Abstract
Powdery mildew is an important foliar disease in wheat, especially in areas with a cool or maritime climate. A dominant powdery mildew resistance gene transferred to the hexaploid germplasm line NC99BGTAG11 from T. timopheevii subsp. armeniacum was mapped distally on the long arm of chromosome 7A. Differential reactions were observed between the resistance gene in NC99BGTAG11 and the alleles of the Pm1 locus that is also located on chromosome arm 7AL. Observed segregation in F2:3 lines from the cross NC99BGTAG11 × Axminster (Pm1a) demonstrate that germplasm line NC99BGTAG11 carries a novel powdery mildew resistance gene, which is now designated as Pm37. This new gene is highly effective against all powdery mildew isolates tested so far. Analyses of the population with molecular markers indicate that Pm37 is located 16 cM proximal to the Pm1 complex. Simple sequence repeat (SSR) markers Xgwm332 and Xwmc790 were located 0.5 cM proximal and distal, respectively, to Pm37. In order to identify new markers in the region, wheat expressed sequence tags (ESTs) located in the distal 10% of 7AL that were orthologous to sequences from chromosome 6 of rice were targeted. The two new EST-derived STS markers were located distal to Pm37 and one marker was closely linked to the Pm1a region. These new markers can be used in marker-assisted selection schemes to develop wheat cultivars with pyramids of powdery mildew resistance genes, including combinations of Pm37 in coupling linkage with alleles of the Pm1 locus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Powdery mildew, caused by Blumeria graminis (DC) Speer f. sp. tritici emend Marchal (Bgt), is an important foliar disease in wheat (Triticum aestivum L.), especially in areas with a cool or maritime climate. Powdery mildew impacts grain yield, significantly reduces flour yield and adversely affects other aspects of grain quality (Everts et al. 2001; Hsam and Zeller 2002). The use of resistant cultivars is an effective, economical, and environmentally safe approach that eliminates the use of fungicides and reduces production losses due to this disease.
The most common breeding strategy for resistance to powdery mildew in wheat has been the use of qualitative (race-specific) resistance conferring hypersensitive foliar reactions (Chen et al. 2005; Huang and Röder 2004). This type of resistance follows the gene-for-gene hypothesis described by Flor (1955), and because of the co-evolution of host and pathogen, race-specific resistance can be overcome by new races of the pathogen possessing corresponding virulence genes. Leath and Murphy (1985) found that the 10 most widely used resistance genes had matching virulence genes in the Southeastern USA. In Europe, 10 powdery mildew resistance genes in wheat cultivars showed corresponding virulence genes in the pathogen (Clarkson 2000). Therefore, the search for and deployment of new powdery mildew resistance genes is necessary to provide wheat growers with resistant cultivars. Currently, 49 powdery mildew resistance genes mapped at 33 loci have been identified in wheat and its wild relatives (McIntosh et al. 2003, 2004, 2005).
The cultivated Triticum timopheevii subsp. timopheevii (Zhuk.) Zhuk. and its wild form T. timopheevii subsp. armeniacum (Jakubz.) van Slageran (AtAtGG, 2n = 28) have been used as sources of pest resistance genes for wheat, including genes for resistance to powdery mildew. Resistance genes Pm6 and Pm27 were transferred to wheat chromosomes 2B and 6B from cultivated timopheevii wheat (Jarve et al. 2000; Jørgensen 1973). A powdery mildew resistance gene was transferred to the long arm of chromosome 7A (7AL) from the wild T. timopheevii subsp. armeniacum to germplasm line NC99BGTAG11 (Srnić et al. 2005). The Pm1 locus is also located on chromosome 7AL and it possesses five dominant alleles, Pm1a to Pm1e. In addition, recessive genes mlRD30 and Pm9 have been reported to be linked to the Pm1 locus (Schneider et al. 1991; Singrün et al. 2004). None of the alleles of the Pm1 locus originate from tetraploid wheat relatives. The gene Pm1b, and two genes that are either new allelic variants of Pm1 or closely linked to the Pm1 locus (Yao et al. 2007) were introgressed from the diploid einkorn Triticum monococcum L. (AmAm, 2n = 14), and Pm1d was introgressed from Triticum aestivum L. subsp. spelta (Hsam et al. 1998). Neu et al. (2002) suggested that Pm1a might have been derived from a wheat relative.
Molecular markers have been used for tagging and mapping powdery mildew resistance genes in wheat, including those on chromosome 7A. Srnić et al. (2005) reported simple sequence repeat (SSR) markers Xgwm332 and Xwmc525 as flanking the resistance gene in NC99BGTAG11 with genetic distances of 2.0 centimorgans (cM) proximal (towards the centromere) and 1.4 cM distal (towards the telomere), respectively. The RFLP markers Xpsr148 and Xpsr680 were reported to co-segregate with the powdery mildew resistance gene Pm1 (Ma et al. 1994; Neu et al. 2002). The Xpsr148 and Xpsr680 markers were converted to STS markers named Xmag1714 and Xmag2185, respectively (Yao et al. 2007).
