reyes JL, Kim YS, Kim SY, Chung KS, Kim JA, Lee M, Lee Y, Kim VN, Chua NH, Park CM (2005) microRNA directed cleavage of ATHB15 mRNA regulates vascular development in Arabidopsis inflorescence stems. Plant J 42:84–94
Kim JH, Woo HR, Kim J, Lim PO, Lee IC, Choi SH, Hwang D, Nam HG (2009) Trifurcate feed-forward regulation of age-dependent cell death involving miR164 in Arabidopsis. Science 323:1053–1057
Kim S-G, Lee S, Ryu J, Park C-M (2010) Probing protein structural requirements for activation of membrane-bound NAC transcription factors in Arabidopsis and rice. Plant Sci 178:239–244
Kim JS, Mizoi J, Yoshida T, Fujita Y, Nakajima J, Ohori T, Todaka D, Nakashima K, Hirayama T, Shinozaki K, Yamaguchi-Shinozaki K (2011) An ABRE promoter sequence is involved in osmotic stress-responsive expression of the DREB2A gene, which encodes a transcription factor regulating drought-inducible genes in Arabidopsis. Plant Cell Physiol 52:2136–2146
Kim MJ, Park M-J, Seo PJ, Song J-S, Kim H-J, Park C-M (2012) Controlled nuclear import of the transcription factor NTL6 reveals a cytoplasmic role of SnRK2.8 in the drought-stress response. Biochem J 448:353–363
Kirigwi FM, Van Ginkel M, Brown-Guedira G, Gill BS, Paulsen GM, Fritz AK (2007) Markers associated with a QTL for grain yield in wheat under drought. Mol Breed 20:401–413
Kobayashi Y, Murata M, Minami H, Yamamoto S, Kagaya Y, Hobo T, Yamamoto A, Hattori T (2005) Abscisic acid-activated SnRK2 protein kinases function in the gene-regulation pathway of ABA signal transduction by phosphorylating ABA response element-binding factors. Plant J 44:939–949
Kobayashi F, Ishibashi M, Takumi S (2008) Transcriptional activation of Cor/Lea genes and increase in abiotic stress tolerance through expression of a wheat DREB2 homolog in transgenic tobacco. Transgenic Res 17:755–767
Kujur A, Bajaj D, Saxena MS, Tripathi S, Upadhyaya HD, Gowda C, Singh S, Jain M, Tyagi AK, Parida SK (2013) Functionally relevant microsatellite markers from chickpea transcription factor genes for efficient genotyping applications and trait association mapping. DNA Res 20:355–374
Kulik A, Wawer I, Krzywińska E, Bucholc M, Dobrowolska G (2011) SnRK2 protein kinases—key regulators of plant response to abiotic stresses. Omics J Integr Biol 15:859–872
Kume S, Kobayashi F, Ishibashi M, Ohno R, Nakamura C, Takumi S (2005) Differential and coordinated expression of Cbf and Cor/Leagenes during long-term cold acclimation in two wheat cultivars showing distinct levels of freezing tolerance. Genes Genet Syst 80:185–197
Lata C, Prasad M (2011) Role of DREBs in regulation of abiotic stress responses in plants. J Exp Bot 62:4731–4748
Lata C, Bhutty S, Bahadur RP, Majee M, Prasad M (2011) Association of an SNP in a novel DREB2-like gene SiDREB2 with stress tolerance in foxtail millet [Setaria italica (L.)]. J Exp Bot 62:3387–3401
Lata C, Muthamilarasan M, Prasad M (2015) Drought stress responses and signal transduction in plants. In: Pandey KG (ed) Elucidation of Abiotic Stress Signaling in Plants: Functional Genomics Perspectives, vol 2. Springer, New York, pp 195–225
Lauter N, Kampani A, Carlson S, Goebel M, Moose SP (2005) MicroRNA172 down-regulates glossy15 to promote vegetative phase change in maize. Proc Natl Acad Sci USA 102:9412–9417
Lee S-J, Kang J-Y, Park H-J, Kim MD, Bae MS, Choi H-I, Kim SY (2010a) DREB2C interacts with ABF2, a bZIP protein regulating abscisic acid-responsive gene expression, and its overexpression affects abscisic acid sensitivity. Plant Physiol 153:716–727
Lee S-J, Park JH, Lee MH, Yu JH, Kim SY (2010b) Isolation and functional characterization of CE1 binding proteins. BMC Plant Biol 10:277
Li W-X, Oono Y, Zhu J, He X-J, Wu J-M, Iida K, Lu X-Y, Cui X, Jin H, Zhu JK (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and post transcriptionally to promote drought resistance. Plant Cell 20:2238–2251
Li W-T, Liu C, Liu Y-X, Pu Z-E, Dai S-F, Wang J-R, Lan X-J, Zheng Y-L, Wei Y-M (2013) Meta-analysis of QTL associated with tolerance to abiotic stresses in barley. Euphytica 189:31–49