The increasing availability of expressed sequence tag (EST) and genomic sequences from wheat is providing a potentially valuable source for marker enrichment. Currently, there are more than 580,000 wheat ESTs with 122,282 unique sequences (http://www.tigr.org/tigr-scripts/tgi/T_index.cgi?species=wheat) deposited in public databases. This provides an excellent resource for mapping genes. A set of wheat deletion lines has been used to locate more than 8,300 unique ESTs into chromosome bin maps (Hossain et al. 2004; Lazo et al. 2004; Qi et al. 2004). The existing synteny between rice and wheat, as well as other cereals, can be exploited to tentatively position ESTs in silico based on orthology with sequences in the rice genome. Hossain et al. (2004) mapped 2148 EST loci to the three homeologous group 7 chromosomes of wheat, showed the distribution of mapped EST loci to the chromosome bins defined by the deletion stocks, and identified putative regions of conserved gene content between the wheat group 7 consensus chromosome and rice chromosome 6 and 8. In addition, several STS markers derived from wheat ESTs homologous to the coding DNA sequences of rice chromosome 6L were linked to two powdery mildew genes mapped on the distal region of 7AL (Yao et al. 2007).
The present study reports the genetic relationship between the resistance gene in NC99BGTAG11, now designated Pm37, and the Pm1 locus, and the identification of additional molecular markers linked to these resistance genes.
Materials and methods
Plant material
Soft red winter wheat germplasm line NC99BGTAG11 (NCAG11 hereafter) (Reg. no. GP-729, PI 615588) is an F7-derived line with the pedigree ‘Saluda’*3/PI 427315 (Murphy et al. 2002). Saluda (PI 480474) is a soft red winter wheat developed and released by Virginia Polytechnic Institute and State University (Starling et al. 1986) and it possesses the powdery mildew resistance gene Pm3a. The Pm3a gene is not effective against naturally occurring powdery mildew populations in North Carolina (Leath and Heun 1990). PI 427315 is a winter growth habit accession of T. timopheevii subsp. armeniacum collected in Iraq. ‘Axminster’ (PI 228307) is a T. aestivum cultivar that possesses the powdery mildew resistance gene Pm1a. A population of 198 F2:3 lines was developed from the cross NCAG11/Axminster to evaluate allelism of the T. timopheevii-derived resistance gene with Pm1a.
The Wheat Genetics Resource Center at Kansas State University supplied the Chinese Spring (CS) wheat and chromosome 7A-related CS aneuploids used in the study. These included: nullisomic 7A-tetrasomic 7B (N7A-T7B), nullisomic 7A-tetrasomic 7D (N7A-T7D), nullisomic 7B-tetrasomic 7A (N7B-T7A), nullisomic 7D-tetrasomic 7A (N7D-T7A), and nullisomic 7D-tetrasomic 7B (N7D-T7B) (Sears 1966); ditelosomic lines 7AS (Dt7AS) and 7AL (Dt7AL) (Sears and Sears 1979); and four CS deletion lines for the terminal region of the long arm of chromosome 7A. The deletion lines are designated by the chromosome arm carrying the deletion and the length of the terminal deletion, expressed as a fraction length (FL) of the whole arm. Deletion lines included were 7AL-16 (FL = 0.86), 7AL-2 (FL = 0.87), 7AL-20 (FL = 0.89), and 7AL-15 (FL = 0.99) (Endo and Gill 1996; Qi et al. 2003). Together, the nullisomic–tetrasomic lines and the deletion lines were used for the chromosome and deletion bin mapping of EST-based STS and SSR markers linked to the powdery mildew resistance gene in NCAG11 (Pm37).
The soft winter (SW) wheat cultivars and breeding lines Ernie, Neuse, McCormick, Roane, Truman, Pioneer brand 2545, VA99W-200, Pioneer brand 2555, Superior, Patterson, Freedom, Patton, GA881130, and Batavia were analyzed with the SSR marker Xgwm332 to determine the level of polymorphism for this marker in elite SRW germplasm.
Powdery mildew evaluations
Powdery mildew resistance was evaluated in the laboratory using the detached leaf technique. Primary leaf segments (1.5-cm) were floated on 0.5% water agar (w/v) amended with 50 mg l−1 benzimidazole in plastic plates. Lines possessing Pm1a, Pm1b, Pm1c, Pm1d, and Pm1e, NCAG11, Saluda, and the control Chancellor were inoculated with 14 different powdery mildew isolates collected from wheat fields in the eastern region of North Carolina. Plates were placed in a growth chamber maintained at 18°C, 85% relative humidity and with a photoperiod of 12 h. The disease severity evaluation was based on a scale from 0 to 9, in which 0–3 = resistant with (0) no visible signs of infection; (1) flecks with no necrosis, (2) necrosis, to (3) chlorosis; 4–6 = intermediate reaction with chlorotic areas decreasing in amount while mycelium and conidia production increased from slight to moderate; and 7–9 = susceptible with increasing amount, size and density of mycelium and conidia to a fully compatible reaction.
Twenty seeds each of 198 F2:3 families from NCAG11/Axminster were grown at 18°C with a photoperiod of 12 h in a growth chamber and inoculated with the Bgt ‘Yuma’ isolate that was avirulent to both NCAG11 and Axminster, but virulent to Saluda (Pm3a). All plants were inoculated when they showed three fully developed leaves about 10–12 days after emergence. The resistant parents (NCAG11 and Axminster) were included as resistant controls and Saluda as the susceptible control. Evaluations were made when the susceptible controls showed distinct disease symptoms and the resistant parents showed no signs of disease. Susceptible and selected segregating families were re-screened with another avirulent Bgt isolate, Arapahoe, to confirm the reactions. In these tests, the reaction of Pm1a (infection type, IT = 0) could not be distinguished from that of Pm37 (IT = 0). The goodness-of-fit to the segregation ratio of 7 resistant:8 segregating:1 susceptible was tested using the χ2 test.
An additional 20 seeds of each of 198 F2:3 families from NCAG11/Axminster was grown at 18–20°C in a greenhouse where natural light was supplemented with artificial high intensity 1,000 W discharge lights to provide 12 h day:12 h night. Plants were inoculated with the Bgt ‘E314’ isolate that is virulent to Pm1a. Thirty plants of each of the parents and Saluda were included in the evaluation. All plants were inoculated at the three- to four-leaf stage about 15 days after emergence. The resistant parent (NCAG11) was considered as the resistant control and Axminster and Saluda were the susceptible controls. Evaluations were based on a 0–9 scale as described above, and data were obtained 15–20 days after inoculation when the susceptible controls showed distinct disease symptoms and the resistant parent showed no signs of disease.
Molecular marker analyses
Leaf tissue was harvested in bulk from 20 young plants each of the 198 F2:3 lines from the NCAG11/Axminster population, the parents, CS wheat, and the CS aneuploids and deletion lines, and stored at −80°C. Frozen leaf tissue was ground in a GenoGrinder (Spex, Metuchen, NJ) and genomic DNA was extracted using a DNeasy Plant Mini Kit (Qiagen, Valencia, CA) following the manufacturer’s instructions. Genomic DNA was amplified with SSR markers and EST-STS markers. Polymerase chain reaction (PCR) amplifications were performed in 12-μl reactions with 1.2 μl of 10× PCR buffer (containing 1.5 mM magnesium chloride), 0.97 μl of dNTPs (2.5 mM each dNTP), 0.5 μl of each forward and reverse primers (10 pmol/μl) and 40–60 ng of DNA in an Eppendorf Mastercycler® Gradient (Brinkmann Instruments, Inc., NY, USA). Forward SSR primers were 5′ labeled with fluorescent dyes (6-FAM and VIC). After initial denaturation at 95°C for 3 min, 40 amplification cycles were performed with 94°C for 45 s, 54–62°C (marker dependent) for 45 s and 72°C for 1 min, and a final extension at 72°C for 10 min. Sizing of the SSR fragments was resolved in an ABI3130 DNA analyzer (Applied Biosystems, Foster, CA) following manufacture’s instructions, and results were analyzed in GeneMarker v1.5 (SoftGenetics LLC, State College, PA). PCR products of EST-STS markers were resolved in 2.3% high resolution agarose (Gene Pure HiRes Agarose, ISC BioExpress) gels with 0.5× TBE buffer and visualized by ethidium bromide staining. After the fragment size of the PCR products from EST-STS markers were verified in agarose gels, the PCR products were denatured and separated on 380 × 500 × 0.4 mm single stranded conformation polymorphism (SSCP) gels using a mutation detection enhancement (MDE) gel solution (Martins-Lopes et al. 2001). The gel mix was made in an 80-ml total volume containing a final concentration of 0.5× gel solution (Cambrex Bio-science Rockland, Rockland, ME) and 0.6× TBE buffer, and polymerized by the addition of 0.16 ml of 20% ammonium persulphate and 24 μl of tetramethylethylenediamine (TEMED). Fragments were electrophoresed for 16 h at a constant power of 4 W at room temperature; and then, silver stained as described by Bassam et al. (1991). The STS marker XstsBE406627 was subsequently assayed using a fluorescent labeled primer and resolved in the ABI3130 sequencer. Genomic DNA of 14 SRW wheat lines was amplified with SSR marker Xgwm332 and electrophoresis was conducted at 110 W for 2.0 h on a 6% denaturing polyacrylamide gel (19 acrylamide:1Bis), 8 M urea and 1× TBE. Amplified fragments were visualized by silver-staining.