Lippold F, Diego H, Sanchez DH, Musialak M, Schlereth A, Scheible WR, Hincha DK, Udvardi MK (2009) At Myb41 regulates transcriptional and metabolic responses to osmotic stress in Arabidopsis. Plant Physiol 149:1761–1772
Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA-binding domain separate two cellular signal transduction pathways in drought and low temperature-responsive gene expression in Arabidopsis. Plant Cell 10:1391–1406
Liu S, Wang N, Zhang P, Cong B, Lin X, Wang S, Xia G, Huang X (2013) Next-generation sequencing-based transcriptome profiling analysis of Pohlia nutans reveals insight into the stress-relevant genes in Antarctic moss. Extremophiles 17:391–403
Liu W, Jia X, Liu Z, Zhang Z, Wang Y, Liu Z, Xie W (2015) Development and characterization of transcription factor gene-derived microsatellite (TFGM) markers in Medicago truncatula and their transferability in leguminous and non-leguminous species. Molecules 20:8759–8771
Lu G, Gao C, Zheng X, Han B (2009) Identification of OsbZIP72 as a positive regulator of ABA response and drought tolerance in rice. Planta 229:605–615
Luan S (2003) Protein phosphatases in plants. Annu Rev Plant Biol 54:63–92
Ma X, Xin Z, Wang Z, Yang Q, Guo S, Guo X, Cao L, Lin T (2015) Identification and comparative analysis of differentially expressed miRNA in leaves of two wheat (Triticum aestivum L.) genotypes during dehydration stress. BMC Plant Biol 15:21
Mallory AC, Dugas DV, Bartel DP, Bartel B (2004) MicroRNA regulation of NAC-domain targets is required for proper formation and separation of adjacent embryonic, vegetative, and floral organs. Curr Biol 14:1035–1046
Mallory AC, Bartel DP, Bartel B (2005) MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17:1360–1375
Mao X, Zhang H, Tian S, Chang X, Jing R (2010) TaSnRK2.4, an SNF1-type serine/threonine protein kinase of wheat (Tricitum aestivum L.), confers enhanced multi-stress tolerance in Arabidopsis. J Exp Bot 61:683–696
Mao X, Jia D, Li A, Zhang H, Tian S, Zhang X, Jia J, Jing R (2011) Transgenic expression of TaMYB2A confers enhanced tolerance to multiple abiotic stresses in Arabidopsis. Funct Integr Genom 11:445–465
Mao X, Zhang H, Qian X, Li A, Zhao G, Jing R (2012) TaNAC2, a NAC-type wheat transcription factor conferring enhanced multiple abiotic stress tolerances in Arabidopsis. J Exp Bot 63:2933–2946
Mao X, Chen S, Li A, Zhai C, Jing R (2014) Novel NAC transcription factor TaNAC67 confers enhanced multi-abiotic stress tolerances in Arabidopsis. PLoS ONE 9:e84359
Maphosa L, Langridge P, Taylor H, Parent B, Emebiri LC, Kuchel H, Reynolds MP, Chalmers KJ, Okada A, Edwards J, Mather DE (2014) Genetic control of grain yield and grain physical characteristics in a bread wheat population grown under a range of environmental conditions. Theor Appl Genet 127:1604–1624
Marcotte Jr WR, Russell SH, Quatrano RS (1989) Abscisic acid responsive sequences from the em gene of wheat. Plant Cell 1989:969–976
Marcotte WR, Bayley CC, Quatrano RS (1988) Regulation of a wheat promoter by abscisic acid in rice protoplasts. Nature 335:454–457
Maruyama K, Sakuma Y, Kasuga M, Ito Y, Seki M, Goda H, Shimada Y, Yoshida S, Shinozaki K, Yamaguchi-Shinozaki K (2004) Identification of cold-inducible downstream genes of the Arabidopsis DREB1A/CBF3 transcriptional factor using two microarray systems. Plant J 38:982–993
Matys V, Kel-Margoulis OV, Pricke E, Liebich I, Land S, Barre-Dirrie A, Reuter I, Chekmenev D, Krull M, Hornischer K, Voss N, Stegmaier P, Lewicki-Potapov B, Saxel H, Kel AE, Wingender E (2006) TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucl Acids Res 2006:D108–D110
Meng Y, Shao C, Wang H, Chen M (2011) The regulatory activities of plant microRNAs: a more dynamic perspective. Plant Physiol 157:1583–1595
Mochida K, Yoshida T, Sakurai T, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2010) LegumeTFDB: an integrative database of Glycine max, Lotus japonicus and Medicago truncatula transcription factors. Bioinformatics 26:290–291