SSR markers of the terminal region of 7AL were evaluated for polymorphism between NCAG11 and Axminster. The physical locations of two wheat SSR markers (Xgwm332 and Xwmc525) that were previously reported to flank the powdery mildew resistance gene in NCAG11 (Srnić et al. 2005) and other SSR markers previously mapped in the terminal region of 7AL, including Xcfa2257, Xcfa2293, Xwmc790, Xgwm63, Xwmc633, Xcfa2019, Xgwm554, Xgwm346, Xwmc273, Xgwm344, and Xcfa2040, were determined by evaluating the markers on the CS aneuploid and deletion line stocks. In addition, two STS markers, Xmag2185 and Xmag1714, developed from RFLP markers (Xpsr680 and Xpsr148, respectively) and one STS marker, Xmag1759 developed from a wheat EST (Yao et al. 2007) were evaluated. The SSR and STS data indicated the physical location of the resistance gene in NCAG11 and were used for subsequent identification of wheat ESTs likely linked to the resistance gene. The terminal region of chromosome 7AL of wheat is syntenic to the distal region of chromosome 6L of rice (La Rota and Sorrells 2004) and was targeted for marker enrichment. Fifty-seven unique wheat EST sequences that mapped distal to the 7AL-18 (FL = 0.90) deletion breakpoint (http://wheat.pw.usda.gov/cgi-bin/westsql/map_locus.cgi) were used to search the rice genome database (http://tigrblast.tigr.org/euk-blast/index.cgi?project=osa1) using the BLASTn program (Altschul et al. 1997). Sequences in the target region of the rice genome were also used as queries in BLASTn searches of the wheat EST database (http://tigrblast.tigr.org/tgi/) to identify additional unmapped wheat ESTs that were potentially linked to Pm37. Wheat EST sequences with high levels of identity (E values < e−15) to sequences from chromosome 6L of rice were used to design primers for EST-derived STS markers. Primer design was conducted with the software Primer3 (Rozen and Skaletsky 2000), and amplicons of 200–500 base pairs (bp) were targeted. All markers were mapped physically using the CS deletion lines. Polymorphic markers that physically mapped in the target region were evaluated on 198 lines of the NCAG11/Axminster F2:3 mapping population. Primer sequences for the two polymorphic STS markers mapped in this study are listed in Table 1.
Chi-squared (χ2) tests were used to test for deviations of observed data from theoretically expected segregation ratios in the allelism tests and in the mapping population. Genetic maps were constructed using the program JoinMap®4 (van Ooijen 2006) and recombination frequencies were converted to cM using the Kosambi mapping function (Kosambi 1944) to estimate genetic distances with a minimum LOD of 3.0.
Results
Differential powdery mildew responses
Differential reactions were observed on lines possessing Pm1a, Pm1b, Pm1c, Pm1d, and Pm1e, NCAG11, Saluda, and Chancellor inoculated with 14 Bgt isolates (Table 2). All the Pm1 alleles could be differentiated from each other and from the gene in NCAG11, indicating that the powdery mildew resistance in NCAG11 was different from the five designated Pm1 alleles. NCAG11 was highly resistant to all isolates. The Chancellor control was fully susceptible to all isolates. The cultivar Saluda used as the recurrent parent in the development of NCAG11 has the Pm3a resistance gene and gave intermediate or resistant reactions with five isolates. MocZlatka (Pm1b) was fully susceptible to only two Bgt isolates and intermediate to two others. In contrast, lines possessing other Pm1 alleles were fully susceptible to four or more Bgt isolates.
Allelism test
Eight of the Bgt isolates were avirulent to the genes in Axminster and NCAG11 and virulent to Pm3a that is present in Saluda (Table 2). The Bgt ‘Yuma’ and Bgt ‘Arapahoe’ isolates were selected to evaluate the segregating population from the cross NCAG11/Axminster to determine if the T. timopheevii-derived gene in NCAG11 was an allele of the Pm1 locus. All of the 198 F2:3 lines showed clear reactions when inoculated with the Bgt ‘Yuma’ isolate. The reaction of susceptible and some segregating families were confirmed in a separate test using the Bgt ‘Arapahoe’ isolate. Three families were fully susceptible when inoculated with both isolates, indicating that a new powdery mildew locus is present in NCAG11 that is not an allele of Pm1. However, the observed ratio of 153 resistant lines:42 segregating lines:3 susceptible lines significantly differed from the 7 resistant:8 segregating:1 susceptible ratio (χ² = 90.78, df = 2, P < 0.0001) expected for two independently segregating dominant genes. Thus, the new resistance gene in NCAG11 is linked to the Pm1 locus. The T. timopheevii-derived resistance gene in accession NCAG11 is designated Pm37.