Mondini L, Nachit M, Porceddu E, Pagnotta MA (2012) Identification of SNP mutations in DREB1, HKT1, and WRKY1 genes involved in drought and salt stress tolerance in durum wheat (Triticum turgidum L. vardurum). OMICS 16:178–187
Mondini L, Nachit M, Pagnotta MA (2015) Allelic variants in durum wheat (Triticum turgidum L.var. durum) DREB genes conferring tolerance to abiotic stresses. Mol Genet Genom 290:531–544
Morse AM, Whetten RW, Dubos C, Campbell MM (2009) Post-translational modification of an R2R3-MYB transcription factor by a MAP Kinase during xylem development. New Phytol 183:1001–1013
Mustilli AC, Merlot S, Vavasseur A, Fenzi F, Giraudat J (2002) Arabidopsis OST1 protein kinase mediates the regulation of stomatal aperture by abscisic acid and acts upstream of reactive oxygen species production. Plant Cell 14:3089–3099
Nakashima K, Ito Y, Yamaguchi-Shinozaki K (2009) Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol 149:88–95
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819:97–103
Nakashima K, Yamaguchi-Shinozaki K, Shinozaki K (2014) The transcriptional regulatory network in the drought response and its crosstalk in abiotic stress responses including drought, cold, and heat. Front Plant Sci 5:170
Narusaka Y, Nakashima K, Shinwari ZK, Sakuma Y, Furihata T, Abe H, Narusaka M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high-salinity stresses. Plant J 34:137–148
Nelson D, Repetti P, Adams T, Creelman R, Wu J, Warner D, Anstrom D, Bensen R, Castiglioni P, Donnarummo M, Hinchey B, Kumimoto R, Maszle D, Canales R, Krolikowski K, Dotson S, Gutterson N, Ratcliffe O, Heard J (2007) Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc Natl Acad Sci USA 104:16450–16455
Nezhad K, Weber WE, Roder MS, Sharma S, Lohwasser U, Meyer RC, Saal B, Borner A (2012) QTL analysis for thousand-grain weight under terminal drought stress in bread wheat (Triticum aestivum L.). Euphytica 186:127–138
Niu X, Helentjaris T, Bate NJ (2002) Maize ABI4 binds coupling element1 in abscisic acid and sugar response genes. Plant Cell 14:2565–2575
Niu CF, Wei W, Zhou QY, Tian AG, Hao YJ, Zhang WK, Mai B, Lin Q, Zhang ZB, Zhang JS, Chen SY (2012) Wheat WRKY genes TaWRKY2 and TaWRKY19 regulate abiotic stress tolerance in transgenic Arabidopsis plants. Plant, Cell Environ 35:1156–1170
O’Malley RC, Huang SC, Song L, Lewsey MG, Bartlett A, Nery JR, Galli M, Gallavotti A, Ecker JR (2016) Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165:1280–1292
Okay S, Derelli E, Unver T (2014) Transcriptome-wide identification of bread wheat WRKY transcription factors in response to drought stress. Mol Genet Genom 289:765–781
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P, Hayashizaki Y, Suzuki K, Kojima K, Takahara Y, Yamamoto K, Kikuchi S (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10:239–247
Palatnik JF, Allen E, Wu X, Schommer C, Schwab R, Carrington JC, Weigel D (2003) Control of leaf morphogenesis by microRNAs. Nature 425:257–263
Pellegrineschi A, Reynolds M, Pacheco M, Brito RM, Almeraya R, Yamaguchi-Shinozaki K, Hoisington D (2004) Stress-induced expression in wheat of the Arabidopsis thaliana DREB1A gene delays water stress symptoms under greenhouse conditions. Genome 47:493–500
Pérez-Rodríguez P, Riaño-Pachón DM, Corrêa LG, Rensing SA, Kersten B, Mueller-Roeber B (2010) PlnTFDB: updated content and new features of the plant transcription factor database. Nucl Acids Res 38:D822–D8227
Pierre CS, Crossa JL, Bonnett D, Yamaguchi-Shinozaki K, Reynolds MP (2012) Phenotyping transgenic wheat for drought resistance. J Exp Bot 63:1799–1808
Pinto RS, Reynolds MP, Mathews KL, McIntyre CL, Olivares-Villegas JJ, Chapman SC (2010) Heat and drought adaptive QTL in a wheat population designed to minimize confounding agronomic effects. Theor Appl Genet 121:1001–1021
Priya P, Jain M (2013) RiceSRTFDB: a database of rice transcription factors containing comprehensive expression, cis-regulatory element and mutant information to facilitate gene function analysis. Database 2013:bat027
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17:369–381
Qin Y, Wang M, Tian Y, He W, Han L, Xia G (2012) Over-expression of TaMYB33 encoding a novel wheat MYB transcription factor increases salt and drought tolerance in Arabidopsis. Mol Biol Rep 39:7183–7192
Quarrie SA, Quarrie SP, Radosevic R, Rancic D, Kaminska AS, Barnes JD, Leverington M, Ceolini C, Dodig D (2006) Dissecting a wheat QTL for yield present in a range of environments: from the QTL to candidate genes. J Exp Bot 57:2627–2637
Ren X, Chen Z, Liu Y, Zhang H, Zhang M, Liu Q, Hong X, Zhu J-K, Gong Z (2010) ABO3, a WRKY transcription factor, mediates plant responses to abscisic acid and drought tolerance in Arabidopsis. Plant J 63:417–429
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP (2002) Prediction of plant microRNA targets. Cell 110:513–520
Riano-Pachon DM, Ruzicic S, Dreyer I, Mueller-Roeber B (2007) PlnTFDB: an integrative plant transcription factor database. BMC Bioinform 8:42
Richardt S, Lang D, Reski R, Frank W, Rensing SA (2007) PlanTAPDB, a phylogeny-based resource of plant transcription-associated proteins. Plant Physiol 143:1452–1466
Riechmann JL, Meyerowitz EM (1998) The AP2/EREBP family of plant transcription factors. Biol Chem 379:633–646
Riechmann JL, Heard J, Martin G, Reuber L, Jiang C, Keddie J, Adam L, Pineda O, Ratcliffe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu G (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110
Romeuf I, Tessier D, Dardevet M, Branlard G, Charmet G, Ravel C (2010) wDBTF: an integrated database resource for studying wheat transcription factor families. BMC Genom 11:185
Rong W, Qi L, Wang A, Ye X, Du L, Liang H, Xin Z, Zhang Z (2014) The ERF transcription factor TaERF3 promotes tolerance to salt and drought stresses in wheat. Plant Biotechnol J 12:468–479
Rubio-Somoza I, Weigel D (2011) MicroRNA networks and developmental plasticity in plants. Trends Plant Sci 16:258–264
Rushton PJ, Bokowiec MT, Laudeman TW, Brannock JF, Chen X, Timko MP (2008) TOBFAC: the database of tobacco transcription factors. BMC Bioinform 9:53
Rushton PJ, Somssich IE, Ringler P, Shen QXJ (2010) WRKY transcription factors. Trends Plant Sci 15:247–258
Sakuraba Y, Kim Y-S, Han S-H, Lee B-D, Paek N-C (2015) The Arabidopsis transcription factor NAC016 promotes drought stress responses by repressing AREB1 transcription through a trifurcate feed-forward regulatory loop involving NAP. Plant Cell 27:1171–1781
Saleh A, Lumreras V, Pages M (2005) Functional role of DRE binding transcription factors in abiotic stress. In: Tuberosa R, Phillips RL, Gale M (eds) Proceedings of the International Congress ‘In the Wake of the Double Helix From the Green Revolution to the Gene Revolution’, 27–31 May 2003, Bologna, Italy, 193–205, 2005 Avenue media. Italy, Bologna
Schwab R, Palatnik JF, Riester M, Schommer C, Schmid M, Weigel D (2005) Specific effects of microRNAs on the plant transcriptome. Dev Cell 8:517–527
Seki M, Narusaka M, Abe H, Kasuga M, Yamaguchi-Shinozaki K, Carninci P, Hayashizaki Y, Shinozaki K (2001) Monitoring the expression pattern of 1300 Arabidopsis genes under drought and cold stresses by using a full-length cDNA microarray. Plant Cell 13:61–72
Seo JS, Joo J, Kim MJ, Kim YK, Nahm BH, Song SI, Cheong JJ, Lee JS, Kim JK, Choi YD (2011) OsbHLH148, a basic helix-loop-helix protein, interacts with OsJAZ proteins in a jasmonate signalling pathway leading to drought tolerance in rice. Plant J 65:907–921
Seo PJ, Lee SB, Suh MC, Park M-J, Go YS, Park CM (2011) The MYB96 transcription factor regulates cuticular wax biosynthesis under drought conditions in Arabidopsis. Plant Cell 23:1138–1152
Septiningsih EM, Pamplona AM, Sanchez DL, Neeraja CN, Vergara GV, Heuer S, Ismail AM, Mackill DJ (2009) Development of submergence-tolerant rice cultivars: the Sub1 locus and beyond. Ann Bot 103:151–160
Shahnejat-Bushehri S, Tarkowska D, Sakuraba Y, Balazadeh S (2016) Arabidopsis NAC transcription factor JUB1 regulates GA/BR metabolism and signalling. Nat Plants 2:16013
Shameer K, Ambika S, Varghese SM, Karaba N, Udayakumar M, Sowdhamini R (2009) STIFDB-Arabidopsis stress responsive transcription factor database. Int J Plant Genom 2009:583429