Mapping the Pm37 powdery mildew resistance gene
Five Bgt isolates were virulent to Pm1a, but only Bgt ‘E314’ was virulent on Saluda as well (Table 2); and therefore, was selected to evaluate the F2:3 lines from the cross NCAG11/Axminster in order to map Pm37. The observed ratio of 44 resistant:101 segregating:48 susceptible did not significantly differ from the 1:2:1 ratio (χ² = 1.94, df = 2, P = 0.38) expected for a single dominant gene for powdery mildew resistance, indicating that resistance to the Bgt ‘E314’ isolate was conferred by a single resistance gene.
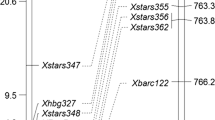
Seven SSR markers Xgwm332, Xwmc790, Xcfa2019, Xwmc346, Xwmc525, Xwmc273, and Xcfa2040 were polymorphic between Axminster, NCAG11 and the recurrent parent Saluda (Table 3). Several other SSR markers that mapped on the distal region of 7AL, such as Xcfa2257, Xcfa2293, Xgwm63, Xwmc633, and Xgwm344, were not polymorphic between NCAG11 and Axminster, but did show polymorphism between NCAG11 and Saluda (data not shown), indicating that the distal portion of 7AL in NCAG11 was derived from the donor parent. In this study, the powdery mildew resistance gene Pm37 was flanked by SSR loci Xgwm332 and Xwmc790 with genetic distances of 0.5 cM proximal and distal, respectively (Fig. 1). This indicates that there were two recombination events in the population of 198 F2 families, one on each side of the gene. Both SSR markers Xgwm332 and Xwmc790 were co-dominant. The 193-bp allele amplified in NCAG11 by the SSR marker Xgwm332 was not present in the 14 soft red winter cultivars and breeding lines tested (Fig. 2). The STS markers Xmag2185 and Xmag1714 were developed from RFLP markers Xpsr680 and Xpsr148, respectively (Yao et al. 2007), and reported to co-segregate with Pm1a (Neu et al. 2002). The STS markers mapped more than 16 cM distal to Pm37 (Fig. 1). The STS markers were dominant and linked in repulsion to the Pm1a allele.
A denaturing polyacrylamide gel pattern of the deletion mapping and screening of 14 soft red wheat cultivars (SRW) of the locus Xgwm332. Amplified fragments are observed in CS, Dt7AL, N7B-T7D, N7D-T7B, and deletion line 7AL-15, but not in Dt7AS, N7A-T7D, and deletion lines 7AL-16, 7AL-20, and 7AL-9. Polymorphism of the A-genome fragment was observed between NCAG11 and Axminster. The 193-bp allele in NCAG11 was not present in the 14 SRW wheat lines surveyed. Size standard HyperLadder IV (Bioline, Randolph, MA), with a 200-bp marker fragment indicated as Marker
Physical mapping and marker enrichment
The SSR markers Xgwm332, Xwmc790, Xwmc525, and Xcfa2040 were located on chromosome arm 7AL in other maps of T. aestivum (Röder et al. 1998, Somers et al. 2004; Sourdille et al. 2004). In our study, the order of these SSR loci agreed well with the established SSR maps of chromosome arm 7AL. Our analysis of the Chinese Spring aneuploid stocks and deletion lines confirmed the presence of these markers on 7AL and their location in the terminal 11% of the chromosome arm between the deletion breakpoints 7AL-20 (FL = 0.89) and 7AL-15 (FL = 0.99).
Wheat ESTs located in the terminal region of 7AL were targeted in an effort to obtain additional markers linked to Pm37. Out of 57 wheat ESTs previously mapped distal to the deletion breakpoint 7AL-18 (FL = 0.90) (Hossain et al. 2004), 22 had a significant homology to sequences on the terminal region of rice chromosome 6L (from 24.90 to 31.10 Mb), 23 had no obvious orthologous sequences in rice, and the remaining 12 had significant hits elsewhere in the rice genome. Twenty-two wheat ESTs, whose only significant orthologous rice sequences were in the terminal region of chromosome 6 were selected for primer design.
Six of the selected EST-based primer pairs amplified fragments that were located in the distal 10% of chromosome 7AL, consistent with the previous assignment of the ESTs based on RFLP analysis (Hossain et al. 2004). The other 16 EST-based primer pairs failed to amplify fragments that could be deletion mapped using SSCP analysis. Of the six markers that were deletion mapped, two STS markers, XstsBE406627 and XstsBE445653 (Table 1), were polymorphic between NCAG11 and Axminster and were evaluated on the mapping population. Marker XstsBE406627 co-segregated with markers Xmag2185 and Xmag1714 and was located 16.2 cM distal to Pm37 (Fig. 1). Marker XstsBE445653 was located 20.4 cM, distal to Pm37.