Shang Y, Yan L, Liu Z-Q, Cao Z, Mei C, Xin Q, Wu F-Q, Wang X-F, Du S-Y, Jiang T, Zhang X-F, Zhao R, Sun H-L, Liu R, Yu Y-T, Zhang D-P (2010) The Mg-chelatase H subunit of Arabidopsis antagonizes a group of WRKY transcription repressors to relieve ABA-responsive genes of inhibition. Plant Cell 22:1909–1935
Sheikh AH, Eschen-Lippold L, Pecher P, Hoehenwarter W, Sinha AK, Scheel D, Lee J (2016) Regulation of WRKY46 transcription factor function by mitogen-activated protein kinases in Arabidopsis thaliana. Front Plant Sci. doi:10.3389/fpls.2016.00061
Shen Q, Zhang P, Ho T-HD (1996) Modular nature of abscisic acid (ABA) response complexes: composite promoter units that are necessary and sufficient for ABA induction of gene expression in barley. Plant Cell 8:1107–1119
Shen YG, Zhang WK, He SJ, Zhang JS, Liu Q, Chen SY (2003) An EREBP/AP2-type protein in Triticum aestivum was a DRE binding transcription factor induced by cold, dehydration and ABA stress. Theor Appl Genet 106:923–930
Shen H, Liu C, Zhang Y, Meng X, Zhou X, Chu C, Wang X (2012) OsWRKY30 is activated by MAP kinases to confer drought tolerance in rice. Plant Mol Biol 80:241–253
Shin D, Moon S-J, Han S, Kim B-G, Park SR, Lee S-K, Yoon H-J, Lee HE, Kwon H-B, Baek D, Yi BY, Byun M-O (2011) Expression of StMYB1R-1, a novel potato single MYB-like domain transcription factor, increases drought tolerance. Plant Physiol 155:421–432
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Shukla S, Singh K, Patil RV, Kadam S, Bharti S, Prasad P, Singh NK, Khanna-Chopra R (2014) Genomic regions associated with grain yield under drought stress in wheat (Triticum aestivum L.). Euphytica 203:449–467
Singh D, Laxmi A (2015) Transcriptional regulation of drought response: a tortuous network of transcriptional factors. Front Plant Sci 6:895
Stefani G, Slack FJ (2008) Small non-coding RNAs in animal development. Nat Rev Mol Cell Biol 9:219–230
Stephenson TJ, McIntyre CL, Collet C, Xue GP (2007) Genome-wide identification and expression analysis of the NF-Y family of transcription factors in Triticum aestivum. Plant Mol Biol 65:77–92
Stockinger EJ, Gilmour SJ, Thomashow MF (1997) Arabidopsis thaliana CBF1 encodes an AP2 domain–containing transcriptional activator that binds to the C-repeat/DRE, a cis-acting DNA regulatory element that stimulates transcription in response to low temperature and water deficit. Proc Natl Acad Sci USA 94:1035–1040
Sun H, Huang X, Xu X, Lan H, Huang J, Zhang HS (2012) ENAC1, a NAC transcription factor, is an early and transient response regulator induced by abiotic stress in rice (Oryza sativa L.). Mol Biotechnol 52:101–110
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16:2001–2019
Swamy BPM, Vikram P, Dixit S, Ahmed HU, Kumar A (2011) Meta-analysis of grain yield QTL identified during agricultural drought in grasses showed consensus. BMC Genom 12:319
Tang Y, Liu M, Gao S, Zhang Z, Zhao X, Zhao C, Zhang F, Chen X (2012) Molecular characterization of novel TaNAC genes in wheat and over-expression of TaNAC2 a confers drought tolerance in tobacco. Physiol Plant 144:210–224
Tran L-SP, Nakashima K, Sakuma Y, Simpson SD, Fujita Y, Maruyama K, Fujita M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2004) Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 16:2481–2498
Tran L-SP, Nakashima K, Sakuma Y, Osakabe Y, Qin F, Simpson SD, Maruyama K, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K (2007) Co-expression of the stress-inducible zinc finger homeodomain ZFHD1 and NAC transcription factors enhances expression of the ERD1 gene in Arabidopsis. Plant J 49:46–63
Tripathi P, Rabara RC, Langum TJ, Boken AK, Rushton DL, Boomsma DD, Rinerson CI, Rabara J, Reese RN, Chen X, Rohila JS, Rushton PJ (2012) The WRKY transcription factor family in Brachypodium distachyon. BMC Genom 13:270
Tripathi P, Rabara RC, Rushton PJ (2014) A systems biology perspective on the role of WRKY transcription factors in drought responses in plants. Planta 239:255–266
Tuberosa R (2012) Phenotyping for drought tolerance of crops in the genomics era. Front Physiol. doi:10.3389/fphys.2012.00347