Discussion
Our differential tests, combined with allelism and molecular marker analyses, indicated that a new dominant powdery mildew resistance gene designated Pm37 was transferred from T. timopheevii subsp. armeniacum to germplasm line NCAG11. Pm37 is proximal to the Pm1 locus. This new gene is highly effective in the field in southeastern U.S. and confers resistance to more than 60 different isolates of Blumeria graminis f. sp. tritici (data not shown).
The Pm37 gene is the first powdery mildew resistance gene transferred from T. timopheevii to the A genome of common wheat. Two other powdery mildew resistance genes, Pm6 and Pm27, were transferred from cultivated T. timopheevii, but these are located on chromosomes 2BL and 6B, respectively (Jørgensen 1973; Jarve et al. 2000).
Several powdery mildew resistance genes have been mapped on chromosome 7A of wheat. Among these, Pm1, Pm9, mlRD30, and two dominant powdery mildew resistance genes introgressed from T. monoccocum (Schneider et al. 1991; Sears and Briggle 1969; Singrün et al. 2004; Yao et al. 2007) are located on the long arm. Our allelism data indicates that Pm37 is not an allele at the Pm1 locus. The map location of the resistance genes introgressed from T. monoccocum by Yao et al. (2007) suggests that they are alleles at the Pm1 locus. The resistance genes Pm9 and mlRD30 were reported to be recessive. The cultivar Normandie carries the recessive powdery mildew resistance gene Pm9, which is 8.5 cM, distal to Pm1a (Schneider et al. 1991; Singrün 2002). The recessive powdery mildew resistance gene mlRD30 was located 1.8 cM distal to SSR marker Xgwm344 (Singrün et al. 2004). We were not able to map Xgwm344 as it was not polymorphic between NCAG11 and Axminster. However, marker Xgwm344 was located 16 cM distal to Xgwm332 (Singrün et al. 2003), and was reported to co-segregate or to map distal to Xmag2185 in four different populations (Yao et al. 2007). In our study, the STS marker Xmag2185 was 16.2 cM distal to Pm37.
In our allelism tests, the reaction of Pm1a could not be distinguished from that of Pm37 using isolates Bgt ‘Yuma’ and Bgt ‘Arapahoe’. The number of lines segregating for resistance was less than expected if Pm37 and Pm1a are 16 cM apart. Given that the population size for each F2:3 line was only 20 plants and that the resistance genes are linked, we likely have overestimated the number of homozygous resistant lines. This hindered our ability to estimate linkage between Pm37 and Pm1a based on the phenotypic data. However, the identification of three homozygous susceptible F2:3 lines in our population did not differ significantly from the 1.26 expected given a genetic distance of 16 cM between Pm37 and the Pm1 locus (χ² = 2.4, df = 1, P < 0.12).
The powdery mildew resistance allele Pm1a was reported to co-segregate with RFLP marker Xcdo347 (Ma et al. 1994; Neu et al. 2002). The Xcdo347 marker also co-segregated with RFLP marker, Xpsr680, which was converted to STS marker Xmag2185 (Yao et al. 2007) that was mapped in this study. These RFLP and STS markers were mapped in different populations at 32.8 cM and more than 30 cM from Xgwm332 and Xcfa2019, respectively (Neu et al. 2002; Yao et al. 2007). Comparison of these marker analyses with our analysis suggests that Pm37 is proximal to the other powdery mildew resistance loci on the long arm of chromosome 7A, Pm1, Pm9 and mlRD30.
Srnić et al. (2005) reported Xgwm332 and Xwmc525 as flanking the resistance gene in NCAG11 with genetic distances of 2.0 cM proximal and 1.4 cM distal, respectively. Although our map positions agree, slightly less recombination was observed between Pm37 and Xgwm332 while more recombination was observed between Pm37 and Xwmc525 in our population. This may be due to the origin of the chromosome segments in the NCAG11/Axminister cross since it has been suggested that Pm1a was derived from an alien introgression (Neu et al. 2002). The distal region of 7AL in NCAG11 was transferred from T. timopheevii subsp. armeniacum. Neu et al. (2002) reported that complete linkage of Xcdo347, Xc607, Xpsr121, Xpsr148, and Xpsr680 with Pm1a was caused by suppressed recombination in hexaploid wheat rather than physical linkage. Although we did not locate any STS marker derived from wheat ESTs in the Pm37 region, we identified a new marker in the Pm1 region that originated from a wheat EST with homology to the syntenic region in rice that might be useful for fine mapping the Pm1 locus.