Ulker B, Somssich IE (2004) WRKY transcription factors: from DNA binding towards biological function. Curr Opin Plant Biol 7:491–498
Umezawa T, Nakashima K, Miyakawa T, Kuromori T, Tanokura M, Shinozaki K, Yamaguchi-Shinozaki K (2010) Molecular basis of the core regulatory network in ABA responses: sensing, signaling and transport. Plant Cell Physiol 51:1821–1839
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidospsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high salinity conditions. Proc Natl Acad Sci USA 97:11632–11637
Vagujfalvi A, Aprile A, Miller A, Dubcovsky J, Delugu G, Galiba G, Cattivelli L (2005) The expression of several Cbf genes at the Fr-A2 locus is linked to frost resistance in wheat. Mol Genet Genom 274:506–514
Van Aken O, Zhang B, Law S, Narsai R, Whelan J (2013) AtWRKY40 and AtWRKY63 modulate the expression of stress-responsive nuclear genes encoding mitochondrial and chloroplast proteins. Plant Physiol 162:254–271
Vazquez F, Gasciolli V, Crete P, Vaucheret H (2004) The nuclear dsRNA binding protein HYL1 is required for microRNA accumulation and plant development, but not post transcriptional transgene silencing. Curr Biol 14:346–351
Vilela B, Moreno-Cortés A, Rabissi A, Leung J, Pagès M, Lumbreras V (2013) The maize OST1 kinase homolog phosphorylates and regulates the maize SNAC1-type transcription factor. PLoS ONE 8:e58105
Waltz E (2014) Beating the heat. Nat Biotech 32:610–613
Wang J, Lu M, Qiu C, Cui Q (2009) TransmiR: a transcription factor-microRNA regulation database. Nucl Acids Res 38:D119–D122
Wang Z, Libault M, Joshi T, Valliyodan B, Nguyen HT, Xu D, Stacey G, Cheng J (2010) SoyDB: a knowledge database of soybean transcription factors. BMC Plant Biol 10:14
Wang H, Maurano MT, Qu H, Varley KE, Gertz J, Pauli F, Lee K, Canfield T, Weaver M, Sandstrom R, Thurman RE, Kaul R, Myers RM, Stamatoyannopoulos JA (2012) Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res 22:1680–1688
Wang J, Zhuang J, Iyer S, Lin X, Whitfield TW, Greven MC, Pierce BG, Dong X, Kundaje A, Cheng Y, Rando OJ, Birney E, Myers RM, Noble WS, Snyder M, Weng Z (2012) Sequence features and chromatin structure around the genomic regions bound by 119 human transcription factors. Genome Res 22:1798–1812
Wang C, Deng P, Chen L, Wang X, Ma H, Hu W, Yao N, Feng Y, Chai R, Yang G, He G (2013) A wheat WRKY transcription factor TaWRKY10 confers tolerance to multiple abiotic stresses in transgenic tobacco. PLoS ONE 8:e65120
Wang X, Zeng J, Li Y, Rong X, Sun J, Sun T, Li M, Wang L, Feng Y, Chai R, Chen M, Chang J, Li K, Yang G, He G (2015) Expression of TaWRKY44, a wheat WRKY gene, in transgenic tobacco confers multiple abiotic stress tolerances. Front Plant Sci 6:615
Wang H, Honglei W, Shao H, Tang X (2016) Recent advances in utilizing transcription factors to improve plant abiotic stress tolerance by transgenic technology. Front Plant Sci 7:1–13
Wang J, Li Q, Mao X, Li A, Jing R (2016) Wheat transcription factor TaAREB3 participates in drought and freezing tolerances in Arabidopsis. Int J Biol Sci 12:257–269
Wei B, Jing R, Wang C, Chen J, Mao X, Chang X, Jia J (2009) Dreb1 genes in wheat (Triticum aestivum L.): development of functional markers and gene mapping based on SNPs. Mol Breed 23:13–22
Wei S, Hu W, Deng X, Zhang Y, Liu X, Zhao X, Luo Q, Jin Z, Li Y, Zhou S, Sun T, Wang L, Yang G, He G (2014) A rice calcium-dependent protein kinase OsCPK9 positively regulates drought stress tolerance and spikelet fertility. BMC Plant Biol 14:133
Wen Y, Li X, Guo C, Ma C, Duan W, Lu W, Xiao K (2015) Characterization and expression analysis of mitogen-activated protein kinase cascade genes in wheat subjected to phosphorus and nitrogen deprivation, high salinity, and drought. J Plant Biochem Biotechnol 24:184–196
Wessler SR (2005) Homing into the origin of the AP2 DNA binding domain. Trends Plant Sci 10:54–56
Wingender E (1988) Compilation of transcription regulating proteins. Nucl Acids Res 16:1879–1902
Wu H, Ni Z, Yao Y, Guo G, Sun Q (2008) Cloning and expression profiles of 15 genes encoding WRKY transcription factor in wheat (Triticum aestivem L.). Prog Nat Sci 18:697–705