Introgression of disease resistance genes from related species into wheat has become crucial in developing resistant genotypes. The resistance gene Pm37 introgressed from T. timopheevii subsp. armeniacum has provided full resistance to all powdery mildew isolates tested in this study, and so far, no virulence to Pm37 has been found. It is now also possible to combine several Pm resistance genes into a single improved wheat genotype for more durable powdery mildew resistance. Co-dominant SSR markers Xgwm332 (proximal) and Xwmc790 (distal) are closely linked to Pm37 and could be used in marker-assisted selection to develop powdery mildew-resistant lines having this gene in combination with other resistance genes. Using the markers linked to Pm37 and Pm1a, we identified recombinant plants having the genes in coupling; such genotypes can be used to develop cultivars with additional resistance genes.
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSIBLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bassam BJ, Caetano-Anolles G, Gresshoff PM (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196:80–83
Briggle LW (1969) Near-isogenic lines of wheat with genes for resistance to Erysiphe graminis tritici. Crop Sci 9:70–72
Chen XM, Luo YH, Xia XC, Xia LQ, Chen X et al (2005) Chromosomal location of powdery mildew resistance gene Pm16 in wheat using SSR marker analysis. Plant Breed 124:225–228
Clarkson JDS (2000) Virulence survey report for wheat powdery mildew in Europe, 1996–1998. http://www.crpmb.org/2000/1204clarkson
Endo TR, Gill BS (1996) The deletion stocks of common wheat. J Hered 87:295–307
Everts KL, Leath S, Finney PL (2001) Impact of powdery mildew on milling and baking quality of soft red winter wheat. Plant Dis 85(4):423–429
Flor HH (1955) Host–parasite interaction in flax rust – its genetics and other implications. Phytopathology 45:680–685
Hossain KG, Kalavacharla V, Lazo GR, Hegstad J, Wentz MJ et al (2004) A chromosome bin map of 2148 expressed sequence tag loci of wheat homeologous group 7. Genetics 168:687–699
Hsam SLK, Zeller FJ (2002) Breeding for powdery mildew resistance in common wheat (Triticum aestivum L.). In: Berlanger RR, Bushnell WR, Dik AJ, Carver DL (eds) The powdery mildews: a comprehensive treatise. Am. Phytopath. Soc., St. Paul, MN, pp 219–238
Hsam SLK, Huang XQ, Earnst F, Hartl L, Zeller FJ (1998) Chromosomal location of genes for resistance to powdery mildew in common wheat (Triticum aestivum L.) 5. Alleles at the Pm1 locus. Theor Appl Genet 96:1129–1134
Huang XQ, Röder MS (2004) Molecular mapping of powdery mildew resistance genes in wheat: a review. Euphytica 137:203–223
Jarve K, Peusha HO, Tsymbalova J, Tamm S, Devos KM, Enno TM (2000) Chromosomal location of a Triticum timopheevii-derived powdery mildew resistance gene transferred to common wheat. Genome 43:377–381
Jørgensen JH (1973) Gene Pm6 for resistance to powdery mildew in wheat. Euphytica 22:43
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
La Rota M, Sorrells ME (2004) Comparative DNA sequence analysis of mapped wheat ESTs reveals the complexity of genome relationships between wheat and rice. Funct Integr Genomics 4:34–46
Lazo GR, Chao S, Hummel DD, Edwards H, Crossman CC, Lui N et al (2004) Development of an expressed sequence tag (EST) resource for wheat (Triticum aestivum L.): EST generation, unigene analysis, probe selection, and bioinformatics for a 16,000-locus bin-delineated map. Genetics 168:585–593
Leath S, Heun M (1990) Identification of powdery mildew resistance genes in cultivars of soft red winter wheat. Plant Dis 74:747–752
Leath S, Murphy JP (1985) Virulence genes of the wheat powdery mildew fungus, Erysiphe graminis f. sp. tritici, in North Carolina. Plant Dis 69:905
Ma ZQ, Sorrells ME, Tanksley SD (1994) RFLP markers linked to powdery mildew resistance genes Pm1, Pm2, Pm3, and Pm4 in wheat. Genome 37:871–875
Martins-Lopes P, Zhang H, Koebner R (2001) Detection of single nucleotide mutations in wheat using single strand conformation polymorphism gels. Plant Mol Biol Rep 19:159–162
McIntosh RA, Yamazaki Y, Devos KM, Dubcovsky J, Rogers WJ, Appels R (2003) Catalogue of gene symbols for wheat. In: Pogna NE, Romano M, Pogna EA, Galterio G (eds) Proc 10th int wheat genet symp, vol 4, pp 1–34
McIntosh RA, Devos KM, Dubcovsky J, Rogers WJ (2004) Catalogue of gene symbols for wheat: 2004 (suppl) http://www.wheat.pw.usda.gov/GG2/pubs.shtml
McIntosh RA, Devos KM, Dubcovsky J, Morris CF, Appels R, Anderson OD (2005) Catalogue of gene symbols for wheat: 2005(suppl) http://www.wheat.pw.usda.gov/GG2/pubs.shtml
Murphy JP, Leath S, Huynh D, Navarro RA (2002) Registration of NC99BGTAG11 wheat germplasm resistant to powdery mildew. Crop Sci 42:1382
Neu C, Stein N, Keller B (2002) Genetic mapping of the Lr20 - Pm1 resistance locus reveals suppressed recombination on chromosome arm 7AL in hexaploid wheat. Genome 45:737–744
Qi L, Echalier B, Friebe B, Gill BS (2003) Molecular characterization of a set of wheat deletion stocks for use in chromosome bin mapping of ESTs. Funct Integr Genomics 3:39–55
Qi LL, Echalier B, Chao S, Lazo GR, Butler GE et al (2004) A chromosome bin map of 16,000 EST loci and distribution of genes among the three genomes of polyploid wheat. Genetics 168:701–712
Röder MS, Korzun V, Wedehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A SSR map of wheat. Genetics 149:2007–2023
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Misener S, Krawetz S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, New Jersey, pp 365–386
Schneider D, Heun M, Fischbeck G (1991) Inheritance of the powdery mildew resistance gene Pm9 in relation to Pm1 and Pm2 of wheat. Plant Breed 107:161–164
Sears ER (1966) Nullisomic–tetrasomic combinations in hexaploid wheat. Univ Mo Agric Exp Stn Bull 572:1–58
Sears ER, Briggle LW (1969) Mapping the Pm1 gene for resistance to Erysiphe graminis f. sp. tritici on chromosome 7A of wheat. Crop Sci 9:96–97
Sears ER, Sears LMS (1979) The telocentric chromosomes of common wheat. In: Ramanujan S (ed) Proc 5th int wheat genet symp. Indian Society of Genetics and Plant Breeding, New Delhi, India, pp 23–28
Singrün C (2002) Untersuchungen zur Lokalisierung und Kartierung von Genen für Resistenz gegen Mehltau und Braunrost in Saatweizen (Triticum aestivum L.) und Dinkel ( Triticum spelta L.). Available via http://deposit.d-nb.de/cgi-bin/dokserv?idn=966108620&dok_var=d1&dok_ext=pdf&filename=966108620.pdf. Accessed 10 July 2007
Singrün CH, Hsam SL, Zeller FJ, Mohler V (2003) Powdery mildew resistance gene Pm22 is a member of the complex Pm1 locus in common wheat (Triticum aestivum L). Theor Appl Genet 106:1420–1424
Singrün CH, Hsam SL, Zeller FJ, Wenzel G, Mohler V (2004) Localization of a novel recessive powdery mildew resistance gene from common wheat line RD30 in the terminal region of chromosome 7AL. Theor Appl Genet 109:210–214
Somers DJ, Peter I, Edwards K (2004) A high-density SSR consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Sourdille P, Singh S, Cadalen T, Brown-Guedira GL, Gay G, Qi L, Gill BS, Dufour P, Murigneux A, Bernard M (2004) SSR-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Funct Integr Genomics 4:12–25
Srnić G, Murphy JP, Lyerly JH, Leath S, Marshall DS (2005) Inheritance and chromosomal assignment of powdery mildew resistance genes in two winter wheat germplasm lines. Crop Sci 45:1578–1586
Starling TM, Roane CW, Camper HM (1986) Registration of ‘Saluda’ wheat. Crop Sci 26:200
van Ooijen JW (2006) JoinMap, software for the calculation of genetic linkage maps. Kyazma BV, Wageningen, The Netherlands, Version 4
Yao G, Zhang J, Yang L, Xu H, Jiang Y, Xiong L, Zhang C, Zhang Z, Ma Z, Sorrels ME (2007) Genetic mapping of two powdery mildew resistance genes in einkorn (Triticum monococcum L.) accessions. Theor Appl Genet 114:351–358
Acknowledgements
The authors would like to thank Dr. Marc Cubeta for his valuable input into this project. We would also like to thank Jared Smith, Kim Howell and Lynda Witcher for their greenhouse and laboratory assistance. This research was supported by the USDA-ARS and the USDA-CSREES National Research Initiative CAP grant 2005-05130.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Dubcovsky.
Rights and permissions
About this article
Cite this article
Perugini, L.D., Murphy, J.P., Marshall, D. et al. Pm37, a new broadly effective powdery mildew resistance gene from Triticum timopheevii. Theor Appl Genet 116, 417–425 (2008). https://doi.org/10.1007/s00122-007-0679-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-007-0679-x