Wu G, Park MY, Conway SR, Wang JW, Weigel D, Poethig RS (2009) The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 138:750–759
Xia N, Zhang G, Liu XY, Deng L, Cai GL, Zhang Y, Wang XJ, Zhao J, Huang LL, Kang ZS (2010a) Characterization of a novel wheat NAC transcription factor gene involved in defense response against stripe rust pathogen infection and abiotic stresses. Mol Biol Rep 37:3703–3712
Xia N, Zhang G, Sun F-Y, Zhu L, Xu L-S, Chen X-M, Liu B, Y-t Yu, Wang X-J, Huang L-L, Kang Z-S (2010b) TaNAC8, a novel NAC transcription factor gene in wheat, responds to stripe rust pathogen infection and abiotic stresses. Physiol Mol Plant Pathol 74:394–402
Xiao BZ, Chen X, Bin Xiang C, Tang N, Zhang QF, Xiong LZ (2009) Evaluation of seven function-known candidate genes for their effects on improving drought resistance of transgenic rice under field conditions. Mol Plant 2:73–83
Xu K, Xu X, Fukao T, Canlas P, Maghirang-Rodriguez R, Heuer S, Ismail AM, Bailey-Serres J, Ronald PC, Mackill DJ (2006) Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 442:705–708
Xu ZY, Kim SY, Hyeon do Y, Kim DH, Dong T, Park Y, Jin JB, Joo SH, Kim SK, Hong JC, Hwang D, Hwang I (2013) The Arabidopsis NAC transcription factor ANAC096 cooperates with bZIP-type transcription factors in dehydration and osmotic stress responses. Plant Cell 25:4708–4724
Xu D-B, Gao S-Q, Ma Y-Z, Xu Z-S, Zhao C-P, Tang Y-M, Li X-Y, Li L-C, Chen Y-F, Chen M (2014) ABI-like transcription factor gene TaABL1 from wheat improves multiple abiotic stress tolerances in transgenic plants. Funct Integr Genom 14:717–730
Xu H, Watanabe KA, Zhang L, Shen QJ (2016) WRKY transcription factor genes in wild rice Oryza nivara. DNA Res. doi:10.1093/dnares/dsw025
Xue G, Way H, Richardson T, Drenth J, Joyce P, McIntyre CL (2011) Overexpression of TaNAC69 leads to enhanced transcript levels of stress up-regulated genes and dehydration tolerance in bread wheat. Mol Plant 4:697–712
Yamaguchi-Shinozaki K, Shinozaki K (1994) A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 6:251–264
Yanez-Cuna JO, Dinh HQ, Kvon EZ, Shlyueva D, Stark A (2012) Uncovering cis-regulatory sequence requirements for context specific transcription factor binding. Genome Res 22:2018–2203
Yang S, Vanderbeld B, Wan J, Huang Y (2010) Narrowing down the targets: towards successful genetic engineering of drought-tolerant crops. Mol Plant 3:469–490
Yanhui C, Xiaoyuan Y, Kun H, Meihua L, Jigang L, Zhaofeng G, Zhiqiang L, Yunfei Z, Xiaoxiao W, Xiaoming Q, Yunping S, Li Z, Xiaohui D, Jingchu L, Xing-Wang D, Zhangliang C, Hongya G, Li-Jia Q (2006) The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60:107–124
Yates A, Akanni W, Amode MA, Barrell D, Billis K et al (2016) Ensembl 2016. Nucl Acids Res 44:D710–D716
Yilmaz A, Nishiyama Jr MY, Fuentes BG, Souza GM, Janies D, Gray J, Grotewold E (2009) GRASSIUS: a platform for comparative regulatory genomics across the grasses. Plant Physiol 149:171–180
Yoshida T, Mogami J, Yamaguchi-Shinozaki K (2014) ABA-dependent and ABA-independent signaling in response to osmotic stress in plants. Curr Opin Plant Biol 21:133–139
You J, Zong W, Hu H, Li X, Xiao J, Xiong L (2014) A STRESS-RESPONSIVE NAC1-regulated protein phosphatase gene rice protein phosphatase18 modulates drought and oxidative stress tolerance through abscisic acid-independent reactive oxygen species scavenging in rice. Plant Physiol 166:2100–2114
Zhang B (2015) MicroRNA: a new target for improving plant tolerance to abiotic stress. J Exp Bot 66:1749–1761
Zhang B, Wang Q (2015) MicroRNA-based biotechnology for plant improvement. J Cell Physiol 230:1–15
Zhang B, Pan X, Cobb GP, Anderson TA (2006) Plant microRNA: a small regulatory molecule with big impact. Dev Biol 289:3–16
Zhang Y, Zhang G, Xia N, Wang XJ, Huang LL, Kang ZS (2009) Cloning and characterization of a bZIP transcription factor gene in wheat and its expression in response to stripe rust pathogen infection and abiotic stresses. Physiol Mol Plant Pathol 73:88–94
Zhang H, Jin JP, Tang L, Zhao Y, Gu XC, Gao G, Luo JC (2011) PlantTFDB 2.0: update and improvement of the comprehensive plant transcription factor database. Nucl Acids Res 39:D1114–D1117
Zhang L, Zhao G, Xia C, Jia J, Liu X, Kong X (2012) A wheat R2R3-MYB gene, TaMYB30-B, improves drought stress tolerance in transgenic Arabidopsis. J Exp Bot 63:5873–5885
Zhang Z, Liu X, Wang X, Zhou M, Zhou X, Ye X, Wei X (2012) An R2R3 MYB transcription factor in wheat, TaPIMP1, mediates host resistance to Bipolaris sorokiniana and drought stresses through regulation of defense- and stress-related genes. New Phytol 196:1155–1170
Zhang L, Liu G, Zhao G, Xia C, Jia J, Liu X, Kong X (2014) Characterization of a wheat R2R3-MYB transcription factor gene, TaMYB19, involved in enhanced abiotic stresses in Arabidopsis. Plant Cell Physiol 55:1802–1812
Zhang L, Zhang L, Xia C, Zhao G, Liu J, Jia J, Kong X (2015) A novel wheat bZIP transcription factor, TabZIP60, confers multiple abiotic stress tolerances in transgenic Arabidopsis. Physiol Plant 153:538–554
Zhang LN, Zhang L, Xia C, Zhao G, Jia J, Kong X (2016) The novel wheat transcription factor TaNAC47 enhances multiple abiotic stress tolerances in transgenic plants. Front Plant Sci. doi:10.3389/fpls.2015.01174
Zhao B, Ge L, Liang R, Li W, Ruan K, Lin H, Jin Y (2009) Members of miR-169 family are induced by high salinity and transiently inhibit the NF-YA transcription factor. BMC Mol Biol 10:29
Zheng X, Liu H, Ji H, Wang Y, Dong B, Qiao Y, Liu M, Li X (2016) The wheat GT factor TaGT2L1D negatively regulates drought tolerance and plant development. Sci Rep 6:27042
Zhou L, Liu Y, Liu Z, Kong D, Duan M, Luo L (2010) Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J Exp Bot 61:4157–4168
Zhu QH, Guo AY, Gao G, Zhong YF, Xu M, Huang M, Luo J (2007) DPTF: a database of poplar transcription factors. Bioinformatics 23:1307–1308
Zhu S-Y, Yu X-C, Wang X-J, Zhao R, Li Y, Fan R-C, Shang Y, Du S-Y, Wang X-F, Wu F-Q, Xu Y-H, Zhang X-Y, Zhang D-P (2007) Two calcium-dependent protein kinases, CPK4 and CPK11, regulate abscisic acid signal transduction in Arabidopsis. Plant Cell 19:3019–3036
Zhu Q, Zhang J, Gao X, Tong J, Xiao L, Li W, Zhang H (2010) The Arabidopsis AP2/ERF transcription factor RAP2.6 participates in ABA, salt and osmotic stress responses. Gene 457:1–12
Zhu X, Liu S, Meng M, Qin L, Kong L, Xia G (2013) WRKY transcription factors in wheat and their induction by biotic and abiotic stress. Plant Mol Biol Rep 31:1053–1067
Zhu Y, Yan J, Liu W, Liu L, Sheng Y, Sun Y, Li Y, Scheller HV, Jiang M, Hou X, Ni L, Zhang A (2016) Phosphorylation of a NAC transcription factor by ZmCCaMK regulates abscisic acid-induced antioxidant defense in maize. Plant Physiol 171:1651–1664
Zong W, Tang N, Yang J, Peng L, Ma S, Xu Y, Li G, Xiong L (2016) Feedback regulation of ABA signaling and biosynthesis by a bZIP transcription factor targets drought resistance related genes. Plant Physiol p 00469. doi:10.1104/pp.16.00469
Acknowledgments
The authors like to thank The Head, Department of Genetics and Plant Breeding, CCS University (Meerut, India) for providing facilities. PKG was awarded a National Academy of Sciences India (NASI) Senior Scientist Platinum Jubilee Fellowship, and INSA Senior Scientist positions during the tenure of which this review was written; VG was awarded a Junior Research Fellowship under the same program, and was later awarded the position of SRF/RA under a DBT project. VJ is awarded with CSIR-Nehru Science Post Doc Fellowship. AK awarded a JRF under the scheme of DBT-BTISnet program and later awarded SRF in same program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by R. K. Varshney.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gahlaut, V., Jaiswal, V., Kumar, A. et al. Transcription factors involved in drought tolerance and their possible role in developing drought tolerant cultivars with emphasis on wheat (Triticum aestivum L.). Theor Appl Genet 129, 2019–2042 (2016). https://doi.org/10.1007/s00122-016-2794-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-016-2794-z